

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0283.14.1
(63 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
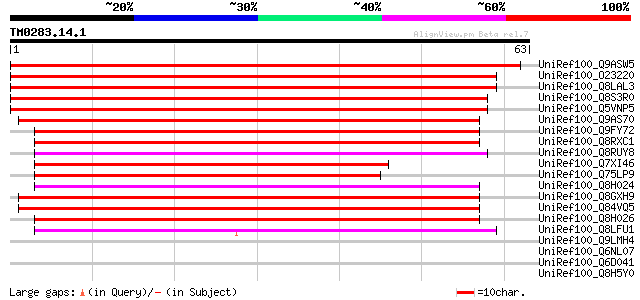
Sequences producing significant alignments: (bits) Value
UniRef100_Q9ASW5 At2g18360/T30D6.13 [Arabidopsis thaliana] 99 3e-20
UniRef100_O23220 Hypothetical protein C7A10.750 [Arabidopsis tha... 94 9e-19
UniRef100_Q8LAL3 Putative hydrolase [Arabidopsis thaliana] 94 9e-19
UniRef100_Q8S3R0 Putative hydrolase [Oryza sativa] 85 4e-16
UniRef100_Q5VNP5 Hydrolase-like [Oryza sativa] 85 4e-16
UniRef100_Q9AS70 P0028E10.22 protein [Oryza sativa] 53 2e-06
UniRef100_Q9FY72 Putative hydrolase [Arabidopsis thaliana] 52 5e-06
UniRef100_Q8RXC1 Hypothetical protein At1g78210 [Arabidopsis tha... 50 1e-05
UniRef100_Q8RUY8 Putative hydrolase [Oryza sativa] 48 6e-05
UniRef100_Q7XI46 Hydrolase-like protein [Oryza sativa] 47 1e-04
UniRef100_Q75LP9 Putative hydrolase [Oryza sativa] 47 1e-04
UniRef100_Q8H024 Hypothetical protein OSJNBa0050H14.6 [Oryza sat... 47 1e-04
UniRef100_Q8GXH9 Hypothetical protein At5g21950/T6G21_60 [Arabid... 47 2e-04
UniRef100_Q84VQ5 Hydrolase, alpha/beta fold family [Arabidopsis ... 47 2e-04
UniRef100_Q8H026 Hypothetical protein OSJNBa0050H14.4 [Oryza sat... 45 5e-04
UniRef100_Q8LFU1 Putative hydrolase [Arabidopsis thaliana] 45 5e-04
UniRef100_Q9LMH4 F16A14.4 [Arabidopsis thaliana] 42 0.003
UniRef100_Q6NL07 At1g13820 [Arabidopsis thaliana] 42 0.003
UniRef100_Q6D041 Putative hydrolase [Erwinia carotovora] 42 0.004
UniRef100_Q8H5Y0 Hydrolase-like protein [Oryza sativa] 41 0.009
>UniRef100_Q9ASW5 At2g18360/T30D6.13 [Arabidopsis thaliana]
Length = 313
Score = 99.0 bits (245), Expect = 3e-20
Identities = 44/62 (70%), Positives = 53/62 (84%)
Query: 1 RIHLLWGENDQIFKRELAKNMKEQLGDGATFQGIKKAGHLVHLERPCVYNRCLKRFIASF 60
+IHLLWGE+DQIF E AK+MKEQLG+ AT + IKKAGHL HLERPCVYNR LK+F+AS
Sbjct: 250 KIHLLWGESDQIFNLEFAKSMKEQLGENATMESIKKAGHLAHLERPCVYNRRLKKFLASV 309
Query: 61 FA 62
++
Sbjct: 310 YS 311
>UniRef100_O23220 Hypothetical protein C7A10.750 [Arabidopsis thaliana]
Length = 317
Score = 94.0 bits (232), Expect = 9e-19
Identities = 42/59 (71%), Positives = 51/59 (86%)
Query: 1 RIHLLWGENDQIFKRELAKNMKEQLGDGATFQGIKKAGHLVHLERPCVYNRCLKRFIAS 59
+IH LWGE+DQIF ELA++MKEQ+G+ AT + IKKAGHLV LERPCVYNR LK+F+AS
Sbjct: 248 KIHFLWGESDQIFDLELARDMKEQIGENATIESIKKAGHLVQLERPCVYNRRLKKFLAS 306
>UniRef100_Q8LAL3 Putative hydrolase [Arabidopsis thaliana]
Length = 317
Score = 94.0 bits (232), Expect = 9e-19
Identities = 42/59 (71%), Positives = 51/59 (86%)
Query: 1 RIHLLWGENDQIFKRELAKNMKEQLGDGATFQGIKKAGHLVHLERPCVYNRCLKRFIAS 59
+IH LWGE+DQIF ELA++MKEQ+G+ AT + IKKAGHLV LERPCVYNR LK+F+AS
Sbjct: 248 KIHFLWGESDQIFDLELARDMKEQIGENATIESIKKAGHLVQLERPCVYNRRLKKFLAS 306
>UniRef100_Q8S3R0 Putative hydrolase [Oryza sativa]
Length = 339
Score = 85.1 bits (209), Expect = 4e-16
Identities = 38/58 (65%), Positives = 46/58 (78%)
Query: 1 RIHLLWGENDQIFKRELAKNMKEQLGDGATFQGIKKAGHLVHLERPCVYNRCLKRFIA 58
+I L+WGE D+IF ELAK MKEQLGDG GI KAGHL+H+ERPC YNR L+RF++
Sbjct: 267 KIMLIWGEEDKIFDIELAKKMKEQLGDGCFLHGIPKAGHLLHVERPCAYNRQLQRFLS 324
>UniRef100_Q5VNP5 Hydrolase-like [Oryza sativa]
Length = 337
Score = 85.1 bits (209), Expect = 4e-16
Identities = 40/58 (68%), Positives = 45/58 (76%)
Query: 1 RIHLLWGENDQIFKRELAKNMKEQLGDGATFQGIKKAGHLVHLERPCVYNRCLKRFIA 58
+I LLWGEND IF ELA MKEQLG+ A Q I KAGHLVH+ERPCVYN+ LK F+A
Sbjct: 269 KILLLWGENDNIFNIELAMTMKEQLGEKAMLQSISKAGHLVHIERPCVYNQHLKEFLA 326
>UniRef100_Q9AS70 P0028E10.22 protein [Oryza sativa]
Length = 336
Score = 53.1 bits (126), Expect = 2e-06
Identities = 26/56 (46%), Positives = 35/56 (62%)
Query: 2 IHLLWGENDQIFKRELAKNMKEQLGDGATFQGIKKAGHLVHLERPCVYNRCLKRFI 57
I ++WGE DQIF E A +KE LG+ AT + I GHL H E P ++N L +F+
Sbjct: 270 ILIIWGEFDQIFPVEKAHKVKEMLGEKATVKIIPNTGHLAHQEDPKMFNDILLKFL 325
>UniRef100_Q9FY72 Putative hydrolase [Arabidopsis thaliana]
Length = 303
Score = 51.6 bits (122), Expect = 5e-06
Identities = 24/54 (44%), Positives = 34/54 (62%)
Query: 4 LLWGENDQIFKRELAKNMKEQLGDGATFQGIKKAGHLVHLERPCVYNRCLKRFI 57
++WGE DQIF EL +K +G+ A IKKAGH V+LE+ + + LK F+
Sbjct: 246 IIWGEEDQIFPLELGYRLKRHIGESAEIVVIKKAGHAVNLEKSKEFVKHLKSFL 299
>UniRef100_Q8RXC1 Hypothetical protein At1g78210 [Arabidopsis thaliana]
Length = 314
Score = 50.4 bits (119), Expect = 1e-05
Identities = 19/54 (35%), Positives = 35/54 (64%)
Query: 4 LLWGENDQIFKRELAKNMKEQLGDGATFQGIKKAGHLVHLERPCVYNRCLKRFI 57
++WGE+DQ+F E+ K +++ +GD IK+ GH+ + E+P + + LK F+
Sbjct: 248 IIWGEHDQVFPLEMGKRLEKHVGDNGKLVIIKRTGHIFNFEKPKKFIKLLKSFL 301
>UniRef100_Q8RUY8 Putative hydrolase [Oryza sativa]
Length = 401
Score = 48.1 bits (113), Expect = 6e-05
Identities = 21/55 (38%), Positives = 32/55 (58%)
Query: 4 LLWGENDQIFKRELAKNMKEQLGDGATFQGIKKAGHLVHLERPCVYNRCLKRFIA 58
++WGE D++F EL +K LGD + +K AGH ++ E+P R +K IA
Sbjct: 275 IIWGEQDRVFPLELGLRLKRHLGDTSELVIVKNAGHAINREKPAELCRLIKNCIA 329
>UniRef100_Q7XI46 Hydrolase-like protein [Oryza sativa]
Length = 327
Score = 47.4 bits (111), Expect = 1e-04
Identities = 19/43 (44%), Positives = 29/43 (67%)
Query: 4 LLWGENDQIFKRELAKNMKEQLGDGATFQGIKKAGHLVHLERP 46
++WGE D++F ELA +K LG+ + I+ AGH V+LE+P
Sbjct: 261 IVWGERDKVFPMELAHRLKRHLGESSRLVVIRNAGHAVNLEKP 303
>UniRef100_Q75LP9 Putative hydrolase [Oryza sativa]
Length = 333
Score = 47.4 bits (111), Expect = 1e-04
Identities = 20/42 (47%), Positives = 29/42 (68%)
Query: 4 LLWGENDQIFKRELAKNMKEQLGDGATFQGIKKAGHLVHLER 45
++WGE DQ+F ELA ++ LG+ + IKKAGH V+LE+
Sbjct: 268 IIWGEQDQVFPMELAHRLERHLGEKSRLVVIKKAGHAVNLEK 309
>UniRef100_Q8H024 Hypothetical protein OSJNBa0050H14.6 [Oryza sativa]
Length = 383
Score = 47.0 bits (110), Expect = 1e-04
Identities = 21/54 (38%), Positives = 32/54 (58%)
Query: 4 LLWGENDQIFKRELAKNMKEQLGDGATFQGIKKAGHLVHLERPCVYNRCLKRFI 57
+LWG+ DQ+F +L ++ LGD + + IK AGH + LE NR +K F+
Sbjct: 327 ILWGDKDQVFPLDLGHRLQRHLGDVSRLEIIKDAGHALQLEGADQVNRFIKSFL 380
>UniRef100_Q8GXH9 Hypothetical protein At5g21950/T6G21_60 [Arabidopsis thaliana]
Length = 204
Score = 46.6 bits (109), Expect = 2e-04
Identities = 20/56 (35%), Positives = 34/56 (60%)
Query: 2 IHLLWGENDQIFKRELAKNMKEQLGDGATFQGIKKAGHLVHLERPCVYNRCLKRFI 57
+ L+WGE DQ+F ++A ++KE LG AT + I+K H+ E+ +N + F+
Sbjct: 142 VMLIWGEQDQVFPLKMAHDLKEMLGTKATLKVIQKTSHIPQTEKSKEFNGFVMSFL 197
>UniRef100_Q84VQ5 Hydrolase, alpha/beta fold family [Arabidopsis thaliana]
Length = 300
Score = 46.6 bits (109), Expect = 2e-04
Identities = 20/56 (35%), Positives = 34/56 (60%)
Query: 2 IHLLWGENDQIFKRELAKNMKEQLGDGATFQGIKKAGHLVHLERPCVYNRCLKRFI 57
+ L+WGE DQ+F ++A ++KE LG AT + I+K H+ E+ +N + F+
Sbjct: 238 VMLIWGEQDQVFPLKMAHDLKEMLGTKATLKVIQKTSHIPQTEKSKEFNGFVMSFL 293
>UniRef100_Q8H026 Hypothetical protein OSJNBa0050H14.4 [Oryza sativa]
Length = 317
Score = 45.1 bits (105), Expect = 5e-04
Identities = 21/54 (38%), Positives = 33/54 (60%)
Query: 4 LLWGENDQIFKRELAKNMKEQLGDGATFQGIKKAGHLVHLERPCVYNRCLKRFI 57
L+WG++DQIF + A +K LGD + IKK GH+ +E P +N+ + F+
Sbjct: 253 LIWGDHDQIFPLDKAFAVKSCLGDHVRLEIIKKTGHVPQMEDPDRFNKIVLDFL 306
>UniRef100_Q8LFU1 Putative hydrolase [Arabidopsis thaliana]
Length = 328
Score = 45.1 bits (105), Expect = 5e-04
Identities = 22/57 (38%), Positives = 34/57 (59%), Gaps = 1/57 (1%)
Query: 4 LLWGENDQIFKRELAKNMKEQLG-DGATFQGIKKAGHLVHLERPCVYNRCLKRFIAS 59
++WGE DQ+F ELA +K LG D A +KK GH ++ E+P + +K F+ +
Sbjct: 243 MIWGEEDQVFPVELAHRLKRYLGEDRAQLVLLKKTGHAINEEKPKEMYKHMKSFLCT 299
>UniRef100_Q9LMH4 F16A14.4 [Arabidopsis thaliana]
Length = 633
Score = 42.4 bits (98), Expect = 0.003
Identities = 19/54 (35%), Positives = 31/54 (57%), Gaps = 1/54 (1%)
Query: 4 LLWGENDQIFKRELAKNMKEQLGDGATFQGIKKAGHLVHLERPCVYNRCLKRFI 57
+LWGE+DQI +LA + +L + A + I GHL H+E+P + + F+
Sbjct: 568 ILWGEDDQIISNKLAWRLHGELSN-ARVKQISNCGHLPHVEKPAAVTKLIAEFV 620
>UniRef100_Q6NL07 At1g13820 [Arabidopsis thaliana]
Length = 339
Score = 42.4 bits (98), Expect = 0.003
Identities = 19/54 (35%), Positives = 31/54 (57%), Gaps = 1/54 (1%)
Query: 4 LLWGENDQIFKRELAKNMKEQLGDGATFQGIKKAGHLVHLERPCVYNRCLKRFI 57
+LWGE+DQI +LA + +L + A + I GHL H+E+P + + F+
Sbjct: 274 ILWGEDDQIISNKLAWRLHGELSN-ARVKQISNCGHLPHVEKPAAVTKLIAEFV 326
>UniRef100_Q6D041 Putative hydrolase [Erwinia carotovora]
Length = 279
Score = 42.0 bits (97), Expect = 0.004
Identities = 23/59 (38%), Positives = 34/59 (56%), Gaps = 6/59 (10%)
Query: 2 IHLLWGENDQIFKRELAKNMKEQLGDGATFQGIKKAGHLVHLERP-----CVYNRCLKR 55
+++LWGE D+ E+ K + +LGD A Q +K AGHLV + P +Y+ LKR
Sbjct: 215 VNMLWGEQDEWIPLEVGKRLAHRLGD-APLQIVKNAGHLVQEDAPEAIVQSMYSLLLKR 272
>UniRef100_Q8H5Y0 Hydrolase-like protein [Oryza sativa]
Length = 336
Score = 40.8 bits (94), Expect = 0.009
Identities = 23/54 (42%), Positives = 30/54 (54%), Gaps = 1/54 (1%)
Query: 4 LLWGENDQIFKRELAKNMKEQLGDGATFQGIKKAGHLVHLERPCVYNRCLKRFI 57
+LWGE+DQIF E A + QLG A + IK GH+ E P +N L F+
Sbjct: 276 VLWGEHDQIFPIEKAFEVA-QLGANARLEIIKNTGHMPQEEDPKRFNEALLNFL 328
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.328 0.142 0.447
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 104,463,826
Number of Sequences: 2790947
Number of extensions: 3108821
Number of successful extensions: 9170
Number of sequences better than 10.0: 156
Number of HSP's better than 10.0 without gapping: 48
Number of HSP's successfully gapped in prelim test: 108
Number of HSP's that attempted gapping in prelim test: 9100
Number of HSP's gapped (non-prelim): 156
length of query: 63
length of database: 848,049,833
effective HSP length: 39
effective length of query: 24
effective length of database: 739,202,900
effective search space: 17740869600
effective search space used: 17740869600
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 68 (30.8 bits)
Lotus: description of TM0283.14.1


