

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0310b.12
(1485 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
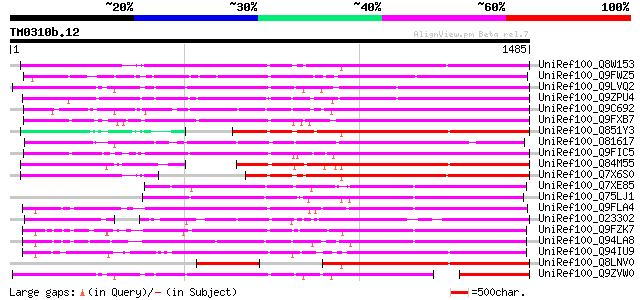
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q8W153 Polyprotein [Oryza sativa] 1085 0.0
UniRef100_Q9FWZ5 Putative retroelement polyprotein [Arabidopsis ... 885 0.0
UniRef100_Q9LVQ2 Retroelement pol polyprotein-like [Arabidopsis ... 880 0.0
UniRef100_Q9ZPU4 Putative retroelement pol polyprotein [Arabidop... 874 0.0
UniRef100_Q9C692 Polyprotein, putative [Arabidopsis thaliana] 843 0.0
UniRef100_Q9FXB7 Putative retroelement polyprotein [Arabidopsis ... 842 0.0
UniRef100_Q851Y3 Putative polyprotein [Oryza sativa] 832 0.0
UniRef100_O81617 F8M12.17 protein [Arabidopsis thaliana] 829 0.0
UniRef100_Q9FIC5 Retroelement pol polyprotein-like [Arabidopsis ... 817 0.0
UniRef100_Q84M55 Putative polyprotein [Oryza sativa] 798 0.0
UniRef100_Q7X6S0 OSJNBb0011N17.2 protein [Oryza sativa] 795 0.0
UniRef100_Q7XE85 Putative pol polyprotein [Oryza sativa] 717 0.0
UniRef100_Q75LJ1 Putative copia-like retrotransposon protein [Or... 695 0.0
UniRef100_Q9FLA4 Polyprotein [Arabidopsis thaliana] 692 0.0
UniRef100_O23302 Retrovirus-related like polyprotein [Arabidopsi... 683 0.0
UniRef100_Q9FZK7 F17L21.7 [Arabidopsis thaliana] 676 0.0
UniRef100_Q94LA8 Polyprotein, putative [Arabidopsis thaliana] 660 0.0
UniRef100_Q94IU9 Copia-like polyprotein [Arabidopsis thaliana] 660 0.0
UniRef100_Q8LNV0 Putative copia-like polyprotein [Oryza sativa] 651 0.0
UniRef100_Q9ZVW0 Putative retroelement pol polyprotein [Arabidop... 625 e-177
>UniRef100_Q8W153 Polyprotein [Oryza sativa]
Length = 1472
Score = 1085 bits (2806), Expect = 0.0
Identities = 623/1503 (41%), Positives = 878/1503 (57%), Gaps = 120/1503 (7%)
Query: 30 LPVLQGSFRLDG-RNYLQWSELVRHTLISRQKISYI--EETAPAETDP-QYDTWREENSL 85
+ ++Q +L+G +NYL WS L ++ Y+ E P T ++ TW NSL
Sbjct: 42 IELMQNEIKLEGVKNYLSWSRRALLILKTKGLEGYVTGEVKEPENTSSVEWKTWSTTNSL 101
Query: 86 IMTWFWYSMIPEISRNYMFFPTAKEIWNNLAQAYSKKQDLSACYELENKVFNSKQGTLSV 145
++ W S+IP I+ +A E+W L + YS + ++ E + K+ +QG SV
Sbjct: 102 VVAWLLTSLIPAIATTVETISSASEMWKTLTKLYSGEGNVMLMVEAQEKISALRQGERSV 161
Query: 146 TDYYGTLNGLWIELDQYQNLKMKCDSDSTTLNKFIEKARIFKFLSGLNSEFDPIRVQILG 205
+Y L LW +LD Y L ++ + K++E+ R+ +FL GLN EF+ R +
Sbjct: 162 AEYVAELKSLWSDLDHYDPLGLEHSDCIAKMKKWVERRRVIEFLKGLNPEFEGRRDAMFH 221
Query: 206 KEQFPSLSEVFSIVRGEETRRTVMVEDKSVDGS---ALASGKGPIK--GSTSFGRPNRDD 260
+ P+L E + + EE ++ V+ S A+ GK + G RD
Sbjct: 222 QTTLPTLDEAIAAMAQEELKKKVLPSAAPCSPSPTYAIVQGKETRECFNCGEMGHLMRDC 281
Query: 261 RPNRDDRFN---GDDRFNRDDRCTYCKRSGHTKEYCFKLYGRENVLARMSKSKGA-PQRR 316
R + G DR Y RS + Y ++ + N + S G P
Sbjct: 282 HAPRKPTYGRGRGVDRGGTRGGRGYAGRSNRGRGYGYRGDYKANAVTLEEGSSGTTPDNV 341
Query: 317 ANHTASDMENDGEVPPVPPAEDVPSLSKAELERLRVFMDSLSKPSGSCSLTMTGKSSTFL 376
AN S SGS + F+
Sbjct: 342 ANFAHST-------------------------------------SGSFN-------QAFM 357
Query: 377 SLNALSTENDWIIDFGATDHMTPHPSHLSSYSTYPGKHYITV--ANGSQNQITGCGNIHL 434
S+N ++ + WI+D GA+ H+T +SY Y H T+ A+G+ Q+ G G +
Sbjct: 358 SMN--TSHSSWILDSGASRHVTGMSGEFTSYKPYSFAHKETIQTADGTSCQVKGEGIVQC 415
Query: 435 SPSLPLKRVLHVPKLSNNLLSIHKLTRDLNCAVTFFHSHGVFQDLATGQIIGIAKEQDGL 494
+PS+ L VL+V NL+SI L +++C V+ + + Q+ TG+ +GI +DGL
Sbjct: 416 TPSITLSSVLYVHSFPVNLISISSLVDNMDCRVSLDRENCLIQERRTGKKLGIGIRRDGL 475
Query: 495 YCLQHE---ESKCHQTSSESWNTSQIWLRHKRLGHPPFSILRTMFPHLFSKVSVESFHCD 551
+ L E C +S S +++ L H RLGH F I+ MFP FSKV CD
Sbjct: 476 WYLDRRGTNEDVCALMASTSKEVTEVLLLHCRLGHISFEIMSKMFPVEFSKVDKHMLICD 535
Query: 552 VCQFSKHHRSTFLPSNNKCAQPFDLVHSDVWGPSSISNVSGAKWFVTFIDDCTRITWVFL 611
C++ KH R++++ + PF L+HSDVW S + ++SG K+FVTFID +R+TW++L
Sbjct: 536 ACEYGKHTRTSYVSRGLRSILPFMLIHSDVW-TSPVVSMSGMKYFVTFIDCYSRMTWLYL 594
Query: 612 MKDKSEVFQLFANFFHMIRTQFGKPIKRLRSDNGKE*VNKNFSEFLTKNGVVHELTCVDA 671
M+ K EV + F NF+ I+ F ++ +R+DNG E +N F FL+ G++H+ +C D
Sbjct: 595 MRHKDEVLKCFQNFYAYIKNHFNARVQFIRTDNGGEYMNSEFGHFLSLEGILHQTSCPDT 654
Query: 672 PQQNGVAERKNRHLLEITRALLFQMNVPKFYWGEAVLTAAYLINRLPSRVLSNVSPVLVM 731
P QNGVAERKNRHLLEI R+L++ MNVPKF W EAV+TAAYLINR PSR+L +P ++
Sbjct: 655 PPQNGVAERKNRHLLEIARSLMYTMNVPKFLWSEAVMTAAYLINRTPSRILGMKTPYEMI 714
Query: 732 TSFFPSVPIKSGLQSRVFGCSAFVHVHGHHRGKLDPRAIKCIFIGYASDKKGYKCYHPPS 791
V + RVFGC+ FV H GKLDPRA+KCIFIGY+S +KGYKC+ P
Sbjct: 715 FGKNEFV-----VPPRVFGCTCFVRDHRPSIGKLDPRAVKCIFIGYSSSQKGYKCWSPSE 769
Query: 792 RRVYISMDVTFQESESFYPNPQLQGGDTQEAESPELSVIPLLQEPLMPSSIPITEDSDDD 851
RR ++SMDVTF+ES FY E ++S + + + L D D
Sbjct: 770 RRTFVSMDVTFRESVPFY------------GEKTDISSLFVDLDDLTRG------DHDQQ 811
Query: 852 DNGPEQVPNQNDDNGLELVPDQNDDNGPELVPDQ-----NNVDRFRIKYQRKVKPVLIQQ 906
G +N+ + ++V + + V +Q + + ++ Y R+++ QQ
Sbjct: 812 KEGEILGLKENEQSKGKIVVGEIPCAIGDPVQEQEWRKPHEEENLQV-YTRRMRLPTTQQ 870
Query: 907 ---QDQPSDPEVSVPNPEPSNSYEHNL----DDLPIALRKGKRSCAKYP----------- 948
DQ SD V S + + +LPIA+RKG RS A P
Sbjct: 871 VEVDDQVSDDLTHVQVSSESGGEQIEIREEESNLPIAIRKGMRSNAGKPPQRYGFEIGDE 930
Query: 949 ------ISQYVSTQNLSVQHQSFISAIDSIRVPTSVQEVLKDKKWIQAMNEEMHALDKNG 1002
I+ YVS +LS +++F+++++S +P +E +D +W QAM +E+ AL+KN
Sbjct: 931 SGDENDIANYVSYTSLSSTYRAFVASLNSAIIPKDWKEAKQDPRWHQAMLDELEALEKNK 990
Query: 1003 TRKIVEKPNDKRTVGCKWIYTVKYRSDGTLDRYKARLVAKGYTQTYGIDYDETFAPVAKM 1062
T +V PN K+ V CKW+Y VK DG ++RYKARLVAKGY+QTYGIDYDETFAPVAKM
Sbjct: 991 TWDLVSYPNGKKVVNCKWVYAVKQNPDGKVERYKARLVAKGYSQTYGIDYDETFAPVAKM 1050
Query: 1063 NTVRIILSLAAHFDWELQQFDVKNAFLHGDLEEEVYMEIPPGYDPTSGRNKVCKLKKALY 1122
+TVR I+S A +FDW L Q DVKNAFLHGDL+EEVYMEIPPG+ + KV +LKK+LY
Sbjct: 1051 STVRTIISCAVNFDWPLHQLDVKNAFLHGDLQEEVYMEIPPGFATLQTKGKVLRLKKSLY 1110
Query: 1123 GLKQSPRAWFGRFTNAMVSLGYKQSQGDHTLFIKHTRNGKLTLLLVYVNDMIIAGNDELE 1182
GLKQSPRAWF RF AM ++GYKQ GDHT+F H+ + +T+L VYV+DMII GND E
Sbjct: 1111 GLKQSPRAWFDRFRRAMCAMGYKQCNGDHTVFYHHSGD-HITILAVYVDDMIITGNDCSE 1169
Query: 1183 KQNLRKELAVQFEMKDLGKLKYFLGMEVAYSKQGIFISQRKYVLDLLKETGKLGCKPMGV 1242
L++ L+ +FE+KDLG+LKYFLG+E+A S +GI +SQRKY LDLL +TG LGC+P
Sbjct: 1170 ITRLKQNLSKEFEVKDLGQLKYFLGIEIARSPRGIVLSQRKYALDLLSDTGMLGCRPAST 1229
Query: 1243 PIEQNHKIGSSEEDLRVHKAQYQRLVGKLIYLSHTRPDIAYPVSVVSQFMHDPQERHLQA 1302
P++QNHK+ +E V+K +YQRLVG+LIYL HTRPDI Y VS+VS++MHDP+ H+ A
Sbjct: 1230 PVDQNHKL-CAESGNPVNKERYQRLVGRLIYLCHTRPDITYAVSMVSRYMHDPRSGHMDA 1288
Query: 1303 VDRILQYLKASPGRGLLFKKSEQLSMEVYTDADYAGSIVDRRSTTGYCMFLGGNLVTWRS 1362
V RIL+YLK SPG+GL FKK+ L +E Y DA +A DRRST+GYC+F+GGNLV+WRS
Sbjct: 1289 VYRILRYLKGSPGKGLWFKKNGHLEVEGYCDAHWASCPDDRRSTSGYCVFVGGNLVSWRS 1348
Query: 1363 KKQNVVARSSAEAEFRAMAQGVCELLRMKIILDDLKIKYEAPMRLFCDNKSAISIAHNPV 1422
KKQ VV+RS+AEAE+RAM+ + ELL ++ +L +L + + PM+L+CDNKSAISIA+NPV
Sbjct: 1349 KKQPVVSRSTAEAEYRAMSVSLSELLWLRNLLSELMLPVDTPMKLWCDNKSAISIANNPV 1408
Query: 1423 QHDRTKHIEIDRHFIKEKLNSSLISTQYVPSGLQLADVLTKGLPLERFRELTCKLGMIDI 1482
QHDRTKH+E+DR FIKEKL+ ++ ++V SG Q+AD TKGL ++ K+GMIDI
Sbjct: 1409 QHDRTKHVELDRFFIKEKLDEGVLELEFVMSGGQVADCFTKGLGVKECNSSCDKMGMIDI 1468
Query: 1483 HSP 1485
+ P
Sbjct: 1469 YHP 1471
>UniRef100_Q9FWZ5 Putative retroelement polyprotein [Arabidopsis thaliana]
Length = 1404
Score = 885 bits (2287), Expect = 0.0
Identities = 527/1473 (35%), Positives = 795/1473 (53%), Gaps = 113/1473 (7%)
Query: 39 LDGRNYLQWSELVRHTLISRQKISYI----------EETAPAETDPQYDTWREENSLIMT 88
L G NYL WS + L R S++ EE P+ + W +E+ ++
Sbjct: 13 LQGGNYLTWSRTTKTVLCGRGLWSHVISSQAPKEDKEEEETETISPEEEKWFQEDQAVLA 72
Query: 89 WFWYSMIPEISRNYMFFPTAKEIWNNLAQAYSKKQDLSACYELENKVFNSKQGTLSVTDY 148
S+ I Y + TAKE+W+ L Y + +L+ +E++ + Q L T +
Sbjct: 73 LLQNSLETSILEGYSYCETAKELWDTLKNVYGNESNLTRVFEVKKAINELSQEDLEFTKH 132
Query: 149 YGTLNGLWIELDQYQNLKMKCDSDSTTLNKFIEKARIFKFLSGLNSEFDPIRVQILGKEQ 208
+G LW EL + + D L++ E+ ++F L LN ++ + +L E+
Sbjct: 133 FGKFRSLWSELKSLRPGTL----DPKILHERREQDKVFGLLLTLNPGYNDLIKHLLRSEK 188
Query: 209 FPSLSEVFSIVRGEETRRTVMVEDKSVDGSALASGKGPIKGSTS-FGRPNRDDRPNRDDR 267
PSL EV S ++ E+ GST FG + N+ +
Sbjct: 189 LPSLDEVCSKIQKEQ-------------------------GSTGLFGGKSELITANKGEV 223
Query: 268 FNGDDRFNRDDR----CTYCKRSGHTKEYCFKLYGRENVLARMSKSKGAPQRRANHTASD 323
+ +DR C +CK+ GHTK+ C+ L+ + +K K + + T +
Sbjct: 224 VANKGVYKNEDRKLLTCDHCKKKGHTKDKCWLLHPH----LKPAKFKDSRAHFSQETHEE 279
Query: 324 MENDGEVPPVPPAEDVPSLSKAELERLRVFMDSLSKPSGSCSLTMTGKSSTFLSLNALST 383
G + K++LE L + SL K SG + T S
Sbjct: 280 QSQAGSSKGETSTSFGDYVRKSDLEALIKSIVSL-KESGITFSSQTSSGSI--------- 329
Query: 384 ENDWIIDFGATDHMTPHPSHLSSYSTYPGKHYITVANGSQNQITGCGNIHLSPSLPLKRV 443
+ID GA+ HM + + L + P ++ +ANG + I G GN+ L +
Sbjct: 330 ----VIDSGASHHMISNSNLLDNIE--PALGHVIIANGDKVPIEGIGNLKLFNKD--SKA 381
Query: 444 LHVPKLSNNLLSIHKLTRDLNCAVTFFHSHGVFQDLATGQIIGIAKEQDGLYCLQH---E 500
+PK ++NLLS+ + TRDLNC F + FQD+ TG++IG + LY L+
Sbjct: 382 FFMPKFTSNLLSVKRTTRDLNCYAIFGPNDVYFQDIETGKVIGEGGSKGELYVLEDLSPN 441
Query: 501 ESKCHQTSSE---SWNTSQIWLRHKRLGHPPFSILRTMFPHLFSKVSVESFHCDVCQFSK 557
S C + S S+NT +W H RLGHP L+ M P+ +S + C+ C K
Sbjct: 442 SSSCFSSKSHLGISFNT--LW--HARLGHPHTRALKLMLPN----ISFDHTSCEACILGK 493
Query: 558 HHRSTFLPSNNKCAQPFDLVHSDVWGPSSISNVSGAKWFVTFIDDCTRITWVFLMKDKSE 617
H +S F S + FDLVHSDVW +S + K+FVTFI++ ++ TW+ L+ K
Sbjct: 494 HCKSVFPKSLTIYEKCFDLVHSDVWTSPCVSRDNN-KYFVTFINEKSKYTWITLLPSKDR 552
Query: 618 VFQLFANFFHMIRTQFGKPIKRLRSDNGKE*VNKNFSEFLTKNGVVHELTCVDAPQQNGV 677
VF+ F NF + QF IK R+DNG E ++ F + L K G++H+ +C PQQNGV
Sbjct: 553 VFEAFTNFETYVTNQFNAKIKVFRTDNGGEYTSQKFRDHLAKRGIIHQTSCPYTPQQNGV 612
Query: 678 AERKNRHLLEITRALLFQMNVPKFYWGEAVLTAAYLINRLPSRVLSNVSPVLVMTSFFPS 737
AERKNRHL+E+ R+++F +VPK +WG+AVLTA YLINR P++VLS++SP V+ + P
Sbjct: 613 AERKNRHLMEVARSMMFHTSVPKRFWGDAVLTACYLINRTPTKVLSDLSPFEVLNNTKPF 672
Query: 738 VPIKSGLQSRVFGCSAFVHVHGHHRGKLDPRAIKCIFIGYASDKKGYKCYHPPSRRVYIS 797
+ RVFGC FV + G R KLD ++ KC+F+GY++ +KGYKC+ P R +IS
Sbjct: 673 ID-----HLRVFGCVCFVLIPGEQRSKLDAKSTKCMFLGYSTTQKGYKCFDPTKNRTFIS 727
Query: 798 MDVTFQESESFYPNPQLQG-GDTQEAESPELSVIPLLQEPLMPSSIPITEDSDDDDNGPE 856
DV F E++ + + D + S + + L + L S T+ + E
Sbjct: 728 RDVKFLENQDYNNKKDWENLKDLTHSTSDRVETLKFLLDHLGNDSTSTTQHQPEMTQDQE 787
Query: 857 QVPNQNDDNGLELVPDQNDDNGPELVPDQNNVDRFRIKYQRKVKPVLIQQQDQPSDPEVS 916
+ +N++ L Q+ +N + D N Q + + +P
Sbjct: 788 DLNQENEEVSL-----QHQENLTHVQEDPPNTQEHSEHVQE-----IQDDSSEDEEPTQV 837
Query: 917 VPNPEPSNSYEHNLDDLPIALRKGKR----SCAKYPISQYVSTQNLSVQHQSFISAIDSI 972
+P P P +R+ K + +P S + + HQ+F+S I
Sbjct: 838 LPPPPPLRRSTR--------IRRKKEFFNSNAVAHPFQATCSLALVPLDHQAFLSKISEH 889
Query: 973 RVPTSVQEVLKDKKWIQAMNEEMHALDKNGTRKIVEKPNDKRTVGCKWIYTVKYRSDGTL 1032
+P + +E ++ K+W A+ +E++A+ +N T + P K+TV +W++T+KY+S+G +
Sbjct: 890 WIPQTYEEAMEVKEWRDAIADEINAMKRNHTWDEDDLPKGKKTVSSRWVFTIKYKSNGDI 949
Query: 1033 DRYKARLVAKGYTQTYGIDYDETFAPVAKMNTVRIILSLAAHFDWELQQFDVKNAFLHGD 1092
+RYK RLVA+G+TQTYG DY ETFAPVAK++TVR++L+LA + W L Q DVKNAFL G+
Sbjct: 950 ERYKTRLVARGFTQTYGSDYMETFAPVAKLHTVRVVLALATNLSWGLWQMDVKNAFLQGE 1009
Query: 1093 LEEEVYMEIPPGYDPTSGRNKVCKLKKALYGLKQSPRAWFGRFTNAMVSLGYKQSQGDHT 1152
LE++VYM PPG + T +KV +L+KA+YGLKQSPRAW+ + + + G+K+S+ DHT
Sbjct: 1010 LEDDVYMTPPPGLEDTIPCDKVLRLRKAIYGLKQSPRAWYHKLSRTLKDHGFKKSESDHT 1069
Query: 1153 LFIKHTRNGKLTLLLVYVNDMIIAGNDELEKQNLRKELAVQFEMKDLGKLKYFLGMEVAY 1212
LF + G + ++L+YV+D+II G+++ + + L F++KDLG+LKYFLG+EV
Sbjct: 1070 LFTLQSPQG-IVVVLIYVDDLIITGDNKDGIDSTKTFLKSCFDIKDLGELKYFLGIEVCR 1128
Query: 1213 SKQGIFISQRKYVLDLLKETGKLGCKPMGVPIEQNHKI---GSSEEDLRVHKAQYQRLVG 1269
S G+F+SQRKY LDLL ETG + KP P+E +K+ G E++ Y++LVG
Sbjct: 1129 SNAGLFLSQRKYTLDLLNETGFMDAKPARTPLEDGYKVNRKGEKEDEKFGDAPLYRKLVG 1188
Query: 1270 KLIYLSHTRPDIAYPVSVVSQFMHDPQERHLQAVDRILQYLKASPGRGLLFKKSEQLSME 1329
KLIYL++TRPDI + V+ VSQ M P H V+RIL+YLK S G+G+ K+ +
Sbjct: 1189 KLIYLTNTRPDICFAVNQVSQHMKVPMVYHWNMVERILRYLKGSSGQGIWMGKNSSTEIV 1248
Query: 1330 VYTDADYAGSIVDRRSTTGYCMFLGGNLVTWRSKKQNVVARSSAEAEFRAMAQGVCELLR 1389
Y DADYAG DRRS TGYC F+GGNL TW++KKQ VV+ SSAE+E+RAM + EL
Sbjct: 1249 GYCDADYAGDRGDRRSKTGYCTFIGGNLATWKTKKQKVVSCSSAESEYRAMRKLTNELTW 1308
Query: 1390 MKIILDDLKIKYEAPMRLFCDNKSAISIAHNPVQHDRTKHIEIDRHFIKEKLNSSLISTQ 1449
+K +L DL I+ P+ + CDNK+AI IA N V H+RTKHIE+D H ++EK+ +
Sbjct: 1309 LKALLKDLGIEQHMPITMHCDNKAAIYIASNSVFHERTKHIEVDCHKVREKIIEGVTLPC 1368
Query: 1450 YVPSGLQLADVLTKGLPLERFRELTCKLGMIDI 1482
Y S QLAD+ TK L+ + KLG++D+
Sbjct: 1369 YTRSEDQLADIFTKAASLKVCNFIHGKLGLVDL 1401
>UniRef100_Q9LVQ2 Retroelement pol polyprotein-like [Arabidopsis thaliana]
Length = 1491
Score = 880 bits (2274), Expect = 0.0
Identities = 542/1534 (35%), Positives = 826/1534 (53%), Gaps = 113/1534 (7%)
Query: 9 PKEKLTSGVKGVSLPPLTHKDLP-VLQGSFRLDGRNYLQWSELVRHTLISRQKISYIEET 67
P+ V VS L D P + S L G NY +WS + + L +++K +I +
Sbjct: 13 PRTNQQPDVTKVSPYTLASSDNPGAMISSVMLTGDNYNEWSTEMLNALQAKRKTGFINGS 72
Query: 68 A--PAETDPQYDTWREENSLIMTWFWYSMIPEISRNYMFFPTAKEIWNNLAQAYSKKQDL 125
P +P Y+ W+ NS+I+ W S+ P++ F A ++W+ L Q +S +
Sbjct: 73 ISKPPLDNPDYENWQAVNSMIVGWIRASIEPKVKSTVTFISDAHQLWSELKQRFSVGNKV 132
Query: 126 SACYELENKVFNSKQGTLSVTDYYGTLNGLWIELDQYQNLKM-KCD--SDSTTL--NKFI 180
++++ ++ +Q V DYYG L LW E Y+ + + KC + TL +K
Sbjct: 133 RV-HQIKAQLAACRQDGQPVIDYYGRLCKLWEEFQIYKPITVCKCGLCTCGATLEPSKER 191
Query: 181 EKARIFKFLSGLN-SEFDPIRVQILGKEQFPSLSEVFS-IVRGEETRRTVMVEDKSVDGS 238
E+ +I +F+ GL+ S F + ++ + FPSL E++S +VR E+ +V + ++
Sbjct: 192 EEEKIHQFVLGLDDSRFGGLSATLIAMDPFPSLGEIYSRVVREEQRLASVQIREQQQSAI 251
Query: 239 ALASGKGPIKGSTSFGRPNRDDRPNRDDRFNGDDRFNRDDRCTYCKRSGHTKEYCFKLYG 298
+ + + T+ GR + +RD R C++C RSGH K+ C+++ G
Sbjct: 252 GFLTRQSEV---TADGRTDSSIIKSRD----------RSVLCSHCGRSGHEKKDCWQIVG 298
Query: 299 -------RENVLARMSKSKGAPQRRANHTASDMENDGEVPPVPPAEDVPSLSKAELERLR 351
R N R S S+G R + S ++ S + ++LR
Sbjct: 299 FPDWWTERTNGGGRGSSSRGRGGRSSGSNNSGRGRGQVTAAHATTSNLSSFPEFTPDQLR 358
Query: 352 VFMDSLSKPSGSCSLTMTGKSSTFLSLNALSTENDWIIDFGATDHMTPHPSHLSSYSTYP 411
V + + S ++GK D I+D GA+ HMT S L++ T P
Sbjct: 359 VITQMIQNKNNGTSDKLSGKMKL----------GDVILDTGASHHMTGQLSLLTNIVTIP 408
Query: 412 GKHYITVANGSQNQITGCGNIHLSPSLPLKRVLHVPKLSNNLLSIHKLTRDLNCAVTFFH 471
+ A+ + G LS ++ L VL+VP L+ +L+S+ KL + + C F
Sbjct: 409 SCS-VGFADDRKTFAISMGTFKLSETVSLSNVLYVPALNCSLISVSKLVKQIKCLALFTD 467
Query: 472 SHGVFQDLATGQIIGIAKEQDGLYCLQHEESKCHQTSSESWNTSQIWLRHKRLGHPPFSI 531
+ V QD + +IG +E+DG+Y L + T + T+ L H+RLGHP FS+
Sbjct: 468 TICVLQDRFSRTLIGTGEERDGVYYLTDAATT---TVHKVDVTTDHALWHQRLGHPSFSV 524
Query: 532 LRTMFPHLFSKVSVESFHCDVCQFSKHHRSTFLPSNNKCAQPFDLVHSDVWGPSSISNVS 591
L ++ S SV S CDVC +K R F S+NK F L+H DVWGP + +
Sbjct: 525 LSSLPLFSGSSCSVSSRSCDVCFRAKQTREVFPDSSNKSTDCFSLIHCDVWGPYRVPSSC 584
Query: 592 GAKWFVTFIDDCTRITWVFLMKDKSEVFQLFANFFHMIRTQFGKPIKRLRSDNGKE*VNK 651
GA +F+T +DD +R W +L+ KSEV + NF QFGK +K +RSDNG E +
Sbjct: 585 GAVYFLTIVDDFSRSVWTYLLLAKSEVRSVLTNFLAYTEKQFGKSVKIIRSDNGTEFMC- 643
Query: 652 NFSEFLTKNGVVHELTCVDAPQQNGVAERKNRHLLEITRALLFQMNVPKFYWGEAVLTAA 711
S + + G+VH+ +CV PQQNG ERK+RH+L ++RALLFQ ++P +WGEAV+TAA
Sbjct: 644 -LSSYFKEQGIVHQTSCVGTPQQNGRVERKHRHILNVSRALLFQASLPIKFWGEAVMTAA 702
Query: 712 YLINRLPSRVLSNVSPVLVMTSFFPSVPIKSGLQSRVFGCSAFVHVHGHHRGKLDPRAIK 771
YLINR PS + + +SP ++ P Q RVFG + + H + K R+
Sbjct: 703 YLINRTPSSIHNGLSPYELLHGCKPDYD-----QLRVFGSACYAHRVTRDKDKFGERSRL 757
Query: 772 CIFIGYASDKKGYKCYHPPSRRVYISMDVTFQESESFYPNPQLQGGDTQEAESPELSVIP 831
CIF+GY +KG+K Y + +S DV F+E+ Y + GDT +P ++
Sbjct: 758 CIFVGYPFGQKGWKVYDLSTNEFIVSRDVVFRENVFPYATNE---GDT--IYTPPVTCPI 812
Query: 832 LLQEPLMP--------------SSIPI----TEDSDDDDNGPEQVPNQNDDN---GLELV 870
E +P S P+ +SD + + P+ +P DD +
Sbjct: 813 TYDEDWLPFTTLEDRGSDENSLSDPPVCVTDVSESDTEHDTPQSLPTPVDDPLSPSTSVT 872
Query: 871 PDQNDDNGPELVPDQNNVD------------------RFRIKYQRKVKP-VLIQQQDQPS 911
P Q N NV + +++ ++K +L P+
Sbjct: 873 PTQTPTNSSSSTSPSTNVSPPQQDTTPIIENTPPRQGKRQVQQLARLKDYILYNASCTPN 932
Query: 912 DPEVSVPNPEPSNSYEHNLDDLPIALRKGKRSCAKYPISQYVSTQNLSVQHQSFISAIDS 971
P V P+ S+S + ++YP++ Y+ + S H+ F++AI +
Sbjct: 933 TPHVLSPSTSQSSS--------------SIQGNSQYPLTDYIFDECFSAGHKVFLAAITA 978
Query: 972 IRVPTSVQEVLKDKKWIQAMNEEMHALDKNGTRKIVEKPNDKRTVGCKWIYTVKYRSDGT 1031
P +E +K K W AM +E+ AL+ N T IV+ P K +G +W+Y K+ +DGT
Sbjct: 979 NDEPKHFKEAVKVKVWNDAMYKEVDALEVNKTWDIVDLPTGKVAIGSQWVYKTKFNADGT 1038
Query: 1032 LDRYKARLVAKGYTQTYGIDYDETFAPVAKMNTVRIILSLAAHFDWELQQFDVKNAFLHG 1091
++RYKARLV +G Q G DY ETFAPV KM TVR +L L A WE+ Q DV NAFLHG
Sbjct: 1039 VERYKARLVVQGNNQIEGEDYTETFAPVVKMTTVRTLLRLVAANQWEVYQMDVHNAFLHG 1098
Query: 1092 DLEEEVYMEIPPGYDPTSGRNKVCKLKKALYGLKQSPRAWFGRFTNAMVSLGYKQSQGDH 1151
DLEEEVYM++PPG+ S +KVC+L+K+LYGLKQ+PR WF + ++A+ G+ Q D+
Sbjct: 1099 DLEEEVYMKLPPGFRH-SHPDKVCRLRKSLYGLKQAPRCWFKKLSDALKRFGFIQGYEDY 1157
Query: 1152 TLFIKHTRNGKLTLLLVYVNDMIIAGNDELEKQNLRKELAVQFEMKDLGKLKYFLGMEVA 1211
+ F ++ G +LVYV+D+II GNDE Q ++ L F MKDLGKLKYFLG+EV+
Sbjct: 1158 SFF-SYSCKGIELRVLVYVDDLIICGNDEYMVQKFKEYLGRCFSMKDLGKLKYFLGIEVS 1216
Query: 1212 YSKQGIFISQRKYVLDLLKETGKLGCKPMGVPIEQNHKIGSSEEDLRVHKAQYQRLVGKL 1271
GIF+SQRKY LD++ ++G LG +P P+EQNH + S + L ++RLVG+L
Sbjct: 1217 RGPDGIFLSQRKYALDIISDSGTLGARPAYTPLEQNHHLASDDGPLLQDPKPFRRLVGRL 1276
Query: 1272 IYLSHTRPDIAYPVSVVSQFMHDPQERHLQAVDRILQYLKASPGRGLLFKKSEQLSMEVY 1331
+YL HTRP+++Y V V+SQFM P+E HL+A RI++YLK SPG+G+L ++ L++EVY
Sbjct: 1277 LYLLHTRPELSYSVHVLSQFMQAPREAHLEAAMRIVRYLKGSPGQGILLSSNKDLTLEVY 1336
Query: 1332 TDADYAGSIVDRRSTTGYCMFLGGNLVTWRSKKQNVVARSSAEAEFRAMAQGVCELLRMK 1391
D+D+ + RRS + Y + LGG+ ++W++KKQ+ V+ SSAEAE+RAM+ + E+ +
Sbjct: 1337 CDSDFQSCPLTRRSLSAYVVLLGGSPISWKTKKQDTVSHSSAEAEYRAMSVALKEIKWLN 1396
Query: 1392 IILDDLKIKYEAPMRLFCDNKSAISIAHNPVQHDRTKHIEIDRHFIKEKLNSSLISTQYV 1451
+L +L I AP RLFCD+K+AISIA NPV H+RTKHIE D H +++ + +I+T +V
Sbjct: 1397 KLLKELGITLAAPTRLFCDSKAAISIAANPVFHERTKHIERDCHSVRDAVRDGIITTHHV 1456
Query: 1452 PSGLQLADVLTKGLPLERFRELTCKLGMIDIHSP 1485
+ QLAD+ TK L +F L KLG+ ++H+P
Sbjct: 1457 RTSEQLADIFTKALGRNQFIYLMSKLGIQNLHTP 1490
>UniRef100_Q9ZPU4 Putative retroelement pol polyprotein [Arabidopsis thaliana]
Length = 1501
Score = 874 bits (2257), Expect = 0.0
Identities = 533/1493 (35%), Positives = 816/1493 (53%), Gaps = 81/1493 (5%)
Query: 36 SFRLDGRNYLQWSELVRHTLISRQKISYIEETAPAE--TDPQYDTWREENSLIMTWFWYS 93
S L+G NY QW+ + + L +++K +I T P DP Y+ W NS+I+ W S
Sbjct: 46 SVELNGDNYNQWATEMLNALQAKRKTGFINGTIPRPPPNDPNYENWTAVNSMIVGWIRTS 105
Query: 94 MIPEISRNYMFFPTAKEIWNNLAQAYSKKQDLSACYELENKVFNSKQGTLSVTDYYGTLN 153
+ P++ F A +W +L Q +S + +++ ++ + +Q +V +YYG L+
Sbjct: 106 IEPKVKATVTFISDAHLLWKDLKQRFSVGNKVRI-HQIRAQLSSCRQDGQAVIEYYGRLS 164
Query: 154 GLWIELDQYQNLKM------KCDSDSTTLNKFIEKARIFKFLSGLN-SEFDPIRVQILGK 206
LW E + Y+ + + +C + S K E+ +I +F+ GL+ S F + ++
Sbjct: 165 NLWEEYNIYKPVTVCTCGLCRCGATSEP-TKEREEEKIHQFVLGLDESRFGGLCATLINM 223
Query: 207 EQFPSLSEVFS-IVRGEETRRTVMVEDKSVDGSALASGKGPIKGSTSFGRPNRDDRPNRD 265
+ PSL E++S ++R E+ +V V ++ + + + + +R D +
Sbjct: 224 DPLPSLGEIYSRVIREEQRLASVHVREQKEEAVGFLARREQLD------HHSRVDASSSR 277
Query: 266 DRFNGDDRFNRDDR----CTYCKRSGHTKEYCFKLYGRENVLARMSKSKGAPQR-RANHT 320
G R N + C+ C R+GH K+ C+++ G + + + +G+ R R
Sbjct: 278 SEHTGGSRSNSIIKGRVTCSNCGRTGHEKKECWQIVGFPDWWSERNGGRGSNGRGRGGRG 337
Query: 321 ASDMENDGEVPPVPPAEDVPSLSKAELERLRVFMDSLSKPSGSCSLTMTGKSSTFLSLNA 380
++ G+V S+ E + L K + T S L+
Sbjct: 338 SNGGRGQGQVMAAHATSSNSSVFPEFTEEHMRVLSQLVKEKSNSGSTSNNNSDR---LSG 394
Query: 381 LSTENDWIIDFGATDHMTPHPSHLSSYSTYPGKHYITVANGSQNQITGCGNIHLSPSLPL 440
+ D I+D GA+ HMT S L++ P + A+GS+ G + LS ++ L
Sbjct: 395 KTKLGDIILDSGASHHMTGTLSSLTNVVPVPPCP-VGFADGSKAFALSVGVLTLSNTVSL 453
Query: 441 KRVLHVPKLSNNLLSIHKLTRDLNCAVTFFHSHGVFQDLATGQIIGIAKEQDGLYCLQH- 499
VL VP L+ L+S+ KL + C TF + QD ++ +IG +E+ G+Y L
Sbjct: 454 TNVLFVPSLNCTLISVSKLLKQTQCLATFTDTLCFLQDRSSKTLIGSGEERGGVYYLTDV 513
Query: 500 EESKCHQTSSESWNTSQIWLRHKRLGHPPFSILRTMFPHLFSKVS--VESFHCDVCQFSK 557
+K H + +S +W H+RLGHP FS+L ++ LFSK S V S CDVC +K
Sbjct: 514 TPAKIHTANVDS--DQALW--HQRLGHPSFSVLSSL--PLFSKTSSTVTSHSCDVCFRAK 567
Query: 558 HHRSTFLPSNNKCAQPFDLVHSDVWGPSSISNVSGAKWFVTFIDDCTRITWVFLMKDKSE 617
R F S NK + F L+H DVWGP + GA +F+T +DD +R W +L+ +KSE
Sbjct: 568 QTREVFPESINKTEECFSLIHCDVWGPYRVPASCGAVYFLTIVDDYSRAVWTYLLLEKSE 627
Query: 618 VFQLFANFFHMIRTQFGKPIKRLRSDNGKE*VNKNFSEFLTKNGVVHELTCVDAPQQNGV 677
V Q+ NF QFGK +K +RSDNG E + S + +NG++H+ +CV PQQNG
Sbjct: 628 VRQVLTNFLKYAEKQFGKTVKMVRSDNGTEFMC--LSSYFRENGIIHQTSCVGTPQQNGR 685
Query: 678 AERKNRHLLEITRALLFQMNVPKFYWGEAVLTAAYLINRLPSRVLSNVSPVLVMTSFFPS 737
ERK+RH+L + RALLFQ ++P +WGE++LTAAYLINR PS +LS +P V+ S
Sbjct: 686 VERKHRHILNVARALLFQASLPIKFWGESILTAAYLINRTPSSILSGRTPYEVLHG---S 742
Query: 738 VPIKSGLQSRVFGCSAFVHVHGHHRGKLDPRAIKCIFIGYASDKKGYKCYHPPSRRVYIS 797
P+ S Q RVFG + +VH + K R+ CIF+GY KKG+K Y +S
Sbjct: 743 KPVYS--QLRVFGSACYVHRVTRDKDKFGQRSRSCIFVGYPFGKKGWKVYDIERNEFLVS 800
Query: 798 MDVTFQESESFYPNPQLQGGDTQEAESPELS-----VIPLLQ-----EPLMPSSIPITED 847
DV F+E +P + P +S IP L+ + + + T D
Sbjct: 801 RDVIFREE--VFPYAGVNSSTLASTSLPTVSEDDDWAIPPLEVRGSIDSVETERVVCTTD 858
Query: 848 SD--DDDNGPEQVPNQNDDNGLELVPDQNDDNGPELVPDQNNVDRFRIKYQRKV-KPVLI 904
D ++PNQ E VPD + P V + + V P+ +
Sbjct: 859 EVVLDTSVSDSEIPNQ------EFVPDDTPPSSPLSVSPSGSPNTPTTPIVVPVASPIPV 912
Query: 905 QQQDQPSDPEVSVPNPE------------PSNSYEHNLDDLPIALRKGKRSCAKYPISQY 952
Q + P P+ PS+ + D + GK + +P++ Y
Sbjct: 913 SPPKQRKSKRATHPPPKLNDYVLYNAMYTPSSIHALPADPSQSSTVPGK---SLFPLTDY 969
Query: 953 VSTQNLSVQHQSFISAIDSIRVPTSVQEVLKDKKWIQAMNEEMHALDKNGTRKIVEKPND 1012
VS S H+++++AI P +E ++ K W AM E+ AL+ N T IV+ P
Sbjct: 970 VSDAAFSSSHRAYLAAITDNVEPKHFKEAVQIKVWNDAMFTEVDALEINKTWDIVDLPPG 1029
Query: 1013 KRTVGCKWIYTVKYRSDGTLDRYKARLVAKGYTQTYGIDYDETFAPVAKMNTVRIILSLA 1072
K +G +W++ KY SDGT++RYKARLV +G Q G DY ETFAPV +M TVR +L
Sbjct: 1030 KVAIGSQWVFKTKYNSDGTVERYKARLVVQGNKQVEGEDYKETFAPVVRMTTVRTLLRNV 1089
Query: 1073 AHFDWELQQFDVKNAFLHGDLEEEVYMEIPPGYDPTSGRNKVCKLKKALYGLKQSPRAWF 1132
A WE+ Q DV NAFLHGDLEEEVYM++PPG+ S +KVC+L+K+LYGLKQ+PR WF
Sbjct: 1090 AANQWEVYQMDVHNAFLHGDLEEEVYMKLPPGFRH-SHPDKVCRLRKSLYGLKQAPRCWF 1148
Query: 1133 GRFTNAMVSLGYKQSQGDHTLFIKHTRNGKLTLLLVYVNDMIIAGNDELEKQNLRKELAV 1192
+ +++++ G+ QS D++LF +TRN +L+YV+D++I GND Q + L+
Sbjct: 1149 KKLSDSLLRFGFVQSYEDYSLF-SYTRNNIELRVLIYVDDLLICGNDGYMLQKFKDYLSR 1207
Query: 1193 QFEMKDLGKLKYFLGMEVAYSKQGIFISQRKYVLDLLKETGKLGCKPMGVPIEQNHKIGS 1252
F MKDLGKLKYFLG+EV+ +GIF+SQRKY LD++ ++G LG +P P+EQNH + S
Sbjct: 1208 CFSMKDLGKLKYFLGIEVSRGPEGIFLSQRKYALDVIADSGNLGSRPAHTPLEQNHHLAS 1267
Query: 1253 SEEDLRVHKAQYQRLVGKLIYLSHTRPDIAYPVSVVSQFMHDPQERHLQAVDRILQYLKA 1312
+ L Y+RLVG+L+YL HTRP+++Y V V++QFM +P+E H A R+++YLK
Sbjct: 1268 DDGPLLSDPKPYRRLVGRLLYLLHTRPELSYSVHVLAQFMQNPREAHFDAALRVVRYLKG 1327
Query: 1313 SPGRGLLFKKSEQLSMEVYTDADYAGSIVDRRSTTGYCMFLGGNLVTWRSKKQNVVARSS 1372
SPG+G+L L++EVY D+D+ + RRS + Y + LGG+ ++W++KKQ+ V+ SS
Sbjct: 1328 SPGQGILLNADPDLTLEVYCDSDWQSCPLTRRSISAYVVLLGGSPISWKTKKQDTVSHSS 1387
Query: 1373 AEAEFRAMAQGVCELLRMKIILDDLKIKYEAPMRLFCDNKSAISIAHNPVQHDRTKHIEI 1432
AEAE+RAM+ + E+ ++ +L +L I+ P RL+CD+K+AI IA NPV H+RTKHIE
Sbjct: 1388 AEAEYRAMSYALKEIKWLRKLLKELGIEQSTPARLYCDSKAAIHIAANPVFHERTKHIES 1447
Query: 1433 DRHFIKEKLNSSLISTQYVPSGLQLADVLTKGLPLERFRELTCKLGMIDIHSP 1485
D H +++ + +I+TQ+V + QLADV TK L +F L KLG+ ++H+P
Sbjct: 1448 DCHSVRDAVRDGIITTQHVRTTEQLADVFTKALGRNQFLYLMSKLGVQNLHTP 1500
>UniRef100_Q9C692 Polyprotein, putative [Arabidopsis thaliana]
Length = 1468
Score = 843 bits (2178), Expect = 0.0
Identities = 520/1499 (34%), Positives = 802/1499 (52%), Gaps = 121/1499 (8%)
Query: 39 LDGRNYLQWSELVRHTLISRQKISYIEETAPAETD--PQYDTWREENSLIMTWFWYSMIP 96
L NY +W+ + L SR+K +++ T P D P + W N+L+++W ++
Sbjct: 38 LKTNNYEEWACGFKTALRSRKKFGFLDGTIPQPLDGSPDLEDWLTINALLVSWMKMTIDS 97
Query: 97 EISRNYMFFPTAKEIWNNLAQAYS---------KKQDLSACYELENKVFNSKQGTLSVTD 147
E+ N A+++W + + +S K DL+ C KQ ++V
Sbjct: 98 ELLTNISHRDVARDLWEQIRKRFSVSNGPKNQKMKADLATC----------KQEGMTVEG 147
Query: 148 YYGTLNGLWIELDQYQNLKM-KCD----SDSTTLNKFIEKARIFKFLSGLN-SEFDPIRV 201
YYG LN +W ++ Y+ L++ KC + T K+ E + ++L GLN ++F IR
Sbjct: 148 YYGKLNKIWDNINSYRPLRICKCGRCICNLGTDQEKYREDDMVHQYLYGLNETKFHTIRS 207
Query: 202 QILGKEQFPSLSEVFSIVRGEETRRTVMVEDKSVDGSALASGKGPIKGSTSFGRPNRDDR 261
+ + P L EV++IVR EE MV ++S + T+F R
Sbjct: 208 SLTSRVPLPGLEEVYNIVRQEED----MVNNRSSNEERT--------DVTAFAVQMRPRS 255
Query: 262 PNRDDRFNGDDRFNRDDRCTYCKRSGHTKEYCFKLYG-----------RENVLARMSKSK 310
++F ++ CT+C R GH+ E CF L G + N S+ +
Sbjct: 256 EVISEKFANSEKLQNKKLCTHCNRGGHSPENCFVLIGYPEWWGDRPRGKSNSNGSTSRGR 315
Query: 311 G--APQRRANHTASDMENDGEVPPVPPAEDVPS-LSKAELERLRVFMDSLSKPSGSCSLT 367
G P N P P +E V ++ ++ + + D + G L
Sbjct: 316 GRFGPGFNGGQPRPTYVNVVMTGPFPSSEHVNRVITDSDRDAVSGLTDEQWR--GVVKLL 373
Query: 368 MTGKSSTFLSLNALSTEN-------DWIIDFGATDHMTPHPSHLSSYSTYPGKHYITVAN 420
G+S NA T++ WI+D GA+ HMT + LS + I +A+
Sbjct: 374 NAGRSDN--KSNAHETQSGTCSLFTSWILDTGASHHMTGNLELLSDMRSM-SPVLIILAD 430
Query: 421 GSQNQITGCGNIHLSPSLPLKRVLHVPKLSNNLLSIHKLTRDLNCAVTFFHSHGVFQDLA 480
G++ G + L L LK V +V +L ++L+S+ ++ + +C D
Sbjct: 431 GNKRVAVSEGTVRLGSHLILKSVFYVKELESDLISVGQMMDENHCV-----------DRT 479
Query: 481 TGQIIGIAKEQDGLYCLQHEESKCHQTSSESWNTSQIWLRHKRLGHPPFSILRTMFPHLF 540
T + I K ++G +C + E+ +S + L H+RLGH I+ + L
Sbjct: 480 TRMVTRIGKRENGSFCFRGMENAAAVHTSVK---APFDLWHRRLGHASDKIVNLLPRELL 536
Query: 541 S--KVSVESFHCDVCQFSKHHRSTFLPSNNKCAQPFDLVHSDVWGPSSISNVSGAKWFVT 598
S K +E+ CD C +K R TF S+N+ F L+H DVWGP + SGA++F+T
Sbjct: 537 SSGKEILENV-CDTCMRAKQTRDTFPLSDNRSMDSFQLIHCDVWGPYRAPSYSGARYFLT 595
Query: 599 FIDDCTRITWVFLMKDKSEVFQLFANFFHMIRTQFGKPIKRLRSDNGKE*VNKNFSEFLT 658
+DD +R WV+LM DKSE + +F ++ QF IK +RSDNG E + E+
Sbjct: 596 IVDDYSRGVWVYLMTDKSETQKHLKDFIALVERQFDTEIKIVRSDNGTEFLCMR--EYFL 653
Query: 659 KNGVVHELTCVDAPQQNGVAERKNRHLLEITRALLFQMNVPKFYWGEAVLTAAYLINRLP 718
G+ HE +CV P QNG ERK+RH+L I RAL FQ +P +WGE +L+AAYLINR P
Sbjct: 654 HKGIAHETSCVGTPHQNGRVERKHRHILNIARALRFQSYLPIQFWGECILSAAYLINRTP 713
Query: 719 SRVLSNVSPVLVMTSFFPSVPIKSGLQSRVFGCSAFVHVHGHHRGKLDPRAIKCIFIGYA 778
S +L SP ++ + + P S L RVFG + H H K R+ +C+F+GY
Sbjct: 714 SMLLQGKSPYEML---YKTAPKYSHL--RVFGSLCYAHNQNHKGDKFAARSRRCVFVGYP 768
Query: 779 SDKKGYKCYHPPSRRVYISMDVTFQESESFYPNPQLQGGDTQEAESPELSVIPLLQEPLM 838
+KG++ + ++ ++S DV FQE+E +P ++ + E + P ++E +
Sbjct: 769 HGQKGWRLFDLEEQKFFVSRDVIFQETE--FPYSKMSCNEEDERVLVDCVGPPFIEEAIG 826
Query: 839 PSSIPITEDSDDDDNGPEQVPNQNDDNGLELVPDQNDDNG-PELVPDQNNVDRFRIKYQR 897
P +I I + + GP ++P+ N ++ P +++D F
Sbjct: 827 PRTI-IGRNIGEATVGPNVATGP-------IIPEINQESSSPSEFVSLSSLDPF------ 872
Query: 898 KVKPVLIQQQDQPSDPEVSVP-----------NPEPSNSYEHNLDDLPIALRKGKRSCAK 946
+ +Q D P P P ++ N + ++ S +
Sbjct: 873 -LASSTVQTADLPLSSTTPAPIQLRRSSRQTQKPMKLKNFVTNTVSVE-SISPEASSSSL 930
Query: 947 YPISQYVSTQNLSVQHQSFISAIDSIRVPTSVQEVLKDKKWIQAMNEEMHALDKNGTRKI 1006
YPI +YV + H++F++A+ + PT+ E + DK W +AM+ E+ +L N T I
Sbjct: 931 YPIEKYVDCHRFTSSHKAFLAAVTAGMEPTTYNEAMVDKAWREAMSAEIESLRVNQTFSI 990
Query: 1007 VEKPNDKRTVGCKWIYTVKYRSDGTLDRYKARLVAKGYTQTYGIDYDETFAPVAKMNTVR 1066
V P KR +G KW+Y +KYRSDG ++RYKARLV G Q G+DYDETFAPVAKM+TVR
Sbjct: 991 VNLPPGKRALGNKWVYKIKYRSDGAIERYKARLVVLGNCQKEGVDYDETFAPVAKMSTVR 1050
Query: 1067 IILSLAAHFDWELQQFDVKNAFLHGDLEEEVYMEIPPGYDPTSGRNKVCKLKKALYGLKQ 1126
+ L +AA DW + Q DV NAFLHGDL+EEVYM++P G+ +KVC+L K+LYGLKQ
Sbjct: 1051 LFLGVAAARDWHVHQMDVHNAFLHGDLKEEVYMKLPQGFQ-CDDPSKVCRLHKSLYGLKQ 1109
Query: 1127 SPRAWFGRFTNAMVSLGYKQSQGDHTLFIKHTRNGKLTLLLVYVNDMIIAGNDELEKQNL 1186
+PR WF + ++A+ G+ QS D++LF + +G +LVYV+D+II+G+
Sbjct: 1110 APRCWFSKLSSALKQYGFTQSLSDYSLF-SYNNDGIFVHVLVYVDDLIISGSCPDAVAQF 1168
Query: 1187 RKELAVQFEMKDLGKLKYFLGMEVAYSKQGIFISQRKYVLDLLKETGKLGCKPMGVPIEQ 1246
+ L F MKDLG LKYFLG+EV+ + QG ++SQRKYVLD++ E G LG +P P+EQ
Sbjct: 1169 KSYLESCFHMKDLGLLKYFLGIEVSRNAQGFYLSQRKYVLDIISEMGLLGARPSAFPLEQ 1228
Query: 1247 NHKIGSSEEDLRVHKAQYQRLVGKLIYLSHTRPDIAYPVSVVSQFMHDPQERHLQAVDRI 1306
NHK+ S L ++Y+RLVG+LIYL TRP+++Y V ++QFM +P++ H A R+
Sbjct: 1229 NHKLSLSTSPLLSDSSRYRRLVGRLIYLVVTRPELSYSVHTLAQFMQNPRQDHWNAAIRV 1288
Query: 1307 LQYLKASPGRGLLFKKSEQLSMEVYTDADYAGSIVDRRSTTGYCMFLGGNLVTWRSKKQN 1366
++YLK++PG+G+L + L + + D+DYA + RRS TGY + LG ++W++KKQ
Sbjct: 1289 VRYLKSNPGQGILLSSTSTLQINGWCDSDYAACPLTRRSLTGYFVQLGDTPISWKTKKQP 1348
Query: 1367 VVARSSAEAEFRAMAQGVCELLRMKIILDDLKIKYEAPMRLFCDNKSAISIAHNPVQHDR 1426
V+RSSAEAE+RAMA EL+ +K +L DL + + MR+F D+KSAI+++ NPVQH+R
Sbjct: 1349 TVSRSSAEAEYRAMAFLTQELMWLKRVLYDLGVSHVQAMRIFSDSKSAIALSVNPVQHER 1408
Query: 1427 TKHIEIDRHFIKEKLNSSLISTQYVPSGLQLADVLTKGLPLERFRELTCKLGMIDIHSP 1485
TKH+E+D HFI++ + +I+T +VPS QLAD+LTK L + R KLG++D+H+P
Sbjct: 1409 TKHVEVDCHFIRDAILDGIIATSFVPSHKQLADILTKALGEKEVRYFLRKLGILDVHAP 1467
>UniRef100_Q9FXB7 Putative retroelement polyprotein [Arabidopsis thaliana]
Length = 1486
Score = 842 bits (2176), Expect = 0.0
Identities = 522/1515 (34%), Positives = 809/1515 (52%), Gaps = 139/1515 (9%)
Query: 39 LDGRNYLQWSELVRHTLISRQKISYIEETAP--AETDPQYDTWREENSLIMTWFWYSMIP 96
L G NY +W+ +R L +R+K + + + P ETDP ++ W N+L+++W ++
Sbjct: 42 LRGPNYDEWATNLRLALKARKKFGFADGSIPQPVETDPDFEDWTANNALVVSWMKLTIDE 101
Query: 97 EISRNYMFFPTAKEIWNNLAQAYSKKQDLSACYELENKVFNSKQGTLSVTDYYGTLNGLW 156
+S + + E+W ++ + + K L+ ++ +Q +++ YYG L+ LW
Sbjct: 102 TVSTSMSHLDDSHELWTHIQKRFGVKNG-QRVQRLKTELATCRQKGVAIETYYGRLSQLW 160
Query: 157 IELDQYQNLKMKCDSDSTTLNKFIEKARIFKFLSGLN-SEFDPIRVQILGKEQFPSLSEV 215
L YQ K D + K E+ ++ +FL GL+ S + ++ +L + PSL E
Sbjct: 161 RSLADYQQAKTMDD-----VRKEREEDKLHQFLMGLDESVYGAVKSALLSRVPLPSLEEA 215
Query: 216 FS-IVRGEETRRTVMVEDKSVDGSALASGKGPIKGSTSFGRPNRDDRPNRDDRFNGDDRF 274
++ + + EE++ + ++ VDG + A + RP RD NR
Sbjct: 216 YNALTQDEESKSLSRLHNERVDGVSFAV--------QTTSRP-RDSSENRV--------- 257
Query: 275 NRDDRCTYCKRSGHTKEYCFKLYGRENVLAR--------------MSKSKGAPQR----R 316
C+ C R GH E CFKL G L +S KG
Sbjct: 258 -----CSNCGRVGHLAEQCFKLIGYPPWLEEKLRLKNTASSSRGGLSSFKGKQSHGRGSS 312
Query: 317 ANHTASD------MENDGEVPPVPPAEDVPSLSKAELERLRVFMDSLSKPSGSCSLTMTG 370
NH AS + N P+ ++D LS + ++ L + + + +G
Sbjct: 313 INHVASSGMAANVVTNSSLTSPLT-SDDRIGLSGLNDSQWKILQTILEERKSTSNDHQSG 371
Query: 371 KSSTFLSLNALSTENDWIIDFGATDHMTPHPSHLSSYSTYPGKHYITVANGSQNQITGCG 430
K FL WIID GAT+HMT + L + P I + +G T G
Sbjct: 372 KY--FLE--------SWIIDSGATNHMTGSLAFLRNVCDMPPV-LIKLPDGRFTTATKQG 420
Query: 431 NIHLSPSLPLKRVLHVPKLSNNLLSIHKLTRDLNCAVTFFHSHGVFQDLATGQIIGIAKE 490
++ L SL L+ VL V L +L+S+ +LTR C + QD T +IG +E
Sbjct: 421 SVQLGSSLDLQDVLFVDGLHCHLISVSQLTRTRRCIFQITDKVCIVQDRTTLMLIGAGRE 480
Query: 491 QDGLYCLQHEESKCHQTSSESWNTSQIWLRHKRLGHPPFSILRTMFPHLFSKVSVESFH- 549
+GLY + E+ TS ++ +SQ+W H+RLGHP L + FS V+ +F
Sbjct: 481 LNGLYFFRGVETAAAVTS-KALPSSQLW--HQRLGHPSSKALHLL---PFSDVTSSTFDS 534
Query: 550 --CDVCQFSKHHRSTFLPSNNKCAQPFDLVHSDVWGPSSISNVSGAKWFVTFIDDCTRIT 607
C++C +K R F S+NK + F+LVH D+WGP +++ G+++F+T +DD +R
Sbjct: 535 KTCEICIQAKQTRDPFPLSSNKTSFAFELVHCDLWGPYRTTSICGSRYFLTLVDDYSRAV 594
Query: 608 WVFLMKDKSEVFQLFANFFHMIRTQFGKPIKRLRSDNGKE*VNKNFSEFLTKNGVVHELT 667
W++L+ K E + NF ++ Q+ IK +RSDNG E + S+F + G++HE +
Sbjct: 595 WLYLLPSKQEAPKHLKNFIALVERQYTTNIKMIRSDNGSEFIC--LSDFFAQKGIIHETS 652
Query: 668 CVDAPQQNGVAERKNRHLLEITRALLFQMNVPKFYWGEAVLTAAYLINRLPSRVLSNVSP 727
CV PQQNG ERK+RH+L + RAL FQ +P +W LTAAYLINR P+ +L +P
Sbjct: 653 CVGTPQQNGRVERKHRHILNVARALRFQSGLPIEFWSYCALTAAYLINRTPTPLLKGKTP 712
Query: 728 VLVMTSFFPSVPIKSGLQSRVFGCSAFVHVHGHHRGKLDPRAIKCIFIGYASDKKGYKCY 787
++ + P + R+FGC +VH H K R+ K IF+GY KKG++ Y
Sbjct: 713 FELIYNRPPPLQ-----HIRIFGCICYVHNLKHGGDKFASRSNKSIFLGYPFAKKGWRVY 767
Query: 788 HPPSRRVYISMDVTFQESESFYP------NPQLQGGDTQEAESPELSVIPLL-------- 833
+ + V +S DV F+E+E +P +P L +E E+S+ P +
Sbjct: 768 NIETGVVSVSRDVVFRETEFHFPISVMDSSPSLDPVLVDSSELEEISMTPPVTPSSPATP 827
Query: 834 QEPLMPSSIPITEDSDDDDNGP-----------------------EQVPNQNDDNGLELV 870
P+ PSS P+T S + P + + D ++
Sbjct: 828 SSPVTPSS-PVTPSSPVSPSSPVTPSSPVTPVSSTTTSAAIDTIEDITTDLEDSTSMDFF 886
Query: 871 PDQNDDNGPELVPDQNNVDRFRIKYQRKVKPVLIQQQDQPSDPEVSVPNPEPSNSYEHNL 930
PD D+ P ++ + V+ L+ + +P P V +
Sbjct: 887 PDDEDEFSPTAT--ESPASSSSPVHPPAVQLELLGKGHRPKRPPVKLA------------ 932
Query: 931 DDLPIALRKGKRSCAKYPISQYVSTQNLSVQHQSFISAIDSIRVPTSVQEVLKDKKWIQA 990
D + L + S YP+ Y+S+ S +Q++I AI S P + E + D W A
Sbjct: 933 DYVTTLLHQPFPSATPYPLDNYISSSRFSDNYQAYILAITSGNEPRNYNEAMLDDHWKGA 992
Query: 991 MNEEMHALDKNGTRKIVEKPNDKRTVGCKWIYTVKYRSDGTLDRYKARLVAKGYTQTYGI 1050
++ E+ +L+ GT + + P K+ +GCKW++ +KY+SDGTL+R+KARLV G QT G+
Sbjct: 993 VSHEIGSLENLGTWTVEDLPPGKKALGCKWVFRLKYKSDGTLERHKARLVVLGNNQTEGL 1052
Query: 1051 DYDETFAPVAKMNTVRIILSLAAHFDWELQQFDVKNAFLHGDLEEEVYMEIPPGYDPTSG 1110
DY ETFAPVAKM TVR L DWE+ Q DV NAFLHGDL+EEVYM+ PPG+ T
Sbjct: 1053 DYTETFAPVAKMVTVRAFLQQVVSLDWEVHQMDVHNAFLHGDLDEEVYMQFPPGFR-TGD 1111
Query: 1111 RNKVCKLKKALYGLKQSPRAWFGRFTNAMVSLGYKQSQGDHTLFIKHTRNGKLTLLLVYV 1170
+ KVC+L+K+LYGLKQ+PR WF + T+A+ + G+ Q D++LFI H +NG +LVYV
Sbjct: 1112 KTKVCRLRKSLYGLKQAPRCWFAKLTSALKNYGFIQDISDYSLFIFH-KNGVRLHVLVYV 1170
Query: 1171 NDMIIAGNDELEKQNLRKELAVQFEMKDLGKLKYFLGMEVAYSKQGIFISQRKYVLDLLK 1230
+D+II G + L+ F MKDLG L+YFLG+EVA S +GI++ QRKY LD++
Sbjct: 1171 DDLIITGTTIAVITEFKHYLSSCFYMKDLGILRYFLGIEVARSPEGIYLCQRKYALDIIT 1230
Query: 1231 ETGKLGCKPMGVPIEQNHKIGSSEEDLRVHKAQYQRLVGKLIYLSHTRPDIAYPVSVVSQ 1290
ETG LG KP P++QNHK+ + + +Y+RLVG++IYL+ TRP+++Y + ++SQ
Sbjct: 1231 ETGLLGVKPASFPLDQNHKLAFATGETIDDPLRYRRLVGRIIYLATTRPELSYVIHILSQ 1290
Query: 1291 FMHDPQERHLQAVDRILQYLKASPGRGLLFKKSEQLSMEVYTDADYAGSIVDRRSTTGYC 1350
FMH+P+ H +A R+++YLK+SPG+G+L + + L + + D+D+ RS TG+
Sbjct: 1291 FMHNPKPAHWEAALRVVRYLKSSPGQGILLRANTPLVLSAWCDSDFGACPHSDRSLTGWF 1350
Query: 1351 MFLGGNLVTWRSKKQNVVARSSAEAEFRAMAQGVCELLRMKIILDDLKIKYEAPMRLFCD 1410
+ LGG+ ++W+++KQNVV+RSSAEAE+RAMA+ V E++ ++ +L L I AP L D
Sbjct: 1351 IQLGGSPLSWKTQKQNVVSRSSAEAEYRAMAETVSEIIWIRELLPALGIPCTAPTTLHSD 1410
Query: 1411 NKSAISIAHNPVQHDRTKHIEIDRHFIKEKLNSSLISTQYVPSGLQLADVLTKGLPLERF 1470
+ SAIS+A NPV H RTKH+ D HFI+++L + I+T++V + QLAD+LTK L + F
Sbjct: 1411 SLSAISLAANPVYHARTKHVRRDVHFIRDELVNGTIATKHVSTTSQLADILTKALGRKEF 1470
Query: 1471 RELTCKLGMIDIHSP 1485
+ KLG+ ++H P
Sbjct: 1471 ADFLAKLGICNLHIP 1485
>UniRef100_Q851Y3 Putative polyprotein [Oryza sativa]
Length = 1299
Score = 832 bits (2149), Expect = 0.0
Identities = 445/876 (50%), Positives = 591/876 (66%), Gaps = 52/876 (5%)
Query: 638 KRLRSDNGKE*VNKNFSEFLTKNGVVHELTCVDAPQQNGVAERKNRHLLEITRALLFQMN 697
K +R+DNG E +N FS FL+ G++H+ +C D P QNGVAERKNRHLLE R+L++ MN
Sbjct: 447 KFIRTDNGGEYINNEFSSFLSSEGILHQTSCPDTPPQNGVAERKNRHLLETARSLMYAMN 506
Query: 698 VPKFYWGEAVLTAAYLINRLPSRVLSNVSPVLVMTSFFPSVPIKSGLQSRVFGCSAFVHV 757
VPKF W EAV+TA YLINR PSR+L +P ++ V + +VFGC+ FV
Sbjct: 507 VPKFLWSEAVMTATYLINRTPSRILGMKTPCEMIFGKNEFV-----VPPKVFGCTCFVRD 561
Query: 758 HGHHRGKLDPRAIKCIFIGYASDKKGYKCYHPPSRRVYISMDVTFQESESFYPNPQLQGG 817
H GKLDPRA+KCIF+GY+S +KGYKC+ P RR ++SMDVTF+ES FY
Sbjct: 562 HRPSIGKLDPRAVKCIFVGYSSGQKGYKCWSPSERRTFVSMDVTFRESVPFY-------- 613
Query: 818 DTQEAESPELSVIPL-LQEPLMPSSIPITEDSDDDDNGPEQVPNQNDDNGLELVPDQNDD 876
E +LS + + L + +D G EQ + G P
Sbjct: 614 ----GEKTDLSSLFIDLDDSTSGHDGHQMKDEILGPKGDEQPKKEKLVVGSIPCP----- 664
Query: 877 NGPELVPDQN----NVDRFRIKYQRKVKPVLIQQQDQPSDPEVSVPNPE---PSNSYEHN 929
G + +Q + ++ Y R+ + +IQQ ++ + E V + S E
Sbjct: 665 MGGPVAQEQEWRKPHEEKNLQVYTRRTRCPVIQQVEEDNQVEEDVSREQGSLESGGEEEE 724
Query: 930 L---DDLPIALRKGKRSCAKYP-----------------ISQYVSTQNLSVQHQSFISAI 969
+ DLPIA+RK RS A P IS YVS +LS +++FI+++
Sbjct: 725 VREESDLPIAIRKSVRSNAGKPPLRYGFEAQDEGDDENNISNYVSYDSLSSTYKAFIASL 784
Query: 970 DSIRVPTSVQEVLKDKKWIQAMNEEMHALDKNGTRKIVEKPNDKRTVGCKWIYTVKYRSD 1029
DS+++P +E +D +W QAM +E+ AL+KN T +V P K+ V CKW+YTVK D
Sbjct: 785 DSVQIPKDWREAKQDPRWHQAMLDELEALEKNKTWDLVPFPKGKKIVNCKWVYTVKQNPD 844
Query: 1030 GTLDRYKARLVAKGYTQTYGIDYDETFAPVAKMNTVRIILSLAAHFDWELQQFDVKNAFL 1089
G ++RYKARLVAKGY+QTYGIDYDETFAPVAKM+TVR ++S AA+FDW L Q DVKNAFL
Sbjct: 845 GKVERYKARLVAKGYSQTYGIDYDETFAPVAKMSTVRTLISCAANFDWPLHQLDVKNAFL 904
Query: 1090 HGDLEEEVYMEIPPGYDPTSGRNKVCKLKKALYGLKQSPRAWFGRFTNAMVSLGYKQSQG 1149
H DL+EEVYM++PPG+ + + KV +LKK+LYGLKQSPRAWF RF AM ++ YKQ G
Sbjct: 905 HRDLQEEVYMDVPPGFATSQTKGKVLRLKKSLYGLKQSPRAWFDRFRRAMCAMDYKQCNG 964
Query: 1150 DHTLFIKHTRNGKLTLLLVYVNDMIIAGNDELEKQNLRKELAVQFEMKDLGKLKYFLGME 1209
DHT+F H+ + +T+L VYV+DMII GND LE L++ L+ +FE+KDLG+L+YFLG+E
Sbjct: 965 DHTVFYHHSGD-HITILAVYVDDMIITGNDCLEITRLKRNLSKEFEVKDLGQLRYFLGIE 1023
Query: 1210 VAYSKQGIFISQRKYVLDLLKETGKLGCKPMGVPIEQNHKIGSSEEDLRVHKAQYQRLVG 1269
+A S +GI ISQRKYVLDLL ETG LGC P+ PI+QNHK+ + D V++ +YQRLVG
Sbjct: 1024 IARSPRGIVISQRKYVLDLLSETGMLGCCPVSTPIDQNHKLCAESGD-PVNRERYQRLVG 1082
Query: 1270 KLIYLSHTRPDIAYPVSVVSQFMHDPQERHLQAVDRILQYLKASPGRGLLFKKSEQLSME 1329
+LIYL HTRPDI Y VS+VS++MHDP+ H++AV RIL+YLK SPG+GL FKK+ L +E
Sbjct: 1083 RLIYLCHTRPDITYAVSMVSRYMHDPRSSHMEAVYRILRYLKGSPGKGLWFKKNGHLKIE 1142
Query: 1330 VYTDADYAGSIVDRRSTTGYCMFLGGNLVTWRSKKQNVVARSSAEAEFRAMAQGVCELLR 1389
Y DAD+A + DRRST+GYC+++GGNLV+WRSKKQ+VV+RS+AEAE+RAMA + ELL
Sbjct: 1143 GYCDADWASCLDDRRSTSGYCVYVGGNLVSWRSKKQSVVSRSTAEAEYRAMAASLSELLW 1202
Query: 1390 MKIILDDLKIKYEAPMRLFCDNKSAISIAHNPVQHDRTKHIEIDRHFIKEKLNSSLISTQ 1449
++ +L +LKI PM+L CDNKSAI+IA+NPVQHDRTKH+EIDR FIKEKL+ ++
Sbjct: 1203 LRNLLVELKILGNTPMKLLCDNKSAINIANNPVQHDRTKHVEIDRFFIKEKLDEGVLELG 1262
Query: 1450 YVPSGLQLADVLTKGLPLERFRELTCKLGMIDIHSP 1485
+V SG Q+AD LTKGL ++ K+GMIDI+ P
Sbjct: 1263 FVTSGGQVADCLTKGLGVKECNCSCDKMGMIDIYHP 1298
Score = 111 bits (277), Expect = 2e-22
Identities = 112/483 (23%), Positives = 188/483 (38%), Gaps = 105/483 (21%)
Query: 30 LPVLQGSFRLDG-RNYLQWSELVRHTLISRQKISYI--EETAPAE-TDPQYDTWREENSL 85
+ ++Q +L+G +NYL WS L ++ Y+ E P + ++ TW NSL
Sbjct: 42 IELMQNDIKLEGVKNYLSWSRRALLILKTKGLEGYVTGEIKEPENISSVEWKTWSTTNSL 101
Query: 86 IMTWFWYSMIPEISRNYMFFPTAKEIWNNLAQAYSKKQDLSACYELENKVFNSKQGTLSV 145
++ W S+IP I+ +A E+W L YS + ++ E + K+ +QG SV
Sbjct: 102 VVAWLLTSLIPAIATTVETISSASEMWKTLTNLYSGEGNVMLMVEAQEKISVLRQGERSV 161
Query: 146 TDYYGTLNGLWIELDQYQNLKMKCDSDSTTLNKFIEKARIFKFLSGLNSEFDPIRVQILG 205
+Y L LW +LD Y L ++ + K+IE+ R+ +FL GLNSEF+ R +
Sbjct: 162 AEYVAELKHLWSDLDHYDPLGLEHPDCIAKMRKWIERRRVIEFLKGLNSEFEGRRDAMFH 221
Query: 206 KEQFPSLSEVFSIVRGEETRRTVMVEDKSVDGSALASGKGPIKGSTSFGRPNRDDRPNRD 265
+ PSL E + + EE ++ V+ SA S P T +++ R
Sbjct: 222 QTTLPSLDEAIAAMAQEELKKKVL-------PSATPSSPSP----TYVVAQSKETR---- 266
Query: 266 DRFNGDDRFNRDDRCTYCKRSGH-TKEYCF---KLYGRENVLAR--MSKSKGAPQRRANH 319
C C GH ++Y YGR R +G R
Sbjct: 267 -------------ECFNCGEMGHLIRDYRAPRKPSYGRGRFGDRGGARGGRGYAGRGNRG 313
Query: 320 TASDMENDGEVPPVPPAEDVPSLSKAELERLRVFMDSLSKPSGSCSLTMTGKSSTFLSLN 379
+ +D V E + ++ L + SG+ + F+S+N
Sbjct: 314 RGYEYRSDHRANVVTLEESCSGSTNVDVANL------VHSSSGN-------SNQAFMSIN 360
Query: 380 ALSTENDWIIDFGATDHMTPHPSHLSSYSTYPGKHYITVANGSQNQITGCGNIHLSPSLP 439
S+ ++WI+D GA+ H+T + +SS
Sbjct: 361 --SSHSNWILDSGASRHVTVNLVSISS--------------------------------- 385
Query: 440 LKRVLHVPKLSNNLLSIHKLTRDLNCAVTFFHSHGVFQDLATGQIIGIAKEQDGLYCLQH 499
L +NC V+ + + Q+ TG+ +GI +DGL+ L
Sbjct: 386 -------------------LVDHMNCRVSLDRENCLIQERETGKKLGIGVRRDGLWYLDR 426
Query: 500 EES 502
+E+
Sbjct: 427 KET 429
>UniRef100_O81617 F8M12.17 protein [Arabidopsis thaliana]
Length = 1633
Score = 829 bits (2142), Expect = 0.0
Identities = 504/1469 (34%), Positives = 774/1469 (52%), Gaps = 139/1469 (9%)
Query: 43 NYLQWSELVRHTLISRQKISYIEETA--PAETDPQYDTWREENSLIMTWFWYSMIPEISR 100
++ W + L R K+ +I+ T P Y W N + TW S+ +I +
Sbjct: 57 DFHSWRRSIWMALNVRNKLGFIDGTIVKPPLDHRDYGAWSRCNDTVSTWLMNSVSKKIGQ 116
Query: 101 NYMFFPTAKEIWNNLAQAYSKKQDLSACYELENKVFNSKQGTLSVTDYYGTLNGLWIELD 160
+ +F PTA+ IW N+ + K+ D Y++E ++ +QG++ ++ YY L LW E
Sbjct: 117 SLLFIPTAEGIWKNMLSRF-KQDDAPRVYDIEQRLSKIEQGSMDISAYYTELQTLWEEHK 175
Query: 161 QYQNLKM----KCDSDSTTL-NKFIEKARIFKFLSGLNSEFDPIRVQILGKEQFPSLSEV 215
Y +L + +C+ D+ + +++ + KFL GLN ++ R IL + ++ E
Sbjct: 176 NYVDLPVCTCGRCECDAAVKWERLQQRSHVTKFLMGLNESYEQTRRHILMLKPIRTIEEA 235
Query: 216 FSIVRGEETRRTVMVEDKSVDGSALASGKGPIKGSTSFGRPNRDDRPNRDDRFNGDDRFN 275
F+IV +E ++ + K + L K P+
Sbjct: 236 FNIVTQDERQKAIRPTPKVDNQDQL---KLPL---------------------------- 264
Query: 276 RDDRCTYCKRSGHTKEYCFKLYG-----------------RENVLARMSKSKGAPQRRAN 318
CT C + GHT + C+K+ G + + +S+ Q+
Sbjct: 265 ----CTNCGKVGHTVQKCYKIIGYPPGYKAATSYRQPQIQTQPRMQMPQQSQPRMQQPIQ 320
Query: 319 HTASDMENDGEVPPVPPAEDV----PSLSKAELERLRVFMDSLSKPSGSCSL-----TMT 369
H S V P A + P+ + E + S + P S SL +T
Sbjct: 321 HLISQFNAQVRVQE-PAATSIYTSSPTATITEHGLMAQTSTSGTIPFPSTSLKYENNNLT 379
Query: 370 GKSSTFLSLNALSTENDWIIDFGATDHMTPHPSHLSSYSTYPGKHYITVANGSQNQITGC 429
++ T SL + + + WIID GA+ H+ + G +T+ NG++ IT
Sbjct: 380 FQNHTLSSLQNVLSSDAWIIDSGASSHVCSDLTMFRELIHVSGVT-VTLPNGTRVAITHT 438
Query: 430 GNIHLSPSLPLKRVLHVPKLSNNLLSIHKLTRDLNCAVTFFHSHGVFQDLATGQIIGIAK 489
G I ++ +L L VL VP NL+S+ C + +L G +IG K
Sbjct: 439 GTICITSTLILHNVLLVPDFKFNLISV--------CCL----------ELTRGLMIGRGK 480
Query: 490 EQDGLYCLQHEESKCHQTSSESWNTSQIWLRHKRLGHPPFSILRTMFPHLFSKVSVESFH 549
+ LY L+ Q +S S + RH L P L + P L S VS + H
Sbjct: 481 TYNNLYILET------QRTSFSPSLPAATSRHPSL--PALQKLVSSIPSLKS-VSSTASH 531
Query: 550 CDVCQFSKHHRSTFLPSNNKCAQPFDLVHSDVWGPSSISNVSGAKWFVTFIDDCTRITWV 609
C + +K R ++ NN + PFDL+H D+WGP SI +V G ++F+T +DDCTR TWV
Sbjct: 532 CRISPLAKQKRLAYVSHNNLASSPFDLIHLDIWGPFSIESVDGFRYFLTLVDDCTRTTWV 591
Query: 610 FLMKDKSEVFQLFANFFHMIRTQFGKPIKRLRSDNGKE*VNKNFSEFLTKNGVVHELTCV 669
++MK+KSEV +F F +I TQ+ IK +RSDN KE F++F+ + G++H+ +C
Sbjct: 592 YMMKNKSEVSNIFPVFVKLIFTQYNAKIKAIRSDNVKELA---FTKFVKEQGMIHQFSCA 648
Query: 670 DAPQQNGVAERKNRHLLEITRALLFQMNVPKFYWGEAVLTAAYLINRLPSRVLSNVSPVL 729
PQQN V ERK++HLL I R+LLFQ NVP YW + VLTAAYLINRLPS +L N +P
Sbjct: 649 YTPQQNSVVERKHQHLLNIARSLLFQSNVPLQYWSDCVLTAAYLINRLPSPLLDNKTPFE 708
Query: 730 VMTSFFPSVPIKSGLQSRVFGCSAFVHVHGHHRGKLDPRAIKCIFIGYASDKKGYKCYHP 789
++ P + C + + H R K PRA C+F+GY S KGYK
Sbjct: 709 LLLKKIPDYTLLKS-------CLCYASTNVHDRNKFSPRARPCVFLGYPSGYKGYKVLDL 761
Query: 790 PSRRVYISMDVTFQESESFYPNPQLQGGDTQEAESPELSVIPLLQEPLMPSSIPITEDSD 849
S + I+ +V F E++ + + + + S++PL S+P+ +D
Sbjct: 762 ESHSISITRNVVFHETKFPFKTSKFL---KESVDMFPNSILPLPAPLHFVESMPLDDDLR 818
Query: 850 DDDNGPEQVPNQNDDNGLELVPDQ-NDDNGPELVPDQNNVDRFRIKYQRKVKPVLIQQQD 908
DDN + + + + +P N N L D N+V R K K P + +
Sbjct: 819 ADDNNASTSNSASSASSIPPLPSTVNTQNTDALDIDTNSVPIARPKRNAKA-PAYLSEYH 877
Query: 909 QPSDPEVSVPNPEPSNSYEHNLDDLPIALRKGKRSCAKYPISQYVSTQNLSVQHQSFISA 968
S P +S +P S S E +P K+ YP+S +S L+ S+I A
Sbjct: 878 CNSVPFLSSLSPTTSTSIETPSSSIP-----PKKITTPYPMSTAISYDKLTPLFHSYICA 932
Query: 969 IDSIRVPTSVQEVLKDKKWIQAMNEEMHALDKNGTRKIVEKPNDKRTVGCKWIYTVKYRS 1028
+ P + + +K +KW +A NEE+HAL++N T + K VGCKW++T+KY
Sbjct: 933 YNVETEPKAFTQAMKSEKWTRAANEELHALEQNKTWIVESLTEGKNVVGCKWVFTIKYNP 992
Query: 1029 DGTLDRYKARLVAKGYTQTYGIDYDETFAPVAKMNTVRIILSLAAHFDWELQQFDVKNAF 1088
DG+++RYKARLVA+G+TQ GIDY ETF+PVAK +V+++L LAA W L Q DV NAF
Sbjct: 993 DGSIERYKARLVAQGFTQQEGIDYMETFSPVAKFGSVKLLLGLAAATGWSLTQMDVSNAF 1052
Query: 1089 LHGDLEEEVYMEIPPGYDPTSG----RNKVCKLKKALYGLKQSPRAWFGRFTNAMVSLGY 1144
LHG+L+EE+YM +P GY P +G VC+L K+LYGLKQ+ R W+ R ++ + +
Sbjct: 1053 LHGELDEEIYMSLPQGYTPPTGISLPSKPVCRLLKSLYGLKQASRQWYKRLSSVFLGANF 1112
Query: 1145 KQSQGDHTLFIKHTRNGKLTLLLVYVNDMIIAGNDELEKQNLRKELAVQFEMKDLGKLKY 1204
QS D+T+F+K + + ++LVYV+D++IA ND +NL++ L +F++KDLG ++
Sbjct: 1113 IQSPADNTMFVKVSCT-SIIVVLVYVDDLMIASNDSSAVENLKELLRSEFKIKDLGPARF 1171
Query: 1205 FLGMEVAYSKQGIFISQRKYVLDLLKETGKLGCKPMGVPIEQNHKIGSSEEDLRVHKAQY 1264
FLG+E+A S +GI + QRKY +LL++ G GCKP +P++ N + L + Y
Sbjct: 1172 FLGLEIARSSEGISVCQRKYAQNLLEDVGLSGCKPSSIPMDPNLHLTKEMGTLLPNATSY 1231
Query: 1265 QRLVGKLIYLSHTRPDIAYPVSVVSQFMHDPQERHLQAVDRILQYLKASPGRGLLFKKSE 1324
+ LVG+L+YL TRPDI + V +SQF+ P + H+QA ++L+YLK +PG+
Sbjct: 1232 RELVGRLLYLCITRPDITFAVHTLSQFLSAPTDIHMQAAHKVLRYLKGNPGQ-------- 1283
Query: 1325 QLSMEVYTDADYAGSIVDRRSTTGYCMFLGGNLVTWRSKKQNVVARSSAEAEFRAMAQGV 1384
DAD+ RRS TG+C++LG +L+TW+SKKQ+VV+RSS E+E+R++AQ
Sbjct: 1284 --------DADWGTCKDSRRSVTGFCIYLGTSLITWKSKKQSVVSRSSTESEYRSLAQAT 1335
Query: 1385 CELLRMKIILDDLKIKYEAPMRLFCDNKSAISIAHNPVQHDRTKHIEIDRHFIKEKLNSS 1444
CE++ ++ +L DL + P +LFCDNKSA+ +A NPV H+RTKHIEID H +++++ +
Sbjct: 1336 CEIIWLQQLLKDLHVTMTCPAKLFCDNKSALHLATNPVFHERTKHIEIDCHTVRDQIKAG 1395
Query: 1445 LISTQYVPSGLQLADVLTKGLPLERFREL 1473
+ T +VP+G QLAD+LTK L F L
Sbjct: 1396 KLKTLHVPTGNQLADILTKPLHPGPFHSL 1424
>UniRef100_Q9FIC5 Retroelement pol polyprotein-like [Arabidopsis thaliana]
Length = 1462
Score = 817 bits (2110), Expect = 0.0
Identities = 504/1477 (34%), Positives = 781/1477 (52%), Gaps = 100/1477 (6%)
Query: 39 LDGRNYLQWSELVRHTLISRQKISYIEETAPA--ETDPQYDTWREENSLIMTWFWYSMIP 96
L G NY +W+ +R L +R+K + + T P ET+P +D W N+L+++W ++
Sbjct: 41 LRGPNYDEWATNLRLALKARKKFGFADGTIPQPDETNPDFDDWIANNALVVSWMKLTIHE 100
Query: 97 EISRNYMFFPTAKEIWNNLAQAYSKKQDLSACYELENKVFNSKQGTLSVTDYYGTLNGLW 156
++ + + ++W ++ + + K L+ ++ +Q + YYG L+ LW
Sbjct: 101 SLATSMSHLDDSHDMWTHIQKRFGVKNG-QRIQRLKTELATCRQKGTPIETYYGKLSQLW 159
Query: 157 IELDQYQNLKMKCDSDSTTLNKFIEKARIFKFLSGLN-SEFDPIRVQILGKEQFPSLSEV 215
L YQ K + + K E+ ++ +FL GL+ S + ++ +L + PSL E
Sbjct: 160 RSLADYQQAKTMEE-----VRKEREEDKLHQFLMGLDESMYGAVKSALLSRVPLPSLEEA 214
Query: 216 FS-IVRGEETRRTVMVEDKSVDGSALASGKGPIKG-STSFGRPNRDDRPNRDDRFNGDDR 273
++ + + EE++ + D+ D + + S S GR + F G
Sbjct: 215 YNTLTQDEESKSLSRLHDERNDEKMRRNATSSSRNQSASMGRGSSVVPATS---FKGKQS 271
Query: 274 FNRDDRCTYCKRSGHTKEYCFKLYGRENVLARMSKSKGAPQRRANHTASDMENDGEVPPV 333
F R + G E+ A S G+ A+
Sbjct: 272 FGRGASANHVANIG------------ESTTAATSSMSGSQLTEADRVG------------ 307
Query: 334 PPAEDVPSLSKAELERLRVFMDSLSKPSGSCSLTMTGKSSTFLSLNALSTENDWIIDFGA 393
+ L+ + ++LR + + S T T S FL WIID GA
Sbjct: 308 -----ISGLNDEQWKQLRQMLKERNFNS-----TNTKSSKFFLE--------SWIIDSGA 349
Query: 394 TDHMTPHPSHLSSYSTYPGKHYITVANGSQNQITGCGNIHLSPSLPLKRVLHVPKLSNNL 453
T+HMT L P I + +G T G ++L SL L+ V V L +L
Sbjct: 350 TNHMTGTLEFLRDVCDMP-PIMIKLPDGRLTTSTKHGRVYLGSSLDLQEVFFVDGLHCHL 408
Query: 454 LSIHKLTRDLNCAVTFFHSHGVFQDLATGQIIGIAKEQDGLYCLQHEESKCHQTSSESWN 513
+S+ +LTR +C + QD T +IG K+Q+GLY + E+ T +S
Sbjct: 409 ISVSQLTRAKSCVFQITDKVCIIQDRITLTLIGAGKQQNGLYFFRGTETVASMTRMDS-- 466
Query: 514 TSQIWLRHKRLGHPPFSILRTMFPHLFSKVSVESFHCDVCQFSKHHRSTFLPSNNKCAQP 573
+SQ+W H RLGHP +L+ + + + +S C++C +K R F SNNK + P
Sbjct: 467 SSQLW--HCRLGHPSSKVLKLLSFSDSTGHAFDSKTCEICIKAKQTRDPFPLSNNKTSSP 524
Query: 574 FDLVHSDVWGPSSISNVSGAKWFVTFIDDCTRITWVFLMKDKSEVFQLFANFFHMIRTQF 633
F++VH D+WGP +++ G+ +F+T +D+ TR W++L+ K NF ++ QF
Sbjct: 525 FEMVHCDLWGPYRTTSICGSNYFLTLVDNYTRAVWLYLLPSKQTAPMHLKNFISLVERQF 584
Query: 634 GKPIKRLRSDNGKE*VNKNFSEFLTKNGVVHELTCVDAPQQNGVAERKNRHLLEITRALL 693
IK +RSDNG E V S F +G++HE +CV PQQNG ERK+RH+L + RAL
Sbjct: 585 STKIKTIRSDNGTEFVC--LSSFFVDHGIIHETSCVGTPQQNGRVERKHRHILNVARALR 642
Query: 694 FQMNVPKFYWGEAVLTAAYLINRLPSRVLSNVSPVLVMTSFFPSVPIKSGLQSRVFGCSA 753
FQ +P +W LTAAYLINR P+ +L +P ++ + P V RVFGC
Sbjct: 643 FQARLPIEFWSYCALTAAYLINRTPTPLLQGKTPFELLYNRPPPVN-----HIRVFGCIC 697
Query: 754 FVHVHGHHRGKLDPRAIKCIFIGYASDKKGYKCYHPPSRRVYISMDVTFQESESFYP--- 810
+VH H K + R+ K IF+GY KKG++ Y+ + + +S DV F+E+E +P
Sbjct: 698 YVHNQKHGGDKFESRSNKSIFLGYPFAKKGWRVYNFETGVISVSRDVVFRETEFPFPASV 757
Query: 811 -----NPQLQGGDTQ-------EAESPE-LSVIPLLQEPLMPSSIPITEDSDDDDNGPEQ 857
+ QL + E ++P +S+ L+ SS + +D+ +
Sbjct: 758 FDSTPDSQLSPSNADQSFFLPSELQAPTPVSITTTLELTQSSSSTNLNDDNFRIPSDESS 817
Query: 858 VPNQNDDNGLELVPDQNDDNGPELVPDQNNVDRFRIKYQRKVKPVLIQQQDQPSDPEVSV 917
N+ DN +L +++ P L P ++ + P PS P+++
Sbjct: 818 SVNEMSDNE-DLNSPTTNESSPFLSPASPSLPLSPASLSLPLSPAA----PSPSLPKIAE 872
Query: 918 PNPEPS---------NSYEHNLDDLPIALRKGKRSCAKYPISQYVSTQNLSVQHQSFISA 968
P PEP D L + S YP+ YVS+ S +Q+++ A
Sbjct: 873 PEPEPELLGKGKRKKTQPVRLADYATTLLHQPHPSVTPYPLDNYVSSSQFSAAYQAYVFA 932
Query: 969 IDSIRVPTSVQEVLKDKKWIQAMNEEMHALDKNGTRKIVEKPNDKRTVGCKWIYTVKYRS 1028
I P S +E + D+ W A+++E+ +L+ GT + + P K+ +GCKW++ +KY+S
Sbjct: 933 ISLGIEPKSYKEAILDENWRCAVSDEIVSLENLGTWTVEDLPPGKKALGCKWVFRLKYKS 992
Query: 1029 DGTLDRYKARLVAKGYTQTYGIDYDETFAPVAKMNTVRIILSLAAHFDWELQQFDVKNAF 1088
DGTL+R+KARLV G QT GIDY ETFAPVAKM TVR L A DWE+ Q DV NAF
Sbjct: 993 DGTLERHKARLVVLGNKQTEGIDYSETFAPVAKMVTVRAFLQQVASLDWEVHQMDVHNAF 1052
Query: 1089 LHGDLEEEVYMEIPPGYDPTSGRNKVCKLKKALYGLKQSPRAWFGRFTNAMVSLGYKQSQ 1148
LHGDL+EEVY++ PPG+ R KVC+L+KALYGLKQ+PR WF + T A+ G+ Q
Sbjct: 1053 LHGDLDEEVYIKFPPGFGSDDNR-KVCRLRKALYGLKQAPRCWFAKLTTALNDYGFIQDI 1111
Query: 1149 GDHTLFIKHTRNGKLTLLLVYVNDMIIAGNDELEKQNLRKELAVQFEMKDLGKLKYFLGM 1208
D++LF RNG +LVYV+D+II G+ + L+ F MKDLG L+YFLG+
Sbjct: 1112 SDYSLFTME-RNGIRLHILVYVDDLIITGSSLDVITKFKGYLSSCFYMKDLGILRYFLGI 1170
Query: 1209 EVAYSKQGIFISQRKYVLDLLKETGKLGCKPMGVPIEQNHKIGSSEEDLRVHKAQYQRLV 1268
EVA S GI++ QRKY +D++ ETG LG P P+EQNHK+ + D ++Y+RLV
Sbjct: 1171 EVARSPAGIYLCQRKYAIDIITETGLLGVWPASHPLEQNHKLALAFGDTISDPSRYRRLV 1230
Query: 1269 GKLIYLSHTRPDIAYPVSVVSQFMHDPQERHLQAVDRILQYLKASPGRGLLFKKSEQLSM 1328
G+LIYL TRP+++Y + ++SQFM DP+ H++A R+++YLK+SPG+G+L + + L +
Sbjct: 1231 GRLIYLGTTRPELSYAIHMLSQFMSDPKADHMEAALRVVRYLKSSPGQGILLRSNTPLVL 1290
Query: 1329 EVYTDADYAGSIVDRRSTTGYCMFLGGNLVTWRSKKQNVVARSSAEAEFRAMAQGVCELL 1388
+ D+ + + +RS TG+ + LGG+ ++W++KK +VV+RSSAEAE+RAMA V ELL
Sbjct: 1291 TGWCDSGFDSCPITQRSLTGWFIQLGGSPISWKTKKHDVVSRSSAEAEYRAMADTVSELL 1350
Query: 1389 RMKIILDDLKIKYEAPMRLFCDNKSAISIAHNPVQHDRTKHIEIDRHFIKEKLNSSLIST 1448
++ +L L I P+ L+ D+ SAIS+A NPV H RTKH+ D HF+++++ I+T
Sbjct: 1351 WLRALLPALGISCNEPIMLYSDSLSAISLAANPVYHARTKHVGRDVHFVRDEIIRGTIAT 1410
Query: 1449 QYVPSGLQLADVLTKGLPLERFRELTCKLGMIDIHSP 1485
++V + QLAD++TK L F KLG+ ++H+P
Sbjct: 1411 KHVSTTSQLADIMTKALGRREFDAFLLKLGICNLHTP 1447
>UniRef100_Q84M55 Putative polyprotein [Oryza sativa]
Length = 1393
Score = 798 bits (2061), Expect = 0.0
Identities = 438/900 (48%), Positives = 583/900 (64%), Gaps = 75/900 (8%)
Query: 649 VNKNFSEFLTKNGVVHELTCVDAPQQNGVAERKNRHLLEITRALLFQMNVPKFYWGEAVL 708
V K S L G++H+ TC P QNGVAERKNRHLLE+ R+L+FQMNVPK+ W EAV+
Sbjct: 505 VEKEIS-LLHYEGIIHQTTCPGTPPQNGVAERKNRHLLEVARSLMFQMNVPKYLWSEAVM 563
Query: 709 TAAYLINRLPSRVLSNVSPV-LVMTSFFPSVPIKSGLQSRVFGCSAFVHVHGHHRGKLDP 767
TAAYLINR+PSR+L SP L++ VP K VFGC FV H GKLDP
Sbjct: 564 TAAYLINRMPSRILGMKSPAELLLGKREFKVPPK------VFGCVCFVRDHRPSVGKLDP 617
Query: 768 RAIKCIFIGYASDKKGYKCYHPPSRRVYISMDVTFQESESFYPNP--------------- 812
A+KC+F+GYAS +KGYKC+ P RR+++SMDVTF+E E +Y +
Sbjct: 618 HAVKCVFVGYASSQKGYKCWDPIGRRLFVSMDVTFREFEPYYKSKGDLDQFLEEFSTVME 677
Query: 813 -QLQGGDTQEAESPELSVIPLLQEPLMPSSIPITEDSDDDDNGPEQVPNQNDDNGLELVP 871
+ G+ + + E +V E ++ SIP S DD + +V D E+VP
Sbjct: 678 VDSREGERERGDIHEKNVGDKNGEAVVIGSIPC---SIDDASKVVEVIEDTQDEDREMVP 734
Query: 872 DQNDDNGPELVPD-----QNNVDRFRIK----YQR--------KVKPVLIQQQDQPSDPE 914
+ D E+V +R + K YQR K K ++ Q ++ P+
Sbjct: 735 HEEDGEEGEVVVGTIPCPMEGAERVKQKDVLVYQRRRFDSQGEKRKGLVQSQIEELPHPK 794
Query: 915 VSVPNPEPSNSYEHNLD---------------DLPIALRKGKRSCAKYP----------- 948
VP S S +L +LP+ R+ RS A P
Sbjct: 795 CPVPESSQSLSPPASLASLETIGNTSPTLEHVELPLVQRRETRSNAGRPPIRLGFEHLSF 854
Query: 949 ---ISQYVSTQNLSVQHQSFISAIDSIRVPTSVQEVLKDKKWIQAMNEEMHALDKNGTRK 1005
I+ Y++ ++S +++FI+++ ++ +P + +D KW AM EE++AL KN T +
Sbjct: 855 MHDIANYITYSHVSPAYKTFIASLQTMPIPKDWKCAKQDPKWKDAMKEELNALVKNKTWE 914
Query: 1006 IVEKPNDKRTVGCKWIYTVKYRSDGTLDRYKARLVAKGYTQTYGIDYDETFAPVAKMNTV 1065
+V+ P +KR VGCKW++TVK +G +DRYKARLVAKGY+QTYGIDYDETFAPVAKM TV
Sbjct: 915 LVKLPPEKRAVGCKWVFTVKQTPEGKVDRYKARLVAKGYSQTYGIDYDETFAPVAKMGTV 974
Query: 1066 RIILSLAAHFDWELQQFDVKNAFLHGDLEEEVYMEIPPGYDPTSGRNKVCKLKKALYGLK 1125
R ++S A +F W L Q DVKNAFLHGDL EEVYMEIPPG+ + KVCKLKK+LYGLK
Sbjct: 975 RALVSCAVNFGWPLHQLDVKNAFLHGDLHEEVYMEIPPGFGNSQTVGKVCKLKKSLYGLK 1034
Query: 1126 QSPRAWFGRFTNAMVSLGYKQSQGDHTLFIKHTRNGKLTLLLVYVNDMIIAGNDELEKQN 1185
QSPRAWF RF +A+ +GY Q GDHT+F KH R +T+L VYV+D++I G+D E +
Sbjct: 1035 QSPRAWFDRFRHAVCDMGYSQCNGDHTVFYKH-RGTHITILAVYVDDIVITGDDVEEIRC 1093
Query: 1186 LRKELAVQFEMKDLGKLKYFLGMEVAYSKQGIFISQRKYVLDLLKETGKLGCKPMGVPIE 1245
L++ L FE+KDLG L+YFLG+E+A S +GI +SQRKYVLDLL +TG LGC+ PI+
Sbjct: 1094 LKERLGKAFEVKDLGPLRYFLGIEIARSSKGIVLSQRKYVLDLLTDTGMLGCRASTTPID 1153
Query: 1246 QNHKIGSSEEDLRVHKAQYQRLVGKLIYLSHTRPDIAYPVSVVSQFMHDPQERHLQAVDR 1305
+NH++ + D V K YQRLVG+LIYL HTRPDI+Y VSVVS++MHDP+ HL V +
Sbjct: 1154 RNHQLCAQSGD-PVDKEAYQRLVGRLIYLCHTRPDISYAVSVVSRYMHDPRTGHLDVVHK 1212
Query: 1306 ILQYLKASPGRGLLFKKSEQLSMEVYTDADYAGSIVDRRSTTGYCMFLGGNLVTWRSKKQ 1365
IL+YLK +PG+GL F+K+ L++E Y DAD+A S+ DRRST+GYC+F+GGNLV+WRSKKQ
Sbjct: 1213 ILRYLKGTPGKGLWFRKNGHLNVEGYCDADWASSMDDRRSTSGYCVFVGGNLVSWRSKKQ 1272
Query: 1366 NVVARSSAEAEFRAMAQGVCELLRMKIILDDLKIKYEAPMRLFCDNKSAISIAHNPVQHD 1425
VVARS+AEAE+RAMA + E+L M+ +L +L++ + L CDNKSAISIA+NPVQHD
Sbjct: 1273 AVVARSTAEAEYRAMALSLSEMLWMRSLLTELRVLRSDTVMLHCDNKSAISIANNPVQHD 1332
Query: 1426 RTKHIEIDRHFIKEKLNSSLISTQYVPSGLQLADVLTKGLPLERFRELTCKLGMIDIHSP 1485
RTKH+EIDR FIKEK++S ++ +Y+ S QLAD LTKGL + + K+GMIDI P
Sbjct: 1333 RTKHVEIDRFFIKEKIDSGVLRLEYIKSCEQLADCLTKGLGPSEIQSICNKMGMIDIFCP 1392
Score = 147 bits (372), Expect = 2e-33
Identities = 122/486 (25%), Positives = 208/486 (42%), Gaps = 51/486 (10%)
Query: 30 LPVLQGSFRLDG-RNYLQWSELVRHTLISRQKISYIEETAPAETDPQ---YDTWREENSL 85
L ++ RL+G +NYL W + L ++ ++ E+ +D + + TW NS
Sbjct: 47 LELMPNELRLEGSKNYLSWCRRAQLMLRAKGVDHFLLESCEEPSDKESQAWRTWNTTNST 106
Query: 86 IMTWFWYSMIPEISRNYMFFPTAKEIWNNLAQAYSKKQDLSACYELENKVFNSKQGTLSV 145
+++W S+ P I R A +W L+ YS + ++ E +NKV N KQ +V
Sbjct: 107 VVSWLMTSVAPSIGRMIEAIQNAAVVWKTLSNMYSGEGNVMMMVEAQNKVENLKQEGRTV 166
Query: 146 TDYYGTLNGLWIELDQYQNLKMKCDSDSTTLNKFIEKARIFKFLSGLNSEFDPIRVQILG 205
+Y L LW +LD Y L++K + D NK++++ R+ FL GLN EF+ R +
Sbjct: 167 QEYASELQQLWADLDHYDPLQLKHEDDIVIGNKWLQRRRVIHFLKGLNKEFEDRRAAMFH 226
Query: 206 KEQFPSLSEVFSIVRGEETRRTVMVEDKSVDGSALASGKGPIKGSTSFGRPNRDDRPNRD 265
+ P++ E S + EE R +M + + +A+ G + + +R+
Sbjct: 227 QATLPTMEEAISAMVQEEMRLRLMRGTNPIRSAYIAADNRECYNCGQVGHVSYNCPTSRN 286
Query: 266 DRFNGDDRFN--------RDDRCTY-CKRSGHTKEYCFKLYGRENVLARMSKSKGAPQRR 316
G R R DR + R G + ++ GR + +G PQ
Sbjct: 287 IGGRGSIRGGHGGTRGGFRGDRGVFGGNRGGRGGDRGGRVGGR-------GRGRGVPQ-- 337
Query: 317 ANHTASDMENDGEVPPVPPAEDVPSLSKAELERLRVFMDSLSKPSGSCSLTMTGKSSTFL 376
A+ ++ DG+ + E V + + + ++ + + G+ +
Sbjct: 338 ----ANAVKEDGKAVTL-IGEQVTQWEEWQKNKTNESSNTTTHFGNFANYAQVGEGTQAQ 392
Query: 377 SL-NALSTENDWIIDFGATDHMTPHPSHLSSYSTYPGKHYITVANGSQNQITGCGNIHLS 435
+L + DWIID GA+ H+T + +SY+ Y I +A+G+ I
Sbjct: 393 ALASTYRHPIDWIIDSGASKHVTGLHNTFTSYTPYIHSETIQIADGTSKPI--------- 443
Query: 436 PSLPLKRVLHVPKLSNNLLSIHKLTRDLNCAVTFFHSHGVFQDLATGQIIGIAKEQDGLY 495
HV NLLSI L C V F + +FQ+ TG+ IG +DGL+
Sbjct: 444 ---------HV-----NLLSISSAIDQLKCIVVFDENSCLFQEKGTGRRIGTGVRRDGLW 489
Query: 496 CLQHEE 501
+ HEE
Sbjct: 490 YINHEE 495
>UniRef100_Q7X6S0 OSJNBb0011N17.2 protein [Oryza sativa]
Length = 1262
Score = 795 bits (2053), Expect = 0.0
Identities = 426/841 (50%), Positives = 566/841 (66%), Gaps = 55/841 (6%)
Query: 674 QNGVAERKNRHLLEITRALLFQMNVPKFYWGEAVLTAAYLINRLPSRVLSNVSPVLVMTS 733
+NGVAERKNRHLLEI R+L++ MNVPKF W EAV+TAAYLINR PSR+L +P ++
Sbjct: 447 ENGVAERKNRHLLEIARSLMYTMNVPKFLWSEAVMTAAYLINRTPSRILGMKTPYEMIFG 506
Query: 734 FFPSVPIKSGLQSRVFGCSAFVHVHGHHRGKLDPRAIKCIFIGYASDKKGYKCYHPPSRR 793
V + RVFGC+ FV H GKLDPRA+KCIFIGY+S +KGYKC+ P RR
Sbjct: 507 KNEFV-----VPPRVFGCTCFVRDHRPSIGKLDPRAVKCIFIGYSSSQKGYKCWSPSERR 561
Query: 794 VYISMDVTFQESESFYPNPQLQGGDTQEAESPELSVIPLLQEPLMPSSIPITEDSDDDDN 853
++SMDVTF+ES FY E ++S + + + L D D
Sbjct: 562 TFVSMDVTFRESVPFY------------GEKTDISSLFVDLDDLTRG------DHDQQKE 603
Query: 854 GPEQVPNQNDDNGLELVPDQNDDNGPELVPDQ-----NNVDRFRIKYQRKVKPVLIQQ-- 906
G +N+ + ++V + + V +Q + + ++ Y R+++ QQ
Sbjct: 604 GEILGLKENEQSKGKIVVGEIPCAIGDPVQEQEWRKPHEEENLQV-YTRRMRLPTTQQVE 662
Query: 907 -QDQPSDPEVSVPNPEPSNSYEHNL----DDLPIALRKGKRSCAKYP------------- 948
DQ SD V S + + +LPIA+RKG RS A P
Sbjct: 663 VDDQVSDDLTHVQVSSESGGEQIEIREEESNLPIAIRKGMRSNAGKPPQRYGFEIGDESG 722
Query: 949 ----ISQYVSTQNLSVQHQSFISAIDSIRVPTSVQEVLKDKKWIQAMNEEMHALDKNGTR 1004
I+ YVS +LS +++F+++++S +P +E +D +W QAM +E+ AL+KN T
Sbjct: 723 DENDIANYVSYTSLSSTYKAFVASLNSAIIPKDWKEAKQDPRWHQAMLDELEALEKNKTW 782
Query: 1005 KIVEKPNDKRTVGCKWIYTVKYRSDGTLDRYKARLVAKGYTQTYGIDYDETFAPVAKMNT 1064
+V PN K+ V CKW+Y VK DG ++RYKARLVAKGY+QTYGIDYDETFAPVAKM+T
Sbjct: 783 DLVSYPNGKKVVNCKWVYAVKQNPDGKVERYKARLVAKGYSQTYGIDYDETFAPVAKMST 842
Query: 1065 VRIILSLAAHFDWELQQFDVKNAFLHGDLEEEVYMEIPPGYDPTSGRNKVCKLKKALYGL 1124
VR I+S A +FDW L Q DVKNAFLHGDL+EEVYMEIPPG+ + KV +LKK+LYGL
Sbjct: 843 VRTIISCAVNFDWPLHQLDVKNAFLHGDLQEEVYMEIPPGFATLQTKGKVLRLKKSLYGL 902
Query: 1125 KQSPRAWFGRFTNAMVSLGYKQSQGDHTLFIKHTRNGKLTLLLVYVNDMIIAGNDELEKQ 1184
KQSPRAWF RF AM ++GYKQ GDHT+F H+ + +T+L VYV+DMII GND E
Sbjct: 903 KQSPRAWFDRFRRAMCAMGYKQCNGDHTVFYHHSGD-HITILAVYVDDMIITGNDCSEIT 961
Query: 1185 NLRKELAVQFEMKDLGKLKYFLGMEVAYSKQGIFISQRKYVLDLLKETGKLGCKPMGVPI 1244
L++ L+ +FE+KDLG+LKYFLG+E+A S +GI +SQRKY LDLL +TG LGC+P P+
Sbjct: 962 RLKQNLSKEFEVKDLGQLKYFLGIEIARSPRGIVLSQRKYALDLLSDTGMLGCRPASTPV 1021
Query: 1245 EQNHKIGSSEEDLRVHKAQYQRLVGKLIYLSHTRPDIAYPVSVVSQFMHDPQERHLQAVD 1304
+QNHK+ +E V+K +YQRLVG+LIYL HTRPDI Y VS+VS++MHDP+ H+ AV
Sbjct: 1022 DQNHKL-CAESGNPVNKERYQRLVGRLIYLCHTRPDITYAVSMVSRYMHDPRSGHMDAVY 1080
Query: 1305 RILQYLKASPGRGLLFKKSEQLSMEVYTDADYAGSIVDRRSTTGYCMFLGGNLVTWRSKK 1364
RIL+YLK SPG+GL FKK+ L +E Y DAD+A DRRST+GYC+F+GGNLV+WRSKK
Sbjct: 1081 RILRYLKGSPGKGLWFKKNGHLEVEGYCDADWASCPDDRRSTSGYCVFVGGNLVSWRSKK 1140
Query: 1365 QNVVARSSAEAEFRAMAQGVCELLRMKIILDDLKIKYEAPMRLFCDNKSAISIAHNPVQH 1424
Q VV+RS+AEAE+RAM+ + ELL ++ +L +L + + PM+L+CDNKSAISIA+NPVQH
Sbjct: 1141 QPVVSRSTAEAEYRAMSVSLSELLWLRNLLSELMLPVDTPMKLWCDNKSAISIANNPVQH 1200
Query: 1425 DRTKHIEIDRHFIKEKLNSSLISTQYVPSGLQLADVLTKGLPLERFRELTCKLGMIDIHS 1484
DRTKH+E+DR FIKEKL+ ++ ++V SG Q+AD TKGL ++ K+GMIDI+
Sbjct: 1201 DRTKHVELDRFFIKEKLDEGVLELEFVMSGGQVADCFTKGLGVKECNSSCDKMGMIDIYH 1260
Query: 1485 P 1485
P
Sbjct: 1261 P 1261
Score = 116 bits (291), Expect = 5e-24
Identities = 103/410 (25%), Positives = 165/410 (40%), Gaps = 60/410 (14%)
Query: 30 LPVLQGSFRLDGRNYLQWSELVRHTLISRQKISYI--EETAPAETDP-QYDTWREENSLI 86
+ ++Q +L +NYL WS L ++ Y+ E P T ++ TW NSL+
Sbjct: 42 IELMQNEIKLGVKNYLSWSRRALLILKTKGLEGYVTGEVKEPENTSSVEWKTWSTTNSLV 101
Query: 87 MTWFWYSMIPEISRNYMFFPTAKEIWNNLAQAYSKKQDLSACYELENKVFNSKQGTLSVT 146
+ W S+IP I+ +A E+W L + YS + ++ E + K+ +QG SV
Sbjct: 102 VAWLLTSLIPAIATTVETISSASEMWKTLTKLYSGEGNVMLMVEAQEKISALRQGERSVA 161
Query: 147 DYYGTLNGLWIELDQYQNLKMKCDSDSTTLNKFIEKARIFKFLSGLNSEFDPIRVQILGK 206
+Y L LW +LD Y L ++ + K++E+ R+ +FL GLN EF+ R + +
Sbjct: 162 EYVAELKSLWSDLDHYDPLGLEHSDCIAKMKKWVERRRVIEFLKGLNPEFEGRRDAMFHQ 221
Query: 207 EQFPSLSEVFSIVRGEETRRTVMVEDKSVDGS---ALASGKGPIK--GSTSFGRPNRDDR 261
P+L E + + EE ++ V+ S A+ GK + G RD
Sbjct: 222 TTLPTLDEAIAAMAQEELKKKVLPSAAPCSPSPTYAIVQGKETRECFNCGEMGHLMRDCH 281
Query: 262 PNRD---DRFNGDDRFNRDDRCTYCKRSGHTKEYCFKLYGRENVLARMSKSKG-APQRRA 317
R R G DR Y RS + Y ++ + N + S G P A
Sbjct: 282 APRKPTYGRGRGVDRGGTRGGRGYAGRSNRGRGYGYRGDYKANAVTLEEGSSGTTPDNVA 341
Query: 318 NHTASDMENDGEVPPVPPAEDVPSLSKAELERLRVFMDSLSKPSGSCSLTMTGKSSTFLS 377
N S SGS + F+S
Sbjct: 342 NFAHS-------------------------------------TSGSF-------NQAFMS 357
Query: 378 LNALSTENDWIIDFGATDHMTPHPSHLSSYSTYPGKHYITV--ANGSQNQ 425
+N ++ + WI+D GA+ H+T +SY Y H T+ A+G+ Q
Sbjct: 358 MN--TSHSSWILDSGASRHVTGMSGEFTSYKPYSFAHKETIQTADGTSCQ 405
>UniRef100_Q7XE85 Putative pol polyprotein [Oryza sativa]
Length = 1688
Score = 717 bits (1850), Expect = 0.0
Identities = 431/1120 (38%), Positives = 624/1120 (55%), Gaps = 70/1120 (6%)
Query: 387 WIIDFGATDHMTPHPSHLSSYSTYPGKHYITVANGSQNQITGCGNIHLSPSLPLKRVLHV 446
WI+D GA+ HM+ S L+S + ANG+ ++T G+I SP + V V
Sbjct: 184 WILDSGASFHMSFDDSWLTSCRLVKNGATVHTANGTLCKVTHQGSIS-SPQFTVPNVSLV 242
Query: 447 PKLSNNLLSIHKLTRDLNCAVTFFHSHGVFQDLATGQIIGIAKEQD---GLYCLQHEESK 503
PKLS NL+S+ +LT D NC V F + QD TG +IG Q GLY L
Sbjct: 243 PKLSMNLISVGQLT-DTNCFVGFDDTSCFVQDRHTGAVIGTGHRQKRSCGLYILDSLSLP 301
Query: 504 CHQTSSESWNTSQI---------WLRHKRLGHPPFSILRTMFPH-LFSKVSVES-FHCDV 552
T++ S + W H RLGH S L T+ + V V++ F C
Sbjct: 302 SSSTNTPSVYSPMCSTACKSFPQW--HHRLGHLCGSRLATLINQGVLGSVPVDTTFVCKG 359
Query: 553 CQFSKHHRSTFLPSNNKCAQPFDLVHSDVWGPSSISNVSGAKWFVTFIDDCTRITWVFLM 612
C+ K + + S ++ ++PFDLVHSDVWG S + G ++V F+DD +R TW++ M
Sbjct: 360 CKLGKQVQLPYPSSTSRSSRPFDLVHSDVWGKSPFPSKGGHNYYVIFVDDYSRYTWIYFM 419
Query: 613 KDKSEVFQLFANFFHMIRTQFGKPIKRLRSDNGKE*VNKNFSEFLTKNGVVHELTCVDAP 672
K +S++ ++ +F MI TQF I+ RSD+G E ++ F EFL G + +L+C A
Sbjct: 420 KHRSQLISIYQSFAQMIHTQFSSAIRIFRSDSGGEYMSNAFREFLVSQGTLPQLSCPGAH 479
Query: 673 QQNGVAERKNRHLLEITRALLFQMNVPKFYWGEAVLTAAYLINRLPSRVLSNVSPVLVMT 732
QNGVAERK+RH++E R LL VP +W EA+ TA YLIN PS L SP V+
Sbjct: 480 AQNGVAERKHRHIIETARTLLIASFVPAHFWAEAISTAVYLINMQPSSSLQGRSPGEVL- 538
Query: 733 SFFPSVPIKSGLQSRVFGCSAFVHVHGHHRGKLDPRAIKCIFIGYASDKKGYKCYHPPSR 792
F S P L RVFGC+ +V + R KL ++++C+F+GY+ + KGY+CY P +R
Sbjct: 539 --FGSPPRYDHL--RVFGCTCYVLLAPRERTKLTAQSVECVFLGYSLEHKGYRCYDPSAR 594
Query: 793 RVYISMDVTFQESESFYPNPQLQGGDTQEAESPELSVIPLLQEPL-MPSSIPITEDSDDD 851
R+ IS DVTF E++ F+ + T + SPE S+ L P+ P S+P + +
Sbjct: 595 RIRISRDVTFDENKPFFYS------STNQPSSPENSISFLYLPPIPSPESLPSSPITPSP 648
Query: 852 DNGPEQVPNQN-------DDNGLELVPDQND---DNGPELVPDQNNVDRFRIKYQRKVKP 901
P VP+ + + P + + P VP +D F Y R+ K
Sbjct: 649 SPIPPSVPSPTYVPPPPPSPSPSPVSPPPSHIPASSSPPHVPSTITLDTFPFHYSRRPKI 708
Query: 902 VLIQQQDQPS--DPEVSVPNPEPSNSYEHNLDDLPIALRKGKRSCAKYPISQYVSTQNLS 959
Q QP+ DP SV + P+ Y D ALR R
Sbjct: 709 PNESQPSQPTLEDPTCSVDDSSPAPRYNLRARD---ALRAPNRD---------------- 749
Query: 960 VQHQSFISAIDSIRVPTSVQEVLKDKKWIQAMNEEMHALDKNGTRKIVEKPNDKRTVGCK 1019
F+ + + P++ QE + W AM+EE+ AL++ T +V P+ + CK
Sbjct: 750 ----DFVVGV--VFEPSTYQEAIVLPHWKLAMSEELAALERTNTWDVVPLPSHAVPITCK 803
Query: 1020 WIYTVKYRSDGTLDRYKARLVAKGYTQTYGIDYDETFAPVAKMNTVRIILSLAAHFDWEL 1079
W+Y VK +SDG ++RYKARLVA+G+ Q +G DYDETFAPVA M TVR ++++AA W +
Sbjct: 804 WVYKVKTKSDGQVERYKARLVARGFQQAHGRDYDETFAPVAHMTTVRTLIAVAATRSWTI 863
Query: 1080 QQFDVKNAFLHGDLEEEVYMEIPPGYDPTSGRNKVCKLKKALYGLKQSPRAWFGRFTNAM 1139
Q DVKNAFLHGDL EEVYM PPG + G V +L++ALYGLKQ+PRAWF RF++ +
Sbjct: 864 SQMDVKNAFLHGDLHEEVYMHPPPGVEAPPGH--VFRLRRALYGLKQAPRAWFARFSSVV 921
Query: 1140 VSLGYKQSQGDHTLFIKHTRNGKLTLLLVYVNDMIIAGNDELEKQNLRKELAVQFEMKDL 1199
++ G+ S D LFI HT + TLLL+YV+DM+I G+D ++ +L+ QF M DL
Sbjct: 922 LAAGFSPSDHDPALFI-HTSSRGRTLLLLYVDDMLITGDDLEYIAFVKGKLSEQFMMSDL 980
Query: 1200 GKLKYFLGMEVAYSKQGIFISQRKYVLDLLKETGKLGCKPMGVPIEQNHKIGSSEEDLRV 1259
G L YFLG+EV + G ++SQ +Y+ DLL ++G + P+E + ++ S++
Sbjct: 981 GPLSYFLGIEVTSTVDGYYLSQHRYIEDLLAQSGLTDSRTTTTPMELHVRLRSTDGTPLD 1040
Query: 1260 HKAQYQRLVGKLIYLSHTRPDIAYPVSVVSQFMHDPQERHLQAVDRILQYLKASPGRGLL 1319
++Y+ LVG L+YL+ TRPDIAY V ++SQF+ P H + R+L+YL+ + + L
Sbjct: 1041 DPSRYRHLVGSLVYLTVTRPDIAYAVHILSQFVSAPISVHYGHLLRVLRYLRGTTTQCLF 1100
Query: 1320 FKKSEQLSMEVYTDADYAGSIVDRRSTTGYCMFLGGNLVTWRSKKQNVVARSSAEAEFRA 1379
+ S L + ++D+ +A +DRRS TGYC+FLG +L+TW+SKKQ V+RSS EAE RA
Sbjct: 1101 YAASSPLQLRAFSDSTWASDPIDRRSVTGYCIFLGTSLLTWKSKKQTAVSRSSTEAELRA 1160
Query: 1380 MAQGVCELLRMKIILDDLKIKYEAPMRLFCDNKSAISIAHNPVQHDRTKHIEIDRHFIKE 1439
+A E++ ++ +L D + + P L CDN AI IA++P++H+ TKHI +D F +
Sbjct: 1161 LATTTSEIVWLRWLLADFGVSCDVPTPLLCDNTGAIQIANDPIKHELTKHIGVDASFTRS 1220
Query: 1440 KLNSSLISTQYVPSGLQLADVLTKGLPLERFRELTCKLGM 1479
S I+ YVPS LQ+AD TK E R KL +
Sbjct: 1221 HCQQSTIALHYVPSELQVADFFTKAQTREHHRLHLLKLNV 1260
>UniRef100_Q75LJ1 Putative copia-like retrotransposon protein [Oryza sativa]
Length = 1399
Score = 695 bits (1794), Expect = 0.0
Identities = 416/1127 (36%), Positives = 629/1127 (54%), Gaps = 64/1127 (5%)
Query: 380 ALSTENDWIIDFGATDHMTPHPSHLSSYSTYPGKHYITVANGSQNQITGCGN--IHL-SP 436
A + +W +D GATDH+T L Y G I A+G+ +I G+ IH +
Sbjct: 281 AYGIDTNWYLDTGATDHITNELDKLDVREKYKGGDKIHTASGAGMEIKHIGDSVIHTPTR 340
Query: 437 SLPLKRVLHVPKLSNNLLSIHKLTRDLNCAVTFFHSHGVFQDLATGQIIGIAKEQDGLYC 496
L LK +LHVP+ NL+S H+L D + ++ + +D AT I + + LY
Sbjct: 341 ELHLKNILHVPQAKKNLISAHRLAMDNFAFLEVHSNYFLIKDRATRNTILKGRCRRRLYS 400
Query: 497 LQHEESK-CHQTSSESWNTSQIWLRHKRLGHPPFSILRTMFPHL---FSKVSVESFHCDV 552
L +K H ++ S++ W H RLGHP I+ + S V+ + CD
Sbjct: 401 LPTSPAKQVHAATTPSFSR---W--HSRLGHPAVPIVTRVLSKNNLPCSTVANKDSICDA 455
Query: 553 CQFSKHHRSTFLPSNNKCAQPFDLVHSDVWGPSSISNVSGAKWFVTFIDDCTRITWVFLM 612
CQ K H+ + S++ +QP +L+ S+VWGP+ IS V G K++V+FIDD ++ TWV+L+
Sbjct: 456 CQKGKSHQLPYPKSSSVSSQPLELIFSNVWGPAPIS-VGGKKFYVSFIDDYSKFTWVYLL 514
Query: 613 KDKSEVFQLFANFFHMIRTQFGKPIKRLRSDNGKE*VNKNFSEFLTKNGVVHELTCVDAP 672
K KSEVFQ F F ++ FG I +++D G E + + F K G+ H ++C A
Sbjct: 515 KHKSEVFQKFQEFQTLVERFFGHKILAVQTDWGGE--YQKLNTFFAKIGISHHVSCPYAH 572
Query: 673 QQNGVAERKNRHLLEITRALLFQMNVPKFYWGEAVLTAAYLINRLPSRVLSNVSPVLVMT 732
QQNG AERK+RHL+E+ LL ++P +W EAVL AAYLINR PS+V++ P +
Sbjct: 573 QQNGSAERKHRHLIEVALTLLAHASMPIKFWDEAVLAAAYLINRTPSKVINFACP---LE 629
Query: 733 SFFPSVPIKSGLQSRVFGCSAFVHVHGHHRGKLDPRAIKCIFIGYASDKKGYKCYHPPSR 792
F P + L R+FGC+ + ++ +++ KL R+ +C+F+GY++ KG+KC +
Sbjct: 630 QLFKEKPNYTAL--RIFGCAVWPNLRPYNKHKLAFRSKRCVFLGYSNLHKGFKCLEIATG 687
Query: 793 RVYISMDVTFQESESFYPNPQLQGGDTQEAESPELSVIPLLQEPLMPSSIPITEDSDDDD 852
RVY+S DVTF ES +P +L + E+S++P PS +P +
Sbjct: 688 RVYVSRDVTF--DESIFPFSELHSNAGARLRA-EISLLP-------PSLVPHLSSLGGEQ 737
Query: 853 NG---------PEQVPNQNDDNGLELVPDQNDDNGPELVPDQNNVDRFRIKYQRKVKPVL 903
N +Q +N + G E+V N + D+N Q V V
Sbjct: 738 NNHVLNYPPNVTDQFGEENAEIGEEIV--ANGEENAAAAADENAAAAANGGAQDDVHGVA 795
Query: 904 IQQQDQPSDPEVSVPNPEPSNSYEHNLDDLPIALRKGKRSCAKYP-------ISQYVSTQ 956
+ S P + + + + + + + + + P + V+T
Sbjct: 796 YDASPEHSSPVTDDAMASAAEQHGNPIQEEHLVQASPQTASSTSPSVASSAGVHDDVTTD 855
Query: 957 NLSVQHQSFISAI-----------DSIRVPTSVQEVLKDKKWIQAMNEEMHALDKNGTRK 1005
Q+ A IR S++E + +K W +AM+ E AL +N T
Sbjct: 856 QSDQTDQAMPEAAVAPIRPKTRLQSGIRKEKSLEEAVNNKHWKEAMDAEYMALIENKTWH 915
Query: 1006 IVEKPNDKRTVGCKWIYTVKYRSDGTLDRYKARLVAKGYTQTYGIDYDETFAPVAKMNTV 1065
+V + + CKW+Y VK ++DG+LDRYKARLVAKG+ Q YGIDY++TF+PV K T+
Sbjct: 916 LVPPQKGRNVIDCKWVYKVKRKADGSLDRYKARLVAKGFKQRYGIDYEDTFSPVVKAATI 975
Query: 1066 RIILSLAAHFDWELQQFDVKNAFLHGDLEEEVYMEIPPGYDPTSGRNKVCKLKKALYGLK 1125
RI+LSLA W L+Q DVKNAFLHG LEEEVYM+ PPGY+ S N VCKL KALYGLK
Sbjct: 976 RIVLSLAVSRGWSLRQLDVKNAFLHGVLEEEVYMKQPPGYEKKSMPNYVCKLDKALYGLK 1035
Query: 1126 QSPRAWFGRFTNAMVSLGYKQSQGDHTLFIKHTRNGKLTL-LLVYVNDMIIAGNDELEKQ 1184
Q+PRAW+ R + + LG+ S+ D +LF + G++++ LL+YV+D+I+A +
Sbjct: 1036 QAPRAWYSRLSTKLSELGFVPSKADTSLFF--YKKGQVSIFLLIYVDDIIVASSVPDATS 1093
Query: 1185 NLRKELAVQFEMKDLGKLKYFLGMEVAYSKQGIFISQRKYVLDLLKETGKLGCKPMGVPI 1244
L +EL+ F +KDLG L YFLG+EV K G+ +SQ KY DLL+ G CKP+ P+
Sbjct: 1094 TLLQELSKDFALKDLGDLHYFLGIEVHKVKDGLMLSQEKYASDLLRRVGMYECKPVSTPL 1153
Query: 1245 EQNHKIGSSEEDL--RVHKAQYQRLVGKLIYLSHTRPDIAYPVSVVSQFMHDPQERHLQA 1302
+ K+ +E L QY+ +VG L YL+ TRPDI++ ++ V QF+H P H A
Sbjct: 1154 STSEKLSVNEGTLLGPQDSTQYRSVVGALQYLTLTRPDISFSINKVCQFLHAPTTTHWAA 1213
Query: 1303 VDRILQYLKASPGRGLLFKKSEQLSMEVYTDADYAGSIVDRRSTTGYCMFLGGNLVTWRS 1362
V RIL+Y+K + GL F ++ L + ++DAD+AGS DRRST G+ +FLG NLV+W +
Sbjct: 1214 VKRILRYVKYTVDTGLKFCRNPSLLVSGFSDADWAGSPDDRRSTGGFAVFLGPNLVSWSA 1273
Query: 1363 KKQNVVARSSAEAEFRAMAQGVCELLRMKIILDDLKIKYEAPMRLFCDNKSAISIAHNPV 1422
+KQ V+RSS EAE++A+A E++ ++ +L +L ++ +L+CDN A ++ NP+
Sbjct: 1274 RKQATVSRSSIEAEYKALANATAEIMWVQTLLQELGVESPRAAKLWCDNLGAKYLSANPI 1333
Query: 1423 QHDRTKHIEIDRHFIKEKLNSSLISTQYVPSGLQLADVLTKGLPLER 1469
H RTKHIE+D HF++E++ L+ Y+ + Q+AD TK +P+ +
Sbjct: 1334 FHARTKHIEVDFHFVRERVARKLLEIAYISTKDQVADGFTKAIPVRQ 1380
Score = 45.4 bits (106), Expect = 0.013
Identities = 41/224 (18%), Positives = 94/224 (41%), Gaps = 30/224 (13%)
Query: 33 LQGSFRLDGRNYLQWSELVRHTLISRQKISYI-----------EETAPAET-----DPQY 76
+Q S +L NY WS V + + +I E+TA +T +P Y
Sbjct: 17 VQVSEKLTKGNYALWSAQVLAAIRGARLDGHITGATAAPSMEIEKTASDKTTEKIVNPAY 76
Query: 77 DTWREENSLIMTWFWYSMIPEISRNYMFFPTAKEIWNNLAQAYSKKQDLSACYELENKVF 136
W + ++ + ++ +I TA + W + ++ Q + + +
Sbjct: 77 QEWFASDQQVLGFLLSTLSRDILTQVATASTAAQAWQQVCAMFTA-QTKARSLNVRLTLT 135
Query: 137 NSKQGTLSVTDYYGTLNGLWIELDQYQNLKMKCDSDSTTLNKFIEKARIFKFLSGLNSEF 196
N+++G +S+++Y G + L D +++ E+ + L+GL+ +F
Sbjct: 136 NTQKGNMSISEYCGKMKAL-------------ADEIASSGKPLDEEDLVAYVLNGLDDDF 182
Query: 197 DPIRVQILGKEQFPSLSEVFSIVRGEETRRTVMVEDKSVDGSAL 240
+P+ I+ + + +++EV+S + E R+ + S + + +
Sbjct: 183 EPVVSAIVARNESTTMAEVYSQLLNFENRQALRQAHASANAAVV 226
>UniRef100_Q9FLA4 Polyprotein [Arabidopsis thaliana]
Length = 1429
Score = 692 bits (1786), Expect = 0.0
Identities = 470/1500 (31%), Positives = 717/1500 (47%), Gaps = 134/1500 (8%)
Query: 38 RLDGRNYLQWSELVRHTLISRQKISY----IEETAPAET-------DPQYDTWREENSLI 86
+L N+L W V L Y IEE T +P+Y W+ ++ LI
Sbjct: 6 KLTSTNFLMWRRQVHALLDGYDLAGYVDGSIEEPHTTVTVHGVTSPNPEYKLWKRQDKLI 65
Query: 87 MTWFWYSMIPEISRNYMFFPTAKEIWNNLAQAYSKKQDLSACYELENKVFNSKQGTLSVT 146
+ ++ + T+ +IW L Y+ ++ ++ K+G+ S+
Sbjct: 66 YSGLIGAISVAVQPLLSQATTSAQIWRKLVDTYANPSR-GHKQQIREQIKQWKKGSRSID 124
Query: 147 DYYGTLNGLWIELDQYQNLKMKCDSDSTTLNKFIEKARIFKFLSGLNSEFDPIRVQILGK 206
DY + GL DQ L+ + +I L GL+ ++ + QI G+
Sbjct: 125 DY---VLGLTTRFDQLALLEEAIPHED----------QIAYILGGLSDDYRRVIDQIEGR 171
Query: 207 EQFPSLSEVFSIVRGEETRRTVMVEDKSVDGSALASG-----KGPIKGSTSFGRPNRDDR 261
+ PS++E+ + E + MV D S +A A+ G S+S G N +
Sbjct: 172 DISPSITELHEKLINFELKLQAMVPDSSTPVTANAASYNNNNNGRNNRSSSRGNQNNQWQ 231
Query: 262 PNRDDRFNGDDRFNR----DDRCTYCKRSGHTKEYCFKLYGRENVLARMSKSKGAPQRRA 317
N+ + ++R ++ RC C GH+ C + S G P
Sbjct: 232 QNQTQQSRSNNRGSQGKGYQGRCQICGVHGHSARRCSQFQPYGGSGGSQSVPSGYPTNGY 291
Query: 318 NHTASDMENDGEVPPVPPAEDVPSLSKAELERLRVFMDSLSKPSGSCSLTMTGKSSTFLS 377
+ P+ P +A + F
Sbjct: 292 S----------------PSPMAPWQPRANIATAPPF------------------------ 311
Query: 378 LNALSTENDWIIDFGATDHMTPHPSHLSSYSTYPGKHYITVANGSQNQITGCGNIHL--- 434
N W++D GAT H+T ++LS + Y G +T+A+GS I+ G+ L
Sbjct: 312 -------NPWVLDSGATHHLTSDLANLSMHQPYTGGEEVTIADGSGLPISHTGSALLPTP 364
Query: 435 SPSLPLKRVLHVPKLSNNLLSIHKLTRDLNCAVTFFHSHGVFQDLATGQIIGIAKEQDGL 494
S SL LK +L+VP +S NL+S+++L +V FF +H +DL TG + + ++ L
Sbjct: 365 SRSLALKDILYVPNVSKNLISVYRLCNANQVSVEFFPAHFQVKDLNTGARLLQGRTRNEL 424
Query: 495 YCLQ-HEESKCHQTSSESWNTS-QIWLRHKRLGHPPFSILRTMFPHL---FSKVSVESFH 549
Y +++S T+S S T W H+RLGHP IL+ + H S +
Sbjct: 425 YEWPVNQKSITILTASPSPKTDLSSW--HQRLGHPALPILKDVVSHFHLPLSNTIPKQLP 482
Query: 550 CDVCQFSKHHRSTFLPSNNKCAQPFDLVHSDVWGPSSISNVSGAKWFVTFIDDCTRITWV 609
C C +K H+ F + +QP + +++DVW IS V K+++ +D TR TW+
Sbjct: 483 CSDCSINKSHKLPFFTNTIVSSQPLEYLYTDVWTSPCIS-VDNYKYYLVIVDHFTRYTWM 541
Query: 610 FLMKDKSEVFQLFANFFHMIRTQFGKPIKRLRSDNGKE*VNKNFSEFLTKNGVVHELTCV 669
+ +K KS+V +F F ++ +F I+ L SDNG E + FL +G+ H +
Sbjct: 542 YPLKQKSQVKDVFVAFKALVENRFQSRIRTLYSDNGGEFIG--LRPFLAAHGISHLTSPP 599
Query: 670 DAPQQNGVAERKNRHLLEITRALLFQMNVPKFYWGEAVLTAAYLINRLPSRVLSNVSPVL 729
P+ NG+AERK+RH++E ALL ++PK +W A TA YLINR+P+ VL SP +
Sbjct: 600 HTPEHNGLAERKHRHIVETGLALLTHASLPKTFWTYAFATAVYLINRMPTEVLQGTSPYV 659
Query: 730 VMTSFFPSVPIKSGLQSRVFGCSAFVHVHGHHRGKLDPRAIKCIFIGYASDKKGYKCYHP 789
+ P+ L+ RVFGC + + ++ KL+ R+ C+F+GY+ + Y C
Sbjct: 660 KLFQMSPNY-----LKLRVFGCLCYPWLRPYNTNKLEARSTMCVFLGYSLTQSAYLCLDI 714
Query: 790 PSRRVYISMDVTFQESESFYPNPQLQGGD-TQEAESPELS-VIPLLQEP---LMPSSIPI 844
+ R+Y S V F ES + +P+ D TQ P + VIPLLQ P P+++P+
Sbjct: 715 ATNRIYTSRHVQFVESSFPFASPRTSETDSTQTMSQPTTTNVIPLLQRPPHIAPPTALPL 774
Query: 845 TEDSDDDDNGP--------EQVPNQNDDNGLELVPDQN------------DDNGPELVPD 884
+ P E VP + + D N P P
Sbjct: 775 CPIFHSPPHSPSSPASPPSEHVPLTAASSSSNAINDDNISSTGQVSVSGPTSQSPHTTPT 834
Query: 885 QNNVDRFRIKYQRKVKPVLIQQQDQPSDPEVSVPNPEPSNSYEHNLDDLPIALRKGK--R 942
N K Q P+ P SV P+ S LP + R
Sbjct: 835 NQNTSPLS-KSPNPTNTNQSQNSTPPTSPTTSVHQHSPTPSPLPQNPPLPPPPQNDHPMR 893
Query: 943 SCAKYPISQYVSTQNLSVQHQSFISAIDSIRVPTSVQEVLKDKKWIQAMNEEMHALDKNG 1002
+ AK I++ + NL+ S +PT+V + LKD W AM+EE++A KN
Sbjct: 894 TRAKNQITKPKTKFNLTTSLTS-----SKPTIPTTVAQALKDPNWRNAMSEEINAQMKNH 948
Query: 1003 TRKIVEKPNDKRTVGCKWIYTVKYRSDGTLDRYKARLVAKGYTQTYGIDYDETFAPVAKM 1062
T +V K + CKWI+T+KY DG++ RYKARLVA+G+ Q YGIDY ETF+PV K
Sbjct: 949 TWDLVSPEEAKHVISCKWIFTLKYNVDGSIARYKARLVARGFNQQYGIDYSETFSPVIKS 1008
Query: 1063 NTVRIILSLAAHFDWELQQFDVKNAFLHGDLEEEVYMEIPPGYDPTSGRNKVCKLKKALY 1122
T+R +L +A +W + Q D+ NAFL G L EEVY+ PPG+ + VC+L KALY
Sbjct: 1009 TTIRTVLEVAVKRNWSIHQVDINNAFLQGTLNEEVYVSQPPGFIDRDRPSHVCRLNKALY 1068
Query: 1123 GLKQSPRAWFGRFTNAMVSLGYKQSQGDHTLFIKHTRNGKLTLLLVYVNDMIIAGNDELE 1182
GLKQ+PRAW+ ++ G+ S D +LFI + R+ +LVYV+D+IIAG + L
Sbjct: 1069 GLKQAPRAWYQELRRFLLQAGFVNSLADASLFI-YNRHNTFMYVLVYVDDIIIAGENALV 1127
Query: 1183 KQNLRKELAVQFEMKDLGKLKYFLGMEVAYSKQGIFISQRKYVLDLLKETGKLGCKPMGV 1242
Q LA +F +KDLG L YFLG+E + +G+ + QRKY+ DLLK+ L KP+
Sbjct: 1128 -QAFNASLASRFSLKDLGPLSYFLGIEATRTSRGLHLMQRKYITDLLKKHNMLDTKPVST 1186
Query: 1243 PIEQNHKIGSSEEDLRVHKAQYQRLVGKLIYLSHTRPDIAYPVSVVSQFMHDPQERHLQA 1302
P+ K+ +Y+ ++G L YL+ TRPDIA+ V+ +SQFMH P H QA
Sbjct: 1187 PMSPTPKLSLLSGTALDDATEYRTVLGSLQYLAFTRPDIAFAVNRLSQFMHRPTNEHWQA 1246
Query: 1303 VDRILQYLKASPGRGLLFKKSEQLSMEVYTDADYAGSIVDRRSTTGYCMFLGGNLVTWRS 1362
RIL+YL + G+ + L++ ++DAD+ + ST Y ++ GG+ V+W S
Sbjct: 1247 AKRILRYLAGTKSHGIFLRSDTPLTIHAFSDADWGCDLDAYLSTNAYIVYFGGSPVSWSS 1306
Query: 1363 KKQNVVARSSAEAEFRAMAQGVCELLRMKIILDDLKIKYEAPMRLFCDNKSAISIAHNPV 1422
KKQ VARSS EAE+RA+A EL + +L ++ I ++CDN A + NPV
Sbjct: 1307 KKQRSVARSSTEAEYRAVANTASELRWLCSLLLEMGISQTTVPVIYCDNIGATYLCANPV 1366
Query: 1423 QHDRTKHIEIDRHFIKEKLNSSLISTQYVPSGLQLADVLTKGLPLERFRELTCKLGMIDI 1482
H R KH+ +D HF++ + S + +V + QLAD LTK LP RF EL K+G+ ++
Sbjct: 1367 FHSRMKHVALDYHFVRGYIQSGALRVSHVSTKDQLADALTKPLPRPRFTELNSKIGVQEL 1426
>UniRef100_O23302 Retrovirus-related like polyprotein [Arabidopsis thaliana]
Length = 1489
Score = 683 bits (1763), Expect = 0.0
Identities = 397/1140 (34%), Positives = 608/1140 (52%), Gaps = 147/1140 (12%)
Query: 371 KSSTFLSLNALSTENDWIIDFGATDHMTPHPSHLSSYSTYPGKHYITVANGSQNQITGCG 430
++ T +L + WIID GA+ H+ S L+ + G
Sbjct: 466 QNHTLSALQKFLPSDAWIIDSGASSHVC---SDLAMFRELKSVS---------------G 507
Query: 431 NIHLSPSLPLKRVLHVPKLSNNLLSIHKLTRDLNCAVTFFHSHGVFQDLATGQIIGIAKE 490
+H++ L L VLHVP NL+S+ L + ++C+ F+ + Q+L+ G +IG +
Sbjct: 508 TVHITQKLILHNVLHVPDFKFNLMSVSSLVKTISCSAHFYVDCCLIQELSQGLMIGRGRL 567
Query: 491 QDGLYCLQHEESK--------CHQTSSESWNTSQIWLRHKRLGHPPFSILRTMFPHLFSK 542
LY L+ E + C T S N +W H+RLGHP +L+
Sbjct: 568 YHNLYILETENTSPSTSTPAACLFTGSVL-NDGHLW--HQRLGHPSSVVLQ--------- 615
Query: 543 VSVESFHCDVCQFSKHHRSTFLPSNNKCAQPFDLVHSDVWGPSSISNVSGAKWFVTFIDD 602
K R ++ NN + PFDLVH D+WGP SI ++ G ++F+T +DD
Sbjct: 616 --------------KLKRLAYISHNNLASNPFDLVHLDIWGPFSIESIEGFRYFLTVVDD 661
Query: 603 CTRITWVFLMKDKSEVFQLFANFFHMIRTQFGKPIKRLRSDNGKE*VNKNFSEFLTKNGV 662
CTR TWV+++++K +V +F F ++ TQF IK +RSDN E F+E + ++G+
Sbjct: 662 CTRTTWVYMLRNKKDVSSVFPEFIKLVSTQFNAKIKAIRSDNAPE---LGFTEIVKEHGM 718
Query: 663 VHELTCVDAPQQNGVAERKNRHLLEITRALLFQMNVPKFYWGEAVLTAAYLINRLPSRVL 722
+H +C PQQN V ERK++H+L + RALLFQ N+P YW + V TA +LINRLPS +L
Sbjct: 719 LHHFSCAYTPQQNSVVERKHQHILNVARALLFQSNIPMQYWSDCVTTAVFLINRLPSPLL 778
Query: 723 SNVSPVLVMTSFFPSVPIKSGLQSRVFGCSAFVHVHGHHRGKLDPRAIKCIFIGYASDKK 782
+N SP ++ + P + FGC FV + H R K PRA C+F+GY S K
Sbjct: 779 NNKSPYELILNKQPDYSLLKN-----FGCLCFVSTNAHERTKFTPRARACVFLGYPSGYK 833
Query: 783 GYKCYHPPSRRVYISMDVTFQE-------------SESFYPNPQLQGGDTQEAESPELSV 829
GYK S V +S +V F+E + +PN L A +
Sbjct: 834 GYKVLDLESHSVTVSRNVVFKEHVFPFKTSELLNKAVDMFPNSILP----LPAPLHFVET 889
Query: 830 IPLLQEPLMPSSIPITEDSDDDDNGPEQVPNQNDDNGLELVPDQNDDNGPELVPDQNNVD 889
+PL+ E S IP T DS DN + ++P ++ ++ D N V
Sbjct: 890 MPLIDED---SLIPTTTDSRTADNHASSSSSALPS----IIPPSSNTETQDI--DSNAVP 940
Query: 890 RFRIKYQRKVKPVLIQQQDQPSDPEVSVPNPEPSNSYEHNLDDLPIALRKGKRSCAKYPI 949
R K + P + + P +S P S+ H L ++ A + YPI
Sbjct: 941 ITRSKRTTRA-PSYLSEYHCSLVPSISTLPPTDSSIPIHPLPEIFTA--SSPKKTTPYPI 997
Query: 950 SQYVSTQNLSVQHQSFISAIDSIRVPTSVQEVLKDKKWIQAMNEEMHALDKNGTRKIVEK 1009
S VS + QS+I A ++ P + + +K +KWI+ EE+ A++ N T +
Sbjct: 998 STVVSYDKYTPLCQSYIFAYNTETEPKTFSQAMKSEKWIRVAVEELQAMELNKTWSVESL 1057
Query: 1010 PNDKRTVGCKWIYTVKYRSDGTLDRYKARLVAKGYTQTYGIDYDETFAPVAKMNTVRIIL 1069
P DK VGCKW++T+KY DGT++RYKARLVA+G+TQ GID+ +TF+PVAK+ + +++L
Sbjct: 1058 PPDKNVVGCKWVFTIKYNPDGTVERYKARLVAQGFTQQEGIDFLDTFSPVAKLTSAKMML 1117
Query: 1070 SLAAHFDWELQQFDVKNAFLHGDLEEEVYMEIPPGYDPTSGR----NKVCKLKKALYGLK 1125
LAA W L Q DV +AFLHGDL+EE++M +P GY P +G N VC+L K++YGLK
Sbjct: 1118 GLAAITGWTLTQMDVSDAFLHGDLDEEIFMSLPQGYTPPAGTILPPNPVCRLLKSIYGLK 1177
Query: 1126 QSPRAWFGRFTNAMVSLGYKQSQGDHTLFIKHTRNGKLTLLLVYVNDMIIAGNDELEKQN 1185
Q+ R W+ RF A LVY++D++IA N++ E +N
Sbjct: 1178 QASRQWYKRFVAA----------------------------LVYIDDIMIASNNDAEVEN 1209
Query: 1186 LRKELAVQFEMKDLGKLKYFLGMEVAYSKQGIFISQRKYVLDLLKETGKLGCKPMGVPIE 1245
L+ L +F++KDLG ++FLG+ LGCKP +P++
Sbjct: 1210 LKALLRSEFKIKDLGPARFFLGL--------------------------LGCKPSSIPMD 1243
Query: 1246 QNHKIGSSEEDLRVHKAQYQRLVGKLIYLSHTRPDIAYPVSVVSQFMHDPQERHLQAVDR 1305
+ + Y++L+G+L+YL+ TRPDI Y V +SQF+ P + HLQA +
Sbjct: 1244 PTLHLVRDMGTPLPNPTAYRKLIGRLLYLTITRPDITYAVHQLSQFISAPSDIHLQAAHK 1303
Query: 1306 ILQYLKASPGRGLLFKKSEQLSMEVYTDADYAGSIVDRRSTTGYCMFLGGNLVTWRSKKQ 1365
+L+Y+KA+PG+GL++ ++ + ++DAD+A RRS +G+C++LG +L++W+SKKQ
Sbjct: 1304 VLRYIKANPGQGLMYSADYEICLNGFSDADWAACKDTRRSISGFCIYLGTSLISWKSKKQ 1363
Query: 1366 NVVARSSAEAEFRAMAQGVCELLRMKIILDDLKIKYEAPMRLFCDNKSAISIAHNPVQHD 1425
V +RSS E+E+R+MAQ CE++ ++ +L DL I P +LFCDNKSA+ + NPV H+
Sbjct: 1364 AVASRSSTESEYRSMAQATCEIIWLQQLLKDLHIPLTCPAKLFCDNKSALHSSLNPVFHE 1423
Query: 1426 RTKHIEIDRHFIKEKLNSSLISTQYVPSGLQLADVLTKGLPLERFRELTCKLGMIDIHSP 1485
RTKHIEID H +++++ + + +VP+ Q AD+LTK L F L ++ + + P
Sbjct: 1424 RTKHIEIDCHTVRDQIKAGNLKALHVPTENQHADILTKALHPGPFHHLLRQMSLSSLFLP 1483
Score = 107 bits (268), Expect = 2e-21
Identities = 69/263 (26%), Positives = 124/263 (46%), Gaps = 21/263 (7%)
Query: 43 NYLQWSELVRHTLISRQKISYIEETA--PAETDPQYDTWREENSLIMTWFWYSMIPEISR 100
++ W + L R K+ +I T P E + W N ++ TW S+ +I +
Sbjct: 56 DFHSWRRSILMALNVRNKLGFINGTITKPPEDHRDFGAWSRCNDIVSTWLMNSVDKKIGQ 115
Query: 101 NYMFFPTAKEIWNNLAQAYSKKQDLSACYELENKVFNSKQGTLSVTDYYGTLNGLWIELD 160
+ ++ T + IWNNL + K+ D +++E K+ +QG++ ++ YY L LW E
Sbjct: 116 SLLYIATVQGIWNNLLSRF-KQDDAPRIFDIEQKLSKIEQGSMDISTYYTALLTLWEEHR 174
Query: 161 QYQNLKM----KCDSDSTTLNKFI-EKARIFKFLSGLNSEFDPIRVQILGKEQFPSLSEV 215
Y L + +C+ D+ + + +++R+ KFL LN FD R IL + P++ E
Sbjct: 175 NYVELPVCTCGRCECDAAVKWEHLQQRSRVTKFLKELNEGFDQTRRHILMLKPIPTIKEA 234
Query: 216 FSIVRGEETRRTVMVEDKSVDGSALASGKGPIKGSTSFGRPNRDDRPNRDDRFNGDDRFN 275
F++V +E +R V + VD A + + ++ RP N
Sbjct: 235 FNMVTQDERQRNVKPLTR-VDSVAFQNTSMINEDENAYVAAYNTVRP------------N 281
Query: 276 RDDRCTYCKRSGHTKEYCFKLYG 298
+ CT+C + GHT + C+K++G
Sbjct: 282 QKPICTHCGKVGHTIQKCYKVHG 304
>UniRef100_Q9FZK7 F17L21.7 [Arabidopsis thaliana]
Length = 1534
Score = 676 bits (1744), Expect = 0.0
Identities = 466/1513 (30%), Positives = 727/1513 (47%), Gaps = 181/1513 (11%)
Query: 38 RLDGRNYLQWSELVRHTLISRQKISYIEET--APAET---------DPQYDTWREENSLI 86
+L N+L W V+ L YI+ + P T +P + W+ ++ LI
Sbjct: 126 KLTSSNFLMWRRQVQALLNGYDLTGYIDGSIVVPPATITANGAVTVNPAFKHWQRQDQLI 185
Query: 87 MTWFW----YSMIPEISRNYMFFPTAKEIWNNLAQAYSKKQDLSACYELENKVFNSKQGT 142
+ S+ P +SR T+ EIW L Y+K S +L ++ K+ T
Sbjct: 186 YSALLGAISISVQPILSRT----TTSAEIWTKLMDTYAKPS-WSHIQQLRQQIKQWKKDT 240
Query: 143 LSVTDYYGTLNGLWIELDQYQNLKMKCDSDSTTLNKFIEKARIFKFLSGLNSEFDPIRVQ 202
S+ +++ GL + DQ L +S+ ++ + GL+ ++ + Q
Sbjct: 241 KSIDEFF---QGLVMRFDQLALLGKPMESEE----------QMEVIVEGLSDDYKQVIDQ 287
Query: 203 ILGKEQFPSLSEVFSIVRGEETRRTVMVEDKSVDGSALASGKGPIKGSTSFGRPNRDDRP 262
I G+E PSL+E+ + E + A+ PI + + RP +++
Sbjct: 288 IQGREVPPSLTEIHEKLLNHEVKLQA------------AASSLPISANAASYRPPANNKH 335
Query: 263 NRDDRFNGDDRFN-----------RDD---------RCTYCKRSGHTKEYCFKLYGRENV 302
N + + G +R N R+D +C C GH+ C +L
Sbjct: 336 NNSNNYRGQNRNNNNRGANSYQQPRNDQPSSRGYQGKCQICGVFGHSARRCSQL------ 389
Query: 303 LARMSKSKGAPQRRANHTASDMENDGEVPPVPPAEDVPSLSKAELERLRVFMDSLSKPSG 362
+MS + P P P VP +A +
Sbjct: 390 --QMSGAYSTPS----------------PSQYPNATVPWQPRANM--------------- 416
Query: 363 SCSLTMTGKSSTFLSLNALSTENDWIIDFGATDHMTPHPSHLSSYSTYPGKHYITVANGS 422
A + N W++D GAT H+T ++L+ + Y G +T+A+GS
Sbjct: 417 -----------------AAMSYNPWLLDSGATHHLTTDLNNLALHQPYNGGEEVTIADGS 459
Query: 423 QNQITGCGNIHLSP---SLPLKRVLHVPKLSNNLLSIHKLTRDLNCAVTFFHSHGVFQDL 479
IT G+ LS SL L +L+VP L NL+S++KL +V FF +H +DL
Sbjct: 460 TLPITHTGSSTLSTQSRSLALNNILYVPNLHKNLISVYKLCNANKVSVEFFPAHFQVKDL 519
Query: 480 ATGQIIGIAKEQDGLY--------CLQHEESKCHQTSSESWNTSQIWLRHKRLGHPPFSI 531
+TG + + +D LY + S +T+ SW H RLGHP +
Sbjct: 520 STGARLLQGRTKDELYEWPVPSNTPISLFASPTPKTTLPSW--------HSRLGHPSPPV 571
Query: 532 LRTMFPHLFSKVSVES---FHCDVCQFSKHHRSTFLPSNNKCAQPFDLVHSDVWGPSSIS 588
L+++ VS S F C C +K H+ F + P + V+SDVW S ++
Sbjct: 572 LKSLVSQFSLPVSNSSQKHFPCSHCLINKSHKLPFYSNTIISYTPLEYVYSDVW-TSPVT 630
Query: 589 NVSGAKWFVTFIDDCTRITWVFLMKDKSEVFQLFANFFHMIRTQFGKPIKRLRSDNGKE* 648
+V K+++ +D TR TW++ +K KS+V + F F ++ +F I+ L SDNG E
Sbjct: 631 SVDNFKYYLILVDHYTRYTWLYPLKQKSQVRETFVAFKALVENRFQTKIRTLYSDNGGEF 690
Query: 649 VNKNFSEFLTKNGVVHELTCVDAPQQNGVAERKNRHLLEITRALLFQMNVPKFYWGEAVL 708
+ +FL +G+ H + P+ NG+AERK+RH+LE LL Q ++P YW A
Sbjct: 691 IA--LRQFLLTHGISHLTSLPHTPEHNGIAERKHRHILETGLTLLTQASIPTSYWTYAFG 748
Query: 709 TAAYLINRLPSRVLSNVSPVLVMTSFFPSVPIKSGLQSRVFGCSAFVHVHGHHRGKLDPR 768
TA YLINRLPS VL+N SP + P+ L+ RVFGCS F + + KL+ R
Sbjct: 749 TAVYLINRLPSSVLNNESPYSKLFKTSPNY-----LKLRVFGCSCFPWLRPYTNHKLERR 803
Query: 769 AIKCIFIGYASDKKGYKCYHPPSRRVYISMDVTFQESESFYPNPQLQGGDTQEAESPELS 828
+ C+F+GY+ + Y C S RVY S V F E + + + + SPE
Sbjct: 804 SQPCVFLGYSLTQSAYLCLDRSSGRVYTSRHVQFVEDQFPF---SISDTHSVSNSSPE-E 859
Query: 829 VIPLLQEPLMPSSIPITEDSDDDDNGPEQVPNQNDDN----GLELVPDQNDDNGPELVPD 884
P +P PS IPI S P +P + D+ E + N +V
Sbjct: 860 ASPSCHQP--PSRIPIQSSSPPLVQAPSSLPPLSSDSHRRPNAETSSSSSSTNNDVVVSK 917
Query: 885 QNNVDRFRIKYQRKVKPVLIQQQDQPSDPEVSV-----PNPEPSNSYEHNLDDLPIALRK 939
N R + Q Q+ S+P S+ PNP PS + +++ + +
Sbjct: 918 DNTQVDNRNNFIGPTSSSSAQSQNN-SNPSSSIQTQNEPNPSPSPTPQNSSPESSPSSST 976
Query: 940 GKRSCAKYPIS--------QYVSTQNLSVQHQSFISAIDSI-----RVPTSVQEVLKDKK 986
S P +N + ++ +S + ++P +V + L+D+K
Sbjct: 977 SATSTVPNPPPPPPTNNHPMRTRAKNHITKPKTKLSLLAKTVQTRPQIPNTVNQALRDEK 1036
Query: 987 WIQAMNEEMHALDKNGTRKIVEKPNDKRTVGCKWIYTVKYRSDGTLDRYKARLVAKGYTQ 1046
W AM EE++A +N T ++V ++ + KWI+T+KY +GTLDRYKARLVA+G+ Q
Sbjct: 1037 WRNAMGEEINAQIRNNTFELVPPKPNQNVISTKWIFTLKYLPNGTLDRYKARLVARGFRQ 1096
Query: 1047 TYGIDYDETFAPVAKMNTVRIILSLAAHFDWELQQFDVKNAFLHGDLEEEVYMEIPPGYD 1106
YG+ Y ETF+PV K T+R++L LA W ++Q DV NAFL G L +EVY+ PPG+
Sbjct: 1097 QYGLHYSETFSPVVKSLTIRLVLQLAVSRSWTIKQLDVNNAFLQGTLTDEVYVTQPPGFI 1156
Query: 1107 PTSGRNKVCKLKKALYGLKQSPRAWFGRFTNAMVSLGYKQSQGDHTLFIKHTRNGKLTLL 1166
+ VC+LKKALYGLKQ+PRAW+ N + SLG+ S D ++F+ + + ++
Sbjct: 1157 DPDRPHHVCRLKKALYGLKQAPRAWYQELRNFVCSLGFTNSLADTSVFV-YINDIQIVYC 1215
Query: 1167 LVYVNDMIIAGNDELEKQNLRKELAVQFEMKDLGKLKYFLGMEVAYSKQGIFISQRKYVL 1226
LVYV+D+I+ G+ + L+ +F +KD L YFLG+E + QG+ + Q KYV
Sbjct: 1216 LVYVDDIIVTGSSDALVMAFITALSRRFSLKDPTDLVYFLGIEATRTSQGLHLMQHKYVY 1275
Query: 1227 DLLKETGKLGCKPMGVPIEQNHKIGSSEEDLRVHKAQYQRLVGKLIYLSHTRPDIAYPVS 1286
DLL L KP+ P+ + K+ +Y+ ++G L YL+ TRPDIAY V+
Sbjct: 1276 DLLSRMKMLDAKPVSTPMATHPKLSLYSGIALDEPGEYRTVIGSLQYLAFTRPDIAYAVN 1335
Query: 1287 VVSQFMHDPQERHLQAVDRILQYLKASPGRGLLFKKSEQLSMEVYTDADYAGSIVDRRST 1346
+SQFMH P + H QA R+L+YL + G+L + + LS+ ++DAD+AG D ST
Sbjct: 1336 RLSQFMHRPTDIHWQAAKRVLRYLAGTATHGILLRSNSPLSLHAFSDADWAGDNDDFVST 1395
Query: 1347 TGYCMFLGGNLVTWRSKKQNVVARSSAEAEFRAMAQGVCELLRMKIILDDLKIKYEAPMR 1406
Y ++LG + W SKKQ VARSS EAE+RA+A E+ + +L +L I
Sbjct: 1396 NAYIVYLGSTPIAWSSKKQKGVARSSTEAEYRAVANTTSEIRWVCSLLTELGITLPKMPV 1455
Query: 1407 LFCDNKSAISIAHNPVQHDRTKHIEIDRHFIKEKLNSSLISTQYVPSGLQLADVLTKGLP 1466
++CDN A ++ NPV H R KH+ +D HFI++ +++ + ++ + QLAD LTK LP
Sbjct: 1456 IYCDNVGATYLSANPVFHSRMKHLALDYHFIRDNVSAGALRVSHISTHDQLADALTKPLP 1515
Query: 1467 LERFRELTCKLGM 1479
+ F + + K+G+
Sbjct: 1516 RQHFLQFSSKIGV 1528
>UniRef100_Q94LA8 Polyprotein, putative [Arabidopsis thaliana]
Length = 1459
Score = 660 bits (1703), Expect = 0.0
Identities = 455/1512 (30%), Positives = 713/1512 (47%), Gaps = 160/1512 (10%)
Query: 38 RLDGRNYLQWSELVRHTLISRQKISYIEETA--PAET---------DPQYDTWREENSLI 86
+L NYL WS + L Y++ + P ET +P + W+ ++ LI
Sbjct: 32 KLTSTNYLMWSIQIHALLDGYDLAGYLDNSVVIPPETTTINSVVSANPSFTLWKRQDKLI 91
Query: 87 MTWFWYSMIPEISRNYMFFPTAKEIWNNLAQAYSKKQDLSACYELENKVFNSKQGTLSVT 146
+ ++ P + + +IW+ L Y+K +L ++ +GT ++
Sbjct: 92 FSALIGAISPAVQSLVSRATNSSQIWSTLNNTYAKPS-YGHIKQLRQQIQRLTKGTKTID 150
Query: 147 DYYGTLNGLWIELDQYQNLKMKCDSDSTTLNKFIEKARIFKFLSGLNSEFDPIRVQILGK 206
+Y + LDQ L + + ++ L GL E+ + QI GK
Sbjct: 151 EY---VQSHTTRLDQLAILGKPMEHEE----------QVEHILKGLPEEYKTVVDQIEGK 197
Query: 207 EQFPSLSEVFSIVRGEETRRTVMVEDKSVDGSALASGKGPIKGSTSFGRPNRDDRPNRDD 266
+ P+++E+ + E++ ++ D+ S+ ++ N++ NR
Sbjct: 198 DNTPTITEIHERLINHESK---LLSDEVPPSSSFPMSANAVQQRNFNNNCNQNQHKNRYQ 254
Query: 267 RFNGDDRFNRDDRCTYCKRSGHTKEYCFKLY-GRENVLARMSKS-KGAPQRRANHTASDM 324
++ N + + + +SG + FK Y G+ + + S + PQ +A +
Sbjct: 255 GNTHNNNTNTNSQPSTYNKSG---QRTFKPYLGKCQICSVQGHSARRCPQLQAMQLPASS 311
Query: 325 ENDGEVPPVPPAEDVPSLSKAELERLRVFMDSLSKPSGSCSLTMTGKSSTFLSLNALSTE 384
P P + L++ +
Sbjct: 312 SAHSPFTPWQPRAN-------------------------------------LAIGSPYAA 334
Query: 385 NDWIIDFGATDHMTPHPSHLSSYSTYPGKHYITVANGSQNQITGCGNIHL---SPSLPLK 441
N W++D GAT H+T + LS + Y G Y+ +A+G+ I G+ L + L L
Sbjct: 335 NPWLLDSGATHHITSDLNALSLHQPYNGGEYVMIADGTGLTIKQTGSTFLPSQNRDLALH 394
Query: 442 RVLHVPKLSNNLLSIHKLTRDLNCAVTFFHSHGVFQDLATGQIIGIAKEQDGLY--CLQH 499
+VL+VP + NL+S+++L +V FF + +DL TG ++ + +D LY + +
Sbjct: 395 KVLYVPDIRKNLISVYRLCNTNQVSVEFFPASFQVKDLNTGTLLLQGRTKDDLYEWPVTN 454
Query: 500 EESKCHQTSSESWNTSQIWLRHKRLGHPPFSILRTMFPHLFSKVSVESFH---CDVCQFS 556
+ TS T W H RLGHP SIL T+ VSV S + C C +
Sbjct: 455 PPATALFTSPSPKTTLSSW--HSRLGHPSASILNTLLSKFSLPVSVASSNKTSCSDCLIN 512
Query: 557 KHHRSTFLPSNNKCAQPFDLVHSDVWGPSSISNVSGAKWFVTFIDDCTRITWVFLMKDKS 616
K H+ F S+ + P + + +DVW IS+ K+++ +D TR TW++ ++ KS
Sbjct: 513 KSHKLPFATSSIHSSSPLEYIFTDVWTSPIISH-DNYKYYLVLVDHYTRYTWLYPLQQKS 571
Query: 617 EVFQLFANFFHMIRTQFGKPIKRLRSDNGKE*VNKNFSEFLTKNGVVHELTCVDAPQQNG 676
+V F F ++ +F I+ L SDNG E + +FL NG+ H + P+ NG
Sbjct: 572 QVKATFIAFKALVENRFQAKIRTLYSDNGGEFIA--LRDFLVSNGISHLTSPPHTPEHNG 629
Query: 677 VAERKNRHLLEITRALLFQMNVPKFYWGEAVLTAAYLINRLPSRVLSNVSPVLVMTSFFP 736
++ERK+RH++E LL Q +VP+ YW A TA YLINR+P+ VL SP F
Sbjct: 630 LSERKHRHIVETGLTLLTQASVPREYWTYAFATAVYLINRMPTPVLCLQSP---FQKLFG 686
Query: 737 SVPIKSGLQSRVFGCSAFVHVHGHHRGKLDPRAIKCIFIGYASDKKGYKCYHPPSRRVYI 796
S P L RVFGC F + + R KL+ R+ +C+F+GY+ + Y C + R+Y
Sbjct: 687 SSPNYQRL--RVFGCLCFPWLRPYTRNKLEERSKRCVFLGYSLTQTAYLCLDVDNNRLYT 744
Query: 797 SMDVTFQESESFYP-----NPQLQGG-----DTQEAESPELSVIPL----LQEPLMPSSI 842
S V F ES YP Q Q ++ + SP S P LQ P P+S
Sbjct: 745 SRHVMFDEST--YPFAASIREQSQSSLVTPPESSSSSSPANSGFPCSVLRLQSP--PASS 800
Query: 843 PITEDSDDDDNGPEQVPNQNDD---------NGLELVPDQNDDNGPELVPDQNNVDRFRI 893
P T N P Q L P + N P +N +
Sbjct: 801 PETPSPPQQQNDSPVSPRQTGSPTPSHHSQVRDSTLSPSPSVSNSEPTAPHENGPEP--- 857
Query: 894 KYQRKVKPVLIQQQDQPSDPEVS-VPNPEPSNSYEHNLDDLPIALRKGKRSCAKYPISQ- 951
+ Q P+ P + +PNP P + +++ P+ K + P +Q
Sbjct: 858 -----------EAQSNPNSPFIGPLPNPNPETNPSSSIEQRPV----DKSTTTALPPNQT 902
Query: 952 -YVSTQNLSVQ-----HQSFISAIDSI------------------RVPTSVQEVLKDKKW 987
+T N Q HQ + ++I P +V + LKDKKW
Sbjct: 903 TIAATSNSRSQPPKNNHQMKTRSKNNITKPKTKTSLTVALTQPHLSEPNTVTQALKDKKW 962
Query: 988 IQAMNEEMHALDKNGTRKIVEKPNDKRTVGCKWIYTVKYRSDGTLDRYKARLVAKGYTQT 1047
AM++E A +N T +V + VGC+W++ +KY +G +D+YKARLVAKG+ Q
Sbjct: 963 RFAMSDEFDAQQRNHTWDLVPPNPTQHLVGCRWVFKLKYLPNGLIDKYKARLVAKGFNQQ 1022
Query: 1048 YGIDYDETFAPVAKMNTVRIILSLAAHFDWELQQFDVKNAFLHGDLEEEVYMEIPPGYDP 1107
YG+DY ETF+PV K T+R++L +A +W L+Q DV NAFL G L EEVYM PPG+
Sbjct: 1023 YGVDYAETFSPVIKATTIRVVLDVAVKKNWPLKQLDVNNAFLQGTLTEEVYMAQPPGFVD 1082
Query: 1108 TSGRNKVCKLKKALYGLKQSPRAWFGRFTNAMVSLGYKQSQGDHTLFIKHTRNGKLTLLL 1167
+ VC+L+KA+YGLKQ+PRAW+ ++++G+ S D +LFI ++ L LL
Sbjct: 1083 KDRPSHVCRLRKAIYGLKQAPRAWYMELKQHLLNIGFVNSLADTSLFI-YSHGTTLLYLL 1141
Query: 1168 VYVNDMIIAGNDELEKQNLRKELAVQFEMKDLGKLKYFLGMEVAYSKQGIFISQRKYVLD 1227
VYV+D+I+ G+D + LA +F +KD L YFLG+E + G+ + QRKY+ D
Sbjct: 1142 VYVDDIIVTGSDHKSVSAVLSSLAERFSIKDPTDLHYFLGIEATRTNTGLHLMQRKYMTD 1201
Query: 1228 LLKETGKLGCKPMGVPIEQNHKIGSSEEDLRVHKAQYQRLVGKLIYLSHTRPDIAYPVSV 1287
LL + L KP+ P+ + K+ ++Y+ +VG L YL+ TRPDIA+ V+
Sbjct: 1202 LLAKHNMLDAKPVATPLPTSPKLTLHGGTKLNDASEYRSVVGSLQYLAFTRPDIAFAVNR 1261
Query: 1288 VSQFMHDPQERHLQAVDRILQYLKASPGRGLLFKKSEQLSMEVYTDADYAGSIVDRRSTT 1347
+SQFMH P H QA R+L+YL + G+ S + + ++DAD+AG D ST
Sbjct: 1262 LSQFMHQPTSDHWQAAKRVLRYLAGTTTHGIFLNSSSPIHLHAFSDADWAGDSADYVSTN 1321
Query: 1348 GYCMFLGGNLVTWRSKKQNVVARSSAEAEFRAMAQGVCELLRMKIILDDLKIKYEAPMRL 1407
Y ++LG N ++W SKKQ V+RSS E+E+RA+A E+ + +L +L I+ +
Sbjct: 1322 AYVIYLGRNPISWSSKKQRGVSRSSTESEYRAVANAASEIRWLCSLLTELHIRLPHGPTI 1381
Query: 1408 FCDNKSAISIAHNPVQHDRTKHIEIDRHFIKEKLNSSLISTQYVPSGLQLADVLTKGLPL 1467
FCDN A I NPV H R KHI +D HF++ + S + +V + QLAD LTK L
Sbjct: 1382 FCDNIGATYICANPVFHSRMKHIALDYHFVRGMIQSRALRVSHVSTNDQLADALTKSLSR 1441
Query: 1468 ERFRELTCKLGM 1479
F K+G+
Sbjct: 1442 PHFLSARSKIGV 1453
>UniRef100_Q94IU9 Copia-like polyprotein [Arabidopsis thaliana]
Length = 1466
Score = 660 bits (1703), Expect = 0.0
Identities = 455/1494 (30%), Positives = 716/1494 (47%), Gaps = 182/1494 (12%)
Query: 36 SFRLDGRNYLQWSELVRHTLISRQKISYIEE--TAPAET-------------DPQYDTWR 80
+ +L+ NYL W L S++ I ++ T PA+T +PQY+ W
Sbjct: 18 TLKLNDSNYLLWKTQFESLLSSQKLIGFVNGVVTPPAQTRLVVNDDVTSEVPNPQYEDWF 77
Query: 81 EENSLIMTWFWYSMIPEISRNYMFFPTAKEIWNNLAQAYSKKQDLSACYELEN--KVFNS 138
+ L+ +W + ++ E+ + T+++IW +LA+ ++ K ++ + L ++
Sbjct: 78 CTDQLVRSWLFGTLSEEVLGHVHNLTTSRQIWISLAENFN-KSSIAREFSLRRNLQLLTK 136
Query: 139 KQGTLSVTDYYGTLNGLWIELDQYQNLKMKCDSDSTTLNKFIEKARIFKFLSGLNSEFDP 198
K +LSV ++ K+ CDS S+ E +IF FL+GL E+DP
Sbjct: 137 KDKSLSV---------------YCRDFKIICDSLSSIGKPVEESMKIFGFLNGLGREYDP 181
Query: 199 IRVQI---LGKEQFPSLSEVFSIVRGEETR-----RTVMVEDKSVDGSALASGKGPIKGS 250
I I L K P+ ++V S V+G +++ TV V + ++ P S
Sbjct: 182 ITTVIQSSLSKLPAPTFNDVISEVQGFDSKLQSYDDTVSVNPHLAFNTERSNSGAPQYNS 241
Query: 251 TSFGRPNRDDRPNRDDRFNGDDRFNRDDR----------CTYCKRSGHTKEYCFKLYGRE 300
S GR R F++ C C R GHT C+ +
Sbjct: 242 NSRGRGRSGQNRGRGGYSTRGRGFSQHQSASPSSGQRPVCQICGRIGHTAIKCYNRF--- 298
Query: 301 NVLARMSKSKGAPQRRANHTASDMENDGEVPPVPPAEDVPSLSKAELERLRVFMDSLSKP 360
D EVP LRV ++ +
Sbjct: 299 ----------------------DNNYQSEVP------------TQAFSALRVSDETGKEW 324
Query: 361 SGSCSLTMTGKSSTFLSLNALSTENDWIIDFGATDHMTPHPSHLSSYSTYPGKHYITVAN 420
+ T +ST NA + E + + G ++ +H+ S + K I +
Sbjct: 325 YPDSAATAHITASTSGLQNATTYEGNDAVLVGDGTYLP--ITHVGSTTISSSKGTIPL-- 380
Query: 421 GSQNQITGCGNIHLSPSLPLKRVLHVPKLSNNLLSIHKLTRDLNCAVTFFHSHGVFQDLA 480
N++ C I K +L V KL + D C V F + DL
Sbjct: 381 ---NEVLVCPAIQ-------KSLLSVSKLCD----------DYPCGVYFDANKVCIIDLT 420
Query: 481 TGQIIGIAKEQDGLYCLQHEESKCHQTSSESWNTSQIWLRHKRLGHPPFSILRTMFPHLF 540
T +++ +GLY L++ E ++ + + + W H RLGH IL+ +
Sbjct: 421 TQKVVSKGPRNNGLYMLENSEFVALYSNRQCAASMETW--HHRLGHSNSKILQQLLTRKE 478
Query: 541 SKV--SVESFHCDVCQFSKHHRSTFLPSNNKCAQPFDLVHSDVWGPSSISNVSGAKWFVT 598
+V S S C+ CQ K R F S+ + +P D VH D+WGPS + + G K++
Sbjct: 479 IQVNKSRTSPVCEPCQMGKSTRLQFFSSDFRALKPLDRVHCDLWGPSPVVSNQGFKYYAV 538
Query: 599 FIDDCTRITWVFLMKDKSEVFQLFANFFHMIRTQFGKPIKRLRSDNGKE*VNKNFSEFLT 658
F+DD +R +W F ++ KS+ +F + ++ Q G IK +SD G E + E
Sbjct: 539 FVDDFSRFSWFFPLRMKSKFISVFIAYQKLVENQLGTKIKEFQSDGGGEFTSNKLKEHFR 598
Query: 659 KNGVVHELTCVDAPQQNGVAERKNRHLLEITRALLFQMNVPKFYWGEAVLTAAYLINRLP 718
++G+ H ++C PQQNGVAERK+RHL+E+ ++L+ + P +W EA TA YL N LP
Sbjct: 599 EHGIHHRISCPYTPQQNGVAERKHRHLVELGLSMLYHSHTPLKFWVEAFFTANYLSNLLP 658
Query: 719 SRVLSNVSPVLVMTSFFPSVPIKSGLQSRVFGCSAFVHVHGHHRGKLDPRAIKCIFIGYA 778
S VL +SP T F V RVFG + + + + K DPR+++C+F+GY
Sbjct: 659 SSVLKEISP--YETLFQQKVDY---TPLRVFGTACYPCLRPLAKNKFDPRSLQCVFLGYH 713
Query: 779 SDKKGYKCYHPPSRRVYISMDVTFQESE--------SFYPNPQ---LQGGDTQEAESPEL 827
+ KGY+C +PP+ +VYIS V F E++ S P Q LQ + P +
Sbjct: 714 NQYKGYRCLYPPTGKVYISRHVIFDEAQFPFKEKYHSLVPKYQTTLLQAWQHTDLTPPSV 773
Query: 828 SVIPLLQEPLMPSSIPI--TEDSDDDDNGPEQVPNQNDDNGLELVPDQNDDNGPELVPDQ 885
L +PL P+ +E+ + E+ N N + + + ND+
Sbjct: 774 PSSQL--QPLARQMTPMATSENQPMMNYETEEAVNVNMETSSDEETESNDE--------- 822
Query: 886 NNVDRFRIKYQRKVKPVLIQQQDQPSDPEVSVPNPEPSNSYEHNLDDLPIALRKGKRSCA 945
+ +V PVL Q + + + S+ N P + + K
Sbjct: 823 ---------FDHEVAPVLNDQNEDNALGQGSLENLHP-------------MITRSKDGIQ 860
Query: 946 KYPISQYVSTQNLSVQHQSFISAIDSIRVPTSVQEVLKDKKWIQAMNEEMHALDKNGTRK 1005
K P +Y + I + S P ++ +K W A+ +E+ + T
Sbjct: 861 K-PNPRY-----------ALIVSKSSFDEPKTITTAMKHPSWNAAVMDEIDRIHMLNTWS 908
Query: 1006 IVEKPNDKRTVGCKWIYTVKYRSDGTLDRYKARLVAKGYTQTYGIDYDETFAPVAKMNTV 1065
+V D + KW++ K + DGT+D+ KARLVAKG+ Q G+DY ETF+PV + T+
Sbjct: 909 LVPATEDMNILTSKWVFKTKLKPDGTIDKLKARLVAKGFDQEEGVDYLETFSPVVRTATI 968
Query: 1066 RIILSLAAHFDWELQQFDVKNAFLHGDLEEEVYMEIPPGYDPTSGRNKVCKLKKALYGLK 1125
R++L A +W L+Q DV NAFLHG+L+E V+M P G+ + N VC+L KALYGLK
Sbjct: 969 RLVLDTATANEWPLKQLDVSNAFLHGELQEPVFMFQPSGFVDPNKPNHVCRLTKALYGLK 1028
Query: 1126 QSPRAWFGRFTNAMVSLGYKQSQGDHTLFIKHTRNGKLTLLLVYVNDMIIAGNDELEKQN 1185
Q+PRAWF F+N ++ G++ S D +LF+ H +NG+ +LL+YV+D+++ G+D+L
Sbjct: 1029 QAPRAWFDTFSNFLLDFGFECSTSDPSLFVCH-QNGQSLILLLYVDDILLTGSDQLLMDK 1087
Query: 1186 LRKELAVQFEMKDLGKLKYFLGMEVAYSKQGIFISQRKYVLDLLKETGKLGCKPMGVPIE 1245
L + L +F MKDLG +YFLG+E+ G+F+ Q Y D+L + G C PM P+
Sbjct: 1088 LLQALNNRFSMKDLGPPRYFLGIEIESYNNGLFLHQHAYASDILHQAGMTECNPMPTPLP 1147
Query: 1246 QNHKIGSSEEDLRVHKAQYQRLVGKLIYLSHTRPDIAYPVSVVSQFMHDPQERHLQAVDR 1305
Q+ + +SE ++ L GKL YL+ TRPDI Y V+ + Q MH P + R
Sbjct: 1148 QHLEDLNSEP--FEEPTYFRSLAGKLQYLTITRPDIQYAVNFICQRMHAPTNSDFGLLKR 1205
Query: 1306 ILQYLKASPGRGLLFKKSEQLSMEVYTDADYAGSIVDRRSTTGYCMFLGGNLVTWRSKKQ 1365
IL+Y+K + GL +K + + D+DYAG RRSTTG+C+ LG L++W +K+Q
Sbjct: 1206 ILRYVKGTINMGLPIRKHHNPVLSGFCDSDYAGCKDTRRSTTGFCILLGSTLISWSAKRQ 1265
Query: 1366 NVVARSSAEAEFRAMAQGVCELLRMKIILDDLKIKYEAPMRLFCDNKSAISIAHNPVQHD 1425
++ SS EAE+RA++ E+ + +L DL I P R+FCDN SA+ ++ NP H
Sbjct: 1266 PTISHSSTEAEYRALSDTAREITWISSLLRDLGISQHQPTRVFCDNLSAVYLSANPALHK 1325
Query: 1426 RTKHIEIDRHFIKEKLNSSLISTQYVPSGLQLADVLTKGLPLERFRELTCKLGM 1479
R+KH + D H+I+E++ LI TQ++P+ +QLADV TK LP F L KLG+
Sbjct: 1326 RSKHFDKDFHYIRERVALGLIETQHIPATIQLADVFTKSLPRRPFITLRAKLGV 1379
>UniRef100_Q8LNV0 Putative copia-like polyprotein [Oryza sativa]
Length = 894
Score = 651 bits (1680), Expect = 0.0
Identities = 329/611 (53%), Positives = 439/611 (71%), Gaps = 23/611 (3%)
Query: 896 QRKVKPVLIQQQDQPSDP--EVSVPNPEPSNSYEHNLDD--LPIALRKGKRSCAKYP--- 948
+++ ++ + D+PSD + + + +E ++ LPIA+RKG RS A P
Sbjct: 285 EQQQSSIVAEVGDKPSDQCDTIQISSDTEGEEFETGGEETNLPIAIRKGVRSNAGKPPQR 344
Query: 949 --------------ISQYVSTQNLSVQHQSFISAIDSIRVPTSVQEVLKDKKWIQAMNEE 994
I+ YVS +L +++F+++++S+ +P +E +D +W QAM EE
Sbjct: 345 YGFEAQGVNDDENNIANYVSYASLLSTYKAFVTSLNSVEIPNDWREAKQDPRWHQAMLEE 404
Query: 995 MHALDKNGTRKIVEKPNDKRTVGCKWIYTVKYRSDGTLDRYKARLVAKGYTQTYGIDYDE 1054
+ AL+KN T +V P K+ V CKW+YTVK D ++RYKARLVAKGY+QTYGIDYDE
Sbjct: 405 LEALEKNKTWDLVPFPKGKKVVNCKWVYTVKQNPDENVERYKARLVAKGYSQTYGIDYDE 464
Query: 1055 TFAPVAKMNTVRIILSLAAHFDWELQQFDVKNAFLHGDLEEEVYMEIPPGYDPTSGRNKV 1114
TFAPVAKM+TVR ++S AA+FDW L Q DVKNAFLHGDL+EEVYMEIPPG+ + KV
Sbjct: 465 TFAPVAKMSTVRTLISCAANFDWPLHQLDVKNAFLHGDLQEEVYMEIPPGFATSQTEGKV 524
Query: 1115 CKLKKALYGLKQSPRAWFGRFTNAMVSLGYKQSQGDHTLFIKHTRNGKLTLLLVYVNDMI 1174
+LKK+LYGLKQSPRAWF RF AM +GYKQ GDHT+F +H R G T+L+VYV+DMI
Sbjct: 525 LRLKKSLYGLKQSPRAWFDRFRRAMCGMGYKQCNGDHTVFYRHNR-GLKTILVVYVDDMI 583
Query: 1175 IAGNDELEKQNLRKELAVQFEMKDLGKLKYFLGMEVAYSKQGIFISQRKYVLDLLKETGK 1234
I G+D LE L++ L+ +FE+KDLG+LKYFLG+E+A S +GI +SQRKYVLDLL +TG
Sbjct: 584 ITGDDCLEISRLKQNLSKEFEVKDLGQLKYFLGIEIARSPRGIVLSQRKYVLDLLSDTGM 643
Query: 1235 LGCKPMGVPIEQNHKIGSSEEDLRVHKAQYQRLVGKLIYLSHTRPDIAYPVSVVSQFMHD 1294
LGC+P IEQNHK+ + D V+K +YQRLVG+LIYL HTRPDI Y VSVVS++MHD
Sbjct: 644 LGCRPASTLIEQNHKLCAESGD-PVNKERYQRLVGRLIYLCHTRPDITYAVSVVSRYMHD 702
Query: 1295 PQERHLQAVDRILQYLKASPGRGLLFKKSEQLSMEVYTDADYAGSIVDRRSTTGYCMFLG 1354
P+ H+ V RIL+YLKASPG+G+ FKK+ L ME Y DAD+ + DRRST+GYC+F+G
Sbjct: 703 PRSGHMDVVYRILRYLKASPGKGIWFKKNGHLDMEGYCDADWGSCLDDRRSTSGYCVFIG 762
Query: 1355 GNLVTWRSKKQNVVARSSAEAEFRAMAQGVCELLRMKIILDDLKIKYEAPMRLFCDNKSA 1414
GNLV+WRSKK++VV+RS+AEAE+R+M+ + ELL +K +L +LK+ M+L+CDNKSA
Sbjct: 763 GNLVSWRSKKESVVSRSTAEAEYRSMSMSLSELLWLKNLLAELKLSTSTSMKLWCDNKSA 822
Query: 1415 ISIAHNPVQHDRTKHIEIDRHFIKEKLNSSLISTQYVPSGLQLADVLTKGLPLERFRELT 1474
I+IA+NPVQHDRTKH+EIDR FIKE+++ ++ +V SG Q+ D LTK L
Sbjct: 823 INIANNPVQHDRTKHVEIDRFFIKERMDEGTLNLGFVNSGEQVVDSLTKALGARECTSSC 882
Query: 1475 CKLGMIDIHSP 1485
K+GMIDI+ P
Sbjct: 883 SKMGMIDIYRP 893
Score = 171 bits (432), Expect = 2e-40
Identities = 86/180 (47%), Positives = 115/180 (63%), Gaps = 1/180 (0%)
Query: 535 MFPHLFSKVSVESFHCDVCQFSKHHRSTFLPSNNKCAQPFDLVHSDVWGPSSISNVSGAK 594
+FP SKV CD C++ KH R+T++ + PF LVHSDVW S I++VSGAK
Sbjct: 4 VFPDEMSKVDKSKLMCDACEYGKHTRTTYISRGLQSILPFVLVHSDVW-TSPIASVSGAK 62
Query: 595 WFVTFIDDCTRITWVFLMKDKSEVFQLFANFFHMIRTQFGKPIKRLRSDNGKE*VNKNFS 654
+FVTFID +R+TW++LM+ KSEV + F NF+ +R Q +K +RSDNG E VN F
Sbjct: 63 YFVTFIDCHSRMTWIYLMRHKSEVLKCFKNFYAWVRNQLNTCVKVIRSDNGGEYVNDEFR 122
Query: 655 EFLTKNGVVHELTCVDAPQQNGVAERKNRHLLEITRALLFQMNVPKFYWGEAVLTAAYLI 714
FL+ G+VH+ +C D P QNGVAERKNRH+LE+T +NV GE +L L+
Sbjct: 123 SFLSAEGIVHQTSCADTPPQNGVAERKNRHILEVTLLGKRDINVGVHLSGELLLVWMLLL 182
>UniRef100_Q9ZVW0 Putative retroelement pol polyprotein [Arabidopsis thaliana]
Length = 1413
Score = 625 bits (1611), Expect = e-177
Identities = 415/1250 (33%), Positives = 632/1250 (50%), Gaps = 93/1250 (7%)
Query: 9 PKEKLTSGVKGVSLPPLTHKDLP-VLQGSFRLDGRNYLQWSELVRHTLISRQKISYIEET 67
P+ V VS L D P + S L G NY +WS + + L +++K +I +
Sbjct: 13 PRTNQQPDVTKVSPYTLASSDNPGAMISSVMLTGDNYNEWSTKMLNALQAKRKTGFINGS 72
Query: 68 A--PAETDPQYDTWREENSLIMTWFWYSMIPEISRNYMFFPTAKEIWNNLAQAYSKKQDL 125
P +P Y+ W+ NS+I+ W S+ P++ F A ++W+ L Q +S +
Sbjct: 73 ISKPPLDNPDYENWQAVNSMIVGWIRASIEPKVKSTVTFICDAHQLWSELKQRFSVGNKV 132
Query: 126 SACYELENKVFNSKQGTLSVTDYYGTLNGLWIELDQYQNLKM-KCD--SDSTTL--NKFI 180
++++ ++ +Q V DYYG L LW E Y+ + + KC + TL +K
Sbjct: 133 HV-HQIKTQLAACRQDGQPVIDYYGRLCKLWEEFQIYKPITVCKCGLCTCGATLEPSKER 191
Query: 181 EKARIFKFLSGLN-SEFDPIRVQILGKEQFPSLSEVFS-IVRGEETRRTVMVEDKSVDGS 238
E+ +I +F+ GL+ S F + ++ + FPSL E++S +VR E+ +V + ++
Sbjct: 192 EEEKIHQFVLGLDDSRFGGLSATLIAMDPFPSLGEIYSRVVREEQRLASVQIREQQQSAI 251
Query: 239 ALASGKGPIKGSTSFGRPNRDDRPNRDDRFNGDDRFNRDDRCTYCKRSGHTKEYCFKLYG 298
+ + + T+ GR + +RD R C++C RSGH K+ C+++ G
Sbjct: 252 GFLTRQSEV---TADGRTDSSIIKSRD----------RSVLCSHCGRSGHEKKDCWQIVG 298
Query: 299 -------RENVLARMSKSKGAPQRRANHTASDMENDGEVPPVPPAEDVPSLSKAELERLR 351
R N R S S+G R + S ++ + ++LR
Sbjct: 299 FPDWWTERTNGGGRGSSSRGRGGRSSGSNNSGRGRGQVTAAHATTSNLSPFPEFTPDQLR 358
Query: 352 VFMDSLSKPSGSCSLTMTGKSSTFLSLNALSTENDWIIDFGATDHMTPHPSHLSSYSTYP 411
V + + S ++GK D I+D GA+ HMT S L++ T P
Sbjct: 359 VITQMIQNKNNGTSDKLSGKMKL----------GDVILDTGASHHMTGQLSLLTNIVTIP 408
Query: 412 GKHYITVANGSQNQITGCGNIHLSPSLPLKRVLHVPKLSNNLLSIHKLTRDLNCAVTFFH 471
+ A+G + G LS ++ L VL+VP L+ +L+S+ KL + + C F
Sbjct: 409 SCS-VGFADGRKTFAISMGTFKLSETVSLSNVLYVPALNCSLISVSKLVKQIKCLALFTD 467
Query: 472 SHGVFQDLATGQIIGIAKEQDGLYCLQHEESKCHQTSSESWNTSQIWLRHKRLGHPPFSI 531
+ V QD + +IG +E+DG+Y L + T + T+ L H+RLGHP FS+
Sbjct: 468 TICVLQDRFSRTLIGTGEERDGVYYLTDAATT---TVHKVDITTDHALWHQRLGHPSFSV 524
Query: 532 LRTMFPHLFSKVSVESFHCDVCQFSKHHRSTFLPSNNKCAQPFDLVHSDVWGPSSISNVS 591
L ++ S SV S CDVC +K R F S+NK F L+H DVWGP + +
Sbjct: 525 LSSLPLFSGSSCSVSSRSCDVCFRAKQTREVFPDSSNKSTDCFSLIHCDVWGPYRVPSSC 584
Query: 592 GAKWFVTFIDDCTRITWVFLMKDKSEVFQLFANFFHMIRTQFGKPIKRLRSDNGKE*VNK 651
GA +F+T +DD +R W +L+ KSEV + NF QFGK +K +RSDNG E +
Sbjct: 585 GAVYFLTIVDDFSRSVWTYLLLAKSEVRSVLTNFLAYTEKQFGKSVKIIRSDNGTEFMC- 643
Query: 652 NFSEFLTKNGVVHELTCVDAPQQNGVAERKNRHLLEITRALLFQMNVPKFYWGEAVLTAA 711
S + + G+VH+ +CV PQQNG ERK+RH+L ++RALLFQ ++P +WGEAV+TAA
Sbjct: 644 -LSSYFKEQGIVHQTSCVGTPQQNGRVERKHRHILNVSRALLFQASLPIKFWGEAVMTAA 702
Query: 712 YLINRLPSRVLSNVSPVLVMTSFFPSVPIKSGLQSRVFGCSAFVHVHGHHRGKLDPRAIK 771
YLINR PS + + +SP ++ P Q RVFG + + H + K R+
Sbjct: 703 YLINRTPSSIHNGLSPYELLHGCKPDYD-----QLRVFGSACYAHRVTRDKDKFGERSRL 757
Query: 772 CIFIGYASDKKGYKCYHPPSRRVYISMDVTFQESESFYPNPQLQGGDTQEAESPELSVIP 831
CIF+GY +KG+K Y + +S DV F+E+ Y + GDT +P ++
Sbjct: 758 CIFVGYPFGQKGWKVYDLSTNEFIVSRDVVFRENVFPYATNE---GDT--IYTPPVTCPI 812
Query: 832 LLQEPLMP--------------SSIPI----TEDSDDDDNGPEQVPNQNDDN---GLELV 870
E +P S P+ +SD + + P+ +P DD +
Sbjct: 813 TYDEDWLPFTTLEDRGSDENYLSDPPVCVTNVSESDTEHDTPQSLPTPVDDPLSPSTSVT 872
Query: 871 PDQNDDNGPELVPDQNNVDRFRIKYQRKVKPVLIQQQDQPSDPEVSVP---------NPE 921
P Q N NV Q+ P++ + +V P N
Sbjct: 873 PTQTPTNSSSSTSPSTNVS----PPQQDTTPIIENTPPRQGKRQVQQPARLKDYILYNAS 928
Query: 922 PSNSYEHNLDDLPIALRKGKRSCAKYPISQYVSTQNLSVQHQSFISAIDSIRVPTSVQEV 981
+ + H L + +YP++ Y+S + S H+ F++AI + P +E
Sbjct: 929 CTPNTPHVLSPSTSQSSSSIQGNLQYPLTDYISDECFSAGHKVFLAAITANDEPKHFKED 988
Query: 982 LKDKKWIQAMNEEMHALDKNGTRKIVEKPNDKRTVGCKWIYTVKYRSDGTLDRYKARLVA 1041
+K K W AM +E+ AL+ N T IV+ P K +G +W+Y K+ +DGT++RYKARLV
Sbjct: 989 VKVKVWNDAMYKEVDALEVNKTWDIVDLPTGKVAIGSQWVYKTKFNADGTVERYKARLVV 1048
Query: 1042 KGYTQTYGIDYDETFAPVAKMNTVRIILSLAAHFDWELQQFDVKNAFLHGDLEEEVYMEI 1101
+G Q G DY ETFAPV KM TVR +L L A WE+ Q DV NAFLHGDLEEEVYM++
Sbjct: 1049 QGNNQIEGEDYTETFAPVVKMTTVRTLLRLVAANQWEVYQMDVHNAFLHGDLEEEVYMKL 1108
Query: 1102 PPGYDPTSGRNKVCKLKKALYGLKQSPRAWFGRFTNAMVSLGYKQSQGDHTLFIKHTRNG 1161
PPG+ S +KVC+L+K+LYGLKQ+PR WF + ++A+ G+ Q D++ F ++ G
Sbjct: 1109 PPGF-RHSHPDKVCRLRKSLYGLKQAPRCWFKKLSDALKRFGFIQGYEDYS-FFSYSCKG 1166
Query: 1162 KLTLLLVYVNDMIIAGNDELEKQNLRKELAVQFEMKDLGKLKYFLGMEVA 1211
+LVYV+D+II GNDE Q ++ L F MKDLGKLKYFLG+EV+
Sbjct: 1167 IELRVLVYVDDLIICGNDEYMVQKFKEYLGRCFSMKDLGKLKYFLGIEVS 1216
Score = 181 bits (460), Expect = 1e-43
Identities = 90/198 (45%), Positives = 137/198 (68%)
Query: 1288 VSQFMHDPQERHLQAVDRILQYLKASPGRGLLFKKSEQLSMEVYTDADYAGSIVDRRSTT 1347
VS+ P+E HL+A RI++YLK SPG+G+L ++ L++EVY D+D+ + RRS +
Sbjct: 1215 VSRGPDAPREAHLEAAMRIVRYLKGSPGQGILLSANKDLTLEVYCDSDFQSCPLTRRSLS 1274
Query: 1348 GYCMFLGGNLVTWRSKKQNVVARSSAEAEFRAMAQGVCELLRMKIILDDLKIKYEAPMRL 1407
Y + LGG+ ++W++KKQ+ V+ SSAEAE+RAM+ + E+ + +L +L I AP RL
Sbjct: 1275 AYVVLLGGSPISWKTKKQDTVSHSSAEAEYRAMSVALKEIKWLNKLLKELGITLAAPTRL 1334
Query: 1408 FCDNKSAISIAHNPVQHDRTKHIEIDRHFIKEKLNSSLISTQYVPSGLQLADVLTKGLPL 1467
FCD+K+AISIA NPV H+RTKHIE D H +++ + +I+T +V + QLAD+ TK L
Sbjct: 1335 FCDSKAAISIAANPVFHERTKHIERDCHSVRDAVRDGIITTHHVRTSEQLADIFTKALGR 1394
Query: 1468 ERFRELTCKLGMIDIHSP 1485
+F L KLG+ ++H+P
Sbjct: 1395 NQFIYLMSKLGIQNLHTP 1412
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.319 0.135 0.405
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,596,755,563
Number of Sequences: 2790947
Number of extensions: 116215020
Number of successful extensions: 308354
Number of sequences better than 10.0: 2383
Number of HSP's better than 10.0 without gapping: 1535
Number of HSP's successfully gapped in prelim test: 856
Number of HSP's that attempted gapping in prelim test: 299348
Number of HSP's gapped (non-prelim): 5010
length of query: 1485
length of database: 848,049,833
effective HSP length: 140
effective length of query: 1345
effective length of database: 457,317,253
effective search space: 615091705285
effective search space used: 615091705285
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 82 (36.2 bits)
Lotus: description of TM0310b.12


