

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0299.14
(88 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
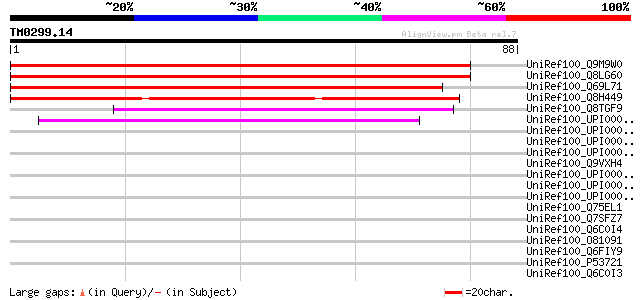
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9M9W0 F18C1.18 protein [Arabidopsis thaliana] 146 1e-34
UniRef100_Q8LG60 Hypothetical protein At5g27760 [Arabidopsis tha... 139 2e-32
UniRef100_Q69L71 Hypoxia-responsive family protein-like [Oryza s... 134 6e-31
UniRef100_Q8H449 Hypothetical protein P0470D12.148 [Oryza sativa] 118 3e-26
UniRef100_Q8TGF9 Hypothetical protein AfA14E5.14c [Aspergillus f... 47 1e-04
UniRef100_UPI000042D5C4 UPI000042D5C4 UniRef100 entry 45 4e-04
UniRef100_UPI00003C1EEF UPI00003C1EEF UniRef100 entry 44 0.001
UniRef100_UPI000042E350 UPI000042E350 UniRef100 entry 42 0.003
UniRef100_UPI0000219FA9 UPI0000219FA9 UniRef100 entry 41 0.006
UniRef100_Q9VXH4 CG9921-PA [Drosophila melanogaster] 41 0.008
UniRef100_UPI000023DD2D UPI000023DD2D UniRef100 entry 40 0.018
UniRef100_UPI0000234C72 UPI0000234C72 UniRef100 entry 39 0.032
UniRef100_UPI000023E79F UPI000023E79F UniRef100 entry 39 0.041
UniRef100_Q75EL1 AAR068Cp [Ashbya gossypii] 38 0.070
UniRef100_Q7SFZ7 Hypothetical protein [Neurospora crassa] 37 0.092
UniRef100_Q6C0I4 Similarities with sp|P53721 Saccharomyces cerev... 37 0.12
UniRef100_O81091 Aie2 protein [Oryza sativa] 37 0.16
UniRef100_Q6FIY9 Similar to sp|P53721 Saccharomyces cerevisiae Y... 35 0.35
UniRef100_P53721 Hypothetical 25.3 kDa protein in TIM23-ARE2 int... 34 0.78
UniRef100_Q6C0I3 Similar to sp|P53721 Saccharomyces cerevisiae Y... 33 1.7
>UniRef100_Q9M9W0 F18C1.18 protein [Arabidopsis thaliana]
Length = 97
Score = 146 bits (368), Expect = 1e-34
Identities = 70/80 (87%), Positives = 74/80 (92%)
Query: 1 MAEAKTQVESIRKWVVDHKLRTVGCLWLSGITGSIAYNWSRPNMKTSVKIIHARLHAQAL 60
M E+KT+ E IRKWV DHKLRTVGCLWLSGITGSIAYNWS+P MKTSVKIIHARLHAQAL
Sbjct: 1 MVESKTKFEEIRKWVSDHKLRTVGCLWLSGITGSIAYNWSQPAMKTSVKIIHARLHAQAL 60
Query: 61 TLAALAGAAVVEYYDHRAEA 80
TLAALAGAAVVEYYDH+ EA
Sbjct: 61 TLAALAGAAVVEYYDHKTEA 80
>UniRef100_Q8LG60 Hypothetical protein At5g27760 [Arabidopsis thaliana]
Length = 96
Score = 139 bits (349), Expect = 2e-32
Identities = 64/80 (80%), Positives = 74/80 (92%)
Query: 1 MAEAKTQVESIRKWVVDHKLRTVGCLWLSGITGSIAYNWSRPNMKTSVKIIHARLHAQAL 60
MAE KT+V IR+W+++HKLRTVGCLWLSGI+GSIAYNWS+P MKTSV+IIHARLHAQAL
Sbjct: 1 MAEPKTKVAEIREWIIEHKLRTVGCLWLSGISGSIAYNWSKPAMKTSVRIIHARLHAQAL 60
Query: 61 TLAALAGAAVVEYYDHRAEA 80
TLAALAGAA VEYYDH++ A
Sbjct: 61 TLAALAGAAAVEYYDHKSGA 80
>UniRef100_Q69L71 Hypoxia-responsive family protein-like [Oryza sativa]
Length = 107
Score = 134 bits (337), Expect = 6e-31
Identities = 62/75 (82%), Positives = 70/75 (92%)
Query: 1 MAEAKTQVESIRKWVVDHKLRTVGCLWLSGITGSIAYNWSRPNMKTSVKIIHARLHAQAL 60
MAEA ++ES+RKWV+DHKLR VGCLWL+GI+ SIAYNWSRPNMKTSVKIIHARLHAQAL
Sbjct: 9 MAEAPGKIESMRKWVIDHKLRAVGCLWLTGISSSIAYNWSRPNMKTSVKIIHARLHAQAL 68
Query: 61 TLAALAGAAVVEYYD 75
TLAAL G+A+VEYYD
Sbjct: 69 TLAALVGSAMVEYYD 83
>UniRef100_Q8H449 Hypothetical protein P0470D12.148 [Oryza sativa]
Length = 99
Score = 118 bits (296), Expect = 3e-26
Identities = 59/78 (75%), Positives = 69/78 (87%), Gaps = 2/78 (2%)
Query: 1 MAEAKTQVESIRKWVVDHKLRTVGCLWLSGITGSIAYNWSRPNMKTSVKIIHARLHAQAL 60
MAE K+ ++S+R+WVVDHKLR V LWL+G+ SIAYNWSRP MKTSVKIIHA LHAQAL
Sbjct: 1 MAEEKSTMQSMREWVVDHKLRAV-TLWLTGVASSIAYNWSRPGMKTSVKIIHA-LHAQAL 58
Query: 61 TLAALAGAAVVEYYDHRA 78
TLAALAG+A+VEYYDHR+
Sbjct: 59 TLAALAGSALVEYYDHRS 76
>UniRef100_Q8TGF9 Hypothetical protein AfA14E5.14c [Aspergillus fumigatus]
Length = 225
Score = 47.0 bits (110), Expect = 1e-04
Identities = 24/59 (40%), Positives = 33/59 (55%)
Query: 19 KLRTVGCLWLSGITGSIAYNWSRPNMKTSVKIIHARLHAQALTLAALAGAAVVEYYDHR 77
K + VG W++ + GS A P + S KI+ AR++AQ LTLA L +A E D R
Sbjct: 116 KYKIVGATWVASMIGSFAMVSRNPYLTGSQKIVQARVYAQGLTLAVLVASAAFEISDQR 174
>UniRef100_UPI000042D5C4 UPI000042D5C4 UniRef100 entry
Length = 226
Score = 45.1 bits (105), Expect = 4e-04
Identities = 20/66 (30%), Positives = 34/66 (51%)
Query: 6 TQVESIRKWVVDHKLRTVGCLWLSGITGSIAYNWSRPNMKTSVKIIHARLHAQALTLAAL 65
TQ+E + WV +HK + W+ + + P T+ K++ AR+ AQ LT+ L
Sbjct: 96 TQMEKAKNWVGNHKYSLISAAWVGSLGLAFGIVARNPYQTTAQKVVQARMWAQGLTVGLL 155
Query: 66 AGAAVV 71
G A++
Sbjct: 156 VGGALL 161
>UniRef100_UPI00003C1EEF UPI00003C1EEF UniRef100 entry
Length = 211
Score = 43.9 bits (102), Expect = 0.001
Identities = 21/58 (36%), Positives = 33/58 (56%), Gaps = 1/58 (1%)
Query: 14 WVVDHKLRTVGCLWLSGITGSIAYNWSRPNMKTSVKIIHARLHAQALTLAALAGAAVV 71
W ++K V W S + G AY ++P M + K++ AR+ AQ +TLA+L G A +
Sbjct: 118 WAKENKFSVVAATWASSMVGIFAYIHTQP-MSFAQKLVQARVWAQGITLASLVGMAAI 174
>UniRef100_UPI000042E350 UPI000042E350 UniRef100 entry
Length = 232
Score = 42.4 bits (98), Expect = 0.003
Identities = 21/68 (30%), Positives = 34/68 (49%)
Query: 3 EAKTQVESIRKWVVDHKLRTVGCLWLSGITGSIAYNWSRPNMKTSVKIIHARLHAQALTL 62
E +++E W HK +G W + + + P TS K++ AR+ AQ LT+
Sbjct: 101 EKMSKLEKAGDWAARHKYGIIGGGWAASMALAFGIVARNPYQSTSQKVVQARMWAQGLTV 160
Query: 63 AALAGAAV 70
A L G+A+
Sbjct: 161 ALLVGSAM 168
>UniRef100_UPI0000219FA9 UPI0000219FA9 UniRef100 entry
Length = 253
Score = 41.2 bits (95), Expect = 0.006
Identities = 21/73 (28%), Positives = 41/73 (55%)
Query: 6 TQVESIRKWVVDHKLRTVGCLWLSGITGSIAYNWSRPNMKTSVKIIHARLHAQALTLAAL 65
+Q++ ++ W +++ V WL+ + ++A + T+ K++ AR++AQ LTLA L
Sbjct: 102 SQMQRLKDWGRENRYTIVFASWLASMGVALAAVGKDKYLTTAQKLVQARVYAQGLTLAVL 161
Query: 66 AGAAVVEYYDHRA 78
AV E D ++
Sbjct: 162 IATAVFETADAKS 174
>UniRef100_Q9VXH4 CG9921-PA [Drosophila melanogaster]
Length = 102
Score = 40.8 bits (94), Expect = 0.008
Identities = 22/77 (28%), Positives = 41/77 (52%)
Query: 1 MAEAKTQVESIRKWVVDHKLRTVGCLWLSGITGSIAYNWSRPNMKTSVKIIHARLHAQAL 60
+AE +T E +++ + ++ L +GCL + + YN+ N K S ++ +R+ AQ
Sbjct: 26 VAEVETTKEKLQRKIKENPLVPLGCLATTAALTAGLYNFRTGNRKMSQLMMRSRIAAQGF 85
Query: 61 TLAALAGAAVVEYYDHR 77
T+ AL V+ Y D +
Sbjct: 86 TVMALVVGVVMTYTDKK 102
>UniRef100_UPI000023DD2D UPI000023DD2D UniRef100 entry
Length = 228
Score = 39.7 bits (91), Expect = 0.018
Identities = 25/75 (33%), Positives = 42/75 (55%), Gaps = 5/75 (6%)
Query: 3 EAKTQVESIRKWVVDHKLRTVGCLWLS--GITGSIAYNWSRPNMKTSVKIIHARLHAQAL 60
E +T E + + +++ + V W++ GI +I SR M T+ K++ AR++AQ L
Sbjct: 99 ENETAYERFKDYAKENRYQIVFASWVASMGIAFAIV---SRQPMSTANKVVQARVYAQGL 155
Query: 61 TLAALAGAAVVEYYD 75
TLA L +A+ E D
Sbjct: 156 TLAVLIVSAIYEMRD 170
>UniRef100_UPI0000234C72 UPI0000234C72 UniRef100 entry
Length = 236
Score = 38.9 bits (89), Expect = 0.032
Identities = 19/64 (29%), Positives = 31/64 (47%)
Query: 14 WVVDHKLRTVGCLWLSGITGSIAYNWSRPNMKTSVKIIHARLHAQALTLAALAGAAVVEY 73
W + + V W++ + GS A P + K++ AR++AQ TLA L +A E
Sbjct: 111 WAREERYTIVFSTWVASMIGSFALVGRNPYLSGPQKLVQARVYAQGATLAVLIASAAFEI 170
Query: 74 YDHR 77
+ R
Sbjct: 171 SERR 174
>UniRef100_UPI000023E79F UPI000023E79F UniRef100 entry
Length = 232
Score = 38.5 bits (88), Expect = 0.041
Identities = 24/73 (32%), Positives = 41/73 (55%), Gaps = 1/73 (1%)
Query: 3 EAKTQVESIRKWVVDHKLRTVGCLWLSGITGSIAYNWSRPNMKTSVKIIHARLHAQALTL 62
E +T +E ++ +++ V WL+ + + A SR M T+ K++ AR++AQ LTL
Sbjct: 99 ENQTAMERFMEYGKENRYSIVFVSWLASMGVAFAMV-SRSPMNTANKVVQARVYAQGLTL 157
Query: 63 AALAGAAVVEYYD 75
A L +A+ E D
Sbjct: 158 AVLIISAIFEVND 170
>UniRef100_Q75EL1 AAR068Cp [Ashbya gossypii]
Length = 221
Score = 37.7 bits (86), Expect = 0.070
Identities = 26/93 (27%), Positives = 46/93 (48%), Gaps = 8/93 (8%)
Query: 1 MAEAKTQVE-----SIRKWVVDHKLRTVGCLWLSGITGSIAYNWSRPNMKTSVKIIHARL 55
+AEAK E + + +++K + +G LW + + GS P M K + AR+
Sbjct: 92 LAEAKRWAELPLGDKLLESGLNNKYKIIGSLWAASMYGSWRLVDRDPIMTKVQKAVQARM 151
Query: 56 HAQALTLAALAGAAVVEYYDHRAEAKRAKNSEQ 88
+AQALT+ L + ++ Y + + +N EQ
Sbjct: 152 YAQALTVVMLLASVMLSMYGAK---RNPENKEQ 181
>UniRef100_Q7SFZ7 Hypothetical protein [Neurospora crassa]
Length = 241
Score = 37.4 bits (85), Expect = 0.092
Identities = 19/73 (26%), Positives = 36/73 (49%)
Query: 6 TQVESIRKWVVDHKLRTVGCLWLSGITGSIAYNWSRPNMKTSVKIIHARLHAQALTLAAL 65
T + +W +++ V W++ + ++A + S K++ AR++AQ LTLA L
Sbjct: 103 TSTQRFMEWGRENRYSIVFASWIAAMGAAMAIVNRNKYLSASQKLVQARVYAQGLTLAVL 162
Query: 66 AGAAVVEYYDHRA 78
A E D ++
Sbjct: 163 IATAAFETADAKS 175
>UniRef100_Q6C0I4 Similarities with sp|P53721 Saccharomyces cerevisiae YNR018w
[Yarrowia lipolytica]
Length = 279
Score = 37.0 bits (84), Expect = 0.12
Identities = 20/65 (30%), Positives = 34/65 (51%)
Query: 1 MAEAKTQVESIRKWVVDHKLRTVGCLWLSGITGSIAYNWSRPNMKTSVKIIHARLHAQAL 60
M K Q ++ +HK + V W+ + GS+ Y + M ++ KI+ AR++AQA
Sbjct: 89 MEHRKEQQAGFMHYLKEHKYKCVFGGWVVSMAGSMWYVFRDKYMTSTQKIVQARMYAQAS 148
Query: 61 TLAAL 65
T+ L
Sbjct: 149 TIILL 153
>UniRef100_O81091 Aie2 protein [Oryza sativa]
Length = 127
Score = 36.6 bits (83), Expect = 0.16
Identities = 15/39 (38%), Positives = 25/39 (63%), Gaps = 1/39 (2%)
Query: 10 SIRKWVVDHKLRTVGCLWLSGITGSIAYNWSR-PNMKTS 47
S++ WV +HKL ++G LW + + S+AY + P M+ S
Sbjct: 7 SVQSWVEEHKLASIGGLWATAVGASVAYGRRKTPQMRDS 45
>UniRef100_Q6FIY9 Similar to sp|P53721 Saccharomyces cerevisiae YNR018w [Candida
glabrata]
Length = 225
Score = 35.4 bits (80), Expect = 0.35
Identities = 18/72 (25%), Positives = 36/72 (50%)
Query: 17 DHKLRTVGCLWLSGITGSIAYNWSRPNMKTSVKIIHARLHAQALTLAALAGAAVVEYYDH 76
++K + + W + + GS P M T+ K + AR++AQ +T+A L + + Y+
Sbjct: 114 NNKYKIITGAWAASLYGSWVLVDRDPIMTTAQKAVQARMYAQFITVALLLASVGLSMYEA 173
Query: 77 RAEAKRAKNSEQ 88
+ + K E+
Sbjct: 174 KLHPDKKKQGEK 185
>UniRef100_P53721 Hypothetical 25.3 kDa protein in TIM23-ARE2 intergenic region
[Saccharomyces cerevisiae]
Length = 224
Score = 34.3 bits (77), Expect = 0.78
Identities = 17/71 (23%), Positives = 36/71 (49%)
Query: 17 DHKLRTVGCLWLSGITGSIAYNWSRPNMKTSVKIIHARLHAQALTLAALAGAAVVEYYDH 76
++K + + W + + GS P M + KI+ AR++AQ +T+ L + + Y++
Sbjct: 114 NNKYKIITGAWAASLYGSWVIVNKDPIMTKAQKIVQARMYAQFITVGLLLASVGLSMYEN 173
Query: 77 RAEAKRAKNSE 87
+ + K +E
Sbjct: 174 KLHPNKQKVNE 184
>UniRef100_Q6C0I3 Similar to sp|P53721 Saccharomyces cerevisiae YNR018w [Yarrowia
lipolytica]
Length = 202
Score = 33.1 bits (74), Expect = 1.7
Identities = 18/69 (26%), Positives = 34/69 (49%)
Query: 9 ESIRKWVVDHKLRTVGCLWLSGITGSIAYNWSRPNMKTSVKIIHARLHAQALTLAALAGA 68
+ I +V +K + + W + + GS+ M K++ AR++AQALT+ L G
Sbjct: 103 QRILYYVSQNKYKMIVGAWAASVGGSLWLINRDKIMTQPQKVVQARMYAQALTVVLLLGT 162
Query: 69 AVVEYYDHR 77
+ Y+ +
Sbjct: 163 LGLTMYEEK 171
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.318 0.127 0.376
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 128,142,941
Number of Sequences: 2790947
Number of extensions: 3624491
Number of successful extensions: 9500
Number of sequences better than 10.0: 35
Number of HSP's better than 10.0 without gapping: 19
Number of HSP's successfully gapped in prelim test: 16
Number of HSP's that attempted gapping in prelim test: 9477
Number of HSP's gapped (non-prelim): 36
length of query: 88
length of database: 848,049,833
effective HSP length: 64
effective length of query: 24
effective length of database: 669,429,225
effective search space: 16066301400
effective search space used: 16066301400
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 68 (30.8 bits)
Lotus: description of TM0299.14


