

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0323a.10
(175 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
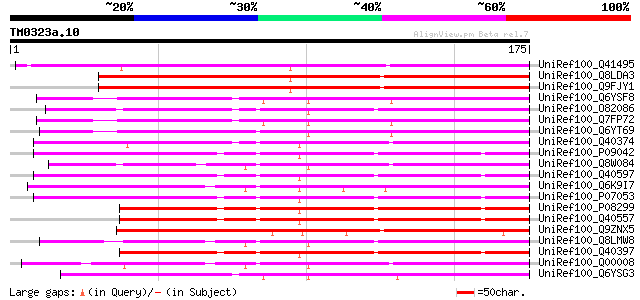
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q41495 STS14 protein precursor [Solanum tuberosum] 164 9e-40
UniRef100_Q8LDA3 Sts14 [Arabidopsis thaliana] 159 2e-38
UniRef100_Q9FJY1 Similarity to pathogenesis-related protein [Ara... 159 3e-38
UniRef100_Q6YSF8 PR-1 type pathogenesis-related protein PR-1a [O... 125 6e-28
UniRef100_O82086 Pathogenesis related protein-1 [Zea mays] 125 6e-28
UniRef100_Q7FP72 PR1a protein [Oryza sativa] 124 8e-28
UniRef100_Q6YT69 Pathogenesis-related protein 1 [Oryza sativa] 122 4e-27
UniRef100_Q40374 Pathogenesis-related protein PR-1 precursor [Me... 122 5e-27
UniRef100_P09042 Pathogenesis-related protein 1C precursor [Nico... 122 5e-27
UniRef100_Q8W084 Putative pathogenesis-related protein [Oryza sa... 120 1e-26
UniRef100_Q40597 Tobacco W38/1 PR-1 pathogenesis-related protein... 120 1e-26
UniRef100_Q6K9I7 Putative pathogenesis related protein-1 [Oryza ... 119 3e-26
UniRef100_P07053 Pathogenesis-related protein 1B precursor [Nico... 118 6e-26
UniRef100_P08299 Pathogenesis-related protein 1A precursor [Nico... 118 7e-26
UniRef100_Q40557 PR1a protein precursor [Nicotiana tabacum] 117 2e-25
UniRef100_Q9ZNX5 PR-1 like protein precursor [Camellia sinensis] 116 3e-25
UniRef100_Q8LMW8 Putative type-1 pathogenesis-related protein [O... 115 4e-25
UniRef100_Q40397 Pathogenesis-related protein-1 [Nicotiana gluti... 113 2e-24
UniRef100_Q00008 Pathogenesis-related protein PRMS precursor [Ze... 112 3e-24
UniRef100_Q6YSG3 Putative pathogenesis-related protein [Oryza sa... 112 4e-24
>UniRef100_Q41495 STS14 protein precursor [Solanum tuberosum]
Length = 214
Score = 164 bits (415), Expect = 9e-40
Identities = 90/180 (50%), Positives = 106/180 (58%), Gaps = 9/180 (5%)
Query: 3 IFMQPFFSLALALFLLSAEAATVLDSVVPQLSAE------AKEFLDAHNWARAEVGVEPL 56
+F Q F L A L A TV P SA A+EFLDAHN AR+EVGV PL
Sbjct: 37 LFFQ-FLLLTTASSLTHISAQTVPPPPPPPTSAATPPSRAAQEFLDAHNKARSEVGVGPL 95
Query: 57 EWSEALAKDAGLFARYQRDRLGCKLANFSHNKYGSNQ-DLGGMGWTPSVAVQDWVNEKEF 115
WS LAK+ L RYQRD+ C AN S+ KYG NQ G TP +AV WV EK+F
Sbjct: 96 TWSPMLAKETSLLVRYQRDKQNCSFANLSNGKYGGNQLWASGTVVTPRMAVDSWVAEKKF 155
Query: 116 YSYRTNSCVAPSFPCWHYTQVVWRKSKQVGCAQLTCVVDKLILTICFYDPPGNINGESPY 175
Y+Y NSC C YTQ+VW+KS ++GCAQ TC LT+CFY+PPGN+ GE PY
Sbjct: 156 YNYENNSCTGDD-KCGVYTQIVWKKSIELGCAQRTCYEGPATLTVCFYNPPGNVIGEKPY 214
>UniRef100_Q8LDA3 Sts14 [Arabidopsis thaliana]
Length = 185
Score = 159 bits (403), Expect = 2e-38
Identities = 78/147 (53%), Positives = 101/147 (68%), Gaps = 3/147 (2%)
Query: 31 PQLSAEAKEFLDAHNWARAEVGVEPLEWSEALAKDAGLFARYQRDRLGCKLANFSHNKYG 90
P +SA AK F DAHN ARA VGV PL WS+ L A ARYQR++ C+ A+ + KYG
Sbjct: 40 PTISAAAKAFTDAHNKARAMVGVPPLVWSQTLEAAASRLARYQRNQKKCEFASLNPGKYG 99
Query: 91 SNQ--DLGGMGWTPSVAVQDWVNEKEFYSYRTNSCVAPSFPCWHYTQVVWRKSKQVGCAQ 148
+NQ G + TPS+AV+ WV EK FY+Y++++C A + C Y QVVWR SK++GCAQ
Sbjct: 100 ANQLWAKGLVAVTPSLAVETWVKEKPFYNYKSDTCAA-NHTCGVYKQVVWRNSKELGCAQ 158
Query: 149 LTCVVDKLILTICFYDPPGNINGESPY 175
TC + +LTICFY+PPGNI G+ PY
Sbjct: 159 ATCTKESTVLTICFYNPPGNIIGQKPY 185
>UniRef100_Q9FJY1 Similarity to pathogenesis-related protein [Arabidopsis thaliana]
Length = 185
Score = 159 bits (402), Expect = 3e-38
Identities = 77/147 (52%), Positives = 101/147 (68%), Gaps = 3/147 (2%)
Query: 31 PQLSAEAKEFLDAHNWARAEVGVEPLEWSEALAKDAGLFARYQRDRLGCKLANFSHNKYG 90
P +SA AK F DAHN ARA VGV PL WS+ L A ARYQR++ C+ A+ + KYG
Sbjct: 40 PTISAAAKAFTDAHNKARAMVGVPPLVWSQTLEAAASRLARYQRNQKKCEFASLNPGKYG 99
Query: 91 SNQ--DLGGMGWTPSVAVQDWVNEKEFYSYRTNSCVAPSFPCWHYTQVVWRKSKQVGCAQ 148
+NQ G + TPS+AV+ WV EK FY+Y++++C A + C Y QVVWR SK++GCAQ
Sbjct: 100 ANQLWAKGLVAVTPSLAVETWVKEKPFYNYKSDTCAA-NHTCGVYKQVVWRNSKELGCAQ 158
Query: 149 LTCVVDKLILTICFYDPPGNINGESPY 175
TC + +LTICFY+PPGN+ G+ PY
Sbjct: 159 ATCTKESTVLTICFYNPPGNVIGQKPY 185
>UniRef100_Q6YSF8 PR-1 type pathogenesis-related protein PR-1a [Oryza sativa]
Length = 168
Score = 125 bits (313), Expect = 6e-28
Identities = 72/170 (42%), Positives = 97/170 (56%), Gaps = 14/170 (8%)
Query: 10 SLALALFLLSAEAATVLDSVVPQLSAEAKEFLDAHNWARAEVGVEPLEWSEALAKDAGLF 69
S L + +A AAT +S A++F+D HN ARA+VGV P+ W + +A A +
Sbjct: 9 SCCLLVLAAAAMAATAQNS--------AQDFVDPHNAARADVGVGPVSWDDTVAAYAESY 60
Query: 70 ARYQRDRLGCKLANF-SHNKYGSNQDLGGMG--WTPSVAVQDWVNEKEFYSYRTNSCVAP 126
A ++ CKL + S KYG N G G WT + AV WV+EK++Y + +NSC AP
Sbjct: 61 AAQRQG--DCKLEHSDSGGKYGENIFWGSAGGDWTAASAVSAWVSEKQWYDHGSNSCSAP 118
Query: 127 S-FPCWHYTQVVWRKSKQVGCAQLTCVVDKLILTICFYDPPGNINGESPY 175
C HYTQVVWR S +GCA++ C D + C Y PPGN G+SPY
Sbjct: 119 EGSSCGHYTQVVWRDSTAIGCARVVCDGDLGVFITCNYSPPGNFVGQSPY 168
>UniRef100_O82086 Pathogenesis related protein-1 [Zea mays]
Length = 163
Score = 125 bits (313), Expect = 6e-28
Identities = 72/165 (43%), Positives = 97/165 (58%), Gaps = 8/165 (4%)
Query: 13 LALFLLSAEAATVLDSVVPQLSAEAKEFLDAHNWARAEVGVEPLEWSEALAKDAGLFARY 72
LA L A AA V+ Q S + +++D HN ARA+VGV P+ W + +A A +A
Sbjct: 5 LACLLALAMAAIVVAPCTAQNSPQ--DYVDPHNAARADVGVGPVSWDDTVAAYAQSYAAQ 62
Query: 73 QRDRLGCKLANFSHNKYGSNQDLGGMG--WTPSVAVQDWVNEKEFYSYRTNSCVAPSFPC 130
++ CKL + S YG N G G W+ S AV WV+EK++Y + TNSC A C
Sbjct: 63 RQG--DCKLIH-SGGPYGENLFWGSAGADWSASDAVGSWVSEKQYYDHDTNSC-AEGQVC 118
Query: 131 WHYTQVVWRKSKQVGCAQLTCVVDKLILTICFYDPPGNINGESPY 175
HYTQVVWR S +GCA++ C + + IC Y+PPGN+ GESPY
Sbjct: 119 GHYTQVVWRDSTAIGCARVVCDNNAGVFIICSYNPPGNVVGESPY 163
>UniRef100_Q7FP72 PR1a protein [Oryza sativa]
Length = 168
Score = 124 bits (312), Expect = 8e-28
Identities = 72/170 (42%), Positives = 97/170 (56%), Gaps = 14/170 (8%)
Query: 10 SLALALFLLSAEAATVLDSVVPQLSAEAKEFLDAHNWARAEVGVEPLEWSEALAKDAGLF 69
S L + +A AAT +S A++F+D HN ARA+VGV P+ W + +A A +
Sbjct: 9 SCCLLVLAAAAMAATAQNS--------AQDFVDPHNAARADVGVGPVSWDDTVAAYAESY 60
Query: 70 ARYQRDRLGCKLANF-SHNKYGSNQDLGGMG--WTPSVAVQDWVNEKEFYSYRTNSCVAP 126
A ++ CKL + S KYG N G G WT + AV WV+EK++Y + +NSC AP
Sbjct: 61 AAQRQG--DCKLEHSDSGGKYGENIFWGSPGGDWTAASAVSAWVSEKQWYDHGSNSCSAP 118
Query: 127 S-FPCWHYTQVVWRKSKQVGCAQLTCVVDKLILTICFYDPPGNINGESPY 175
C HYTQVVWR S +GCA++ C D + C Y PPGN G+SPY
Sbjct: 119 EGSSCGHYTQVVWRDSTAIGCARVVCDGDLGVFITCNYSPPGNFVGQSPY 168
>UniRef100_Q6YT69 Pathogenesis-related protein 1 [Oryza sativa]
Length = 165
Score = 122 bits (306), Expect = 4e-27
Identities = 67/168 (39%), Positives = 97/168 (56%), Gaps = 12/168 (7%)
Query: 11 LALALFLLSAEAATVLDSVVPQLSAEAKEFLDAHNWARAEVGVEPLEWSEALAKDAGLFA 70
L A+ +A AAT +S A++++DAHN AR++VGV P+ W + +A A +A
Sbjct: 7 LVAAVLAAAAMAATAQNS--------AQDYVDAHNAARSDVGVGPVSWDDTVAAYAESYA 58
Query: 71 RYQRDRLGCKLANFSHNKYGSNQDLGGMG--WTPSVAVQDWVNEKEFYSYRTNSCVAPS- 127
++ + ++ S KYG N G G WT + AV WV EK++Y + +NSC AP+
Sbjct: 59 AQRQGDCALEHSD-SGGKYGENIFWGSAGGDWTAASAVSSWVAEKQWYDHDSNSCSAPAG 117
Query: 128 FPCWHYTQVVWRKSKQVGCAQLTCVVDKLILTICFYDPPGNINGESPY 175
C HYTQVVW S +GCA++ C + C Y PPGN++GESPY
Sbjct: 118 SSCGHYTQVVWSNSTAIGCARVVCDNSLGVFITCNYSPPGNVDGESPY 165
>UniRef100_Q40374 Pathogenesis-related protein PR-1 precursor [Medicago truncatula]
Length = 173
Score = 122 bits (305), Expect = 5e-27
Identities = 73/172 (42%), Positives = 96/172 (55%), Gaps = 9/172 (5%)
Query: 9 FSLALALFLLSAEAATVLDSVVPQLSAEAK----EFLDAHNWARAEVGVEPLEWSEALAK 64
FS AL L LL+ + + + S +++ +FL N ARA VG+ PL W + L
Sbjct: 6 FSFALFLLLLTFISHVSASYIPNKKSFKSRSFKNQFLIPQNIARAAVGLRPLVWDDKLTH 65
Query: 65 DAGLFARYQRDRLGCKLANFSHNKYGSNQDLG-GMGWTPSVAVQDWVNEKEFYSYRTNSC 123
A +A +R+ C L + S+ YG N G G+GW P+ AV WV+EK+FY+Y NSC
Sbjct: 66 YAQWYANQRRN--DCALEH-SNGPYGENIFWGSGVGWNPAQAVSAWVDEKQFYNYWHNSC 122
Query: 124 VAPSFPCWHYTQVVWRKSKQVGCAQLTCVVDKLILTICFYDPPGNINGESPY 175
V C HYTQVVW + +VGCA + C DK C YDPPGN GE PY
Sbjct: 123 VDGEM-CGHYTQVVWGSTTKVGCASVVCSDDKGTFMTCNYDPPGNYYGERPY 173
>UniRef100_P09042 Pathogenesis-related protein 1C precursor [Nicotiana tabacum]
Length = 168
Score = 122 bits (305), Expect = 5e-27
Identities = 72/168 (42%), Positives = 101/168 (59%), Gaps = 6/168 (3%)
Query: 9 FSLALALFLLSAEAATVLDSVVPQLSAEAKEFLDAHNWARAEVGVEPLEWSEALAKDAGL 68
FS + FL+S ++ S +++LDAHN ARA+VGVEPL W + +A A
Sbjct: 6 FSQMSSFFLVSTLLLFLIISHSCHAQNSQQDYLDAHNTARADVGVEPLTWDDQVAAYAQN 65
Query: 69 FARYQRDRLGCKLANFSHNKYGSNQDLG-GMGWTPSVAVQDWVNEKEFYSYRTNSCVAPS 127
+A + C L + SH +YG N G G T + AV+ WVNEK++Y++ +N+C A
Sbjct: 66 YA--SQLAADCNLVH-SHGQYGENLAWGSGDFLTAAKAVEMWVNEKQYYAHDSNTC-AQG 121
Query: 128 FPCWHYTQVVWRKSKQVGCAQLTCVVDKLILTICFYDPPGNINGESPY 175
C HYTQVVWR S +VGCA++ C I++ C YDPPGN+ G+SPY
Sbjct: 122 QVCGHYTQVVWRNSVRVGCARVQCNNGGYIVS-CNYDPPGNVIGKSPY 168
>UniRef100_Q8W084 Putative pathogenesis-related protein [Oryza sativa]
Length = 167
Score = 120 bits (301), Expect = 1e-26
Identities = 72/165 (43%), Positives = 95/165 (56%), Gaps = 10/165 (6%)
Query: 14 ALFLLSAEAATVLDSVVPQLSAEAKEFLDAHNWARAEVGVEPLEWSEALAKDAGLFARYQ 73
+LF+L+ AAT+ Q S + ++L N AR+ VGV P+ WS L G Y
Sbjct: 10 SLFVLAVVAATMFHCSDAQNSPQ--DYLSPQNAARSAVGVGPMSWSTKLQ---GFAEDYA 64
Query: 74 RDRLG-CKLANFSHNKYGSNQDLGGMG--WTPSVAVQDWVNEKEFYSYRTNSCVAPSFPC 130
R R G C+L + S YG N G G WT + AV+ WV+EK++Y+Y +NSC A C
Sbjct: 65 RQRKGDCRLQH-SGGPYGENIFWGSAGADWTAADAVRSWVDEKKYYNYASNSCAAGKV-C 122
Query: 131 WHYTQVVWRKSKQVGCAQLTCVVDKLILTICFYDPPGNINGESPY 175
HYTQVVWR S VGCA++ C ++ I IC Y+P GNI G PY
Sbjct: 123 GHYTQVVWRDSTNVGCARVRCDANRGIFIICNYEPRGNIVGRRPY 167
>UniRef100_Q40597 Tobacco W38/1 PR-1 pathogenesis-related protein [Nicotiana tabacum]
Length = 184
Score = 120 bits (301), Expect = 1e-26
Identities = 71/168 (42%), Positives = 99/168 (58%), Gaps = 6/168 (3%)
Query: 9 FSLALALFLLSAEAATVLDSVVPQLSAEAKEFLDAHNWARAEVGVEPLEWSEALAKDAGL 68
FS + FL+S ++ S+ +++LDAHN ARA+VGVEPL W + +A A
Sbjct: 6 FSQMPSFFLVSTLLLFLIISLSCGAQNSPQDYLDAHNTARADVGVEPLTWDDQVAAYAQN 65
Query: 69 FARYQRDRLGCKLANFSHNKYGSNQDLG-GMGWTPSVAVQDWVNEKEFYSYRTNSCVAPS 127
+A + C L + SH +YG N G G T + AV+ WVNEK++Y + +N+C A
Sbjct: 66 YA--SQLAADCMLVH-SHGQYGENLAWGSGDFMTAAKAVEMWVNEKQYYDHDSNTC-AQG 121
Query: 128 FPCWHYTQVVWRKSKQVGCAQLTCVVDKLILTICFYDPPGNINGESPY 175
C HYTQVVWR S +VGCA+ C +++ C YDPPGN G+SPY
Sbjct: 122 QVCGHYTQVVWRNSVRVGCARAQCNSGGYVVS-CNYDPPGNFVGQSPY 168
>UniRef100_Q6K9I7 Putative pathogenesis related protein-1 [Oryza sativa]
Length = 178
Score = 119 bits (298), Expect = 3e-26
Identities = 74/176 (42%), Positives = 92/176 (52%), Gaps = 11/176 (6%)
Query: 7 PFFSLALALFLLSAEAATVLDSVVPQLSAEAKEFLDAHNWARAEVGVEPLEWSEALAKDA 66
PF ++ + LL A S + + A FLDAHN AR +VGV PL W E LA A
Sbjct: 7 PFAAVTAVILLLHGAADAKSSSSSGKTKSLASGFLDAHNAARRQVGVPPLRWDERLASYA 66
Query: 67 GLFARYQRDRLG----CKLANFSHNKYGSNQDLG-GMGWTPSVAVQDWVN-EKEFYSYRT 120
ARY R G C L + SH YG N G G+GW P+ V WV+ E+ Y +
Sbjct: 67 ---ARYAAARSGAGGGCALVH-SHGPYGENLFHGSGVGWAPADVVAAWVSRERALYDAAS 122
Query: 121 NSCVA-PSFPCWHYTQVVWRKSKQVGCAQLTCVVDKLILTICFYDPPGNINGESPY 175
NSC + C HYTQVVWR++ VGCA TC + +C Y+PPGN G PY
Sbjct: 123 NSCRGGDAAACGHYTQVVWRRTTAVGCALATCAGGRGTYGVCSYNPPGNYVGVRPY 178
>UniRef100_P07053 Pathogenesis-related protein 1B precursor [Nicotiana tabacum]
Length = 168
Score = 118 bits (296), Expect = 6e-26
Identities = 69/168 (41%), Positives = 98/168 (58%), Gaps = 6/168 (3%)
Query: 9 FSLALALFLLSAEAATVLDSVVPQLSAEAKEFLDAHNWARAEVGVEPLEWSEALAKDAGL 68
FS + FL+S ++ S +++LDAHN ARA+VGVEPL W +A A
Sbjct: 6 FSQMPSFFLVSTLLLFLIISHSSHAQNSQQDYLDAHNTARADVGVEPLTWDNGVAAYAQN 65
Query: 69 FARYQRDRLGCKLANFSHNKYGSNQDLG-GMGWTPSVAVQDWVNEKEFYSYRTNSCVAPS 127
+ + C L + SH +YG N G G T + AV+ WV+EK++Y + +N+C A
Sbjct: 66 YV--SQLAADCNLVH-SHGQYGENLAQGSGDFMTAAKAVEMWVDEKQYYDHDSNTC-AQG 121
Query: 128 FPCWHYTQVVWRKSKQVGCAQLTCVVDKLILTICFYDPPGNINGESPY 175
C HYTQVVWR S +VGCA++ C +++ C YDPPGN+ G+SPY
Sbjct: 122 QVCGHYTQVVWRNSVRVGCARVKCNNGGYVVS-CNYDPPGNVIGQSPY 168
>UniRef100_P08299 Pathogenesis-related protein 1A precursor [Nicotiana tabacum]
Length = 168
Score = 118 bits (295), Expect = 7e-26
Identities = 65/139 (46%), Positives = 89/139 (63%), Gaps = 6/139 (4%)
Query: 38 KEFLDAHNWARAEVGVEPLEWSEALAKDAGLFARYQRDRLGCKLANFSHNKYGSNQDLG- 96
+++LDAHN ARA+VGVEPL W + +A A +A + C L + SH +YG N G
Sbjct: 35 QDYLDAHNTARADVGVEPLTWDDQVAAYAQNYA--SQLAADCNLVH-SHGQYGENLAEGS 91
Query: 97 GMGWTPSVAVQDWVNEKEFYSYRTNSCVAPSFPCWHYTQVVWRKSKQVGCAQLTCVVDKL 156
G T + AV+ WV+EK++Y + +N+C A C HYTQVVWR S +VGCA++ C
Sbjct: 92 GDFMTAAKAVEMWVDEKQYYDHDSNTC-AQGQVCGHYTQVVWRNSVRVGCARVQCNNGGY 150
Query: 157 ILTICFYDPPGNINGESPY 175
+++ C YDPPGN GESPY
Sbjct: 151 VVS-CNYDPPGNYRGESPY 168
>UniRef100_Q40557 PR1a protein precursor [Nicotiana tabacum]
Length = 168
Score = 117 bits (292), Expect = 2e-25
Identities = 64/139 (46%), Positives = 89/139 (63%), Gaps = 6/139 (4%)
Query: 38 KEFLDAHNWARAEVGVEPLEWSEALAKDAGLFARYQRDRLGCKLANFSHNKYGSNQDLG- 96
+++LDAHN ARA+VGVEPL W + +A A +A + C L + SH +YG N G
Sbjct: 35 QDYLDAHNTARADVGVEPLTWDDQVAAYAQNYA--SQLAADCNLVH-SHGQYGENLAEGS 91
Query: 97 GMGWTPSVAVQDWVNEKEFYSYRTNSCVAPSFPCWHYTQVVWRKSKQVGCAQLTCVVDKL 156
G T + AV+ WV+EK++Y + +N+C + C HYTQVVWR S +VGCA++ C
Sbjct: 92 GDFMTAAKAVEMWVDEKQYYDHDSNTC-SQGQVCGHYTQVVWRNSVRVGCARVQCNNGGY 150
Query: 157 ILTICFYDPPGNINGESPY 175
+++ C YDPPGN GESPY
Sbjct: 151 VVS-CNYDPPGNYRGESPY 168
>UniRef100_Q9ZNX5 PR-1 like protein precursor [Camellia sinensis]
Length = 191
Score = 116 bits (290), Expect = 3e-25
Identities = 66/143 (46%), Positives = 88/143 (61%), Gaps = 5/143 (3%)
Query: 37 AKEFLDAHNWARAEVGVEPLEWSEALAKDAGLFARYQRDRLGCKLANFSHN-KYGSNQDL 95
A++F+DAHN ARAEVGV+PL+WS +LA A RYQ++ + C+ A+ + +YGSNQ
Sbjct: 50 ARQFVDAHNSARAEVGVDPLKWSYSLANAASRLVRYQKNYMHCEFADMTGQLQYGSNQMW 109
Query: 96 GG-MGWTPSVAVQDWVNE-KEFYSYRTNSCVAPSFPCWHYTQVVWRKSKQVGCAQLTCVV 153
P V+ WVN K+ Y Y N CV + C Y QVVW K++ VGCAQ C
Sbjct: 110 SDYSAKPPREVVEYWVNSGKKHYRYTHNYCVR-NQNCGPYKQVVWEKTEMVGCAQGVCGN 168
Query: 154 DKLILTICFYDP-PGNINGESPY 175
+ L+ICFY P PGN+ G+ PY
Sbjct: 169 NNGSLSICFYYPHPGNLGGQRPY 191
>UniRef100_Q8LMW8 Putative type-1 pathogenesis-related protein [Oryza sativa]
Length = 176
Score = 115 bits (289), Expect = 4e-25
Identities = 68/168 (40%), Positives = 90/168 (53%), Gaps = 14/168 (8%)
Query: 11 LALALFLLSAEAATVLDSVVPQLSAEAKEFLDAHNWARAEVGVEPLEWSEALAKDAGLFA 70
LA+A+ L A T PQ ++++ HN AR GV P+ W +A A
Sbjct: 12 LAVAISLAMAATTTTSAQNTPQ------DYVNLHNSARRADGVGPVSWDPKVASFA---Q 62
Query: 71 RYQRDRLG-CKLANFSHNKYGSNQDLGGMG--WTPSVAVQDWVNEKEFYSYRTNSCVAPS 127
Y R G C+L + S YG N G G W+ + AV WV EK+ Y Y TN+C P
Sbjct: 63 SYAAKRAGDCRLQH-SGGPYGENIFWGSAGRAWSAADAVASWVGEKKNYHYDTNTC-DPG 120
Query: 128 FPCWHYTQVVWRKSKQVGCAQLTCVVDKLILTICFYDPPGNINGESPY 175
C HYTQVVWRKS ++GCA++ C ++ + C YDPPGN NGE P+
Sbjct: 121 KVCGHYTQVVWRKSVRIGCARVVCAANRGVFITCNYDPPGNFNGERPF 168
>UniRef100_Q40397 Pathogenesis-related protein-1 [Nicotiana glutinosa]
Length = 168
Score = 113 bits (283), Expect = 2e-24
Identities = 62/139 (44%), Positives = 87/139 (61%), Gaps = 6/139 (4%)
Query: 38 KEFLDAHNWARAEVGVEPLEWSEALAKDAGLFARYQRDRLGCKLANFSHNKYGSNQDLG- 96
+++L+AHN ARA+VGVEPL W + +A A + + C L SH +YG N +G
Sbjct: 35 QDYLNAHNTARADVGVEPLTWDDEVAAYAQNYV--SQLAADCNLVT-SHGQYGENLAMGS 91
Query: 97 GMGWTPSVAVQDWVNEKEFYSYRTNSCVAPSFPCWHYTQVVWRKSKQVGCAQLTCVVDKL 156
G T + AV+ WV+EK++Y + +N C A C HYTQVVWR S +VGCA++ C
Sbjct: 92 GDFMTAAKAVEMWVDEKQYYDHGSNHC-AQGQVCGHYTQVVWRNSVRVGCARVQCNNGGY 150
Query: 157 ILTICFYDPPGNINGESPY 175
+++ C YDPPGN G+SPY
Sbjct: 151 VVS-CNYDPPGNFVGQSPY 168
>UniRef100_Q00008 Pathogenesis-related protein PRMS precursor [Zea mays]
Length = 167
Score = 112 bits (281), Expect = 3e-24
Identities = 72/175 (41%), Positives = 97/175 (55%), Gaps = 12/175 (6%)
Query: 5 MQPFFSLALALFLLSAEAATVLDSVVPQLSAEA-KEFLDAHNWARAEVGVEPLEWSEALA 63
M+ LA+ L L AAT +V P S + +++L N ARA VGV P+ WS L
Sbjct: 1 MEASNKLAVLLLWLVMAAAT---AVHPSYSENSPQDYLTPQNSARAAVGVGPVTWSTKLQ 57
Query: 64 KDAGLFARYQRDRLG-CKLANFSHNKYGSNQDLGGMG--WTPSVAVQDWVNEKEFYSYRT 120
+ A +Y R G C+L + S YG N G G W AV+ WV+EK++Y+Y T
Sbjct: 58 QFA---EKYAAQRAGDCRLQH-SGGPYGENIFWGSAGFDWKAVDAVRSWVDEKQWYNYAT 113
Query: 121 NSCVAPSFPCWHYTQVVWRKSKQVGCAQLTCVVDKLILTICFYDPPGNINGESPY 175
NSC A C HYTQVVWR + +GCA++ C ++ + IC Y+P GNI G PY
Sbjct: 114 NSCAAGKV-CGHYTQVVWRATTSIGCARVVCRDNRGVFIICNYEPRGNIAGMKPY 167
>UniRef100_Q6YSG3 Putative pathogenesis-related protein [Oryza sativa]
Length = 172
Score = 112 bits (280), Expect = 4e-24
Identities = 63/162 (38%), Positives = 87/162 (52%), Gaps = 6/162 (3%)
Query: 18 LSAEAATVLDSVVPQLSAEAKEFLDAHNWARAEVGVEPLEWSEALAKDAGLFARYQRDRL 77
LS A L + ++F+ HN ARA V W++ +A A +A ++
Sbjct: 13 LSLAAVLALAAAPSMAQNSPQDFVSPHNAARANVSAAAAAWNDTVAAYAQSYAAQRQG-- 70
Query: 78 GCKLANF-SHNKYGSNQDLGGMG--WTPSVAVQDWVNEKEFYSYRTNSCVAPSFP-CWHY 133
CKL + S +YG N G G WT + AV WV+EK++Y++ +NSC APS C HY
Sbjct: 71 DCKLVHSDSGGRYGENLFWGSAGGNWTAASAVSAWVSEKQWYNHTSNSCSAPSGQSCGHY 130
Query: 134 TQVVWRKSKQVGCAQLTCVVDKLILTICFYDPPGNINGESPY 175
TQVVWR S +GCA++ C + C Y PPGN G+SPY
Sbjct: 131 TQVVWRSSTAIGCARVVCNGSLGVFITCNYSPPGNYIGQSPY 172
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.321 0.135 0.435
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 300,931,327
Number of Sequences: 2790947
Number of extensions: 11511862
Number of successful extensions: 22506
Number of sequences better than 10.0: 487
Number of HSP's better than 10.0 without gapping: 301
Number of HSP's successfully gapped in prelim test: 186
Number of HSP's that attempted gapping in prelim test: 21639
Number of HSP's gapped (non-prelim): 506
length of query: 175
length of database: 848,049,833
effective HSP length: 119
effective length of query: 56
effective length of database: 515,927,140
effective search space: 28891919840
effective search space used: 28891919840
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 70 (31.6 bits)
Lotus: description of TM0323a.10


