

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0278.5.1
(30 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
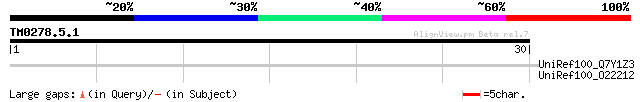
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q7Y1Z3 Putative small nuclear ribonucleoprotein Prp4p ... 36 0.25
UniRef100_O22212 Hypothetical Trp-Asp repeats containing protein... 36 0.25
>UniRef100_Q7Y1Z3 Putative small nuclear ribonucleoprotein Prp4p [Arabidopsis
thaliana]
Length = 554
Score = 36.2 bits (82), Expect = 0.25
Identities = 17/30 (56%), Positives = 22/30 (72%), Gaps = 3/30 (10%)
Query: 4 IVTVSHDRTIKLWSSNMTNDNE---HTMDV 30
I TVSHDRTIKLW+S+ +D + TMD+
Sbjct: 523 IATVSHDRTIKLWTSSGNDDEDEEKETMDI 552
>UniRef100_O22212 Hypothetical Trp-Asp repeats containing protein At2g41500
[Arabidopsis thaliana]
Length = 554
Score = 36.2 bits (82), Expect = 0.25
Identities = 17/30 (56%), Positives = 22/30 (72%), Gaps = 3/30 (10%)
Query: 4 IVTVSHDRTIKLWSSNMTNDNE---HTMDV 30
I TVSHDRTIKLW+S+ +D + TMD+
Sbjct: 523 IATVSHDRTIKLWTSSGNDDEDEEKETMDI 552
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.313 0.128 0.388
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 53,362,297
Number of Sequences: 2790947
Number of extensions: 802904
Number of successful extensions: 5750
Number of sequences better than 10.0: 2
Number of HSP's better than 10.0 without gapping: 2
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 5742
Number of HSP's gapped (non-prelim): 8
length of query: 30
length of database: 848,049,833
effective HSP length: 6
effective length of query: 24
effective length of database: 831,304,151
effective search space: 19951299624
effective search space used: 19951299624
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 69 (31.2 bits)
Lotus: description of TM0278.5.1


