

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0283.2
(142 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
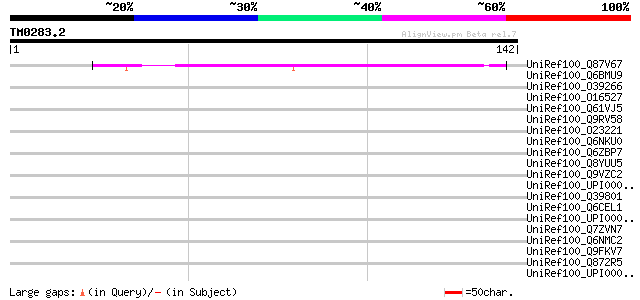
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q87V67 Amine oxidase, flavin-containing [Pseudomonas s... 44 6e-04
UniRef100_Q6BMU9 Similarities with ca|CA5907|CaSSN6 Candida albi... 39 0.019
UniRef100_O39266 24 [Equid herpesvirus 4] 39 0.024
UniRef100_O16527 Ce-LEA [Caenorhabditis elegans] 39 0.024
UniRef100_Q61VJ5 Hypothetical protein CBG04813 [Caenorhabditis b... 39 0.032
UniRef100_Q9RV58 Protein DR1172 [Deinococcus radiodurans] 39 0.032
UniRef100_O23221 Hypothetical protein AT4g36600 [Arabidopsis tha... 37 0.071
UniRef100_Q6NKU0 At4g36600 [Arabidopsis thaliana] 37 0.071
UniRef100_Q6ZBP7 Hypothetical protein P0605H02.32 [Oryza sativa] 37 0.12
UniRef100_Q8YUU5 Ferrichrome-iron receptor [Anabaena sp.] 36 0.16
UniRef100_Q9VZC2 CG15021-PA [Drosophila melanogaster] 36 0.16
UniRef100_UPI000021B378 UPI000021B378 UniRef100 entry 36 0.21
UniRef100_Q39801 51 kDa seed maturation protein precursor [Glyci... 36 0.21
UniRef100_Q6CEL1 Similarity [Yarrowia lipolytica] 36 0.21
UniRef100_UPI0000437936 UPI0000437936 UniRef100 entry 35 0.27
UniRef100_Q7ZVN7 Zgc:55839 [Brachydanio rerio] 35 0.27
UniRef100_Q6NMC2 At5g44310 [Arabidopsis thaliana] 35 0.27
UniRef100_Q9FKV7 Late embryogenesis abundant protein-like [Arabi... 35 0.27
UniRef100_Q872R5 Related to RNA-directed RNA polymerase [Neurosp... 35 0.27
UniRef100_UPI0000257CE2 UPI0000257CE2 UniRef100 entry 35 0.35
>UniRef100_Q87V67 Amine oxidase, flavin-containing [Pseudomonas syringae]
Length = 625
Score = 44.3 bits (103), Expect = 6e-04
Identities = 40/119 (33%), Positives = 54/119 (44%), Gaps = 13/119 (10%)
Query: 24 SPATSPSP-SQKSDSPPSFASWAYDKFSHVFGPYRNDEPVLPRTTENLQTGFGTGL--GA 80
+P +P+P SQKS+ + FS++FG D+P P T + + GF + L GA
Sbjct: 467 APPKAPAPASQKSEG---------NFFSNLFGGGSADKPAEPTTPQASKPGFFSRLFGGA 517
Query: 81 PVEAPAPGPFASAPQPSSEAAQQAFSYGKDKAYDIKDKAEVAYEDAKDKAESAYQSAKK 139
APAP P AP+P+ A A K V E AK A A +AKK
Sbjct: 518 QEPAPAPAPLEKAPEPAQAPAPVAAPVPVVAPVVQPAKPPVKAESAKKPAAKA-DAAKK 575
>UniRef100_Q6BMU9 Similarities with ca|CA5907|CaSSN6 Candida albicans CaSSN6
transcriptional repressor [Debaryomyces hansenii]
Length = 708
Score = 39.3 bits (90), Expect = 0.019
Identities = 32/126 (25%), Positives = 53/126 (41%), Gaps = 18/126 (14%)
Query: 19 ICRPDSPATSPSPSQKSDSPPSFASWAYDKFSHVFGPYRNDEPVLPRTTENLQTGFGTGL 78
I +P+ PA SP P PP A P + P +P+ + +Q+ T
Sbjct: 505 ISQPNQPAVSPIPPVHQ--PPPLAQ---------SNPQTSYLPSIPQPSSQIQSSTHTPN 553
Query: 79 GAPVEAPAPGPFASAPQPSSEAAQQAFSY-------GKDKAYDIKDKAEVAYEDAKDKAE 131
A V+ P+P P +S P P + Q++ S G + + K +V + +K A+
Sbjct: 554 QASVKPPSPPPQSSLPPPQNSPPQKSTSIPSIQQNNGTKRDLEDKKDTKVTKKQSKKSAK 613
Query: 132 SAYQSA 137
S S+
Sbjct: 614 SQQNSS 619
>UniRef100_O39266 24 [Equid herpesvirus 4]
Length = 3534
Score = 38.9 bits (89), Expect = 0.024
Identities = 39/130 (30%), Positives = 56/130 (43%), Gaps = 15/130 (11%)
Query: 22 PDSPATSPSPSQKSDSP-PSFASWAYDKFSHVFGPYRNDEPVLPRTTENLQTGFGTGLGA 80
P PA +P+PS+ + +P PS + A P + P ++
Sbjct: 2827 PSKPAAAPAPSKPAAAPAPSKPAAAPAPSKPAAAPAPSKPAAAPAPSK---PAAAPAPSK 2883
Query: 81 PVEAPAPGPFASAPQPSSEAAQQAFS---------YGKDKAYD-IKDKA-EVAYEDAKDK 129
P APAP A+AP PS AA A S KD+A D KD+A + A + AKD+
Sbjct: 2884 PAAAPAPSKPAAAPAPSKPAAAPAPSKPQNTLVAIVAKDQAKDQAKDQAKDQAKDQAKDQ 2943
Query: 130 AESAYQSAKK 139
A+ + K
Sbjct: 2944 AKDQAKDQAK 2953
Score = 33.1 bits (74), Expect = 1.3
Identities = 33/113 (29%), Positives = 44/113 (38%), Gaps = 6/113 (5%)
Query: 22 PDSPATSPSPSQKSDSP-PSFASWAYDKFSHVFGPYRNDEPVLPRTTENLQTGFGTGLGA 80
P PA +P+PS+ + +P PS + A P + P ++
Sbjct: 2755 PSKPAAAPAPSKPAAAPAPSKPAAAPAPSKPAAAPAPSKPAAAPAPSK---PAAAPAPSK 2811
Query: 81 PVEAPAPGPFASAPQPSSEAAQQAFSYGKDKAYDIKDKAEVAYEDAKDKAESA 133
P APAP A+AP PS AA A S K A K A +K A A
Sbjct: 2812 PAAAPAPSKPAAAPAPSKPAAAPAPS--KPAAAPAPSKPAAAPAPSKPAAAPA 2862
Score = 32.7 bits (73), Expect = 1.8
Identities = 33/113 (29%), Positives = 44/113 (38%), Gaps = 15/113 (13%)
Query: 22 PDSPATSPSPSQKSDSP-PSFASWAYDKFSHVFGPYRNDEPVLPRTTENLQTGFGTGLGA 80
P PA +P+PS+ + +P PS + A P + P ++
Sbjct: 2719 PSKPAAAPAPSKPAAAPAPSKPAAAPAPSKPAAAPAPSKPAAAPAPSK------------ 2766
Query: 81 PVEAPAPGPFASAPQPSSEAAQQAFSYGKDKAYDIKDKAEVAYEDAKDKAESA 133
P APAP A+AP PS AA A S K A K A +K A A
Sbjct: 2767 PAAAPAPSKPAAAPAPSKPAAAPAPS--KPAAAPAPSKPAAAPAPSKPAAAPA 2817
>UniRef100_O16527 Ce-LEA [Caenorhabditis elegans]
Length = 733
Score = 38.9 bits (89), Expect = 0.024
Identities = 21/48 (43%), Positives = 27/48 (55%)
Query: 91 ASAPQPSSEAAQQAFSYGKDKAYDIKDKAEVAYEDAKDKAESAYQSAK 138
A A + + A A+ KDKA D KDKA A++ KDKA A+ S K
Sbjct: 486 ADAYNSAKDKASDAWDKTKDKAGDAKDKAADAWDTTKDKAGDAWDSTK 533
Score = 37.0 bits (84), Expect = 0.093
Identities = 19/48 (39%), Positives = 29/48 (59%)
Query: 91 ASAPQPSSEAAQQAFSYGKDKAYDIKDKAEVAYEDAKDKAESAYQSAK 138
+ A + + A A+ KDKA + KDKA A+++ KDKA +A+ S K
Sbjct: 599 SDAYNSAKDKASDAWDKTKDKAGEAKDKAGDAWDNTKDKAGNAWDSTK 646
Score = 34.7 bits (78), Expect = 0.46
Identities = 20/52 (38%), Positives = 28/52 (53%)
Query: 91 ASAPQPSSEAAQQAFSYGKDKAYDIKDKAEVAYEDAKDKAESAYQSAKKTVT 142
A A + + A A+ KD A D KDKA A DAKDK++S + A ++
Sbjct: 515 ADAWDTTKDKAGDAWDSTKDHAADAKDKASDAAGDAKDKSKSLTEKAGDAIS 566
Score = 33.9 bits (76), Expect = 0.79
Identities = 16/51 (31%), Positives = 27/51 (52%)
Query: 88 GPFASAPQPSSEAAQQAFSYGKDKAYDIKDKAEVAYEDAKDKAESAYQSAK 138
G + S + +S+ A ++ D ++++KA AY AKDKA A+ K
Sbjct: 265 GAYDSVKEKASDVADSFKAHSTDSKDNVENKAADAYNTAKDKASDAWDKTK 315
Score = 33.9 bits (76), Expect = 0.79
Identities = 18/48 (37%), Positives = 26/48 (53%)
Query: 91 ASAPQPSSEAAQQAFSYGKDKAYDIKDKAEVAYEDAKDKAESAYQSAK 138
A A + + A A+ KDKA + KDK A++ KDKA A+ + K
Sbjct: 297 ADAYNTAKDKASDAWDKTKDKAGEAKDKMGDAWDTTKDKAGDAWDTTK 344
Score = 33.1 bits (74), Expect = 1.3
Identities = 17/38 (44%), Positives = 22/38 (57%)
Query: 101 AQQAFSYGKDKAYDIKDKAEVAYEDAKDKAESAYQSAK 138
A A++ KDKA D DK + DAKDKA A+ + K
Sbjct: 485 AADAYNSAKDKASDAWDKTKDKAGDAKDKAADAWDTTK 522
Score = 32.7 bits (73), Expect = 1.8
Identities = 15/51 (29%), Positives = 27/51 (52%)
Query: 88 GPFASAPQPSSEAAQQAFSYGKDKAYDIKDKAEVAYEDAKDKAESAYQSAK 138
G + + + +S+ A ++ D ++++KA AY AKDKA A+ K
Sbjct: 454 GAYDTVKEKASDIADSFKAHSTDSKDNVENKAADAYNSAKDKASDAWDKTK 504
Score = 32.3 bits (72), Expect = 2.3
Identities = 15/51 (29%), Positives = 27/51 (52%)
Query: 88 GPFASAPQPSSEAAQQAFSYGKDKAYDIKDKAEVAYEDAKDKAESAYQSAK 138
G + S + +S+ A ++ + ++++KA AY AKDKA A+ K
Sbjct: 567 GAYDSVKEKASDIADSFKAHSTNSKDNVENKASDAYNSAKDKASDAWDKTK 617
Score = 32.0 bits (71), Expect = 3.0
Identities = 17/42 (40%), Positives = 24/42 (56%)
Query: 97 SSEAAQQAFSYGKDKAYDIKDKAEVAYEDAKDKAESAYQSAK 138
+ + A A+ KDKA D K KA A++ KDKA A+ + K
Sbjct: 332 TKDKAGDAWDTTKDKAGDGKGKAGDAWDTTKDKASDAWDTTK 373
Score = 31.6 bits (70), Expect = 3.9
Identities = 16/38 (42%), Positives = 22/38 (57%)
Query: 101 AQQAFSYGKDKAYDIKDKAEVAYEDAKDKAESAYQSAK 138
A A++ KDKA D DK + +AKDKA A+ + K
Sbjct: 598 ASDAYNSAKDKASDAWDKTKDKAGEAKDKAGDAWDNTK 635
Score = 31.2 bits (69), Expect = 5.1
Identities = 18/50 (36%), Positives = 24/50 (48%)
Query: 88 GPFASAPQPSSEAAQQAFSYGKDKAYDIKDKAEVAYEDAKDKAESAYQSA 137
G A + + A A+ KDKA + KDK A++ KDKA A A
Sbjct: 352 GKAGDAWDTTKDKASDAWDTTKDKAGEAKDKMGEAWDHTKDKAGEAKDKA 401
Score = 31.2 bits (69), Expect = 5.1
Identities = 23/64 (35%), Positives = 31/64 (47%), Gaps = 7/64 (10%)
Query: 82 VEAPAPGPFASAPQPSSEA---AQQAFSYGKDKAYDI----KDKAEVAYEDAKDKAESAY 134
VE A + SA +S+A + KDKA D KDKA A++ KDKA A+
Sbjct: 594 VENKASDAYNSAKDKASDAWDKTKDKAGEAKDKAGDAWDNTKDKAGNAWDSTKDKASDAW 653
Query: 135 QSAK 138
+ K
Sbjct: 654 DTTK 657
Score = 30.4 bits (67), Expect = 8.7
Identities = 14/30 (46%), Positives = 22/30 (72%)
Query: 103 QAFSYGKDKAYDIKDKAEVAYEDAKDKAES 132
+A+ + KDKA + KDKA A +DA+ K++S
Sbjct: 385 EAWDHTKDKAGEAKDKASDAADDAQGKSKS 414
>UniRef100_Q61VJ5 Hypothetical protein CBG04813 [Caenorhabditis briggsae]
Length = 732
Score = 38.5 bits (88), Expect = 0.032
Identities = 22/58 (37%), Positives = 34/58 (57%), Gaps = 10/58 (17%)
Query: 91 ASAPQPSSEAAQQAFSYGKDKAYDIKDKAE----------VAYEDAKDKAESAYQSAK 138
A A + +AA+ A+ + K+KA ++KDKAE +E AKDKAE+A+ + K
Sbjct: 599 ADAYNKAKDAAEDAWDHTKNKAEEVKDKAEDKKDDYKERSGPWETAKDKAETAWDNTK 656
Score = 32.0 bits (71), Expect = 3.0
Identities = 17/42 (40%), Positives = 26/42 (61%)
Query: 91 ASAPQPSSEAAQQAFSYGKDKAYDIKDKAEVAYEDAKDKAES 132
A A + AA A+ KDKA ++K+KA+ +DAKD+ +S
Sbjct: 297 ADAYNSAKGAAGDAWDATKDKAGEMKEKAQDKADDAKDEGKS 338
Score = 30.4 bits (67), Expect = 8.7
Identities = 16/40 (40%), Positives = 23/40 (57%)
Query: 99 EAAQQAFSYGKDKAYDIKDKAEVAYEDAKDKAESAYQSAK 138
EA ++ D K KA AY+DAKDKA +A+++ K
Sbjct: 352 EATKERAQGAADTYNTAKYKAADAYDDAKDKAGNAWEATK 391
>UniRef100_Q9RV58 Protein DR1172 [Deinococcus radiodurans]
Length = 298
Score = 38.5 bits (88), Expect = 0.032
Identities = 20/46 (43%), Positives = 27/46 (58%)
Query: 95 QPSSEAAQQAFSYGKDKAYDIKDKAEVAYEDAKDKAESAYQSAKKT 140
Q + AQQA S KDK D+K A A + AKDKA+ Q+ K++
Sbjct: 197 QNVKQGAQQAASDAKDKVQDVKADASRAADQAKDKAQDVAQNVKQS 242
Score = 35.4 bits (80), Expect = 0.27
Identities = 18/39 (46%), Positives = 24/39 (61%)
Query: 101 AQQAFSYGKDKAYDIKDKAEVAYEDAKDKAESAYQSAKK 139
AQQA + KDK D+K A A + AKDKA+ Q+ K+
Sbjct: 163 AQQAAANVKDKVQDVKADASKAADQAKDKAQDVAQNVKQ 201
Score = 31.6 bits (70), Expect = 3.9
Identities = 16/43 (37%), Positives = 24/43 (55%)
Query: 97 SSEAAQQAFSYGKDKAYDIKDKAEVAYEDAKDKAESAYQSAKK 139
+S+AA QA +D A ++K A+ A DAKDK + A +
Sbjct: 181 ASKAADQAKDKAQDVAQNVKQGAQQAASDAKDKVQDVKADASR 223
>UniRef100_O23221 Hypothetical protein AT4g36600 [Arabidopsis thaliana]
Length = 347
Score = 37.4 bits (85), Expect = 0.071
Identities = 31/111 (27%), Positives = 48/111 (42%), Gaps = 17/111 (15%)
Query: 30 SPSQKSDSPPSFASWAYDKFSHVF-GPYRNDEPVLPRTTENLQTGFGTGLGAPVEAPAPG 88
S S+K+ FA YDK +H Y E V+ T+ + A G
Sbjct: 197 SASEKAGQAKDFA---YDKAAHAKDAAYNKAEDVIKMATDT---------SGEAKDSAYG 244
Query: 89 PFASAPQPSSEAAQQAFSYGKDKAYDIKDKAEVAYEDAKDKAESAYQSAKK 139
+ + E ++ A DKA+D+++ A A + AKDKA AY+S +
Sbjct: 245 TY----ERFKEGSKNAKDIASDKAHDVRETAGRAVDYAKDKANDAYESGSE 291
Score = 35.4 bits (80), Expect = 0.27
Identities = 20/41 (48%), Positives = 24/41 (57%)
Query: 97 SSEAAQQAFSYGKDKAYDIKDKAEVAYEDAKDKAESAYQSA 137
+ EAA+ A +Y DKA D A A + A DKA SAY SA
Sbjct: 75 AEEAAESAKNYAYDKAGSAYDNAGYAKDFASDKAGSAYDSA 115
>UniRef100_Q6NKU0 At4g36600 [Arabidopsis thaliana]
Length = 350
Score = 37.4 bits (85), Expect = 0.071
Identities = 31/111 (27%), Positives = 48/111 (42%), Gaps = 17/111 (15%)
Query: 30 SPSQKSDSPPSFASWAYDKFSHVF-GPYRNDEPVLPRTTENLQTGFGTGLGAPVEAPAPG 88
S S+K+ FA YDK +H Y E V+ T+ + A G
Sbjct: 200 SASEKAGQAKDFA---YDKAAHAKDAAYNKAEDVIKMATDT---------SGEAKDSAYG 247
Query: 89 PFASAPQPSSEAAQQAFSYGKDKAYDIKDKAEVAYEDAKDKAESAYQSAKK 139
+ + E ++ A DKA+D+++ A A + AKDKA AY+S +
Sbjct: 248 TY----ERFKEGSKNAKDIASDKAHDVRETAGRAVDYAKDKANDAYESGSE 294
Score = 35.4 bits (80), Expect = 0.27
Identities = 20/41 (48%), Positives = 24/41 (57%)
Query: 97 SSEAAQQAFSYGKDKAYDIKDKAEVAYEDAKDKAESAYQSA 137
+ EAA+ A +Y DKA D A A + A DKA SAY SA
Sbjct: 78 AEEAAESAKNYAYDKAGSAYDNAGYAKDFASDKAGSAYDSA 118
Score = 30.4 bits (67), Expect = 8.7
Identities = 34/126 (26%), Positives = 45/126 (34%), Gaps = 18/126 (14%)
Query: 28 SPSPSQKSDSPPSFASWAYDKFSHVFGPYRNDEPVLPRTTENLQTGFGTGLGAPVEAPAP 87
S + D S+A W DK S G + + + +N + A A
Sbjct: 46 SDAAHDTKDKTASWAGWVSDKISTGLGGKKAEAEEAAESAKNY--AYDKAGSAYDNAGYA 103
Query: 88 GPFASAPQPSS-EAAQQAFSYGKDKAYDIKD---------------KAEVAYEDAKDKAE 131
FAS S+ ++A A Y DKA D KD KA AYE A +
Sbjct: 104 KDFASDKAGSAYDSAHNAKHYAYDKAGDAKDMAYDKTGQAKYMAYDKAGSAYEKAGQAKD 163
Query: 132 SAYQSA 137
AY A
Sbjct: 164 MAYDKA 169
>UniRef100_Q6ZBP7 Hypothetical protein P0605H02.32 [Oryza sativa]
Length = 301
Score = 36.6 bits (83), Expect = 0.12
Identities = 33/143 (23%), Positives = 57/143 (39%), Gaps = 8/143 (5%)
Query: 1 MGNAKIVLVVLCLAVCVGICRPDSPATSPSPSQKSDSPPSFASWAYDKFSHVFGPYRNDE 60
M A + LV + LA C + + A +P+ + KS S + +S ++ S P ++ E
Sbjct: 103 MARADLALVAVLLAACAAVALA-AEAQAPAAAPKSSSSSNSSSGSHTSPSKAPSPSKSPE 161
Query: 61 P------VLPRTTENLQTGFGTGLGAPVEAPAPGPFASAPQPSSEAAQQAFSYGKDKAYD 114
P + G AP EAP A +P+ E+ + S KD +
Sbjct: 162 KSGKAPAAAPPKAAAAKAPSGKS-EAPSEAPDAESGAESPEAGEESGKSPASAPKDSSSS 220
Query: 115 IKDKAEVAYEDAKDKAESAYQSA 137
++ E + D+ D + A
Sbjct: 221 SSEEEEASSPDSGDMEDETAAEA 243
>UniRef100_Q8YUU5 Ferrichrome-iron receptor [Anabaena sp.]
Length = 858
Score = 36.2 bits (82), Expect = 0.16
Identities = 16/41 (39%), Positives = 21/41 (51%)
Query: 33 QKSDSPPSFASWAYDKFSHVFGPYRNDEPVLPRTTENLQTG 73
Q +DS +ASW +FG RN+EP P T E + G
Sbjct: 633 QPTDSTSIYASWTNSFNPQIFGKTRNNEPFKPETAEQFEVG 673
>UniRef100_Q9VZC2 CG15021-PA [Drosophila melanogaster]
Length = 420
Score = 36.2 bits (82), Expect = 0.16
Identities = 24/75 (32%), Positives = 30/75 (40%), Gaps = 11/75 (14%)
Query: 22 PDSPATSPSPSQKSDSPPSFASWAYDKFSHVFGPYRNDEPVLPRTTENLQTGFGTGLGAP 81
PD P P+PS+ PP + + +GP P P+ T G G P
Sbjct: 194 PDQPKPRPTPSRPQPPPPPPPR---PQPTPGYGPPPPPPPPKPQPTP--------GYGPP 242
Query: 82 VEAPAPGPFASAPQP 96
P PGP APQP
Sbjct: 243 TPPPGPGPAQPAPQP 257
>UniRef100_UPI000021B378 UPI000021B378 UniRef100 entry
Length = 1285
Score = 35.8 bits (81), Expect = 0.21
Identities = 29/108 (26%), Positives = 45/108 (40%), Gaps = 6/108 (5%)
Query: 22 PDSPATSPSPSQKSDSPPSFASWAYDKFSHVFGPYRNDE--PVLPRTTENLQTGFGTGLG 79
P +P +PS S+ S PP + F + P+ P L G + +
Sbjct: 268 PPAPKVAPSASKPSGPPPVAEKPTGNAFKDRIAAFNRAAAPPITPFKPSGL--GGSSFIK 325
Query: 80 APVEAPAPGPFASAPQPSSEAAQQAFSYGKDKAYDIKDKAEVAYEDAK 127
P AP P A PQP S A + Y +D+ +IK++ + E A+
Sbjct: 326 KPFVAPPPSRNAFVPQPQSAPAPRV--YRRDEDPEIKEREQENQETAE 371
>UniRef100_Q39801 51 kDa seed maturation protein precursor [Glycine max]
Length = 473
Score = 35.8 bits (81), Expect = 0.21
Identities = 27/103 (26%), Positives = 46/103 (44%), Gaps = 2/103 (1%)
Query: 36 DSPPSFASWAYDKFSHVFGPYRNDEPVLPRTTENLQTGFGTGLGAPVEAPAPGPFASAPQ 95
++ S+ WA +K S G +++D+ TT N + + T + A +
Sbjct: 61 EAAESWTEWAKEKLSEGLG-FKHDQESKESTT-NKVSDYATDTAQKSKDYATDTAQKSKD 118
Query: 96 PSSEAAQQAFSYGKDKAYDIKDKAEVAYEDAKDKAESAYQSAK 138
+ +AAQ++ Y D A KD A + +KD A A Q +K
Sbjct: 119 YAGDAAQKSKDYAGDAAQKTKDYASDTAQTSKDYAGDAAQKSK 161
>UniRef100_Q6CEL1 Similarity [Yarrowia lipolytica]
Length = 914
Score = 35.8 bits (81), Expect = 0.21
Identities = 29/115 (25%), Positives = 51/115 (44%), Gaps = 2/115 (1%)
Query: 26 ATSPSPSQKSDSPP-SFASWAYDKFSHVFGPYRNDEPVLPRTTENLQTGFGTGLGAPVEA 84
A + PS SP + A A + +HV P + T+ T T L +P A
Sbjct: 442 ARNYKPSGARRSPQITRAEMASPRSTHVTPASPRGSPAVADTSPAAPTAPATALASPASA 501
Query: 85 PAPGPFASAPQPSSEAAQQAFSYGKDKAYDIKDKAEVAYEDAKDKAESAYQSAKK 139
A P A+ QP+ +AA + ++ ++KA++ E+ + E ++A+K
Sbjct: 502 SA-SPVAAPSQPAMDAAAALLAQQEETMRQAREKAKLRKEEEQRVEEERRKAARK 555
>UniRef100_UPI0000437936 UPI0000437936 UniRef100 entry
Length = 801
Score = 35.4 bits (80), Expect = 0.27
Identities = 25/95 (26%), Positives = 36/95 (37%), Gaps = 13/95 (13%)
Query: 22 PDSPATSPSPSQKSDSPPSFASWAYDK---FSHVFGPYRNDEPVLPRTTENLQTGFGTGL 78
P P PSP S P S +S F P + P RT ++
Sbjct: 462 PPPPQPQPSPQPPSSQPNSVSSGPTPSPGGFQPSPSPQPSQSPASSRTPQSY-------- 513
Query: 79 GAPVEAPAPGPFASAPQPSSEAAQQAFSYGKDKAY 113
P++ P+PGP + PSS + S +D+ Y
Sbjct: 514 --PLQVPSPGPLNTPGNPSSVMSPAGASQSEDQLY 546
>UniRef100_Q7ZVN7 Zgc:55839 [Brachydanio rerio]
Length = 809
Score = 35.4 bits (80), Expect = 0.27
Identities = 25/95 (26%), Positives = 36/95 (37%), Gaps = 13/95 (13%)
Query: 22 PDSPATSPSPSQKSDSPPSFASWAYDK---FSHVFGPYRNDEPVLPRTTENLQTGFGTGL 78
P P PSP S P S +S F P + P RT ++
Sbjct: 470 PPPPQPQPSPQPPSSQPNSVSSGPTPSPGGFQPSPSPQPSQSPASSRTPQSY-------- 521
Query: 79 GAPVEAPAPGPFASAPQPSSEAAQQAFSYGKDKAY 113
P++ P+PGP + PSS + S +D+ Y
Sbjct: 522 --PLQVPSPGPLNTPGNPSSVMSPAGASQSEDQLY 554
>UniRef100_Q6NMC2 At5g44310 [Arabidopsis thaliana]
Length = 342
Score = 35.4 bits (80), Expect = 0.27
Identities = 21/54 (38%), Positives = 32/54 (58%), Gaps = 11/54 (20%)
Query: 97 SSEAAQQAFSY---GKDKAYDIKDKAEVAYEDAKDK--------AESAYQSAKK 139
+S AA +A+ KDKAYD+K+K + E+AKDK A+ AY++ +K
Sbjct: 119 ASRAADKAYETKEKAKDKAYDVKEKTKDYAEEAKDKVNEGASRAADKAYETKEK 172
Score = 32.7 bits (73), Expect = 1.8
Identities = 20/47 (42%), Positives = 28/47 (59%), Gaps = 4/47 (8%)
Query: 97 SSEAAQQAFSY---GKDKAYDIKDKAEVAYEDAKDKA-ESAYQSAKK 139
+S AA +A+ KDKAYD+K+K + E KDK E A ++A K
Sbjct: 199 ASRAADKAYETKEKAKDKAYDVKEKTKNYAEQTKDKVNEGASRAADK 245
Score = 32.3 bits (72), Expect = 2.3
Identities = 19/54 (35%), Positives = 31/54 (57%), Gaps = 11/54 (20%)
Query: 97 SSEAAQQAFSY---GKDKAYDIKDKAEVAYEDAKDK--------AESAYQSAKK 139
+S AA +A+ KDKAYD+K+K + E+ K+K A+ AY++ +K
Sbjct: 159 ASRAADKAYETKEKAKDKAYDVKEKTKDFAEETKEKVNEGASRAADKAYETKEK 212
>UniRef100_Q9FKV7 Late embryogenesis abundant protein-like [Arabidopsis thaliana]
Length = 331
Score = 35.4 bits (80), Expect = 0.27
Identities = 21/54 (38%), Positives = 32/54 (58%), Gaps = 11/54 (20%)
Query: 97 SSEAAQQAFSY---GKDKAYDIKDKAEVAYEDAKDK--------AESAYQSAKK 139
+S AA +A+ KDKAYD+K+K + E+AKDK A+ AY++ +K
Sbjct: 119 ASRAADKAYETKEKAKDKAYDVKEKTKDYAEEAKDKVNEGASRAADKAYETKEK 172
Score = 32.3 bits (72), Expect = 2.3
Identities = 20/55 (36%), Positives = 30/55 (54%), Gaps = 11/55 (20%)
Query: 97 SSEAAQQAFSY---GKDKAYDIKDKAEVAYEDAKDK--------AESAYQSAKKT 140
+S AA +A+ KDKAYD+K+K + E+ K+K A+ AY +KT
Sbjct: 159 ASRAADKAYETKEKAKDKAYDVKEKTKDFAEETKEKVNEGASRAADKAYDVKEKT 213
Score = 31.6 bits (70), Expect = 3.9
Identities = 17/46 (36%), Positives = 25/46 (53%), Gaps = 1/46 (2%)
Query: 95 QPSSEAAQQAFSYGKDKAYDIKDKAEVAYEDAKDKA-ESAYQSAKK 139
+ + E + S DKAYD+K+K + E KDK E A ++A K
Sbjct: 189 EETKEKVNEGASRAADKAYDVKEKTKNYAEQTKDKVNEGASRAADK 234
>UniRef100_Q872R5 Related to RNA-directed RNA polymerase [Neurospora crassa]
Length = 1351
Score = 35.4 bits (80), Expect = 0.27
Identities = 29/97 (29%), Positives = 41/97 (41%), Gaps = 3/97 (3%)
Query: 18 GICRPDSPATSPSPSQKSDSPPSFASWAYDK-FSHVFGPYRNDEPVLPRTTENLQTGFGT 76
G+ P S + +PS S+ PS S D F+ GPY +P L +T +
Sbjct: 1254 GLGSPSSTSAPTTPSSGSNYTPSSGSDMRDGIFNSPTGPYAGYQPELVHAQAGARTAGTS 1313
Query: 77 GLGAPVEAPAPGPFASAPQPSSEAAQQAFSYGKDKAY 113
G P APA P A P+ + A A Y + + Y
Sbjct: 1314 GGQVPSAAPANMPVRLA--PADKVAHLAALYAQMQIY 1348
>UniRef100_UPI0000257CE2 UPI0000257CE2 UniRef100 entry
Length = 1584
Score = 35.0 bits (79), Expect = 0.35
Identities = 27/85 (31%), Positives = 35/85 (40%), Gaps = 2/85 (2%)
Query: 26 ATSPSPSQKSDSPPSFASWAYDKFSH--VFGPYRNDEPVLPRTTENLQTGFGTGLGAPVE 83
A SP+ S S P+ ++ A + V P P P T TG G+ +P
Sbjct: 867 AASPTASTSGSSTPAASTSAPVNVTSGSVAAPSAAPAPTYPTTPATPSTGTAFGIFSPSG 926
Query: 84 APAPGPFASAPQPSSEAAQQAFSYG 108
A P AS P P + AA A S G
Sbjct: 927 TTASQPAASLPVPPAPAATTATSSG 951
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.311 0.128 0.379
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 263,767,248
Number of Sequences: 2790947
Number of extensions: 12314057
Number of successful extensions: 54702
Number of sequences better than 10.0: 368
Number of HSP's better than 10.0 without gapping: 86
Number of HSP's successfully gapped in prelim test: 290
Number of HSP's that attempted gapping in prelim test: 53697
Number of HSP's gapped (non-prelim): 1034
length of query: 142
length of database: 848,049,833
effective HSP length: 118
effective length of query: 24
effective length of database: 518,718,087
effective search space: 12449234088
effective search space used: 12449234088
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 67 (30.4 bits)
Lotus: description of TM0283.2


