

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0338.8
(61 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
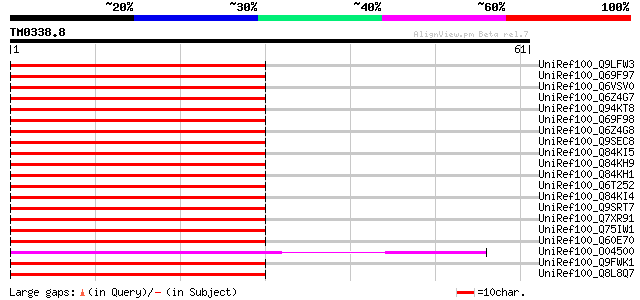
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9LFW3 COBRA-like protein 4 precursor [Arabidopsis tha... 67 2e-10
UniRef100_Q69F97 Phytochelatin synthetase-like protein [Phaseolu... 65 6e-10
UniRef100_Q6VSV0 BRITTLE CULM1 [Oryza sativa] 64 8e-10
UniRef100_Q6Z4G7 Putative phytochelatin synthetase [Oryza sativa] 64 1e-09
UniRef100_Q94KT8 COBRA protein precursor [Arabidopsis thaliana] 60 1e-08
UniRef100_Q69F98 Phytochelatin synthetase-like protein [Phaseolu... 60 2e-08
UniRef100_Q6Z4G8 Putative phytochelatin synthetase [Oryza sativa] 58 7e-08
UniRef100_Q9SEC8 Phytochelatin synthetase-like protein [Zea mays] 57 2e-07
UniRef100_Q84KI5 Phytochelatin synthetase-like protein 1 [Sorghu... 57 2e-07
UniRef100_Q84KH9 Phytochelatin synthetase [Hordeum vulgare var. ... 57 2e-07
UniRef100_Q84KH1 Phytochelatin synthetase [Triticum monococcum] 57 2e-07
UniRef100_Q6T252 Phytochelatin synthetase [Triticum aestivum] 57 2e-07
UniRef100_Q84KI4 Phytochelatin synthetase-like protein 2 [Sorghu... 56 3e-07
UniRef100_Q9SRT7 COBRA-like protein 1 precursor [Arabidopsis tha... 56 3e-07
UniRef100_Q7XR91 OSJNBa0011L07.10 protein [Oryza sativa] 55 4e-07
UniRef100_Q75IW1 Putative phytochelatin synthetase [Oryza sativa] 55 5e-07
UniRef100_Q60E70 Putative phytochelatin synthetase [Oryza sativa] 55 6e-07
UniRef100_O04500 COBRA-like protein 6 precursor [Arabidopsis tha... 54 8e-07
UniRef100_Q9FWK1 Hypothetical protein F21N10.4 [Arabidopsis thal... 54 1e-06
UniRef100_Q8L8Q7 COBRA-like protein 2 precursor [Arabidopsis tha... 54 1e-06
>UniRef100_Q9LFW3 COBRA-like protein 4 precursor [Arabidopsis thaliana]
Length = 431
Score = 66.6 bits (161), Expect = 2e-10
Identities = 26/30 (86%), Positives = 29/30 (96%)
Query: 1 KDKNTFSFKQGWAFPRKVYFNGDECMMPPP 30
KD+ TF+FKQGWAFPRKVYFNGDECM+PPP
Sbjct: 372 KDQKTFTFKQGWAFPRKVYFNGDECMLPPP 401
>UniRef100_Q69F97 Phytochelatin synthetase-like protein [Phaseolus vulgaris]
Length = 463
Score = 64.7 bits (156), Expect = 6e-10
Identities = 25/30 (83%), Positives = 29/30 (96%)
Query: 1 KDKNTFSFKQGWAFPRKVYFNGDECMMPPP 30
KDK+TF+ KQGWAFPRKVYFNG+ECM+PPP
Sbjct: 403 KDKDTFTLKQGWAFPRKVYFNGEECMLPPP 432
>UniRef100_Q6VSV0 BRITTLE CULM1 [Oryza sativa]
Length = 468
Score = 64.3 bits (155), Expect = 8e-10
Identities = 25/30 (83%), Positives = 27/30 (89%)
Query: 1 KDKNTFSFKQGWAFPRKVYFNGDECMMPPP 30
KD NTF+F QGWAFPRK+YFNGDEC MPPP
Sbjct: 406 KDYNTFTFSQGWAFPRKIYFNGDECKMPPP 435
>UniRef100_Q6Z4G7 Putative phytochelatin synthetase [Oryza sativa]
Length = 451
Score = 63.9 bits (154), Expect = 1e-09
Identities = 25/30 (83%), Positives = 27/30 (89%)
Query: 1 KDKNTFSFKQGWAFPRKVYFNGDECMMPPP 30
KD TF+F+QGWAFPRKVYFNGDEC MPPP
Sbjct: 389 KDARTFTFRQGWAFPRKVYFNGDECQMPPP 418
>UniRef100_Q94KT8 COBRA protein precursor [Arabidopsis thaliana]
Length = 456
Score = 60.1 bits (144), Expect = 1e-08
Identities = 21/30 (70%), Positives = 29/30 (96%)
Query: 1 KDKNTFSFKQGWAFPRKVYFNGDECMMPPP 30
KD++TF+F++GWAFPR++YFNGD C+MPPP
Sbjct: 394 KDQSTFTFEKGWAFPRRIYFNGDNCVMPPP 423
>UniRef100_Q69F98 Phytochelatin synthetase-like protein [Phaseolus vulgaris]
Length = 448
Score = 59.7 bits (143), Expect = 2e-08
Identities = 22/30 (73%), Positives = 27/30 (89%)
Query: 1 KDKNTFSFKQGWAFPRKVYFNGDECMMPPP 30
KDK TF+F +GWAFPR++YFNGD C+MPPP
Sbjct: 385 KDKATFTFDKGWAFPRRIYFNGDNCVMPPP 414
>UniRef100_Q6Z4G8 Putative phytochelatin synthetase [Oryza sativa]
Length = 446
Score = 57.8 bits (138), Expect = 7e-08
Identities = 21/30 (70%), Positives = 27/30 (90%)
Query: 1 KDKNTFSFKQGWAFPRKVYFNGDECMMPPP 30
KD +F+F++GWAFPR+VYFNGD C+MPPP
Sbjct: 382 KDPKSFTFEKGWAFPRRVYFNGDNCVMPPP 411
>UniRef100_Q9SEC8 Phytochelatin synthetase-like protein [Zea mays]
Length = 407
Score = 56.6 bits (135), Expect = 2e-07
Identities = 21/30 (70%), Positives = 26/30 (86%)
Query: 1 KDKNTFSFKQGWAFPRKVYFNGDECMMPPP 30
KD TF+F++GWAFPR+VYFNGD C+MP P
Sbjct: 297 KDSRTFTFEKGWAFPRRVYFNGDNCVMPSP 326
>UniRef100_Q84KI5 Phytochelatin synthetase-like protein 1 [Sorghum bicolor]
Length = 425
Score = 56.6 bits (135), Expect = 2e-07
Identities = 21/30 (70%), Positives = 26/30 (86%)
Query: 1 KDKNTFSFKQGWAFPRKVYFNGDECMMPPP 30
KD TF+F++GWAFPR+VYFNGD C+MP P
Sbjct: 386 KDSRTFTFEKGWAFPRRVYFNGDNCVMPAP 415
>UniRef100_Q84KH9 Phytochelatin synthetase [Hordeum vulgare var. distichum]
Length = 112
Score = 56.6 bits (135), Expect = 2e-07
Identities = 21/30 (70%), Positives = 26/30 (86%)
Query: 1 KDKNTFSFKQGWAFPRKVYFNGDECMMPPP 30
KD TF+F++GWAFPR+VYFNGD C+MP P
Sbjct: 78 KDSETFTFQKGWAFPRRVYFNGDNCVMPSP 107
>UniRef100_Q84KH1 Phytochelatin synthetase [Triticum monococcum]
Length = 457
Score = 56.6 bits (135), Expect = 2e-07
Identities = 21/30 (70%), Positives = 26/30 (86%)
Query: 1 KDKNTFSFKQGWAFPRKVYFNGDECMMPPP 30
KD TF+F++GWAFPR+VYFNGD C+MP P
Sbjct: 392 KDSETFTFQKGWAFPRRVYFNGDNCVMPSP 421
>UniRef100_Q6T252 Phytochelatin synthetase [Triticum aestivum]
Length = 456
Score = 56.6 bits (135), Expect = 2e-07
Identities = 21/30 (70%), Positives = 26/30 (86%)
Query: 1 KDKNTFSFKQGWAFPRKVYFNGDECMMPPP 30
KD TF+F++GWAFPR+VYFNGD C+MP P
Sbjct: 391 KDSETFTFQKGWAFPRRVYFNGDNCVMPSP 420
>UniRef100_Q84KI4 Phytochelatin synthetase-like protein 2 [Sorghum bicolor]
Length = 449
Score = 55.8 bits (133), Expect = 3e-07
Identities = 21/30 (70%), Positives = 25/30 (83%)
Query: 1 KDKNTFSFKQGWAFPRKVYFNGDECMMPPP 30
KD TF+F +GWAFPR+VYFNGD C+MP P
Sbjct: 385 KDSRTFTFDKGWAFPRRVYFNGDNCVMPSP 414
>UniRef100_Q9SRT7 COBRA-like protein 1 precursor [Arabidopsis thaliana]
Length = 452
Score = 55.8 bits (133), Expect = 3e-07
Identities = 19/30 (63%), Positives = 27/30 (89%)
Query: 1 KDKNTFSFKQGWAFPRKVYFNGDECMMPPP 30
K+ + F+F++GWAFPR++YFNGD C+MPPP
Sbjct: 389 KEASAFTFEKGWAFPRRIYFNGDNCVMPPP 418
>UniRef100_Q7XR91 OSJNBa0011L07.10 protein [Oryza sativa]
Length = 439
Score = 55.5 bits (132), Expect = 4e-07
Identities = 21/30 (70%), Positives = 26/30 (86%)
Query: 1 KDKNTFSFKQGWAFPRKVYFNGDECMMPPP 30
KDK+ F+F GWAFPR+VYF+G EC+MPPP
Sbjct: 375 KDKSDFTFSGGWAFPRRVYFDGHECVMPPP 404
>UniRef100_Q75IW1 Putative phytochelatin synthetase [Oryza sativa]
Length = 458
Score = 55.1 bits (131), Expect = 5e-07
Identities = 19/30 (63%), Positives = 27/30 (89%)
Query: 1 KDKNTFSFKQGWAFPRKVYFNGDECMMPPP 30
KD++TF+F +GWAFPR++YFNG+ C+MP P
Sbjct: 384 KDRSTFTFDKGWAFPRRIYFNGESCVMPSP 413
>UniRef100_Q60E70 Putative phytochelatin synthetase [Oryza sativa]
Length = 457
Score = 54.7 bits (130), Expect = 6e-07
Identities = 20/30 (66%), Positives = 24/30 (79%)
Query: 1 KDKNTFSFKQGWAFPRKVYFNGDECMMPPP 30
KD F+F +GWAFP +VYFNGD C+MPPP
Sbjct: 393 KDSKDFTFDKGWAFPHRVYFNGDNCVMPPP 422
>UniRef100_O04500 COBRA-like protein 6 precursor [Arabidopsis thaliana]
Length = 454
Score = 54.3 bits (129), Expect = 8e-07
Identities = 25/56 (44%), Positives = 32/56 (56%), Gaps = 12/56 (21%)
Query: 1 KDKNTFSFKQGWAFPRKVYFNGDECMMPPPTPTHFCQILLLQA*FPSRHSSSHYSS 56
KD F+F++GWAFPR++ FNGDEC+MP P FP S+H SS
Sbjct: 387 KDMGNFTFREGWAFPRRILFNGDECVMPSPDD------------FPRLPKSAHSSS 430
>UniRef100_Q9FWK1 Hypothetical protein F21N10.4 [Arabidopsis thaliana]
Length = 404
Score = 53.5 bits (127), Expect = 1e-06
Identities = 19/30 (63%), Positives = 26/30 (86%)
Query: 1 KDKNTFSFKQGWAFPRKVYFNGDECMMPPP 30
K+ F+F++GWAFPR++YFNGD C+MPPP
Sbjct: 343 KNPLEFTFEKGWAFPRRIYFNGDNCVMPPP 372
>UniRef100_Q8L8Q7 COBRA-like protein 2 precursor [Arabidopsis thaliana]
Length = 441
Score = 53.5 bits (127), Expect = 1e-06
Identities = 19/30 (63%), Positives = 26/30 (86%)
Query: 1 KDKNTFSFKQGWAFPRKVYFNGDECMMPPP 30
K+ F+F++GWAFPR++YFNGD C+MPPP
Sbjct: 380 KNPLEFTFEKGWAFPRRIYFNGDNCVMPPP 409
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.335 0.145 0.512
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 118,218,267
Number of Sequences: 2790947
Number of extensions: 3678403
Number of successful extensions: 9154
Number of sequences better than 10.0: 29
Number of HSP's better than 10.0 without gapping: 29
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 9125
Number of HSP's gapped (non-prelim): 29
length of query: 61
length of database: 848,049,833
effective HSP length: 37
effective length of query: 24
effective length of database: 744,784,794
effective search space: 17874835056
effective search space used: 17874835056
T: 11
A: 40
X1: 15 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.7 bits)
S2: 68 (30.8 bits)
Lotus: description of TM0338.8


