

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0304.15
(697 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
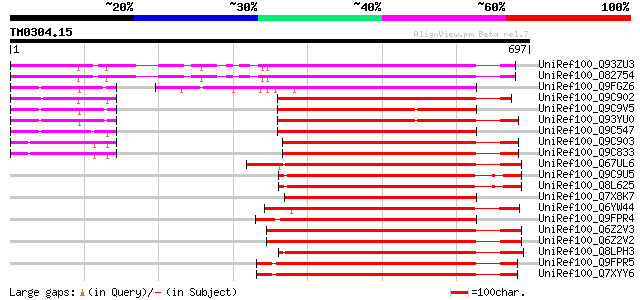
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q93ZU3 Putative serine/threonine kinase [Arabidopsis t... 516 e-145
UniRef100_O82754 Putative serine/threonine kinase [Arabidopsis t... 511 e-143
UniRef100_Q9FGZ6 Protein kinase [Arabidopsis thaliana] 361 4e-98
UniRef100_Q9C902 Protein kinase, putative; 19229-23534 [Arabidop... 358 2e-97
UniRef100_Q9C9V5 Hypothetical protein T23K23.26 [Arabidopsis tha... 356 1e-96
UniRef100_Q93YU0 Hypothetical protein At1g67890 [Arabidopsis tha... 356 1e-96
UniRef100_Q9C547 Protein kinase, putative; 12576-15979 [Arabidop... 348 4e-94
UniRef100_Q9C903 Protein kinase, putative; 8050-11829 [Arabidops... 342 2e-92
UniRef100_Q9C833 Protein kinase, putative; 42705-46677 [Arabidop... 342 2e-92
UniRef100_Q67UL6 Putative CTR1-like kinase kinase kinase [Oryza ... 330 7e-89
UniRef100_Q9C9U5 Hypothetical protein F25P22.8 [Arabidopsis thal... 330 9e-89
UniRef100_Q8L625 Hypothetical protein At1g73660 [Arabidopsis tha... 330 9e-89
UniRef100_Q7X8K7 CTR1-like kinase kinase kinase [Brassica juncea] 328 3e-88
UniRef100_Q6YW44 Putative MAP3K delta-1 protein kinase [Oryza sa... 328 3e-88
UniRef100_Q9FPR4 EDR1 [Hordeum vulgare] 324 6e-87
UniRef100_Q6Z2V3 Putative MAP kinase kinase kinase [Oryza sativa] 324 6e-87
UniRef100_Q6Z2V2 Putative MAP kinase kinase kinase [Oryza sativa] 324 6e-87
UniRef100_Q8LPH3 MAP kinase, putative [Arabidopsis thaliana] 323 8e-87
UniRef100_Q9FPR5 EDR1 [Oryza sativa] 321 4e-86
UniRef100_Q7XYY6 EDR1 [Oryza sativa] 321 4e-86
>UniRef100_Q93ZU3 Putative serine/threonine kinase [Arabidopsis thaliana]
Length = 735
Score = 516 bits (1329), Expect = e-145
Identities = 311/710 (43%), Positives = 414/710 (57%), Gaps = 132/710 (18%)
Query: 1 NHSAETLYGWKDYEIIGKRVTKFLITEEYYAPLQKILERLITTGETWSGKFPYKKKCGEV 60
+ SAE LY W E++G R L+TEEY L I R + GETW+G+FP++KK GE+
Sbjct: 117 SRSAENLYHWYAEEVVGYRTIDVLVTEEYRNSLTGIRNR-VCRGETWTGQFPFQKKTGEL 175
Query: 61 FMVMVTKAPLYEDGVLVGVISVSSEAAKFNT---IDSEFQRTCQSGASGQPGVQKLNSKG 117
FM +VTK+P+YE+G LVGV++VSS+A FN + +E Q+ +S + ++K
Sbjct: 176 FMALVTKSPVYENGELVGVVTVSSDATLFNRMHPLSNEHQQQARSNNRHESNLRK----- 230
Query: 118 NQWPL-RPLIAP------MPQIASSV-SNLASKILPNRHMDDTVSRNTSTDANDEKLEKC 169
+QW L RP IA +PQ +S+V SNLASK+LP R+ DD+ + N ++ + DE +
Sbjct: 231 HQWHLPRPQIAAASQVPVVPQYSSAVASNLASKLLPQRNGDDSFNGNHNSRSRDENVPVV 290
Query: 170 SIYETKLHSRHHHKEQNATIREASEKDESTTEFGRASKIAARFMAKLQIGGTGKWG-KDT 228
+ +T F + +A +F+ KLQ TG G +D
Sbjct: 291 A----------------------------STTFEKYGSLADKFLGKLQRKITGSQGTEDN 322
Query: 229 QSVKTNCRVDNPESDGVNNGNHSSGGSA------ALTSHQDVANGVDREENLHKCDSLFS 282
+ + N G+N SGGS+ T+ +D NG + + D
Sbjct: 323 EPILRN---------GINKSACGSGGSSKASNAVTCTAFRDNGNGKPKRAEVRISD---- 369
Query: 283 MKTAHTNVYARRTSGVFEVSSAKPFSRECYECLGSLVPQDPLLRLHRQFDPKKLEP---- 338
VY G+ + + ++ +G+L P P+ LE
Sbjct: 370 -------VYGNGAEGLIH-------NGDRFQYIGNLGQSKP---------PRGLESGLVS 406
Query: 339 --EATHMA-----IEDEVQKQQGRLQLPSSRESVGSHESSSSKGDNDSVVD--CEIHWED 389
T M+ IED + LP + G +S ++ +N V D CEI WED
Sbjct: 407 GMRGTKMSDLNGEIEDAWNTRLSVDPLPILGVNSGRQQSPVNQRNNRLVTDSSCEIRWED 466
Query: 390 LHLRDEIGQGSYAVVYHGIWNGSDVAIKVYFGNGYTEETLQDYKKEIDIMKRLRHPNVLL 449
L L +E+G+GS+A V+ G+WNGSDVAIKVYF Y TL + KKEI+IMK+LRHPNVLL
Sbjct: 467 LQLGEEVGRGSFAAVHRGVWNGSDVAIKVYFDGDYNAMTLTECKKEINIMKKLRHPNVLL 526
Query: 450 FMGAVYSLERLAIVTELLPRGSLFKTLHKSNQTLDIRRRLRMALDIARGMNYLHHRNPPI 509
FMGAV + E+ AI+ E +PRGSLFK LH +NQ LD +RRLRMALD+ARGMNYLH RNPPI
Sbjct: 527 FMGAVCTEEKSAIIMEYMPRGSLFKILHNTNQPLDKKRRLRMALDVARGMNYLHRRNPPI 586
Query: 510 VHRDLKSSNLLVDKNWTVKVGDFGLSRLKDTTLLTTKSGRGTPQWMAPEILRNEPSNEKS 569
VHRDLKSSNLLVDKNW VKVGDFGLS+ K+ T L+TKSG+GTPQWMAPE+LR+EPSNEK
Sbjct: 587 VHRDLKSSNLLVDKNWNVKVGDFGLSKWKNATFLSTKSGKGTPQWMAPEVLRSEPSNEKC 646
Query: 570 DVYSYGVVLWELMTQSIPWENLNSLQVVGVVGFMDRRLDLPEGLDPHVASIINDCWRRCF 629
DV+S+GV+LWELMT +PW+ LNS+QVVGVVGFMDRRLDLPEGL+P +ASII DCW+
Sbjct: 647 DVFSFGVILWELMTTLVPWDRLNSIQVVGVVGFMDRRLDLPEGLNPRIASIIQDCWQ--- 703
Query: 630 YMYSSLQLYYGRNHLLPQTESLLVLNMSSDPEQRPSFEELIQRMLFVLNR 679
+DP +RPSFEELI +M+ + +
Sbjct: 704 ----------------------------TDPAKRPSFEELISQMMSLFRK 725
>UniRef100_O82754 Putative serine/threonine kinase [Arabidopsis thaliana]
Length = 736
Score = 511 bits (1317), Expect = e-143
Identities = 311/711 (43%), Positives = 414/711 (57%), Gaps = 133/711 (18%)
Query: 1 NHSAETLYGWKDYEIIGKRVTKFLITEEYYAPLQKILERLITTGETWSGKFPYKKKCGEV 60
+ SAE LY W E++G R L+TEEY L I R + GETW+G+FP++KK GE+
Sbjct: 117 SRSAENLYHWYAEEVVGYRTIDVLVTEEYRNSLTGIRNR-VCRGETWTGQFPFQKKTGEL 175
Query: 61 FMVMVTKAPLYEDGVLVGVISVSSEAAKFNT---IDSEFQRTCQSGASGQPGVQKLNSKG 117
FM +VTK+P+YE+G LVGV++VSS+A FN + +E Q+ +S + ++K
Sbjct: 176 FMALVTKSPVYENGELVGVVTVSSDATLFNRMHPLSNEHQQQARSNNRHESNLRK----- 230
Query: 118 NQWPL-RPLIAP------MPQIASSV-SNL-ASKILPNRHMDDTVSRNTSTDANDEKLEK 168
+QW L RP IA +PQ +S+V SNL ASK+LP R+ DD+ + N ++ + DE +
Sbjct: 231 HQWHLPRPQIAAASQVPVVPQYSSAVASNLKASKLLPQRNGDDSFNGNHNSRSRDENVPV 290
Query: 169 CSIYETKLHSRHHHKEQNATIREASEKDESTTEFGRASKIAARFMAKLQIGGTGKWG-KD 227
+ +T F + +A +F+ KLQ TG G +D
Sbjct: 291 VA----------------------------STTFEKYGSLADKFLGKLQRKITGSQGTED 322
Query: 228 TQSVKTNCRVDNPESDGVNNGNHSSGGSA------ALTSHQDVANGVDREENLHKCDSLF 281
+ + N G+N SGGS+ T+ +D NG + + D
Sbjct: 323 NEPILRN---------GINKSACGSGGSSKASNAVTCTAFRDNGNGKPKRAEVRISD--- 370
Query: 282 SMKTAHTNVYARRTSGVFEVSSAKPFSRECYECLGSLVPQDPLLRLHRQFDPKKLEP--- 338
VY G+ + + ++ +G+L P P+ LE
Sbjct: 371 --------VYGNGAEGLIH-------NGDRFQYIGNLGQSKP---------PRGLESGLV 406
Query: 339 ---EATHMA-----IEDEVQKQQGRLQLPSSRESVGSHESSSSKGDNDSVVD--CEIHWE 388
T M+ IED + LP + G +S ++ +N V D CEI WE
Sbjct: 407 SGMRGTKMSDLNGEIEDAWNTRLSVDPLPILGVNSGRQQSPVNQRNNRLVTDSSCEIRWE 466
Query: 389 DLHLRDEIGQGSYAVVYHGIWNGSDVAIKVYFGNGYTEETLQDYKKEIDIMKRLRHPNVL 448
DL L +E+G+GS+A V+ G+WNGSDVAIKVYF Y TL + KKEI+IMK+LRHPNVL
Sbjct: 467 DLQLGEEVGRGSFAAVHRGVWNGSDVAIKVYFDGDYNAMTLTECKKEINIMKKLRHPNVL 526
Query: 449 LFMGAVYSLERLAIVTELLPRGSLFKTLHKSNQTLDIRRRLRMALDIARGMNYLHHRNPP 508
LFMGAV + E+ AI+ E +PRGSLFK LH +NQ LD +RRLRMALD+ARGMNYLH RNPP
Sbjct: 527 LFMGAVCTEEKSAIIMEYMPRGSLFKILHNTNQPLDKKRRLRMALDVARGMNYLHRRNPP 586
Query: 509 IVHRDLKSSNLLVDKNWTVKVGDFGLSRLKDTTLLTTKSGRGTPQWMAPEILRNEPSNEK 568
IVHRDLKSSNLLVDKNW VKVGDFGLS+ K+ T L+TKSG+GTPQWMAPE+LR+EPSNEK
Sbjct: 587 IVHRDLKSSNLLVDKNWNVKVGDFGLSKWKNATFLSTKSGKGTPQWMAPEVLRSEPSNEK 646
Query: 569 SDVYSYGVVLWELMTQSIPWENLNSLQVVGVVGFMDRRLDLPEGLDPHVASIINDCWRRC 628
DV+S+GV+LWELMT +PW+ LNS+QVVGVVGFMDRRLDLPEGL+P +ASII DCW+
Sbjct: 647 CDVFSFGVILWELMTTLVPWDRLNSIQVVGVVGFMDRRLDLPEGLNPRIASIIQDCWQ-- 704
Query: 629 FYMYSSLQLYYGRNHLLPQTESLLVLNMSSDPEQRPSFEELIQRMLFVLNR 679
+DP +RPSFEELI +M+ + +
Sbjct: 705 -----------------------------TDPAKRPSFEELISQMMSLFRK 726
>UniRef100_Q9FGZ6 Protein kinase [Arabidopsis thaliana]
Length = 730
Score = 361 bits (927), Expect = 4e-98
Identities = 219/502 (43%), Positives = 281/502 (55%), Gaps = 73/502 (14%)
Query: 197 ESTTEFGRA-SKIAARFMAKLQIGGTGKWGKDTQ--------------SVKTNCRVDNPE 241
E T F R S +A++ + K G D+Q + K +V +
Sbjct: 229 EGETSFSRVTSSVASKLGLDSKEAVVSKLGLDSQQPIQVAIASKISDLASKVGNKVKSKM 288
Query: 242 SDGVNNGNHSSGGSAALTSHQDVANGVDREENLHKCDSLFSMKTAHTNVYARRTSGVF-- 299
G NN + GGS SHQ D + D+ S + + GVF
Sbjct: 289 RAGDNNAANLEGGSG--DSHQSDQGFFDAAFADRREDAATSGADTPRGDFIQSPFGVFLR 346
Query: 300 --EVSSAKPFSRECYECLGSLVPQDPLLRLHRQFDPKK-------------LEPEATHMA 344
E +S KPF E G+ V L ++ KK LE +H
Sbjct: 347 SDEKASTKPFRDSSDESDGNSVVPKTLTSKAEEWMVKKGLSWPWKGNEREGLEGRRSHSV 406
Query: 345 ---IEDEVQKQQGR-------------------------LQLPSSRESVGSHESSSSKGD 376
+ +E QKQQ + SS +V S SSSS G
Sbjct: 407 WPWVRNEQQKQQAYQSNSNHSVKSESQACESIKASSNEPMGYWSSSVNVNSTSSSSSCGS 466
Query: 377 NDSVV-----------DCEIHWEDLHLRDEIGQGSYAVVYHGIWNGSDVAIKVYFGNGYT 425
S V D EI WEDL + ++IGQGS VYHG+W GSDVA+KV+ Y+
Sbjct: 467 TSSSVMNKVDMDSDCLDYEILWEDLTIGEQIGQGSCGTVYHGLWFGSDVAVKVFSKQEYS 526
Query: 426 EETLQDYKKEIDIMKRLRHPNVLLFMGAVYSLERLAIVTELLPRGSLFKTLHKSNQTLDI 485
EE + +++E+ +MKRLRHPNVLLFMGAV S +RL IVTE LPRGSLF+ L ++ LD
Sbjct: 527 EEIITSFRQEVSLMKRLRHPNVLLFMGAVTSPQRLCIVTEFLPRGSLFRLLQRNTSKLDW 586
Query: 486 RRRLRMALDIARGMNYLHHRNPPIVHRDLKSSNLLVDKNWTVKVGDFGLSRLKDTTLLTT 545
RRR+ MA DIARGMNYLHH PPI+HRDLKSSNLLVDKNWTVKV DFGLSR+K T LTT
Sbjct: 587 RRRIHMASDIARGMNYLHHCTPPIIHRDLKSSNLLVDKNWTVKVADFGLSRIKHETYLTT 646
Query: 546 KSGRGTPQWMAPEILRNEPSNEKSDVYSYGVVLWELMTQSIPWENLNSLQVVGVVGFMDR 605
K+GRGTPQWMAPE+LRNE ++EKSDVYS+GV+LWEL+T+ IPWE+LN++QV+G VGFM++
Sbjct: 647 KTGRGTPQWMAPEVLRNEAADEKSDVYSFGVILWELVTEKIPWESLNAMQVIGAVGFMNQ 706
Query: 606 RLDLPEGLDPHVASIINDCWRR 627
RL++P+ +DP S++ CW R
Sbjct: 707 RLEVPKNVDPQWISLMESCWHR 728
Score = 70.9 bits (172), Expect = 1e-10
Identities = 52/159 (32%), Positives = 81/159 (50%), Gaps = 20/159 (12%)
Query: 1 NHSAETLYGWKDYEIIGKRVTKFLITEEYYAPLQKILERLITTGETWSGKFPYKKKCGEV 60
N AE LYG+ E +GK L+ + + Q I R ++GE+W+G+FP K K GE
Sbjct: 126 NSMAEKLYGFSASEALGKDPIDILVDVQDASVAQNITRRC-SSGESWTGEFPVKNKAGER 184
Query: 61 FMVMVTKAPLY-EDGVLVGVISVSSEAAKFNTI-------------DSEFQRTCQSGAS- 105
F V+ T +P Y +DG L+G+I +++++A F ++ F R S AS
Sbjct: 185 FSVVTTMSPSYDDDGCLIGIICITNDSALFQDPRGSPAKTRRGQEGETSFSRVTSSVASK 244
Query: 106 -GQPGVQKLNSKGNQWPLRPLIAPMPQIASSVSNLASKI 143
G + + SK +P+ IAS +S+LASK+
Sbjct: 245 LGLDSKEAVVSKLGLDSQQPI---QVAIASKISDLASKV 280
>UniRef100_Q9C902 Protein kinase, putative; 19229-23534 [Arabidopsis thaliana]
Length = 773
Score = 358 bits (920), Expect = 2e-97
Identities = 179/316 (56%), Positives = 227/316 (71%), Gaps = 33/316 (10%)
Query: 360 SSRESVGSHESS-SSKGDNDSV-VDCEIHWEDLHLRDEIGQGSYAVVYHGIWNGSDVAIK 417
SS S GS SS +K D DS ++ EI W+DL + +++GQGS VYHG+W GSDVA+K
Sbjct: 462 SSASSCGSTSSSVMNKVDTDSEGLEYEILWDDLTIGEQVGQGSCGTVYHGLWFGSDVAVK 521
Query: 418 VYFGNGYTEETLQDYKKEIDIMKRLRHPNVLLFMGAVYSLERLAIVTELLPRGSLFKTLH 477
V+ Y+ E ++ +K+E+ +MKRLRHPNVLLFMGAV S +RL IV+E LPRGSLF+ L
Sbjct: 522 VFSKQEYSAEVIESFKQEVLLMKRLRHPNVLLFMGAVTSPQRLCIVSEFLPRGSLFRLLQ 581
Query: 478 KSNQTLDIRRRLRMALDIARGMNYLHHRNPPIVHRDLKSSNLLVDKNWTVKVGDFGLSRL 537
KS LD RRR+ MALDIARGMNYLHH +PPI+HRDLKSSNLLVDKNWTVKV DFGLSR+
Sbjct: 582 KSTSKLDWRRRIHMALDIARGMNYLHHCSPPIIHRDLKSSNLLVDKNWTVKVADFGLSRI 641
Query: 538 KDTTLLTTKSGRGTPQWMAPEILRNEPSNEKSDVYSYGVVLWELMTQSIPWENLNSLQVV 597
K T LT+KSG+GTPQWMAPE+LRNE ++EKSD+YS+GVVLWEL T+ IPWE LNS+QV+
Sbjct: 642 KHETYLTSKSGKGTPQWMAPEVLRNESADEKSDIYSFGVVLWELATEKIPWETLNSMQVI 701
Query: 598 GVVGFMDRRLDLPEGLDPHVASIINDCWRRCFYMYSSLQLYYGRNHLLPQTESLLVLNMS 657
G VGFMD+RL++P+ +DP S++ CW
Sbjct: 702 GAVGFMDQRLEIPKDIDPRWISLMESCWH------------------------------- 730
Query: 658 SDPEQRPSFEELIQRM 673
SD + RP+F+EL+ ++
Sbjct: 731 SDTKLRPTFQELMDKL 746
Score = 58.2 bits (139), Expect = 8e-07
Identities = 44/156 (28%), Positives = 72/156 (45%), Gaps = 14/156 (8%)
Query: 1 NHSAETLYGWKDYEIIGKRVTKFLITEEYYAPLQKILERLITTGETWSGKFPYKKKCGEV 60
N AE +YG+ E +G+ + + A I R + GE+W+G+FP K K G+
Sbjct: 129 NAMAEKVYGYSAAEALGENPINVIADDRDAAFAMNIARRCVR-GESWTGEFPVKSKSGDR 187
Query: 61 FMVMVTKAPLY-EDGVLVGVISVSSEAAKFNT---------IDSEFQRTCQSGASGQPGV 110
F + T +P Y +DG L+G+I ++S A + E + + +
Sbjct: 188 FSAVTTCSPFYDDDGALMGIICITSNTAPYLNPRISLAKLKAQEEGETSSIPARNSFASK 247
Query: 111 QKLNSKGNQWPLRPLIAPMP---QIASSVSNLASKI 143
L+S+G L + P IAS +S+LASK+
Sbjct: 248 LGLDSRGAVISKLGLDSDQPIQVAIASKISDLASKV 283
>UniRef100_Q9C9V5 Hypothetical protein T23K23.26 [Arabidopsis thaliana]
Length = 738
Score = 356 bits (914), Expect = 1e-96
Identities = 173/270 (64%), Positives = 216/270 (79%), Gaps = 3/270 (1%)
Query: 360 SSRESVGSHESS-SSKGDNDS-VVDCEIHWEDLHLRDEIGQGSYAVVYHGIWNGSDVAIK 417
SS S GS SS +K D DS +D EI WEDL + ++IGQGS VYHG+W GSDVA+K
Sbjct: 455 SSASSCGSTSSSVMNKVDMDSDCLDYEILWEDLTIGEQIGQGSCGTVYHGLWFGSDVAVK 514
Query: 418 VYFGNGYTEETLQDYKKEIDIMKRLRHPNVLLFMGAVYSLERLAIVTELLPRGSLFKTLH 477
V+ Y+EE + +K+E+ +MKRLRHPNVLLFMGAV S +RL IVTE LPRGSLF+ L
Sbjct: 515 VFSKQEYSEEIITSFKQEVSLMKRLRHPNVLLFMGAVASPQRLCIVTEFLPRGSLFRLLQ 574
Query: 478 KSNQTLDIRRRLRMALDIARGMNYLHHRNPPIVHRDLKSSNLLVDKNWTVKVGDFGLSRL 537
++ LD+RRR+ MA DIARGMNYLHH +PPI+HRDLKSSNLLVD+NWTVKV DFGLSR+
Sbjct: 575 RNKSKLDLRRRIHMASDIARGMNYLHHCSPPIIHRDLKSSNLLVDRNWTVKVADFGLSRI 634
Query: 538 KDTTLLTTKSGRGTPQWMAPEILRNEPSNEKSDVYSYGVVLWELMTQSIPWENLNSLQVV 597
K T LTT +GRGTPQWMAPE+LRNE ++EKSDVYS+GVVLWEL+T+ IPWENLN++QV+
Sbjct: 635 KHETYLTT-NGRGTPQWMAPEVLRNEAADEKSDVYSFGVVLWELVTEKIPWENLNAMQVI 693
Query: 598 GVVGFMDRRLDLPEGLDPHVASIINDCWRR 627
G VGFM++RL++P+ +DP +++ CW R
Sbjct: 694 GAVGFMNQRLEVPKDVDPQWIALMESCWHR 723
Score = 72.8 bits (177), Expect = 3e-11
Identities = 54/158 (34%), Positives = 83/158 (52%), Gaps = 19/158 (12%)
Query: 1 NHSAETLYGWKDYEIIGKRVTKFLITEEYYAPLQKILERLITTGETWSGKFPYKKKCGEV 60
N AE LYG+ E +GK L+ + A + I +R ++GE+W+G+FP K K GE
Sbjct: 127 NSMAEKLYGFSAAEALGKDSINILVDGQDAAVAKNIFQRC-SSGESWTGEFPVKNKMGER 185
Query: 61 FMVMVTKAPLY-EDGVLVGVISVSSEAAKFNTI------------DSEFQRTCQSGAS-- 105
F V+ T +P Y +DG+L+G+I +++++A F DS F R AS
Sbjct: 186 FSVVTTISPFYDDDGLLIGIICITNDSALFQRPRVPPAKNRWQEGDSSFCRGTNGVASRL 245
Query: 106 GQPGVQKLNSKGNQWPLRPLIAPMPQIASSVSNLASKI 143
G + + SK +P+ A IAS +S+LASK+
Sbjct: 246 GFDSKEAVVSKLGLDSQQPIQA---AIASKISDLASKV 280
>UniRef100_Q93YU0 Hypothetical protein At1g67890 [Arabidopsis thaliana]
Length = 765
Score = 356 bits (913), Expect = 1e-96
Identities = 181/326 (55%), Positives = 232/326 (70%), Gaps = 34/326 (10%)
Query: 360 SSRESVGSHESS-SSKGDNDS-VVDCEIHWEDLHLRDEIGQGSYAVVYHGIWNGSDVAIK 417
SS S GS SS +K D DS +D EI WEDL ++IGQGS VYHG+W GSDVA+K
Sbjct: 455 SSASSCGSTSSSVMNKVDMDSDCLDYEILWEDLTNGEQIGQGSCGTVYHGLWFGSDVAVK 514
Query: 418 VYFGNGYTEETLQDYKKEIDIMKRLRHPNVLLFMGAVYSLERLAIVTELLPRGSLFKTLH 477
V+ Y+EE + +K+E+ +MKRLRHPNVLLFMGAV S +RL IVTE LPRGSLF+ L
Sbjct: 515 VFSKQEYSEEIITSFKQEVSLMKRLRHPNVLLFMGAVASPQRLCIVTEFLPRGSLFRLLQ 574
Query: 478 KSNQTLDIRRRLRMALDIARGMNYLHHRNPPIVHRDLKSSNLLVDKNWTVKVGDFGLSRL 537
++ LD+RRR+ MA DIARGMNYLHH +PPI+HRDLKSSNLLVD+NWTVKV DFGLSR+
Sbjct: 575 RNKSKLDLRRRIHMASDIARGMNYLHHCSPPIIHRDLKSSNLLVDRNWTVKVADFGLSRI 634
Query: 538 KDTTLLTTKSGRGTPQWMAPEILRNEPSNEKSDVYSYGVVLWELMTQSIPWENLNSLQVV 597
K T LTT +GRGTPQWMAPE+LRNE ++EKSDVYS+GVVLWEL+T+ IPWENLN++QV+
Sbjct: 635 KHETYLTT-NGRGTPQWMAPEVLRNEAADEKSDVYSFGVVLWELVTEKIPWENLNAMQVI 693
Query: 598 GVVGFMDRRLDLPEGLDPHVASIINDCWRRCFYMYSSLQLYYGRNHLLPQTESLLVLNMS 657
G VGFM++RL++P+ +DP +++ CW
Sbjct: 694 GAVGFMNQRLEVPKDVDPQWIALMESCWH------------------------------- 722
Query: 658 SDPEQRPSFEELIQRMLFVLNRVTAE 683
S+P+ RPSF+EL+ ++ + + T +
Sbjct: 723 SEPQCRPSFQELMDKLRELQRKYTIQ 748
Score = 73.2 bits (178), Expect = 3e-11
Identities = 54/158 (34%), Positives = 83/158 (52%), Gaps = 19/158 (12%)
Query: 1 NHSAETLYGWKDYEIIGKRVTKFLITEEYYAPLQKILERLITTGETWSGKFPYKKKCGEV 60
N AE LYG+ E +GK L+ + A + I +R ++GE+W+G+FP K K GE
Sbjct: 127 NSMAEKLYGFSAAEALGKDSINILVDGQDAAVAKNIFQRC-SSGESWTGEFPVKNKMGES 185
Query: 61 FMVMVTKAPLY-EDGVLVGVISVSSEAAKFNTI------------DSEFQRTCQSGAS-- 105
F V+ T +P Y +DG+L+G+I +++++A F DS F R AS
Sbjct: 186 FSVVTTISPFYDDDGLLIGIICITNDSALFQRPRVPPAKNRWQEGDSSFCRGTNGVASRL 245
Query: 106 GQPGVQKLNSKGNQWPLRPLIAPMPQIASSVSNLASKI 143
G + + SK +P+ A IAS +S+LASK+
Sbjct: 246 GFDSKEAVVSKLGLDSQQPIQA---AIASKISDLASKV 280
>UniRef100_Q9C547 Protein kinase, putative; 12576-15979 [Arabidopsis thaliana]
Length = 671
Score = 348 bits (892), Expect = 4e-94
Identities = 169/270 (62%), Positives = 212/270 (77%), Gaps = 2/270 (0%)
Query: 360 SSRESVGSHESS-SSKGDNDS-VVDCEIHWEDLHLRDEIGQGSYAVVYHGIWNGSDVAIK 417
+S S GS S K D DS ++ EI W+DL + ++IG+GS VYHGIW GSDVA+K
Sbjct: 402 NSASSCGSTSRSVMDKVDIDSDPLEHEILWDDLTIGEQIGRGSCGTVYHGIWFGSDVAVK 461
Query: 418 VYFGNGYTEETLQDYKKEIDIMKRLRHPNVLLFMGAVYSLERLAIVTELLPRGSLFKTLH 477
V+ Y+E ++ ++KE+ +MKRLRHPNVLLFMGAV S +RL IV+E +PRGSLF+ L
Sbjct: 462 VFSKQEYSESVIKSFEKEVSLMKRLRHPNVLLFMGAVTSPQRLCIVSEFVPRGSLFRLLQ 521
Query: 478 KSNQTLDIRRRLRMALDIARGMNYLHHRNPPIVHRDLKSSNLLVDKNWTVKVGDFGLSRL 537
+S LD RRR+ MALDIARGMNYLH +PPI+HRDLKSSNLLVD+NWTVKV DFGLSR+
Sbjct: 522 RSMSKLDWRRRINMALDIARGMNYLHCCSPPIIHRDLKSSNLLVDRNWTVKVADFGLSRI 581
Query: 538 KDTTLLTTKSGRGTPQWMAPEILRNEPSNEKSDVYSYGVVLWELMTQSIPWENLNSLQVV 597
K T LT+KSG+GTPQWMAPE+LRNE ++EKSD+YS+GVVLWEL T+ IPWENLNS+QV+
Sbjct: 582 KHQTYLTSKSGKGTPQWMAPEVLRNESADEKSDIYSFGVVLWELATEKIPWENLNSMQVI 641
Query: 598 GVVGFMDRRLDLPEGLDPHVASIINDCWRR 627
G VGFM++RL++P+ DP S+I CW R
Sbjct: 642 GAVGFMNQRLEIPKDTDPDWISLIESCWHR 671
Score = 69.7 bits (169), Expect = 3e-10
Identities = 46/147 (31%), Positives = 76/147 (51%), Gaps = 8/147 (5%)
Query: 1 NHSAETLYGWKDYEIIGKRVTKFLITEEYYAPLQKILERLITTGETWSGKFPYKKKCGEV 60
N +E LYG+ E++G+ ++ ++ A + R GE+W+G+FP K K G++
Sbjct: 90 NAMSEKLYGYSAAEVVGRNPVHVIVDDQNAAFALNVARRC-ANGESWTGEFPVKTKSGKI 148
Query: 61 FMVMVTKAPLYED-GVLVGVISVSSEAAKFNTIDSEFQRTCQSGASGQPGVQKLNSKGNQ 119
F + T +P Y+D G +VG+IS++S+ A + R + G L+SKG
Sbjct: 149 FSAVTTCSPFYDDNGTVVGIISITSDIAPYLNPRLSLPRLKPQEPERKLG---LDSKGAV 205
Query: 120 WPLRPLIAPMP---QIASSVSNLASKI 143
L + P IAS +S+LASK+
Sbjct: 206 ISKPGLDSDQPIQVSIASKISSLASKL 232
>UniRef100_Q9C903 Protein kinase, putative; 8050-11829 [Arabidopsis thaliana]
Length = 763
Score = 342 bits (878), Expect = 2e-92
Identities = 167/317 (52%), Positives = 222/317 (69%), Gaps = 31/317 (9%)
Query: 367 SHESSSSKGDNDSVVDCEIHWEDLHLRDEIGQGSYAVVYHGIWNGSDVAIKVYFGNGYTE 426
SH + + N + ++ EI W+DL + ++IGQGS VYHG+W GSDVA+K+ Y+E
Sbjct: 423 SHSTMNKVDTNSNCLEYEILWDDLTIGEQIGQGSCGTVYHGLWFGSDVAVKLISKQEYSE 482
Query: 427 ETLQDYKKEIDIMKRLRHPNVLLFMGAVYSLERLAIVTELLPRGSLFKTLHKSNQTLDIR 486
E +Q +++E+ +M+RLRHPNVLLFMGAV + L IV+E LPRGSLF+ L ++ LD R
Sbjct: 483 EVIQSFRQEVSLMQRLRHPNVLLFMGAVTLPQGLCIVSEFLPRGSLFRLLQRNMSKLDWR 542
Query: 487 RRLRMALDIARGMNYLHHRNPPIVHRDLKSSNLLVDKNWTVKVGDFGLSRLKDTTLLTTK 546
RR+ MALDIARGMNYLH +PPI+HRDLKSSNLLVDKN TVKV DFGLSR+K T LT+K
Sbjct: 543 RRINMALDIARGMNYLHRCSPPIIHRDLKSSNLLVDKNLTVKVADFGLSRIKHHTYLTSK 602
Query: 547 SGRGTPQWMAPEILRNEPSNEKSDVYSYGVVLWELMTQSIPWENLNSLQVVGVVGFMDRR 606
SG+G PQWMAPE+LRNE ++EKSD+YS+GVVLWEL T+ IPWENLNS+QV+G VGFM++R
Sbjct: 603 SGKGMPQWMAPEVLRNESADEKSDIYSFGVVLWELATEKIPWENLNSMQVIGAVGFMNQR 662
Query: 607 LDLPEGLDPHVASIINDCWRRCFYMYSSLQLYYGRNHLLPQTESLLVLNMSSDPEQRPSF 666
L++P+ +DP S+I CW R D + RP+F
Sbjct: 663 LEIPKDIDPDWISLIESCWHR-------------------------------DAKLRPTF 691
Query: 667 EELIQRMLFVLNRVTAE 683
+EL++R+ + + T +
Sbjct: 692 QELMERLRDLQRKYTIQ 708
Score = 72.0 bits (175), Expect = 6e-11
Identities = 48/155 (30%), Positives = 81/155 (51%), Gaps = 13/155 (8%)
Query: 1 NHSAETLYGWKDYEIIGKRVTKFLITEEYYAPLQKILERLITTGETWSGKFPYKKKCGEV 60
N AE +YG+ E +G+ ++ ++ AP + +L + GE+W+GKFP K++ GE
Sbjct: 90 NAMAEKVYGYSAAEAVGQNPIDVMV-DDRDAPFAMTIAQLCSNGESWTGKFPVKRRTGEK 148
Query: 61 FMVMVTKAPLY-EDGVLVGVISVSSEAAKF--NTIDSEFQRTCQSGASGQPGVQK----- 112
F + T +P Y +DG L+G++S++S+ A + TI + + S P
Sbjct: 149 FSAVTTCSPFYADDGSLIGIVSITSDVAPYLNPTISLAKLKASEVETSSTPARNSFAFKL 208
Query: 113 -LNSKGNQWPLRPLIAPMP---QIASSVSNLASKI 143
L++KG L + P IAS +S+LASK+
Sbjct: 209 GLDTKGAVVSKLGLDSDQPIQVAIASKISDLASKV 243
>UniRef100_Q9C833 Protein kinase, putative; 42705-46677 [Arabidopsis thaliana]
Length = 777
Score = 342 bits (878), Expect = 2e-92
Identities = 167/317 (52%), Positives = 222/317 (69%), Gaps = 31/317 (9%)
Query: 367 SHESSSSKGDNDSVVDCEIHWEDLHLRDEIGQGSYAVVYHGIWNGSDVAIKVYFGNGYTE 426
SH + + N + ++ EI W+DL + ++IGQGS VYHG+W GSDVA+K+ Y+E
Sbjct: 423 SHSTMNKVDTNSNCLEYEILWDDLTIGEQIGQGSCGTVYHGLWFGSDVAVKLISKQEYSE 482
Query: 427 ETLQDYKKEIDIMKRLRHPNVLLFMGAVYSLERLAIVTELLPRGSLFKTLHKSNQTLDIR 486
E +Q +++E+ +M+RLRHPNVLLFMGAV + L IV+E LPRGSLF+ L ++ LD R
Sbjct: 483 EVIQSFRQEVSLMQRLRHPNVLLFMGAVTLPQGLCIVSEFLPRGSLFRLLQRNMSKLDWR 542
Query: 487 RRLRMALDIARGMNYLHHRNPPIVHRDLKSSNLLVDKNWTVKVGDFGLSRLKDTTLLTTK 546
RR+ MALDIARGMNYLH +PPI+HRDLKSSNLLVDKN TVKV DFGLSR+K T LT+K
Sbjct: 543 RRINMALDIARGMNYLHRCSPPIIHRDLKSSNLLVDKNLTVKVADFGLSRIKHHTYLTSK 602
Query: 547 SGRGTPQWMAPEILRNEPSNEKSDVYSYGVVLWELMTQSIPWENLNSLQVVGVVGFMDRR 606
SG+G PQWMAPE+LRNE ++EKSD+YS+GVVLWEL T+ IPWENLNS+QV+G VGFM++R
Sbjct: 603 SGKGMPQWMAPEVLRNESADEKSDIYSFGVVLWELATEKIPWENLNSMQVIGAVGFMNQR 662
Query: 607 LDLPEGLDPHVASIINDCWRRCFYMYSSLQLYYGRNHLLPQTESLLVLNMSSDPEQRPSF 666
L++P+ +DP S+I CW R D + RP+F
Sbjct: 663 LEIPKDIDPDWISLIESCWHR-------------------------------DAKLRPTF 691
Query: 667 EELIQRMLFVLNRVTAE 683
+EL++R+ + + T +
Sbjct: 692 QELMERLRDLQRKYTIQ 708
Score = 72.0 bits (175), Expect = 6e-11
Identities = 48/155 (30%), Positives = 81/155 (51%), Gaps = 13/155 (8%)
Query: 1 NHSAETLYGWKDYEIIGKRVTKFLITEEYYAPLQKILERLITTGETWSGKFPYKKKCGEV 60
N AE +YG+ E +G+ ++ ++ AP + +L + GE+W+GKFP K++ GE
Sbjct: 90 NAMAEKVYGYSAAEAVGQNPIDVMV-DDRDAPFAMTIAQLCSNGESWTGKFPVKRRTGEK 148
Query: 61 FMVMVTKAPLY-EDGVLVGVISVSSEAAKF--NTIDSEFQRTCQSGASGQPGVQK----- 112
F + T +P Y +DG L+G++S++S+ A + TI + + S P
Sbjct: 149 FSAVTTCSPFYADDGSLIGIVSITSDVAPYLNPTISLAKLKASEVETSSTPARNSFAFKL 208
Query: 113 -LNSKGNQWPLRPLIAPMP---QIASSVSNLASKI 143
L++KG L + P IAS +S+LASK+
Sbjct: 209 GLDTKGAVVSKLGLDSDQPIQVAIASKISDLASKV 243
>UniRef100_Q67UL6 Putative CTR1-like kinase kinase kinase [Oryza sativa]
Length = 1078
Score = 330 bits (847), Expect = 7e-89
Identities = 175/385 (45%), Positives = 236/385 (60%), Gaps = 47/385 (12%)
Query: 318 LVPQDPLLRLHRQFDPKKLEPEATHMAIEDEVQKQQGRLQLPS---------------SR 362
L P LL L +L P+ H +++++ G+ +P S
Sbjct: 718 LEPGCQLLSLPSSSGANELIPKGRHDFWDNQLEIDHGQTSVPEKEKDLVEVPQEAERVSD 777
Query: 363 ESVGSHESSSSKGDNDSVVDCEIHWEDLHLRDEIGQGSYAVVYHGIWNGSDVAIKVYFGN 422
+SVG+ ESS S D V + EI WE++ L + +G GS+ VY G W+G++VA+K +
Sbjct: 778 KSVGT-ESSRSDIALDGVAEFEIQWEEITLGERVGLGSFGEVYKGEWHGTEVAVKKFLQQ 836
Query: 423 GYTEETLQDYKKEIDIMKRLRHPNVLLFMGAVYSLERLAIVTELLPRGSLFKTLHKSNQT 482
+ + L +++ E IMKRLRHPNV+LFMGAV + L+IVTE LPRGSLF+ +H+ N
Sbjct: 837 DISSDALDEFRTEFQIMKRLRHPNVVLFMGAVTRVPNLSIVTEFLPRGSLFRLIHRPNNQ 896
Query: 483 LDIRRRLRMALDIARGMNYLHHRNPPIVHRDLKSSNLLVDKNWTVKVGDFGLSRLKDTTL 542
LD RRRLRMALD+ARGMNYLH+ +P +VHRDLKS NLLVDKNW VKV DFGLSR+K++T
Sbjct: 897 LDERRRLRMALDVARGMNYLHNCSPVVVHRDLKSPNLLVDKNWVVKVCDFGLSRMKNSTF 956
Query: 543 LTTKSGRGTPQWMAPEILRNEPSNEKSDVYSYGVVLWELMTQSIPWENLNSLQVVGVVGF 602
L+++S GT +WMAPE+LRNEPS+EK DV+SYGV+LWEL T PWE +N +QVVG VGF
Sbjct: 957 LSSRSTAGTAEWMAPEVLRNEPSDEKCDVFSYGVILWELFTLLQPWEGMNPMQVVGAVGF 1016
Query: 603 MDRRLDLPEGLDPHVASIINDCWRRCFYMYSSLQLYYGRNHLLPQTESLLVLNMSSDPEQ 662
RRLD+P +DP +A II CW+ +DP+
Sbjct: 1017 QQRRLDIPAHVDPTIAEIIRRCWQ-------------------------------TDPKM 1045
Query: 663 RPSFEELIQRMLFVLNRVTAESVKR 687
RPSF E++ + +L A +R
Sbjct: 1046 RPSFSEIMSSLKPLLKNTLANQPQR 1070
>UniRef100_Q9C9U5 Hypothetical protein F25P22.8 [Arabidopsis thaliana]
Length = 1030
Score = 330 bits (846), Expect = 9e-89
Identities = 169/327 (51%), Positives = 218/327 (65%), Gaps = 34/327 (10%)
Query: 361 SRESVGSHESSSSKGDNDSVVDCEIHWEDLHLRDEIGQGSYAVVYHGIWNGSDVAIKVYF 420
S +S+G+ SSK D D V DCEI WE++ + + IG GSY VY G W+G++VA+K +
Sbjct: 722 SDKSIGNE---SSKSDCDDVSDCEILWEEITVGERIGLGSYGEVYRGDWHGTEVAVKKFL 778
Query: 421 GNGYTEETLQDYKKEIDIMKRLRHPNVLLFMGAVYSLERLAIVTELLPRGSLFKTLHKSN 480
T E L++++ E+ IMK+LRHPN++LFMGAV L+IVTE LPRGSL++ +H+ N
Sbjct: 779 DQDLTGEALEEFRSEVRIMKKLRHPNIVLFMGAVTRPPNLSIVTEFLPRGSLYRLIHRPN 838
Query: 481 QTLDIRRRLRMALDIARGMNYLHHRNPPIVHRDLKSSNLLVDKNWTVKVGDFGLSRLKDT 540
LD RRRLRMALD ARGMNYLH NP IVHRDLKS NLLVDKNW VKV DFGLSR+K +
Sbjct: 839 NQLDERRRLRMALDAARGMNYLHSCNPMIVHRDLKSPNLLVDKNWVVKVCDFGLSRMKHS 898
Query: 541 TLLTTKSGRGTPQWMAPEILRNEPSNEKSDVYSYGVVLWELMTQSIPWENLNSLQVVGVV 600
T L++KS GT +WMAPE+LRNEP++EK DVYSYGV+LWEL T PW +N +QVVG V
Sbjct: 899 TYLSSKSTAGTAEWMAPEVLRNEPADEKCDVYSYGVILWELFTLQQPWGKMNPMQVVGAV 958
Query: 601 GFMDRRLDLPEGLDPHVASIINDCWRRCFYMYSSLQLYYGRNHLLPQTESLLVLNMSSDP 660
GF RRLD+P+ +DP +A +I+ CW QT+S L
Sbjct: 959 GFQHRRLDIPDFVDPAIADLISKCW---------------------QTDSKL-------- 989
Query: 661 EQRPSFEELIQRMLFVLNRVTAESVKR 687
RPSF E++ + + VT ++ R
Sbjct: 990 --RPSFAEIMASLKRLQKPVTGSNIPR 1014
>UniRef100_Q8L625 Hypothetical protein At1g73660 [Arabidopsis thaliana]
Length = 1030
Score = 330 bits (846), Expect = 9e-89
Identities = 169/327 (51%), Positives = 218/327 (65%), Gaps = 34/327 (10%)
Query: 361 SRESVGSHESSSSKGDNDSVVDCEIHWEDLHLRDEIGQGSYAVVYHGIWNGSDVAIKVYF 420
S +S+G+ SSK D D V DCEI WE++ + + IG GSY VY G W+G++VA+K +
Sbjct: 722 SDKSIGNE---SSKSDCDDVSDCEILWEEITVGERIGLGSYGEVYRGDWHGTEVAVKKFL 778
Query: 421 GNGYTEETLQDYKKEIDIMKRLRHPNVLLFMGAVYSLERLAIVTELLPRGSLFKTLHKSN 480
T E L++++ E+ IMK+LRHPN++LFMGAV L+IVTE LPRGSL++ +H+ N
Sbjct: 779 DQDLTGEALEEFRSEVRIMKKLRHPNIVLFMGAVTRPPNLSIVTEFLPRGSLYRLIHRPN 838
Query: 481 QTLDIRRRLRMALDIARGMNYLHHRNPPIVHRDLKSSNLLVDKNWTVKVGDFGLSRLKDT 540
LD RRRLRMALD ARGMNYLH NP IVHRDLKS NLLVDKNW VKV DFGLSR+K +
Sbjct: 839 NQLDERRRLRMALDAARGMNYLHSCNPMIVHRDLKSPNLLVDKNWVVKVCDFGLSRMKHS 898
Query: 541 TLLTTKSGRGTPQWMAPEILRNEPSNEKSDVYSYGVVLWELMTQSIPWENLNSLQVVGVV 600
T L++KS GT +WMAPE+LRNEP++EK DVYSYGV+LWEL T PW +N +QVVG V
Sbjct: 899 TYLSSKSTAGTAEWMAPEVLRNEPADEKCDVYSYGVILWELFTLQQPWGKMNPMQVVGAV 958
Query: 601 GFMDRRLDLPEGLDPHVASIINDCWRRCFYMYSSLQLYYGRNHLLPQTESLLVLNMSSDP 660
GF RRLD+P+ +DP +A +I+ CW QT+S L
Sbjct: 959 GFQHRRLDIPDFVDPAIADLISKCW---------------------QTDSKL-------- 989
Query: 661 EQRPSFEELIQRMLFVLNRVTAESVKR 687
RPSF E++ + + VT ++ R
Sbjct: 990 --RPSFAEIMASLKRLQKPVTGSNIPR 1014
>UniRef100_Q7X8K7 CTR1-like kinase kinase kinase [Brassica juncea]
Length = 970
Score = 328 bits (842), Expect = 3e-88
Identities = 154/258 (59%), Positives = 196/258 (75%)
Query: 369 ESSSSKGDNDSVVDCEIHWEDLHLRDEIGQGSYAVVYHGIWNGSDVAIKVYFGNGYTEET 428
ESS S G D V DCEI WE++ L + IG GSY VY G W+G++VA K + T E
Sbjct: 666 ESSKSDGTLDDVSDCEILWEEITLGERIGLGSYGEVYRGDWHGTEVAAKKFLDQDLTGEA 725
Query: 429 LQDYKKEIDIMKRLRHPNVLLFMGAVYSLERLAIVTELLPRGSLFKTLHKSNQTLDIRRR 488
L++++ E+ IMK+LRHPN++LFMGAV L+I+TE LPRGSL++ +H+ N LD RRR
Sbjct: 726 LEEFRSEVQIMKKLRHPNIVLFMGAVTRPPNLSIITEFLPRGSLYRLIHRPNNQLDERRR 785
Query: 489 LRMALDIARGMNYLHHRNPPIVHRDLKSSNLLVDKNWTVKVGDFGLSRLKDTTLLTTKSG 548
LRMALD ARGMNYLH +P IVHRDLKS NLLVDKNW VKV DFGLSR+K++T L++KS
Sbjct: 786 LRMALDAARGMNYLHSCSPMIVHRDLKSPNLLVDKNWVVKVCDFGLSRMKNSTYLSSKST 845
Query: 549 RGTPQWMAPEILRNEPSNEKSDVYSYGVVLWELMTQSIPWENLNSLQVVGVVGFMDRRLD 608
GT +WMAPE+LRNEP++EK DVYSYGV+LWEL T PW +N++QVVG VGF RRLD
Sbjct: 846 AGTAEWMAPEVLRNEPADEKCDVYSYGVILWELFTLQQPWGRMNAMQVVGAVGFQHRRLD 905
Query: 609 LPEGLDPHVASIINDCWR 626
+P+ +DP +A +I+ CW+
Sbjct: 906 IPDFVDPAIAELISKCWQ 923
>UniRef100_Q6YW44 Putative MAP3K delta-1 protein kinase [Oryza sativa]
Length = 864
Score = 328 bits (841), Expect = 3e-88
Identities = 168/346 (48%), Positives = 222/346 (63%), Gaps = 35/346 (10%)
Query: 343 MAIEDEVQKQQGRLQLP-SSRESVGSHESSSSKGDN---DSVVDCEIHWEDLHLRDEIGQ 398
+A ED+ + +++P SS ++ SS+K + D V D EI WEDLH+ + IG
Sbjct: 549 IANEDQRFSEDSLVKMPGSSNGNLDKSSCSSTKTISSVIDDVADYEIPWEDLHIGERIGL 608
Query: 399 GSYAVVYHGIWNGSDVAIKVYFGNGYTEETLQDYKKEIDIMKRLRHPNVLLFMGAVYSLE 458
GSY VYH WNG++VA+K + + L +K E+ IM RLRHPNV+LF+G V
Sbjct: 609 GSYGEVYHADWNGTEVAVKKFLDQDLSGVALDQFKCEVGIMSRLRHPNVVLFLGYVTQPP 668
Query: 459 RLAIVTELLPRGSLFKTLHKSNQTLDIRRRLRMALDIARGMNYLHHRNPPIVHRDLKSSN 518
L+I+TE LPRGSL++ LH+ N +D RRL+MALD+A+GMNYLH +P IVHRDLKS N
Sbjct: 669 NLSILTEYLPRGSLYRLLHRPNSQIDETRRLKMALDVAKGMNYLHASHPTIVHRDLKSPN 728
Query: 519 LLVDKNWTVKVGDFGLSRLKDTTLLTTKSGRGTPQWMAPEILRNEPSNEKSDVYSYGVVL 578
LLVDKNW VKV DFG+SRLK T L++KS GTP+WMAPE+LRNEPSNEK DVYS+GV+L
Sbjct: 729 LLVDKNWVVKVSDFGMSRLKHHTFLSSKSTAGTPEWMAPEVLRNEPSNEKCDVYSFGVIL 788
Query: 579 WELMTQSIPWENLNSLQVVGVVGFMDRRLDLPEGLDPHVASIINDCWRRCFYMYSSLQLY 638
WEL T +PW LN +QVVG VGF +RRL++P+ +DP VA+II+ CW
Sbjct: 789 WELATMRVPWSGLNPMQVVGAVGFQNRRLEIPKEIDPLVATIISSCW------------- 835
Query: 639 YGRNHLLPQTESLLVLNMSSDPEQRPSFEELIQRMLFVLNRVTAES 684
+DP +RPSF +L+ + + V E+
Sbjct: 836 ------------------ENDPSKRPSFSQLLSPLKQLQRLVVPEN 863
>UniRef100_Q9FPR4 EDR1 [Hordeum vulgare]
Length = 957
Score = 324 bits (830), Expect = 6e-87
Identities = 159/299 (53%), Positives = 211/299 (70%), Gaps = 8/299 (2%)
Query: 331 FDPKKLEPEATHMAIEDEVQKQQGRLQLPSSRESVGSHESSSSKGDN--DSVVDCEIHWE 388
+D + L P+ ++ + + + + E V + SSK D D V +CEI WE
Sbjct: 623 YDHRMLHPDPRKSPLDRFMDRPRQNI------ECVSPSQVGSSKVDLVLDEVSECEILWE 676
Query: 389 DLHLRDEIGQGSYAVVYHGIWNGSDVAIKVYFGNGYTEETLQDYKKEIDIMKRLRHPNVL 448
DL + + IG GSY VYH WNG++VA+K + + + L++++ E+ IM+RLRHPN++
Sbjct: 677 DLVIDERIGIGSYGEVYHADWNGTEVAVKKFLDQEFYGDALEEFRCEVRIMRRLRHPNIV 736
Query: 449 LFMGAVYSLERLAIVTELLPRGSLFKTLHKSNQTLDIRRRLRMALDIARGMNYLHHRNPP 508
LFMGAV L+IV+E LPRGSL+K +H+ N +D +RR++MALD+ARGMN LH P
Sbjct: 737 LFMGAVTRPPHLSIVSEYLPRGSLYKIIHRPNCQIDEKRRIKMALDVARGMNCLHTSVPT 796
Query: 509 IVHRDLKSSNLLVDKNWTVKVGDFGLSRLKDTTLLTTKSGRGTPQWMAPEILRNEPSNEK 568
IVHRDLKS NLLVD NWTVKV DFGLSRLK +T L++KS GTP+WMAPE+LRNE SNEK
Sbjct: 797 IVHRDLKSPNLLVDDNWTVKVCDFGLSRLKHSTFLSSKSTAGTPEWMAPEVLRNEQSNEK 856
Query: 569 SDVYSYGVVLWELMTQSIPWENLNSLQVVGVVGFMDRRLDLPEGLDPHVASIINDCWRR 627
D+YS+GV+LWEL T PW +N +QVVG VGF DRRLD+P+ +DP VASII DCW++
Sbjct: 857 CDIYSFGVILWELATLRKPWHGMNQMQVVGAVGFQDRRLDIPKEVDPIVASIIRDCWQK 915
>UniRef100_Q6Z2V3 Putative MAP kinase kinase kinase [Oryza sativa]
Length = 1111
Score = 324 bits (830), Expect = 6e-87
Identities = 158/343 (46%), Positives = 225/343 (65%), Gaps = 31/343 (9%)
Query: 345 IEDEVQKQQGRLQLPSSRESVGSHESSSSKGDNDSVVDCEIHWEDLHLRDEIGQGSYAVV 404
++ E + R + + ES+ S+ D V + EI WE++ + + IG GS+ V
Sbjct: 793 LDQEKDSAEVRQDAERTSDKSSGTESAKSEITLDDVAEFEIQWEEITIGERIGLGSFGEV 852
Query: 405 YHGIWNGSDVAIKVYFGNGYTEETLQDYKKEIDIMKRLRHPNVLLFMGAVYSLERLAIVT 464
Y G W+G++VA+K + + + L++++ E+ I+KRLRHPNV+LFMGA+ + L+IVT
Sbjct: 853 YRGEWHGTEVAVKKFLQQDISSDALEEFRTEVRIIKRLRHPNVVLFMGAITRVPNLSIVT 912
Query: 465 ELLPRGSLFKTLHKSNQTLDIRRRLRMALDIARGMNYLHHRNPPIVHRDLKSSNLLVDKN 524
E LPRGSLF+ +H+ N LD R+RLRMALD+ARGMNYLH+ P IVHRDLKS NLLVDKN
Sbjct: 913 EFLPRGSLFRLIHRPNNQLDERKRLRMALDVARGMNYLHNCTPVIVHRDLKSPNLLVDKN 972
Query: 525 WTVKVGDFGLSRLKDTTLLTTKSGRGTPQWMAPEILRNEPSNEKSDVYSYGVVLWELMTQ 584
W VKV DFGLS++K+ T L+++S GT +WMAPE+LRNEPS+EK DV+SYGV+LWEL T
Sbjct: 973 WVVKVCDFGLSKMKNKTFLSSRSTAGTAEWMAPEVLRNEPSDEKCDVFSYGVILWELCTL 1032
Query: 585 SIPWENLNSLQVVGVVGFMDRRLDLPEGLDPHVASIINDCWRRCFYMYSSLQLYYGRNHL 644
PWE +N++QVVG VGF +RRLD+P+ DP +A II CW+
Sbjct: 1033 LQPWEGMNAMQVVGAVGFQNRRLDIPDNTDPAIAEIIAKCWQ------------------ 1074
Query: 645 LPQTESLLVLNMSSDPEQRPSFEELIQRMLFVLNRVTAESVKR 687
+DP+ RPSF +++ + +L +TA++ ++
Sbjct: 1075 -------------TDPKLRPSFADIMASLKPLLKNMTAQAPRQ 1104
>UniRef100_Q6Z2V2 Putative MAP kinase kinase kinase [Oryza sativa]
Length = 991
Score = 324 bits (830), Expect = 6e-87
Identities = 158/343 (46%), Positives = 225/343 (65%), Gaps = 31/343 (9%)
Query: 345 IEDEVQKQQGRLQLPSSRESVGSHESSSSKGDNDSVVDCEIHWEDLHLRDEIGQGSYAVV 404
++ E + R + + ES+ S+ D V + EI WE++ + + IG GS+ V
Sbjct: 673 LDQEKDSAEVRQDAERTSDKSSGTESAKSEITLDDVAEFEIQWEEITIGERIGLGSFGEV 732
Query: 405 YHGIWNGSDVAIKVYFGNGYTEETLQDYKKEIDIMKRLRHPNVLLFMGAVYSLERLAIVT 464
Y G W+G++VA+K + + + L++++ E+ I+KRLRHPNV+LFMGA+ + L+IVT
Sbjct: 733 YRGEWHGTEVAVKKFLQQDISSDALEEFRTEVRIIKRLRHPNVVLFMGAITRVPNLSIVT 792
Query: 465 ELLPRGSLFKTLHKSNQTLDIRRRLRMALDIARGMNYLHHRNPPIVHRDLKSSNLLVDKN 524
E LPRGSLF+ +H+ N LD R+RLRMALD+ARGMNYLH+ P IVHRDLKS NLLVDKN
Sbjct: 793 EFLPRGSLFRLIHRPNNQLDERKRLRMALDVARGMNYLHNCTPVIVHRDLKSPNLLVDKN 852
Query: 525 WTVKVGDFGLSRLKDTTLLTTKSGRGTPQWMAPEILRNEPSNEKSDVYSYGVVLWELMTQ 584
W VKV DFGLS++K+ T L+++S GT +WMAPE+LRNEPS+EK DV+SYGV+LWEL T
Sbjct: 853 WVVKVCDFGLSKMKNKTFLSSRSTAGTAEWMAPEVLRNEPSDEKCDVFSYGVILWELCTL 912
Query: 585 SIPWENLNSLQVVGVVGFMDRRLDLPEGLDPHVASIINDCWRRCFYMYSSLQLYYGRNHL 644
PWE +N++QVVG VGF +RRLD+P+ DP +A II CW+
Sbjct: 913 LQPWEGMNAMQVVGAVGFQNRRLDIPDNTDPAIAEIIAKCWQ------------------ 954
Query: 645 LPQTESLLVLNMSSDPEQRPSFEELIQRMLFVLNRVTAESVKR 687
+DP+ RPSF +++ + +L +TA++ ++
Sbjct: 955 -------------TDPKLRPSFADIMASLKPLLKNMTAQAPRQ 984
>UniRef100_Q8LPH3 MAP kinase, putative [Arabidopsis thaliana]
Length = 992
Score = 323 bits (829), Expect = 8e-87
Identities = 165/329 (50%), Positives = 214/329 (64%), Gaps = 32/329 (9%)
Query: 361 SRESVGSHESSSSKGDNDSVVDCEIHWEDLHLRDEIGQGSYAVVYHGIWNGSDVAIKVYF 420
S S+G+ ESS S D V +CEI WE++ + + IG GSY VY G W+G+ VA+K +
Sbjct: 687 SDRSIGN-ESSKSDAAIDDVAECEILWEEITVAERIGLGSYGEVYRGDWHGTAVAVKKFI 745
Query: 421 GNGYTEETLQDYKKEIDIMKRLRHPNVLLFMGAVYSLERLAIVTELLPRGSLFKTLHKSN 480
T E L++++ E+ +M+RLRHPN++LFMGAV L+IVTE LPRGSL++ +H+ N
Sbjct: 746 DQDITGEALEEFRSEVRMMRRLRHPNIVLFMGAVTRPPNLSIVTEFLPRGSLYRLIHRPN 805
Query: 481 QTLDIRRRLRMALDIARGMNYLHHRNPPIVHRDLKSSNLLVDKNWTVKVGDFGLSRLKDT 540
LD R+RLRMALD ARGMNYLH NP IVHRDLKS NLLVDKNW VKV DFGLSR+K +
Sbjct: 806 NQLDERKRLRMALDAARGMNYLHSCNPVIVHRDLKSPNLLVDKNWVVKVCDFGLSRMKVS 865
Query: 541 TLLTTKSGRGTPQWMAPEILRNEPSNEKSDVYSYGVVLWELMTQSIPWENLNSLQVVGVV 600
T L++KS GT +WMAPE+LRNEP+++K DVYSYGV+LWEL T PW +N +QVVG V
Sbjct: 866 TYLSSKSTAGTAEWMAPEVLRNEPADKKCDVYSYGVILWELFTLQQPWGKMNPMQVVGAV 925
Query: 601 GFMDRRLDLPEGLDPHVASIINDCWRRCFYMYSSLQLYYGRNHLLPQTESLLVLNMSSDP 660
GF RRLD+PE +DP +A II CW+ +DP
Sbjct: 926 GFQHRRLDIPEFVDPGIADIIRKCWQ-------------------------------TDP 954
Query: 661 EQRPSFEELIQRMLFVLNRVTAESVKRSS 689
RPSF E++ + + + +V SS
Sbjct: 955 RLRPSFGEIMDSLKQLQKPIQRAAVPSSS 983
>UniRef100_Q9FPR5 EDR1 [Oryza sativa]
Length = 903
Score = 321 bits (823), Expect = 4e-86
Identities = 170/353 (48%), Positives = 221/353 (62%), Gaps = 39/353 (11%)
Query: 332 DPKKLEPEATHMAIEDEVQKQQGRLQLPSSR-ESVGSHESSSSKGDN--DSVVDCEIHWE 388
D KKL P+ ++ + +PS ESV + S K D D V +CEIHWE
Sbjct: 565 DNKKLHPDPKKSPLDRFMDTS-----MPSRNPESVSPSFARSHKLDTMFDDVSECEIHWE 619
Query: 389 DLHLRDEIGQGSYAVVYHGIWNGSDVAIKVYFGNGYTEETLQDYKKEIDIMKRLRHPNVL 448
DL + + IG GSY VY WNG++VA+K + + + L +++ E+ IM+RLRHPN++
Sbjct: 620 DLVIGERIGLGSYGEVYRADWNGTEVAVKKFLDQDFYGDALDEFRSEVRIMRRLRHPNIV 679
Query: 449 LFMGAVYSLERLAIVTELLPRGSLFKTLHKSNQTLDIRRRLRMALDIARGMNYLHHRNPP 508
LFMGAV L+IV+E LPRGSL+K LH+ N +D +RR++MALD+A+GMN LH P
Sbjct: 680 LFMGAVTRPPNLSIVSEYLPRGSLYKILHRPNCQIDEKRRIKMALDVAKGMNCLHISVPT 739
Query: 509 IVHRDLKSSNLLVDKNWTVKVGDFGLSRLKDTTLLTTKSGRGTPQWMAPEILRNEPSNEK 568
IVHRDLKS NLLVD NW VKV DFGLSRLK +T L++KS GTP+WMAPE+LRNE SNEK
Sbjct: 740 IVHRDLKSPNLLVDNNWNVKVCDFGLSRLKHSTFLSSKSTAGTPEWMAPEVLRNEQSNEK 799
Query: 569 SDVYSYGVVLWELMTQSIPWENLNSLQVVGVVGFMDRRLDLPEGLDPHVASIINDCWRRC 628
DVYS+GV+LWEL T +PW +N +QVVG VGF D+RLD+P+ +DP VA II +CW++
Sbjct: 800 CDVYSFGVILWELATLRMPWSGMNPMQVVGAVGFQDKRLDIPKEIDPLVARIIWECWQK- 858
Query: 629 FYMYSSLQLYYGRNHLLPQTESLLVLNMSSDPEQRPSFEELIQRMLFVLNRVT 681
DP RPSF +L + V VT
Sbjct: 859 ------------------------------DPNLRPSFAQLTSALKTVQRLVT 881
>UniRef100_Q7XYY6 EDR1 [Oryza sativa]
Length = 1017
Score = 321 bits (823), Expect = 4e-86
Identities = 170/353 (48%), Positives = 221/353 (62%), Gaps = 39/353 (11%)
Query: 332 DPKKLEPEATHMAIEDEVQKQQGRLQLPSSR-ESVGSHESSSSKGDN--DSVVDCEIHWE 388
D KKL P+ ++ + +PS ESV + S K D D V +CEIHWE
Sbjct: 679 DNKKLHPDPKKSPLDRFMDTS-----MPSRNPESVSPSFARSHKLDTMFDDVSECEIHWE 733
Query: 389 DLHLRDEIGQGSYAVVYHGIWNGSDVAIKVYFGNGYTEETLQDYKKEIDIMKRLRHPNVL 448
DL + + IG GSY VY WNG++VA+K + + + L +++ E+ IM+RLRHPN++
Sbjct: 734 DLVIGERIGLGSYGEVYRADWNGTEVAVKKFLDQDFYGDALDEFRSEVRIMRRLRHPNIV 793
Query: 449 LFMGAVYSLERLAIVTELLPRGSLFKTLHKSNQTLDIRRRLRMALDIARGMNYLHHRNPP 508
LFMGAV L+IV+E LPRGSL+K LH+ N +D +RR++MALD+A+GMN LH P
Sbjct: 794 LFMGAVTRPPNLSIVSEYLPRGSLYKILHRPNCQIDEKRRIKMALDVAKGMNCLHISVPT 853
Query: 509 IVHRDLKSSNLLVDKNWTVKVGDFGLSRLKDTTLLTTKSGRGTPQWMAPEILRNEPSNEK 568
IVHRDLKS NLLVD NW VKV DFGLSRLK +T L++KS GTP+WMAPE+LRNE SNEK
Sbjct: 854 IVHRDLKSPNLLVDNNWNVKVCDFGLSRLKHSTFLSSKSTAGTPEWMAPEVLRNEQSNEK 913
Query: 569 SDVYSYGVVLWELMTQSIPWENLNSLQVVGVVGFMDRRLDLPEGLDPHVASIINDCWRRC 628
DVYS+GV+LWEL T +PW +N +QVVG VGF D+RLD+P+ +DP VA II +CW++
Sbjct: 914 CDVYSFGVILWELATLRMPWSGMNPMQVVGAVGFQDKRLDIPKEIDPLVARIIWECWQK- 972
Query: 629 FYMYSSLQLYYGRNHLLPQTESLLVLNMSSDPEQRPSFEELIQRMLFVLNRVT 681
DP RPSF +L + V VT
Sbjct: 973 ------------------------------DPNLRPSFAQLTSALKTVQRLVT 995
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.316 0.132 0.390
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,176,795,357
Number of Sequences: 2790947
Number of extensions: 51081177
Number of successful extensions: 190823
Number of sequences better than 10.0: 18528
Number of HSP's better than 10.0 without gapping: 11952
Number of HSP's successfully gapped in prelim test: 6576
Number of HSP's that attempted gapping in prelim test: 144985
Number of HSP's gapped (non-prelim): 23640
length of query: 697
length of database: 848,049,833
effective HSP length: 134
effective length of query: 563
effective length of database: 474,062,935
effective search space: 266897432405
effective search space used: 266897432405
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 78 (34.7 bits)
Lotus: description of TM0304.15


