

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0338.9
(434 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
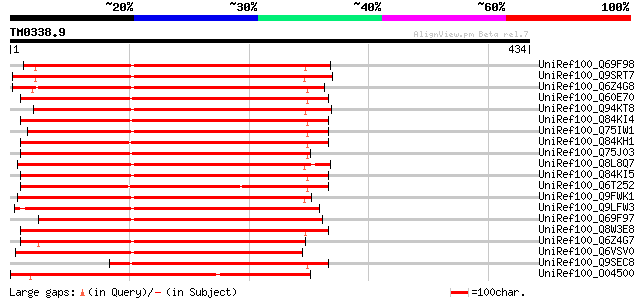
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q69F98 Phytochelatin synthetase-like protein [Phaseolu... 419 e-115
UniRef100_Q9SRT7 COBRA-like protein 1 precursor [Arabidopsis tha... 419 e-115
UniRef100_Q6Z4G8 Putative phytochelatin synthetase [Oryza sativa] 399 e-109
UniRef100_Q60E70 Putative phytochelatin synthetase [Oryza sativa] 398 e-109
UniRef100_Q94KT8 COBRA protein precursor [Arabidopsis thaliana] 398 e-109
UniRef100_Q84KI4 Phytochelatin synthetase-like protein 2 [Sorghu... 392 e-107
UniRef100_Q75IW1 Putative phytochelatin synthetase [Oryza sativa] 386 e-106
UniRef100_Q84KH1 Phytochelatin synthetase [Triticum monococcum] 385 e-105
UniRef100_Q75J03 Putative phytochelatin synthetase, 3'-partial [... 385 e-105
UniRef100_Q8L8Q7 COBRA-like protein 2 precursor [Arabidopsis tha... 380 e-104
UniRef100_Q84KI5 Phytochelatin synthetase-like protein 1 [Sorghu... 376 e-103
UniRef100_Q6T252 Phytochelatin synthetase [Triticum aestivum] 376 e-103
UniRef100_Q9FWK1 Hypothetical protein F21N10.4 [Arabidopsis thal... 375 e-102
UniRef100_Q9LFW3 COBRA-like protein 4 precursor [Arabidopsis tha... 373 e-102
UniRef100_Q69F97 Phytochelatin synthetase-like protein [Phaseolu... 364 3e-99
UniRef100_Q8W3E8 Putative phytochelatin synthetase [Oryza sativa] 348 2e-94
UniRef100_Q6Z4G7 Putative phytochelatin synthetase [Oryza sativa] 343 4e-93
UniRef100_Q6VSV0 BRITTLE CULM1 [Oryza sativa] 334 3e-90
UniRef100_Q9SEC8 Phytochelatin synthetase-like protein [Zea mays] 275 2e-72
UniRef100_O04500 COBRA-like protein 6 precursor [Arabidopsis tha... 252 1e-65
>UniRef100_Q69F98 Phytochelatin synthetase-like protein [Phaseolus vulgaris]
Length = 448
Score = 419 bits (1076), Expect = e-115
Identities = 189/267 (70%), Positives = 223/267 (82%), Gaps = 11/267 (4%)
Query: 12 PCTLLLLLP-----SSSEAYDPLDPNGNITIRWDILSWTGDGYVAAVTLNNFQQYRHIQE 66
PC LLL+L +S++AYDPLDPNGNITI+WDI++WT DGYVA VT+NNFQQYRHI
Sbjct: 11 PCILLLVLLYCTCFTSTDAYDPLDPNGNITIKWDIITWTPDGYVAVVTMNNFQQYRHIAS 70
Query: 67 PGWSLGWTWAKNEIIWQMIGGQTTEQGDCSKFKGQVSPHCCKKDPVAVDLIPGAPYSEQV 126
PGW LGWTWAK E+IW M+GGQTTEQGDCSKFK + PHCCKKDP VDL+PG PY++Q+
Sbjct: 71 PGWRLGWTWAKKEVIWSMMGGQTTEQGDCSKFKAGI-PHCCKKDPTVVDLLPGTPYNQQI 129
Query: 127 ANCCKGGVLTSLKQDPANAVASFRVVVGRAGTTTKSVRLPKNFTFKSPGPGYTCGPARIV 186
ANCCKGGVL S QDP NAV+SF+V VGRAGTT K+V++PKNFT +PGPGYTCGPA+IV
Sbjct: 130 ANCCKGGVLGSWVQDPTNAVSSFQVSVGRAGTTNKTVKVPKNFTLSAPGPGYTCGPAKIV 189
Query: 187 KPTRFLRYDKRRVTQAMMSWNVTCTYSQFLAAKAPTCCVSLSTFYNDTIVSCPTCACGCQ 246
KPT+F++ DKRRVTQA+M+WNVTC YSQFLA K P+CCVSLS+FYNDTIV CPTCACGCQ
Sbjct: 190 KPTQFIQADKRRVTQALMTWNVTCQYSQFLAQKTPSCCVSLSSFYNDTIVPCPTCACGCQ 249
Query: 247 -----SGNCVDPNSPNLTSVISNPGNN 268
SG+CV+ +P+L SV+S G N
Sbjct: 250 GNSSLSGSCVNSQTPHLASVVSGSGKN 276
>UniRef100_Q9SRT7 COBRA-like protein 1 precursor [Arabidopsis thaliana]
Length = 452
Score = 419 bits (1076), Expect = e-115
Identities = 188/274 (68%), Positives = 225/274 (81%), Gaps = 7/274 (2%)
Query: 3 SLLLFRFVLPCTLLLLLP--SSSEAYDPLDPNGNITIRWDILSWTGDGYVAAVTLNNFQQ 60
S + F+F + L+ + SEAYDPLDP+GNIT++WDI++WTGDGYVA VT+ NFQQ
Sbjct: 9 SSIFFKFGISIIFLVSFSGLTPSEAYDPLDPSGNITVKWDIITWTGDGYVATVTVYNFQQ 68
Query: 61 YRHIQEPGWSLGWTWAKNEIIWQMIGGQTTEQGDCSKFKGQVSPHCCKKDPVAVDLIPGA 120
YRHIQ PGW+LGW+WAK E+IW M GGQTTEQGDCSKFKG + PHCCKK P VDL+PG+
Sbjct: 69 YRHIQAPGWTLGWSWAKREVIWGMNGGQTTEQGDCSKFKGTI-PHCCKKTPSVVDLLPGS 127
Query: 121 PYSEQVANCCKGGVLTSLKQDPANAVASFRVVVGRAGTTTKSVRLPKNFTFKSPGPGYTC 180
PY++Q+ANCC+GGVL S QDPA AV++F++ VG+AGTT K+VR+PKNFT K+PGPGYTC
Sbjct: 128 PYNQQIANCCRGGVLNSWAQDPATAVSAFQLTVGQAGTTNKTVRVPKNFTLKAPGPGYTC 187
Query: 181 GPARIVKPTRFLRYDKRRVTQAMMSWNVTCTYSQFLAAKAPTCCVSLSTFYNDTIVSCPT 240
PA+IVKPTRF+ DKRRVTQA+M+WNVTCTYSQFLA K PTCCVSLS+FYN TIVSCPT
Sbjct: 188 SPAKIVKPTRFIGTDKRRVTQALMTWNVTCTYSQFLAQKTPTCCVSLSSFYNKTIVSCPT 247
Query: 241 CACGC----QSGNCVDPNSPNLTSVISNPGNNGH 270
C+CGC Q GNCVDP P + SVI NPG N +
Sbjct: 248 CSCGCRNTSQPGNCVDPKGPRIASVIPNPGKNAY 281
>UniRef100_Q6Z4G8 Putative phytochelatin synthetase [Oryza sativa]
Length = 446
Score = 399 bits (1024), Expect = e-109
Identities = 184/269 (68%), Positives = 218/269 (80%), Gaps = 10/269 (3%)
Query: 3 SLLLFRFVLPCTLLL-----LLPSSSEAYDPLDPNGNITIRWDILSWTGDGYVAAVTLNN 57
+LLL R + LL+ L+PSS EAYDPLDPNGNITI+WD+L WT DGYVA V+L N
Sbjct: 2 ALLLLRMGVSVALLVAFFSSLIPSS-EAYDPLDPNGNITIKWDVLQWTPDGYVAVVSLYN 60
Query: 58 FQQYRHIQEPGWSLGWTWAKNEIIWQMIGGQTTEQGDCSKFKGQVSPHCCKKDPVAVDLI 117
+QQYRHIQ PGW LGW WAK EIIW M GGQ TEQGDCSKFK + PHCCKKDP VDL+
Sbjct: 61 YQQYRHIQSPGWKLGWVWAKKEIIWAMNGGQATEQGDCSKFKSNI-PHCCKKDPEIVDLL 119
Query: 118 PGAPYSEQVANCCKGGVLTSLKQDPANAVASFRVVVGRAGTTTKSVRLPKNFTFKSPGPG 177
PG PY+ Q+ANCCKGGVL S QDPANA+ASF+V VG+AGTT K+VR+P+NFT KSPGPG
Sbjct: 120 PGTPYNMQIANCCKGGVLNSWAQDPANAIASFQVSVGQAGTTNKTVRVPRNFTLKSPGPG 179
Query: 178 YTCGPARIVKPTRFLRYDKRRVTQAMMSWNVTCTYSQFLAAKAPTCCVSLSTFYNDTIVS 237
YTCG A++V+PT+F D RR TQA M+WNVTCTYSQ +A ++PTCCVSLS+FYNDTIV+
Sbjct: 180 YTCGSAKVVRPTKFFSQDGRRTTQAHMTWNVTCTYSQIVAQRSPTCCVSLSSFYNDTIVN 239
Query: 238 CPTCACGCQS---GNCVDPNSPNLTSVIS 263
CPTC+CGCQ+ G+CV+ NSP L SV++
Sbjct: 240 CPTCSCGCQNNKPGSCVEGNSPYLASVVN 268
>UniRef100_Q60E70 Putative phytochelatin synthetase [Oryza sativa]
Length = 457
Score = 398 bits (1023), Expect = e-109
Identities = 176/262 (67%), Positives = 212/262 (80%), Gaps = 6/262 (2%)
Query: 10 VLPCTLLLLLPSSSEAYDPLDPNGNITIRWDILSWTGDGYVAAVTLNNFQQYRHIQEPGW 69
+L LL P+++EAYD LDPNGNITI+WD++SWT DGYVA VT+ N+QQ+RHIQ PGW
Sbjct: 21 LLAAALLFSAPATTEAYDALDPNGNITIKWDVMSWTPDGYVAVVTMFNYQQFRHIQAPGW 80
Query: 70 SLGWTWAKNEIIWQMIGGQTTEQGDCSKFKGQVSPHCCKKDPVAVDLIPGAPYSEQVANC 129
LGWTWAK E+IW M+G QTTEQGDCSKFKG +PHCCKKDP VDL+PG PY+ Q+ANC
Sbjct: 81 QLGWTWAKKEVIWSMVGAQTTEQGDCSKFKGG-TPHCCKKDPTVVDLLPGTPYNMQIANC 139
Query: 130 CKGGVLTSLKQDPANAVASFRVVVGRAGTTTKSVRLPKNFTFKSPGPGYTCGPARIVKPT 189
CK GV+ + QDP+NA +SF++ VG AGTT K+V+LPKNFT K+PGPGYTCG A IV+PT
Sbjct: 140 CKAGVINTFNQDPSNAASSFQISVGLAGTTNKTVKLPKNFTLKAPGPGYTCGRAMIVRPT 199
Query: 190 RFLRYDKRRVTQAMMSWNVTCTYSQFLAAKAPTCCVSLSTFYNDTIVSCPTCACGCQS-- 247
+F D RR TQA+M+WNVTCTYSQFLA K P+CCVSLS+FYNDTIV+CPTC+CGCQ+
Sbjct: 200 KFFTGDGRRATQALMTWNVTCTYSQFLAQKTPSCCVSLSSFYNDTIVNCPTCSCGCQNNG 259
Query: 248 ---GNCVDPNSPNLTSVISNPG 266
G+CV+ NSP L S I PG
Sbjct: 260 TSPGSCVNENSPYLQSAIDGPG 281
>UniRef100_Q94KT8 COBRA protein precursor [Arabidopsis thaliana]
Length = 456
Score = 398 bits (1023), Expect = e-109
Identities = 172/254 (67%), Positives = 210/254 (81%), Gaps = 6/254 (2%)
Query: 21 SSSEAYDPLDPNGNITIRWDILSWTGDGYVAAVTLNNFQQYRHIQEPGWSLGWTWAKNEI 80
+S+EAYD LDP GNIT++WD++SWT DGYVA VT+ NFQ+YRHIQ PGW+LGW WAK E+
Sbjct: 32 TSTEAYDALDPEGNITMKWDVMSWTPDGYVAVVTMFNFQKYRHIQSPGWTLGWKWAKKEV 91
Query: 81 IWQMIGGQTTEQGDCSKFKGQVSPHCCKKDPVAVDLIPGAPYSEQVANCCKGGVLTSLKQ 140
IW M+G QTTEQGDCSK+KG + PHCCKKDP VDL+PG PY++Q+ANCCKGGV+ S Q
Sbjct: 92 IWSMVGAQTTEQGDCSKYKGNI-PHCCKKDPTVVDLLPGTPYNQQIANCCKGGVMNSWVQ 150
Query: 141 DPANAVASFRVVVGRAGTTTKSVRLPKNFTFKSPGPGYTCGPARIVKPTRFLRYDKRRVT 200
DPA A +SF++ VG AGTT K+VR+P+NFT PGPGYTCGPA+IV+PT+F+ D RR T
Sbjct: 151 DPATAASSFQISVGAAGTTNKTVRVPRNFTLMGPGPGYTCGPAKIVRPTKFVTTDTRRTT 210
Query: 201 QAMMSWNVTCTYSQFLAAKAPTCCVSLSTFYNDTIVSCPTCACGCQ-----SGNCVDPNS 255
QAMM+WN+TCTYSQFLA + PTCCVSLS+FYN+TIV CPTCACGCQ SG C+DP++
Sbjct: 211 QAMMTWNITCTYSQFLAQRTPTCCVSLSSFYNETIVGCPTCACGCQNNRTESGACLDPDT 270
Query: 256 PNLTSVISNPGNNG 269
P+L SV+S P G
Sbjct: 271 PHLASVVSPPTKKG 284
>UniRef100_Q84KI4 Phytochelatin synthetase-like protein 2 [Sorghum bicolor]
Length = 449
Score = 392 bits (1006), Expect = e-107
Identities = 173/260 (66%), Positives = 209/260 (79%), Gaps = 4/260 (1%)
Query: 10 VLPCTLLLLLPSSSEAYDPLDPNGNITIRWDILSWTGDGYVAAVTLNNFQQYRHIQEPGW 69
+L LLL P+++EAYD LDP GNITI+WDI+ WT DGYVA VT+ NFQQ+RHI PGW
Sbjct: 15 LLAAALLLSAPTTTEAYDSLDPTGNITIKWDIMQWTPDGYVAVVTMYNFQQFRHIGAPGW 74
Query: 70 SLGWTWAKNEIIWQMIGGQTTEQGDCSKFKGQVSPHCCKKDPVAVDLIPGAPYSEQVANC 129
LGWTWAK E+IW M+G QTTEQGDCSKFKG +PHCCKKDP VDL+PG PY+ Q+ANC
Sbjct: 75 QLGWTWAKKEVIWSMVGAQTTEQGDCSKFKGN-TPHCCKKDPTIVDLLPGTPYNMQIANC 133
Query: 130 CKGGVLTSLKQDPANAVASFRVVVGRAGTTTKSVRLPKNFTFKSPGPGYTCGPARIVKPT 189
CK GV+ + QDPANA +SF++ VG AGTT K+V++PKNFT K+PGPGYTCG A + +PT
Sbjct: 134 CKAGVINTFNQDPANAASSFQISVGLAGTTNKTVKVPKNFTLKTPGPGYTCGRAIVGRPT 193
Query: 190 RFLRYDKRRVTQAMMSWNVTCTYSQFLAAKAPTCCVSLSTFYNDTIVSCPTCACGCQS-- 247
+F D RR TQA+M+WNVTCTYSQFLA K P+CCVSLS+FYNDTIV+CPTC+CGCQ+
Sbjct: 194 KFWSADGRRATQALMTWNVTCTYSQFLAQKTPSCCVSLSSFYNDTIVNCPTCSCGCQNPS 253
Query: 248 -GNCVDPNSPNLTSVISNPG 266
NCV+ +SPNL + I PG
Sbjct: 254 GSNCVNEDSPNLQAAIDGPG 273
>UniRef100_Q75IW1 Putative phytochelatin synthetase [Oryza sativa]
Length = 458
Score = 386 bits (991), Expect = e-106
Identities = 176/255 (69%), Positives = 207/255 (81%), Gaps = 5/255 (1%)
Query: 16 LLLLPSSSEAYDPLDPNGNITIRWDILSWTGDGYVAAVTLNNFQQYRHIQEPGWSLGWTW 75
LLL+ +EAYDPLDPNGNITI+WDI WT DGYVA VT+ NFQ+YRHIQ PGWSLGW W
Sbjct: 20 LLLMVPFAEAYDPLDPNGNITIKWDITQWTPDGYVAVVTIYNFQKYRHIQAPGWSLGWAW 79
Query: 76 AKNEIIWQMIGGQTTEQGDCSKFKGQVSPHCCKKDPVAVDLIPGAPYSEQVANCCKGGVL 135
AK EIIW M GGQ TEQGDCS FK + PHCCK+DP VDL+PGAPY+ Q NCCKGGVL
Sbjct: 80 AKKEIIWSMAGGQATEQGDCSAFKANI-PHCCKRDPRVVDLVPGAPYNMQFGNCCKGGVL 138
Query: 136 TSLKQDPANAVASFRVVVGRAGTTTKSVRLPKNFTFKSPGPGYTCGPARIVK-PTRFLRY 194
TS QDP NAVASF++ VG +GT+ K+V+ PKNFT K+PGPGY+CG A+ VK PTRF+
Sbjct: 139 TSWVQDPLNAVASFQITVGHSGTSNKTVKAPKNFTLKAPGPGYSCGLAQEVKPPTRFISL 198
Query: 195 DKRRVTQAMMSWNVTCTYSQFLAAKAPTCCVSLSTFYNDTIVSCPTCACGCQS---GNCV 251
D RR TQA ++WNVTCTYSQF+A +APTCCVSLS+FYN+TIV+CP CACGCQ+ G+CV
Sbjct: 199 DGRRTTQAHVTWNVTCTYSQFVAQRAPTCCVSLSSFYNETIVNCPKCACGCQNKKPGSCV 258
Query: 252 DPNSPNLTSVISNPG 266
+ NSP L SV++ PG
Sbjct: 259 EGNSPYLASVVNGPG 273
>UniRef100_Q84KH1 Phytochelatin synthetase [Triticum monococcum]
Length = 457
Score = 385 bits (988), Expect = e-105
Identities = 168/262 (64%), Positives = 208/262 (79%), Gaps = 6/262 (2%)
Query: 10 VLPCTLLLLLPSSSEAYDPLDPNGNITIRWDILSWTGDGYVAAVTLNNFQQYRHIQEPGW 69
+L L P+++EAYD LDPNGNITI+WDI+SWT DGYVA VT+ N+QQ+RHI PGW
Sbjct: 20 LLAAVLFFSAPATTEAYDSLDPNGNITIKWDIISWTPDGYVATVTMFNYQQFRHIPAPGW 79
Query: 70 SLGWTWAKNEIIWQMIGGQTTEQGDCSKFKGQVSPHCCKKDPVAVDLIPGAPYSEQVANC 129
LGW+WAK E+IW M+G Q TEQGDCSKFK PHCCK+DP VDL+PG P+++Q+ANC
Sbjct: 80 QLGWSWAKKEVIWSMVGAQATEQGDCSKFKS-APPHCCKRDPTIVDLLPGTPFNQQIANC 138
Query: 130 CKGGVLTSLKQDPANAVASFRVVVGRAGTTTKSVRLPKNFTFKSPGPGYTCGPARIVKPT 189
CK GV+ + QDP NA +SF++ VG AGTT K+V++PKNFT ++PGPGYTCG A + +PT
Sbjct: 139 CKAGVIKTFNQDPGNAASSFQISVGLAGTTNKTVKMPKNFTLRAPGPGYTCGRALVGRPT 198
Query: 190 RFLRYDKRRVTQAMMSWNVTCTYSQFLAAKAPTCCVSLSTFYNDTIVSCPTCACGCQS-- 247
++ D RRVTQA+MSWNVTCTYSQFLA K PTCCVSLS+FYNDTIV+CPTC+CGCQ+
Sbjct: 199 KYYSSDGRRVTQALMSWNVTCTYSQFLAQKTPTCCVSLSSFYNDTIVNCPTCSCGCQNNI 258
Query: 248 ---GNCVDPNSPNLTSVISNPG 266
G+CV+ NSP L S I+ PG
Sbjct: 259 TRPGSCVNDNSPYLQSAINGPG 280
>UniRef100_Q75J03 Putative phytochelatin synthetase, 3'-partial [Oryza sativa]
Length = 266
Score = 385 bits (988), Expect = e-105
Identities = 170/247 (68%), Positives = 201/247 (80%), Gaps = 6/247 (2%)
Query: 10 VLPCTLLLLLPSSSEAYDPLDPNGNITIRWDILSWTGDGYVAAVTLNNFQQYRHIQEPGW 69
+L LLL PS++EAYD LDPNGNITI+WD++ WT DGYVA VT+ N+QQ+RHIQ PGW
Sbjct: 21 LLVAALLLSAPSATEAYDSLDPNGNITIKWDVMQWTPDGYVAVVTMFNYQQFRHIQAPGW 80
Query: 70 SLGWTWAKNEIIWQMIGGQTTEQGDCSKFKGQVSPHCCKKDPVAVDLIPGAPYSEQVANC 129
LGWTWAK E+IW M+G QTTEQGDCSKFKG PHCCKKDP VDL+PG PY+ Q+ANC
Sbjct: 81 QLGWTWAKKEVIWSMVGAQTTEQGDCSKFKGN-PPHCCKKDPTIVDLLPGTPYNMQIANC 139
Query: 130 CKGGVLTSLKQDPANAVASFRVVVGRAGTTTKSVRLPKNFTFKSPGPGYTCGPARIVKPT 189
CK GV+ + QDP NA +SF++ VG AGTT K+V+LPKNFT K+PGPGYTCG A IV+PT
Sbjct: 140 CKAGVINTFNQDPLNAASSFQISVGLAGTTNKTVKLPKNFTLKAPGPGYTCGRAMIVRPT 199
Query: 190 RFLRYDKRRVTQAMMSWNVTCTYSQFLAAKAPTCCVSLSTFYNDTIVSCPTCACGCQS-- 247
+F D RR TQA+M+WNVTCTYSQFLA K PTCCVSLS+FYNDTIV+CPTC+CGCQ+
Sbjct: 200 KFFTNDGRRATQALMTWNVTCTYSQFLAQKTPTCCVSLSSFYNDTIVNCPTCSCGCQNNG 259
Query: 248 ---GNCV 251
G+CV
Sbjct: 260 TSPGSCV 266
>UniRef100_Q8L8Q7 COBRA-like protein 2 precursor [Arabidopsis thaliana]
Length = 441
Score = 380 bits (976), Expect = e-104
Identities = 165/266 (62%), Positives = 210/266 (78%), Gaps = 8/266 (3%)
Query: 7 FRFVLPCTLLLLLPSSSEAYDPLDPNGNITIRWDILSWTGDGYVAAVTLNNFQQYRHIQE 66
F + C+ +++EAYD LDP GNITI+WDI+SWTGDGYVA VT+ NFQQYRHI+
Sbjct: 10 FLLLFLCSWTSFTFTTTEAYDALDPYGNITIKWDIMSWTGDGYVAVVTIFNFQQYRHIEA 69
Query: 67 PGWSLGWTWAKNEIIWQMIGGQTTEQGDCSKFKGQVSPHCCKKDPVAVDLIPGAPYSEQV 126
PGW LGW+W K E+IW M+GGQ TEQGDCSKFKG + PHCCKK P VDL+PG PY++Q+
Sbjct: 70 PGWQLGWSWMKKEVIWSMVGGQATEQGDCSKFKGNI-PHCCKKTPAIVDLLPGTPYNQQI 128
Query: 127 ANCCKGGVLTSLKQDPANAVASFRVVVGRAGTTTKSVRLPKNFTFKSPGPGYTCGPARIV 186
+NCC+GGV+++ QDPA A++SF++ VG++GTT +VR P+N T K+PGPGYTCGPA++V
Sbjct: 129 SNCCRGGVISAWAQDPATAISSFQISVGQSGTTNTTVRAPRNITLKAPGPGYTCGPAKLV 188
Query: 187 KPTRFLRYDKRRVTQAMMSWNVTCTYSQFLAAKAPTCCVSLSTFYNDTIVSCPTCACGC- 245
KP+RF+ DKRR TQ++++WN+TCTYSQFLA K PTCCVSLS FYN+TIV CPTC+CGC
Sbjct: 189 KPSRFISADKRRKTQSLLTWNITCTYSQFLARKTPTCCVSLSAFYNETIVPCPTCSCGCQ 248
Query: 246 ---QSGNCVDPNSPNLTSVISNPGNN 268
Q+G CVD P + SV+ G N
Sbjct: 249 NSSQAGTCVD---PKIASVVPALGKN 271
>UniRef100_Q84KI5 Phytochelatin synthetase-like protein 1 [Sorghum bicolor]
Length = 425
Score = 376 bits (966), Expect = e-103
Identities = 165/260 (63%), Positives = 205/260 (78%), Gaps = 4/260 (1%)
Query: 10 VLPCTLLLLLPSSSEAYDPLDPNGNITIRWDILSWTGDGYVAAVTLNNFQQYRHIQEPGW 69
+L LL+ P ++EAYD LDPNGNITI+WD++ WT DGYVA VT+ N+QQ+RHI PGW
Sbjct: 16 LLAVALLISAPDAAEAYDSLDPNGNITIKWDVMQWTPDGYVAVVTMYNYQQFRHIGPPGW 75
Query: 70 SLGWTWAKNEIIWQMIGGQTTEQGDCSKFKGQVSPHCCKKDPVAVDLIPGAPYSEQVANC 129
LGWTWAK E+IW M+G Q TEQGDCSKFKG + PH CKKDPV VDL+P PY Q+ANC
Sbjct: 76 QLGWTWAKKEVIWSMVGAQATEQGDCSKFKGNI-PHSCKKDPVIVDLLPSTPYDMQIANC 134
Query: 130 CKGGVLTSLKQDPANAVASFRVVVGRAGTTTKSVRLPKNFTFKSPGPGYTCGPARIVKPT 189
CK GV++++ QDPANA +SF++ VG +G+TTK+V++PKNFT ++PGPGYTCG A + KPT
Sbjct: 135 CKAGVISTISQDPANAASSFQLSVGLSGSTTKTVKVPKNFTLRTPGPGYTCGRAIVGKPT 194
Query: 190 RFLRYDKRRVTQAMMSWNVTCTYSQFLAAKAPTCCVSLSTFYNDTIVSCPTCACGCQS-- 247
F D RR T+A+M+WNVTCTYSQFLA K P+CCVSLS+FYN+T V+CPTC+CGCQ+
Sbjct: 195 IFFSADGRRATKALMTWNVTCTYSQFLAQKTPSCCVSLSSFYNNTTVNCPTCSCGCQNPS 254
Query: 248 -GNCVDPNSPNLTSVISNPG 266
NCV+ SP L S I PG
Sbjct: 255 GSNCVNKGSPRLRSAIDGPG 274
>UniRef100_Q6T252 Phytochelatin synthetase [Triticum aestivum]
Length = 456
Score = 376 bits (965), Expect = e-103
Identities = 167/262 (63%), Positives = 207/262 (78%), Gaps = 7/262 (2%)
Query: 10 VLPCTLLLLLPSSSEAYDPLDPNGNITIRWDILSWTGDGYVAAVTLNNFQQYRHIQEPGW 69
+L L P+++EAYD LDPNGNITI+WDI+SWT DGYVA VT+ N+QQ+RHI PGW
Sbjct: 20 LLAAVLFFSAPATTEAYDSLDPNGNITIKWDIISWTPDGYVATVTMFNYQQFRHIPAPGW 79
Query: 70 SLGWTWAKNEIIWQMIGGQTTEQGDCSKFKGQVSPHCCKKDPVAVDLIPGAPYSEQVANC 129
LG +WAK E+IW M+G Q TEQGDCSKFK PHCCK+DP VDL+PG P+++Q+ANC
Sbjct: 80 QLGCSWAKKEVIWSMVGAQATEQGDCSKFK-SAPPHCCKRDPTIVDLLPGTPFNQQIANC 138
Query: 130 CKGGVLTSLKQDPANAVASFRVVVGRAGTTTKSVRLPKNFTFKSPGPGYTCGPARIVKPT 189
CK GV+ + QDP NA +SF++ VG AGTT K+V++PKNFT ++PGPGYTCG A + +PT
Sbjct: 139 CKAGVIKTFNQDPGNAASSFQISVGLAGTTNKAVKMPKNFTLRAPGPGYTCGRALVGRPT 198
Query: 190 RFLRYDKRRVTQAMMSWNVTCTYSQFLAAKAPTCCVSLSTFYNDTIVSCPTCACGCQS-- 247
++ D RRVTQA+MSWNVTCTYSQFLA K PTCCVSLS+FYNDTIV+CPTC+CGCQ+
Sbjct: 199 KYYS-DGRRVTQALMSWNVTCTYSQFLAQKTPTCCVSLSSFYNDTIVNCPTCSCGCQNNI 257
Query: 248 ---GNCVDPNSPNLTSVISNPG 266
G+CV+ NSP L S I+ PG
Sbjct: 258 TRPGSCVNDNSPYLQSAINGPG 279
>UniRef100_Q9FWK1 Hypothetical protein F21N10.4 [Arabidopsis thaliana]
Length = 404
Score = 375 bits (964), Expect = e-102
Identities = 160/250 (64%), Positives = 203/250 (81%), Gaps = 5/250 (2%)
Query: 7 FRFVLPCTLLLLLPSSSEAYDPLDPNGNITIRWDILSWTGDGYVAAVTLNNFQQYRHIQE 66
F + C+ +++EAYD LDP GNITI+WDI+SWTGDGYVA VT+ NFQQYRHI+
Sbjct: 10 FLLLFLCSWTSFTFTTTEAYDALDPYGNITIKWDIMSWTGDGYVAVVTIFNFQQYRHIEA 69
Query: 67 PGWSLGWTWAKNEIIWQMIGGQTTEQGDCSKFKGQVSPHCCKKDPVAVDLIPGAPYSEQV 126
PGW LGW+W K E+IW M+GGQ TEQGDCSKFKG + PHCCKK P VDL+PG PY++Q+
Sbjct: 70 PGWQLGWSWMKKEVIWSMVGGQATEQGDCSKFKGNI-PHCCKKTPAIVDLLPGTPYNQQI 128
Query: 127 ANCCKGGVLTSLKQDPANAVASFRVVVGRAGTTTKSVRLPKNFTFKSPGPGYTCGPARIV 186
+NCC+GGV+++ QDPA A++SF++ VG++GTT +VR P+N T K+PGPGYTCGPA++V
Sbjct: 129 SNCCRGGVISAWAQDPATAISSFQISVGQSGTTNTTVRAPRNITLKAPGPGYTCGPAKLV 188
Query: 187 KPTRFLRYDKRRVTQAMMSWNVTCTYSQFLAAKAPTCCVSLSTFYNDTIVSCPTCACGC- 245
KP+RF+ DKRR TQ++++WN+TCTYSQFLA K PTCCVSLS FYN+TIV CPTC+CGC
Sbjct: 189 KPSRFISADKRRKTQSLLTWNITCTYSQFLARKTPTCCVSLSAFYNETIVPCPTCSCGCQ 248
Query: 246 ---QSGNCVD 252
Q+G CVD
Sbjct: 249 NSSQAGTCVD 258
>UniRef100_Q9LFW3 COBRA-like protein 4 precursor [Arabidopsis thaliana]
Length = 431
Score = 373 bits (958), Expect = e-102
Identities = 170/256 (66%), Positives = 202/256 (78%), Gaps = 5/256 (1%)
Query: 5 LLFRFVLPCTLLLLLPSSSEAYDPLDPNGNITIRWDILSWTGDGYVAAVTLNNFQQYRHI 64
LLF F C ++ ++ AYDPLDP+GNITI+WDI+SWT DGYVA VT+NNFQ YRHI
Sbjct: 3 LLFSF---CFFFFMIIFTATAYDPLDPSGNITIKWDIMSWTADGYVATVTMNNFQIYRHI 59
Query: 65 QEPGWSLGWTWAKNEIIWQMIGGQTTEQGDCSKFKGQVSPHCCKKDPVAVDLIPGAPYSE 124
Q PGW+LGWTWAK E+IW M+G QTTEQGDCSKFKG V PHCCKK P VDL+PG PY++
Sbjct: 60 QNPGWTLGWTWAKKEVIWSMVGAQTTEQGDCSKFKGNV-PHCCKKTPTVVDLLPGVPYNQ 118
Query: 125 QVANCCKGGVLTSLKQDPANAVASFRVVVGRAGTTTKSVRLPKNFTFKSPGPGYTCGPAR 184
Q +NCCKGGV+ + QDP+ AV+ F+V G AGTT K+V+LPKNFT PGPGYTCGPA+
Sbjct: 119 QFSNCCKGGVIGAWGQDPSAAVSQFQVSAGLAGTTNKTVKLPKNFTLLGPGPGYTCGPAK 178
Query: 185 IVKPTRFLRYDKRRVTQAMMSWNVTCTYSQFLAAKAPTCCVSLSTFYNDTIVSCPTCACG 244
IV T FL DKRR TQA+M+WNVTCTYSQFLA K P+CCVS S+FYNDTI CP+CACG
Sbjct: 179 IVPSTVFLTTDKRRKTQALMTWNVTCTYSQFLARKHPSCCVSFSSFYNDTITPCPSCACG 238
Query: 245 CQS-GNCVDPNSPNLT 259
C++ +CV +S LT
Sbjct: 239 CENKKSCVKADSKILT 254
>UniRef100_Q69F97 Phytochelatin synthetase-like protein [Phaseolus vulgaris]
Length = 463
Score = 364 bits (934), Expect = 3e-99
Identities = 158/238 (66%), Positives = 194/238 (81%), Gaps = 2/238 (0%)
Query: 25 AYDPLDPNGNITIRWDILSWTGDGYVAAVTLNNFQQYRHIQEPGWSLGWTWAKNEIIWQM 84
AYDPLDPNGN+TI+WD++SWT DGYVA VT++NFQ +RHI PGW+L W+WAK E+IW M
Sbjct: 51 AYDPLDPNGNVTIKWDVVSWTPDGYVAVVTMSNFQMFRHIMNPGWTLSWSWAKKEVIWSM 110
Query: 85 IGGQTTEQGDCSKFKGQVSPHCCKKDPVAVDLIPGAPYSEQVANCCKGGVLTSLKQDPAN 144
+G QTTEQGDCSKFKG V PHCCKK P VDL+PG PY++Q +NCCKGGV+ + QDP++
Sbjct: 111 VGAQTTEQGDCSKFKGNV-PHCCKKTPTVVDLLPGVPYNQQFSNCCKGGVVAAWGQDPSS 169
Query: 145 AVASFRVVVGRAGTTTKSVRLPKNFTFKSPGPGYTCGPARIVKPTRFLRYDKRRVTQAMM 204
AV+SF+V +G AGT+ K+V+LPKNFT PGPGYTCGPA++V T FL DKRR TQA+M
Sbjct: 170 AVSSFQVSIGLAGTSNKTVKLPKNFTLLGPGPGYTCGPAKVVPSTVFLTADKRRKTQALM 229
Query: 205 SWNVTCTYSQFLAAKAPTCCVSLSTFYNDTIVSCPTCACGCQS-GNCVDPNSPNLTSV 261
+WNVTCTYSQFLA K P CCVSLS+FYN+TI CP+CACGCQ+ NC+ +S + V
Sbjct: 230 TWNVTCTYSQFLARKNPGCCVSLSSFYNETITPCPSCACGCQNKRNCIKSDSKRINMV 287
>UniRef100_Q8W3E8 Putative phytochelatin synthetase [Oryza sativa]
Length = 425
Score = 348 bits (892), Expect = 2e-94
Identities = 155/263 (58%), Positives = 193/263 (72%), Gaps = 6/263 (2%)
Query: 10 VLPCTLLLLLPSSSEAYDPLDPNGNITIRWDILSWTGDGYVAAVTLNNFQQYRHIQEPGW 69
+L LL P+++ AYD LDPNGNITI+WD++ WT DGY A VTL+N+QQ+RHIQ PGW
Sbjct: 12 LLAAALLFSSPATTYAYDSLDPNGNITIKWDVMQWTPDGYAAVVTLSNYQQFRHIQPPGW 71
Query: 70 SLGWTWAKNEIIWQMIGGQTTEQGDCSKFK-GQVSPHCCKKDPVAVDLIPGAPYSEQVAN 128
LGWTW + E+IW M G Q EQGDCS K G PH CKK P VDL+PGAP Q+AN
Sbjct: 72 QLGWTWQQKEVIWSMYGAQAIEQGDCSMSKEGSNVPHSCKKHPTVVDLLPGAPIDLQIAN 131
Query: 129 CCKGGVLTSLKQDPANAVASFRVVVGRAGTTTKSVRLPKNFTFKSPGPGYTCGPARIVKP 188
CCK G L++ QDPAN+ ASF+++VG +G + ++VR+PKNF+ +PGPGYTC A IVKP
Sbjct: 132 CCKAGSLSAFSQDPANSAASFQIIVGHSGNSNETVRVPKNFSLMAPGPGYTCSRAMIVKP 191
Query: 189 TRFLRYDKRRVTQAMMSWNVTCTYSQFLAAKAPTCCVSLSTFYNDTIVSCPTCACGCQ-- 246
+RFL D RR TQ +M+WNV CTYSQFLA K P+CCVSLS+F ND V CPTC+CGC+
Sbjct: 192 SRFLSPDGRRATQVLMTWNVICTYSQFLAQKVPSCCVSLSSFDNDKTVDCPTCSCGCRNE 251
Query: 247 ---SGNCVDPNSPNLTSVISNPG 266
+G CV N+P+L S+I PG
Sbjct: 252 KSTTGKCVKKNAPDLQSIIHGPG 274
>UniRef100_Q6Z4G7 Putative phytochelatin synthetase [Oryza sativa]
Length = 451
Score = 343 bits (881), Expect = 4e-93
Identities = 151/241 (62%), Positives = 191/241 (78%), Gaps = 4/241 (1%)
Query: 10 VLPCTLLLLLPSS---SEAYDPLDPNGNITIRWDILSWTGDGYVAAVTLNNFQQYRHIQE 66
VL LLL+L ++ + AYDPLDPNGNITI+WD++SWT DGYVA VT+NN+Q YR I
Sbjct: 3 VLGSLLLLILAATLSVAVAYDPLDPNGNITIKWDVMSWTPDGYVAMVTINNYQTYRQIMA 62
Query: 67 PGWSLGWTWAKNEIIWQMIGGQTTEQGDCSKFKGQVSPHCCKKDPVAVDLIPGAPYSEQV 126
PGW++GWTWA+ E+IW M+G Q T+QGDCS+FK + PHCC++ P VDL+PG PY++Q+
Sbjct: 63 PGWTVGWTWARQEVIWSMVGAQATDQGDCSRFKANL-PHCCRRTPAVVDLLPGVPYNQQI 121
Query: 127 ANCCKGGVLTSLKQDPANAVASFRVVVGRAGTTTKSVRLPKNFTFKSPGPGYTCGPARIV 186
ANCC+GGVL + Q P+ A A+F+V VG+AGTT ++VRLP+NFT PGPGYTCG AR+V
Sbjct: 122 ANCCRGGVLPAYGQAPSAAAAAFQVSVGQAGTTNRTVRLPRNFTLLGPGPGYTCGRARVV 181
Query: 187 KPTRFLRYDKRRVTQAMMSWNVTCTYSQFLAAKAPTCCVSLSTFYNDTIVSCPTCACGCQ 246
T FL D+RR TQA+M+WNVTCTYSQ LA+K P+CCVS S+FYNDTIV C CACGC
Sbjct: 182 PSTVFLTADRRRKTQALMTWNVTCTYSQHLASKYPSCCVSFSSFYNDTIVPCAKCACGCD 241
Query: 247 S 247
+
Sbjct: 242 A 242
>UniRef100_Q6VSV0 BRITTLE CULM1 [Oryza sativa]
Length = 468
Score = 334 bits (856), Expect = 3e-90
Identities = 149/240 (62%), Positives = 185/240 (77%), Gaps = 1/240 (0%)
Query: 6 LFRFVLPCTLLLLLPSSSEAYDPLDPNGNITIRWDILSWTGDGYVAAVTLNNFQQYRHIQ 65
L R L LL + S + AYDPLDP GNITI+WD++SWT DGYVA VT++N+Q YR I
Sbjct: 3 LHRCSLLALLLAVTCSVAVAYDPLDPKGNITIKWDVISWTPDGYVAMVTMSNYQMYRQIL 62
Query: 66 EPGWSLGWTWAKNEIIWQMIGGQTTEQGDCSKFKGQVSPHCCKKDPVAVDLIPGAPYSEQ 125
PGW++GW+WAK E+IW ++G Q TEQGDCSKFKG + PH CK+ P VDL+PG PY++Q
Sbjct: 63 APGWTVGWSWAKKEVIWSIVGAQATEQGDCSKFKGGI-PHSCKRTPAIVDLLPGVPYNQQ 121
Query: 126 VANCCKGGVLTSLKQDPANAVASFRVVVGRAGTTTKSVRLPKNFTFKSPGPGYTCGPARI 185
+ANCCK GV+++ QDPA +V++F+V VG AGTT K+V+LP NFT PGPGYTCGPA I
Sbjct: 122 IANCCKAGVVSAYGQDPAGSVSAFQVSVGLAGTTNKTVKLPTNFTLAGPGPGYTCGPATI 181
Query: 186 VKPTRFLRYDKRRVTQAMMSWNVTCTYSQFLAAKAPTCCVSLSTFYNDTIVSCPTCACGC 245
V T +L D+RR TQA+M+W VTCTYSQ LA++ PTCCVS S+FYN TIV C CACGC
Sbjct: 182 VPSTVYLTPDRRRRTQALMTWTVTCTYSQQLASRYPTCCVSFSSFYNSTIVPCARCACGC 241
>UniRef100_Q9SEC8 Phytochelatin synthetase-like protein [Zea mays]
Length = 407
Score = 275 bits (703), Expect = 2e-72
Identities = 122/186 (65%), Positives = 149/186 (79%), Gaps = 4/186 (2%)
Query: 84 MIGGQTTEQGDCSKFKGQVSPHCCKKDPVAVDLIPGAPYSEQVANCCKGGVLTSLKQDPA 143
M+G QTTEQGDCSKFK PHCCKKDP VDL+PG PY+ Q+ANCCK GV+ + QDPA
Sbjct: 1 MVGAQTTEQGDCSKFKSS-PPHCCKKDPTIVDLLPGTPYNMQIANCCKAGVVNTFNQDPA 59
Query: 144 NAVASFRVVVGRAGTTTKSVRLPKNFTFKSPGPGYTCGPARIVKPTRFLRYDKRRVTQAM 203
NA +SF++ VG AGTT K+V++P+NFT K+PGPGYTCG A + +PT+F D RR TQA+
Sbjct: 60 NAASSFQISVGLAGTTNKTVKVPRNFTLKTPGPGYTCGRAIVGRPTKFFTADGRRATQAL 119
Query: 204 MSWNVTCTYSQFLAAKAPTCCVSLSTFYNDTIVSCPTCACGCQS---GNCVDPNSPNLTS 260
M+WNVTCTYSQFLA K P+CCVSLS+FYNDTIV+CPTC+CGCQ+ NCV+ +SPNL +
Sbjct: 120 MTWNVTCTYSQFLAQKTPSCCVSLSSFYNDTIVNCPTCSCGCQNPSGSNCVNEDSPNLQA 179
Query: 261 VISNPG 266
I PG
Sbjct: 180 AIDGPG 185
>UniRef100_O04500 COBRA-like protein 6 precursor [Arabidopsis thaliana]
Length = 454
Score = 252 bits (644), Expect = 1e-65
Identities = 121/259 (46%), Positives = 161/259 (61%), Gaps = 10/259 (3%)
Query: 1 MDSLLLFRFVLPCTLL-------LLLPSSSEAYDPLDPNGNITIRWDILSWTGDGYVAAV 53
M +LLL V+ C++L ++ ++ YDPLDP G I I+WD+L + + V
Sbjct: 4 MLNLLLVVTVILCSILSPTRFMIMIDKMVADGYDPLDPFGKIIIKWDLLLSSPGQHHVQV 63
Query: 54 TLNNFQQYRHIQEPGWSLGWTWAKNEIIWQMIGGQTTEQGDCSKFKGQVS-PHCCKKDPV 112
TL N Q+YRH+++PGW L W W E+IW M G +TTEQG+CS F + PHCC + P
Sbjct: 64 TLENMQEYRHVEKPGWKLSWHWLNQEVIWDMKGAETTEQGNCSAFASSGNLPHCCLERPT 123
Query: 113 AVDLIPGAPYSEQVANCCKGGVLTSLKQDPANAVASFRVVVGRAGTTTKSVRLPKNFTFK 172
VDL+PGA + QVANCC+GGVLTS+ QD AN V++F + VG + + +P NF
Sbjct: 124 IVDLLPGASLNVQVANCCRGGVLTSMSQDHANHVSAFHMTVGSSPDGPEEFNMPSNFDIG 183
Query: 173 SPGPGYTCGPARIVKPTRFLRYDKRRVTQAMMSWNVTCTYSQFLAAKAPTCCVSLSTFYN 232
PGY+C A V PT+F RR TQA+ +W C YSQF ++ +P CCVSLS FY
Sbjct: 184 V--PGYSCDNATSVSPTKFSTDKGRRKTQALATWEAVCVYSQFRSSPSPKCCVSLSAFYY 241
Query: 233 DTIVSCPTCACGCQSGNCV 251
IV CPTC+CGC S +CV
Sbjct: 242 QNIVPCPTCSCGCSSSHCV 260
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.334 0.145 0.504
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 737,086,696
Number of Sequences: 2790947
Number of extensions: 30121201
Number of successful extensions: 86394
Number of sequences better than 10.0: 46
Number of HSP's better than 10.0 without gapping: 35
Number of HSP's successfully gapped in prelim test: 11
Number of HSP's that attempted gapping in prelim test: 86248
Number of HSP's gapped (non-prelim): 77
length of query: 434
length of database: 848,049,833
effective HSP length: 130
effective length of query: 304
effective length of database: 485,226,723
effective search space: 147508923792
effective search space used: 147508923792
T: 11
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.6 bits)
S2: 76 (33.9 bits)
Lotus: description of TM0338.9


