

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0344.6
(242 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
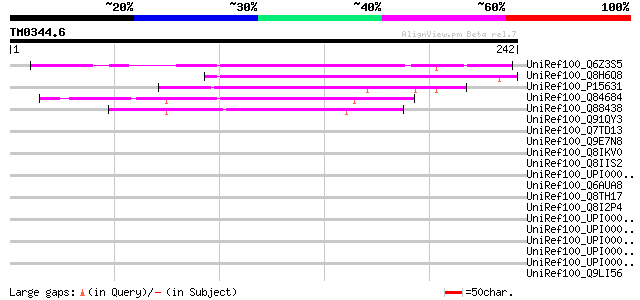
UniRef100_Q6Z3S5 Hypothetical protein OSJNBa0025J22.29 [Oryza sa... 104 2e-21
UniRef100_Q8H6Q8 CTV.20 [Poncirus trifoliata] 75 1e-12
UniRef100_P15631 Movement protein [Soybean chlorotic mottle virus] 54 3e-06
UniRef100_Q84684 Putative cell-to-cell transport protein [Peanut... 49 1e-04
UniRef100_Q88438 ORF I protein [Strawberry vein banding virus] 48 2e-04
UniRef100_Q91QY3 Movement protein [Grapevine berry inner necrosi... 44 0.005
UniRef100_Q7TD13 Putative movement protein [Cestrum yellow leaf ... 43 0.008
UniRef100_Q9E7N8 4b protein [Lettuce necrotic yellows virus] 37 0.44
UniRef100_Q8IKV0 Hypothetical protein [Plasmodium falciparum] 37 0.58
UniRef100_Q8IIS2 Hypothetical protein [Plasmodium falciparum] 36 0.98
UniRef100_UPI0000468B31 UPI0000468B31 UniRef100 entry 35 1.7
UniRef100_Q6AUA8 Hypothetical protein P0017D10.13 [Oryza sativa] 35 1.7
UniRef100_Q8TH17 DEXX-box atpase [Pyrococcus furiosus] 34 2.9
UniRef100_Q8I2P4 Hypothetical protein PFI1300c [Plasmodium falci... 34 3.7
UniRef100_UPI00003AC55E UPI00003AC55E UniRef100 entry 33 4.9
UniRef100_UPI0000499DD9 UPI0000499DD9 UniRef100 entry 33 4.9
UniRef100_UPI000046623E UPI000046623E UniRef100 entry 33 4.9
UniRef100_UPI0000465CBA UPI0000465CBA UniRef100 entry 33 4.9
UniRef100_UPI000029EC7C UPI000029EC7C UniRef100 entry 33 4.9
UniRef100_Q9LI56 EST AU055734(S20025) corresponds to a region of... 33 4.9
>UniRef100_Q6Z3S5 Hypothetical protein OSJNBa0025J22.29 [Oryza sativa]
Length = 1275
Score = 104 bits (260), Expect = 2e-21
Identities = 67/231 (29%), Positives = 111/231 (48%), Gaps = 34/231 (14%)
Query: 11 NNSEENDSNLEQKLQNWSIPKIRTNQVYKTSIFEFNPDYEIKTVEKSFAVQGPYQSISIL 70
N S + ++E+ + NW IPK+ ++Y + EI ++
Sbjct: 5 NASSISFEDIEKSISNWKIPKVNIKEIYHV-------EREINSIRAK------------- 44
Query: 71 DSRIANQHVRKYGYLHIGLVQVAVKPLTRLRYCDLPILICLRDKRMLRFHESVVAVLESN 130
+Y YLH GL+QVA++ LTR + ++ +L CLRD R RF +S++ ++E++
Sbjct: 45 ---------HQYHYLHFGLIQVAIRSLTR-KGLNVSVLACLRDCRNKRFKDSLLGMVEAS 94
Query: 131 IQNGPAYFNCFSSYPVCLRDPHVTTCVDLDIKIPNEMFIDGIPPVSIMYRFCYKVTDNLF 190
+ NGP YFN F + V L D ++ + L+++ G +S+ YR YK L
Sbjct: 95 LSNGPVYFNTFPDFSVSLSDTNIHKVLTLNLQTSGYDLEPGSENISVTYRVYYKAMTTL- 153
Query: 191 GIKAVRDYTVNE-TEVTEINAENSSRITRKILKHDEITYPQEWLQGCSSQP 240
+ YT T + + N N I K LK DEIT P++W+ + +P
Sbjct: 154 -APCAKHYTPKGLTTLLQTNPNNRHTIP-KTLKWDEITLPEKWVLSQAVEP 202
>UniRef100_Q8H6Q8 CTV.20 [Poncirus trifoliata]
Length = 3148
Score = 75.1 bits (183), Expect = 1e-12
Identities = 43/150 (28%), Positives = 80/150 (52%), Gaps = 2/150 (1%)
Query: 94 VKPLTRLRYCDLPILICLRDKRMLRFHESVVAVLESNIQNGPAYFNCFSSYPVCLRDPHV 153
+KPLT+ D IL LRD R + F +S+++ +ES++ GP F+C+ + + L+D +V
Sbjct: 102 IKPLTK-EGLDTSILAVLRDGRFISFDDSLLSSIESSLCKGPIPFDCYPNITISLKDKNV 160
Query: 154 TTCVDLDIKIPNEMFIDGIPPVSIMYRFCYKVTDNLFGIKAVRDYTVNETEVTEINAENS 213
+ L IK N I G PV+++++ YK + F + +ET + + + +
Sbjct: 161 LKSMILQIKTHNYKMIKGSIPVALIFKISYKAMVSAFSTQHRFQSKRDETLLLQTDLSRA 220
Query: 214 SRITRKILKHDEITYPQEW-LQGCSSQPAP 242
+ + K ++ +I P+EW L+G + P
Sbjct: 221 NTVIPKPIQWKDINLPEEWILEGAAPPTIP 250
>UniRef100_P15631 Movement protein [Soybean chlorotic mottle virus]
Length = 303
Score = 53.9 bits (128), Expect = 3e-06
Identities = 43/152 (28%), Positives = 75/152 (49%), Gaps = 6/152 (3%)
Query: 72 SRIANQHVRKYGYLHIGLVQVAVKPLTRLRYCDLPILICLRDKRMLRFHESVVAVLESNI 131
S+I + R Y+HI +QV +K T L+ D P+ + LRD R+L ES +AV N+
Sbjct: 89 SKIEEKQRRLIRYVHISTLQVLIKS-TFLKGLDTPLELTLRDNRLLNLEESKIAVGHGNL 147
Query: 132 QNGPAYFNCFSSYPVCLRDPHVTTCVDLDIKIPNEMFI-DGIPPVSIMYRFCYKVTDNLF 190
+ G F+ + L+D + + L+ K F+ +G SI YR Y ++++
Sbjct: 148 KYGKMKFDVNLQLGLSLKDLDLDRSIILNYKFLRRNFMKEGNHAFSISYRINYALSNSHH 207
Query: 191 GI--KAVRDYTVNE--TEVTEINAENSSRITR 218
+ K ++E +EV E+ S++T+
Sbjct: 208 SVEFKQKEKIYIDELFSEVLELKHPVFSKLTK 239
>UniRef100_Q84684 Putative cell-to-cell transport protein [Peanut chlorotic streak
virus]
Length = 320
Score = 48.9 bits (115), Expect = 1e-04
Identities = 39/183 (21%), Positives = 77/183 (41%), Gaps = 11/183 (6%)
Query: 15 ENDSNLEQKLQNWSIPKIRTNQVYKTSIFEFNPDYEIKTVEKSFAVQGPYQSISILDSR- 73
E++ + QK+ + KI + I I+ F Q P IS + R
Sbjct: 46 ESNEGINQKI----LEKINRKNIIYNGIMCGEQSVPIEQATAKF--QAPVVKISQVQERL 99
Query: 74 --IANQHVRKYGYLHIGLVQVAVKPLTRLRYCDLPILICLRDKRMLRFHESVVAVLESNI 131
I +K G++H+ +VQ+ ++ R P++I + D R+ S++ +E ++
Sbjct: 100 KKIKETDRKKIGFIHVNVVQIVIRSTFR-EGITTPVIIRVEDNRIQDKRYSLLGQIEGDL 158
Query: 132 QNGPAYFNCFSSYPVCLRDPHVTTCVDLDIKI-PNEMFIDGIPPVSIMYRFCYKVTDNLF 190
G FN YP+ L C+ + + ++ G P+ + YR Y +T++
Sbjct: 159 GYGVIKFNVTLQYPIPLITRSFNNCIGVICEFRKQDLMKQGDIPLVVCYRIAYALTNSAI 218
Query: 191 GIK 193
++
Sbjct: 219 SLQ 221
>UniRef100_Q88438 ORF I protein [Strawberry vein banding virus]
Length = 328
Score = 48.1 bits (113), Expect = 2e-04
Identities = 34/145 (23%), Positives = 69/145 (47%), Gaps = 5/145 (3%)
Query: 48 DYEIKTVEKSFAVQGPYQSISILDSR---IANQHVRKYGYLHIGLVQVAVKPLTRLRYCD 104
+Y I+ + + + P+ + ++S+ + + +K ++HIG V++ +K R D
Sbjct: 78 EYPIEIAQANGLTEIPFFNREEIESKKRVLKPEDRKKIDFIHIGSVKIMIKSTFRTGI-D 136
Query: 105 LPILICLRDKRMLRFHESVVAVLESNIQNGPAYFNCFSSYPVCLRDPHVTTCVDL-DIKI 163
PI + L D+RM ++V ++ N+ G F V LRDP + + L
Sbjct: 137 SPISVALLDRRMKNAKDAVFGGVKGNLSYGKLIFTYNPKISVSLRDPTINKTLTLAHFFE 196
Query: 164 PNEMFIDGIPPVSIMYRFCYKVTDN 188
E+ +G P +I Y+ Y ++++
Sbjct: 197 KEELMHEGNHPYTISYKIGYTLSNS 221
>UniRef100_Q91QY3 Movement protein [Grapevine berry inner necrosis virus]
Length = 350
Score = 43.5 bits (101), Expect = 0.005
Identities = 33/167 (19%), Positives = 69/167 (40%), Gaps = 13/167 (7%)
Query: 33 RTNQVYKTSIFEFNPDYEIKTVEKSFAVQGPYQSISILDSRIANQHVRKYGYLHIGLVQV 92
R VY+ + ++ + T+ P + + +L+ N Y+H G + +
Sbjct: 46 RNEIVYQVEPSSISDEFRVTTIPIV-----PMEQLKLLNGSNMN-------YIHFGALSI 93
Query: 93 AVKPLTRLRYCDLPILICLRDKRMLRFHESVVAVLESNIQNGPAYFNCFSSYPVCLRDPH 152
++ PL + R + + D R ++++ ++ +G A C +Y V L DP
Sbjct: 94 SIDPLFK-RNSGVKGKAFVYDSRWNNAEQALLQAFSFDLNSGTASLICSPNYSVQLTDPR 152
Query: 153 VTTCVDLDIKIPNEMFIDGIPPVSIMYRFCYKVTDNLFGIKAVRDYT 199
++TC+ + N F +G +S+ Y+ ++ G D T
Sbjct: 153 LSTCLSTVLVFENLNFREGSFAISVRIGVMYRPYNSYIGDSLELDQT 199
>UniRef100_Q7TD13 Putative movement protein [Cestrum yellow leaf curling virus]
Length = 312
Score = 42.7 bits (99), Expect = 0.008
Identities = 30/124 (24%), Positives = 58/124 (46%), Gaps = 3/124 (2%)
Query: 72 SRIANQHVRK-YGYLHIGLVQVAVKPLTRLRYCDLPILICLRDKRMLRFHESVVAVLESN 130
S+I NQ RK +HI +Q+ +K T L+ D PI + + D+R+ E ++ ++ N
Sbjct: 93 SKIRNQEKRKTLSCIHISTIQIVLKS-TFLKGLDYPISLAITDERINNPKEKIIGIVHGN 151
Query: 131 IQNGPAYFNCFSSYPVCLRDPHVTTCVDLDIK-IPNEMFIDGIPPVSIMYRFCYKVTDNL 189
+ F+ + + L + ++ + L K N++ D SI Y Y + ++
Sbjct: 152 LATVTLKFSVHLGFAIPLTEEDLSRSISLTYKAYRNDLMNDQKQGFSITYAVSYALANSH 211
Query: 190 FGIK 193
I+
Sbjct: 212 HSIQ 215
>UniRef100_Q9E7N8 4b protein [Lettuce necrotic yellows virus]
Length = 302
Score = 37.0 bits (84), Expect = 0.44
Identities = 28/149 (18%), Positives = 66/149 (43%), Gaps = 7/149 (4%)
Query: 84 YLHIGLVQVAVKPLTRLRYC-----DLPILICLRDKRMLRFHESVVAVLESNIQNGPAYF 138
YLHIG + V+++PL RY + + D + +S++++ + ++ G A +
Sbjct: 77 YLHIGCITVSIEPLLHQRYMKNFGKTIAGNCAIIDSTFRKVDQSIISLHKYDLSRGRADY 136
Query: 139 NCFSSYPVCLRDPHVTTCVDLDIKIPNEMFIDGIPPVSIMYRFCYKVTDNLFGIK--AVR 196
+ ++ + L DP + + + + I G+ SI + + L + ++
Sbjct: 137 VSYPNHCLSLTDPMIQKRLSVLLGIKGIDVEPGVELFSICIGYIVSSVNTLHPVSQLGIQ 196
Query: 197 DYTVNETEVTEINAENSSRITRKILKHDE 225
+N TE +I+ + I + L +++
Sbjct: 197 GVAINGTESADIDELGAEDIDQLSLSYND 225
>UniRef100_Q8IKV0 Hypothetical protein [Plasmodium falciparum]
Length = 3347
Score = 36.6 bits (83), Expect = 0.58
Identities = 32/141 (22%), Positives = 66/141 (46%), Gaps = 20/141 (14%)
Query: 3 NLTLNQHANNSEENDSNLEQKLQNWSIPKIRTNQVYKTSIFEFNPDYEIKTVEKSFAVQG 62
N+ +N H NN+ EN++ LE + +N + N+ + T I + D ++K +SF +
Sbjct: 2523 NININIHNNNNIENNNLLEDENKNCDLYDY-VNENFSTIIIHHSRDSKLKQWIESFILII 2581
Query: 63 PYQSISILDSRIANQHVRKYGYLHIGLVQVAVKPLTRLRYCDLPILICLRDKRMLRFHES 122
+ +LD + +K+ Y + +++ +K + + Y +K M H S
Sbjct: 2582 EGCNRILLD----QDNYKKFDYFYDIFIKI-LKTILFIEY---------NNKNMFELHIS 2627
Query: 123 VVAVLESNIQN-----GPAYF 138
++ +L + + GP+YF
Sbjct: 2628 IIQILNNIYMSKLNYYGPSYF 2648
>UniRef100_Q8IIS2 Hypothetical protein [Plasmodium falciparum]
Length = 568
Score = 35.8 bits (81), Expect = 0.98
Identities = 20/58 (34%), Positives = 30/58 (51%), Gaps = 6/58 (10%)
Query: 6 LNQHANNSEENDSNLEQKLQNWSIPKIRTNQVYKTSIFE------FNPDYEIKTVEKS 57
L+ H + +EEN ++ +QNW P I N YK + E NP+Y K +EK+
Sbjct: 427 LHSHKHETEENKKLIKNIMQNWMGPIIGINTNYKQYLKERQKRISENPEYHKKVLEKA 484
>UniRef100_UPI0000468B31 UPI0000468B31 UniRef100 entry
Length = 543
Score = 35.0 bits (79), Expect = 1.7
Identities = 30/145 (20%), Positives = 64/145 (43%), Gaps = 12/145 (8%)
Query: 7 NQHANNSEENDSNLEQKLQNWSIPKIRTNQVYKTSIFEFNPDYEIKTVEKSFAVQGPYQS 66
NQH ++ E S +EQ + +T+ + S N D + + S + G
Sbjct: 222 NQHPSHPENISSQIEQNIDTTKSDSKQTDSSFSNSS---NDDSQSGSRSSSNSKPGDIS- 277
Query: 67 ISILDSRIANQHVRKYGYLHIGLVQVAVKPLTRLRYCDLP---ILICLRDKRMLRFHESV 123
L+ I N++ +K G L ++ + L L +C++ ++ CL + ++ F + +
Sbjct: 278 ---LNPLIENKNAKKLGILKKIILNKYINTL--LNFCNIYNDIVIECLHVQLLMFFFKYI 332
Query: 124 VAVLESNIQNGPAYFNCFSSYPVCL 148
+ V++S + NC+S + +
Sbjct: 333 LQVIKSKFRKEYVKDNCYSKQQILI 357
>UniRef100_Q6AUA8 Hypothetical protein P0017D10.13 [Oryza sativa]
Length = 950
Score = 35.0 bits (79), Expect = 1.7
Identities = 23/91 (25%), Positives = 39/91 (42%), Gaps = 2/91 (2%)
Query: 150 DPHVTTCVDLDIKIPNEMFIDGIPPVSIMYRFCYKVTDNLFGIKAVRDYTVNETEVTEIN 209
D ++ + L+++ + G +S+ YR YK L + YT
Sbjct: 35 DTNIHKVLTLNLQTSSYELEPGSENISVTYRVYYKAMTTL--APCAKHYTPKGLTTLLQT 92
Query: 210 AENSSRITRKILKHDEITYPQEWLQGCSSQP 240
N+ T K LK DEIT P++W+ + +P
Sbjct: 93 NPNNRCTTPKTLKWDEITLPEKWVLSQAVEP 123
>UniRef100_Q8TH17 DEXX-box atpase [Pyrococcus furiosus]
Length = 435
Score = 34.3 bits (77), Expect = 2.9
Identities = 25/81 (30%), Positives = 40/81 (48%), Gaps = 12/81 (14%)
Query: 154 TTCVDLDIKIPNEMFIDGIPPVSIMYRFCYKVTDNLFGIKAVRDYTVNETEVTEINAENS 213
TT + L +K ++ +GI P SI Y C ++D + + +Y E+ EI S
Sbjct: 58 TTLLKLIVK---KLLDEGINPRSIFYLRCDYISDYKELMNVIDEYL----EIREIEGIRS 110
Query: 214 SRITRKILKHDEITYPQEWLQ 234
S I DE+T+P+EW +
Sbjct: 111 SYILL-----DEVTFPREWFR 126
>UniRef100_Q8I2P4 Hypothetical protein PFI1300c [Plasmodium falciparum]
Length = 1922
Score = 33.9 bits (76), Expect = 3.7
Identities = 20/67 (29%), Positives = 34/67 (49%), Gaps = 11/67 (16%)
Query: 12 NSEENDSNLEQKLQNWSIPKIRTNQVYKTSIFEFNPDYEIKTVEKSFAVQGPYQSISILD 71
N ++ + +L NWS P +R N S+F++N Y E+S + Y++++ D
Sbjct: 1551 NDKDTNKSLNNNYNNWSEPPLRNN----NSLFDYNNTYR----EQSNVL---YKNLNYQD 1599
Query: 72 SRIANQH 78
S I N H
Sbjct: 1600 SDIINHH 1606
>UniRef100_UPI00003AC55E UPI00003AC55E UniRef100 entry
Length = 247
Score = 33.5 bits (75), Expect = 4.9
Identities = 16/64 (25%), Positives = 31/64 (48%), Gaps = 3/64 (4%)
Query: 120 HESVV-AVLESNIQNGPAYFNCF--SSYPVCLRDPHVTTCVDLDIKIPNEMFIDGIPPVS 176
H +++ +L N+ + P + C + VCLR PH+T D+ + F+ +P
Sbjct: 14 HSAILYTLLSQNLSHSPPHQRCLCHQRWTVCLRTPHITCQAASDVLVKTISFMKNLPSFH 73
Query: 177 IMYR 180
++ R
Sbjct: 74 LLPR 77
>UniRef100_UPI0000499DD9 UPI0000499DD9 UniRef100 entry
Length = 861
Score = 33.5 bits (75), Expect = 4.9
Identities = 29/122 (23%), Positives = 59/122 (47%), Gaps = 16/122 (13%)
Query: 101 RYCDLPILICLRDKRMLRFHESVVAVLESNIQNGPAYF------NCFSSYPVCLRDPHVT 154
R D P + CLR+ + L ++++ + + G + N F+++P+ +T
Sbjct: 332 RLKDFPNIKCLREIKTLILQKNMLGSIPLEMFTGTSLTALDLSSNSFNAFPMS-----IT 386
Query: 155 TCVDLDIKIPNEMFIDGIPPVSIMYRFCYKVTDNLFGIKAV--RDYTVNE-TEVTEINAE 211
TC +L + + ++D +P +S Y K+ L GI + T+NE T +T ++ E
Sbjct: 387 TCTNLVVLNMSNNYLDSLPDIS--YSCFAKLEALLLGINIIDRLPETMNELTNLTTLHLE 444
Query: 212 NS 213
++
Sbjct: 445 HN 446
>UniRef100_UPI000046623E UPI000046623E UniRef100 entry
Length = 282
Score = 33.5 bits (75), Expect = 4.9
Identities = 20/58 (34%), Positives = 28/58 (47%), Gaps = 6/58 (10%)
Query: 6 LNQHANNSEENDSNLEQKLQNWSIPKIRTNQVYKTSIFE------FNPDYEIKTVEKS 57
L+ H + + EN + + LQNW P I N YK E NP+Y K +EK+
Sbjct: 141 LHTHKDETSENKNMIRNILQNWMGPIIGINSNYKQFAKERQKRIIENPEYHKKILEKA 198
>UniRef100_UPI0000465CBA UPI0000465CBA UniRef100 entry
Length = 455
Score = 33.5 bits (75), Expect = 4.9
Identities = 20/58 (34%), Positives = 28/58 (47%), Gaps = 6/58 (10%)
Query: 6 LNQHANNSEENDSNLEQKLQNWSIPKIRTNQVYKTSIFE------FNPDYEIKTVEKS 57
L+ H + + EN + + LQNW P I N YK E NP+Y K +EK+
Sbjct: 325 LHTHKDETSENKNMIRNILQNWMGPIIGINSNYKQFAKERQKRIIENPEYHKKILEKA 382
>UniRef100_UPI000029EC7C UPI000029EC7C UniRef100 entry
Length = 119
Score = 33.5 bits (75), Expect = 4.9
Identities = 25/98 (25%), Positives = 48/98 (48%), Gaps = 2/98 (2%)
Query: 130 NIQNGPAYFNCFSSYPVCLRDPH-VTTCVDLDIKIPNEMFIDGIPPVSIMYRFCYKVTDN 188
N+ GP +F+ + YP+ + D + + DIK+ E+ I G+ + K+T+N
Sbjct: 18 NLILGPIFFDVLN-YPIGVLDTRKFSYSNETDIKLNVELTIKGVSKLLFTDLKVIKLTNN 76
Query: 189 LFGIKAVRDYTVNETEVTEINAENSSRITRKILKHDEI 226
L IK+ ++ N+ + N N++ + KI + I
Sbjct: 77 LVQIKSNLEFNRNDFNIGTGNWSNTTILKNKIRVNSNI 114
>UniRef100_Q9LI56 EST AU055734(S20025) corresponds to a region of the predicted gene
[Oryza sativa]
Length = 719
Score = 33.5 bits (75), Expect = 4.9
Identities = 21/70 (30%), Positives = 30/70 (42%), Gaps = 2/70 (2%)
Query: 171 GIPPVSIMYRFCYKVTDNLFGIKAVRDYTVNETEVTEINAENSSRITRKILKHDEITYPQ 230
G +S+ YR YK L + YT N+ T K LK DEIT P+
Sbjct: 109 GSENISVTYRVYYKAMTTL--APCAKHYTPKGLTTLLQTNPNNRCTTPKTLKWDEITLPE 166
Query: 231 EWLQGCSSQP 240
+W+ + +P
Sbjct: 167 KWVLSQAVEP 176
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.320 0.136 0.409
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 400,078,703
Number of Sequences: 2790947
Number of extensions: 15409660
Number of successful extensions: 37063
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 7
Number of HSP's successfully gapped in prelim test: 23
Number of HSP's that attempted gapping in prelim test: 37048
Number of HSP's gapped (non-prelim): 34
length of query: 242
length of database: 848,049,833
effective HSP length: 124
effective length of query: 118
effective length of database: 501,972,405
effective search space: 59232743790
effective search space used: 59232743790
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 73 (32.7 bits)
Lotus: description of TM0344.6


