

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0327.1
(91 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
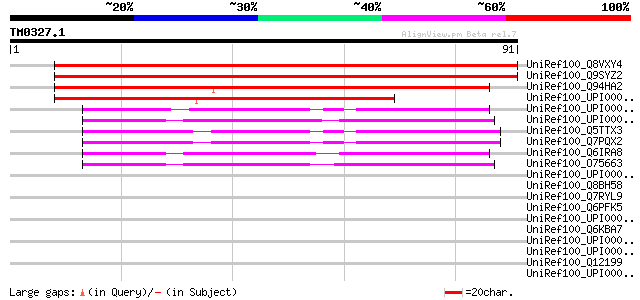
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q8VXY4 Hypothetical protein At4g34270 [Arabidopsis tha... 105 2e-22
UniRef100_Q9SYZ2 Hypothetical protein AT4g34270 [Arabidopsis tha... 105 2e-22
UniRef100_Q94HA2 Hypothetical protein OSJNBb0048A17.5 [Oryza sat... 96 2e-19
UniRef100_UPI00002EC848 UPI00002EC848 UniRef100 entry 51 6e-06
UniRef100_UPI00003A9B8B UPI00003A9B8B UniRef100 entry 48 5e-05
UniRef100_UPI000013D8D5 UPI000013D8D5 UniRef100 entry 47 1e-04
UniRef100_Q5TTX3 ENSANGP00000028166 [Anopheles gambiae str. PEST] 46 3e-04
UniRef100_Q7PQX2 ENSANGP00000021091 [Anopheles gambiae str. PEST] 46 3e-04
UniRef100_Q6IRA8 MGC79019 protein [Xenopus laevis] 45 4e-04
UniRef100_O75663 Hypothetical protein [Homo sapiens] 44 7e-04
UniRef100_UPI00001D0A77 UPI00001D0A77 UniRef100 entry 43 0.002
UniRef100_Q8BH58 CG9578-like (Mus musculus ES cells cDNA, RIKEN ... 43 0.002
UniRef100_Q7RYL9 Hypothetical protein [Neurospora crassa] 41 0.006
UniRef100_Q6PFK5 Hypothetical protein zgc:66327 [Brachydanio rerio] 40 0.011
UniRef100_UPI000023604F UPI000023604F UniRef100 entry 40 0.014
UniRef100_Q6KBA7 TipA protein [Emericella nidulans] 40 0.014
UniRef100_UPI000001A97E UPI000001A97E UniRef100 entry 39 0.041
UniRef100_UPI0000318EDE UPI0000318EDE UniRef100 entry 39 0.041
UniRef100_Q12199 Hypothetical protein YPR037W [Saccharomyces cer... 38 0.053
UniRef100_UPI000042FFBA UPI000042FFBA UniRef100 entry 35 0.34
>UniRef100_Q8VXY4 Hypothetical protein At4g34270 [Arabidopsis thaliana]
Length = 290
Score = 105 bits (263), Expect = 2e-22
Identities = 52/83 (62%), Positives = 63/83 (75%)
Query: 9 LVLMQLRVDGVLIRLRDTRMHCVFGGSTNPVILRESCWRESTFQALSAKGTPFDPAVYND 68
L+ LRVDGVL+RLR+TRMH FG P +LRE+CWRE+TFQ+LSAKG P D AV++D
Sbjct: 207 LLRFWLRVDGVLMRLRETRMHYRFGEDEAPTVLRENCWREATFQSLSAKGYPVDLAVWSD 266
Query: 69 PSIISQRLPIVKQMTQNLVISPK 91
PS ISQRLP++K TQ L I K
Sbjct: 267 PSSISQRLPVIKHTTQKLKIPSK 289
>UniRef100_Q9SYZ2 Hypothetical protein AT4g34270 [Arabidopsis thaliana]
Length = 231
Score = 105 bits (263), Expect = 2e-22
Identities = 52/83 (62%), Positives = 63/83 (75%)
Query: 9 LVLMQLRVDGVLIRLRDTRMHCVFGGSTNPVILRESCWRESTFQALSAKGTPFDPAVYND 68
L+ LRVDGVL+RLR+TRMH FG P +LRE+CWRE+TFQ+LSAKG P D AV++D
Sbjct: 148 LLRFWLRVDGVLMRLRETRMHYRFGEDEAPTVLRENCWREATFQSLSAKGYPVDLAVWSD 207
Query: 69 PSIISQRLPIVKQMTQNLVISPK 91
PS ISQRLP++K TQ L I K
Sbjct: 208 PSSISQRLPVIKHTTQKLKIPSK 230
>UniRef100_Q94HA2 Hypothetical protein OSJNBb0048A17.5 [Oryza sativa]
Length = 290
Score = 95.9 bits (237), Expect = 2e-19
Identities = 45/79 (56%), Positives = 58/79 (72%), Gaps = 1/79 (1%)
Query: 9 LVLMQLRVDGVLIRLRDTRMHCVFGGS-TNPVILRESCWRESTFQALSAKGTPFDPAVYN 67
L+ LRVDGVL+RLRDTR++C FG P++LRE CWRE+TF +LSAKG P D A Y
Sbjct: 208 LLRFWLRVDGVLMRLRDTRVYCSFGSDEAKPIVLRECCWREATFASLSAKGYPSDSAAYG 267
Query: 68 DPSIISQRLPIVKQMTQNL 86
DP++I+ +LP+V Q Q L
Sbjct: 268 DPNLIAHKLPVVMQKIQKL 286
>UniRef100_UPI00002EC848 UPI00002EC848 UniRef100 entry
Length = 228
Score = 51.2 bits (121), Expect = 6e-06
Identities = 29/64 (45%), Positives = 39/64 (60%), Gaps = 3/64 (4%)
Query: 9 LVLMQLRVDGVLIRLRDTRMHCVF---GGSTNPVILRESCWRESTFQALSAKGTPFDPAV 65
L+ LRVDGVLIRLR+TR C S +++RE+ RE T+ AL A+G P +P
Sbjct: 137 LLRFWLRVDGVLIRLRETRFFCDTTERDKSGAVIVVRETQHREETWDALRARGAPCEPRQ 196
Query: 66 YNDP 69
+ DP
Sbjct: 197 FPDP 200
>UniRef100_UPI00003A9B8B UPI00003A9B8B UniRef100 entry
Length = 256
Score = 48.1 bits (113), Expect = 5e-05
Identities = 32/73 (43%), Positives = 44/73 (59%), Gaps = 7/73 (9%)
Query: 14 LRVDGVLIRLRDTRMHCVFGGSTNPVILRESCWRESTFQALSAKGTPFDPAVYNDPSIIS 73
LRVDGVLIR+ DTR+H S +LRE RES +L K P P+++ +P+ IS
Sbjct: 182 LRVDGVLIRMNDTRLH---HESDKAYMLREYTSRESKISSL--KHVP--PSLFTEPNEIS 234
Query: 74 QRLPIVKQMTQNL 86
Q LPI + + + L
Sbjct: 235 QYLPIKETICEKL 247
>UniRef100_UPI000013D8D5 UPI000013D8D5 UniRef100 entry
Length = 273
Score = 46.6 bits (109), Expect = 1e-04
Identities = 28/74 (37%), Positives = 44/74 (58%), Gaps = 6/74 (8%)
Query: 14 LRVDGVLIRLRDTRMHCVFGGSTNPVILRESCWRESTFQALSAKGTPFDPAVYNDPSIIS 73
LR+DGVLIR+ DTR+ + + +LRE RES +L A P+++ +P+ IS
Sbjct: 187 LRIDGVLIRMNDTRL---YHEADKTYMLREYTSRESKISSLMA---TVPPSLFTEPNEIS 240
Query: 74 QRLPIVKQMTQNLV 87
Q LPI + + + L+
Sbjct: 241 QYLPIKEAVCEKLI 254
>UniRef100_Q5TTX3 ENSANGP00000028166 [Anopheles gambiae str. PEST]
Length = 273
Score = 45.8 bits (107), Expect = 3e-04
Identities = 29/75 (38%), Positives = 43/75 (56%), Gaps = 7/75 (9%)
Query: 14 LRVDGVLIRLRDTRMHCVFGGSTNPVILRESCWRESTFQALSAKGTPFDPAVYNDPSIIS 73
LRVD VL+R DTR H G N +L+E RE+ + L K P PA++ +P I
Sbjct: 204 LRVDNVLLRSNDTRFHYEKG---NDYVLKEFTHREAKVEQL--KHIP--PALFTNPDHIV 256
Query: 74 QRLPIVKQMTQNLVI 88
+LP+V+++ + L I
Sbjct: 257 DKLPVVRKVNEKLTI 271
>UniRef100_Q7PQX2 ENSANGP00000021091 [Anopheles gambiae str. PEST]
Length = 238
Score = 45.8 bits (107), Expect = 3e-04
Identities = 29/75 (38%), Positives = 43/75 (56%), Gaps = 7/75 (9%)
Query: 14 LRVDGVLIRLRDTRMHCVFGGSTNPVILRESCWRESTFQALSAKGTPFDPAVYNDPSIIS 73
LRVD VL+R DTR H G N +L+E RE+ + L K P PA++ +P I
Sbjct: 167 LRVDNVLLRSNDTRFHYEKG---NDYVLKEFTHREAKVEQL--KHIP--PALFTNPDHIV 219
Query: 74 QRLPIVKQMTQNLVI 88
+LP+V+++ + L I
Sbjct: 220 DKLPVVRKVNEKLTI 234
>UniRef100_Q6IRA8 MGC79019 protein [Xenopus laevis]
Length = 259
Score = 45.1 bits (105), Expect = 4e-04
Identities = 28/73 (38%), Positives = 42/73 (57%), Gaps = 7/73 (9%)
Query: 14 LRVDGVLIRLRDTRMHCVFGGSTNPVILRESCWRESTFQALSAKGTPFDPAVYNDPSIIS 73
LRVDGVLIR+ DTR+ + + +LRE +ES LS P +Y +P+ IS
Sbjct: 187 LRVDGVLIRMNDTRL---YHEADKTFMLREYTSKESKISNLS----HVPPPLYTEPNEIS 239
Query: 74 QRLPIVKQMTQNL 86
Q LP+++ + + L
Sbjct: 240 QYLPVIQTIYEKL 252
>UniRef100_O75663 Hypothetical protein [Homo sapiens]
Length = 272
Score = 44.3 bits (103), Expect = 7e-04
Identities = 27/74 (36%), Positives = 43/74 (57%), Gaps = 7/74 (9%)
Query: 14 LRVDGVLIRLRDTRMHCVFGGSTNPVILRESCWRESTFQALSAKGTPFDPAVYNDPSIIS 73
LR+DGVLIR+ DTR+ + + +LRE RES +L P+++ +P+ IS
Sbjct: 187 LRIDGVLIRMNDTRL---YHEADKTYMLREYTSRESKISSL----MHVPPSLFTEPNEIS 239
Query: 74 QRLPIVKQMTQNLV 87
Q LPI + + + L+
Sbjct: 240 QYLPIKEAVCEKLI 253
>UniRef100_UPI00001D0A77 UPI00001D0A77 UniRef100 entry
Length = 271
Score = 43.1 bits (100), Expect = 0.002
Identities = 28/74 (37%), Positives = 42/74 (55%), Gaps = 7/74 (9%)
Query: 14 LRVDGVLIRLRDTRMHCVFGGSTNPVILRESCWRESTFQALSAKGTPFDPAVYNDPSIIS 73
LR+DGVLIR+ DTR+ + + +LRE RES L P+++ +P+ IS
Sbjct: 187 LRIDGVLIRMNDTRL---YHEADKTYMLREYTSRESKIANL----MHVPPSLFTEPNEIS 239
Query: 74 QRLPIVKQMTQNLV 87
Q LPI + + + LV
Sbjct: 240 QYLPIKEAVCEKLV 253
>UniRef100_Q8BH58 CG9578-like (Mus musculus ES cells cDNA, RIKEN full-length enriched
library, clone:C330003J07 product:DJ69E11.3 (YEAST
YPR037W AND WORM C02C2.6 PREDICTED PROTEINS LIKE)
homolog) (Mus musculus 12 days embryo spinal cord cDNA,
RIKEN full-length enriched library, clone:C530049L15
product:DJ69E11.3 (YEAST YPR037W AND WORM C02C2.6
PREDICTED PROTEINS LIKE) homolog) (Mus musculus 10 day
old male pancreas cDNA, RIKEN full-length enriched
library, clone:1810011K17 product:DJ69E11.3 (YEAST
YPR037W AND WORM C02C2.6 PREDICTED PROTEINS LIKE)
homolog) [Mus musculus]
Length = 271
Score = 43.1 bits (100), Expect = 0.002
Identities = 28/74 (37%), Positives = 42/74 (55%), Gaps = 7/74 (9%)
Query: 14 LRVDGVLIRLRDTRMHCVFGGSTNPVILRESCWRESTFQALSAKGTPFDPAVYNDPSIIS 73
LR+DGVLIR+ DTR+ + + +LRE RES L P+++ +P+ IS
Sbjct: 187 LRIDGVLIRMNDTRL---YHEADKTYMLREYTSRESKIANL----MHVPPSLFTEPNEIS 239
Query: 74 QRLPIVKQMTQNLV 87
Q LPI + + + LV
Sbjct: 240 QYLPIKEAVCEKLV 253
>UniRef100_Q7RYL9 Hypothetical protein [Neurospora crassa]
Length = 322
Score = 41.2 bits (95), Expect = 0.006
Identities = 26/88 (29%), Positives = 46/88 (51%), Gaps = 9/88 (10%)
Query: 9 LVLMQLRVDGVLIRLRDTRMHCVFGGSTNPVILRESCWRESTFQALSAK----GTPFDPA 64
L M +R+D VL+R+RDTR++ F ++RE RE F+ + K G D
Sbjct: 208 LCRMFMRLDNVLVRVRDTRVYVDF---DKEEVIREYTEREDKFENVKTKLFMQGLKLDQV 264
Query: 65 --VYNDPSIISQRLPIVKQMTQNLVISP 90
D + ++ LP++KQ +++++ P
Sbjct: 265 TIALRDANQVAPLLPLIKQGVESVILGP 292
>UniRef100_Q6PFK5 Hypothetical protein zgc:66327 [Brachydanio rerio]
Length = 269
Score = 40.4 bits (93), Expect = 0.011
Identities = 28/63 (44%), Positives = 34/63 (53%), Gaps = 7/63 (11%)
Query: 14 LRVDGVLIRLRDTRMHCVFGGSTNPVILRESCWRESTFQALSAKGTPFDPAVYNDPSIIS 73
LRVDGVLIR+ DTR++ G +LRE RES L PAVY DP+ I+
Sbjct: 190 LRVDGVLIRINDTRLYHEAG---KKHMLREFSTRESNMADLQ----HIPPAVYTDPNEIA 242
Query: 74 QRL 76
L
Sbjct: 243 PHL 245
>UniRef100_UPI000023604F UPI000023604F UniRef100 entry
Length = 261
Score = 40.0 bits (92), Expect = 0.014
Identities = 26/78 (33%), Positives = 43/78 (54%), Gaps = 6/78 (7%)
Query: 14 LRVDGVLIRLRDTRMHCVFGGSTNPVILRESCWRESTFQALS---AKGTPFDPAVYNDPS 70
LR+D VL RLRDTR++ F S ++RE +E + + A PA+ DP+
Sbjct: 185 LRLDNVLFRLRDTRVYVDFEKSE---VIREYQSKECDYGIVRQKLASARDDIPAIMRDPN 241
Query: 71 IISQRLPIVKQMTQNLVI 88
+S+ LP+V + + +V+
Sbjct: 242 RLSELLPLVDRRLERVVL 259
>UniRef100_Q6KBA7 TipA protein [Emericella nidulans]
Length = 280
Score = 40.0 bits (92), Expect = 0.014
Identities = 26/78 (33%), Positives = 43/78 (54%), Gaps = 6/78 (7%)
Query: 14 LRVDGVLIRLRDTRMHCVFGGSTNPVILRESCWRESTFQALS---AKGTPFDPAVYNDPS 70
LR+D VL RLRDTR++ F S ++RE +E + + A PA+ DP+
Sbjct: 204 LRLDNVLFRLRDTRVYVDFEKSE---VIREYQSKECDYGIVRQKLASARDDIPAIMRDPN 260
Query: 71 IISQRLPIVKQMTQNLVI 88
+S+ LP+V + + +V+
Sbjct: 261 RLSELLPLVDRRLERVVL 278
>UniRef100_UPI000001A97E UPI000001A97E UniRef100 entry
Length = 260
Score = 38.5 bits (88), Expect = 0.041
Identities = 26/65 (40%), Positives = 37/65 (56%), Gaps = 4/65 (6%)
Query: 14 LRVDGVLIRLRDTRMHCVFGGSTNPVILRESCWRESTFQALSAKGTPFDPAVYNDPSIIS 73
LRVD VLIR+ DTR++ + +LRE RES + L + P A+Y DP+ I+
Sbjct: 187 LRVDRVLIRINDTRLYHEASRAGKNYMLREFSTRESKIEEL--QNVP--AALYTDPNEIA 242
Query: 74 QRLPI 78
Q L +
Sbjct: 243 QYLTL 247
>UniRef100_UPI0000318EDE UPI0000318EDE UniRef100 entry
Length = 245
Score = 38.5 bits (88), Expect = 0.041
Identities = 28/65 (43%), Positives = 38/65 (58%), Gaps = 7/65 (10%)
Query: 14 LRVDGVLIRLRDTRMHCVFGGSTNPVILRESCWRESTFQALSAKGTPFDPAVYNDPSIIS 73
LRVD VLIR+ DTR++ G + +LRE RES + L K P A+Y DP+ I+
Sbjct: 187 LRVDRVLIRINDTRLYHEAGKN---YMLREFSTRESKIEEL--KNVP--AALYTDPNEIA 239
Query: 74 QRLPI 78
Q L +
Sbjct: 240 QYLTL 244
>UniRef100_Q12199 Hypothetical protein YPR037W [Saccharomyces cerevisiae]
Length = 356
Score = 38.1 bits (87), Expect = 0.053
Identities = 26/75 (34%), Positives = 42/75 (55%), Gaps = 7/75 (9%)
Query: 14 LRVDGVLIRLRDTRMHCVFGGSTNPVILRESCWRESTFQALSAK---GTPFDP-AVYNDP 69
LRVD VL+R+ DTR++ F + V++RES E +Q + AK DP A D
Sbjct: 281 LRVDDVLVRVYDTRIYVEFDEN---VVIRESKEFEGKYQDVLAKHRLSQSHDPKAALRDS 337
Query: 70 SIISQRLPIVKQMTQ 84
+ ++Q P++K+ +
Sbjct: 338 NWVAQNTPMIKRQCE 352
>UniRef100_UPI000042FFBA UPI000042FFBA UniRef100 entry
Length = 393
Score = 35.4 bits (80), Expect = 0.34
Identities = 20/80 (25%), Positives = 42/80 (52%), Gaps = 7/80 (8%)
Query: 14 LRVDGVLIRLRDTRMHCVFGGSTNPVILRESCWRESTF----QALSAKGTPFDPAVYNDP 69
LR+D V+ R+RDTR + F +T ++RE +E + + K ++ D
Sbjct: 314 LRIDDVIFRVRDTRCYIDFETNT---LIREYKVQEQEYDKVMSLVKVKNNKDPKKLFRDS 370
Query: 70 SIISQRLPIVKQMTQNLVIS 89
+ +SQ +P++ + + +V++
Sbjct: 371 NWVSQNIPVLNKEIETIVVN 390
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.331 0.144 0.435
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 136,744,362
Number of Sequences: 2790947
Number of extensions: 4684844
Number of successful extensions: 11725
Number of sequences better than 10.0: 26
Number of HSP's better than 10.0 without gapping: 8
Number of HSP's successfully gapped in prelim test: 18
Number of HSP's that attempted gapping in prelim test: 11702
Number of HSP's gapped (non-prelim): 26
length of query: 91
length of database: 848,049,833
effective HSP length: 67
effective length of query: 24
effective length of database: 661,056,384
effective search space: 15865353216
effective search space used: 15865353216
T: 11
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.9 bits)
S2: 68 (30.8 bits)
Lotus: description of TM0327.1


