

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0314.4
(199 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
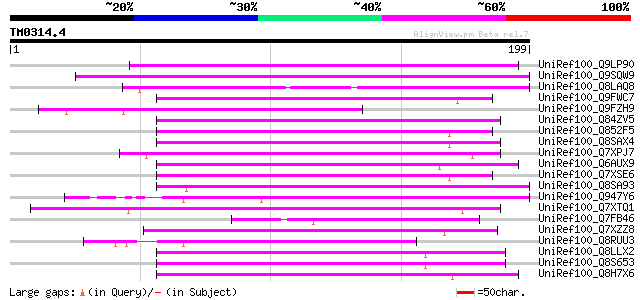
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9LP90 T32E20.30 [Arabidopsis thaliana] 107 1e-22
UniRef100_Q9SQW9 Putative retroelement pol polyprotein [Arabidop... 94 2e-18
UniRef100_Q8LAQ8 Hypothetical protein [Arabidopsis thaliana] 83 5e-15
UniRef100_Q9FWC7 Putative plant disease resistance polyprotein [... 74 2e-12
UniRef100_Q9FZH9 F1O19.7 protein [Arabidopsis thaliana] 74 2e-12
UniRef100_Q84ZV5 Polyprotein [Glycine max] 74 2e-12
UniRef100_Q852F5 Putative polyprotein [Oryza sativa] 70 4e-11
UniRef100_Q8SAX4 Putative polyprotein [Oryza sativa] 67 2e-10
UniRef100_Q7XPJ7 OSJNBa0087O24.13 protein [Oryza sativa] 66 5e-10
UniRef100_Q6AUX9 Putative retrotransposon gag protein [Oryza sat... 65 1e-09
UniRef100_Q7XSE6 OSJNBb0033G08.8 protein [Oryza sativa] 65 1e-09
UniRef100_Q8SA93 Putative polyprotein [Zea mays] 64 3e-09
UniRef100_Q947Y6 Putative retroelement [Oryza sativa] 62 7e-09
UniRef100_Q7XTQ1 OSJNBa0041A02.23 protein [Oryza sativa] 60 4e-08
UniRef100_Q7FB46 Hypothetical protein T9L3_130 [Arabidopsis thal... 59 6e-08
UniRef100_Q7XZZ8 Putative polyprotein [Oryza sativa] 57 4e-07
UniRef100_Q8RUU3 Putative gag-pol polyprotein [Oryza sativa] 55 8e-07
UniRef100_Q8LLX2 Putative retroelement [Oryza sativa] 55 8e-07
UniRef100_Q8S653 Putative retroelement [Oryza sativa] 55 8e-07
UniRef100_Q8H7X6 Putative retroelement [Oryza sativa] 55 1e-06
>UniRef100_Q9LP90 T32E20.30 [Arabidopsis thaliana]
Length = 1397
Score = 107 bits (268), Expect = 1e-22
Identities = 55/149 (36%), Positives = 86/149 (56%)
Query: 47 RVHLRAVTGRRVDIPMFDGSDAYGWINRVERFFQLSRVGEEERLEMVMIAMEGKALGWFQ 106
R +R + V++PMFDGS YGWI RVERFF+ E E+L +V +++ G+AL W+
Sbjct: 119 RPRIRENLLKNVEMPMFDGSGIYGWIARVERFFRSGGYNEAEQLALVSVSVSGEALSWYN 178
Query: 107 WWEEQAPERAWEPFKHALIRRFQPTLVQNPFGPLLSIKQKGSVMAYRESFEEAAAPMRGA 166
W + +W K L+ RF ++ P L IKQ GSV Y + FE+ ++ + G
Sbjct: 179 WAISRGDFVSWLKLKSGLMLRFGNLKLRGPSQSLFCIKQTGSVAEYVQRFEDLSSQVGGL 238
Query: 167 DREILKGVFLNGLQEEIRAEMKLYPAADL 195
D + L+G+FLNGL E++ + ++ +L
Sbjct: 239 DDQKLEGIFLNGLTGEMQELVHMHKPQNL 267
>UniRef100_Q9SQW9 Putative retroelement pol polyprotein [Arabidopsis thaliana]
Length = 1661
Score = 94.4 bits (233), Expect = 2e-18
Identities = 50/174 (28%), Positives = 90/174 (50%)
Query: 26 SGSRTGSRFSDHSRERFDHPHRVHLRAVTGRRVDIPMFDGSDAYGWINRVERFFQLSRVG 85
+G +T F + PH+ A R VD P ++G +A W+ R+E+ F +R
Sbjct: 199 TGFQTTPTFQTQTFPPQSAPHQPRFEAAPRRTVDYPAYEGGNADDWLFRLEQCFLSNRTL 258
Query: 86 EEERLEMVMIAMEGKALGWFQWWEEQAPERAWEPFKHALIRRFQPTLVQNPFGPLLSIKQ 145
EEE+LE + + G ++ W++ +++ W F+ + RF+P+ + LL+++Q
Sbjct: 259 EEEKLEKAVSCLTGASVTWWRCSKDREQIYTWREFQEKFMLRFRPSRGSSAVDHLLNVRQ 318
Query: 146 KGSVMAYRESFEEAAAPMRGADREILKGVFLNGLQEEIRAEMKLYPAADLAGLM 199
G+V YRE FEE + +IL+ FLNGL+ +R ++ +LA ++
Sbjct: 319 TGTVEEYRERFEELTVDLPHVTSDILESAFLNGLRRSLRDQVVRCRPVNLADIV 372
>UniRef100_Q8LAQ8 Hypothetical protein [Arabidopsis thaliana]
Length = 272
Score = 82.8 bits (203), Expect = 5e-15
Identities = 45/157 (28%), Positives = 88/157 (55%), Gaps = 4/157 (2%)
Query: 44 HPHRV-HLRAVTGRRVDIPMFDGSDAYGWINRVERFFQLSRVGEEERLEMVMIAMEGKAL 102
HPHR+ + ++++P+F+G + W+ R ER+F+ E++++V +++EG AL
Sbjct: 99 HPHRMSRYIQLMKPKIEMPVFEGPNVNSWLTRAERYFEFGSFTNAEKIQLVYMSVEGPAL 158
Query: 103 GWFQWWEEQAPERAWEPFKHALIRRFQPTLVQNPFGPLLSIKQKGSVMAYRESFEEAAAP 162
WF E + P W FK +++RF ++ LL++KQ SV++Y FE+ +
Sbjct: 159 CWFN-LENRNPFVDWNDFKARVLQRFGDP--RSAMERLLTLKQVDSVISYLGEFEDLSTQ 215
Query: 163 MRGADREILKGVFLNGLQEEIRAEMKLYPAADLAGLM 199
+L+ VF+ GL+EEI+ ++++ L+ ++
Sbjct: 216 CLDDLDTVLEVVFVKGLKEEIQEMLRVFQPKGLSDII 252
>UniRef100_Q9FWC7 Putative plant disease resistance polyprotein [Oryza sativa]
Length = 894
Score = 74.3 bits (181), Expect = 2e-12
Identities = 38/133 (28%), Positives = 69/133 (51%), Gaps = 4/133 (3%)
Query: 57 RVDIPMFDGSDAYGWINRVERFFQLSRVGEEERLEMVMIAMEGKALGWFQWWEEQAPERA 116
++D P FDGSD W+N E +F + ++ + ++ V++ M G A W+ ++ +
Sbjct: 121 KMDFPRFDGSDVRIWLNMCETYFDMYQITQNFKVSAVVLHMSGNAAQWYHSYKLVNEVNS 180
Query: 117 WEPFKHALIRRFQPTLVQNPFGPLLSIKQKGSVMAYRESFEEAAAPMRGADREI----LK 172
W+ F+ A+ F+ + + L ++ Q G+V Y++ F+ +R D + L
Sbjct: 181 WDQFRMAVATEFEGVVEREKMSALDTLTQTGTVTEYKQQFDYLVYQIRVFDPSVGGKMLV 240
Query: 173 GVFLNGLQEEIRA 185
F+NGL EEIRA
Sbjct: 241 TRFMNGLTEEIRA 253
>UniRef100_Q9FZH9 F1O19.7 protein [Arabidopsis thaliana]
Length = 659
Score = 74.3 bits (181), Expect = 2e-12
Identities = 42/127 (33%), Positives = 66/127 (51%), Gaps = 3/127 (2%)
Query: 12 RQASRERDR-EASSHSGSRTGSRFSDHSRERF--DHPHRVHLRAVTGRRVDIPMFDGSDA 68
R RE R E HS FS +R + P + R+ RR+++P+FDGS
Sbjct: 61 RMEIREGKRVEQGEHSCQSPRHSFSSSMPQRGAGESPILLDNRSSLIRRIEMPVFDGSGV 120
Query: 69 YGWINRVERFFQLSRVGEEERLEMVMIAMEGKALGWFQWWEEQAPERAWEPFKHALIRRF 128
Y W ++VERFF++ R + ++L++V +++EG AL WF R W F+ L+ RF
Sbjct: 121 YEWFSKVERFFRVGRYQDSDKLDLVALSLEGVALKWFLREMSTLEFRDWNSFEQRLLARF 180
Query: 129 QPTLVQN 135
P + +
Sbjct: 181 DPVKIHS 187
>UniRef100_Q84ZV5 Polyprotein [Glycine max]
Length = 1552
Score = 73.9 bits (180), Expect = 2e-12
Identities = 37/132 (28%), Positives = 63/132 (47%)
Query: 57 RVDIPMFDGSDAYGWINRVERFFQLSRVGEEERLEMVMIAMEGKALGWFQWWEEQAPERA 116
++D P FDG + WI + E+FF + +RL + + ++ + W+Q ++ P +
Sbjct: 99 KLDFPRFDGKNVMDWIFKAEQFFDYYATPDADRLIIASVHLDQDVVPWYQMLQKTEPFSS 158
Query: 117 WEPFKHALIRRFQPTLVQNPFGPLLSIKQKGSVMAYRESFEEAAAPMRGADREILKGVFL 176
W+ F AL F P+ P L + Q +V Y F + G E + F+
Sbjct: 159 WQAFTRALELDFGPSAYDCPRATLFKLNQSATVNEYYMQFTALVNRVDGLSAEAILDCFV 218
Query: 177 NGLQEEIRAEMK 188
+GLQEEI ++K
Sbjct: 219 SGLQEEISRDVK 230
>UniRef100_Q852F5 Putative polyprotein [Oryza sativa]
Length = 1155
Score = 69.7 bits (169), Expect = 4e-11
Identities = 35/133 (26%), Positives = 69/133 (51%), Gaps = 4/133 (3%)
Query: 57 RVDIPMFDGSDAYGWINRVERFFQLSRVGEEERLEMVMIAMEGKALGWFQWWEEQAPERA 116
++D P FDGSD W++ E +F + ++ + ++ ++ M G A W+ ++ +
Sbjct: 121 KMDFPRFDGSDVRIWLDMCETYFDMYQITQNFKVSAAVLHMSGNAAQWYHSYKLVNEVNS 180
Query: 117 WEPFKHALIRRFQPTLVQNPFGPLLSIKQKGSVMAYRESFEEAAAPMRGAD----REILK 172
W+ F+ A+ F+ + + L ++ Q G++ Y++ F+ +R D ++L
Sbjct: 181 WDQFRMAVATEFECVVEREKMSALDTLTQTGTMTEYKQQFDSLVYQIRVFDPSMGGKMLV 240
Query: 173 GVFLNGLQEEIRA 185
F+NGL EEIRA
Sbjct: 241 THFMNGLTEEIRA 253
>UniRef100_Q8SAX4 Putative polyprotein [Oryza sativa]
Length = 1087
Score = 67.4 bits (163), Expect = 2e-10
Identities = 39/136 (28%), Positives = 63/136 (45%), Gaps = 4/136 (2%)
Query: 57 RVDIPMFDGSDAYGWINRVERFFQLSRVGEEERLEMVMIAMEGKALGWFQWWEEQAPERA 116
++D P F GSD W++ E FF ++ + ++ ++ M G A W+Q W+ +
Sbjct: 144 KMDFPKFSGSDVRVWLDNCETFFLFYQIADGYKVSAAVLNMSGDAANWYQEWKFEQGWHD 203
Query: 117 WEPFKHALIRRFQPTLVQNPFGPLLSIKQKGSVMAYRESFEEAAAPMRGAD----REILK 172
W K AL+ F+ L LL + Q G V YR F + +R D L
Sbjct: 204 WPMLKAALLGEFEVHLKYVKMDELLLLTQTGFVSDYRSKFNQLVYQIRLYDPLLSPNFLI 263
Query: 173 GVFLNGLQEEIRAEMK 188
FL GL++E+R ++
Sbjct: 264 RQFLLGLKDELRVPVQ 279
>UniRef100_Q7XPJ7 OSJNBa0087O24.13 protein [Oryza sativa]
Length = 1311
Score = 66.2 bits (160), Expect = 5e-10
Identities = 41/152 (26%), Positives = 73/152 (47%), Gaps = 6/152 (3%)
Query: 43 DHPHRVHLR--AVTGRRVDIPMFDGSDAYGWINRVERFFQLSRVGEEERLEMVMIAMEGK 100
++ HR H A R VD P F+G WI + E++F L + EE+++ + + + G+
Sbjct: 273 NYAHRPHSADPAKRSRNVDFPTFEGDYPESWIRKAEKYFSLYQTPEEDKVLLAEVHISGR 332
Query: 101 ALGWFQWWEEQAPERAWEPFKHALIRRFQPTLVQNPFGPLLSIKQKGSVMAYRESFEEAA 160
A W + +W FK + +RF ++KQ GSV +Y + FEE
Sbjct: 333 ADQWIESSAVPTASLSWPEFKTMVCQRFAAKSKIEITETFRNLKQYGSVDSYIDKFEETM 392
Query: 161 APMRGADREILKGVFL----NGLQEEIRAEMK 188
A ++ ++ + + FL +GL+ I+ +K
Sbjct: 393 ALVKRSNPTLTEDYFLDYFISGLKGHIKRPLK 424
>UniRef100_Q6AUX9 Putative retrotransposon gag protein [Oryza sativa]
Length = 748
Score = 65.1 bits (157), Expect = 1e-09
Identities = 42/143 (29%), Positives = 62/143 (42%), Gaps = 4/143 (2%)
Query: 57 RVDIPMFDGSDAYGWINRVERFFQLSRVGEEERLEMVMIAMEGKALGWFQWWEEQAPERA 116
++D P+FDGSD W + E +F + + + I G+A W
Sbjct: 208 KLDFPIFDGSDPQDWRMKCEHYFDVCDTSPNPWVRIATIYFRGRAASWLHSSRAHVCYLG 267
Query: 117 WEPFKHALIRRFQPTLVQNPFGPLLSIKQKGSVMAYRESFEEAAAPM----RGADREILK 172
W F A+ +F L G L +IKQ G+V Y E F+E A + A+
Sbjct: 268 WAEFCGAVACKFDINLHDWVLGQLDTIKQLGTVQEYYEKFDELANQLLVYDLAANDRYFT 327
Query: 173 GVFLNGLQEEIRAEMKLYPAADL 195
F NGL++EIR + L+ DL
Sbjct: 328 HKFTNGLRKEIRGAVILHRPRDL 350
>UniRef100_Q7XSE6 OSJNBb0033G08.8 protein [Oryza sativa]
Length = 368
Score = 64.7 bits (156), Expect = 1e-09
Identities = 37/133 (27%), Positives = 62/133 (45%), Gaps = 4/133 (3%)
Query: 57 RVDIPMFDGSDAYGWINRVERFFQLSRVGEEERLEMVMIAMEGKALGWFQWWEEQAPERA 116
++D P FDG+ WI+ E +F ++ E + + + G A W+Q W+ +
Sbjct: 9 KMDFPKFDGNGVKVWIDNCETYFAFYQITEAFLVSATSMNLMGDAANWYQTWKLEVGWHD 68
Query: 117 WEPFKHALIRRFQPTLVQNPFGPLLSIKQKGSVMAYRESFEEAAAPMRGAD----REILK 172
WE K A+ R F L LL +KQ +V+ Y F E +R D +L
Sbjct: 69 WEMLKTAISREFDVHLWSVKMDELLLLKQTSTVIEYCSKFSELVYQIRLYDPLLCGSVLV 128
Query: 173 GVFLNGLQEEIRA 185
F+ GL++++R+
Sbjct: 129 SHFMLGLKDDLRS 141
>UniRef100_Q8SA93 Putative polyprotein [Zea mays]
Length = 2749
Score = 63.5 bits (153), Expect = 3e-09
Identities = 38/144 (26%), Positives = 62/144 (42%), Gaps = 1/144 (0%)
Query: 57 RVDIPMFDGS-DAYGWINRVERFFQLSRVGEEERLEMVMIAMEGKALGWFQWWEEQAPER 115
++D P +DGS D W+N+ E+FF+ R +R + ++G A W+ E+
Sbjct: 472 KIDFPTYDGSVDPLNWLNQCEQFFRGQRTLVTDRTWLASYHLKGAAQTWYYALEQDEGMP 531
Query: 116 AWEPFKHALIRRFQPTLVQNPFGPLLSIKQKGSVMAYRESFEEAAAPMRGADREILKGVF 175
W FK RF P + L + +V Y + F R D + +F
Sbjct: 532 TWGRFKEVCTLRFGPPVRGTRLSELARLPFTSTVQDYADRFNAMLGHTRKLDAQQKAELF 591
Query: 176 LNGLQEEIRAEMKLYPAADLAGLM 199
+ GL + IRA++ + DL M
Sbjct: 592 VGGLPDHIRADVAIRDPQDLQSAM 615
>UniRef100_Q947Y6 Putative retroelement [Oryza sativa]
Length = 1461
Score = 62.4 bits (150), Expect = 7e-09
Identities = 55/180 (30%), Positives = 82/180 (45%), Gaps = 14/180 (7%)
Query: 22 ASSHSGSRTGSRFSDHSRERFDHPHRVHLRAVTGRRVDIPMFDG-SDAYGWINRVERFFQ 80
+SS SG+R G F D +R P + H R D P +DG SD +INR E FF
Sbjct: 45 SSSTSGARPG--FGDQPPDR---PPK-HWRP------DFPHYDGKSDPLIFINRCESFFL 92
Query: 81 LSRVGEEERLEMVMI-AMEGKALGWFQWWEEQAPERAWEPFKHALIRRFQPTLVQNPFGP 139
R+ +EE++ M +EG L + Q E++ W FK L R+ P L P
Sbjct: 93 QQRIMQEEKVWMASHNLLEGAQLWYMQVQEDERGTPTWTRFKELLNLRYGPPLRSAPLFE 152
Query: 140 LLSIKQKGSVMAYRESFEEAAAPMRGADREILKGVFLNGLQEEIRAEMKLYPAADLAGLM 199
L S ++ G+V Y++ F+ D E +F GL + ++++ LA M
Sbjct: 153 LSSCRRTGTVEDYQDRFQALLPRAGRLDEEQRVQLFTGGLLPPLSLQVQMQNPQSLAAAM 212
>UniRef100_Q7XTQ1 OSJNBa0041A02.23 protein [Oryza sativa]
Length = 1373
Score = 59.7 bits (143), Expect = 4e-08
Identities = 41/185 (22%), Positives = 84/185 (45%), Gaps = 5/185 (2%)
Query: 9 PRFRQASRERDREASSHSGSRTGSRFSDHSRERFDH-PHRVHLRAVTGRRVDIPMFDGSD 67
P+ + +R H + G + + R F P+ + +R+++ +F+G D
Sbjct: 240 PQMQPTARPHLNATGIHMANGGGPVWPEGPRHLFGGMPNPYADMTLRSQRLELTIFNGED 299
Query: 68 AYGWINRVERFFQLSRVGEEERLEMVMIAMEGKALGWFQWWEEQAPERAWEPFKHALIRR 127
A GW+ + E+FF+++ E+ + + + G+A WF+ W F + R
Sbjct: 300 AIGWLQQCEKFFEMTGTPVEQWVNLATGHLVGRAGKWFRNLGIPWHYINWPQFYLMISDR 359
Query: 128 FQPTLVQNPFGPLLSIKQKGSVMAYRESFEEAAAPMRGADREILK----GVFLNGLQEEI 183
F L +++Q GSVM Y + FE++ A ++ +++ F+ GL+ +I
Sbjct: 360 FTEANAHEAVEQLQNVRQTGSVMHYIDKFEDSVALVKRDHPYLIEYYILSCFIGGLRADI 419
Query: 184 RAEMK 188
+ + K
Sbjct: 420 KHDHK 424
>UniRef100_Q7FB46 Hypothetical protein T9L3_130 [Arabidopsis thaliana]
Length = 757
Score = 59.3 bits (142), Expect = 6e-08
Identities = 33/97 (34%), Positives = 50/97 (51%), Gaps = 4/97 (4%)
Query: 86 EEERLEMVMIAMEGKALGWFQWWEEQAPER--AWEPFKHALIRRFQPTLVQNPFGPLLSI 143
E ++LE + A+ G A+ W W Q PE+ W+ F+ RF+P L +I
Sbjct: 620 ETQKLEQAITALTGTAINW--WQIAQYPEKISTWKDFRDKFKVRFKPARGAATIDQLFTI 677
Query: 144 KQKGSVMAYRESFEEAAAPMRGADREILKGVFLNGLQ 180
Q+GSV YR FEE A + ++L+ FLNG++
Sbjct: 678 FQRGSVEEYRNRFEEIAVELPHVTDDVLESAFLNGVE 714
>UniRef100_Q7XZZ8 Putative polyprotein [Oryza sativa]
Length = 1246
Score = 56.6 bits (135), Expect = 4e-07
Identities = 32/140 (22%), Positives = 69/140 (48%), Gaps = 4/140 (2%)
Query: 52 AVTGRRVDIPMFDGSDAYGWINRVERFFQLSRVGEEERLEMVMIAMEGKALGWFQWWEEQ 111
++ +R+++ +F+G DA GW+ + E+FF+++ ++ + + + G+A WF+
Sbjct: 294 SLRSQRLELTLFNGEDAVGWLQQCEKFFEMTGTPVDQWVNLASGHLVGRAGKWFRNLAIP 353
Query: 112 APERAWEPFKHALIRRFQPTLVQNPFGPLLSIKQKGSVMAYRESFEEAAAPMRG----AD 167
W+ F ++ RF L + KQ GSVM Y + FE+ + ++
Sbjct: 354 WYCLNWQQFYLMVLDRFTEANAHEAVELLQNAKQTGSVMQYIDKFEDCVSLVKRDHPYLT 413
Query: 168 REILKGVFLNGLQEEIRAEM 187
+ F+ GL+ +I+ ++
Sbjct: 414 ESYILSCFIGGLRADIKHDV 433
>UniRef100_Q8RUU3 Putative gag-pol polyprotein [Oryza sativa]
Length = 1338
Score = 55.5 bits (132), Expect = 8e-07
Identities = 40/136 (29%), Positives = 58/136 (42%), Gaps = 15/136 (11%)
Query: 29 RTGSRFSDHSR---ERFD----HPHRVHLRAVTGRRVDIPMFDG-SDAYGWINRVERFFQ 80
R G F H R RFD P R H ++D P FDG D ++NR E+FF+
Sbjct: 82 RGGDGFHGHGRPGGHRFDGDGDRPPRYH-------KLDFPKFDGRGDPLPFLNRCEQFFR 134
Query: 81 LSRVGEEERLEMVMIAMEGKALGWFQWWEEQAPERAWEPFKHALIRRFQPTLVQNPFGPL 140
R E+ ++ + + A W+ E +W F L R+ P L P G L
Sbjct: 135 GQRTPEDNKVWLASYHLLDGAQQWYTRLERDHEPPSWHRFSELLNMRYGPPLHSTPLGEL 194
Query: 141 LSIKQKGSVMAYRESF 156
+ ++ +V Y E F
Sbjct: 195 AACRRTTTVDDYAERF 210
>UniRef100_Q8LLX2 Putative retroelement [Oryza sativa]
Length = 813
Score = 55.5 bits (132), Expect = 8e-07
Identities = 37/138 (26%), Positives = 57/138 (40%), Gaps = 4/138 (2%)
Query: 57 RVDIPMFDGSDAYGWINRVERFFQLSRVGEEERLEMVMIAMEGKALGWFQWWEEQAPERA 116
+ D P F G + W E++F + V E + + G A W Q +EE +
Sbjct: 111 KTDFPRFSGENPKLWKKNAEKYFGMYNVAYETWAQFATLHFTGNAALWLQTYEELHSVES 170
Query: 117 WEPFKHALIRRFQPTLVQNPFGPLLSIKQKGSVMAYRESFEE----AAAPMRGADREILK 172
W A+ +F Q+ L ++KQ GSV FEE RG D
Sbjct: 171 WAELCVAVNSKFGRDPYQHHLEELENLKQTGSVEECHNKFEELMHLVLVYNRGYDETFFV 230
Query: 173 GVFLNGLQEEIRAEMKLY 190
F+ GL+ +I+ +KL+
Sbjct: 231 TRFVGGLKADIKLAIKLH 248
>UniRef100_Q8S653 Putative retroelement [Oryza sativa]
Length = 1043
Score = 55.5 bits (132), Expect = 8e-07
Identities = 37/138 (26%), Positives = 57/138 (40%), Gaps = 4/138 (2%)
Query: 57 RVDIPMFDGSDAYGWINRVERFFQLSRVGEEERLEMVMIAMEGKALGWFQWWEEQAPERA 116
+ D P F G + W E++F + V E + + G A W Q +EE +
Sbjct: 111 KTDFPRFSGENPKLWKKNAEKYFGMYNVAYETWAQFATLHFTGNAALWLQTYEELHSVES 170
Query: 117 WEPFKHALIRRFQPTLVQNPFGPLLSIKQKGSVMAYRESFEE----AAAPMRGADREILK 172
W A+ +F Q+ L ++KQ GSV FEE RG D
Sbjct: 171 WAELCVAVNSKFGRDPYQHHLEELENLKQTGSVEECHNKFEELMHLVLVYNRGYDETFFV 230
Query: 173 GVFLNGLQEEIRAEMKLY 190
F+ GL+ +I+ +KL+
Sbjct: 231 TRFVGGLKADIKLAIKLH 248
>UniRef100_Q8H7X6 Putative retroelement [Oryza sativa]
Length = 1021
Score = 55.1 bits (131), Expect = 1e-06
Identities = 41/143 (28%), Positives = 60/143 (41%), Gaps = 4/143 (2%)
Query: 57 RVDIPMFDGSDAYGWINRVERFFQLSRVGEEERLEMVMIAMEGKALGWFQWWEEQAPERA 116
++D P FDG+D W R E +F ++ + + I G+A W + +
Sbjct: 232 KLDFPKFDGTDPEDWRMRCEHYFDVNNTFPGLWVRIATIYFSGRAASWLRSTKAHVRFPI 291
Query: 117 WEPFKHALIRRFQPTLVQNPFGPLLSIKQKGSVMAYRESFEEAAAPMRGADR----EILK 172
WE F A+ +F + + SIKQ GSV Y E F+E + D L
Sbjct: 292 WENFCVAMSNKFDRDQHEVLIRQMDSIKQIGSVWEYYEKFDELMNQLLVYDPLLNIRYLT 351
Query: 173 GVFLNGLQEEIRAEMKLYPAADL 195
F GL+ EIR + L DL
Sbjct: 352 HRFTAGLRREIRNAVLLQRPKDL 374
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.321 0.136 0.413
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 316,085,246
Number of Sequences: 2790947
Number of extensions: 12566633
Number of successful extensions: 32420
Number of sequences better than 10.0: 317
Number of HSP's better than 10.0 without gapping: 91
Number of HSP's successfully gapped in prelim test: 226
Number of HSP's that attempted gapping in prelim test: 32173
Number of HSP's gapped (non-prelim): 421
length of query: 199
length of database: 848,049,833
effective HSP length: 121
effective length of query: 78
effective length of database: 510,345,246
effective search space: 39806929188
effective search space used: 39806929188
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 71 (32.0 bits)
Lotus: description of TM0314.4


