

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0336.12
(267 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
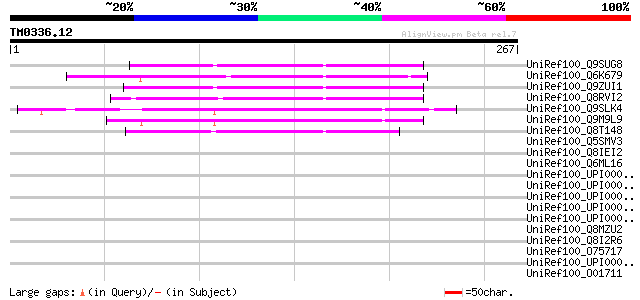
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9SUG8 Hypothetical protein T10C21.130 [Arabidopsis th... 115 2e-24
UniRef100_Q6K679 Hypothetical protein P0700F06.23 [Oryza sativa] 112 8e-24
UniRef100_Q9ZUI1 At2g24100/F27D4.1 [Arabidopsis thaliana] 108 1e-22
UniRef100_Q8RVI2 Hypothetical protein [Pinus pinaster] 108 2e-22
UniRef100_Q9SLK4 F20D21.12 protein [Arabidopsis thaliana] 98 3e-19
UniRef100_Q9M9L9 F10A16.6 protein [Arabidopsis thaliana] 91 2e-17
UniRef100_Q8T148 Similar to Dictyostelium discoideum (Slime mold... 74 5e-12
UniRef100_Q5SMV3 Hypothetical protein P0470C02.5-1 [Oryza sativa] 45 0.002
UniRef100_Q8IEI2 Hypothetical protein MAL13P1.66 [Plasmodium fal... 35 1.5
UniRef100_Q6ML16 ABC-type transporter [Bdellovibrio bacteriovorus] 35 2.0
UniRef100_UPI0000452BD9 UPI0000452BD9 UniRef100 entry 35 2.6
UniRef100_UPI000042F312 UPI000042F312 UniRef100 entry 35 2.6
UniRef100_UPI000030AED5 UPI000030AED5 UniRef100 entry 34 4.5
UniRef100_UPI0000437153 UPI0000437153 UniRef100 entry 33 5.8
UniRef100_UPI000024816C UPI000024816C UniRef100 entry 33 5.8
UniRef100_Q8MZU2 Gal/GalNAc lectin heavy subunit region A [Entam... 33 5.8
UniRef100_Q8I2R6 Hypothetical protein PFI1185c [Plasmodium falci... 33 5.8
UniRef100_O75717 WD repeat and HMG-box DNA binding protein 1 [Ho... 33 5.8
UniRef100_UPI00003451DE UPI00003451DE UniRef100 entry 33 7.6
UniRef100_O01711 Hypothetical protein Y6G8.2 [Caenorhabditis ele... 33 7.6
>UniRef100_Q9SUG8 Hypothetical protein T10C21.130 [Arabidopsis thaliana]
Length = 589
Score = 115 bits (287), Expect = 2e-24
Identities = 61/156 (39%), Positives = 89/156 (56%), Gaps = 4/156 (2%)
Query: 64 KFAPATFYAKLLNVGKYNFIPTTVNQLVVKCQFARRKLIWEILDSSKNKKFKAEVPWDRI 123
K + F A LL +G++ + LV KC FA+ KL+WE+L+ + K K E+ W I
Sbjct: 135 KLKASNFPASLLKIGQWEYKSRYEGDLVAKCYFAKHKLVWEVLE--QGLKSKIEIQWSDI 192
Query: 124 SAIRAFIEENKTEILEIELDEQPIFHKEINSQST-HTTWEISDHDFTGGHAMTYRIHHLE 182
A++A E+ L + L QP+F +E N Q HT W+ + DFT G A R H L+
Sbjct: 193 MALKANCPEDGPGTLTLVLARQPLFFRETNPQPRKHTLWQATS-DFTDGQASMNRQHFLQ 251
Query: 183 FASGDLGQHYENMLKCDERLMKLSQQPFPRLDSPYF 218
A G + +H+E +++CD RL LS+QP +DSPYF
Sbjct: 252 CAQGIMNKHFEKLVQCDHRLFHLSRQPEIAIDSPYF 287
>UniRef100_Q6K679 Hypothetical protein P0700F06.23 [Oryza sativa]
Length = 523
Score = 112 bits (281), Expect = 8e-24
Identities = 70/197 (35%), Positives = 107/197 (53%), Gaps = 11/197 (5%)
Query: 31 PLGLK-RTTSAMKEREKMWRKMQKSKKQNSSTTLKFAP-----ATFYAKLLNVGKYNFIP 84
PLGL+ R + ++ E +M M+ +KK++ + A + F A L +G +
Sbjct: 99 PLGLRLRKSPSLLELIQMKLAMENTKKEDIKSRSLIASERVKASNFAADFLKIGTWECTS 158
Query: 85 TTVNQLVVKCQFARRKLIWEILDSSKNKKFKAEVPWDRISAIRAFIEENKTEILEIELDE 144
LV KC FA+ KL+WE+LD+ +K E+ W I A++A EN L++ L
Sbjct: 159 QYEGDLVAKCYFAKHKLVWEVLDAGLKRKI--EIQWSDIIALKATCPENGIGTLDLVLAR 216
Query: 145 QPIFHKEINSQST-HTTWEISDHDFTGGHAMTYRIHHLEFASGDLGQHYENMLKCDERLM 203
P F KE + Q HT W+++ DFTGG A R H L+ S L +++E +++CD+RL
Sbjct: 217 PPTFFKETDPQPRKHTLWQVAS-DFTGGQASIKRRHILQCQSSLLSKNFEKLIQCDQRLN 275
Query: 204 KLSQQPFPRLDSPYFGP 220
LS QP+ +DSP F P
Sbjct: 276 YLSLQPY-MIDSPVFRP 291
>UniRef100_Q9ZUI1 At2g24100/F27D4.1 [Arabidopsis thaliana]
Length = 466
Score = 108 bits (270), Expect = 1e-22
Identities = 58/159 (36%), Positives = 90/159 (56%), Gaps = 4/159 (2%)
Query: 61 TTLKFAPATFYAKLLNVGKYNFIPTTVNQLVVKCQFARRKLIWEILDSSKNKKFKAEVPW 120
T K + F A +L +G++ + LV KC FA+ KL+WE+L+ + K K E+ W
Sbjct: 105 TVEKLKASNFPATILRIGQWEYKSRYEGDLVAKCYFAKHKLVWEVLE--QGLKSKIEIQW 162
Query: 121 DRISAIRAFIEENKTEILEIELDEQPIFHKEINSQST-HTTWEISDHDFTGGHAMTYRIH 179
I A++A + E++ L I L +P+F +E N Q HT W+ + DFT G A R H
Sbjct: 163 SDIMALKANLPEDEPGTLTIVLARRPLFFRETNPQPRKHTLWQATS-DFTDGQASMNRQH 221
Query: 180 HLEFASGDLGQHYENMLKCDERLMKLSQQPFPRLDSPYF 218
L+ G + +H+E +++CD RL LS+QP L +P+F
Sbjct: 222 FLQCPPGIMNKHFEKLVQCDHRLFCLSRQPEINLAAPFF 260
>UniRef100_Q8RVI2 Hypothetical protein [Pinus pinaster]
Length = 197
Score = 108 bits (269), Expect = 2e-22
Identities = 64/166 (38%), Positives = 90/166 (53%), Gaps = 6/166 (3%)
Query: 54 SKKQNSSTTLKFAPATFYAKLLNVGKYNFIPTTVNQLVVKCQFARRKLIWEILDSSKNKK 113
S QN+ LK + F L +G + I LV KC FA+ KL+WE+LD K
Sbjct: 8 SAAQNNPDKLK--ASNFPVSNLRIGTWECISRYEGDLVAKCYFAKHKLVWEVLDGGL--K 63
Query: 114 FKAEVPWDRISAIRAFIEENKTEILEIELDEQPIFHKEINSQST-HTTWEISDHDFTGGH 172
K E+ W I+A++A E++ L+IE+ P+F +E N Q HT W+ + DFTGG
Sbjct: 64 SKIEIQWSDITALKASYLEDEPGTLDIEVSRPPLFFRETNPQPRKHTLWQATS-DFTGGQ 122
Query: 173 AMTYRIHHLEFASGDLGQHYENMLKCDERLMKLSQQPFPRLDSPYF 218
A R H L+ G L +HYE +++CD RL LS++ F SP F
Sbjct: 123 ATICRRHFLQCPHGLLNRHYEKLIQCDPRLNLLSKKGFLSEASPLF 168
>UniRef100_Q9SLK4 F20D21.12 protein [Arabidopsis thaliana]
Length = 444
Score = 97.8 bits (242), Expect = 3e-19
Identities = 75/240 (31%), Positives = 113/240 (46%), Gaps = 27/240 (11%)
Query: 5 PPFGLELSSST--MNEREPHLRNVCPKPPLGLKRTTSAMKEREKMWRKMQKSKKQNSSTT 62
P L L+ S +N+ E +L C P + T +R + + +K K N
Sbjct: 6 PHLNLSLTKSPELINKIESYLNGHCTCP----HQQTENSSKRSTLPKSPEKLKAMN---- 57
Query: 63 LKFAPATFYAKLLNVGKYNFIPTTVNQLVVKCQFARRKLIWEIL-----DSSKNKKFKAE 117
F + +G + + + +V K FA++KLIWE L ++ K K E
Sbjct: 58 -------FPISTIRIGGWVVVAKNPDDIVAKFYFAKKKLIWEFLFGEPETNTLRLKRKIE 110
Query: 118 VPWDRISAIRAFIEE-NKTEILEIELDEQPIFHKEINSQS-THTTWEISDHDFTGGHAMT 175
+ W+ +S+ I ++T IL+IEL ++P F E N Q+ HT W+ DHDFTG HA
Sbjct: 111 IQWNDVSSFEESISSRDETGILKIELKKRPTFFIETNPQAGKHTQWKQLDHDFTGDHASN 170
Query: 176 YRIHHLEFASGDLGQHYENMLKCDERLMKLSQQPFPRLDSPYFGPPVLFKNIRDYNTVQH 235
YR H L F G L ++ E ++ D KL + PFP +S YF F+N N+ H
Sbjct: 171 YRRHTLHFPPGVLQKNLEKLV-TDSFWSKLYEVPFPVHESRYFDSG--FENNSSRNSHSH 227
>UniRef100_Q9M9L9 F10A16.6 protein [Arabidopsis thaliana]
Length = 410
Score = 91.3 bits (225), Expect = 2e-17
Identities = 60/179 (33%), Positives = 93/179 (51%), Gaps = 13/179 (7%)
Query: 52 QKSKKQNSSTTLKFAPA-----TFYAKLLNVGKYNFIPTTVNQLVVKCQFARRKLIWEIL 106
Q+++ + ++TL +P F + +G F+ + +V K FA++KL+WE L
Sbjct: 53 QQTENSSKTSTLPKSPEKLKAMNFPISTIKIGDCVFVAKNPDDIVAKFYFAKKKLLWEFL 112
Query: 107 -----DSSKNKKFKAEVPWDRISAIRAFIEE-NKTEILEIELDEQPIFHKEINSQS-THT 159
+ K K E+ W+ +S+ I ++T IL+IEL ++P F E N Q+ HT
Sbjct: 113 FGEPVANMPRLKSKIEIQWNDVSSFEESINSRDETGILKIELKKRPTFFTETNPQAGKHT 172
Query: 160 TWEISDHDFTGGHAMTYRIHHLEFASGDLGQHYENMLKCDERLMKLSQQPFPRLDSPYF 218
W+ D+DFTG A YR H L F G L ++ E +L D KL + PFP +S YF
Sbjct: 173 QWKQLDYDFTGDQASYYRRHTLHFPPGVLQKNLEKLL-TDSFWSKLYKVPFPVHESLYF 230
>UniRef100_Q8T148 Similar to Dictyostelium discoideum (Slime mold).
Homeobox-containing protein [Dictyostelium discoideum]
Length = 1108
Score = 73.6 bits (179), Expect = 5e-12
Identities = 44/146 (30%), Positives = 76/146 (51%), Gaps = 5/146 (3%)
Query: 62 TLKFAPATFYAKLLNVGKYNFIPTTVNQLVVKCQFARRKLIWEILDSSKNKKFKAEVPWD 121
T K P+ A + VG++ + ++ K F +++++WE+L K K + +
Sbjct: 476 TSKLKPSNLAATKIKVGEWECSASFPGDIIAKFYFTKKQIVWELL--KLGLKSKMVISFS 533
Query: 122 RISAIRAFIEENKTEILEIELDEQPIFHKEINSQ-STHTTWEISDHDFTGGH-AMTYRIH 179
I+A++ + + L IE+ + P F+KE+N Q +TTW +S+ DFT A TYR H
Sbjct: 534 EITAVKVEDTSDDSATLSIEISKAPKFYKEVNPQPKKNTTWNLSN-DFTDNQSASTYRRH 592
Query: 180 HLEFASGDLGQHYENMLKCDERLMKL 205
L F L ++ + K D++L KL
Sbjct: 593 ILTFKKSSLAKNIVKLHKSDQKLKKL 618
>UniRef100_Q5SMV3 Hypothetical protein P0470C02.5-1 [Oryza sativa]
Length = 299
Score = 45.1 bits (105), Expect = 0.002
Identities = 22/44 (50%), Positives = 30/44 (68%)
Query: 177 RIHHLEFASGDLGQHYENMLKCDERLMKLSQQPFPRLDSPYFGP 220
R H L+ S L +++E +L+CD+RL +LSQQP LDSP F P
Sbjct: 3 RRHFLQCPSSLLSKNFEKLLQCDQRLNQLSQQPDIILDSPVFEP 46
>UniRef100_Q8IEI2 Hypothetical protein MAL13P1.66 [Plasmodium falciparum]
Length = 2900
Score = 35.4 bits (80), Expect = 1.5
Identities = 24/86 (27%), Positives = 42/86 (47%), Gaps = 4/86 (4%)
Query: 79 KYNFIPTTVNQLVVKCQFARRKLIWEILDSSKNKKFKAEVPWDRISAIRAFIE----ENK 134
KY T NQ ++K ++K I + SKNK F+ + + I+ + IE E K
Sbjct: 2750 KYKLEQTNNNQYILKIIAIKKKKIHKTFSLSKNKHFRKCLEIELINTEKKNIENEMYERK 2809
Query: 135 TEILEIELDEQPIFHKEINSQSTHTT 160
+L+ ++ + +FHK+ +TT
Sbjct: 2810 ENVLDKKIYGKQVFHKKSFDNLYNTT 2835
>UniRef100_Q6ML16 ABC-type transporter [Bdellovibrio bacteriovorus]
Length = 466
Score = 35.0 bits (79), Expect = 2.0
Identities = 19/87 (21%), Positives = 42/87 (47%), Gaps = 2/87 (2%)
Query: 33 GLKRTTSAMKEREKMWRKMQKSKKQNSSTTLKFAPATFY--AKLLNVGKYNFIPTTVNQL 90
G+ + + +E+ W+++ +S+ QN S L + ++L++ +F+P N+L
Sbjct: 43 GVTTALKSKESKEETWKQLLQSQAQNISAELGELKGSLMKAGQMLSMYGEHFLPPEANEL 102
Query: 91 VVKCQFARRKLIWEILDSSKNKKFKAE 117
+ Q L WE ++ + K+ E
Sbjct: 103 LKSLQHDSPPLSWEAIEPTLKKQLPPE 129
>UniRef100_UPI0000452BD9 UPI0000452BD9 UniRef100 entry
Length = 577
Score = 34.7 bits (78), Expect = 2.6
Identities = 23/113 (20%), Positives = 50/113 (43%), Gaps = 23/113 (20%)
Query: 37 TTSAMKEREKMWRK----MQKSKKQNSSTTLKFAPATFYAKLLNVGKYNFIPTTVNQLVV 92
T+ MK + +W + +Q + + + ++++ +LLN+ + IP + QL+
Sbjct: 464 TSELMKRDKNLWTRGLYMLQNTDRSELESIVRWS-----CRLLNISNFRNIPIQIGQLLA 518
Query: 93 KCQFARRKLIWEILDSSKNKKFKAEVPWDRISAIRAFIEENKTEILEIELDEQ 145
K + I I +S+ +KK + + N E++ IEL ++
Sbjct: 519 KKSTSATPKITHIFESASSKKISS--------------QNNSQEVISIELHDR 557
>UniRef100_UPI000042F312 UPI000042F312 UniRef100 entry
Length = 684
Score = 34.7 bits (78), Expect = 2.6
Identities = 23/113 (20%), Positives = 50/113 (43%), Gaps = 23/113 (20%)
Query: 37 TTSAMKEREKMWRK----MQKSKKQNSSTTLKFAPATFYAKLLNVGKYNFIPTTVNQLVV 92
T+ MK + +W + +Q + + + ++++ +LLN+ + IP + QL+
Sbjct: 464 TSELMKRDKNLWTRGLYMLQNTDRSELESIVRWS-----CRLLNISNFRNIPIQIGQLLA 518
Query: 93 KCQFARRKLIWEILDSSKNKKFKAEVPWDRISAIRAFIEENKTEILEIELDEQ 145
K + I I +S+ +KK + + N E++ IEL ++
Sbjct: 519 KKSTSATPKITHIFESASSKKISS--------------QNNSQEVISIELHDR 557
>UniRef100_UPI000030AED5 UPI000030AED5 UniRef100 entry
Length = 250
Score = 33.9 bits (76), Expect = 4.5
Identities = 31/119 (26%), Positives = 54/119 (45%), Gaps = 11/119 (9%)
Query: 42 KEREKMWRK----MQKSKKQNSSTTLKFAPATFYAKLLNVGKYNFIPTTVNQLVVKCQFA 97
K ++K+W+ K ST + F+ ++ L N F+P T ++
Sbjct: 35 KYKQKIWQNSLNIFLKLPDPIKSTIMIFSGKRSFSNLFNDYNVKFLPKTQ---LIDLNLK 91
Query: 98 RRKLIWEILDSSKNKKFKAEVPWDRISAIRAFIEENKTEILEIELDEQPIFHKEINSQS 156
R+K + D N KF E+ DR+ F E +T I +I DE+ I K++N+++
Sbjct: 92 RKKTNFNKKD---NSKFFIEIYNDRVIVANVFGEFYETYIDDI-TDEKKIIFKKLNTEN 146
>UniRef100_UPI0000437153 UPI0000437153 UniRef100 entry
Length = 528
Score = 33.5 bits (75), Expect = 5.8
Identities = 43/173 (24%), Positives = 65/173 (36%), Gaps = 33/173 (19%)
Query: 108 SSKNKKFKAEVPWDRISAIRAFIE----ENKTEILEIELDEQPIFHKEINSQSTHTTWEI 163
SS+ K + E+ DR S E E+K + LEIELDE+ + +N + T ++
Sbjct: 312 SSRLKNMETELQTDRSSQTDRSREIRLLEDKVKTLEIELDEEKSGAELLNERITRCREQV 371
Query: 164 SD------------HDFT-GGHAMTYRIHHLEFASGDLG----------------QHYEN 194
HD A+ +I L+ D+G Q E+
Sbjct: 372 DQLRSELMQERSARHDLEMDKSALERQIKELKSRIADMGTQSRPSAGVTMLENKVQELED 431
Query: 195 MLKCDERLMKLSQQPFPRLDSPYFGPPVLFKNIRDYNTVQHDELMEQVMALDR 247
L+ +ER Q RLD R+ + Q D+L +V AL R
Sbjct: 432 RLRSEEREKNTIQAAQRRLDRKLKDVTATLDQERNQHAEQRDQLSLRVKALKR 484
>UniRef100_UPI000024816C UPI000024816C UniRef100 entry
Length = 517
Score = 33.5 bits (75), Expect = 5.8
Identities = 43/173 (24%), Positives = 65/173 (36%), Gaps = 33/173 (19%)
Query: 108 SSKNKKFKAEVPWDRISAIRAFIE----ENKTEILEIELDEQPIFHKEINSQSTHTTWEI 163
SS+ K + E+ DR S E E+K + LEIELDE+ + +N + T ++
Sbjct: 300 SSRLKNMETELQTDRSSQTDRSREIRLLEDKVKTLEIELDEEKSGAELLNERITRCREQV 359
Query: 164 SD------------HDFT-GGHAMTYRIHHLEFASGDLG----------------QHYEN 194
HD A+ +I L+ D+G Q E+
Sbjct: 360 DQLRSELMQERSARHDLEMDKSALERQIKELKSRIADMGTQSRPSAGVTMLENKVQELED 419
Query: 195 MLKCDERLMKLSQQPFPRLDSPYFGPPVLFKNIRDYNTVQHDELMEQVMALDR 247
L+ +ER Q RLD R+ + Q D+L +V AL R
Sbjct: 420 RLRSEEREKNTIQAAQRRLDRKLKDVTATLDQERNQHAEQRDQLSLRVKALKR 472
>UniRef100_Q8MZU2 Gal/GalNAc lectin heavy subunit region A [Entamoeba histolytica]
Length = 194
Score = 33.5 bits (75), Expect = 5.8
Identities = 24/107 (22%), Positives = 51/107 (47%), Gaps = 24/107 (22%)
Query: 87 VNQLVVKCQFARRKLIWEILDSSKNKKFKAEVPWDRISAIRAFIEENKTEILEIE----- 141
+N+ +K F R++ + I K EV WD + ++E + E ++I
Sbjct: 93 INKTNIKQDFCRKEYAYPIE--------KYEVDWDNVP-----VDEQRIESVDINGKTCF 139
Query: 142 --LDEQPIFHKEINSQSTHTT----WEISDHDFTGGHAMTYRIHHLE 182
++P+ + +N++ T+ T +++ DF GG ++T+R + E
Sbjct: 140 KYAAKRPLAYVYLNTKMTYATKTEAYDVCRMDFIGGRSITFRSFNTE 186
>UniRef100_Q8I2R6 Hypothetical protein PFI1185c [Plasmodium falciparum]
Length = 698
Score = 33.5 bits (75), Expect = 5.8
Identities = 18/66 (27%), Positives = 35/66 (52%), Gaps = 3/66 (4%)
Query: 79 KYNFIPTTVNQLVVKCQFARRKLIWEILDSSKNKKFKAEVPWDRISAIRAFIEENKTEIL 138
K N++ TT+NQ V KC+ + L + + +K V +I + A+++ K I+
Sbjct: 631 KSNYLETTINQDVSKCKIVNKNLKHPM---KGDNLYKEIVELKKIKSNHAYVQIRKKYIM 687
Query: 139 EIELDE 144
++ LD+
Sbjct: 688 KVSLDD 693
>UniRef100_O75717 WD repeat and HMG-box DNA binding protein 1 [Homo sapiens]
Length = 1129
Score = 33.5 bits (75), Expect = 5.8
Identities = 17/58 (29%), Positives = 28/58 (47%)
Query: 11 LSSSTMNEREPHLRNVCPKPPLGLKRTTSAMKEREKMWRKMQKSKKQNSSTTLKFAPA 68
LS +T NE+ P ++ + PKP S ++R K ++ K++N L PA
Sbjct: 949 LSRTTNNEKSPIIKPLIPKPKPKQASAASYFQKRNSQTNKTEEVKEENLKNVLSETPA 1006
>UniRef100_UPI00003451DE UPI00003451DE UniRef100 entry
Length = 280
Score = 33.1 bits (74), Expect = 7.6
Identities = 18/49 (36%), Positives = 27/49 (54%), Gaps = 2/49 (4%)
Query: 122 RISAIRAFIEENK--TEILEIELDEQPIFHKEINSQSTHTTWEISDHDF 168
+I I F EENK +++L +++ P + E +S H W ISD DF
Sbjct: 14 QIGPIYVFFEENKKVSDLLNSIINKSPDYDFESFDKSVHKLWCISDKDF 62
>UniRef100_O01711 Hypothetical protein Y6G8.2 [Caenorhabditis elegans]
Length = 964
Score = 33.1 bits (74), Expect = 7.6
Identities = 37/175 (21%), Positives = 77/175 (43%), Gaps = 18/175 (10%)
Query: 46 KMWRKMQKSKKQNSSTTLKFAPATFYAKL---LNVGKYNFIPTTVNQLVVKCQFARRKLI 102
+ W ++ K +N ST LKF +AK+ ++ Y+ + T + + + +R L
Sbjct: 284 EQWLNGKEVKMKNLSTDLKFDAFFHFAKVSTGMDPNTYSEL-TLLRDVTAVPNYIQRALT 342
Query: 103 WEILDSSKNKK--FKAEVPWDRISAIRAFIEENKTEILEIELDEQPIFHKEINSQSTHTT 160
+ LD +KN + +K W I + E E+ + + + + H E +S
Sbjct: 343 TQTLDETKNIRISYKPRGDWCIIES------ERMQEVSQSPVCKVSVLHLEYLLRSIPVL 396
Query: 161 W-----EISDHDFTGGHAMTYRIHH-LEFASGDLGQHYENMLKCDERLMKLSQQP 209
W I++ + G + + +IH L+ + + + LK + ++ LS++P
Sbjct: 397 WSFSIASITNLELDGSNEILLQIHKTLQDTVEAMLRERSSKLKVNTFVVTLSEKP 451
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.318 0.133 0.394
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 448,401,707
Number of Sequences: 2790947
Number of extensions: 18138077
Number of successful extensions: 58204
Number of sequences better than 10.0: 26
Number of HSP's better than 10.0 without gapping: 9
Number of HSP's successfully gapped in prelim test: 17
Number of HSP's that attempted gapping in prelim test: 58180
Number of HSP's gapped (non-prelim): 28
length of query: 267
length of database: 848,049,833
effective HSP length: 125
effective length of query: 142
effective length of database: 499,181,458
effective search space: 70883767036
effective search space used: 70883767036
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 73 (32.7 bits)
Lotus: description of TM0336.12


