

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0338.2
(282 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
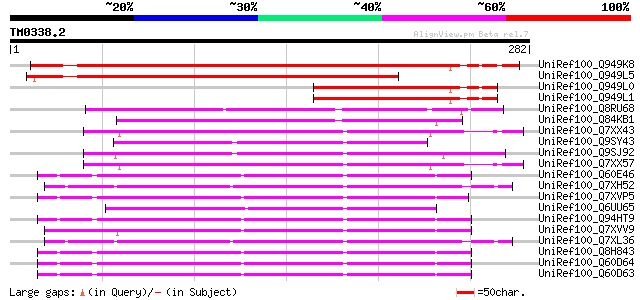
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q949K8 Gag polyprotein [Cicer arietinum] 269 5e-71
UniRef100_Q949L5 Gag polyprotein [Cicer arietinum] 192 8e-48
UniRef100_Q949L0 Putative gag polyprotein [Cicer arietinum] 120 5e-26
UniRef100_Q949L1 Putative gag polyprotein [Cicer arietinum] 115 1e-24
UniRef100_Q8RU68 Putative 22 kDa kafirin cluster; Ty3-Gypsy type... 108 1e-22
UniRef100_Q84KB1 Gag-protease polyprotein [Cucumis melo] 102 1e-20
UniRef100_Q7XX43 OSJNBa0059D20.6 protein [Oryza sativa] 100 3e-20
UniRef100_Q9SY43 Putative reverse transcriptase [Arabidopsis tha... 96 8e-19
UniRef100_Q9SJ92 Putative retroelement pol polyprotein [Arabidop... 96 1e-18
UniRef100_Q7XX57 OSJNBb0049I21.6 protein [Oryza sativa] 96 1e-18
UniRef100_Q60E46 Putative polyprotein [Oryza sativa] 95 2e-18
UniRef100_Q7XH52 Putative polyprotein [Oryza sativa] 95 2e-18
UniRef100_Q7XVP5 OSJNBa0023J03.1 protein [Oryza sativa] 94 3e-18
UniRef100_Q6UU65 Putative gag protein [Oryza sativa] 94 5e-18
UniRef100_Q94HT9 Putative retroelement [Oryza sativa] 94 5e-18
UniRef100_Q7XVV9 OSJNBa0065J03.13 protein [Oryza sativa] 93 7e-18
UniRef100_Q7XL36 OSJNBa0056L23.21 protein [Oryza sativa] 93 7e-18
UniRef100_Q8H843 Putative polyprotein [Oryza sativa] 93 7e-18
UniRef100_Q60D64 Putative polyprotein [Oryza sativa] 93 7e-18
UniRef100_Q60D63 Putative polyprotein [Oryza sativa] 93 7e-18
>UniRef100_Q949K8 Gag polyprotein [Cicer arietinum]
Length = 277
Score = 269 bits (688), Expect = 5e-71
Identities = 141/269 (52%), Positives = 182/269 (67%), Gaps = 16/269 (5%)
Query: 12 NQLAEMVATLVQAMTVQTNDNAQRRAAEDARELHRLQREAALDQNRGLNDFRRQDPPKFT 71
+Q+AE + + ++ QT AA+ R+L + RE ++RGL DFRR +PPKF
Sbjct: 16 DQMAEAMNNMAASVAAQT-------AAKTQRDLEKRGREIRAAESRGLVDFRRYNPPKFK 68
Query: 72 GGTDPDEADLWIQEIEKIFEVLQTSEGAKVGLATYLLLGDAEYWWRGARGMMEANHVEVN 131
G ++AD WIQE+EKIF+++ G KV ATY+LLGDAEYWWR AR +M A H EVN
Sbjct: 69 GDEGSEKADQWIQEVEKIFDMINCQAGVKVSYATYMLLGDAEYWWRSARLLMGAAHEEVN 128
Query: 132 WNSFRVAFLEKYFPDSARDERESQFLTLRQGSMTIPEYAAKLESLAKHFRFFRDQVDEPY 191
W SF+ FL+KYFP S R + FL L QGS+T+ EYAAK ESL++HFRFFR+++DEP+
Sbjct: 129 WESFKRKFLDKYFPMSTRTKLGDDFLKLHQGSLTVGEYAAKFESLSRHFRFFREEIDEPF 188
Query: 192 MCKRFVRGLRADIEDSVRPLGIMRFQALVEKATEVELMKNRRMDRAG---TGGPMRSSPR 248
MC RF GL+ +I+DSV PLGI RFQALVEK EVE MKN+R R G +GGP R+
Sbjct: 189 MCHRFQDGLKYEIQDSVLPLGIQRFQALVEKCREVEDMKNKRASRVGNFNSGGPSRA--- 245
Query: 249 SYQGKGKLQMKKPYQRPTGEGFTPGPYRP 277
+YQ +GK Q+ KPY RP + GP RP
Sbjct: 246 TYQNRGK-QVAKPYNRP--QNNNGGPSRP 271
>UniRef100_Q949L5 Gag polyprotein [Cicer arietinum]
Length = 209
Score = 192 bits (488), Expect = 8e-48
Identities = 100/205 (48%), Positives = 132/205 (63%), Gaps = 10/205 (4%)
Query: 10 NAN---QLAEMVATLVQAMTVQTNDNAQRRAAEDARELHRLQREAALDQNRGLNDFRRQD 66
NAN Q+AE + + ++ QT AA+ R+L + RE ++RGL DFRR +
Sbjct: 12 NANRNDQMAEAMNNMAASVAAQT-------AAKTQRDLEKRGREIRAAESRGLEDFRRYN 64
Query: 67 PPKFTGGTDPDEADLWIQEIEKIFEVLQTSEGAKVGLATYLLLGDAEYWWRGARGMMEAN 126
PPKF G ++AD WIQE+EKI ++++ G V ATY+LLGDAEYW R R +M A
Sbjct: 65 PPKFKGDESSEKADQWIQEVEKIIDMIKCQAGVTVSYATYMLLGDAEYWRRSGRLLMGAA 124
Query: 127 HVEVNWNSFRVAFLEKYFPDSARDERESQFLTLRQGSMTIPEYAAKLESLAKHFRFFRDQ 186
H EVNW SF+ FL+KYFP S R + FL L QGSMT+ EYAAK SL++HFRF R++
Sbjct: 125 HEEVNWESFKRKFLDKYFPMSTRTKLGDDFLKLHQGSMTVGEYAAKFGSLSRHFRFVREE 184
Query: 187 VDEPYMCKRFVRGLRADIEDSVRPL 211
DEP+M F GL+ +I+DSV PL
Sbjct: 185 TDEPFMIHCFQDGLKYEIQDSVLPL 209
>UniRef100_Q949L0 Putative gag polyprotein [Cicer arietinum]
Length = 115
Score = 120 bits (300), Expect = 5e-26
Identities = 62/103 (60%), Positives = 77/103 (74%), Gaps = 7/103 (6%)
Query: 166 IPEYAAKLESLAKHFRFFRDQVDEPYMCKRFVRGLRADIEDSVRPLGIMRFQALVEKATE 225
+ EYAAK ESL++HFRFFR+++DEP+MC RF GL+ +I+DSV PLGI RFQALVEK +
Sbjct: 1 VGEYAAKFESLSRHFRFFREEIDEPFMCHRFQDGLKYEIQDSVLPLGIQRFQALVEKCRK 60
Query: 226 VELMKNRRMDRAG---TGGPMRSSPRSYQGKGKLQMKKPYQRP 265
VE MKN+R R G +GGP R +YQ KGK Q+ KPY RP
Sbjct: 61 VEDMKNKRASRVGNFNSGGPSRV---NYQNKGK-QVAKPYNRP 99
>UniRef100_Q949L1 Putative gag polyprotein [Cicer arietinum]
Length = 116
Score = 115 bits (288), Expect = 1e-24
Identities = 61/103 (59%), Positives = 76/103 (73%), Gaps = 7/103 (6%)
Query: 166 IPEYAAKLESLAKHFRFFRDQVDEPYMCKRFVRGLRADIEDSVRPLGIMRFQALVEKATE 225
+ EYAAK ESL++HFRFFR++ EP+MC RF GL+ +I+DSV PLGI RFQALVEK E
Sbjct: 1 VGEYAAKFESLSRHFRFFREKDYEPFMCHRFQDGLKYEIQDSVLPLGIQRFQALVEKCRE 60
Query: 226 VELMKNRRMDRAG---TGGPMRSSPRSYQGKGKLQMKKPYQRP 265
VE MKN+R R G +GGP R+ +YQ +GK Q+ KPY RP
Sbjct: 61 VEDMKNKRASRVGNFNSGGPSRA---NYQNRGK-QVAKPYNRP 99
>UniRef100_Q8RU68 Putative 22 kDa kafirin cluster; Ty3-Gypsy type [Oryza sativa]
Length = 1230
Score = 108 bits (271), Expect = 1e-22
Identities = 74/233 (31%), Positives = 120/233 (50%), Gaps = 13/233 (5%)
Query: 42 RELHRLQREAALDQNR-GLNDFRRQDPPKFTGGTDPDEADLWIQEIEKIFEVLQTSEGAK 100
R++ L+R D NR GL +F++ PP F+G +P EA+ WI +EK FE + ++ K
Sbjct: 53 RQMEILERITKSDANRNGLGEFQKLKPPTFSGTANPLEAEEWIVAMEKSFEAMGCTDKEK 112
Query: 101 VGLATYLLLGDAEYWWRGARGMMEANHVEVNWNSFRVAFLEKYFPDSARDERESQFLTLR 160
+ ATY+L A WW A + + + W F+ AF +KYFP+S + +E +FL L+
Sbjct: 113 IIYATYMLQSSAFEWW-DAHKKSYSERIFITWELFKEAFYKKYFPESVKRMKEKEFLELK 171
Query: 161 QGSMTIPEYAAKLESLAKHFRFFRDQVD-EPYMCKRFVRGLRADIEDSVRPLGIMRFQAL 219
QG+ ++ EY + LA RF + V + +RF GLR ++ V + F+ +
Sbjct: 172 QGNKSVAEYEIEFSRLA---RFAPEFVQTDGSKARRFESGLRQPLKRRVEAFELTIFREV 228
Query: 220 VEKATEVELMKNRRMDRAGTGGPMR----SSPRSYQGKGKLQMKKPYQRPTGE 268
V KA +E K R G P + ++P++ QG+ + QR + E
Sbjct: 229 VSKAQLLE--KGYHEQRIEHGQPQKKFKTNNPQN-QGRFRGNYSGQMQRKSSE 278
>UniRef100_Q84KB1 Gag-protease polyprotein [Cucumis melo]
Length = 429
Score = 102 bits (254), Expect = 1e-20
Identities = 59/194 (30%), Positives = 93/194 (47%), Gaps = 9/194 (4%)
Query: 59 LNDFRRQDPPKFTGGT-DPDEADLWIQEIEKIFEVLQTSEGAKVGLATYLLLGDAEYWWR 117
L DFR+ +P F G DP A +W+ +E IF ++ E KV A ++L WW
Sbjct: 60 LRDFRKYNPTTFDGSLEDPTRAQMWLSSLETIFRYMKCPEDQKVQCAVFMLTDRGTAWWE 119
Query: 118 GARGMMEANHVEVNWNSFRVAFLEKYFPDSARDERESQFLTLRQGSMTIPEYAAKLESLA 177
M+ + ++ W F+ +F K+F S RD + +FL L QG MT+ +Y A+ + L+
Sbjct: 120 TTERMLGGDVSQITWQQFKESFYAKFFSASLRDAKRQEFLNLEQGDMTVEQYDAEFDMLS 179
Query: 178 KHFRFFRDQV-DEPYMCKRFVRGLRADIEDSVRPLGIMRFQALVEKATEVELMK----NR 232
RF + + E +FVRGLR DI+ VR + A ++ L + ++
Sbjct: 180 ---RFAPEMIATEAARADKFVRGLRLDIQGLVRAFRPATHADALRLAVDLSLQERANSSK 236
Query: 233 RMDRAGTGGPMRSS 246
R T G R +
Sbjct: 237 TAGRGSTSGQKRKA 250
>UniRef100_Q7XX43 OSJNBa0059D20.6 protein [Oryza sativa]
Length = 1470
Score = 100 bits (250), Expect = 3e-20
Identities = 70/246 (28%), Positives = 113/246 (45%), Gaps = 26/246 (10%)
Query: 41 ARELHRLQREAALDQNRG---LNDFRRQDPPKFTGGTDPDEADLWIQEIEKIFEVLQTSE 97
A + LQ A+ NRG +F R PP F +P +A+ W++ IEK +++ E
Sbjct: 27 ATQTQLLQAIASNQGNRGGSSFGEFMRTKPPTFATAEEPMDAEDWLRIIEKKLTLVRVRE 86
Query: 98 GAKVGLATYLLLGDAEYWWRGARGMMEANHVEVNWNSFRVAFLEKYFPDSARDERESQFL 157
KV A L G A WW + E + E NW F AF + + P + ++++F
Sbjct: 87 ADKVIFAANQLEGPAGDWWDTYKEAREEDAGEPNWEEFTTAFRDNFVPAAVMRMKKNEFR 146
Query: 158 TLRQGSMTIPEYAAKLESLAKHFRFFRDQVDEPYMCKRFVRGLRADIEDSVRPLGIMRFQ 217
LRQG+ T+ EY + LA++ RD DE +F+ GL ++ + FQ
Sbjct: 147 KLRQGNTTVQEYLNRFTQLARYAT--RDLADEEEKIDKFIEGLNDELRGPMIGQDHDSFQ 204
Query: 218 ALVEKATEVE----LMKNRRMDRAGTGGPMRSSPRSYQGKGKLQMKKPYQRPTGEGFTPG 273
+L+ K +E ++++ R R P +++P QRP +G TP
Sbjct: 205 SLINKVVRLENDRKVVEHNRKRRLAMNRPPQTAP---------------QRP--KGTTPS 247
Query: 274 PYRPTI 279
+RPT+
Sbjct: 248 AWRPTV 253
>UniRef100_Q9SY43 Putative reverse transcriptase [Arabidopsis thaliana]
Length = 839
Score = 96.3 bits (238), Expect = 8e-19
Identities = 58/171 (33%), Positives = 81/171 (46%), Gaps = 4/171 (2%)
Query: 57 RGLNDFRRQDPPKFTGGTDPDEADLWIQEIEKIFEVLQTSEGAKVGLATYLLLGDAEYWW 116
R + +R F+GGT P+EAD W ++E+ F + +V L + L GDA WW
Sbjct: 143 RMMKQLQRIGTEYFSGGTSPEEADSWRSQVERNFGSSRCPAEYRVDLTVHFLEGDAHLWW 202
Query: 117 RGARGMMEANHVEVNWNSFRVAFLEKYFPDSARDERESQFLTLRQGSMTIPEYAAKLESL 176
R +++W F F KYFP A D E++FL L QG ++ EY K L
Sbjct: 203 RSVTA--RRRQADMSWADFMAEFNAKYFPREALDRMEARFLELTQGVRSVREYDRKFNRL 260
Query: 177 AKHFRFFRDQVDEPYMCKRFVRGLRADIEDSVRPLGIMRFQALVEKATEVE 227
+ R D+ +RF+RGLR ++ R ALVE A EVE
Sbjct: 261 LVYAG--RGMEDDQAQMRRFLRGLRPNLRVRCRVSQYATKAALVETAAEVE 309
>UniRef100_Q9SJ92 Putative retroelement pol polyprotein [Arabidopsis thaliana]
Length = 1411
Score = 95.5 bits (236), Expect = 1e-18
Identities = 69/238 (28%), Positives = 103/238 (42%), Gaps = 13/238 (5%)
Query: 41 ARELHRLQREAALDQN-----RGLNDFRRQDPPKFTGGTDPDEADLWIQEIEKIFEVLQT 95
A + +Q+ AA+ + R + +R F+ GT P+EAD W +++ F +
Sbjct: 104 APRMAEVQQRAAVAEEVPSYLRMMEQLQRIGTWYFSSGTSPEEADSWRSRVQRNFGSSRC 163
Query: 96 SEGAKVGLATYLLLGDAEYWWRGARGMMEANHVEVNWNSFRVAFLEKYFPDSARDERESQ 155
+V LA + L GDA WWR +++W F F KYFP A D E++
Sbjct: 164 PAEYRVDLAVHFLEGDAHLWWRSVTA--RRRQADMSWADFVAEFNAKYFPQEALDRMEAR 221
Query: 156 FLTLRQGSMTIPEYAAKLESLAKHFRFFRDQVDEPYMCKRFVRGLRADIEDSVRPLGIMR 215
FL L QG ++ EY + L + R D+ +RF+RGLR D+ R
Sbjct: 222 FLELTQGEWSVREYDREFNRLLAYAG--RGMEDDQAQMRRFLRGLRPDLRVQCRVSQYAT 279
Query: 216 FQALVEKATEVELMKNRRM----DRAGTGGPMRSSPRSYQGKGKLQMKKPYQRPTGEG 269
ALVE A EVE R++ T + S GK K+ + P+ G
Sbjct: 280 KAALVETAAEVEEDLQRQVVGVSPAVQTKNTQQQVTPSKGGKPAQGQKRKWDHPSRAG 337
>UniRef100_Q7XX57 OSJNBb0049I21.6 protein [Oryza sativa]
Length = 1606
Score = 95.5 bits (236), Expect = 1e-18
Identities = 68/246 (27%), Positives = 111/246 (44%), Gaps = 26/246 (10%)
Query: 41 ARELHRLQREAALDQNRG---LNDFRRQDPPKFTGGTDPDEADLWIQEIEKIFEVLQTSE 97
A + LQ A NRG +F R PP +P +A+ W++ IEK +++ E
Sbjct: 2 ATQTQLLQAIANNQGNRGGSSFGEFMRTKPPTLATAEEPMDAEDWLRIIEKKLTLVRVRE 61
Query: 98 GAKVGLATYLLLGDAEYWWRGARGMMEANHVEVNWNSFRVAFLEKYFPDSARDERESQFL 157
KV A L G A WW + E + E NW F AF + + P + ++++F
Sbjct: 62 ADKVIFAANQLEGPAGDWWDTYKEAREEDAGEPNWEEFTTAFRDNFVPAAVMRMKKNEFR 121
Query: 158 TLRQGSMTIPEYAAKLESLAKHFRFFRDQVDEPYMCKRFVRGLRADIEDSVRPLGIMRFQ 217
LRQG+ T+ EY + LA++ RD DE +F+ GL ++ + + FQ
Sbjct: 122 RLRQGNTTVQEYLNRFTQLARYAT--RDLADEEEKIDKFIEGLNDELREPMIGQDHDSFQ 179
Query: 218 ALVEKATEVE----LMKNRRMDRAGTGGPMRSSPRSYQGKGKLQMKKPYQRPTGEGFTPG 273
+L+ K +E ++++ R R P +++P QRP +G T
Sbjct: 180 SLINKVVRLENDRKVVEHNRKRRLAMNRPPQTAP---------------QRP--KGTTTS 222
Query: 274 PYRPTI 279
+RPT+
Sbjct: 223 AWRPTV 228
>UniRef100_Q60E46 Putative polyprotein [Oryza sativa]
Length = 1850
Score = 95.1 bits (235), Expect = 2e-18
Identities = 67/236 (28%), Positives = 110/236 (46%), Gaps = 7/236 (2%)
Query: 16 EMVATLVQAMTVQTNDNAQRRAAEDARELHRLQREAALDQNRGLNDFRRQDPPKFTGGTD 75
+M+A ++Q M Q + +R + + H Q+ L +F R PP F+ T+
Sbjct: 284 QMMAAMMQQMQ-QQHQQMHQRMMQHVEQQH--QQFGPPPPQSKLPEFLRVRPPTFSSTTN 340
Query: 76 PDEADLWIQEIEKIFEVLQTSEGAKVGLATYLLLGDAEYWWRGARGMMEANHVEVNWNSF 135
P EA+ W+ IEK +LQ ++ KV AT+ L G A WW EV W F
Sbjct: 341 PMEANDWLHAIEKKLNLLQCNDQEKVAFATHQLQGPASAWWDNHMATRPPG-TEVTWAEF 399
Query: 136 RVAFLEKYFPDSARDERESQFLTLRQGSMTIPEYAAKLESLAKHFRFFRDQVDEPYMCKR 195
+F + PD +++ +F L QG+ T+ EY + LA++ D + ++
Sbjct: 400 CCSFRKAQVPDGVVAQKKREFRALHQGNRTVTEYLHEFNRLARYAP--EDVRTDAEKQEK 457
Query: 196 FVRGLRADIEDSVRPLGIMRFQALVEKATEVELMKNRRMDRAGTGGPMRSSPRSYQ 251
F+ GL ++ + + F+ LV+KA E +N +MDR R+ P S+Q
Sbjct: 458 FMAGLDDELTNQLISGDYADFERLVDKAIRQEDQRN-KMDRKRKAAQFRAPPGSHQ 512
>UniRef100_Q7XH52 Putative polyprotein [Oryza sativa]
Length = 371
Score = 95.1 bits (235), Expect = 2e-18
Identities = 75/254 (29%), Positives = 118/254 (45%), Gaps = 11/254 (4%)
Query: 20 TLVQAMTVQTNDNAQRRAAEDARELHRLQREAALDQNRGLNDFRRQDPPKFTGGTDPDEA 79
TL Q + QT + +L++ A QN+ L DF R PP F+ T+P EA
Sbjct: 19 TLAQVLAQQTQ-LMNMMIQQMQNQLNQGNNNAPPPQNK-LADFLRVRPPTFSSTTNPVEA 76
Query: 80 DLWIQEIEKIFEVLQTSEGAKVGLATYLLLGDAEYWWRGARGMMEANHVEVNWNSFRVAF 139
W+ +EK E+LQ ++ KV A++ L G A WW R M A + W F AF
Sbjct: 77 GDWLHAVEKKLELLQCTDQEKVVFASHQLQGPASEWWDHFR-MNRAEGQPITWADFTEAF 135
Query: 140 LEKYFPDSARDERESQFLTLRQGSMTIPEYAAKLESLAKHFRFFRDQVDEPYMCKRFVRG 199
+ + P ++ +F L+Q T+ EY + LA++ D + ++F+ G
Sbjct: 136 EKTHIPTGVVSLKKREFRALKQRDHTVAEYLHEFNRLARYAP--EDVRTDEERQEKFLEG 193
Query: 200 LRADIEDSVRPLGIMRFQALVEKATEVELMKNRRMDRAGTGGPMRSSPRSYQGKGKLQMK 259
L+ ++ ++ + FQ LV+KA +E KNR +R + + QG + Q
Sbjct: 194 LKDELSVTLISHDYVDFQQLVDKAIRLEDKKNRMDNRKRKMTVFQEA----QGSSQRQRI 249
Query: 260 KPYQRPTGEGFTPG 273
KP Q GE F+ G
Sbjct: 250 KPLQ--IGESFSTG 261
>UniRef100_Q7XVP5 OSJNBa0023J03.1 protein [Oryza sativa]
Length = 1827
Score = 94.4 bits (233), Expect = 3e-18
Identities = 68/234 (29%), Positives = 110/234 (46%), Gaps = 7/234 (2%)
Query: 16 EMVATLVQAMTVQTNDNAQRRAAEDARELHRLQREAALDQNRGLNDFRRQDPPKFTGGTD 75
+M+A ++Q M Q + +R + A + H Q+ L +F R PP F+ T+
Sbjct: 358 QMMAAMMQQMQ-QQHQQMHQRMMQHAEQQH--QQFGPPPPQSKLPEFLRVRPPTFSSTTN 414
Query: 76 PDEADLWIQEIEKIFEVLQTSEGAKVGLATYLLLGDAEYWWRGARGMMEANHVEVNWNSF 135
P EA+ W+ IEK +LQ ++ KV AT+ L G A WW EV W F
Sbjct: 415 PMEANDWLHAIEKKLNLLQCNDQEKVAFATHQLQGPASAWWDNHMATRPPG-TEVTWAEF 473
Query: 136 RVAFLEKYFPDSARDERESQFLTLRQGSMTIPEYAAKLESLAKHFRFFRDQVDEPYMCKR 195
+F + PD +++ +F L QG+MT+ EY + LA++ D + ++
Sbjct: 474 CRSFRKAQVPDGVVAQKKREFRALHQGNMTVTEYLHEFNRLARYAP--EDVRTDAEKQEK 531
Query: 196 FVRGLRADIEDSVRPLGIMRFQALVEKATEVELMKNRRMDRAGTGGPMRSSPRS 249
F+ GL ++ + + F+ LV+KA E +N +MDR RS+ S
Sbjct: 532 FMAGLDDELTNQLISGDYADFERLVDKAIRQEDQRN-KMDRKRKAAQFRSNQGS 584
>UniRef100_Q6UU65 Putative gag protein [Oryza sativa]
Length = 695
Score = 93.6 bits (231), Expect = 5e-18
Identities = 53/180 (29%), Positives = 89/180 (49%), Gaps = 3/180 (1%)
Query: 53 LDQNRGLNDFRRQDPPKFTGGTDPDEADLWIQEIEKIFEVLQTSEGAKVGLATYLLLGDA 112
+ Q L++F + +PP F DP EA W++ IEK ++Q ++ +VG AT+ L+G A
Sbjct: 70 VQQQCKLSEFLKTEPPTFAIAVDPMEASEWLRTIEKKLGLIQCTDQERVGFATHQLVGPA 129
Query: 113 EYWWRGARGMMEANHVEVNWNSFRVAFLEKYFPDSARDERESQFLTLRQGSMTIPEYAAK 172
WW H + WN F F + P R+ +F L+QG +T+ EY +
Sbjct: 130 SEWWDSYEASRPEGHT-ITWNEFSSVFRRSHVPAGMITLRKREFRYLKQGDLTVTEYLHE 188
Query: 173 LESLAKHFRFFRDQVDEPYMCKRFVRGLRADIEDSVRPLGIMRFQALVEKATEVELMKNR 232
LA+H D + ++F++GL+ ++ + F+ LV+KA ++E NR
Sbjct: 189 FYRLARHAP--EDVETDEEKQEKFLQGLKDELAVQLLLEDYENFEKLVDKAMQLESEHNR 246
>UniRef100_Q94HT9 Putative retroelement [Oryza sativa]
Length = 1714
Score = 93.6 bits (231), Expect = 5e-18
Identities = 67/236 (28%), Positives = 110/236 (46%), Gaps = 7/236 (2%)
Query: 16 EMVATLVQAMTVQTNDNAQRRAAEDARELHRLQREAALDQNRGLNDFRRQDPPKFTGGTD 75
+M+A ++Q M Q + +R + A + H Q+ L +F R PP F+ T+
Sbjct: 245 QMIAAMMQQMQ-QQHQQMHQRMMQHAEQQH--QQFGPPPPQSKLPEFLRVRPPTFSSTTN 301
Query: 76 PDEADLWIQEIEKIFEVLQTSEGAKVGLATYLLLGDAEYWWRGARGMMEANHVEVNWNSF 135
P EA+ W+ IEK +LQ ++ KV AT+ L G A WW EV W F
Sbjct: 302 PMEANDWLHAIEKKLNLLQCNDQEKVAFATHQLQGPASAWWDNHMATRPPG-TEVTWAEF 360
Query: 136 RVAFLEKYFPDSARDERESQFLTLRQGSMTIPEYAAKLESLAKHFRFFRDQVDEPYMCKR 195
+F + PD +++ +F L QG+ T+ EY + LA++ D + ++
Sbjct: 361 CRSFRKAQVPDGVVAQKKREFRALHQGNRTVTEYLHEFNRLARYAP--EDVRTDAEKQEK 418
Query: 196 FVRGLRADIEDSVRPLGIMRFQALVEKATEVELMKNRRMDRAGTGGPMRSSPRSYQ 251
F+ GL ++ + + F+ LV+KA E +N +MDR R+ S+Q
Sbjct: 419 FMAGLDDELTNQLISGDYADFERLVDKAIRQEDQRN-KMDRKRKAAQFRAHQGSHQ 473
>UniRef100_Q7XVV9 OSJNBa0065J03.13 protein [Oryza sativa]
Length = 759
Score = 93.2 bits (230), Expect = 7e-18
Identities = 69/240 (28%), Positives = 110/240 (45%), Gaps = 12/240 (5%)
Query: 20 TLVQAMTVQTNDNAQRRAAEDARELHRLQREAALDQNR--------GLNDFRRQDPPKFT 71
TL Q M QT A + + R+Q++ Q + L +F R PP F+
Sbjct: 350 TLAQVMAHQTQMMAAMMQQQHQQMHQRMQQQDEQQQQQFGPPPPQSKLPEFLRVRPPTFS 409
Query: 72 GGTDPDEADLWIQEIEKIFEVLQTSEGAKVGLATYLLLGDAEYWWRGARGMMEANHVEVN 131
T+P EA+ W+ IEK +LQ ++ KV AT+ L G A WW EV
Sbjct: 410 CTTNPMEANDWLHAIEKKLNLLQCNDQEKVAFATHQLQGSASAWWDNHMATRPPG-TEVT 468
Query: 132 WNSFRVAFLEKYFPDSARDERESQFLTLRQGSMTIPEYAAKLESLAKHFRFFRDQVDEPY 191
W F +F + PD+ +++ +F L QG+ T+ EY + LA++ D +
Sbjct: 469 WAEFCRSFRKAQVPDAVVAQKKREFRALHQGNRTVTEYLHEFNRLARYAP--EDVRTDAE 526
Query: 192 MCKRFVRGLRADIEDSVRPLGIMRFQALVEKATEVELMKNRRMDRAGTGGPMRSSPRSYQ 251
++F+ GL ++ + + F+ LV+KA E +N +MDR RS+ S+Q
Sbjct: 527 KQEKFLGGLDDELTNQLISGDYADFERLVDKAIRQEDQRN-KMDRKRKAAQFRSNQGSHQ 585
>UniRef100_Q7XL36 OSJNBa0056L23.21 protein [Oryza sativa]
Length = 449
Score = 93.2 bits (230), Expect = 7e-18
Identities = 75/254 (29%), Positives = 117/254 (45%), Gaps = 11/254 (4%)
Query: 20 TLVQAMTVQTNDNAQRRAAEDARELHRLQREAALDQNRGLNDFRRQDPPKFTGGTDPDEA 79
TL Q + QT + +L++ A QN+ L DF R PP F+ T+P EA
Sbjct: 99 TLAQVLAQQTQ-LMNMMIQQMQNQLNQGNNNAPPPQNK-LADFLRVRPPTFSSTTNPVEA 156
Query: 80 DLWIQEIEKIFEVLQTSEGAKVGLATYLLLGDAEYWWRGARGMMEANHVEVNWNSFRVAF 139
W+ IEK E+LQ ++ KV A++ L G A WW R M A + W F AF
Sbjct: 157 GDWLHAIEKKLELLQCTDQEKVVFASHQLQGPASEWWDHFR-MNRAEGQPITWVEFTEAF 215
Query: 140 LEKYFPDSARDERESQFLTLRQGSMTIPEYAAKLESLAKHFRFFRDQVDEPYMCKRFVRG 199
+ + P ++ +F L+Q T+ EY + LA++ D + ++F+ G
Sbjct: 216 KKTHIPTGVVSLKKREFRALKQKDQTVAEYLHEFNRLARYAP--EDVRTDEERQEKFLEG 273
Query: 200 LRADIEDSVRPLGIMRFQALVEKATEVELMKNRRMDRAGTGGPMRSSPRSYQGKGKLQMK 259
L+ ++ ++ + FQ LV+KA +E KNR +R + + QG + Q
Sbjct: 274 LKDELSVTLISHDYVDFQQLVDKAIRLEDKKNRMDNRKRKMTVFQEA----QGSSQRQRI 329
Query: 260 KPYQRPTGEGFTPG 273
KP Q GE + G
Sbjct: 330 KPLQ--IGESSSAG 341
>UniRef100_Q8H843 Putative polyprotein [Oryza sativa]
Length = 1796
Score = 93.2 bits (230), Expect = 7e-18
Identities = 67/236 (28%), Positives = 110/236 (46%), Gaps = 7/236 (2%)
Query: 16 EMVATLVQAMTVQTNDNAQRRAAEDARELHRLQREAALDQNRGLNDFRRQDPPKFTGGTD 75
+M+A ++Q M Q + +R + A + H Q+ L +F R PP F+ T+
Sbjct: 327 QMMAAMMQQMQ-QQHQQMHQRMMQHAEQQH--QQFGPPPPQSKLPEFLRVRPPTFSSTTN 383
Query: 76 PDEADLWIQEIEKIFEVLQTSEGAKVGLATYLLLGDAEYWWRGARGMMEANHVEVNWNSF 135
P EA+ W+ IEK +LQ ++ KV AT+ L G A WW EV W F
Sbjct: 384 PMEANDWLHAIEKKLNLLQCNDQEKVAFATHQLQGPASAWWDNHMATRPPG-TEVTWAEF 442
Query: 136 RVAFLEKYFPDSARDERESQFLTLRQGSMTIPEYAAKLESLAKHFRFFRDQVDEPYMCKR 195
+F + PD +++ +F L QG+ T+ EY + LA++ D + ++
Sbjct: 443 CRSFRKAQVPDGVVAQKKREFRALHQGNRTVTEYLQEFNRLARYAP--EDVRTDAEKQEK 500
Query: 196 FVRGLRADIEDSVRPLGIMRFQALVEKATEVELMKNRRMDRAGTGGPMRSSPRSYQ 251
F+ GL ++ + + F+ LV+KA E +N +MDR R+ S+Q
Sbjct: 501 FMAGLDDELTNQLISGDYADFERLVDKAIRQEDQRN-KMDRKRKAAQFRAPQGSHQ 555
>UniRef100_Q60D64 Putative polyprotein [Oryza sativa]
Length = 1777
Score = 93.2 bits (230), Expect = 7e-18
Identities = 67/236 (28%), Positives = 110/236 (46%), Gaps = 7/236 (2%)
Query: 16 EMVATLVQAMTVQTNDNAQRRAAEDARELHRLQREAALDQNRGLNDFRRQDPPKFTGGTD 75
+M+A ++Q M Q + +R + A + H Q+ L +F R PP F+ T+
Sbjct: 308 QMMAAMMQQMQ-QQHQQMHQRMMQHAEQQH--QQFGPPPPQSKLPEFLRVRPPTFSSTTN 364
Query: 76 PDEADLWIQEIEKIFEVLQTSEGAKVGLATYLLLGDAEYWWRGARGMMEANHVEVNWNSF 135
P EA+ W+ IEK +LQ ++ KV AT+ L G A WW EV W F
Sbjct: 365 PMEANDWLHAIEKKLNLLQCNDQEKVAFATHQLQGPASAWWDNHMATRPPG-TEVTWAEF 423
Query: 136 RVAFLEKYFPDSARDERESQFLTLRQGSMTIPEYAAKLESLAKHFRFFRDQVDEPYMCKR 195
+F + PD +++ +F L QG+ T+ EY + LA++ D + ++
Sbjct: 424 CRSFRKAQVPDGVVAQKKREFRALHQGNRTVTEYLHEFNGLARYAP--EDVRTDAEKQEK 481
Query: 196 FVRGLRADIEDSVRPLGIMRFQALVEKATEVELMKNRRMDRAGTGGPMRSSPRSYQ 251
F+ GL ++ + + F+ LV+KA E +N +MDR R+ S+Q
Sbjct: 482 FMAGLDDELTNQLISGDYADFERLVDKAIRQEDQRN-KMDRKRKAAQFRAPQGSHQ 536
>UniRef100_Q60D63 Putative polyprotein [Oryza sativa]
Length = 1765
Score = 93.2 bits (230), Expect = 7e-18
Identities = 67/236 (28%), Positives = 110/236 (46%), Gaps = 7/236 (2%)
Query: 16 EMVATLVQAMTVQTNDNAQRRAAEDARELHRLQREAALDQNRGLNDFRRQDPPKFTGGTD 75
+M+A ++Q M Q + +R + A + H Q+ L +F R PP F+ T+
Sbjct: 308 QMMAAMMQQMQ-QQHQQMHQRMMQHAEQQH--QQFGPPPPQSKLPEFLRVRPPTFSSTTN 364
Query: 76 PDEADLWIQEIEKIFEVLQTSEGAKVGLATYLLLGDAEYWWRGARGMMEANHVEVNWNSF 135
P EA+ W+ IEK +LQ ++ KV AT+ L G A WW EV W F
Sbjct: 365 PMEANDWLHAIEKKLNLLQCNDQEKVAFATHQLQGPASAWWDNHMATRPPG-TEVTWAEF 423
Query: 136 RVAFLEKYFPDSARDERESQFLTLRQGSMTIPEYAAKLESLAKHFRFFRDQVDEPYMCKR 195
+F + PD +++ +F L QG+ T+ EY + LA++ D + ++
Sbjct: 424 CRSFRKAQVPDGVVAQKKREFRALHQGNRTVTEYLHEFNGLARYAP--EDVRTDAEKQEK 481
Query: 196 FVRGLRADIEDSVRPLGIMRFQALVEKATEVELMKNRRMDRAGTGGPMRSSPRSYQ 251
F+ GL ++ + + F+ LV+KA E +N +MDR R+ S+Q
Sbjct: 482 FMAGLDDELTNQLISGDYADFERLVDKAIRQEDQRN-KMDRKRKAAQFRAPQGSHQ 536
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.319 0.134 0.394
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 466,157,085
Number of Sequences: 2790947
Number of extensions: 19540835
Number of successful extensions: 43300
Number of sequences better than 10.0: 566
Number of HSP's better than 10.0 without gapping: 416
Number of HSP's successfully gapped in prelim test: 151
Number of HSP's that attempted gapping in prelim test: 42382
Number of HSP's gapped (non-prelim): 601
length of query: 282
length of database: 848,049,833
effective HSP length: 126
effective length of query: 156
effective length of database: 496,390,511
effective search space: 77436919716
effective search space used: 77436919716
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 74 (33.1 bits)
Lotus: description of TM0338.2


