

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0296b.2
(1291 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
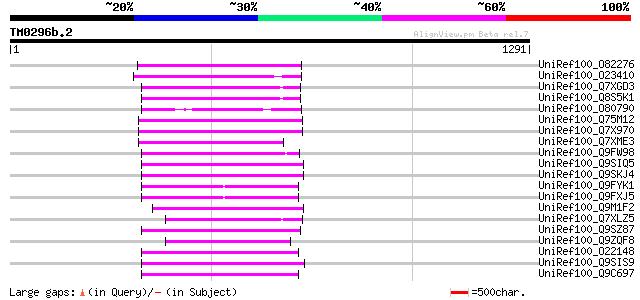
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_O82276 Putative non-LTR retroelement reverse transcrip... 327 1e-87
UniRef100_O23410 STRONG homology to reverse transcriptase [Arabi... 309 3e-82
UniRef100_Q7XGD3 Putative retroelement [Oryza sativa] 241 8e-62
UniRef100_Q8S5K1 Putative retroelement [Oryza sativa] 241 8e-62
UniRef100_O80790 Putative non-LTR retroelement reverse transcrip... 236 4e-60
UniRef100_Q75M12 Hypothetical protein P0676G05.2 [Oryza sativa] 229 4e-58
UniRef100_Q7X970 BZIP-like protein [Oryza sativa] 229 5e-58
UniRef100_Q7XME3 OSJNBa0061G20.16 protein [Oryza sativa] 228 7e-58
UniRef100_Q9FW98 Putative non-LTR retroelement reverse transcrip... 223 3e-56
UniRef100_Q9SIQ5 Putative non-LTR retroelement reverse transcrip... 223 3e-56
UniRef100_Q9SKJ4 Putative non-LTR retroelement reverse transcrip... 223 4e-56
UniRef100_Q9FYK1 F21J9.30 [Arabidopsis thaliana] 223 4e-56
UniRef100_Q9FXJ5 F5A9.24 [Arabidopsis thaliana] 223 4e-56
UniRef100_Q9M1F2 Hypothetical protein F9K21.130 [Arabidopsis tha... 222 5e-56
UniRef100_Q7XLZ5 OSJNBa0086O06.14 protein [Oryza sativa] 221 1e-55
UniRef100_Q9SZ87 RNA-directed DNA polymerase-like protein [Arabi... 220 2e-55
UniRef100_Q9ZQF8 Putative non-LTR retroelement reverse transcrip... 220 2e-55
UniRef100_O22148 Putative non-LTR retroelement reverse transcrip... 219 3e-55
UniRef100_Q9SIS9 Putative non-LTR retroelement reverse transcrip... 219 6e-55
UniRef100_Q9C697 Reverse transcriptase, putative; 16838-20266 [A... 215 8e-54
>UniRef100_O82276 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 1231
Score = 327 bits (838), Expect = 1e-87
Identities = 172/411 (41%), Positives = 249/411 (59%), Gaps = 1/411 (0%)
Query: 317 TLTNTQAVIIRKRNSVHRLKLSDGSWCSDTSKLQEEALGYFQALFAGNQEAS-QTPFPGD 375
T +T +I R+RN + LK D W +D +L+ AL Y++ L++ + + P
Sbjct: 216 TFFHTSTIIRRRRNRIESLKGDDDRWVTDKVELEAMALTYYKRLYSLEDVSEVRNMLPTG 275
Query: 376 NVPKLKEVVRFLLVQPVSKEEVRQALMSMKSFTAPGPDGFQPFFFKQYWEQVGDTVWKVV 435
+ E + L+Q +K EV A+ SM F APGPDG+QP F++Q WE VG +V + V
Sbjct: 276 GFASISEAEKAALLQAFTKAEVVSAVKSMGRFKAPGPDGYQPVFYQQCWETVGPSVTRFV 335
Query: 436 SEAFESGKVDERLLEILVVLIPKVNHPTSIKEFRPISLCNVTYKLITKLLVNRIRPFLND 495
E FE+G + + L+VLI KV P I++FRP+SLCNV +K+ITK++V R++ ++
Sbjct: 336 LEFFETGVLPASTNDALLVLIAKVAKPERIQQFRPVSLCNVLFKIITKMMVTRLKNVISK 395
Query: 496 IVGPMQNSFLPGRGTMDNAFLA*EIIHQMHRSMARKGNLAFKIDLEKAYDSISWSFLRET 555
++GP Q SF+PGR ++DN L E +H M R RKG + K+DLEKAYD + W FL+ET
Sbjct: 396 LIGPAQASFIPGRLSIDNIVLVQEAVHSMRRKKGRKGWMLLKLDLEKAYDRVRWDFLQET 455
Query: 556 LELYEFPLVIIDLIMCYVSSSQISLLWNGAPLPSFKPGRGLRQGDPKSPYFFVLCMERLS 615
LE IM V+ +S+LWNG SF P RGLRQGDP SPY FVLC+ERL
Sbjct: 456 LEAAGLSEGWTSRIMAGVTDPSMSVLWNGERTDSFVPARGLRQGDPLSPYLFVLCLERLC 515
Query: 616 IHIQNLVEPGRWKPLHRTAGGTGISHLFFAGDVLLFCKASVAQVTVLVDALQFLCEGSVL 675
I+ V WKP+ + GG+ +SH+ FA D++LF +ASVAQ+ ++ L+ CE S
Sbjct: 516 HLIEASVGKREWKPIAVSCGGSKLSHVCFADDLILFAEASVAQIRIIRRVLERFCEASGQ 575
Query: 676 RINVQKSKAITSIGVAPEVRTEIGLRALIPFVNDLGKYLGFPLQGRRLNSE 726
+++++KSK S V+ E+ I + I +LGKYLG P+ +R+N E
Sbjct: 576 KVSLEKSKIFFSHNVSREMEQLISEESGIGCTKELGKYLGMPILQKRMNKE 626
>UniRef100_O23410 STRONG homology to reverse transcriptase [Arabidopsis thaliana]
Length = 929
Score = 309 bits (792), Expect = 3e-82
Identities = 171/421 (40%), Positives = 248/421 (58%), Gaps = 18/421 (4%)
Query: 307 KLERTGSDWGTLTNTQAVIIRKRNSVHRLKLSDGSWCSDTSKLQEEALGYFQALFA-GNQ 365
KL G T +T VI R+RN + LK S+ W ++ L++ A+ Y++ L++ +
Sbjct: 158 KLLALGDRNTTFFHTSTVIRRRRNRIEMLKDSEDRWVTEKEALEKLAMDYYRKLYSLEDV 217
Query: 366 EASQTPFPGDNVPKLKEVVRFLLVQPVSKEEVRQALMSMKSFTAPGPDGFQPFFFKQYWE 425
+ P + P+L + L +P +++EV A+ SM F APGPDG+QP F++Q WE
Sbjct: 218 SVVRGTLPTEGFPRLTREEKNNLNRPFTRDEVVVAVRSMGRFKAPGPDGYQPVFYQQCWE 277
Query: 426 QVGDTVWKVVSEAFESGKVDERLLEILVVLIPKVNHPTSIKEFRPISLCNVTYKLITKLL 485
VG++V K V E FESG + + ++L+VL+ KV P I +FRP+SLCNV +K+ITK++
Sbjct: 278 TVGESVSKFVMEFFESGVLPKSTNDVLLVLLAKVAKPERITQFRPVSLCNVLFKIITKMM 337
Query: 486 VNRIRPFLNDIVGPMQNSFLPGRGTMDNAFLA*EIIHQMHRSMARKGNLAFKIDLEKAYD 545
V R++ ++ ++GP Q SF+PGR + DN + E +H M R RKG + K+DLEKAYD
Sbjct: 338 VIRLKNVISKLIGPAQASFIPGRLSFDNIVVVQEAVHSMRRKKGRKGWMLLKLDLEKAYD 397
Query: 546 SISWSFLRETLELYEFPLVIIDLIMCYVSSSQISLLWNGAPLPSFKPGRGLRQGDPKSPY 605
I W FL ETLE I IM V+ ++SLLWNG SF P RGLRQGDP SPY
Sbjct: 398 RIRWDFLAETLEAAGLSEGWIKRIMECVAGPEMSLLWNGEKTDSFTPERGLRQGDPISPY 457
Query: 606 FFVLCMERLSIHIQNLVEPGRWKPLHRTAGGTGISHLFFAGDVLLFCKASVAQVTVLVDA 665
FVLC+ERL I+ V G WK + + GG +SH+ FA D++LF +ASVAQ
Sbjct: 458 LFVLCIERLCHQIETAVGRGDWKSISISQGGPKVSHVCFADDLILFAEASVAQ------- 510
Query: 666 LQFLCEGSVLRINVQKSKAITSIGVAPEVRTEIGLRALIPFVNDLGKYLGFPLQGRRLNS 725
+++++KSK S V+ ++ I I +LGKYLG P+ +R+N
Sbjct: 511 ----------KVSLEKSKIFFSNNVSRDLEGLITAETGIGSTRELGKYLGMPVLQKRINK 560
Query: 726 E 726
+
Sbjct: 561 D 561
>UniRef100_Q7XGD3 Putative retroelement [Oryza sativa]
Length = 1694
Score = 241 bits (616), Expect = 8e-62
Identities = 138/400 (34%), Positives = 209/400 (51%), Gaps = 8/400 (2%)
Query: 328 KRNSVHRLKLSDGSWCSDTSKLQEEALGYFQALFAGNQEASQTPFPGDNVPKLKEVVRFL 387
K+N + +L+ SD + S T +L+ A YFQ LF + + PK+ + +
Sbjct: 978 KKNKITKLRDSDDTVHSTTKELERMATEYFQRLFTADPSIDHSRVTSLMKPKVTDAMNEE 1037
Query: 388 LVQPVSKEEVRQALMSMKSFTAPGPDGFQPFFFKQYWEQVGDTVWKVVSEAFESGKVDER 447
L + S+EE+ AL + APGPDGF F+++ W + D + + V E F G +
Sbjct: 1038 LCKTFSEEEIANALFQIGPLKAPGPDGFPGRFYQRNWAILKDDIVRAVQEFFSLGTMPSG 1097
Query: 448 LLEILVVLIPKVNHPTSIKEFRPISLCNVTYKLITKLLVNRIRPFLNDIVGPMQNSFLPG 507
+ E +VLIPK P +K+FRPISLCNV YK+++K LVNR+RP L+D+V Q++F+PG
Sbjct: 1098 VNETAIVLIPKTEQPQELKDFRPISLCNVVYKIVSKCLVNRLRPILDDLVSQNQSAFVPG 1157
Query: 508 RGTMDNAFLA*EIIHQMHRSM-ARKGNLAFKIDLEKAYDSISWSFLRETLELYEFPLVII 566
R DNA +A E H + R+ A+K+DL KAYD + W FL + + F +
Sbjct: 1158 RLITDNALIAFEYFHHIQRNKNPENAYSAYKLDLSKAYDRVDWEFLEQAMVKLGFAHCWV 1217
Query: 567 DLIMCYVSSSQISLLWNGAPLPSFKPGRGLRQGDPKSPYFFVLCMERLSIHIQNLVEPGR 626
IM ++ + ++ NG L +F P RGLRQGDP SP+ F+ + LS ++ V+ G
Sbjct: 1218 KWIMACITLVRYAVKLNGTLLNTFAPSRGLRQGDPLSPFLFLFVADGLSFLLEEKVDQGA 1277
Query: 627 WKPLHRTAGGTGISHLFFAGDVLLFCKASVAQVTVLVDALQFL--CEGSVLRINVQKSKA 684
P+H + HL FA D LF KA+ Q V+ + L+ C G + + SK
Sbjct: 1278 MNPIHICRHAPAVLHLLFADDTPLFFKAANDQAVVMKEVLETYASCTGQL----INPSKC 1333
Query: 685 ITSIGVAPEVRTEIGLRALIPFVNDL-GKYLGFPLQGRRL 723
G + + ++ + N KYLGFP R+
Sbjct: 1334 SIMFGQSSPTAVQNQIKQTLQISNSFEDKYLGFPTPEGRM 1373
>UniRef100_Q8S5K1 Putative retroelement [Oryza sativa]
Length = 1888
Score = 241 bits (616), Expect = 8e-62
Identities = 138/400 (34%), Positives = 209/400 (51%), Gaps = 8/400 (2%)
Query: 328 KRNSVHRLKLSDGSWCSDTSKLQEEALGYFQALFAGNQEASQTPFPGDNVPKLKEVVRFL 387
K+N + +L+ SD + S T +L+ A YFQ LF + + PK+ + +
Sbjct: 935 KKNKITKLRDSDDTVHSTTKELERMATEYFQRLFTADPSIDHSRVTSLMKPKVTDAMNEE 994
Query: 388 LVQPVSKEEVRQALMSMKSFTAPGPDGFQPFFFKQYWEQVGDTVWKVVSEAFESGKVDER 447
L + S+EE+ AL + APGPDGF F+++ W + D + + V E F G +
Sbjct: 995 LCKTFSEEEIANALFQIGPLKAPGPDGFPGRFYQRNWAILKDDIVRAVQEFFSLGTMPSG 1054
Query: 448 LLEILVVLIPKVNHPTSIKEFRPISLCNVTYKLITKLLVNRIRPFLNDIVGPMQNSFLPG 507
+ E +VLIPK P +K+FRPISLCNV YK+++K LVNR+RP L+D+V Q++F+PG
Sbjct: 1055 VNETAIVLIPKTEQPQELKDFRPISLCNVVYKIVSKCLVNRLRPILDDLVSQNQSAFVPG 1114
Query: 508 RGTMDNAFLA*EIIHQMHRSM-ARKGNLAFKIDLEKAYDSISWSFLRETLELYEFPLVII 566
R DNA +A E H + R+ A+K+DL KAYD + W FL + + F +
Sbjct: 1115 RLITDNALIAFEYFHHIQRNKNPENAYSAYKLDLSKAYDRVDWEFLEQAMVKLGFAHCWV 1174
Query: 567 DLIMCYVSSSQISLLWNGAPLPSFKPGRGLRQGDPKSPYFFVLCMERLSIHIQNLVEPGR 626
IM ++ + ++ NG L +F P RGLRQGDP SP+ F+ + LS ++ V+ G
Sbjct: 1175 KWIMACITLVRYAVKLNGTLLNTFAPSRGLRQGDPLSPFLFLFVADGLSFLLEEKVDQGA 1234
Query: 627 WKPLHRTAGGTGISHLFFAGDVLLFCKASVAQVTVLVDALQFL--CEGSVLRINVQKSKA 684
P+H + HL FA D LF KA+ Q V+ + L+ C G + + SK
Sbjct: 1235 MNPIHICRHAPAVLHLLFADDTPLFFKAANDQAVVMKEVLETYASCTGQL----INPSKC 1290
Query: 685 ITSIGVAPEVRTEIGLRALIPFVNDL-GKYLGFPLQGRRL 723
G + + ++ + N KYLGFP R+
Sbjct: 1291 SIMFGQSSPTAVQNQIKQTLQISNSFEDKYLGFPTPEGRM 1330
>UniRef100_O80790 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 970
Score = 236 bits (601), Expect = 4e-60
Identities = 145/401 (36%), Positives = 208/401 (51%), Gaps = 51/401 (12%)
Query: 327 RKRNSVHRLKLSDGSWCSDTSKLQEEALGYFQALFA-GNQEASQTPFPGDNVPKLKEVVR 385
+KRN + K D W +D +L+ AL Y+ L++ + A P L + +
Sbjct: 108 KKRNRIESPKDGDNRWMTDKVELETMALNYYTRLYSLEDVSAVWNMLPIGGFATLTDGEK 167
Query: 386 FLLVQPVSKEEVRQALMSMKSFTAPGPDGFQPFFFKQYWEQVGDTVWKVVSEAFESGKVD 445
L++P + EEV A+ M F A GP + ++A
Sbjct: 168 AALLKPFTSEEVVVAVKGMGRFKASGP-------------------YAATNDA------- 201
Query: 446 ERLLEILVVLIPKVNHPTSIKEFRPISLCNVTYKLITKLLVNRIRPFLNDIVGPMQNSFL 505
L+VLI KV P I +FRP+SLCNV +K+ITK++VNR++ + ++GP Q +F+
Sbjct: 202 ------LLVLIAKVAKPERITQFRPMSLCNVLFKIITKMIVNRLKKVILKLIGPAQANFI 255
Query: 506 PGRGTMDNAFLA*EIIHQMHRSMARKGNLAFKIDLEKAYDSISWSFLRETLELYEFPLVI 565
PGR ++DN L E +H M R RKG + K+DLEKAYD I W+FL+ETL V
Sbjct: 256 PGRLSIDNIVLVQEAVHSMRRKKGRKGWMLLKLDLEKAYDRIRWNFLQETLVATGLSEVW 315
Query: 566 IDLIMCYVSSSQISLLWNGAPLPSFKPGRGLRQGDPKSPYFFVLCMERLSIHIQNLVEPG 625
IM V+ +S+LWNG SF P RGLRQGDP SPY FVLC+ERL I+ V
Sbjct: 316 THRIMAGVTDPSMSVLWNGGKTDSFVPARGLRQGDPLSPYLFVLCLERLCHLIEASVGTW 375
Query: 626 RWKPLHRTAGGTGISHLFFAGDVLLFCKASVAQVTVLVDALQFLCEGSVLRINVQKSKAI 685
WKP+ L C+ASVAQ+ ++ L+ CE S ++++KSK +
Sbjct: 376 DWKPM------------------ALSCEASVAQILIIRRVLEQFCEASGQNVSLEKSKIV 417
Query: 686 TSIGVAPEVRTEIGLRALIPFVNDLGKYLGFPLQGRRLNSE 726
S V+ ++ I + I +LGKYLG P+ +R+N E
Sbjct: 418 FSNNVSRDMERLISGESGIGCTRELGKYLGMPILQKRMNKE 458
>UniRef100_Q75M12 Hypothetical protein P0676G05.2 [Oryza sativa]
Length = 1936
Score = 229 bits (584), Expect = 4e-58
Identities = 137/409 (33%), Positives = 214/409 (51%), Gaps = 1/409 (0%)
Query: 320 NTQAVIIRKRNSVHRLKLSDGSWCSDTSKLQEEALGYFQALFAGNQEASQTPFPGDNVPK 379
+++AV K+N + +L+ +G+ S TS L+ A YFQ ++ + + K
Sbjct: 996 HSRAVWRAKKNKISKLRDENGAIHSTTSVLETMATEYFQGVYKADPSLNPESVTRLFQEK 1055
Query: 380 LKEVVRFLLVQPVSKEEVRQALMSMKSFTAPGPDGFQPFFFKQYWEQVGDTVWKVVSEAF 439
+ + + L Q +EE+ QA+ + +P PDGF F+++ W + + V F
Sbjct: 1056 VTDAMNEKLCQEFKEEEIAQAIFQIGPLKSPRPDGFPARFYQRNWGTLKSDIILAVRNFF 1115
Query: 440 ESGKVDERLLEILVVLIPKVNHPTSIKEFRPISLCNVTYKLITKLLVNRIRPFLNDIVGP 499
+SG + + + + +VLIPK + P +K++RPISLCNV YK+++K LVNR+RP L+D+V
Sbjct: 1116 QSGLMPKGVNDTAIVLIPKKDQPIDLKDYRPISLCNVVYKVVSKCLVNRLRPILDDLVSK 1175
Query: 500 MQNSFLPGRGTMDNAFLA*EIIHQMHRS-MARKGNLAFKIDLEKAYDSISWSFLRETLEL 558
Q++F+ GR DNA LA E H + ++ A A+K+DL KAYD + W FL L
Sbjct: 1176 EQSAFIQGRMITDNALLAFECFHSIQKNKKANSAACAYKLDLSKAYDRVDWRFLELALNK 1235
Query: 559 YEFPLVIIDLIMCYVSSSQISLLWNGAPLPSFKPGRGLRQGDPKSPYFFVLCMERLSIHI 618
F + IM V++ + S+ +NG L SF P RGLRQG+P SP+ F+ + LS+ +
Sbjct: 1236 LGFAHRWVSWIMLCVTTVRYSVKFNGTLLRSFAPTRGLRQGEPLSPFLFLFVADGLSLLL 1295
Query: 619 QNLVEPGRWKPLHRTAGGTGISHLFFAGDVLLFCKASVAQVTVLVDALQFLCEGSVLRIN 678
+ V PL GIS+L FA D LLF KA + V+ + L +G+ IN
Sbjct: 1296 KEKVAQNSLTPLKICRQAPGISYLLFADDTLLFFKAEKKEAEVVKEVLTNYAQGTGQLIN 1355
Query: 679 VQKSKAITSIGVAPEVRTEIGLRALIPFVNDLGKYLGFPLQGRRLNSER 727
K + V +I + N +YLGFP R++ R
Sbjct: 1356 PAKCSILFGEASPSSVSEDIRNTLQVERDNFEDRYLGFPTPEGRMHKGR 1404
>UniRef100_Q7X970 BZIP-like protein [Oryza sativa]
Length = 2367
Score = 229 bits (583), Expect = 5e-58
Identities = 135/409 (33%), Positives = 212/409 (51%), Gaps = 1/409 (0%)
Query: 320 NTQAVIIRKRNSVHRLKLSDGSWCSDTSKLQEEALGYFQALFAGNQEASQTPFPGDNVPK 379
+++AV K+N + +L+ ++ + S T KL+ A YFQ ++ + + K
Sbjct: 1090 HSRAVWRAKKNKISKLRDANETVHSSTMKLESMATEYFQDVYTADPNLNPETVTRLIQEK 1149
Query: 380 LKEVVRFLLVQPVSKEEVRQALMSMKSFTAPGPDGFQPFFFKQYWEQVGDTVWKVVSEAF 439
+ +++ L + +++E+ QA+ + +PGPDGF F+++ W + + V F
Sbjct: 1150 VTDIMNEKLCEDFTEDEISQAIFQIGPLKSPGPDGFPARFYQRNWGTIKADIIGAVRRFF 1209
Query: 440 ESGKVDERLLEILVVLIPKVNHPTSIKEFRPISLCNVTYKLITKLLVNRIRPFLNDIVGP 499
++G + E + + +VLIPK P +++FRPISLCNV YK+++K LVNR+RP L+D+V
Sbjct: 1210 QTGLMPEGVNDTAIVLIPKKEQPVDLRDFRPISLCNVVYKVVSKCLVNRLRPILDDLVSV 1269
Query: 500 MQNSFLPGRGTMDNAFLA*EIIHQMHRS-MARKGNLAFKIDLEKAYDSISWSFLRETLEL 558
Q++F+ GR DNA LA E H M ++ A A+K+DL KAYD + W FL +
Sbjct: 1270 EQSAFVQGRMITDNALLAFECFHAMQKNKKANHAACAYKLDLSKAYDRVDWRFLEMAMNK 1329
Query: 559 YEFPLVIIDLIMCYVSSSQISLLWNGAPLPSFKPGRGLRQGDPKSPYFFVLCMERLSIHI 618
F ++ IM V+S + + +NG L SF P RGLRQGDP P+ F+ + LS+ +
Sbjct: 1330 LGFARRWVNWIMKCVTSVRYMVKFNGTLLQSFAPTRGLRQGDPLLPFLFLFVADGLSLLL 1389
Query: 619 QNLVEPGRWKPLHRTAGGTGISHLFFAGDVLLFCKASVAQVTVLVDALQFLCEGSVLRIN 678
+ V P GISHL FA D LLF KA + V+ + L G+ IN
Sbjct: 1390 KEKVAQNSLTPFKVCRAAPGISHLLFADDTLLFFKAHQREAEVVKEVLSSYAMGTGQLIN 1449
Query: 679 VQKSKAITSIGVAPEVRTEIGLRALIPFVNDLGKYLGFPLQGRRLNSER 727
K + P V I + +YLGFP R++ R
Sbjct: 1450 PAKCSILMGGASTPAVSEAISEILQVERDRFEDRYLGFPTPEGRMHKGR 1498
>UniRef100_Q7XME3 OSJNBa0061G20.16 protein [Oryza sativa]
Length = 1285
Score = 228 bits (582), Expect = 7e-58
Identities = 131/363 (36%), Positives = 199/363 (54%), Gaps = 1/363 (0%)
Query: 320 NTQAVIIRKRNSVHRLKLSDGSWCSDTSKLQEEALGYFQALFAGNQEASQTPFPGDNVPK 379
+++AV K+N + +L+ +G+ S TS L+ A YFQ ++ + + K
Sbjct: 908 HSRAVWRAKKNKISKLRDENGAIHSMTSVLETMATEYFQEVYKADPSLNPESVTRLFQEK 967
Query: 380 LKEVVRFLLVQPVSKEEVRQALMSMKSFTAPGPDGFQPFFFKQYWEQVGDTVWKVVSEAF 439
+ + + L Q +EE+ QA+ + +PGPDGF F+++ W + + V F
Sbjct: 968 VTDAMNEKLCQEFKEEEIAQAIFQIGPLKSPGPDGFPARFYQRNWGTLKSDIILAVRNFF 1027
Query: 440 ESGKVDERLLEILVVLIPKVNHPTSIKEFRPISLCNVTYKLITKLLVNRIRPFLNDIVGP 499
+SG + E + + +VLIPK + P +K++RPISLCNV YK+++K LVNR+RP L+D+V
Sbjct: 1028 QSGLMPEGVNDTAIVLIPKKDQPIDLKDYRPISLCNVVYKVVSKCLVNRLRPILDDLVSK 1087
Query: 500 MQNSFLPGRGTMDNAFLA*EIIHQMHRS-MARKGNLAFKIDLEKAYDSISWSFLRETLEL 558
Q++F+ GR DNA LA E H + ++ A A+K+DL KAYD + W FL L
Sbjct: 1088 EQSAFIQGRMITDNALLAFECFHSIQKNKKANSAACAYKLDLSKAYDRVDWRFLELALNK 1147
Query: 559 YEFPLVIIDLIMCYVSSSQISLLWNGAPLPSFKPGRGLRQGDPKSPYFFVLCMERLSIHI 618
F + IM V++ + S+ +NG L SF P RGLRQGDP SP+ F+ + LS+ +
Sbjct: 1148 LGFAHRWVSWIMSCVTTVRYSVKFNGTLLRSFAPTRGLRQGDPLSPFLFLFVADGLSLLL 1207
Query: 619 QNLVEPGRWKPLHRTAGGTGISHLFFAGDVLLFCKASVAQVTVLVDALQFLCEGSVLRIN 678
+ V PL GISHL FA D LLF KA + V+ + L +G+ IN
Sbjct: 1208 KEKVAQNSLTPLKICCQAPGISHLLFADDTLLFFKAEKKEAEVVKEVLTNYAQGTGQLIN 1267
Query: 679 VQK 681
K
Sbjct: 1268 PAK 1270
>UniRef100_Q9FW98 Putative non-LTR retroelement reverse transcriptase [Oryza sativa]
Length = 1382
Score = 223 bits (568), Expect = 3e-56
Identities = 134/395 (33%), Positives = 204/395 (50%), Gaps = 4/395 (1%)
Query: 327 RKRNSVHRLKLSDGSWCSDTSKLQEEALGYFQALFAGNQEASQTPFPGDNVPKLKEVVRF 386
R+ N + L DGS C ++ A +++ LF+ S K+ + +
Sbjct: 379 RRTNRIKELVRDDGSRCISQEGIKRMAEVFYENLFSSEPCDSMEEVLDAIPNKVGDFING 438
Query: 387 LLVQPVSKEEVRQALMSMKSFTAPGPDGFQPFFFKQYWEQVGDTVWKVVSEAFESGKVDE 446
L + + EE++ AL M S APGPDGF F++ +W + + + V ++ E
Sbjct: 439 ELGKQYTNEEIKTALFQMGSTKAPGPDGFPALFYQTHWGILEEHICNAVRGFLLGEEIPE 498
Query: 447 RLLEILVVLIPKVNHPTSIKEFRPISLCNVTYKLITKLLVNRIRPFLNDIVGPMQNSFLP 506
L + +VVLIPKVN+ + + +FRPISLCNV YK+ +K+L NR++PFL DIV Q++F+P
Sbjct: 499 GLCDSVVVLIPKVNNASHLSKFRPISLCNVLYKIASKVLANRLKPFLPDIVSEFQSAFVP 558
Query: 507 GRGTMDNAFLA*EIIHQMHRSMARKGNLAFKIDLEKAYDSISWSFLRETLELYEFPLVII 566
GR D+A +A E +H + + + A KID+ KAYD + W++L L F I
Sbjct: 559 GRLITDSALVAYECLHTIRKQHNKNPFFALKIDMMKAYDRVEWAYLSGCLSKLGFSQDWI 618
Query: 567 DLIMCYVSSSQISLLWNGAPLPSFKPGRGLRQGDPKSPYFFVLCMERLSIHIQNLVEPGR 626
+ +M VSS + ++ NG P RG+RQGDP SPY F+LC E LS + G
Sbjct: 619 NTVMRCVSSVRYAVKINGELTKPVVPSRGIRQGDPISPYLFLLCTEGLSCLLHKKEVAGE 678
Query: 627 WKPLHRTAGGTGISHLFFAGDVLLFCKASVAQVTVLVDALQFLCEGSVLRINVQKSKAIT 686
+ + G ISHL FA D + F KA V L + L+ C S +IN+ KS
Sbjct: 679 LQGIKNGRHGPPISHLLFADDSIFFAKADSRNVQALKNTLRSYCSASGQKINLHKSSIF- 737
Query: 687 SIGVAPEVRTEIGLRALIPFVNDL--GKYLGFPLQ 719
G +I +++ + N++ YLG P +
Sbjct: 738 -FGKRCPDAVKISVKSCLQVDNEVLQDSYLGMPTE 771
Score = 40.8 bits (94), Expect = 0.27
Identities = 29/107 (27%), Positives = 43/107 (40%), Gaps = 1/107 (0%)
Query: 1 VVILMEPKCQSLRVRRFWTSLGFSFAFVMEAVGFSGGIWVLTNNSSNLSFRLVDLHAQVV 60
+V L E K + + R SLGFS +F + G SGG+ + + +S R + H +
Sbjct: 36 LVFLSETKMRDKQARNLMWSLGFSGSFAVSCEGLSGGLALFWTTAYTVSLRGFNSHF-ID 94
Query: 61 TFEMWRDSLSWVCSAVYTSPIPAQRESMWRYLIRVREDVSALGSCWG 107
+ W S VY P R W L R+ + C G
Sbjct: 95 VLVSTEELPPWRISFVYGEPKRELRHFFWNLLRRLHDQWRGPWLCCG 141
>UniRef100_Q9SIQ5 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 1524
Score = 223 bits (568), Expect = 3e-56
Identities = 142/404 (35%), Positives = 202/404 (49%), Gaps = 3/404 (0%)
Query: 329 RNSVHRLKLSDGSWCSDTSKLQEEALGYFQALFAGNQ-EASQTPFPGDNVPKLKEVVRFL 387
+N V+ + G + ++ A +F +F+ N + S F D + V
Sbjct: 521 QNRVNTIMDDQGRMFTGDKEIGNHAQDFFTNIFSTNGIKVSPIDF-ADFKSTVTNTVNLD 579
Query: 388 LVQPVSKEEVRQALMSMKSFTAPGPDGFQPFFFKQYWEQVGDTVWKVVSEAFESGKVDER 447
L + S E+ A+ + APGPDG F+K W+ VG V V + FE+ +
Sbjct: 580 LTKEFSDTEIYDAICQIGDDKAPGPDGLTARFYKNCWDIVGYDVILEVKKFFETSFMKPS 639
Query: 448 LLEILVVLIPKVNHPTSIKEFRPISLCNVTYKLITKLLVNRIRPFLNDIVGPMQNSFLPG 507
+ + +IPK+ +PT++ ++RPI+LCNV YK+I+K LVNR++ LN IV Q +F+PG
Sbjct: 640 INHTNICMIPKITNPTTLSDYRPIALCNVLYKVISKCLVNRLKSHLNSIVSDSQAAFIPG 699
Query: 508 RGTMDNAFLA*EIIHQMH-RSMARKGNLAFKIDLEKAYDSISWSFLRETLELYEFPLVII 566
R DN +A E++H + R K +A K D+ KAYD + W FL T+ L+ F I
Sbjct: 700 RIINDNVMIAHEVMHSLKVRKRVSKTYMAVKTDVSKAYDRVEWDFLETTMRLFGFCNKWI 759
Query: 567 DLIMCYVSSSQISLLWNGAPLPSFKPGRGLRQGDPKSPYFFVLCMERLSIHIQNLVEPGR 626
IM V S S+L NG+P P RG+RQGDP SPY F+LC + LS I G
Sbjct: 760 GWIMAAVKSVHYSVLINGSPHGYITPTRGIRQGDPLSPYLFILCGDILSHLINGRASSGD 819
Query: 627 WKPLHRTAGGTGISHLFFAGDVLLFCKASVAQVTVLVDALQFLCEGSVLRINVQKSKAIT 686
+ + G I+HL FA D L FC+A+V L D S +INVQKS
Sbjct: 820 LRGVRIGNGAPAITHLQFADDSLFFCQANVRNCQALKDVFDVYEYYSGQKINVQKSMITF 879
Query: 687 SIGVAPEVRTEIGLRALIPFVNDLGKYLGFPLQGRRLNSERCEY 730
V ++++ IP GKYLG P Q R E EY
Sbjct: 880 GSRVYGSTQSKLKQILEIPNQGGGGKYLGLPEQFGRKKKEMFEY 923
Score = 36.6 bits (83), Expect = 5.2
Identities = 29/102 (28%), Positives = 49/102 (47%), Gaps = 12/102 (11%)
Query: 5 MEPKCQSLRVRRFWTSLGFSFAFVMEAVGFSGGIWVLTNNSSNLSF-----RLVDLHAQV 59
+E Q +V + LGF G SGG+ ++ N+ +LS RL+D H
Sbjct: 190 LETLNQCDKVCKLAYDLGFPNVITQPPNGRSGGLALMWKNNVSLSLISQDERLIDSH--- 246
Query: 60 VTFEMWRDSLSWVCSAVYTSPIPAQRESMWRYLIRVREDVSA 101
VTF ++ S+ S VY P ++R +W+ L + ++ +A
Sbjct: 247 VTF----NNKSFYLSCVYGHPTQSERHQLWQTLEHISDNRNA 284
>UniRef100_Q9SKJ4 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 1750
Score = 223 bits (567), Expect = 4e-56
Identities = 142/404 (35%), Positives = 201/404 (49%), Gaps = 3/404 (0%)
Query: 329 RNSVHRLKLSDGSWCSDTSKLQEEALGYFQALFAGNQ-EASQTPFPGDNVPKLKEVVRFL 387
+N V+ + G + ++ A +F +F+ N + S F D + V
Sbjct: 747 QNRVNTIMDDQGRMFTGDKEIGNHAQDFFTNIFSTNGIKVSPIDF-ADFKSTVTNTVNLD 805
Query: 388 LVQPVSKEEVRQALMSMKSFTAPGPDGFQPFFFKQYWEQVGDTVWKVVSEAFESGKVDER 447
L + S E+ A+ + APGPDG F+K W+ VG V V + FE+ +
Sbjct: 806 LTKEFSDTEIYDAICQIGDDKAPGPDGLTARFYKNCWDIVGYDVILEVKKFFETSFMKPS 865
Query: 448 LLEILVVLIPKVNHPTSIKEFRPISLCNVTYKLITKLLVNRIRPFLNDIVGPMQNSFLPG 507
+ + +IPK+ +PT++ ++RPI+LCNV YK+I+K LVNR++ LN IV Q +F+PG
Sbjct: 866 INHTNICMIPKITNPTTLSDYRPIALCNVLYKVISKCLVNRLKSHLNSIVSDSQAAFIPG 925
Query: 508 RGTMDNAFLA*EIIHQMH-RSMARKGNLAFKIDLEKAYDSISWSFLRETLELYEFPLVII 566
R DN +A E++H + R K +A K D+ KAYD + W FL T+ L+ F I
Sbjct: 926 RIINDNVMIAHEVMHSLKVRKRVSKTYMAVKTDVSKAYDRVEWDFLETTMRLFGFCNKWI 985
Query: 567 DLIMCYVSSSQISLLWNGAPLPSFKPGRGLRQGDPKSPYFFVLCMERLSIHIQNLVEPGR 626
IM V S S+L NG+P P RG+RQGDP SPY F+LC + LS I G
Sbjct: 986 GWIMAAVKSVHYSVLINGSPHGYITPTRGIRQGDPLSPYLFILCGDILSHLINGRASSGD 1045
Query: 627 WKPLHRTAGGTGISHLFFAGDVLLFCKASVAQVTVLVDALQFLCEGSVLRINVQKSKAIT 686
+ + G I+HL FA D L FC+A+V L D S +INVQKS
Sbjct: 1046 LRGVRIGNGAPAITHLQFADDSLFFCQANVRNCQALKDVFDVYEYYSGQKINVQKSMITF 1105
Query: 687 SIGVAPEVRTEIGLRALIPFVNDLGKYLGFPLQGRRLNSERCEY 730
V ++ + IP GKYLG P Q R E EY
Sbjct: 1106 GSRVYGSTQSRLKQILEIPNQGGGGKYLGLPEQFGRKKKEMFEY 1149
Score = 39.3 bits (90), Expect = 0.80
Identities = 30/106 (28%), Positives = 52/106 (48%), Gaps = 12/106 (11%)
Query: 1 VVILMEPKCQSLRVRRFWTSLGFSFAFVMEAVGFSGGIWVLTNNSSNLSF-----RLVDL 55
++ L+E Q +V + LGF G SGG+ ++ N+ +LS RL+D
Sbjct: 412 ILFLVETLNQCDKVCKLAYDLGFPNVITQPPNGRSGGLALMWKNNVSLSLISQDERLIDS 471
Query: 56 HAQVVTFEMWRDSLSWVCSAVYTSPIPAQRESMWRYLIRVREDVSA 101
H VTF ++ S+ S VY P ++R +W+ L + ++ +A
Sbjct: 472 H---VTF----NNKSFYLSCVYGHPTQSERHQLWQTLEHISDNRNA 510
>UniRef100_Q9FYK1 F21J9.30 [Arabidopsis thaliana]
Length = 1270
Score = 223 bits (567), Expect = 4e-56
Identities = 139/393 (35%), Positives = 205/393 (51%), Gaps = 3/393 (0%)
Query: 327 RKRNSVHRLKLSDGSWCSDTSKLQEEALGYFQALFAGNQEASQTPFPGDNVPKLKEVVRF 386
R+R + +LK +G+ + + E A YF LF + + T + VP++ EV+
Sbjct: 326 RQRKRIEKLKDVNGNMQTSEAAKGEVAAAYFGNLFKSSNPSGFTDWFSGLVPRVSEVMNE 385
Query: 387 LLVQPVSKEEVRQALMSMKSFTAPGPDGFQPFFFKQYWEQVGDTVWKVVSEAFESGKVDE 446
LV VS +E+++A+ S+K +APGPDG FF+ YW VG+ V V + F G +
Sbjct: 386 SLVGEVSAQEIKEAVFSIKPASAPGPDGMSALFFQHYWSTVGNQVTSEVKKFFADGIMPA 445
Query: 447 RLLEILVVLIPKVNHPTSIKEFRPISLCNVTYKLITKLLVNRIRPFLNDIVGPMQNSFLP 506
+ LIPK HPT + + RPISLC+V YK+I+K++ R++P+L +IV Q++F+
Sbjct: 446 EWNYTHLCLIPKTQHPTEMVDLRPISLCSVLYKIISKIMAKRLQPWLPEIVSDTQSAFVS 505
Query: 507 GRGTMDNAFLA*EIIHQM--HRSMARKGNLAFKIDLEKAYDSISWSFLRETLELYEFPLV 564
R DN +A E++H + H ++ + +A K D+ KAYD + WS+LR L F L
Sbjct: 506 ERLITDNILVAHELVHSLKVHPRISSEF-MAVKSDMSKAYDRVEWSYLRSLLLSLGFHLK 564
Query: 565 IIDLIMCYVSSSQISLLWNGAPLPSFKPGRGLRQGDPKSPYFFVLCMERLSIHIQNLVEP 624
++ IM VSS S+L N P RGLRQGDP SP+ FVLC E L+ +
Sbjct: 565 WVNWIMVCVSSVTYSVLINDCPFGLIILQRGLRQGDPLSPFLFVLCTEGLTHLLNKAQWE 624
Query: 625 GRWKPLHRTAGGTGISHLFFAGDVLLFCKASVAQVTVLVDALQFLCEGSVLRINVQKSKA 684
G + + + G + HL FA D L CKAS Q VL L+ + IN+ KS
Sbjct: 625 GALEGIQFSENGPMVHHLLFADDSLFLCKASREQSLVLQKILKVYGNATGQTINLNKSSI 684
Query: 685 ITSIGVAPEVRTEIGLRALIPFVNDLGKYLGFP 717
V +++ I I G YLG P
Sbjct: 685 TFGEKVDEQLKGTIRTCLGIFTEGGAGTYLGLP 717
>UniRef100_Q9FXJ5 F5A9.24 [Arabidopsis thaliana]
Length = 1254
Score = 223 bits (567), Expect = 4e-56
Identities = 139/393 (35%), Positives = 205/393 (51%), Gaps = 3/393 (0%)
Query: 327 RKRNSVHRLKLSDGSWCSDTSKLQEEALGYFQALFAGNQEASQTPFPGDNVPKLKEVVRF 386
R+R + +LK +G+ + + E A YF LF + + T + VP++ EV+
Sbjct: 329 RQRKRIEKLKDVNGNMQTSEAAKGEVAAAYFGNLFKSSNPSGFTDWFSGLVPRVSEVMNE 388
Query: 387 LLVQPVSKEEVRQALMSMKSFTAPGPDGFQPFFFKQYWEQVGDTVWKVVSEAFESGKVDE 446
LV VS +E+++A+ S+K +APGPDG FF+ YW VG+ V V + F G +
Sbjct: 389 SLVGEVSAQEIKEAVFSIKPASAPGPDGMSALFFQHYWSTVGNQVTSEVKKFFADGIMPA 448
Query: 447 RLLEILVVLIPKVNHPTSIKEFRPISLCNVTYKLITKLLVNRIRPFLNDIVGPMQNSFLP 506
+ LIPK HPT + + RPISLC+V YK+I+K++ R++P+L +IV Q++F+
Sbjct: 449 EWNYTHLCLIPKTQHPTEMVDLRPISLCSVLYKIISKIMAKRLQPWLPEIVSDTQSAFVS 508
Query: 507 GRGTMDNAFLA*EIIHQM--HRSMARKGNLAFKIDLEKAYDSISWSFLRETLELYEFPLV 564
R DN +A E++H + H ++ + +A K D+ KAYD + WS+LR L F L
Sbjct: 509 ERLITDNILVAHELVHSLKVHPRISSEF-MAVKSDMSKAYDRVEWSYLRSLLLSLGFHLK 567
Query: 565 IIDLIMCYVSSSQISLLWNGAPLPSFKPGRGLRQGDPKSPYFFVLCMERLSIHIQNLVEP 624
++ IM VSS S+L N P RGLRQGDP SP+ FVLC E L+ +
Sbjct: 568 WVNWIMVCVSSVTYSVLINDCPFGLIILQRGLRQGDPLSPFLFVLCTEGLTHLLNKAQWE 627
Query: 625 GRWKPLHRTAGGTGISHLFFAGDVLLFCKASVAQVTVLVDALQFLCEGSVLRINVQKSKA 684
G + + + G + HL FA D L CKAS Q VL L+ + IN+ KS
Sbjct: 628 GALEGIQFSENGPMVHHLLFADDSLFLCKASREQSLVLQKILKVYGNATGQTINLNKSSI 687
Query: 685 ITSIGVAPEVRTEIGLRALIPFVNDLGKYLGFP 717
V +++ I I G YLG P
Sbjct: 688 TFGEKVDEQLKGTIRTCLGIFTEGGAGTYLGLP 720
>UniRef100_Q9M1F2 Hypothetical protein F9K21.130 [Arabidopsis thaliana]
Length = 851
Score = 222 bits (566), Expect = 5e-56
Identities = 139/377 (36%), Positives = 192/377 (50%), Gaps = 3/377 (0%)
Query: 356 YFQALFAGNQ-EASQTPFPGDNVPKLKEVVRFLLVQPVSKEEVRQALMSMKSFTAPGPDG 414
+F +F N + S F D + ++ L Q E+ +A+ + APGPDG
Sbjct: 13 FFTDIFTTNGIQVSPIDF-ADFPSSVTNIINSELTQDFRDSEIFEAICQIGDDKAPGPDG 71
Query: 415 FQPFFFKQYWEQVGDTVWKVVSEAFESGKVDERLLEILVVLIPKVNHPTSIKEFRPISLC 474
F+KQ W+ VG+ V K V FES + + + +IPK+ +P ++ ++RPI+LC
Sbjct: 72 LTARFYKQCWDIVGNDVIKEVKLFFESSHMKTSVNHTNICMIPKIQNPQTLSDYRPIALC 131
Query: 475 NVTYKLITKLLVNRIRPFLNDIVGPMQNSFLPGRGTMDNAFLA*EIIHQMH-RSMARKGN 533
NV YK+I+K +VNR++ LN IV Q +F+PGR DN +A EI+H + R K
Sbjct: 132 NVLYKVISKCMVNRLKAHLNSIVSDSQAAFIPGRIINDNVMIAHEIMHSLKVRKRVSKTY 191
Query: 534 LAFKIDLEKAYDSISWSFLRETLELYEFPLVIIDLIMCYVSSSQISLLWNGAPLPSFKPG 593
+A K D+ KAYD + W FL T+ L+ F I IM V S S+L NG+P P
Sbjct: 192 MAVKTDVSKAYDRVEWDFLETTMRLFGFCDKWIGWIMAAVKSVHYSVLINGSPHGYISPT 251
Query: 594 RGLRQGDPKSPYFFVLCMERLSIHIQNLVEPGRWKPLHRTAGGTGISHLFFAGDVLLFCK 653
RG+RQGDP SPY F+LC + LS I+ G + + G I+HL FA D L FC+
Sbjct: 252 RGIRQGDPLSPYLFILCGDILSHLIKVKASSGDIRGVRIGNGAPAITHLQFADDSLFFCQ 311
Query: 654 ASVAQVTVLVDALQFLCEGSVLRINVQKSKAITSIGVAPEVRTEIGLRALIPFVNDLGKY 713
A+V L D S +INVQKS V +T + IP GKY
Sbjct: 312 ANVRNCQALKDVFDVYEYYSGQKINVQKSLITFGSRVYGSTQTRLKTLLNIPNQGGGGKY 371
Query: 714 LGFPLQGRRLNSERCEY 730
LG P Q R E Y
Sbjct: 372 LGLPEQFGRKKKEMFNY 388
>UniRef100_Q7XLZ5 OSJNBa0086O06.14 protein [Oryza sativa]
Length = 1205
Score = 221 bits (563), Expect = 1e-55
Identities = 129/346 (37%), Positives = 192/346 (55%), Gaps = 9/346 (2%)
Query: 387 LLVQPVSKEEVRQALMSMKSFTAPGPDGFQPFFFKQYWEQVGDTVWKVVSEAFESGKVDE 446
+L +P + EE+ AL + APGPDGF FF++ W + V + V E FE+G+ E
Sbjct: 411 MLTKPFTDEEISDALFQIGPLKAPGPDGFPARFFQRNWGVLKRDVIEGVREFFETGEWKE 470
Query: 447 RLLEILVVLIPKVNHPTSIKEFRPISLCNVTYKLITKLLVNRIRPFLNDIVGPMQNSFLP 506
+ + ++V+IPK N P +K+FRP+SLCNV YK++ K LVNR+RP L +I+ Q++F+P
Sbjct: 471 GMNDTVIVMIPKTNAPVEMKDFRPVSLCNVIYKVVAKCLVNRLRPLLQEIISETQSAFVP 530
Query: 507 GRGTMDNAFLA*EIIHQMHRSMARKGNL-AFKIDLEKAYDSISWSFLRETLELYEFPLVI 565
GR DNA +A E H +H+ + A K+DL KAYD + W FL L+ F +
Sbjct: 531 GRMITDNALVAFECFHSIHKCTRESQDFCALKLDLSKAYDRVDWGFLDGALQKLGFGNIW 590
Query: 566 IDLIMCYVSSSQISLLWNGAPLPSFKPGRGLRQGDPKSPYFFVLCMERLSIHIQNLVEPG 625
IM V+S + S+ NG L F P RGLR+GDP +PY F+ + LS +Q +
Sbjct: 591 RKWIMSCVTSVRYSVRLNGNMLEPFYPTRGLREGDPLNPYLFLFIADGLSNILQRRRDER 650
Query: 626 RWKPLHRTAGGTGISHLFFAGDVLLFCKASVAQVTVLVDALQFL--CEGSVLRINVQKSK 683
+ +PL G+SHL FA D LLF KA V Q T + +AL C G ++ K
Sbjct: 651 QIQPLKVCRSAPGVSHLLFADDSLLFFKAEVIQATRIKEALDLYERCTGQLIN---PKEC 707
Query: 684 AITSIGVAPEVRTEIGLRALIPFVNDL--GKYLGFPLQGRRLNSER 727
++ + P+ R + G++A++ K LG P R+ +E+
Sbjct: 708 SLLFSALCPQERQD-GIKAVLQVERTCFDDKCLGLPTPDGRMKAEQ 752
>UniRef100_Q9SZ87 RNA-directed DNA polymerase-like protein [Arabidopsis thaliana]
Length = 1274
Score = 220 bits (561), Expect = 2e-55
Identities = 134/398 (33%), Positives = 203/398 (50%), Gaps = 2/398 (0%)
Query: 327 RKRNSVHRLKLSDGSWCSDTSKLQEEALGYFQALFAGNQEASQTPFPGDNVPKLKEVVRF 386
R N++ ++ G + ++ YFQ +F + + P +
Sbjct: 304 RMLNNLSVIEDGSGQEFHEEEQIASTISSYFQNIFTTSNNSDLQVVQEALSPIISSHCNE 363
Query: 387 LLVQPVSKEEVRQALMSMKSFTAPGPDGFQPFFFKQYWEQVGDTVWKVVSEAFESGKVDE 446
L++ S E+++AL S+ + APGPDGF FF YW+ + V + + F +
Sbjct: 364 ELIKISSLLEIKEALFSISADKAPGPDGFSASFFHAYWDIIEADVSRDIRSFFVDSCLSP 423
Query: 447 RLLEILVVLIPKVNHPTSIKEFRPISLCNVTYKLITKLLVNRIRPFLNDIVGPMQNSFLP 506
RL E V LIPK++ P + ++RPI+LCNV YK++ K+L R++P+L++++ Q++F+P
Sbjct: 424 RLNETHVTLIPKISAPRKVSDYRPIALCNVQYKIVAKILTRRLQPWLSELISLHQSAFVP 483
Query: 507 GRGTMDNAFLA*EIIHQMHRSMARK-GNLAFKIDLEKAYDSISWSFLRETLELYEFPLVI 565
GR DN + EI+H + S A+K ++A K D+ KAYD I W+FL+E L F
Sbjct: 484 GRAIADNVLITHEILHFLRVSGAKKYCSMAIKTDMSKAYDRIKWNFLQEVLMRLGFHDKW 543
Query: 566 IDLIMCYVSSSQISLLWNGAPLPSFKPGRGLRQGDPKSPYFFVLCMERLSIHIQNLVEPG 625
I +M V + S L NG+P S P RGLRQGDP SPY F+LC E LS + E G
Sbjct: 544 IRWVMQCVCTVSYSFLINGSPQGSVVPSRGLRQGDPLSPYLFILCTEVLSGLCRKAQEKG 603
Query: 626 RWKPLHRTAGGTGISHLFFAGDVLLFCKASVAQVTVLVDALQFLCEGSVLRINVQKSKAI 685
+ G ++HL FA D + FCK + L + L+ S IN+ KS
Sbjct: 604 VMVGIRVARGSPQVNHLLFADDTMFFCKTNPTCCGALSNILKKYELASGQSINLAKSAIT 663
Query: 686 TSIGVAPEVRTEIGLRALIPFVNDLGKYLGFPLQ-GRR 722
S +++ + L I +GKYLG P GRR
Sbjct: 664 FSSKTPQDIKRRVKLSLRIDNEGGIGKYLGLPEHFGRR 701
>UniRef100_Q9ZQF8 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 1225
Score = 220 bits (561), Expect = 2e-55
Identities = 121/312 (38%), Positives = 175/312 (55%), Gaps = 1/312 (0%)
Query: 388 LVQPVSKEEVRQALMSMKSFTAPGPDGFQPFFFKQYWEQVGDTVWKVVSEAFESGKVDER 447
L + +S EEV++AL S+ APGPDG FF++ YW+ G + K+V +G DER
Sbjct: 371 LTKVISPEEVKRALFSLNPDKAPGPDGMTAFFYQHYWDLTGPDLIKLVQNFHSTGFFDER 430
Query: 448 LLEILVVLIPKVNHPTSIKEFRPISLCNVTYKLITKLLVNRIRPFLNDIVGPMQNSFLPG 507
L E + LIPK P + EFRPISLCNV+YK+I+K+L +R++ L +++ Q++F+
Sbjct: 431 LNETNICLIPKTERPRKMAEFRPISLCNVSYKVISKVLSSRLKRLLPELISETQSAFVAE 490
Query: 508 RGTMDNAFLA*EIIHQMHRSMA-RKGNLAFKIDLEKAYDSISWSFLRETLELYEFPLVII 566
R DN +A E H + + A +K +A K D+ KAYD + WSFLR + F +
Sbjct: 491 RLITDNILIAQENFHALRTNPACKKKYMAIKTDMSKAYDRVEWSFLRALMLKMGFAQKWV 550
Query: 567 DLIMCYVSSSQISLLWNGAPLPSFKPGRGLRQGDPKSPYFFVLCMERLSIHIQNLVEPGR 626
D I+ +SS +L NG+P KP RG+RQGDP SP+ F+LC E L +++ GR
Sbjct: 551 DWIIFCISSVSYKILLNGSPKGFIKPSRGIRQGDPISPFLFILCTEALVAKLKDAEWHGR 610
Query: 627 WKPLHRTAGGTGISHLFFAGDVLLFCKASVAQVTVLVDALQFLCEGSVLRINVQKSKAIT 686
+ L + SHL FA D L FCKA Q ++D L+ E S ++N KS +
Sbjct: 611 IQGLQISRASPSTSHLLFADDSLFFCKADPLQGKEIIDILRLYGEASGQQLNPDKSSVMF 670
Query: 687 SIGVAPEVRTEI 698
V +R I
Sbjct: 671 GHEVDNSIRNTI 682
>UniRef100_O22148 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 1374
Score = 219 bits (559), Expect = 3e-55
Identities = 132/393 (33%), Positives = 201/393 (50%), Gaps = 3/393 (0%)
Query: 327 RKRNSVHRLKLSDG-SWCSDTSKLQEEALGYFQALFAGNQEASQTPFPGDNVPKLKEVVR 385
R +N + +L +G W SD L A YF+ LFA + P + + +
Sbjct: 363 RAQNRIQKLIDEEGREWTSDED-LGRVAEAYFKKLFASEDVGYTVEELENLTPLVSDQMN 421
Query: 386 FLLVQPVSKEEVRQALMSMKSFTAPGPDGFQPFFFKQYWEQVGDTVWKVVSEAFESGKVD 445
L+ P++KEEV++A S+ PGPDG F ++Q+WE +GD + ++V F SG ++
Sbjct: 422 NNLLAPITKEEVQRATFSINPHKCPGPDGMNGFLYQQFWETMGDQITEMVQAFFRSGSIE 481
Query: 446 ERLLEILVVLIPKVNHPTSIKEFRPISLCNVTYKLITKLLVNRIRPFLNDIVGPMQNSFL 505
E + + + LIPK+ + +FRPISLCNV YK+I KL+ NR++ L ++ Q +F+
Sbjct: 482 EGMNKTNICLIPKILKAEKMTDFRPISLCNVIYKVIGKLMANRLKKILPSLISETQAAFV 541
Query: 506 PGRGTMDNAFLA*EIIHQM-HRSMARKGNLAFKIDLEKAYDSISWSFLRETLELYEFPLV 564
GR DN +A E++H + + + +A K D+ KAYD + W FL + + F
Sbjct: 542 KGRLISDNILIAHELLHALSSNNKCSEEFIAIKTDISKAYDRVEWPFLEKAMRGLGFADH 601
Query: 565 IIDLIMCYVSSSQISLLWNGAPLPSFKPGRGLRQGDPKSPYFFVLCMERLSIHIQNLVEP 624
I LIM V S + +L NG P P RGLRQGDP SPY FV+C E L +Q+ +
Sbjct: 602 WIRLIMECVKSVRYQVLINGTPHGEIIPSRGLRQGDPLSPYLFVICTEMLVKMLQSAEQK 661
Query: 625 GRWKPLHRTAGGTGISHLFFAGDVLLFCKASVAQVTVLVDALQFLCEGSVLRINVQKSKA 684
+ L G ISHL FA D + +CK + + ++ ++ S R+N KS
Sbjct: 662 NQITGLKVARGAPPISHLLFADDSMFYCKVNDEALGQIIRIIEEYSLASGQRVNYLKSSI 721
Query: 685 ITSIGVAPEVRTEIGLRALIPFVNDLGKYLGFP 717
++ E R + + I G YLG P
Sbjct: 722 YFGKHISEERRCLVKRKLGIEREGGEGVYLGLP 754
>UniRef100_Q9SIS9 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 1715
Score = 219 bits (557), Expect = 6e-55
Identities = 135/408 (33%), Positives = 204/408 (49%), Gaps = 1/408 (0%)
Query: 327 RKRNSVHRLKLSDGSWCSDTSKLQEEALGYFQALFAGNQEASQTPFPGDNVPKLKEVVRF 386
+ RN + + + G + + A YF LF Q + PK+ E +
Sbjct: 725 KSRNRIKAITDAQGIENFRDDTIGKVAENYFADLFTTTQTSDWEEIISGIAPKVTEQMNH 784
Query: 387 LLVQPVSKEEVRQALMSMKSFTAPGPDGFQPFFFKQYWEQVGDTVWKVVSEAFESGKVDE 446
L+Q V+ +EVR A+ ++ + APG DGF F+ W+ +G+ V +V FES +D
Sbjct: 785 ELLQSVTDQEVRDAVFAIGADRAPGFDGFTAAFYHHLWDLIGNDVCLMVRHFFESDVMDN 844
Query: 447 RLLEILVVLIPKVNHPTSIKEFRPISLCNVTYKLITKLLVNRIRPFLNDIVGPMQNSFLP 506
++ + + LIPK+ P + ++RPISLC +YK+I+K+L+ R++ L D++ Q +F+P
Sbjct: 845 QINQTQICLIPKIIDPKHMSDYRPISLCTASYKIISKILIKRLKQCLGDVISDSQAAFVP 904
Query: 507 GRGTMDNAFLA*EIIHQM-HRSMARKGNLAFKIDLEKAYDSISWSFLRETLELYEFPLVI 565
G+ DN +A E++H + R + G +A K D+ KAYD + W+FL + + F
Sbjct: 905 GQNISDNVLVAHELLHSLKSRRECQSGYVAVKTDISKAYDRVEWNFLEKVMIQLGFAPRW 964
Query: 566 IDLIMCYVSSSQISLLWNGAPLPSFKPGRGLRQGDPKSPYFFVLCMERLSIHIQNLVEPG 625
+ IM V+S +L NG+P P RG+RQGDP SPY F+ C E LS ++
Sbjct: 965 VKWIMTCVTSVSYEVLINGSPYGKIFPSRGIRQGDPLSPYLFLFCAEVLSNMLRKAEVNK 1024
Query: 626 RWKPLHRTAGGTGISHLFFAGDVLLFCKASVAQVTVLVDALQFLCEGSVLRINVQKSKAI 685
+ + T ISHL FA D L FC+AS + L + E S +IN KS I
Sbjct: 1025 QIHGMKITKDCLAISHLLFADDSLFFCRASNQNIEQLALIFKKYEEASGQKINYAKSSII 1084
Query: 686 TSIGVAPEVRTEIGLRALIPFVNDLGKYLGFPLQGRRLNSERCEYNFT 733
+ R + I V GKYLG P Q R E EY T
Sbjct: 1085 FGQKIPTMRRQRLHRLLGIDNVRGGGKYLGLPEQLGRRKVELFEYIVT 1132
Score = 43.5 bits (101), Expect = 0.042
Identities = 36/143 (25%), Positives = 60/143 (41%), Gaps = 14/143 (9%)
Query: 1 VVILMEPKCQSLRVRRFWTSLGFSFAFVMEAVGFSGGIWVLTNNSSNLSF-----RLVDL 55
++ L+E K Q R +GF ++ G SGG+ V ++ RLVDL
Sbjct: 393 MLFLIETKQQDNYTRDLGVKMGFEDMCIISPRGLSGGLVVYWKKHLSIQVISHDVRLVDL 452
Query: 56 HAQVVTFEMWRDSLSWVCSAVYTSPIPAQRESMWRYLIRVREDVSALGSCWGTLTKSYNL 115
+ + F + S +Y PIP++R +W L RV S G + NL
Sbjct: 453 YVEYKNFNFY-------LSCIYGHPIPSERHHLWEKLQRVSAHRSGPWMMCGDFNEILNL 505
Query: 116 MRSEEELSYQAEQQNLQRFLMLV 138
+E++ + +LQ F ++
Sbjct: 506 --NEKKGGRRRSIGSLQNFTNMI 526
>UniRef100_Q9C697 Reverse transcriptase, putative; 16838-20266 [Arabidopsis thaliana]
Length = 1142
Score = 215 bits (547), Expect = 8e-54
Identities = 131/392 (33%), Positives = 195/392 (49%), Gaps = 1/392 (0%)
Query: 327 RKRNSVHRLKLSDGSWCSDTSKLQEEALGYFQALFAGNQEASQTPFPGDNVPKLKEVVRF 386
R RN + L +G W + +Q A+ YFQ LF G+ + + +
Sbjct: 147 RARNRITGLHDENGIWSIEDDDIQNIAVSYFQNLFTTANPQVFDEALGEVQVLITDRIND 206
Query: 387 LLVQPVSKEEVRQALMSMKSFTAPGPDGFQPFFFKQYWEQVGDTVWKVVSEAFESGKVDE 446
LL ++ EVR AL + APGPDG FF++ W + + +V+ + G D+
Sbjct: 207 LLTADATECEVRAALFMIHPEKAPGPDGMTALFFQKSWAIIKSDLLSLVNSFLQEGVFDK 266
Query: 447 RLLEILVVLIPKVNHPTSIKEFRPISLCNVTYKLITKLLVNRIRPFLNDIVGPMQNSFLP 506
RL + LIPK PT + E RPISLCNV YK+I+K+L R++ L +++ Q++F+
Sbjct: 267 RLNTTNICLIPKTERPTRMTELRPISLCNVGYKVISKILCQRLKTVLPNLISETQSAFVD 326
Query: 507 GRGTMDNAFLA*EIIHQMHRSMARKGN-LAFKIDLEKAYDSISWSFLRETLELYEFPLVI 565
GR DN +A E+ H + + + K +A K D+ KAYD + W+F+ L F
Sbjct: 327 GRLISDNILIAQEMFHGLRTNSSCKDKFMAIKTDMSKAYDQVEWNFIEALLRKMGFCEKW 386
Query: 566 IDLIMCYVSSSQISLLWNGAPLPSFKPGRGLRQGDPKSPYFFVLCMERLSIHIQNLVEPG 625
I IM +++ Q +L NG P P RGLRQGDP SPY F+LC E L +I+
Sbjct: 387 ISWIMWCITTVQYKVLINGQPKGLIIPERGLRQGDPLSPYLFILCTEVLIANIRKAERQN 446
Query: 626 RWKPLHRTAGGTGISHLFFAGDVLLFCKASVAQVTVLVDALQFLCEGSVLRINVQKSKAI 685
+ +SHL FA D L FCKA+ Q ++++ L+ S +IN KS
Sbjct: 447 LITGIKVATPSPAVSHLLFADDSLFFCKANKEQCGIILEILKQYESVSGQQINFSKSSIQ 506
Query: 686 TSIGVAPEVRTEIGLRALIPFVNDLGKYLGFP 717
V ++ +I L I + +G YLG P
Sbjct: 507 FGHKVEDSIKADIKLILGIHNLGGMGSYLGLP 538
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.340 0.149 0.508
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,018,407,115
Number of Sequences: 2790947
Number of extensions: 81253839
Number of successful extensions: 231509
Number of sequences better than 10.0: 947
Number of HSP's better than 10.0 without gapping: 603
Number of HSP's successfully gapped in prelim test: 344
Number of HSP's that attempted gapping in prelim test: 229431
Number of HSP's gapped (non-prelim): 1301
length of query: 1291
length of database: 848,049,833
effective HSP length: 139
effective length of query: 1152
effective length of database: 460,108,200
effective search space: 530044646400
effective search space used: 530044646400
T: 11
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.9 bits)
S2: 81 (35.8 bits)
Lotus: description of TM0296b.2


