

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0265.5
(404 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
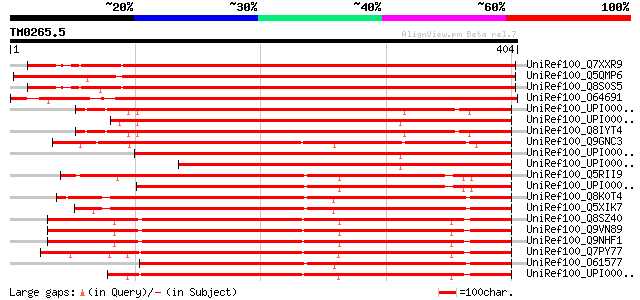
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q7XXR9 Katanin [Oryza sativa] 581 e-164
UniRef100_Q5QMP6 Vacuolar protein sorting factor 4B-like [Oryza ... 579 e-164
UniRef100_Q8S0S5 Katanin p60 subunit A 1-like [Oryza sativa] 572 e-162
UniRef100_O64691 Putative katanin [Arabidopsis thaliana] 555 e-157
UniRef100_UPI000036A419 UPI000036A419 UniRef100 entry 360 4e-98
UniRef100_UPI00003AAC53 UPI00003AAC53 UniRef100 entry 360 5e-98
UniRef100_Q8IYT4 Similar to mouse 4933439B08Rik protein [Homo sa... 360 5e-98
UniRef100_Q9GNC3 Probable AAA ATPase [Leishmania major] 344 3e-93
UniRef100_UPI0000360C40 UPI0000360C40 UniRef100 entry 329 7e-89
UniRef100_UPI000024AF1B UPI000024AF1B UniRef100 entry 301 2e-80
UniRef100_Q5RII9 Novel protein similar to vertebrate katanin p60... 289 8e-77
UniRef100_UPI0000437799 UPI0000437799 UniRef100 entry 289 1e-76
UniRef100_Q8K0T4 Katanin p60 subunit A-like 1 [Mus musculus] 287 4e-76
UniRef100_Q5XIK7 Hypothetical protein MGC94518 [Rattus norvegicus] 287 4e-76
UniRef100_Q8SZ40 RE17942p [Drosophila melanogaster] 285 2e-75
UniRef100_Q9VN89 CG10229-PA [Drosophila melanogaster] 284 4e-75
UniRef100_Q9NHF1 Putative microtubule severing protein katanin p... 284 4e-75
UniRef100_Q7PY77 ENSANGP00000018492 [Anopheles gambiae str. PEST] 282 1e-74
UniRef100_O61577 Katanin p60 subunit [Strongylocentrotus purpura... 282 1e-74
UniRef100_UPI0000244E4F UPI0000244E4F UniRef100 entry 281 2e-74
>UniRef100_Q7XXR9 Katanin [Oryza sativa]
Length = 386
Score = 581 bits (1498), Expect = e-164
Identities = 302/390 (77%), Positives = 334/390 (85%), Gaps = 11/390 (2%)
Query: 15 DFKLYYDSKFGRKKVVQNGENADKAVGNGSSMSVVSNGNVHSKRSSDMAIYEQLRSQG-Q 73
DFK +++S+FG KK + +N NG V+NG+V KR+SD+A+YEQ Q Q
Sbjct: 2 DFKGFWESRFGGKKEQEPEQNGH---ANG-----VANGSVR-KRTSDLAVYEQFEQQARQ 52
Query: 74 NGIHTNDVSPNNMDERPQKSLLPPFESAEMRTLAESLSRDIIRGSPDVKWESIKGLENAK 133
+ + N D QK LLP FESAEMR LAE+L RDIIRGSPDVKWESIKGLENAK
Sbjct: 53 TEVRAAAIRDGNADAI-QKPLLPSFESAEMRNLAETLLRDIIRGSPDVKWESIKGLENAK 111
Query: 134 RLLKEAVVMPIKYPKYFTGLLSPWKGILLFGPPGTGKTMLAKAVATECKTTFFNISASSV 193
RLLKEAVVMPIKYPKYFTGLLSPWKGILLFGPPGTGKTMLAKAVATECKTTFFNISASS+
Sbjct: 112 RLLKEAVVMPIKYPKYFTGLLSPWKGILLFGPPGTGKTMLAKAVATECKTTFFNISASSI 171
Query: 194 VSKWRGDSEKLVKVLFQLARHHAPSTIFLDEIDAIISQRGEARSEHEASRRLKTELLIQM 253
VSKWRGDSEKLVKVLF+LARHHAPSTIFLDEIDAIISQRGEARSEHEASRRLKTELLIQM
Sbjct: 172 VSKWRGDSEKLVKVLFELARHHAPSTIFLDEIDAIISQRGEARSEHEASRRLKTELLIQM 231
Query: 254 DGLTRTDELVFVLAATNLPWELDAAMLRRLEKRILVPLPEPEARVAMFEELLPPQPDEES 313
DGLT+T++LVFVLAATNLPWELDAAMLRRLEKRILVPLPE EAR AMFEELLP +
Sbjct: 232 DGLTKTNDLVFVLAATNLPWELDAAMLRRLEKRILVPLPEAEARHAMFEELLPSTTSKLE 291
Query: 314 IPYDLLVNQTEGYSGSDIRLLCKEVAMQPLRRLMSQLEQREDLVPEEELPKVGPIRPEDI 373
+PYD LV +TEGYSGSDIRL+CKE AMQPLRRLMS LE R++LVPEEELP+VGP++PEDI
Sbjct: 292 VPYDTLVEKTEGYSGSDIRLVCKEAAMQPLRRLMSVLEARDELVPEEELPEVGPLKPEDI 351
Query: 374 QAALKNTRPSAHLHAHKYDKFNADYGSQIL 403
+ AL+NTRPSAHLHAH+Y+KFN DYGSQIL
Sbjct: 352 EVALRNTRPSAHLHAHRYEKFNQDYGSQIL 381
>UniRef100_Q5QMP6 Vacuolar protein sorting factor 4B-like [Oryza sativa]
Length = 410
Score = 579 bits (1492), Expect = e-164
Identities = 297/409 (72%), Positives = 337/409 (81%), Gaps = 13/409 (3%)
Query: 4 EEPMPTRWSFQDFKLYYDSKFGRKKVVQNGENADKAVGNGSSMSVVSNGNVHSKRSS--- 60
+EP TRW+F+DF++YY+ + G ++ E+ D G G + + +G+ S R S
Sbjct: 5 DEPSITRWTFEDFEVYYEVRLGIRREPGGDEDGDGDGGGGRGYAPLGSGSAGSTRPSAAH 64
Query: 61 -----DMAIYEQL-RSQGQNGIHTNDVSPNNMDERPQKSLLPPFESAEMRTLAESLSRDI 114
D+A++EQ R + + + + PQKSLLP FESAEMR LAE+L RDI
Sbjct: 65 ANGGADLAVFEQFERLERKVELRNGAIEAGP----PQKSLLPSFESAEMRNLAETLLRDI 120
Query: 115 IRGSPDVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGLLSPWKGILLFGPPGTGKTMLA 174
IRGSPDVKWESIKGLENAKRLLKEAVVMPIKYPKYF GLLSPWKGILLFGPPGTGKTMLA
Sbjct: 121 IRGSPDVKWESIKGLENAKRLLKEAVVMPIKYPKYFKGLLSPWKGILLFGPPGTGKTMLA 180
Query: 175 KAVATECKTTFFNISASSVVSKWRGDSEKLVKVLFQLARHHAPSTIFLDEIDAIISQRGE 234
KAVATECKTTFFNISASS+VSKWRGDSEKLVKVLF+LARHHAPSTIFLDEIDAIISQRGE
Sbjct: 181 KAVATECKTTFFNISASSIVSKWRGDSEKLVKVLFELARHHAPSTIFLDEIDAIISQRGE 240
Query: 235 ARSEHEASRRLKTELLIQMDGLTRTDELVFVLAATNLPWELDAAMLRRLEKRILVPLPEP 294
ARSEHEASRRLKTELLIQMDGLT+TD+LVFVLAATNLPWELDAAMLRRLEKRILVPLPE
Sbjct: 241 ARSEHEASRRLKTELLIQMDGLTKTDDLVFVLAATNLPWELDAAMLRRLEKRILVPLPEQ 300
Query: 295 EARVAMFEELLPPQPDEESIPYDLLVNQTEGYSGSDIRLLCKEVAMQPLRRLMSQLEQRE 354
EAR AMFEELLP P +IPYD+LV +TEGYSGSDIRL+CKE AMQPLRRLMS LE R+
Sbjct: 301 EARHAMFEELLPSVPGTMNIPYDVLVEKTEGYSGSDIRLVCKEAAMQPLRRLMSVLEGRQ 360
Query: 355 DLVPEEELPKVGPIRPEDIQAALKNTRPSAHLHAHKYDKFNADYGSQIL 403
+ VPE+ELP+VGP+ EDI+ AL+NTRPSAHLH H+Y+KFN DYGS +L
Sbjct: 361 EEVPEDELPEVGPVTTEDIELALRNTRPSAHLHVHRYEKFNQDYGSHVL 409
>UniRef100_Q8S0S5 Katanin p60 subunit A 1-like [Oryza sativa]
Length = 428
Score = 572 bits (1474), Expect = e-162
Identities = 302/412 (73%), Positives = 334/412 (80%), Gaps = 33/412 (8%)
Query: 15 DFKLYYDSKFGRKKVVQNGENADKAVGNGSSMSVVSNGNVHSKRSSDMAIYEQLRSQ--- 71
DFK +++S+FG KK + +N NG V+NG+V KR+SD+A+YEQ Q
Sbjct: 22 DFKGFWESRFGGKKEQEPEQNGH---ANG-----VANGSVR-KRTSDLAVYEQFEQQVFL 72
Query: 72 --------------------GQNGIHTNDVSPNNMDERPQKSLLPPFESAEMRTLAESLS 111
Q + + N D QK LLP FESAEMR LAE+L
Sbjct: 73 DRRLVSCVVLDPRSVLLFQARQTEVRAAAIRDGNADAI-QKPLLPSFESAEMRNLAETLL 131
Query: 112 RDIIRGSPDVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGLLSPWKGILLFGPPGTGKT 171
RDIIRGSPDVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGLLSPWKGILLFGPPGTGKT
Sbjct: 132 RDIIRGSPDVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGLLSPWKGILLFGPPGTGKT 191
Query: 172 MLAKAVATECKTTFFNISASSVVSKWRGDSEKLVKVLFQLARHHAPSTIFLDEIDAIISQ 231
MLAKAVATECKTTFFNISASS+VSKWRGDSEKLVKVLF+LARHHAPSTIFLDEIDAIISQ
Sbjct: 192 MLAKAVATECKTTFFNISASSIVSKWRGDSEKLVKVLFELARHHAPSTIFLDEIDAIISQ 251
Query: 232 RGEARSEHEASRRLKTELLIQMDGLTRTDELVFVLAATNLPWELDAAMLRRLEKRILVPL 291
RGEARSEHEASRRLKTELLIQMDGLT+T++LVFVLAATNLPWELDAAMLRRLEKRILVPL
Sbjct: 252 RGEARSEHEASRRLKTELLIQMDGLTKTNDLVFVLAATNLPWELDAAMLRRLEKRILVPL 311
Query: 292 PEPEARVAMFEELLPPQPDEESIPYDLLVNQTEGYSGSDIRLLCKEVAMQPLRRLMSQLE 351
PE EAR AMFEELLP + +PYD LV +TEGYSGSDIRL+CKE AMQPLRRLMS LE
Sbjct: 312 PEAEARHAMFEELLPSTTSKLEVPYDTLVEKTEGYSGSDIRLVCKEAAMQPLRRLMSVLE 371
Query: 352 QREDLVPEEELPKVGPIRPEDIQAALKNTRPSAHLHAHKYDKFNADYGSQIL 403
R++LVPEEELP+VGP++PEDI+ AL+NTRPSAHLHAH+Y+KFN DYGSQIL
Sbjct: 372 ARDELVPEEELPEVGPLKPEDIEVALRNTRPSAHLHAHRYEKFNQDYGSQIL 423
>UniRef100_O64691 Putative katanin [Arabidopsis thaliana]
Length = 384
Score = 555 bits (1431), Expect = e-157
Identities = 289/409 (70%), Positives = 329/409 (79%), Gaps = 30/409 (7%)
Query: 1 MADEEPMPTRWSFQDFKLYYDSKFGRKKV----VQNGENADKAVGNGSSMSVVSNGNVHS 56
MA +EP TRWSF FGRKK+ V N + + NG++ V +N + +
Sbjct: 1 MATDEPSQTRWSFL---------FGRKKLPEEDVSNKDQPEDGSSNGNNGDVNNNSSPVT 51
Query: 57 KRSSDMAIYEQLRSQGQNGIHTNDVSPNNMDERPQKSLLPPFESAEMRTLAESLSRDIIR 116
+ + A+ NG N + E+P+KS+ PPFESAE RTLAESLSRDIIR
Sbjct: 52 NQDGNTAL--------ANG--------NVIREKPKKSMFPPFESAETRTLAESLSRDIIR 95
Query: 117 GSPDVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGLLSPWKGILLFGPPGTGKTMLAKA 176
G+P++KWESIKGLENAK+LLKEAVVMPIKYP YF GLL+PWKGILLFGPPGTGKTMLAKA
Sbjct: 96 GNPNIKWESIKGLENAKKLLKEAVVMPIKYPTYFNGLLTPWKGILLFGPPGTGKTMLAKA 155
Query: 177 VATECKTTFFNISASSVVSKWRGDSEKLVKVLFQLARHHAPSTIFLDEIDAIISQRG-EA 235
VATEC TTFFNISASSVVSKWRGDSEKL++VLF LARHHAPSTIFLDEIDAIISQRG E
Sbjct: 156 VATECNTTFFNISASSVVSKWRGDSEKLIRVLFDLARHHAPSTIFLDEIDAIISQRGGEG 215
Query: 236 RSEHEASRRLKTELLIQMDGLTRTDELVFVLAATNLPWELDAAMLRRLEKRILVPLPEPE 295
RSEHEASRRLKTELLIQMDGL +T+ELVFVLAATNLPWELDAAMLRRLEKRILVPLP+PE
Sbjct: 216 RSEHEASRRLKTELLIQMDGLQKTNELVFVLAATNLPWELDAAMLRRLEKRILVPLPDPE 275
Query: 296 ARVAMFEELLPPQPDEESIPYDLLVNQTEGYSGSDIRLLCKEVAMQPLRRLMSQLEQRED 355
AR MFE L+P QP +E +P+D+LV ++EGYSGSDIR+LCKE AMQPLRR ++ LE RED
Sbjct: 276 ARRGMFEMLIPSQPGDEPLPHDVLVEKSEGYSGSDIRILCKEAAMQPLRRTLAILEDRED 335
Query: 356 LVPEEELPKVGPIRPEDIQAALKNTRPSAHLHAHKYDKFNADYGSQILQ 404
+VPE+ELPK+GPI PEDI AL NTRPSAHLHAH YDKFN DYGSQIL+
Sbjct: 336 VVPEDELPKIGPILPEDIDRALSNTRPSAHLHAHLYDKFNDDYGSQILK 384
>UniRef100_UPI000036A419 UPI000036A419 UniRef100 entry
Length = 426
Score = 360 bits (924), Expect = 4e-98
Identities = 194/364 (53%), Positives = 256/364 (70%), Gaps = 20/364 (5%)
Query: 53 NVHSKRSSDMAIYEQLRSQGQNGIHTNDVSPNNMDERPQKS--LLPPFES-----AEMRT 105
N H +R + ++ L + G T++++ N D P S LL P + +EMR
Sbjct: 66 NAHPRRGQ-IIDFQGLLTDAIKGA-TSELALNTFDHNPDPSERLLKPLSAFIGMNSEMRE 123
Query: 106 LAESLSRDIIRGSPDVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGLLSPWKGILLFGP 165
LA +SRDI +P++KW I GL+ AK+L+KEAVV PI+YP+ FTG+LSPWKG+LL+GP
Sbjct: 124 LAAMVSRDIYLHNPNIKWNDIIGLDAAKQLVKEAVVYPIRYPQLFTGILSPWKGLLLYGP 183
Query: 166 PGTGKTMLAKAVATECKTTFFNISASSVVSKWRGDSEKLVKVLFQLARHHAPSTIFLDEI 225
PGTGKT+LAKAVATECKTTFFNISAS++VSKWRGDSEKLV+VLF+LAR+HAPSTIFLDE+
Sbjct: 184 PGTGKTLLAKAVATECKTTFFNISASTIVSKWRGDSEKLVRVLFELARYHAPSTIFLDEL 243
Query: 226 DAIISQRGEAR-SEHEASRRLKTELLIQMDGLTRTDELVFVLAATNLPWELDAAMLRRLE 284
++++SQRG A EHE S R+KTELL+QMDGL R+++LVFVLAA+NLPWELD AMLRRLE
Sbjct: 244 ESVMSQRGTASGGEHEGSLRMKTELLVQMDGLARSEDLVFVLAASNLPWELDCAMLRRLE 303
Query: 285 KRILVPLPEPEARVAMFEELLPPQPDEES------IPYDLLVNQTEGYSGSDIRLLCKEV 338
KRILV LP EAR AM LPP + + Y +L +TEGYSGSDI+L+C+E
Sbjct: 304 KRILVDLPSREARQAMIYHWLPPVSKSRALELHTELEYSVLSQETEGYSGSDIKLVCREA 363
Query: 339 AMQPLRRLMSQLEQREDLVPEEELPKV--GPIRPEDIQAALKNTRPSAHLHAHKYDKFNA 396
AM+P+R++ LE + +LP++ + D L +T+PSA A +Y +
Sbjct: 364 AMRPVRKIFDALESHQS--ESSDLPRIQLDIVTTADFLDVLTHTKPSAKNLAQRYSDWQR 421
Query: 397 DYGS 400
++ S
Sbjct: 422 EFES 425
>UniRef100_UPI00003AAC53 UPI00003AAC53 UniRef100 entry
Length = 464
Score = 360 bits (923), Expect = 5e-98
Identities = 190/339 (56%), Positives = 242/339 (71%), Gaps = 19/339 (5%)
Query: 81 VSPNNM-------DERPQKSLLPPFES-----AEMRTLAESLSRDIIRGSPDVKWESIKG 128
VSPN + D P + LL P + EMR LA +S+DI +P+VKW+ I G
Sbjct: 125 VSPNGIGLSSLTGDPDPSERLLKPLSAFIGMNGEMRELATVVSKDIYLHNPNVKWDDIIG 184
Query: 129 LENAKRLLKEAVVMPIKYPKYFTGLLSPWKGILLFGPPGTGKTMLAKAVATECKTTFFNI 188
L+ AKRL+KEAVV PI+YP+ FTG+LSPWKG+LL+GPPGTGKT+LAKAVATEC TTFFNI
Sbjct: 185 LDAAKRLVKEAVVYPIRYPQLFTGILSPWKGLLLYGPPGTGKTLLAKAVATECNTTFFNI 244
Query: 189 SASSVVSKWRGDSEKLVKVLFQLARHHAPSTIFLDEIDAIISQRGE-ARSEHEASRRLKT 247
SAS++VSKWRGDSEKLV+VLF+LAR+HAPSTIFLDE+++++SQRG + EHE SRR+KT
Sbjct: 245 SASTIVSKWRGDSEKLVRVLFELARYHAPSTIFLDELESVMSQRGTISGGEHEGSRRMKT 304
Query: 248 ELLIQMDGLTRTDELVFVLAATNLPWELDAAMLRRLEKRILVPLPEPEARVAMFEELLPP 307
ELL+QMDGL R+D+LVFVLAA+NLPWELD+AMLRRLEKRILV LP EAR AM LPP
Sbjct: 305 ELLVQMDGLARSDDLVFVLAASNLPWELDSAMLRRLEKRILVDLPNQEARQAMIRHWLPP 364
Query: 308 QPD------EESIPYDLLVNQTEGYSGSDIRLLCKEVAMQPLRRLMSQLEQREDLVPEEE 361
+ + Y LL +T+GYSGSDI+L+CKE AM+P+R++ LE +
Sbjct: 365 LSNSGGVELRTDLDYSLLGRETDGYSGSDIKLVCKEAAMRPVRKVFDALENHQPGNSNLA 424
Query: 362 LPKVGPIRPEDIQAALKNTRPSAHLHAHKYDKFNADYGS 400
+ I D + +T+PSA + KY + ++ S
Sbjct: 425 AVHLDMITTADFLDVIAHTKPSAKKLSQKYTAWQREFES 463
>UniRef100_Q8IYT4 Similar to mouse 4933439B08Rik protein [Homo sapiens]
Length = 466
Score = 360 bits (923), Expect = 5e-98
Identities = 194/364 (53%), Positives = 256/364 (70%), Gaps = 20/364 (5%)
Query: 53 NVHSKRSSDMAIYEQLRSQGQNGIHTNDVSPNNMDERPQKS--LLPPFES-----AEMRT 105
N H +R + ++ L + G T++++ N D P S LL P + +EMR
Sbjct: 106 NAHPRRGQ-IIDFQGLLTDAIKGA-TSELALNTFDHNPDPSERLLKPLSAFIGMNSEMRE 163
Query: 106 LAESLSRDIIRGSPDVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGLLSPWKGILLFGP 165
LA +SRDI +P++KW I GL+ AK+L+KEAVV PI+YP+ FTG+LSPWKG+LL+GP
Sbjct: 164 LAAVVSRDIYLHNPNIKWNDIIGLDAAKQLVKEAVVYPIRYPQLFTGILSPWKGLLLYGP 223
Query: 166 PGTGKTMLAKAVATECKTTFFNISASSVVSKWRGDSEKLVKVLFQLARHHAPSTIFLDEI 225
PGTGKT+LAKAVATECKTTFFNISAS++VSKWRGDSEKLV+VLF+LAR+HAPSTIFLDE+
Sbjct: 224 PGTGKTLLAKAVATECKTTFFNISASTIVSKWRGDSEKLVRVLFELARYHAPSTIFLDEL 283
Query: 226 DAIISQRGEAR-SEHEASRRLKTELLIQMDGLTRTDELVFVLAATNLPWELDAAMLRRLE 284
++++SQRG A EHE S R+KTELL+QMDGL R+++LVFVLAA+NLPWELD AMLRRLE
Sbjct: 284 ESVMSQRGTASGGEHEGSLRMKTELLVQMDGLARSEDLVFVLAASNLPWELDCAMLRRLE 343
Query: 285 KRILVPLPEPEARVAMFEELLPPQPDEES------IPYDLLVNQTEGYSGSDIRLLCKEV 338
KRILV LP EAR AM LPP + + Y +L +TEGYSGSDI+L+C+E
Sbjct: 344 KRILVDLPSREARQAMIYHWLPPVSKSRALELHTELEYSVLSQETEGYSGSDIKLVCREA 403
Query: 339 AMQPLRRLMSQLEQREDLVPEEELPKV--GPIRPEDIQAALKNTRPSAHLHAHKYDKFNA 396
AM+P+R++ LE + +LP++ + D L +T+PSA A +Y +
Sbjct: 404 AMRPVRKIFDALENHQS--ESSDLPRIQLDIVTTADFLDVLTHTKPSAKNLAQRYSDWQR 461
Query: 397 DYGS 400
++ S
Sbjct: 462 EFES 465
>UniRef100_Q9GNC3 Probable AAA ATPase [Leishmania major]
Length = 565
Score = 344 bits (882), Expect = 3e-93
Identities = 198/390 (50%), Positives = 254/390 (64%), Gaps = 28/390 (7%)
Query: 35 NADKAVGNGSSMSVVSNGNVH----SKRSSDMAIYEQLRSQGQNGIHTNDVSPNNMDERP 90
N D NG +V NG+ S + A + R +G+ G N D+
Sbjct: 176 NGDSGSANGDKRAVALNGDTSALGLSLQGESAAAHTSAR-EGEKGRRDNGRDGEGSDDAA 234
Query: 91 QKSL-----------LPPFESAEMRTLAESLSRDIIRGSPDVKWESIKGLENAKRLLKEA 139
+ L LPPF ++E+ LA ++ R+I+ P V+W I LENAK LL+EA
Sbjct: 235 EDPLGSLMSRRILKPLPPFPTSELSELAATILREILDVDPSVRWRDIADLENAKHLLREA 294
Query: 140 VVMPIKYPKYFTGLLSPWKGILLFGPPGTGKTMLAKAVATECKTTFFNISASSVVSKWRG 199
VVMP+KYP F G+L PWKGILLFGPPGTGKT+LAKAVATEC+TTFFNI+ASSVVSKWRG
Sbjct: 295 VVMPVKYPGLFQGILRPWKGILLFGPPGTGKTLLAKAVATECRTTFFNIAASSVVSKWRG 354
Query: 200 DSEKLVKVLFQLARHHAPSTIFLDEIDAIISQRGEARSEHEASRRLKTELLIQMDGLT-- 257
DSEKLV++LF LA H+APSTIF+DEID+++S R + EHE SRR+KTELL QMDGL+
Sbjct: 355 DSEKLVRMLFDLAVHYAPSTIFIDEIDSLMSAR-SSDGEHEGSRRMKTELLTQMDGLSKR 413
Query: 258 RTDELVFVLAATNLPWELDAAMLRRLEKRILVPLPEPEARVAMFEELLPPQPDEESIPYD 317
R E+VFVLAA+N+PW+LD AMLRRLEKRILV LP +ARV MF LLP ++ Y+
Sbjct: 414 RGGEVVFVLAASNVPWDLDTAMLRRLEKRILVSLPTRDARVLMFRRLLPNSFASDA-DYE 472
Query: 318 LLVNQTEGYSGSDIRLLCKEVAMQPLRRLMSQLEQR-EDLVPEEELPKVGPIRP-----E 371
TEG SG+DI ++C+E M+P+R+L+SQLE D LP P+RP E
Sbjct: 473 ACAALTEGMSGADIDVVCREAMMRPVRKLISQLEAAGNDRNAHARLPS-EPLRPPAATLE 531
Query: 372 DIQAALKNTRPSAHL-HAHKYDKFNADYGS 400
D+QA++ TR S + KYD + ++GS
Sbjct: 532 DVQASVACTRSSVRVADLDKYDVWTREHGS 561
>UniRef100_UPI0000360C40 UPI0000360C40 UniRef100 entry
Length = 320
Score = 329 bits (844), Expect = 7e-89
Identities = 166/307 (54%), Positives = 217/307 (70%), Gaps = 6/307 (1%)
Query: 100 SAEMRTLAESLSRDIIRGSPDVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGLLSPWKG 159
S+EM LA +SRDI V+W++I GLE AK+L+KEAVV PIKYP+ FTG++SPWK
Sbjct: 14 SSEMSELAAVISRDIYMDDLSVQWDNIIGLEEAKKLVKEAVVYPIKYPQLFTGIVSPWKA 73
Query: 160 ILLFGPPGTGKTMLAKAVATECKTTFFNISASSVVSKWRGDSEKLVKVLFQLARHHAPST 219
+LL+GPPGTGKT+LAKAVA EC+TTFFNISASS+ SKWRGDSEKLV+VLF+LA +HAPST
Sbjct: 74 LLLYGPPGTGKTLLAKAVAAECRTTFFNISASSITSKWRGDSEKLVRVLFELAWYHAPST 133
Query: 220 IFLDEIDAIISQRGEARSEHEASRRLKTELLIQMDGLTRTDELVFVLAATNLPWELDAAM 279
IFLDE+++++ RG EHE SRR+K ELL+QMDGL +++LVFVLAA+NLPWELD A+
Sbjct: 134 IFLDELESLMGGRGSGGGEHEGSRRMKAELLVQMDGLKSSEQLVFVLAASNLPWELDQAV 193
Query: 280 LRRLEKRILVPLPEPEARVAMFEELLPPQPD------EESIPYDLLVNQTEGYSGSDIRL 333
LRRLEKRILV LP AR M LPP+ + + Y+ L +TEGYSGSDIRL
Sbjct: 194 LRRLEKRILVGLPSWSARCTMISHWLPPRSSTGGLELQTQLDYEALAEETEGYSGSDIRL 253
Query: 334 LCKEVAMQPLRRLMSQLEQREDLVPEEELPKVGPIRPEDIQAALKNTRPSAHLHAHKYDK 393
+CKE M+ +R++ LE ++ + ++ + D + ++PS KY
Sbjct: 254 VCKEAVMRSVRKIFDALESHDEGGVDVATVQLESVTTADFLEVMSRSKPSCRNLKDKYAW 313
Query: 394 FNADYGS 400
+ + Y S
Sbjct: 314 WESRYQS 320
>UniRef100_UPI000024AF1B UPI000024AF1B UniRef100 entry
Length = 274
Score = 301 bits (772), Expect = 2e-80
Identities = 154/273 (56%), Positives = 199/273 (72%), Gaps = 7/273 (2%)
Query: 135 LLKEAVVMPIKYPKYFTGLLSPWKGILLFGPPGTGKTMLAKAVATECKTTFFNISASSVV 194
L+ + V++ I++ TG+LSPWKG+LL+GPPGTGKTMLAKAVATEC TTFFNISASS+V
Sbjct: 1 LIHDLVILYIRFIYKITGILSPWKGLLLYGPPGTGKTMLAKAVATECNTTFFNISASSIV 60
Query: 195 SKWRGDSEKLVKVLFQLARHHAPSTIFLDEIDAIISQRGEAR-SEHEASRRLKTELLIQM 253
SKWRGDSEKLV+VLF+LAR+HAPSTIFLDE+++++SQRG + +HE SRR+KTELL+QM
Sbjct: 61 SKWRGDSEKLVRVLFELARYHAPSTIFLDELESVMSQRGVGQGGDHEGSRRMKTELLVQM 120
Query: 254 DGLTRTDELVFVLAATNLPWELDAAMLRRLEKRILVPLPEPEARVAMFEELLPPQPD--- 310
DGL R+++LVFVLAA+NLPWELD AMLRRLEKRILV LP AR AM LPP +
Sbjct: 121 DGLARSNDLVFVLAASNLPWELDHAMLRRLEKRILVSLPSAPARQAMISHWLPPVSNTGG 180
Query: 311 ---EESIPYDLLVNQTEGYSGSDIRLLCKEVAMQPLRRLMSQLEQREDLVPEEELPKVGP 367
+ YD L +T GYSGSDIRL+CKE AM+P+R++ LE + + ++
Sbjct: 181 VELRTELDYDSLAQETGGYSGSDIRLVCKEAAMRPVRKIFDALENHIEGQSNMPVIELET 240
Query: 368 IRPEDIQAALKNTRPSAHLHAHKYDKFNADYGS 400
+ D + +T+PSA +Y + +Y S
Sbjct: 241 VTTADFLDVIVHTKPSARCLMDRYTAWEKEYES 273
>UniRef100_Q5RII9 Novel protein similar to vertebrate katanin p60 (ATPase-containing)
subunit A 1 [Brachydanio rerio]
Length = 485
Score = 289 bits (740), Expect = 8e-77
Identities = 167/374 (44%), Positives = 237/374 (62%), Gaps = 23/374 (6%)
Query: 41 GNGSSMSVVSNGNVHSKRSSDMAIYEQLRSQGQNGIHTNDVSPN---NMDERPQKSLLPP 97
G+G+ +SV G ++ S +A ++ + Q N P N E + +
Sbjct: 120 GHGNRLSVGPRGQ--ARPSPRVANGDKGKPQKSKEKKENPSKPKEDKNKAEAVETEVKRF 177
Query: 98 FESAEMRTLAESLSRDIIRGSPDVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGLLSPW 157
E + L ++L RDII +P+V W+ I LE AK+LLKEAVV+P+ P++F G+ PW
Sbjct: 178 DRGGEDKDLIDALERDIISQNPNVTWDDIADLEEAKKLLKEAVVLPMWMPEFFKGIRRPW 237
Query: 158 KGILLFGPPGTGKTMLAKAVATECKTTFFNISASSVVSKWRGDSEKLVKVLFQLARHHAP 217
KG+L+ GPPGTGKT+LAKAVATEC+TTFFN+S+S++ SK+RG+SEKLV++LF++AR +AP
Sbjct: 238 KGVLMVGPPGTGKTLLAKAVATECRTTFFNVSSSTLTSKYRGESEKLVRLLFEMARFYAP 297
Query: 218 STIFLDEIDAIISQRGEARSEHEASRRLKTELLIQMDGLTRTDE-----LVFVLAATNLP 272
+TIF+DEID+I S+RG + EHEASRR+K ELL+QMDG+ T E +V VLAATN P
Sbjct: 298 TTIFIDEIDSICSRRGTS-EEHEASRRVKAELLVQMDGVGGTSENDPSKMVMVLAATNFP 356
Query: 273 WELDAAMLRRLEKRILVPLPEPEARVAMFEELLPPQPDEESIPYDLLVNQTEGYSGSDIR 332
W++D A+ RRLEKRI +PLP + RV + + L + D + Q EGYSG+DI
Sbjct: 357 WDIDEALRRRLEKRIYIPLPSAKGRVDLLKINLKELDLANDVNMDKIAEQMEGYSGADIT 416
Query: 333 LLCKEVAMQPLRRLMSQLEQREDLVPEE--ELPKVG---PIRPEDIQAALKNTRPS-AHL 386
+C++ ++ +RR + E L PEE LPK P ED + ALK S +
Sbjct: 417 NVCRDASLMAMRRRI------EGLTPEEIRNLPKDEMHMPTTMEDFETALKKVSKSVSAA 470
Query: 387 HAHKYDKFNADYGS 400
KY+K+ A++GS
Sbjct: 471 DLEKYEKWIAEFGS 484
>UniRef100_UPI0000437799 UPI0000437799 UniRef100 entry
Length = 482
Score = 289 bits (739), Expect = 1e-76
Identities = 155/310 (50%), Positives = 214/310 (69%), Gaps = 18/310 (5%)
Query: 102 EMRTLAESLSRDIIRGSPDVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGLLSPWKGIL 161
E + L ++L RDII +P+V W+ I LE AK+LLKEAVV+P+ P++F G+ PWKG+L
Sbjct: 180 EDKDLIDALERDIISQNPNVTWDDIADLEEAKKLLKEAVVLPMWMPEFFKGIRRPWKGVL 239
Query: 162 LFGPPGTGKTMLAKAVATECKTTFFNISASSVVSKWRGDSEKLVKVLFQLARHHAPSTIF 221
+ GPPGTGKT+LAKAVATEC+TTFFN+S+S++ SK+RG+SEKLV++LF++AR +AP+TIF
Sbjct: 240 MVGPPGTGKTLLAKAVATECRTTFFNVSSSTLTSKYRGESEKLVRLLFEMARFYAPTTIF 299
Query: 222 LDEIDAIISQRGEARSEHEASRRLKTELLIQMDGLTRTDE-----LVFVLAATNLPWELD 276
+DEID+I S+RG + EHEASRR+K ELL+QMDG+ T E +V VLAATN PW++D
Sbjct: 300 IDEIDSICSRRGTS-EEHEASRRVKAELLVQMDGVGGTSENDPSKMVMVLAATNFPWDID 358
Query: 277 AAMLRRLEKRILVPLPEPEARVAMFEELLPPQPDEESIPYDLLVNQTEGYSGSDIRLLCK 336
A+ RRLEKRI +PLP + RV + + L + D + Q EGYSG+DI +C+
Sbjct: 359 EALRRRLEKRIYIPLPSAKGRVDLLKINLKELDLANDVNMDKIAEQMEGYSGADITNVCR 418
Query: 337 EVAMQPLRRLMSQLEQREDLVPEE--ELPKVG---PIRPEDIQAALKNTRPS-AHLHAHK 390
+ ++ +RR + E L PEE LPK P ED + ALK S + K
Sbjct: 419 DASLMAMRRRI------EGLTPEEIRNLPKDEMHMPTTMEDFETALKKVSKSVSAADLEK 472
Query: 391 YDKFNADYGS 400
Y+K+ A++GS
Sbjct: 473 YEKWIAEFGS 482
>UniRef100_Q8K0T4 Katanin p60 subunit A-like 1 [Mus musculus]
Length = 488
Score = 287 bits (734), Expect = 4e-76
Identities = 160/372 (43%), Positives = 241/372 (64%), Gaps = 18/372 (4%)
Query: 38 KAVGNGSSMSVVSNGNVHSKRSSDMAIYEQLRSQGQNGIHTNDVSPNNMDERPQKSLLPP 97
K VG G+ +V + SK + + R++G++ D + N+ + S +P
Sbjct: 125 KDVGAGAR-GLVGRAHQISKSDKPASRDKDYRARGRD-----DKARKNVQDGASDSEIPK 178
Query: 98 FESAEM-RTLAESLSRDIIRGSPDVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGLLSP 156
F+ A + L E+L RDI+ +P + W+ I LE AK+LL+EAVV+P+ P +F G+ P
Sbjct: 179 FDGAGYDKDLVEALERDIVSRNPSIHWDDIADLEEAKKLLREAVVLPMWMPDFFKGIRRP 238
Query: 157 WKGILLFGPPGTGKTMLAKAVATECKTTFFNISASSVVSKWRGDSEKLVKVLFQLARHHA 216
WKG+L+ GPPGTGKTMLAKAVATEC TTFFN+S+S++ SK+RG+SEKLV++LF++AR +A
Sbjct: 239 WKGVLMVGPPGTGKTMLAKAVATECGTTFFNVSSSTLTSKYRGESEKLVRLLFEMARFYA 298
Query: 217 PSTIFLDEIDAIISQRGEARSEHEASRRLKTELLIQMDGL------TRTDELVFVLAATN 270
P+TIF+DEID+I S+RG + EHEASRR+K+ELLIQMDG+ ++V VLAATN
Sbjct: 299 PTTIFIDEIDSICSRRGTS-DEHEASRRVKSELLIQMDGVGGALENDDPSKMVMVLAATN 357
Query: 271 LPWELDAAMLRRLEKRILVPLPEPEARVAMFEELLPPQPDEESIPYDLLVNQTEGYSGSD 330
PW++D A+ RRLEKRI +PLP + R + + L + + + + ++TEGYSG+D
Sbjct: 358 FPWDIDEALRRRLEKRIYIPLPTAKGRAELLKISLREVELDPDVHLEDIADKTEGYSGAD 417
Query: 331 IRLLCKEVAMQPLRRLMSQLEQRE-DLVPEEELPKVGPIRPEDIQAALKNTRPS-AHLHA 388
I +C++ ++ +RR ++ L E + +EEL P+ D++ ALK S +
Sbjct: 418 ITNICRDASLMAMRRRINGLSPEEIRALSKEELQM--PVTRGDLELALKKIAKSVSAADL 475
Query: 389 HKYDKFNADYGS 400
KY+K+ ++GS
Sbjct: 476 EKYEKWMVEFGS 487
>UniRef100_Q5XIK7 Hypothetical protein MGC94518 [Rattus norvegicus]
Length = 488
Score = 287 bits (734), Expect = 4e-76
Identities = 159/360 (44%), Positives = 233/360 (64%), Gaps = 19/360 (5%)
Query: 52 GNVHSKRSSDMAIY--EQLRSQGQNGIHTNDVSPNNMDERPQKSLLPPFESAEM-RTLAE 108
G H SD A + R++G++ D + NM + +P F+ A + L E
Sbjct: 136 GRAHQISKSDKAASRDKDYRARGRD-----DKARKNMQDGASDGEIPKFDGAGYDKDLVE 190
Query: 109 SLSRDIIRGSPDVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGLLSPWKGILLFGPPGT 168
+L RDI+ +P + W+ I LE AK+LL+EAVV+P+ P +F G+ PWKG+L+ GPPGT
Sbjct: 191 ALERDIVSRNPSIHWDDIADLEEAKKLLREAVVLPMWMPDFFKGIRRPWKGVLMVGPPGT 250
Query: 169 GKTMLAKAVATECKTTFFNISASSVVSKWRGDSEKLVKVLFQLARHHAPSTIFLDEIDAI 228
GKTMLAKAVATEC TTFFN+S+S++ SK+RG+SEKLV++LF++AR +AP+TIF+DEID+I
Sbjct: 251 GKTMLAKAVATECGTTFFNVSSSTLTSKYRGESEKLVRLLFEMARFYAPTTIFIDEIDSI 310
Query: 229 ISQRGEARSEHEASRRLKTELLIQMDGL------TRTDELVFVLAATNLPWELDAAMLRR 282
S+RG + EHEASRR+K+ELLIQMDG+ ++V VLAATN PW++D A+ RR
Sbjct: 311 CSRRGTS-DEHEASRRVKSELLIQMDGVGGALENDDPSKMVMVLAATNFPWDIDEALRRR 369
Query: 283 LEKRILVPLPEPEARVAMFEELLPPQPDEESIPYDLLVNQTEGYSGSDIRLLCKEVAMQP 342
LEKRI +PLP + R + + L + I + + +TEGYSG+DI +C++ ++
Sbjct: 370 LEKRIYIPLPTAKGRAELLKISLREVELDPDIHLEDIAEKTEGYSGADITNICRDASLMA 429
Query: 343 LRRLMSQLEQRE-DLVPEEELPKVGPIRPEDIQAALKNTRPS-AHLHAHKYDKFNADYGS 400
+RR ++ L E + +EEL P+ D++ ALK S + KY+K+ ++GS
Sbjct: 430 MRRRINGLSPEEIRALSKEELQM--PVTRGDLELALKKIAKSVSAADLEKYEKWMVEFGS 487
>UniRef100_Q8SZ40 RE17942p [Drosophila melanogaster]
Length = 572
Score = 285 bits (728), Expect = 2e-75
Identities = 164/380 (43%), Positives = 238/380 (62%), Gaps = 18/380 (4%)
Query: 31 QNGENADKAVGNGSSMSVVSNGNVHSKRSSDMAIYEQLRSQGQNGIHTNDVS---PNNMD 87
+N +A G + + + G S +++ A + + G NG D P
Sbjct: 200 RNSTSATAPSGGARTTNGRAGGRKLSTSNTNEARDDDSTAAGNNGGAAGDGENGDPQAAQ 259
Query: 88 ERPQKSLLPPFESAEMRTLAESLSRDIIRGSPDVKWESIKGLENAKRLLKEAVVMPIKYP 147
E +K AE L + L RDI++ P V+W I L +AKRLL+EAVV+P+ P
Sbjct: 260 EEERKFQTNNHIEAE---LVDILERDILQKDPKVRWSDIADLHDAKRLLEEAVVLPMLMP 316
Query: 148 KYFTGLLSPWKGILLFGPPGTGKTMLAKAVATECKTTFFNISASSVVSKWRGDSEKLVKV 207
YF G+ PWKG+L+ GPPGTGKTMLAKAVATEC TTFFN+S++++ SK+RG+SEK+V++
Sbjct: 317 DYFKGIRRPWKGVLMVGPPGTGKTMLAKAVATECGTTFFNVSSATLTSKYRGESEKMVRL 376
Query: 208 LFQLARHHAPSTIFLDEIDAIISQRGEARSEHEASRRLKTELLIQMDGLTRTDE---LVF 264
LF++AR +APSTIF+DEID++ S+RG + SEHEASRR+K+ELL+QMDG+ +E +V
Sbjct: 377 LFEMARFYAPSTIFIDEIDSLCSRRG-SESEHEASRRVKSELLVQMDGVGGGEEQAKVVM 435
Query: 265 VLAATNLPWELDAAMLRRLEKRILVPLPEPEARVAMFEELLPPQPDEESIPYDLLVNQTE 324
VLAATN PW++D A+ RRLEKRI +PLP E R A+ + L ++S+ + N+ +
Sbjct: 436 VLAATNFPWDIDEALRRRLEKRIYIPLPSDEGREALLKINLREVKVDDSVDLTYVANELK 495
Query: 325 GYSGSDIRLLCKEVAMQPLRRLMSQL--EQREDLVPEE-ELPKVGPIRPEDIQAALKNTR 381
GYSG+DI +C+E +M +RR ++ L EQ L EE +L P+ +D A+
Sbjct: 496 GYSGADITNVCREASMMSMRRKIAGLTPEQIRQLATEEVDL----PVSNKDFNEAMSRCN 551
Query: 382 PS-AHLHAHKYDKFNADYGS 400
S + KY+K+ ++GS
Sbjct: 552 KSVSRADLDKYEKWMREFGS 571
>UniRef100_Q9VN89 CG10229-PA [Drosophila melanogaster]
Length = 572
Score = 284 bits (726), Expect = 4e-75
Identities = 164/380 (43%), Positives = 238/380 (62%), Gaps = 18/380 (4%)
Query: 31 QNGENADKAVGNGSSMSVVSNGNVHSKRSSDMAIYEQLRSQGQNGIHTNDVS---PNNMD 87
+N +A G + + + G S +++ A + + G NG D P
Sbjct: 200 RNSTSATAPSGGARTTNGRAGGRKLSTSNTNEARDDDSTAAGNNGGAAGDGENGDPQAAQ 259
Query: 88 ERPQKSLLPPFESAEMRTLAESLSRDIIRGSPDVKWESIKGLENAKRLLKEAVVMPIKYP 147
E +K AE L + L RDI++ P V+W I L +AKRLL+EAVV+P+ P
Sbjct: 260 EEERKFQPNNHIEAE---LVDILERDILQKDPKVRWSDIADLHDAKRLLEEAVVLPMLMP 316
Query: 148 KYFTGLLSPWKGILLFGPPGTGKTMLAKAVATECKTTFFNISASSVVSKWRGDSEKLVKV 207
YF G+ PWKG+L+ GPPGTGKTMLAKAVATEC TTFFN+S++++ SK+RG+SEK+V++
Sbjct: 317 DYFKGIRRPWKGVLMVGPPGTGKTMLAKAVATECGTTFFNVSSATLTSKYRGESEKMVRL 376
Query: 208 LFQLARHHAPSTIFLDEIDAIISQRGEARSEHEASRRLKTELLIQMDGLTRTDE---LVF 264
LF++AR +APSTIF+DEID++ S+RG + SEHEASRR+K+ELL+QMDG+ +E +V
Sbjct: 377 LFEMARFYAPSTIFIDEIDSLCSRRG-SESEHEASRRVKSELLVQMDGVGGGEEQAKVVM 435
Query: 265 VLAATNLPWELDAAMLRRLEKRILVPLPEPEARVAMFEELLPPQPDEESIPYDLLVNQTE 324
VLAATN PW++D A+ RRLEKRI +PLP E R A+ + L ++S+ + N+ +
Sbjct: 436 VLAATNFPWDIDEALRRRLEKRIYIPLPSDEGREALLKINLREVKVDDSVDLTYVANELK 495
Query: 325 GYSGSDIRLLCKEVAMQPLRRLMSQL--EQREDLVPEE-ELPKVGPIRPEDIQAALKNTR 381
GYSG+DI +C+E +M +RR ++ L EQ L EE +L P+ +D A+
Sbjct: 496 GYSGADITNVCREASMMSMRRKIAGLTPEQIRQLATEEVDL----PVSNKDFNEAMSRCN 551
Query: 382 PS-AHLHAHKYDKFNADYGS 400
S + KY+K+ ++GS
Sbjct: 552 KSVSRADLDKYEKWMREFGS 571
>UniRef100_Q9NHF1 Putative microtubule severing protein katanin p60 subunit
[Drosophila melanogaster]
Length = 571
Score = 284 bits (726), Expect = 4e-75
Identities = 163/379 (43%), Positives = 238/379 (62%), Gaps = 17/379 (4%)
Query: 31 QNGENADKAVGNGSSMSVVSNGNVHSKRSSDMAIYEQLRSQGQNGIHTNDVS---PNNMD 87
+N +A G + + + G S +++ A + + G NG D P
Sbjct: 200 RNSTSATAPSGGARTTNGRAGGRKLSTSNTNEARDDDSTAAGNNGGAAGDGENGDPQAAQ 259
Query: 88 ERPQKSLLPPFESAEMRTLAESLSRDIIRGSPDVKWESIKGLENAKRLLKEAVVMPIKYP 147
E +K AE L + L RDI++ P V+W I L +AKRLL+EAVV+P+ P
Sbjct: 260 EEERKFQPNNHIEAE---LVDILERDILQKDPKVRWSDIADLHDAKRLLEEAVVLPMLMP 316
Query: 148 KYFTGLLSPWKGILLFGPPGTGKTMLAKAVATECKTTFFNISASSVVSKWRGDSEKLVKV 207
YF G+ PWKG+L+ GP GTGKTMLAKAVATEC TTFFN+S++++ SK+RG+SEK+V++
Sbjct: 317 DYFKGIRRPWKGVLMVGPSGTGKTMLAKAVATECGTTFFNVSSATLTSKYRGESEKMVRL 376
Query: 208 LFQLARHHAPSTIFLDEIDAIISQRGEARSEHEASRRLKTELLIQMDGLTRTDE--LVFV 265
LF++AR +APSTIF+DEID++ S+RG + SEHEASRR+K+ELL+QMDG+ R ++ +V V
Sbjct: 377 LFEMARFYAPSTIFIDEIDSLCSRRG-SESEHEASRRVKSELLVQMDGVAREEQAKVVMV 435
Query: 266 LAATNLPWELDAAMLRRLEKRILVPLPEPEARVAMFEELLPPQPDEESIPYDLLVNQTEG 325
LAATN PW++D A+ RRLEKRI +PLP E R A+ + L ++S+ + N+ +G
Sbjct: 436 LAATNFPWDIDEALRRRLEKRIYIPLPSDEGREALLKINLREVKVDDSVDLTYVANELKG 495
Query: 326 YSGSDIRLLCKEVAMQPLRRLMSQL--EQREDLVPEE-ELPKVGPIRPEDIQAALKNTRP 382
YSG+DI +C+E +M +RR ++ L EQ L EE +L P+ +D A+
Sbjct: 496 YSGADITNVCREASMMSMRRKIAGLTPEQIRQLATEEVDL----PVSNKDFNEAMSRCNK 551
Query: 383 S-AHLHAHKYDKFNADYGS 400
S + KY+K+ ++GS
Sbjct: 552 SVSRADLDKYEKWMREFGS 570
>UniRef100_Q7PY77 ENSANGP00000018492 [Anopheles gambiae str. PEST]
Length = 605
Score = 282 bits (721), Expect = 1e-74
Identities = 163/389 (41%), Positives = 244/389 (61%), Gaps = 19/389 (4%)
Query: 25 GRKKVVQNGENADKAVGNGSSMS--VVSNGNVHSKRSSDMAIYEQLRSQGQNGIHT---N 79
GRK + Q + NG+S + + + S+ + G+ G +
Sbjct: 222 GRKTISQTARAGGTSGMNGASRTGTLPRTKAAGQRGSTAPGASDATNGNGEKGDKEKLGD 281
Query: 80 DVSPNNMDERPQK--SLLPPFESAEMRTLAESLSRDIIRGSPDVKWESIKGLENAKRLLK 137
D NN + P++ P A++ L + L RDI++ +P++ W+ I L AKRLL+
Sbjct: 282 DEEGNNGGDTPEEVERKFEPASHADV-DLVDMLERDILQKNPNIHWDDIADLHEAKRLLE 340
Query: 138 EAVVMPIKYPKYFTGLLSPWKGILLFGPPGTGKTMLAKAVATECKTTFFNISASSVVSKW 197
EAVV+P+ P YF G+ PWKG+L+ GPPGTGKTMLAKAVATEC TTFFN+S+S++ SK+
Sbjct: 341 EAVVLPMWMPDYFKGIRRPWKGVLMVGPPGTGKTMLAKAVATECGTTFFNVSSSTLTSKY 400
Query: 198 RGDSEKLVKVLFQLARHHAPSTIFLDEIDAIISQRGEARSEHEASRRLKTELLIQMDGLT 257
RG+SEKLV++LF++AR +APSTIF+DEID++ S+RG + SEHEASRR+K+ELL+QMDG++
Sbjct: 401 RGESEKLVRLLFEMARFYAPSTIFIDEIDSLCSRRG-SESEHEASRRVKSELLVQMDGVS 459
Query: 258 RTD--ELVFVLAATNLPWELDAAMLRRLEKRILVPLPEPEARVAMFEELLPPQPDEESIP 315
+ ++V VLAATN PW++D A+ RRLEKRI +PLP E R A+ + L +ES+
Sbjct: 460 NDEATKIVMVLAATNFPWDIDEALRRRLEKRIYIPLPNSEGREALLKINLREVKVDESVD 519
Query: 316 YDLLVNQTEGYSGSDIRLLCKEVAMQPLRRLMSQL--EQREDLVPEE-ELPKVGPIRPED 372
+ ++ +GYSG+DI +C++ +M +RR ++ L EQ L EE +L P+ +D
Sbjct: 520 MRDIADRLDGYSGADITNVCRDASMMSMRRKIAGLRPEQIRQLAKEELDL----PVSKQD 575
Query: 373 IQAALKNTRPSAHL-HAHKYDKFNADYGS 400
+ A+ S KY ++ ++GS
Sbjct: 576 FKEAISKCNKSVSKDDLAKYQQWMKEFGS 604
>UniRef100_O61577 Katanin p60 subunit [Strongylocentrotus purpuratus]
Length = 516
Score = 282 bits (721), Expect = 1e-74
Identities = 151/305 (49%), Positives = 208/305 (67%), Gaps = 11/305 (3%)
Query: 104 RTLAESLSRDIIRGSPDVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGLLSPWKGILLF 163
+ L E+L RDI++ +P+V W I GL AKRLL+EAVV+P+ P YF G+ PWKG+L+
Sbjct: 214 KDLVENLERDIVQRNPNVHWADIAGLTEAKRLLEEAVVLPLWMPDYFKGIRRPWKGVLMV 273
Query: 164 GPPGTGKTMLAKAVATECKTTFFNISASSVVSKWRGDSEKLVKVLFQLARHHAPSTIFLD 223
GPPGTGKTMLAKAVATEC TTFFN+S++S+ SK+ G+SEKLV++LF++AR +APSTIF+D
Sbjct: 274 GPPGTGKTMLAKAVATECGTTFFNVSSASLTSKYHGESEKLVRLLFEMARFYAPSTIFID 333
Query: 224 EIDAIISQRGEARSEHEASRRLKTELLIQMDGLT------RTDELVFVLAATNLPWELDA 277
EID+I S+RG SEHEASRR+K+ELLIQMDG++ + ++V VLAATN PW++D
Sbjct: 334 EIDSICSKRGTG-SEHEASRRVKSELLIQMDGVSGPSAGEESSKMVMVLAATNFPWDIDE 392
Query: 278 AMLRRLEKRILVPLPEPEARVAMFEELLPPQPDEESIPYDLLVNQTEGYSGSDIRLLCKE 337
A+ RRLEKRI +PLPE + R + L P + I + + +GYSG+DI +C++
Sbjct: 393 ALRRRLEKRIYIPLPEIDGREQLLRINLKEVPLADDIDLKSIAEKMDGYSGADITNVCRD 452
Query: 338 VAMQPLRRLMSQLEQRE-DLVPEEELPKVGPIRPEDIQAALKNTRPSAHLH-AHKYDKFN 395
+M +RR + L E +P+EEL + P P D AL+ S KY +
Sbjct: 453 ASMMAMRRRIQGLRPEEIRHIPKEELNQ--PSTPADFLLALQKVSKSVGKEDLVKYMAWM 510
Query: 396 ADYGS 400
++GS
Sbjct: 511 EEFGS 515
>UniRef100_UPI0000244E4F UPI0000244E4F UniRef100 entry
Length = 492
Score = 281 bits (720), Expect = 2e-74
Identities = 153/330 (46%), Positives = 225/330 (67%), Gaps = 14/330 (4%)
Query: 79 NDVSPNNMDERPQK--SLLPPFESAEMRTLAESLSRDIIRGSPDVKWESIKGLENAKRLL 136
+D NN + P++ P A++ L + L RDI++ +P++ W+ I L AKRLL
Sbjct: 168 DDEEGNNGGDTPEEVERKFEPASHADV-DLVDMLERDILQKNPNIHWDDIADLHEAKRLL 226
Query: 137 KEAVVMPIKYPKYFTGLLSPWKGILLFGPPGTGKTMLAKAVATECKTTFFNISASSVVSK 196
+EAVV+P+ P YF G+ PWKG+L+ GPPGTGKTMLAKAVATEC TTFFN+S+S++ SK
Sbjct: 227 EEAVVLPMWMPDYFKGIRRPWKGVLMVGPPGTGKTMLAKAVATECGTTFFNVSSSTLTSK 286
Query: 197 WRGDSEKLVKVLFQLARHHAPSTIFLDEIDAIISQRGEARSEHEASRRLKTELLIQMDGL 256
+RG+SEKLV++LF++AR +APSTIF+DEID++ S+RG + SEHEASRR+K+ELL+QMDG+
Sbjct: 287 YRGESEKLVRLLFEMARFYAPSTIFIDEIDSLCSRRG-SESEHEASRRVKSELLVQMDGV 345
Query: 257 TRTD--ELVFVLAATNLPWELDAAMLRRLEKRILVPLPEPEARVAMFEELLPPQPDEESI 314
+ + ++V VLAATN PW++D A+ RRLEKRI +PLP E R A+ + L +ES+
Sbjct: 346 SNDEATKIVMVLAATNFPWDIDEALRRRLEKRIYIPLPNSEGREALLKINLREVKVDESV 405
Query: 315 PYDLLVNQTEGYSGSDIRLLCKEVAMQPLRRLMSQL--EQREDLVPEE-ELPKVGPIRPE 371
+ ++ +GYSG+DI +C++ +M +RR ++ L EQ L EE +L P+ +
Sbjct: 406 DMRDIADRLDGYSGADITNVCRDASMMSMRRKIAGLRPEQIRQLAKEELDL----PVSKQ 461
Query: 372 DIQAALKNTRPSAHL-HAHKYDKFNADYGS 400
D + A+ S KY ++ ++GS
Sbjct: 462 DFKEAISKCNKSVSKDDLAKYQQWMKEFGS 491
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.315 0.133 0.381
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 678,683,270
Number of Sequences: 2790947
Number of extensions: 28158897
Number of successful extensions: 113032
Number of sequences better than 10.0: 4735
Number of HSP's better than 10.0 without gapping: 2700
Number of HSP's successfully gapped in prelim test: 2036
Number of HSP's that attempted gapping in prelim test: 103373
Number of HSP's gapped (non-prelim): 5790
length of query: 404
length of database: 848,049,833
effective HSP length: 130
effective length of query: 274
effective length of database: 485,226,723
effective search space: 132952122102
effective search space used: 132952122102
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 76 (33.9 bits)
Lotus: description of TM0265.5


