

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0265.11
(1543 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
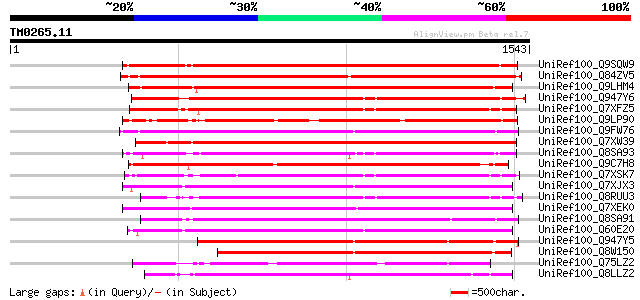
Sequences producing significant alignments: (bits) Value
UniRef100_Q9SQW9 Putative retroelement pol polyprotein [Arabidop... 1103 0.0
UniRef100_Q84ZV5 Polyprotein [Glycine max] 1086 0.0
UniRef100_Q9LHM4 Retroelement pol polyprotein [Arabidopsis thali... 1008 0.0
UniRef100_Q947Y6 Putative retroelement [Oryza sativa] 980 0.0
UniRef100_Q7XFZ5 Putative retroelement [Oryza sativa] 974 0.0
UniRef100_Q9LP90 T32E20.30 [Arabidopsis thaliana] 971 0.0
UniRef100_Q9FW76 Putative gag-pol polyprotein [Oryza sativa] 948 0.0
UniRef100_Q7XW39 OSJNBb0062H02.17 protein [Oryza sativa] 934 0.0
UniRef100_Q8SA93 Putative polyprotein [Zea mays] 932 0.0
UniRef100_Q9C7H8 Gypsy/Ty3 element polyprotein, putative [Arabid... 931 0.0
UniRef100_Q7XSK7 OSJNBa0059D20.8 protein [Oryza sativa] 930 0.0
UniRef100_Q7XJX3 OSJNBa0004L19.16 protein [Oryza sativa] 899 0.0
UniRef100_Q8RUU3 Putative gag-pol polyprotein [Oryza sativa] 893 0.0
UniRef100_Q7XEK0 Hypothetical protein [Oryza sativa] 888 0.0
UniRef100_Q8SA91 Putative gag-pol polyprotein [Zea mays] 881 0.0
UniRef100_Q60E20 Putative polyprotein [Oryza sativa] 879 0.0
UniRef100_Q947Y5 Putative retroelement [Oryza sativa] 863 0.0
UniRef100_Q8W150 Polyprotein [Oryza sativa] 836 0.0
UniRef100_Q75LZ2 Putative gag-pol polyprotein [Oryza sativa] 782 0.0
UniRef100_Q8LLZ2 Putative retroelement [Oryza sativa] 754 0.0
>UniRef100_Q9SQW9 Putative retroelement pol polyprotein [Arabidopsis thaliana]
Length = 1661
Score = 1103 bits (2854), Expect = 0.0
Identities = 567/1187 (47%), Positives = 775/1187 (64%), Gaps = 25/1187 (2%)
Query: 336 HKCPERAMRVLILGDGETMDEDGEIVMLEEEAEEIEEEE-----TVECKLMGVLGCMGEH 390
HKC + ++ L + T + + E ++EE E + EEE K+M + E
Sbjct: 453 HKCKPQKLKGLAI----TEEVEEESPLIEELNEPLTEEEGDPEPAEGFKVMTLSSLNDES 508
Query: 391 QTQTMKLAGQVANVELLILIDSGASHNFISPKITSALGLVITPITTRSIKLGDGHKVVTR 450
Q Q+MK+ G + N ++++L+DSGA+ NFIS + G ++T + +K+G G + +
Sbjct: 509 QEQSMKMRGYIGNTKVVLLVDSGATCNFISEALVREKGWLVTQTRSFGVKVGGGRIIKSS 568
Query: 451 GICRGIKAKLGSIEITIDALVLELGGLDLVLGVSWLSTLGKVIMDWKLLTMQFVYGEQVV 510
G C I ++ IE D + +LG LDLVLG SWL+ LG+ +W+ L + + G V
Sbjct: 569 GKCVDIPLEVQGIEFVQDYYLFDLGDLDLVLGFSWLAGLGETRANWRDLRISWQIGRTWV 628
Query: 511 QLQG----MRGKDSYQSNLHSFLSDTHGREQMEWWGPQLQVVEATKKGHE---IPQELRA 563
L G RG+ S +S + + T +E L + +KK E + ++
Sbjct: 629 SLYGDPDLCRGQISMRS-MERVIKYTGTAYLLE-----LASLFESKKQEEQTALQPAIQR 682
Query: 564 VLADFHEVFTKKIQLPPVRSRVHQITLNPEHGPINVRPYRYPHHQKEEIERQVTELLEAG 623
+L + VF LPPVR+R H ITL P+N+RPYRY QK EIE+ V E+L A
Sbjct: 683 LLDQYQGVFQTPQLLPPVRNREHAITLQEGSSPVNIRPYRYSFAQKNEIEKLVREMLNAQ 742
Query: 624 IIRPSMSSFSSPVILVKKKDKSWRMCVDYRALNKATIPDKYPIPIVDELLDELNGAVIFS 683
IIRPS+S +SSPV+LVKKKD WR CVDYRALN+ATIPDKYPIP+++ELLDEL GA +FS
Sbjct: 743 IIRPSVSPYSSPVLLVKKKDGGWRFCVDYRALNEATIPDKYPIPVIEELLDELKGATVFS 802
Query: 684 KIDLKSGYHQIRVHEEDIPKTTFRTHNGHYEYLVMPFGLMNAPATFQAIMNDIFRPYLRK 743
K+DLKSGY QIR+ D+ KT F+TH GHYE+LVMPFGL NAP+TFQ++MND+FRPYLRK
Sbjct: 803 KLDLKSGYFQIRMKLSDVEKTAFKTHEGHYEFLVMPFGLTNAPSTFQSVMNDLFRPYLRK 862
Query: 744 FVLVFFDDILIYSKSLLEHRDHLKLVLTVLLSNSFVANQSKCKFGCAQIDYLGHIISGAG 803
FVLVFFDDIL+YS + H HL+ VL +L + F AN KC FG +I YLGHIIS G
Sbjct: 863 FVLVFFDDILVYSPDMKTHLKHLETVLQLLHLHQFYANFKKCTFGSTRISYLGHIISEQG 922
Query: 804 VSVDPEKVQCIVEWPEPKNVKGVRGFLGLTGYYRKFIKNYGKMAKPLTELTKKDNFTWGE 863
V+ DPEKV+ +++WP PK+V +RGFLG TGYYR+F+KNYG++A+PL + KK++F W E
Sbjct: 923 VATDPEKVEAMLQWPLPKSVTELRGFLGFTGYYRRFVKNYGQIARPLRDQLKKNSFDWNE 982
Query: 864 EAKQAFHKLKTVMTSSPVLALPDFDKSFEVECDAAGRGIGAVLMQQRQPIAYFSKALSNG 923
A AF LK +++ PVL LPDF + F VE DA+G GIGAVL Q ++ IA+ S+A S+
Sbjct: 983 AATSAFQALKAAVSALPVLVLPDFQQEFTVETDASGMGIGAVLSQNKRLIAFLSQAFSSQ 1042
Query: 924 NLTKSVYEKELMALVLCIQHWRHYLLGKAFTVYTDHKSLKHFLQQKISSPDQQCWLAKLL 983
+SVYE+EL+A+V + W+HYL K F + TD +SL+H L+QK S QQ W +KL
Sbjct: 1043 GRIRSVYERELLAIVKAVTKWKHYLSSKEFIIKTDQRSLRHLLEQKSVSTIQQRWASKLS 1102
Query: 984 GYQFEVKYKPGQENKAADALSRRGDESDFFSL-VSFPLWDDRKKLLEEIASDPYIKNLLQ 1042
G ++ ++YKPG +NK ADALSRR L ++ P D L EI D + +L+
Sbjct: 1103 GLKYRIEYKPGVDNKVADALSRRPPTEALSQLTITGPPTIDLTALKAEIQQDHELSQILK 1162
Query: 1043 DVQQDPQSKPGFSVLQGVLLYQGRLVLSPTSPSIPWLLQEYHGSPTGGHSGFLRTYRRLA 1102
+ Q F+V G++ +G LV+ SP IP +L+++H SP GGH G L+T++RL
Sbjct: 1163 NWAQGDHHDSDFTVADGLIYRKGCLVIPVGSPFIPKMLEKFHTSPIGGHEGALKTFKRLT 1222
Query: 1103 ESLYWVGMQRTVRDFVRACDTCQRQKYSAMSPGGLLQPFPVPNAVWEDLSLDFIIGLPKS 1162
+YW G+++ V ++++ C CQ KYS +SP GLL P P+P +W D+SLDF+ GLP S
Sbjct: 1223 SEVYWRGLRKDVVNYIKGCQICQENKYSTLSPAGLLSPLPIPQQIWSDVSLDFVEGLPSS 1282
Query: 1163 KGYEAILVVVDRLSKYSHFILLKHPYTAKSIAEVFVREIVRLHGIPNTVKSDRDPIFVSH 1222
+ ILVVVDRLSKYSHFI LKHP+TAK++ E F+R++V+LHG PNT+ SDRD IF+S
Sbjct: 1283 NRFNCILVVVDRLSKYSHFIPLKHPFTAKTVVEAFIRDVVKLHGFPNTLVSDRDRIFLSG 1342
Query: 1223 FWSELFKLQGTKLKMSSSYHPETDGQTEVINRCLESYLRCFASDHPKTWSFWVPWAEFWY 1282
FWSELFKLQGT L+ S++YHP+TDGQTEV+NRCLESYLRCFA P +W W+PWAE+WY
Sbjct: 1343 FWSELFKLQGTGLQKSTAYHPQTDGQTEVVNRCLESYLRCFAGRRPTSWFQWLPWAEYWY 1402
Query: 1283 NSTFHCSIGQTPFEVVYGRKAPPIIRFLTNETKVVAVALELSERDEALKQLRTHLQRAQE 1342
N+++H + TPF+ VYGR+ P ++R+ T V L +RD L +LR +L+ AQ
Sbjct: 1403 NTSYHSATKTTPFQAVYGREPPVLLRYGDIPTNNANVEELLKDRDGMLVELRENLEIAQA 1462
Query: 1343 QMASYANKKRRDLSFDIGEWVFLKLRPHRQHSVVKRINQKLAARFYGPFQIEDKIGTVAY 1402
QM A+K RRD++F+I EWV+LKLRP+RQ SV R N+KL+ R++GPF++ +IG VAY
Sbjct: 1463 QMKKAADKSRRDVAFEIDEWVYLKLRPYRQSSVAHRKNEKLSQRYFGPFKVLHRIGQVAY 1522
Query: 1403 KLKLPVESRIHPVFHVSLLKRAIGNYQVQGQLPTDLGIEEATDIYPEAILGTRIIRQGDS 1462
KL+LP S IHPVFHVS LKRA+ +LP L + PE +L R + +
Sbjct: 1523 KLQLPEHSTIHPVFHVSQLKRAVPPSFTPQELPKILSPTLEWNTGPEKLLDIR--QSNTN 1580
Query: 1463 EVHQSLIKWKNRSIDDVTWEDNEVLAGQFPEFCLEDKAVSKEGGIER 1509
+ L++W S + TWE L Q+P+F LEDK G I+R
Sbjct: 1581 SGPEVLVQWSGLSTLESTWEPLLTLVQQYPDFDLEDKVSLLRGSIDR 1627
>UniRef100_Q84ZV5 Polyprotein [Glycine max]
Length = 1552
Score = 1086 bits (2808), Expect = 0.0
Identities = 563/1198 (46%), Positives = 769/1198 (63%), Gaps = 25/1198 (2%)
Query: 330 KISPTLHKCPERAMRVLILGDGETMDEDGEIVMLEEEAEEIEEEETVECKLMGVLGCMGE 389
K SP HKCP R + +L L + + D+ E VM+ EEA ++ + M G
Sbjct: 333 KFSPA-HKCPNRQVMLLQLEETDE-DQTDEQVMVTEEANMDDDTHHLSLNAM-----RGS 385
Query: 390 HQTQTMKLAGQVANVELLILIDSGASHNFISPKITSALGLVITPITTRSIKLGDGHKVVT 449
+ T++ GQV + + IL+D G+S NFI P++ L L + P + +G+G +
Sbjct: 386 NGVGTIRFTGQVGGIAVKILVDGGSSDNFIQPRVAQVLKLPVEPAPNLRVLVGNGQILSA 445
Query: 450 RGICRGIKAKLGSIEITIDALVLELGGLDLVLGVSWLSTLGKVIMDWKLLTMQFVYGEQV 509
GI + + + E+ + +L++ G D++LG +WL+TLG + D+ LT++F ++
Sbjct: 446 EGIVQQLPLHIQGQEVKVPVYLLQISGADVILGSTWLATLGPHVADYAALTLKFFQNDKF 505
Query: 510 VQLQGMRGKDSYQSNLHSFLSDTHGREQMEWWGPQL---QVVEATKKG--HEIPQELRAV 564
+ LQG ++ Q+ LH F + + E + QL +V E T K I EL +
Sbjct: 506 ITLQGEGNSEATQAQLHHFRRLQNTKSIEECFAIQLIQKEVPEDTLKDLPTNIDPELAIL 565
Query: 565 LADFHEVFTKKIQLPPVRSRVHQITLNPEHGPINVRPYRYPHHQKEEIERQVTELLEAGI 624
L + +VF LPP R + H I L GP+ VRPYRYPH QK++IE+ + E+L GI
Sbjct: 566 LHTYAQVFAVPASLPPQREQDHAIPLKQGSGPVKVRPYRYPHTQKDQIEKMIQEMLVQGI 625
Query: 625 IRPSMSSFSSPVILVKKKDKSWRMCVDYRALNKATIPDKYPIPIVDELLDELNGAVIFSK 684
I+PS S FS P++LVKKKD SWR C DYRALN T+ D +P+P VDELLDEL+GA FSK
Sbjct: 626 IQPSNSPFSLPILLVKKKDGSWRFCTDYRALNAITVKDSFPMPTVDELLDELHGAQYFSK 685
Query: 685 IDLKSGYHQIRVHEEDIPKTTFRTHNGHYEYLVMPFGLMNAPATFQAIMNDIFRPYLRKF 744
+DL+SGYHQI V ED KT FRTH+GHYE+LVMPFGL NAPATFQ +MN IF+ LRKF
Sbjct: 686 LDLRSGYHQILVQPEDREKTAFRTHHGHYEWLVMPFGLTNAPATFQCLMNKIFQFALRKF 745
Query: 745 VLVFFDDILIYSKSLLEHRDHLKLVLTVLLSNSFVANQSKCKFGCAQIDYLGHIISGAGV 804
VLVFFDDILIYS S +H HL+ VL L + A SKC FG ++DYLGH +SG GV
Sbjct: 746 VLVFFDDILIYSASWKDHLKHLESVLQTLKQHQLFARLSKCSFGDTEVDYLGHKVSGLGV 805
Query: 805 SVDPEKVQCIVEWPEPKNVKGVRGFLGLTGYYRKFIKNYGKMAKPLTELTKKDNFTWGEE 864
S++ KVQ +++WP P NVK +RGFLGLTGYYR+FIK+Y +A PLT+L +KD+F W E
Sbjct: 806 SMENTKVQAVLDWPTPNNVKQLRGFLGLTGYYRRFIKSYANIAGPLTDLLQKDSFLWNNE 865
Query: 865 AKQAFHKLKTVMTSSPVLALPDFDKSFEVECDAAGRGIGAVLMQQRQPIAYFSKALSNGN 924
A+ AF KLK MT +PVL+LPDF + F +E DA+G G+GAVL Q PIAYFSK L+
Sbjct: 866 AEAAFVKLKKAMTEAPVLSLPDFSQPFILETDASGIGVGAVLGQNGHPIAYFSKKLAPRM 925
Query: 925 LTKSVYEKELMALVLCIQHWRHYLLGKAFTVYTDHKSLKHFLQQKISSPDQQCWLAKLLG 984
+S Y +EL+A+ + +RHYLLG F + TD +SLK + Q + +P+QQ WL K LG
Sbjct: 926 QKQSAYTRELLAITEALSKFRHYLLGNKFIIRTDQRSLKSLMDQSLQTPEQQAWLHKFLG 985
Query: 985 YQFEVKYKPGQENKAADALSRRGDESDFFSLVSFPLWDDRKKLLEEIASDPYIKNLLQDV 1044
Y F+++YKPG++N+AADALSR F S P ++L + SDP++K L++
Sbjct: 986 YDFKIEYKPGKDNQAADALSRM-----FMLAWSEPHSIFLEELRARLISDPHLKQLMETY 1040
Query: 1045 QQDPQSKPGFSVLQGVLLYQGRLVLSPTSPSIPWLLQEYHGSPTGGHSGFLRTYRRLAES 1104
+Q + ++V +G+L ++ R+V+ + +LQEYH SP GGH+G RT RL
Sbjct: 1041 KQGADAS-HYTVREGLLYWKDRVVIPAEEEIVNKILQEYHSSPIGGHAGITRTLARLKAQ 1099
Query: 1105 LYWVGMQRTVRDFVRACDTCQRQKYSAMSPGGLLQPFPVPNAVWEDLSLDFIIGLPKSKG 1164
YW MQ V+ +++ C CQ+ K + P GLLQP P+P VWED+++DFI GLP S G
Sbjct: 1100 FYWPKMQEDVKAYIQKCLICQQAKSNNTLPAGLLQPLPIPQQVWEDVAMDFITGLPNSFG 1159
Query: 1165 YEAILVVVDRLSKYSHFILLKHPYTAKSIAEVFVREIVRLHGIPNTVKSDRDPIFVSHFW 1224
I+VV+DRL+KY+HFI LK Y +K +AE F+ IV+LHGIP ++ SDRD +F S FW
Sbjct: 1160 LSVIMVVIDRLTKYAHFIPLKADYNSKVVAEAFMSHIVKLHGIPRSIVSDRDRVFTSTFW 1219
Query: 1225 SELFKLQGTKLKMSSSYHPETDGQTEVINRCLESYLRCFASDHPKTWSFWVPWAEFWYNS 1284
LFKLQGT L MSS+YHP++DGQ+EV+N+CLE YLRCF +HPK W +PWAEFWYN+
Sbjct: 1220 QHLFKLQGTTLAMSSAYHPQSDGQSEVLNKCLEMYLRCFTYEHPKGWVKALPWAEFWYNT 1279
Query: 1285 TFHCSIGQTPFEVVYGRKAPPIIRFLTNETKVVAVALELSERDEALKQLRTHLQRAQEQM 1344
+H S+G TPF +YGR+ P + R + V +L++RD L +L+ +L RAQ+ M
Sbjct: 1280 AYHMSLGMTPFRALYGREPPTLTRQACSIDDPAEVREQLTDRDALLAKLKINLTRAQQVM 1339
Query: 1345 ASYANKKRRDLSFDIGEWVFLKLRPHRQHSVVKRINQKLAARFYGPFQIEDKIGTVAYKL 1404
A+KKR D+SF IG+ V +KL+P+RQHS V R NQKL+ R++GPF++ KIG VAYKL
Sbjct: 1340 KRQADKKRLDVSFQIGDEVLVKLQPYRQHSAVLRKNQKLSMRYFGPFKVLAKIGDVAYKL 1399
Query: 1405 KLPVESRIHPVFHVSLLKRAIGNYQVQGQLPTDLGIEEATDI-YPEAILGTRIIRQGDSE 1463
+LP +RIHPVFHVS LK G Q LP L + E + P IL +RII +G ++
Sbjct: 1400 ELPSAARIHPVFHVSQLKPFNGTAQ-DPYLPLPLTVTEMGPVMQPVKILASRIIIRGHNQ 1458
Query: 1464 VHQSLIKWKNRSIDDVTWEDNEVLAGQFPEFCLEDKAVSKEGGIERDSNTKWGMGLGK 1521
+ Q L++W+N D+ TWED E + +P F LEDK V K G N GM G+
Sbjct: 1459 IEQILVQWENGLQDEATWEDIEDIKASYPTFNLEDKVVFKGEG-----NVTNGMSRGE 1511
>UniRef100_Q9LHM4 Retroelement pol polyprotein [Arabidopsis thaliana]
Length = 1499
Score = 1008 bits (2607), Expect = 0.0
Identities = 506/1152 (43%), Positives = 730/1152 (62%), Gaps = 17/1152 (1%)
Query: 354 MDEDGEIVMLEEEAEEIEEEETVECKLMGVLGCMGEHQTQTMKLAGQVANVELLILIDSG 413
MD D E EE ++E + + V G G +TM++ G + ILIDSG
Sbjct: 352 MDVDEEFEDAREELVNDDDEHMPQISVNAVSGIAGY---KTMRVKGTYDKKIIFILIDSG 408
Query: 414 ASHNFISPKITSALGLVITPITTRSIKLGDGHKVVTRGICRGIKAKLGSIEITIDALVLE 473
++HNF+ P + LG + + + DG K+ G KL + D L++
Sbjct: 409 STHNFLDPNTAAKLGCKVDTAGLTRVSVADGRKLRVEGKVTDFSWKLQTTTFQSDILLIP 468
Query: 474 LGGLDLVLGVSWLSTLGKVIMDWKLLTMQFVYGEQVVQLQGMRGKDSYQSNLHSFLSDTH 533
L G+D+VLGV WL TLG++ ++K L M+F + Q V L G+ +
Sbjct: 469 LQGIDMVLGVQWLETLGRISWEFKKLEMRFKFNNQKVLLHGLTSGSVREVKAQKLQKLQE 528
Query: 534 GREQMEWWGPQLQVVEATKKG--------HEIPQE--LRAVLADFHEVFTKKIQLPPVRS 583
+ Q+ Q +V E+T+ E+ +E + VL ++ ++F + LPP R
Sbjct: 529 DQVQLAMLCVQ-EVSESTEGELCTINALTSELGEESVVEEVLNEYPDIFIEPTALPPFRE 587
Query: 584 RV-HQITLNPEHGPINVRPYRYPHHQKEEIERQVTELLEAGIIRPSMSSFSSPVILVKKK 642
+ H+I L P+N RPYRY HQK EI++ V +LL G ++ S S ++SPV+LVKKK
Sbjct: 588 KHNHKIKLLEGSNPVNQRPYRYSIHQKNEIDKLVEDLLTNGTVQASSSPYASPVVLVKKK 647
Query: 643 DKSWRMCVDYRALNKATIPDKYPIPIVDELLDELNGAVIFSKIDLKSGYHQIRVHEEDIP 702
D +WR+CVDYR LN T+ D +PIP++++L+DEL GAVIFSKIDL++GYHQ+R+ +DI
Sbjct: 648 DGTWRLCVDYRELNGMTVKDSFPIPLIEDLMDELGGAVIFSKIDLRAGYHQVRMDPDDIQ 707
Query: 703 KTTFRTHNGHYEYLVMPFGLMNAPATFQAIMNDIFRPYLRKFVLVFFDDILIYSKSLLEH 762
KT F+TH+GH+EYLVMPFGL NAPATFQ +MN IF+P+LRKFVLVFFDDIL+YS SL EH
Sbjct: 708 KTAFKTHSGHFEYLVMPFGLTNAPATFQGLMNFIFKPFLRKFVLVFFDDILVYSSSLEEH 767
Query: 763 RDHLKLVLTVLLSNSFVANQSKCKFGCAQIDYLGHIISGAGVSVDPEKVQCIVEWPEPKN 822
R HLK V V+ +N A SKC F +++YLGH IS G+ DP K++ + EWP+P
Sbjct: 768 RQHLKQVFEVMRANKLFAKLSKCAFAVPKVEYLGHFISAQGIETDPAKIKAVKEWPQPTT 827
Query: 823 VKGVRGFLGLTGYYRKFIKNYGKMAKPLTELTKKDNFTWGEEAKQAFHKLKTVMTSSPVL 882
+K +RGFLGL GYYR+F++++G +A PL LTK D F W A+QAF LK + +PVL
Sbjct: 828 LKQLRGFLGLAGYYRRFVRSFGVIAGPLHALTKTDAFEWTAVAQQAFEDLKAALCQAPVL 887
Query: 883 ALPDFDKSFEVECDAAGRGIGAVLMQQRQPIAYFSKALSNGNLTKSVYEKELMALVLCIQ 942
+LP FDK F VE DA G+GIGAVLMQ+ P+AY S+ L L S+YEKEL+A++ ++
Sbjct: 888 SLPLFDKQFVVETDACGQGIGAVLMQEGHPLAYISRQLKGKQLHLSIYEKELLAVIFAVR 947
Query: 943 HWRHYLLGKAFTVYTDHKSLKHFLQQKISSPDQQCWLAKLLGYQFEVKYKPGQENKAADA 1002
WRHYLL F + TD +SLK+ L+Q++++P QQ WL KLL + +E++Y+ G+EN ADA
Sbjct: 948 KWRHYLLQSHFIIKTDQRSLKYLLEQRLNTPIQQQWLPKLLEFDYEIQYRQGKENVVADA 1007
Query: 1003 LSRRGDESDFFSLVSFPLWDDRKKLLEEIASDPYIKNLLQDVQQDPQSKPGFSVLQGVLL 1062
LSR ++ D K + A+D +++++ +Q+DP SK FS Q +L
Sbjct: 1008 LSRVEGSEVLHMAMTVVECDLLKDIQAGYANDSQLQDIITALQRDPDSKKYFSWSQNILR 1067
Query: 1063 YQGRLVLSPTSPSIPWLLQEYHGSPTGGHSGFLRTYRRLAESLYWVGMQRTVRDFVRACD 1122
+ ++V+ +L HGS GGHSG T++R+ YW GM + ++ ++R+C
Sbjct: 1068 RKSKIVVPANDNIKNTILLWLHGSGVGGHSGRDVTHQRVKGLFYWKGMIKDIQAYIRSCG 1127
Query: 1123 TCQRQKYSAMSPGGLLQPFPVPNAVWEDLSLDFIIGLPKSKGYEAILVVVDRLSKYSHFI 1182
TCQ+ K + GLLQP P+P+ +W ++S+DFI GLP S G I+VVVDRLSK +HFI
Sbjct: 1128 TCQQCKSDPAASPGLLQPLPIPDTIWSEVSMDFIEGLPVSGGKTVIMVVVDRLSKAAHFI 1187
Query: 1183 LLKHPYTAKSIAEVFVREIVRLHGIPNTVKSDRDPIFVSHFWSELFKLQGTKLKMSSSYH 1242
L HPY+A ++A ++ + +LHG P ++ SDRD +F S FW E F LQG LK++S+YH
Sbjct: 1188 ALSHPYSALTVAHAYLDNVFKLHGCPTSIVSDRDVVFTSEFWREFFTLQGVALKLTSAYH 1247
Query: 1243 PETDGQTEVINRCLESYLRCFASDHPKTWSFWVPWAEFWYNSTFHCSIGQTPFEVVYGRK 1302
P++DGQTEV+NRCLE+YLRC D P+ WS W+ AE+WYN+ +H S TPFE+VYG+
Sbjct: 1248 PQSDGQTEVVNRCLETYLRCMCHDRPQLWSKWLALAEYWYNTNYHSSSRMTPFEIVYGQV 1307
Query: 1303 APPIIRFLTNETKVVAVALELSERDEALKQLRTHLQRAQEQMASYANKKRRDLSFDIGEW 1362
P + +L E+KV VA L ER++ L L+ HL RAQ +M +A++ R + F+IG++
Sbjct: 1308 PPVHLPYLPGESKVAVVARSLQEREDMLLFLKFHLMRAQHRMKQFADQHRTEREFEIGDY 1367
Query: 1363 VFLKLRPHRQHSVVKRINQKLAARFYGPFQIEDKIGTVAYKLKLPVESRIHPVFHVSLLK 1422
V++KL+P+RQ SVV R NQKL+ +++GP++I D+ G VAYKL LP S++HPVFHVS LK
Sbjct: 1368 VYVKLQPYRQQSVVMRANQKLSPKYFGPYKIIDRCGEVAYKLALPSYSQVHPVFHVSQLK 1427
Query: 1423 RAIGNYQVQGQLPTDLGIEEATDIYPEAILGTRIIRQGDSEVHQSLIKWKNRSIDDVTWE 1482
+GN LP+ + ++ + PE ++ +++ + V + L+KW N +++ TWE
Sbjct: 1428 VLVGNVSTTVHLPSVM--QDVFEKVPEKVVERKMVNRQGKAVTKVLVKWSNEPLEEATWE 1485
Query: 1483 DNEVLAGQFPEF 1494
L FPEF
Sbjct: 1486 FLFDLQKTFPEF 1497
>UniRef100_Q947Y6 Putative retroelement [Oryza sativa]
Length = 1461
Score = 980 bits (2533), Expect = 0.0
Identities = 516/1172 (44%), Positives = 704/1172 (60%), Gaps = 53/1172 (4%)
Query: 363 LEEEAEEIEEEET-VECKLMGVLGCMGEHQTQTMKLAGQVANVELLILIDSGASHNFISP 421
+E+E EE E+E E + + G + L Q+ L+ L+D+G++HNFI
Sbjct: 311 VEDEDEEAPEDEVDAEAPVFSLHAVAGVAVGHPILLRVQLGATTLVALVDTGSTHNFIGE 370
Query: 422 KITSALGLVITPITTRSIKLGDGHKVVTRGICRGIKAKLGSIEITIDALVLELGGLDLVL 481
+ GL + P + + +G KV G+ R + + +D V+ L G D+VL
Sbjct: 371 SAAARTGLSVQPRPRMTATVANGEKVACPGVLRHAPITIEGMPFHVDLYVMPLAGYDIVL 430
Query: 482 GVSWLSTLGKVIMDWKLLTMQFVYGEQVVQLQGMRGKDSYQSNLHSFLSDTHGREQMEWW 541
G W++ LG+ M W + T + D GR + W
Sbjct: 431 GTQWMAKLGR--MSWDVTTRALTF-------------------------DLEGRT-ICWQ 462
Query: 542 GPQLQVVEATKKGHEIPQELRAVLADFHEVFTKKIQLPPVRSRVHQITLNPEHGPINVRP 601
G Q A + L +L F +VFT+ LPP R R H I L P+ VRP
Sbjct: 463 GAPNQDGPAVRAASADDSLLGGLLDSFADVFTEPTGLPPQRGRDHAIVLKQGTSPVAVRP 522
Query: 602 YRYPHHQKEEIERQVTELLEAGIIRPSMSSFSSPVILVKKKDKSWRMCVDYRALNKATIP 661
YRYP K+E+ERQ ++ GI+R S S+FSSPV+LVKK D SWR CVDYRALN T+
Sbjct: 523 YRYPAAHKDELERQCAAMISQGIVRRSDSAFSSPVLLVKKADSSWRFCVDYRALNALTVK 582
Query: 662 DKYPIPIVDELLDELNGAVIFSKIDLKSGYHQIRVHEEDIPKTTFRTHNGHYEYLVMPFG 721
D +PIP+VDELLDEL+GA FSK+DL+SGYHQ+R+ EDI KT FRTH+G YE+LVMPFG
Sbjct: 583 DAFPIPVVDELLDELHGARFFSKLDLRSGYHQVRMRPEDIHKTAFRTHDGLYEFLVMPFG 642
Query: 722 LMNAPATFQAIMNDIFRPYLRKFVLVFFDDILIYSKSLLEHRDHLKLVLTVLLSNSFVAN 781
L NAPATFQA+MND+ R +LR+FVLVFFDDILIYS + +H HL+ VLTVL +
Sbjct: 643 LCNAPATFQALMNDVLRSFLRRFVLVFFDDILIYSDTWADHLRHLRAVLTVLREHKLFIK 702
Query: 782 QSKCKFGCAQIDYLGHIISGAGVSVDPEKVQCIVEWPEPKNVKGVRGFLGLTGYYRKFIK 841
+SKC FG + YLGH+IS AGV++DP KVQ I EWP+P++ + VRGFLGL GYYRKF+
Sbjct: 703 RSKCAFGVDSVAYLGHVISAAGVAMDPAKVQAIREWPQPRSARAVRGFLGLAGYYRKFVH 762
Query: 842 NYGKMAKPLTELTKKDNFTWGEEAKQAFHKLKTVMTSSPVLALPDFDKSFEVECDAAGRG 901
NYG +A PLT L KK+ F W E A AF LK ++S+P+LA+PDF K+F VECDA+ G
Sbjct: 763 NYGTIAAPLTALLKKEGFAWTEAATAAFDALKAAVSSAPILAMPDFTKAFTVECDASSHG 822
Query: 902 IGAVLMQQRQPIAYFSKALSNGNLTKSVYEKELMALVLCIQHWRHYLLGKAFTVYTDHKS 961
GAVL+Q P+A+FS+ ++ + + YE+EL+ LVL ++HWR YL G+ FTV TDH S
Sbjct: 823 FGAVLIQDGHPLAFFSRPVAPRHRALAAYERELIGLVLAVRHWRPYLWGRHFTVKTDHYS 882
Query: 962 LKHFLQQKISSPDQQCWLAKLLGYQFEVKYKPGQENKAADALSRRGDESDFFSLV-SFPL 1020
LK+ L Q++S+ Q W+ KLLG+ F V+YKPG N ADALSRR D LV S P
Sbjct: 883 LKYLLDQRLSTIPQHHWVGKLLGFDFTVEYKPGAANTVADALSRRDTTEDASVLVLSAPR 942
Query: 1021 WDDRKKLLEEIASDPYIKNLLQDVQQDPQSKPGFSVLQGVLLYQGRLVLSPTSPSIPWLL 1080
+D ++L + DP + L +++ ++ P +S+ G++L+ GRL L P SP + +L
Sbjct: 943 FDFIERLRQAQDVDPALVALQAEIRSGTRAGP-WSMADGMVLFAGRLYLPPASPLLQEVL 1001
Query: 1081 QEYHGSPTGGHSGFLRTYRRLAESLYWVGMQRTVRDFVRACDTCQRQKYSAMSPGGLLQP 1140
+ H GH G RT RL ++ M+ V+DFVR C+ CQR K + P GLL P
Sbjct: 1002 RAVHEE---GHEGVQRTLHRLRRDFHFPNMKSVVQDFVRTCEVCQRYKAEHLQPAGLLLP 1058
Query: 1141 FPVPNAVWEDLSLDFIIGLPKSKGYEAILVVVDRLSKYSHFILLKHPYTAKSIAEVFVRE 1200
PVP VW D++LDF+ LP+ +G IL VVDR SKY HFI L HPY+A+S+A+VF E
Sbjct: 1059 LPVPQGVWTDVALDFVEALPRVRGKSVILTVVDRFSKYCHFIPLAHPYSAESVAQVFFAE 1118
Query: 1201 IVRLHGIPNTVKSDRDPIFVSHFWSELFKLQGTKLKMSSSYHPETDGQTEVINRCLESYL 1260
IVRLHG+P ++ SDRDP+F S FWSEL +L GTKL M++++HP++DGQ+E NR + YL
Sbjct: 1119 IVRLHGVPQSMVSDRDPVFTSAFWSELMRLVGTKLHMTTAFHPQSDGQSEAANRVIIMYL 1178
Query: 1261 RCFASDHPKTWSFWVPWAEFWYNSTFHCSIGQTPFEVVYGRKAPPIIRFLTNETKVVAVA 1320
RC D P+ W W+PWAEF +N+ + S+ TPF VVYGR P I + +T+V AVA
Sbjct: 1179 RCLTGDRPRQWLRWLPWAEFVFNTAYQTSLRDTPFRVVYGRDPPSIRSYEPGDTRVAAVA 1238
Query: 1321 LELSERDEALKQLRTHLQRAQEQMASYANKKRRDLSFDIGEWVFLKLRPHRQHSVVKRIN 1380
+ ER E L+ +R L++AQ Y +K R +SF +G+WV L+LR S+ ++
Sbjct: 1239 KSMEERSEFLEDIRYRLEQAQAIQKKYYDKSHRAVSFQVGDWVLLRLRQRAPASLSLAVS 1298
Query: 1381 QKLAARFYGPFQIEDKIGTVAYKLKLPVESRIHPVFHVSLLKRAIGNYQVQGQLPTDLGI 1440
KL R++GP++I + I VA +L LP +R+H VFH+ LLK+ G P
Sbjct: 1299 GKLKPRYFGPYRIAEMINEVAARLALPAGARLHDVFHIGLLKKWHG-APPDAPPPLPNVH 1357
Query: 1441 EEATDIYPEAILGTRIIRQGDSEVHQSLIKWKNRSIDDVTWEDNEVLAGQFPEFCLEDKA 1500
A PE ++ R+ R V Q L++WK S TWED E ++P LED+
Sbjct: 1358 HGAVACEPERVIKARLAR----GVRQVLVQWKGTSAASATWEDREPFFARYPALQLEDEL 1413
Query: 1501 VSKEGGIERDSNTKWGMGLGKPKIWKVYVRKK 1532
GG + WG + YVR++
Sbjct: 1414 PLDGGG-----DVMWG---------RTYVRRR 1431
>UniRef100_Q7XFZ5 Putative retroelement [Oryza sativa]
Length = 1476
Score = 974 bits (2519), Expect = 0.0
Identities = 513/1157 (44%), Positives = 702/1157 (60%), Gaps = 31/1157 (2%)
Query: 356 EDGEIVMLEEEAEEIEEEETVECKLMGVLGCMGEHQTQTMKLAGQVANVELLILIDSGAS 415
E G I ++ E+ EE TVE + + G + + L + L+ L+D+G++
Sbjct: 300 EGGAIEEGDDTVEDDTEEATVEAPVFSLHAVAGIPLGKPILLQVTLGAASLVALVDTGST 359
Query: 416 HNFISPKITSALGLVITPITTRSIKLGDGHKVVTRGICRGIKAKLGSIEITIDALVLELG 475
HNFI GL + P + + +G KV G+ R + + +D V+ L
Sbjct: 360 HNFIGEDAALRTGLPVQPRPRLTATVANGEKVSCPGVLRRAPITIQGMAFDVDLYVMPLA 419
Query: 476 GLDLVLGVSWLSTLGKVIMDWKLLTMQFVYGEQVVQLQGMRGKDSYQSNLHSFLSDTHGR 535
G D+VLG W++ LG I W + T V Q S+QS H R
Sbjct: 420 GYDMVLGTQWMAHLGTTIA-WDVTT-------GTVSFQHQGRTVSWQS------LPPHQR 465
Query: 536 EQMEWWGPQLQVVEATKKGHEIPQE------LRAVLADFHEVFTKKIQLPPVRSRVHQIT 589
+ +V AT P L +L F +VF + LPP R R H I
Sbjct: 466 ADVHAVSTGTSLVAATGSSSSTPAPTTEPALLDGLLGSFDDVFAEPRGLPPPRGRDHAIH 525
Query: 590 LNPEHGPINVRPYRYPHHQKEEIERQVTELLEAGIIRPSMSSFSSPVILVKKKDKSWRMC 649
L P P+ VRPYRYP K+E+ERQ ++E G+IR S S+FSSPV+LVKK D SWR C
Sbjct: 526 LLPGAPPVAVRPYRYPVAHKDELERQCAVMMEQGLIRRSTSAFSSPVLLVKKADGSWRFC 585
Query: 650 VDYRALNKATIPDKYPIPIVDELLDELNGAVIFSKIDLKSGYHQIRVHEEDIPKTTFRTH 709
VDYRALN TI D YPIP+VDELLDEL+GA F+K+DL+SGYHQ+R+ ED+ KT FRTH
Sbjct: 586 VDYRALNAITIKDAYPIPVVDELLDELHGAKFFTKLDLRSGYHQVRMRAEDVAKTAFRTH 645
Query: 710 NGHYEYLVMPFGLMNAPATFQAIMNDIFRPYLRKFVLVFFDDILIYSKSLLEHRDHLKLV 769
+G YE+LVMPFGL NAPATFQA+MNDI R YLR+FVLVFFDDILIYS + +H H++ V
Sbjct: 646 DGLYEFLVMPFGLCNAPATFQALMNDILRIYLRRFVLVFFDDILIYSNTWADHLRHIRAV 705
Query: 770 LTVLLSNSFVANQSKCKFGCAQIDYLGHIISGAGVSVDPEKVQCIVEWPEPKNVKGVRGF 829
L +L + +SKC FG + I YLGHII GVS+DP KVQ +V+WP+P++ + VRGF
Sbjct: 706 LLLLRQHRLFVKRSKCAFGVSSISYLGHIIGATGVSMDPAKVQAVVDWPQPRSARTVRGF 765
Query: 830 LGLTGYYRKFIKNYGKMAKPLTELTKKDNFTWGEEAKQAFHKLKTVMTSSPVLALPDFDK 889
LGL GYYRKF+ +YG +A PLT LTKK+ F W +E AFH LK +T++PVLALPDF K
Sbjct: 766 LGLAGYYRKFVHDYGTIAAPLTALTKKEGFRWSDEVATAFHALKHAVTTAPVLALPDFVK 825
Query: 890 SFEVECDAAGRGIGAVLMQQRQPIAYFSKALSNGNLTKSVYEKELMALVLCIQHWRHYLL 949
F VECDA+ G GAVL+Q + P+A+FS+ ++ + + YE+EL+ LVL I+HWR YL
Sbjct: 826 PFVVECDASTHGFGAVLLQDKHPLAFFSRPVAPRHRALAAYERELIGLVLAIRHWRPYLW 885
Query: 950 GKAFTVYTDHKSLKHFLQQKISSPDQQCWLAKLLGYQFEVKYKPGQENKAADALSRRGDE 1009
G+AF V TDH SLK+ L Q++++ Q W+ KLLG+ F V+YK G N ADALSRR +
Sbjct: 886 GRAFVVRTDHYSLKYLLDQRLATIPQHHWVGKLLGFDFTVEYKSGASNVVADALSRRDTD 945
Query: 1010 SDFFSLVSFPLWDDRKKLLEEIASDPYIKNLLQDVQQDPQSKPGFSVLQGVLLYQGRLVL 1069
+S P +D ++L ++P + + +Q +S P +++ G++++ RL +
Sbjct: 946 EGAVLALSAPRFDYIERLRAAQTTEPALVAIRDAIQAGTRSAP-WALRDGMVMFDSRLYI 1004
Query: 1070 SPTSPSIPWLLQEYHGSPTGGHSGFLRTYRRLAESLYWVGMQRTVRDFVRACDTCQRQKY 1129
P+SP + +L H T GH G RT RL + M+R V++FVRACDTCQR K
Sbjct: 1005 PPSSPLLHEILAAIH---TDGHEGVQRTLHRLRRDFHSPAMRRVVQEFVRACDTCQRNKS 1061
Query: 1130 SAMSPGGLLQPFPVPNAVWEDLSLDFIIGLPKSKGYEAILVVVDRLSKYSHFILLKHPYT 1189
+ PGGLL P PVP VW D+ LDF+ LP+ G IL VVDR SKY HFI L HPYT
Sbjct: 1062 EHLHPGGLLLPLPVPTTVWADIGLDFVEALPRVGGKTVILTVVDRFSKYCHFIPLAHPYT 1121
Query: 1190 AKSIAEVFVREIVRLHGIPNTVKSDRDPIFVSHFWSELFKLQGTKLKMSSSYHPETDGQT 1249
A+S+A+ F +IVRLHGIP ++ SDRDP+F S FW EL +L GTK+ M+++ HP++DGQT
Sbjct: 1122 AESVAQAFYADIVRLHGIPQSMVSDRDPVFTSSFWRELMRLTGTKMHMTTAIHPQSDGQT 1181
Query: 1250 EVINRCLESYLRCFASDHPKTWSFWVPWAEFWYNSTFHCSIGQTPFEVVYGRKAPPIIRF 1309
E N+ + YLRCF D P+ W W+PWAE+ YN+ + S+ TPF VVYGR P I +
Sbjct: 1182 EAANKVIVMYLRCFTGDRPRQWVRWLPWAEYIYNTAYQTSLRDTPFRVVYGRDPPIIRSY 1241
Query: 1310 LTNETKVVAVALELSERDEALKQLRTHLQRAQEQMASYANKKRRDLSFDIGEWVFLKLRP 1369
ET+V AVA +++RDE L +R L++AQ Y +K R +S+++G+ V L+LR
Sbjct: 1242 EPGETRVAAVARSMADRDEFLADVRYRLEQAQATHKKYYDKGHRAVSYEVGDLVLLRLRH 1301
Query: 1370 HRQHSVVKRINQKLAARFYGPFQIEDKIGTVAYKLKLPVESRIHPVFHVSLLKRAIGNYQ 1429
S+ + KL R++GP+++ + I VA +L+LP +++H VFHV LLK+ +G
Sbjct: 1302 RAPASLPQVSKGKLKPRYFGPYRVVEVINPVAVRLELPPRAKLHDVFHVGLLKKFVG--A 1359
Query: 1430 VQGQLPTDLGIEE-ATDIYPEAILGTRIIRQGDSEVHQSLIKWKNRSIDDVTWEDNEVLA 1488
P + A D PE + +R+ R V Q L+ WK S TWED +
Sbjct: 1360 APPSPPALPAVHHGAIDPEPERVTRSRLAR----GVRQVLVHWKGESAASATWEDLDTFK 1415
Query: 1489 GQFPEFCLEDKAVSKEG 1505
++P F LED+ +EG
Sbjct: 1416 ERYPAFQLEDELALEEG 1432
>UniRef100_Q9LP90 T32E20.30 [Arabidopsis thaliana]
Length = 1397
Score = 971 bits (2509), Expect = 0.0
Identities = 530/1183 (44%), Positives = 727/1183 (60%), Gaps = 116/1183 (9%)
Query: 336 HKCPERAMRVLILGDGETMDEDGEIVMLEEEAEEIEEEETVECKLMGVLGCMGEHQTQTM 395
HKCP + +RVL + +G M+ V+ EE + + + MG T+
Sbjct: 315 HKCPNKELRVLTVINGFEME-----VLESNSVEEEFHDSVAQFAELSFSSYMGLPSYTTI 369
Query: 396 KLAGQVANVELLILIDSGASHNFISPKITSALGLVITPITTRSIKLGDGHKVVTRGICRG 455
K+ G + E F P I+LG G V G+C
Sbjct: 370 KMKGSICKGE-------WCHTQFYFPNF--------------HIRLGTGITVQGLGLCDK 408
Query: 456 IKAKL-----GSIEITIDALVLELGGLDLVLGVSWLSTLGKVIMDWKLLTMQFVYGEQVV 510
+ L +E+T + L+LG +D++LG++WL TLG ++W+ + F+Y + V
Sbjct: 409 VTMTLPVGCGQELELTTHFITLDLGPVDVILGIAWLRTLGDCKVNWERHELSFLYHGRTV 468
Query: 511 QLQGMRGKDSYQSNLHSFLSDTHGREQMEWWGPQLQVVEATKKGHEIPQELRAVLADFHE 570
L+G D+++ +L SF + + Q + +E + H Q L+
Sbjct: 469 TLRGDPELDTFKMSLKSFSTK---------FRLQNKELEVSLNSH---QNLKG------- 509
Query: 571 VFTKKIQLPPVRSRVHQITLNPEHGPINVRPYRYPHHQKEEIERQVTELLEAGIIRPSMS 630
LPP++ H I+L P I+VRPYRYPH KE +E V+E+L+ GIIR S S
Sbjct: 510 -------LPPIKGNEHAISLLPGTRAISVRPYRYPHAHKEAMEGLVSEMLDNGIIRASKS 562
Query: 631 SFSSPVILVKKKDKSWRMCVDYRALNKATIPDKYPIPIVDELLDELNGAVIFSKIDLKSG 690
FSSPV+LVKKKD+SWR CVDYRALN+ATIP+K+PIP++D+LLDEL+GA+IFSK+DL++G
Sbjct: 563 PFSSPVLLVKKKDQSWRFCVDYRALNRATIPNKFPIPMIDQLLDELHGAIIFSKLDLRAG 622
Query: 691 YHQIRVHEEDIPKTTFRTHNGHYEYLVMPFGLMNAPATFQAIMNDIFRPYLRKFVLVFFD 750
YHQIR+ EDI KTTFRTH+GH+E+LVMPFGL NAPATFQ+ MND+ RP+LRKFVLVFFD
Sbjct: 623 YHQIRMKVEDIEKTTFRTHDGHFEFLVMPFGLSNAPATFQSSMNDMLRPFLRKFVLVFFD 682
Query: 751 DILIYSKSLLEHRDHLKLVLTVLLSNSFVANQSKCKFGCAQIDYLGHIISGAGVSVDPEK 810
DILIYS++ EH +HL +VL VL + F AN+ K HI GVS DP K
Sbjct: 683 DILIYSRNEQEHEEHLAMVLKVLEEHQFYANRKKPY----------HITQ--GVSTDPTK 730
Query: 811 VQCIVEWPEPKNVKGVRGFLGLTGYYRKFIKNYGKMAKPLTELTKKDNFTWGEEAKQAFH 870
+ +W P++VK +RGFLGLTGYYR+F+K YG +A+PLTEL KKD+F W E A++AF
Sbjct: 731 TVAMTKWVTPQSVKELRGFLGLTGYYRRFLKGYGTLARPLTELLKKDSFVWSESAQEAFD 790
Query: 871 KLKTVMTSSPVLALPDFDKSFEVECDAAGRGIGAVLMQQRQPIAYFSKALSNGNLTKSVY 930
LK M+++PVLALPDF K L++ K VY
Sbjct: 791 ALKRAMSTAPVLALPDFGK---------------------------VHGLTSKEQLKPVY 823
Query: 931 EKELMALVLCIQHWRHYLLGKAFTVYTDHKSLKHFLQQKISSPDQQCWLAKLLGYQFEVK 990
E+ELMA+VL IQ W+HYL+G+ F ++TD KSLK +Q+ S D Q WL KLL Y+F++
Sbjct: 824 ERELMAIVLSIQKWKHYLMGRRFVLHTDQKSLKFLQEQREVSMDYQKWLTKLLHYEFDIL 883
Query: 991 YKPGQENKAADALSRRGDESDFFS---LVSF--PLWDDRKKLLEEIASDPYIKNLLQDVQ 1045
YK G +NKAAD LSR + FS L++F P L EEI S+ ++++L+++
Sbjct: 884 YKLGVDNKAADGLSRMVQPTGSFSSMLLMAFTVPTVLQLHDLYEEIDSNAHLQHLVKECL 943
Query: 1046 QDPQSKPGFSVLQGVLLYQGRLVLSPTSPSIPWLLQEYHGSPTGGHSGFLRTYRRLAESL 1105
Q ++V +G L + RL++ S +P +L EYH GGHSG L+T +R+ +S
Sbjct: 944 SAKQGTSAYTVKEGRLWKKQRLIIPKDSKFLPLILAEYHSGLLGGHSGVLKTMKRIQQSF 1003
Query: 1106 YWVGMQRTVRDFVRACDTCQRQKYSAMSPGGLLQPFPVPNAVWEDLSLDFIIGLPKSKGY 1165
+W GM + ++ FV C+ CQRQKYS +SP GLLQP P+P VWED+SLDF+ GLP
Sbjct: 1004 HWEGMMKDIQKFVAKCEMCQRQKYSTLSPAGLLQPLPIPTQVWEDISLDFVEGLP----- 1058
Query: 1166 EAILVVVDRLSKYSHFILLKHPYTAKSIAEVFVREIVRLHGIPNTVKSDRDPIFVSHFWS 1225
DRLSKY HFI LKHP+ A +A +F+ E+V+LHG P ++ SDRD F+S FW
Sbjct: 1059 -------DRLSKYGHFIGLKHPFNAVDVARIFIHEVVKLHGFPASIVSDRDNTFLSSFWK 1111
Query: 1226 ELFKLQGTKLKMSSSYHPETDGQTEVINRCLESYLRCFASDHPKTWSFWVPWAEFWYNST 1285
+ FKL GTKLK S+++HP+TDGQTEV+NR LE+YLRCFAS HPKTW ++P AE WYNS+
Sbjct: 1112 DCFKLSGTKLKYSTAFHPQTDGQTEVLNRTLETYLRCFASAHPKTWFQYLPRAELWYNSS 1171
Query: 1286 FHCSIGQTPFEVVYGRKAPPIIRFLTNETKVVAVALELSERDEALKQLRTHLQRAQEQMA 1345
FH +I TPF+V+YGR PPI+RF N TK + L +RD L ++ HL AQ+ M
Sbjct: 1172 FHTTIKTTPFKVLYGRDPPPIMRFEANSTKNCELEGLLKQRDLMLADIKEHLVNAQQLMK 1231
Query: 1346 SYANKKRRDLSFDIGEWVFLKLRPHRQHSVVKRINQKLAARFYGPFQIEDKIGTVAYKLK 1405
+ +K RR++ FD VFLKLRP+RQ+SV KR+ QKLAA+++GPF+I ++IG VAY+LK
Sbjct: 1232 NNDDKHRREVEFDTRNRVFLKLRPYRQNSVTKRVCQKLAAKYFGPFEIMERIGKVAYRLK 1291
Query: 1406 LPVESRIHPVFHVSLLKRAIGNYQVQGQLPTDLGIEEATDIYPEAILGTRIIRQGDSEVH 1465
LP S+IH VFHVS LK+ +G++ LP L + + PEA+L TR G +
Sbjct: 1292 LPEGSKIHLVFHVSQLKQVLGDHHQVIPLPEVLTADNEFVVVPEAVLETRYNEDG---LL 1348
Query: 1466 QSLIKWKNRSIDDVTWEDNEVLAGQFPEFCLEDKAVSKEGGIE 1508
++L+ W+ + + TWE + L QFP L+DK + GG E
Sbjct: 1349 EALVHWQGLPVHEDTWEIAKDLKKQFPGLALQDKLHVEGGGGE 1391
>UniRef100_Q9FW76 Putative gag-pol polyprotein [Oryza sativa]
Length = 1608
Score = 948 bits (2451), Expect = 0.0
Identities = 506/1188 (42%), Positives = 717/1188 (59%), Gaps = 19/1188 (1%)
Query: 328 WGKISPTLHKCPERAMRVLILGDGETMDEDGEIVMLEEEAEEIEEEETVECKLMGVLGCM 387
WG+ HKC ++ + D E E A E EE+ + +
Sbjct: 379 WGRD----HKCATTVQLHVVEELINALKTDPEENCNSEGAPESEEDSLMAISFQAL---N 431
Query: 388 GEHQTQTMKLAGQVANVELLILIDSGASHNFISPKITSALGLVITPITTRSIKLGDGHKV 447
G +++++L G V N ELL+L+DSG++H+FI K+ + L + +++ DG ++
Sbjct: 432 GTDSSKSIRLRGWVQNTELLMLVDSGSTHSFIDAKLGAQLCGLQKLNQAIKVQVADGSQL 491
Query: 448 VTRGICRGIKAKLGSIEITIDALVLELGGLDLVLGVSWLSTLGKVIMDWKLLTMQFVYGE 507
T D +L LG D +LG+ WL + +DW + F +
Sbjct: 492 FCDSFLPNCSWWSQGHSFTSDFRLLPLGSYDAILGMDWLEQFSPMQVDWVHKWIAFQHHG 551
Query: 508 QVVQLQGMRGKDSYQSNLHSFLSDTHGREQMEWWGPQLQVVEATKKGHEIPQELRAVLAD 567
Q VQLQG+ + S + + ++ L V E T +P+ ++ +L +
Sbjct: 552 QAVQLQGIHPQLSTCFPISNDQLQGMSKKGAVMCLVHLNVAE-TLTATTVPEIVQPILNE 610
Query: 568 FHEVFTKKIQLPPVRSRVHQITLNPEHGPINVRPYRYPHHQKEEIERQVTELLEAGIIRP 627
F E+F++ +LPP R+ H I L P+N+RPYRY K+EIERQV E+L +G+I+P
Sbjct: 611 FQEIFSEPTELPPKRNCDHHIPLVEGAKPVNLRPYRYKPALKDEIERQVAEMLRSGVIQP 670
Query: 628 SMSSFSSPVILVKKKDKSWRMCVDYRALNKATIPDKYPIPIVDELLDELNGAVIFSKIDL 687
S S FSSP +LVKKKD +WR+C+DYR LN T+ KYP+P++DELLDEL G+ FSK+DL
Sbjct: 671 SSSPFSSPALLVKKKDGTWRLCIDYRQLNDVTVKSKYPVPVIDELLDELAGSKWFSKLDL 730
Query: 688 KSGYHQIRVHEEDIPKTTFRTHNGHYEYLVMPFGLMNAPATFQAIMNDIFRPYLRKFVLV 747
++GYHQIR+ E D KT F+TH+GHYEY VM FGL APATF + MN+ P LRKF LV
Sbjct: 731 RAGYHQIRMAEGDEYKTAFQTHSGHYEYKVMSFGLTGAPATFLSAMNETLSPVLRKFALV 790
Query: 748 FFDDILIYSKSLLEHRDHLKLVLTVLLSNSFVANQSKCKFGCAQIDYLGHIISGAGVSVD 807
FFDDILIYS +L H H++ VL +L ++ + SKC F +I YLGH+I AGV+ D
Sbjct: 791 FFDDILIYSPTLELHLQHVRTVLQLLSAHQWKVKLSKCSFAQQEISYLGHVIGAAGVATD 850
Query: 808 PEKVQCIVEWPEPKNVKGVRGFLGLTGYYRKFIKNYGKMAKPLTELTKKD-NFTWGEEAK 866
P K+Q +V WP+P +K +RGFLGL GYYRKF++++G ++KPLT+L KK F W E +
Sbjct: 851 PAKIQDVVSWPQPTTIKKLRGFLGLAGYYRKFVRHFGLISKPLTQLLKKGIPFKWTPEIE 910
Query: 867 QAFHKLKTVMTSSPVLALPDFDKSFEVECDAAGRGIGAVLMQQRQPIAYFSKALSNGNLT 926
AF +LK + ++PVLALPDF K F +E DA+ GIGAVL Q++ PIAY S+AL
Sbjct: 911 SAFQQLKQALVAAPVLALPDFSKHFTIETDASDVGIGAVLSQEKHPIAYLSRALGPKTRG 970
Query: 927 KSVYEKELMALVLCIQHWRHYLLGKAFTVYTDHKSLKHFLQQKISSPDQQCWLAKLLGYQ 986
S YEKE MA++L ++HWR YL F + TDH SL H +Q++ +P QQ KLLG Q
Sbjct: 971 LSTYEKEYMAIILAVEHWRPYLQQGEFIILTDHHSLMHLTEQRLHTPWQQKAFTKLLGLQ 1030
Query: 987 FEVKYKPGQENKAADALSRRGDESDFFSLVS--FPLWDDRKKLLEEIASDPYIKNLLQDV 1044
+++ Y+ G N AADALSRR + +S P W ++L++ D K LL ++
Sbjct: 1031 YKICYRKGVSNAAADALSRRESPISEVAAISECIPSW--MQELMQGYQLDGQSKQLLAEL 1088
Query: 1045 QQDPQSKPGFSVLQGVLLYQGRLVLSPTSPSIPWLLQEYHGSPTGGHSGFLRTYRRLAES 1104
P S+ + + QG+L Y+G++ + + L+ E H +P GGHSGF TYR++
Sbjct: 1089 AISPNSRKDYQLCQGILKYKGKIWVGNNTALQHKLVNELHATPLGGHSGFPVTYRKVKSL 1148
Query: 1105 LYWVGMQRTVRDFVRACDTCQRQKYSAMSPGGLLQPFPVPNAVWEDLSLDFIIGLPKSKG 1164
W GM++ +++ +++C C + K GLLQP PVP W+ +SLDFI GLP+S
Sbjct: 1149 FAWPGMKKLIKEQLQSCQVCLQAKPDRARYPGLLQPLPVPAGAWQTISLDFIEGLPRSSH 1208
Query: 1165 YEAILVVVDRLSKYSHFILLKHPYTAKSIAEVFVREIVRLHGIPNTVKSDRDPIFVSHFW 1224
Y ILVVVD+ SKYSHFI L HP+ A +A+ F++ I +LHG+P + SDRD IF S FW
Sbjct: 1209 YNCILVVVDKFSKYSHFIPLSHPFNAGGVAQEFMKNIYKLHGLPRAIISDRDKIFTSQFW 1268
Query: 1225 SELFKLQGTKLKMSSSYHPETDGQTEVINRCLESYLRCFASDHPKTWSFWVPWAEFWYNS 1284
+LF GT L MSS+YHP++DGQTE +N+CLE YLRCF P WS W+ AEFWYN+
Sbjct: 1269 DQLFSKFGTDLHMSSAYHPQSDGQTERVNQCLEIYLRCFVHAAPHKWSSWLYLAEFWYNT 1328
Query: 1285 TFHCSIGQTPFEVVYGRKAPPIIRFLTNETKVVAVALELSERDEALKQLRTHLQRAQEQM 1344
+FH ++ +TPFEV+YG P ++ ++ + +ER + L+ HL RAQ+QM
Sbjct: 1329 SFHSTLNKTPFEVLYG-YTPSHFGIGLDDCQIADLHEWHTERKFMQQLLQQHLNRAQQQM 1387
Query: 1345 ASYANKKRRDLSFDIGEWVFLKLRPHRQHSVVKRINQKLAARFYGPFQIEDKIGTVAYKL 1404
A+KKR F +G+WV+LKL+P+ Q V +R KLA R+YGPFQ+ ++GTVAY +
Sbjct: 1388 KHQADKKRSFRQFAVGDWVYLKLQPYVQTFVAQRACHKLAFRYYGPFQVMSRVGTVAYHI 1447
Query: 1405 KLPVESRIHPVFHVSLLKRAIG-NYQVQGQLPTDLGIEEATDIYPEAILGTRIIRQGDSE 1463
+LP S IHPVFHVS LK A+G + +VQ +LPT LG + P L R++++G+
Sbjct: 1448 QLPATSSIHPVFHVSQLKAAVGFSKKVQDELPTSLGALQV----PFQFLDKRLVKKGNRS 1503
Query: 1464 VHQSLIKWKNRSIDDVTWEDNEVLAGQFPEFCLEDKAVSKEGGIERDS 1511
V Q L W + S + TWED E L +FP +A ++ GI R S
Sbjct: 1504 VLQLLTHWYHSSPSESTWEDMEDLFARFPRALAWGQASFQDWGIVRPS 1551
>UniRef100_Q7XW39 OSJNBb0062H02.17 protein [Oryza sativa]
Length = 1629
Score = 934 bits (2413), Expect = 0.0
Identities = 508/1143 (44%), Positives = 706/1143 (61%), Gaps = 14/1143 (1%)
Query: 373 EETVECKLMGVL--GCMGEHQTQTMKLAGQVANVELLILIDSGASHNFISPKITSAL-GL 429
+ VE LM V G M++ GQ+ E+LIL+DSG+S +FIS ++ S+L G+
Sbjct: 454 DHVVEQTLMAVSLQAVQGTETGGCMRMLGQIQGKEILILVDSGSSASFISKRVASSLMGV 513
Query: 430 VITPITTRSIKLGDGHKVVTRGICRGIKAKLGSIEITIDALVLELGGLDLVLGVSWLSTL 489
+ P+ + + G I G + T + VLEL D++LG+ WL
Sbjct: 514 LEQPVHVQVMVAGGAKLHCCSEILNCEWTIQGHVFFT-NLKVLELNNYDMILGMDWLMQH 572
Query: 490 GKVIMDWKLLTMQFVYGEQVVQLQGMRGKDSYQSNLHSF-LSDTHGREQMEWWGPQLQVV 548
+ +DW ++ Y +QL G+R +++ S L + + R + Q V
Sbjct: 573 SPMTVDWTTKSLIIAYAGTQIQLYGVRSDTEQCAHISSKQLRELNDRTAVSNL-VQFCSV 631
Query: 549 EATKKGHEIPQELRAVLADFHEVFTKKIQLPPVRSRVHQITLNPEHGPINVRPYRYPHHQ 608
A + +IP+ ++ VL +F VF + LPP+R H I L P GP+NVRPYRY Q
Sbjct: 632 FALEYQEQIPEVVQTVLTEFSSVFDEPKGLPPIRQFDHTIPLLPGAGPVNVRPYRYTPIQ 691
Query: 609 KEEIERQVTELLEAGIIRPSMSSFSSPVILVKKKDKSWRMCVDYRALNKATIPDKYPIPI 668
K EIE QV E+L GII+PS S FSSPV+LVKKKD SWR CVDYR LN T+ +KYP+P+
Sbjct: 692 KNEIESQVQEMLSKGIIQPSSSPFSSPVLLVKKKDGSWRFCVDYRHLNAITVKNKYPLPV 751
Query: 669 VDELLDELNGAVIFSKIDLKSGYHQIRVHEEDIPKTTFRTHNGHYEYLVMPFGLMNAPAT 728
+DELLDEL GA FSK+DL+SGYHQIR+H +D KT F+TH+GH+E+ V+PFGL +APAT
Sbjct: 752 IDELLDELAGAQWFSKLDLRSGYHQIRMHPDDEHKTAFQTHHGHFEFRVLPFGLTSAPAT 811
Query: 729 FQAIMNDIFRPYLRKFVLVFFDDILIYSKSLLEHRDHLKLVLTVLLSNSFVANQSKCKFG 788
FQ +MN + LR+ VLVF DDILIYSKSL EH HLK V +LL + ++KC F
Sbjct: 812 FQGVMNSVLATLLRRCVLVFVDDILIYSKSLEEHVQHLKTVFQILLKHQLKVKRTKCSFA 871
Query: 789 CAQIDYLGHIISGAGVSVDPEKVQCIVEWPEPKNVKGVRGFLGLTGYYRKFIKNYGKMAK 848
++ YLGHII GVS DPEK+Q I WP P +VK +R FLGL+GYYRKF++NYG ++K
Sbjct: 872 QQELAYLGHIIQPNGVSTDPEKIQVIQHWPAPTSVKELRSFLGLSGYYRKFVRNYGILSK 931
Query: 849 PLTELTKKDN-FTWGEEAKQAFHKLKTVMTSSPVLALPDFDKSFEVECDAAGRGIGAVLM 907
PLT L +K + W E + AF LK + ++ VLA+PDF F VE DA+ +GIGAVLM
Sbjct: 932 PLTNLLRKGQLYIWTAETEDAFQALKQALITALVLAMPDFQTPFVVETDASDKGIGAVLM 991
Query: 908 QQRQPIAYFSKALSNGNLTKSVYEKELMALVLCIQHWRHYLLGKAFTVYTDHKSLKHFLQ 967
Q P+A+ S+AL + S YEKE +A++L + HWR YL F + TDH+SL +
Sbjct: 992 QNNHPLAFLSRALGLRHPGLSTYEKESLAIMLAVDHWRPYLQHDEFFIRTDHRSLAFLTE 1051
Query: 968 QKISSPDQQCWLAKLLGYQFEVKYKPGQENKAADALSR--RGDESDFFSL-VSFPLWDDR 1024
Q++++P Q L KLLG ++++ +K G +N AADALSR D + +L V+ P W +
Sbjct: 1052 QRLTTPWQHKALTKLLGLRYKIIFKKGIDNSAADALSRYPGSDRVELSALSVAVPEWIN- 1110
Query: 1025 KKLLEEIASDPYIKNLLQDVQQDPQSKPGFSVLQGVLLYQGRLVLSPTSPSIPWLLQEYH 1084
++ +SDP + +Q + + + P FS+ GVL +Q RL + +L H
Sbjct: 1111 -DIVAGYSSDPDACSKVQTLCINSGAVPNFSLRNGVLYFQNRLWVGHNVDVQQRILANLH 1169
Query: 1085 GSPTGGHSGFLRTYRRLAESLYWVGMQRTVRDFVRACDTCQRQKYSAMSPGGLLQPFPVP 1144
+ GGHSG TY+R+ + W ++ TV +V+AC CQ+ K + G+LQP PVP
Sbjct: 1170 TAAVGGHSGIQVTYQRVKQLFAWPRLRATVVQYVQACSVCQQAKSEHVKYPGMLQPLPVP 1229
Query: 1145 NAVWEDLSLDFIIGLPKSKGYEAILVVVDRLSKYSHFILLKHPYTAKSIAEVFVREIVRL 1204
+ W+ +SLDF+ GLPKS + ILVVVD+ SKYSHF+ L HP++A +AE +++ I RL
Sbjct: 1230 DHAWQIVSLDFVEGLPKSASFNCILVVVDKFSKYSHFVPLTHPFSALDVAEAYMQHIHRL 1289
Query: 1205 HGIPNTVKSDRDPIFVSHFWSELFKLQGTKLKMSSSYHPETDGQTEVINRCLESYLRCFA 1264
HG+P ++ SDRD IF S W+ LF+L GT+L+MSSSYHP+TDGQTE +N+CLE++LRCF
Sbjct: 1290 HGLPQSLISDRDRIFTSTLWTTLFRLAGTQLRMSSSYHPQTDGQTERVNQCLETFLRCFV 1349
Query: 1265 SDHPKTWSFWVPWAEFWYNSTFHCSIGQTPFEVVYGRKAPPIIRFLTNETKVVAVALELS 1324
P WS W+ AE+WYN++FH ++G TPFEV+YG K + + + L
Sbjct: 1350 HACPSQWSRWLALAEYWYNTSFHSALGTTPFEVLYGHKPRYFGLSASAACRSDDLVEWLH 1409
Query: 1325 ERDEALKQLRTHLQRAQEQMASYANKKRRDLSFDIGEWVFLKLRPHRQHSVVKRINQKLA 1384
ER++ +R HL RAQ +M A++ R + SF +G+WV+LKL+P Q SVV R N+KL+
Sbjct: 1410 EREKMQALIRDHLLRAQTRMKQQADQHRSERSFAVGDWVYLKLQPFVQQSVVTRANRKLS 1469
Query: 1385 ARFYGPFQIEDKIGTVAYKLKLPVESRIHPVFHVSLLKRAIG-NYQVQGQLPTDLGIEEA 1443
RFYGPFQ+ DK+GTVAY+L LP S IHPV HVS LK+A+ QV LP L A
Sbjct: 1470 FRFYGPFQVLDKVGTVAYRLDLPSSSLIHPVVHVSQLKKALAPTEQVHSPLPV-LDPTNA 1528
Query: 1444 TDIYPEAILGTRIIRQGDSEVHQSLIKWKNRSIDDVTWEDNEVLAGQFPEFCLEDKAVSK 1503
T + P IL R IR+G V Q ++W + TWE+ + L +FP +A ++
Sbjct: 1529 THVCPAQILDRRFIRKGSKLVEQIQVRWTGDAPAATTWENPQELRRRFPTAPAWGQAGTQ 1588
Query: 1504 EGG 1506
GG
Sbjct: 1589 GGG 1591
>UniRef100_Q8SA93 Putative polyprotein [Zea mays]
Length = 2749
Score = 932 bits (2408), Expect = 0.0
Identities = 502/1193 (42%), Positives = 706/1193 (59%), Gaps = 68/1193 (5%)
Query: 336 HKCPERAMRVLILGDGETMDEDGEIVMLEEEAE-EIEEEETVECKLMGVLGCMGEHQT-- 392
H CP R+ L + + +D++ + EE A+ +I E+ + ++ + H
Sbjct: 699 HVCP----RLFYLENDDYIDDEPQ----EEGADLQIALEQEPPSRAAAIIPTVSLHALAG 750
Query: 393 ----QTMKLAGQVANVELLILIDSGASHNFISPKITSALGLVITPITTRSIKLGDGHKVV 448
M L + L+ L+DSG++ NF+S + S L L TP T +++ +G +
Sbjct: 751 VRTPNAMLLPVSINGHRLVALVDSGSTTNFMSVGLMSRLQLPSTPHPTIKVQVANGDNIP 810
Query: 449 TRGICRGIKAKLGSIEITIDALVLELGGLDLVLGVSWLSTLGKVIMDWKLLTMQFVYGEQ 508
+G+ R + ++G+ + +ID + L LG D++LG +L LG ++ D L+M F G +
Sbjct: 811 CQGMARSVDLRVGTEQFSIDCIGLTLGTFDVILGFEFLRLLGPILWDCDRLSMSFTKGGR 870
Query: 509 VVQLQGMRGKDSYQSNLHSFLSDTHGREQMEWWGPQLQVVEATKKGHEIPQELRAVLADF 568
+ G+ + P VV +T + LR F
Sbjct: 871 HIIWSGLGAPGAVPPQ------------------PAACVVSSTPTQPLLDDLLR----QF 908
Query: 569 HEVFTKKIQLPPVRSRVHQITLNPEHGPINVRPYRYPHHQKEEIERQVTELLEAGIIRPS 628
VF + LPP R H+I L P P+ VRPYRYP QK+E+ERQ + +L GIIRPS
Sbjct: 909 ELVFAEPQGLPPARPYDHRIHLLPGAAPVAVRPYRYPQLQKDELERQCSAMLAQGIIRPS 968
Query: 629 MSSFSSPVILVKKKDKSWRMCVDYRALNKATIPDKYPIPIVDELLDELNGAVIFSKIDLK 688
S FS+PV+LV+K D SWR C+DYRALN T DK+PIP+VDELLDEL+GA F+K+DL+
Sbjct: 969 TSPFSAPVLLVRKPDNSWRFCIDYRALNAKTSKDKFPIPVVDELLDELHGAHFFTKLDLR 1028
Query: 689 SGYHQIRVHEEDIPKTTFRTHNGHYEYLVMPFGLMNAPATFQAIMNDIFRPYLRKFVLVF 748
SGYHQ+R+H D+ KT FRTH GHYE+LVMPFGL NAPATFQA+MND+ RPYLRK+VLVF
Sbjct: 1029 SGYHQVRMHPADVEKTAFRTHEGHYEFLVMPFGLSNAPATFQALMNDVLRPYLRKYVLVF 1088
Query: 749 FDDILIYSKSLLEHRDHLKLVLTVLLSNSFVANQSKCKFGCAQIDYLGHIISGAGVSVDP 808
FDDILIYSK+ EH H+ +VL L + +SKC FG + YLGH+IS AGV++D
Sbjct: 1089 FDDILIYSKTWAEHLQHISIVLHALRDHQLHLKRSKCSFGARSVAYLGHVISAAGVAMDA 1148
Query: 809 EKVQCIVEWPEPKNVKGVRGFLGLTGYYRKFIKNYGKMAKPLTELTKKDNFTWGEEAKQA 868
KV+ + WP P + +G+RGFLGL GYYRKFI+++G +A PLT L ++D FTW ++ + A
Sbjct: 1149 AKVEAVSSWPAPHSARGLRGFLGLAGYYRKFIRDFGVIAAPLTRLLRRDAFTWDDDTQAA 1208
Query: 869 FHKLKTVMTSSPVLALPDFDKSFEVECDAAGRGIGAVLMQQRQPIAYFSKALSNGNLTKS 928
F +LKT +T+ PVL +P+F+K+F V+CDA+G G GAVL Q P+A+FS+ +L +
Sbjct: 1209 FQQLKTALTTGPVLQMPNFEKTFVVDCDASGTGFGAVLHQGAGPVAFFSRPFVTRHLKLA 1268
Query: 929 VYEKELMALVLCIQHWRHYLLGKAFTVYTDHKSLKHFLQQKISSPDQQCWLAKLLGYQFE 988
YE+EL+ LV ++HWR YL G+ F V TDH SLK+ L Q++S+ Q WL+KL G+ FE
Sbjct: 1269 AYERELIGLVQAVRHWRPYLWGRHFAVRTDHYSLKYLLDQRLSTVPQHQWLSKLFGFDFE 1328
Query: 989 VKYKPGQENKAADALSRRGDE-------------SDFFSLVSFPLWDDRKKLLEEIASDP 1035
V+Y+PG+ N AADALSRR E + S SF DD ++ A+ P
Sbjct: 1329 VEYRPGRLNVAADALSRRDAELLQPSAGELGAAAALALSGPSFAFLDDIRR---ATATSP 1385
Query: 1036 YIKNLLQDVQQDPQSKPGFSVLQGVLLYQGRLVLSPTSPSIPWLLQEYHGSPTGGHSGFL 1095
L Q +Q + P + + G+LL+ R+ + + H + GH G
Sbjct: 1386 DSSRLCQQLQDGTLTAP-WRLEDGLLLHGSRIYVPNHGDLRHQAILLAH---SAGHEGIQ 1441
Query: 1096 RTYRRLAESLYWVGMQRTVRDFVRACDTCQRQKYSAMSPGGLLQPFPVPNAVWEDLSLDF 1155
+T RL Y G + V D+VR C TCQR K + P GLLQP VP+ VW D+S+DF
Sbjct: 1442 KTLHRLRAEFYVPGDRTLVADWVRTCTTCQRNKTETLQPAGLLQPLQVPSQVWADISMDF 1501
Query: 1156 IIGLPKSKGYEAILVVVDRLSKYSHFILLKHPYTAKSIAEVFVREIVRLHGIPNTVKSDR 1215
I GLPK G IL VVDR SKY+HFI L HPYTA S+A F IVRLHG P+++ SDR
Sbjct: 1502 IEGLPKVGGKSVILTVVDRFSKYAHFIPLGHPYTAASVARAFFDGIVRLHGFPSSIVSDR 1561
Query: 1216 DPIFVSHFWSELFKLQGTKLKMSSSYHPETDGQTEVINRCLESYLRCFASDHPKTWSFWV 1275
DP+F H W +LFK G L+MS+++HP+TDGQ+EV+N+ + YLRC D P+ W W+
Sbjct: 1562 DPVFTGHVWRDLFKCAGVSLRMSTAFHPQTDGQSEVVNKVIAMYLRCVTGDRPRAWVDWL 1621
Query: 1276 PWAEFWYNSTFHCSIGQTPFEVVYGRKAPPIIRFLTNETKVVAVALELSERDEALKQLRT 1335
WAE+ YN++FH ++ TPFEVVYGR PPI+ + + A L +RD L ++R
Sbjct: 1622 SWAEYCYNTSFHTALRATPFEVVYGRPPPPILPYQAGSARTAAAEELLRDRDNILAEVRQ 1681
Query: 1336 HLQRAQEQMASYANKKRRDLSFDIGEWVFLKLRPHRQHSVVKRINQKLAARFYGPFQIED 1395
L +AQ+ Y + RD+ G+WV+L+L S+ R KL R+ GPF++ +
Sbjct: 1682 RLVQAQQLSKRYYDAGHRDMELADGDWVWLRLLHRPVQSLEPRAKGKLGPRYAGPFRVLE 1741
Query: 1396 KIGTVAYKLKLPVESRIHPVFHVSLLKRAIGN--YQVQGQLPTDLGIEEATDIYPEAILG 1453
+IG VAY+L+LP +R+H VFHV LLKR G Q G P G + P +
Sbjct: 1742 RIGKVAYRLELPEGARLHDVFHVGLLKRHKGEPPEQRAGLPPVQNG-----RLLPAPL-- 1794
Query: 1454 TRIIR-QGDSEVHQSLIKWKNRSIDDVTWEDNEVLAGQFPEFCLEDKAVSKEG 1505
+++R Q L++W+ S ++ TWE + G +P+F LED+ + G
Sbjct: 1795 -KVLRAQQRRGTWHILVQWQGLSPEEATWEPLDDFRGLYPDFQLEDELFAHAG 1846
>UniRef100_Q9C7H8 Gypsy/Ty3 element polyprotein, putative [Arabidopsis thaliana]
Length = 1447
Score = 931 bits (2405), Expect = 0.0
Identities = 486/1142 (42%), Positives = 701/1142 (60%), Gaps = 50/1142 (4%)
Query: 354 MDEDGEIVMLEEEAEEIEEEETVECKLM---GVLGCMGEHQTQTMKLAGQVANVELLILI 410
MD D E E+A E+ ++ E K M V G +TM + G V +L ILI
Sbjct: 329 MDVDEEF----EDAVEVLSDDDHEQKPMPQISVNAVSGISGYKTMGVKGTVDKRDLFILI 384
Query: 411 DSGASHNFISPKITSALGLVITPITTRSIKLGDGHKVVTRGICRGIKAKLGSIEITIDAL 470
DSG++HNFI + + LG + + + DG K+ G +G KL S D L
Sbjct: 385 DSGSTHNFIDSTVAAKLGCHVESAGLTKVAVADGRKLNVDGQIKGFTWKLQSTTFQSDIL 444
Query: 471 VLELGGLDLVLGVSWLSTLGKVIMDWKLLTMQFVYGEQVVQLQGMRGKDSYQSNLHSF-- 528
++ L G+D+VLGV WL TLG++ ++K L MQF Y Q V L G+ H
Sbjct: 445 LIPLQGVDMVLGVQWLETLGRISWEFKKLEMQFFYKNQRVWLHGIITGSVRDIKAHKLQK 504
Query: 529 -------LSDTHGREQMEWWGPQLQVVEATKKGHEIPQELRAVLADFHEVFTKKIQLPPV 581
L+ RE + ++ + A ++ ++ +F +VF + LPP
Sbjct: 505 TQADQIQLAMVCVREVVSDEEQEIGSISALTSDVVEESVVQNIVEEFPDVFAEPTDLPPF 564
Query: 582 RSRV-HQITLNPEHGPINVRPYRYPHHQKEEIERQVTELLEAGIIRPSMSSFSSPVILVK 640
R + H+I L P+N RPYRY HQK+EI++ V +++++G I+ S S F+SPV+LVK
Sbjct: 565 REKHDHKIKLLEGANPVNQRPYRYVVHQKDEIDKIVQDMIKSGTIQVSSSPFASPVVLVK 624
Query: 641 KKDKSWRMCVDYRALNKATIPDKYPIPIVDELLDELNGAVIFSKIDLKSGYHQIRVHEED 700
KKD +WR+CVDY LN T+ D++ IP++++L+DEL G+V+FSKIDL++GYHQ+R+ +D
Sbjct: 625 KKDGTWRLCVDYTELNGMTVKDRFLIPLIEDLMDELGGSVVFSKIDLRAGYHQVRMDPDD 684
Query: 701 IPKTTFRTHNGHYEYLVMPFGLMNAPATFQAIMNDIFRPYLRKFVLVFFDDILIYSKSLL 760
I KT F+THNGH+EYLVM FGL NAPATFQ++MN +FR +LRKFVLVFFDDILIYS S+
Sbjct: 685 IQKTAFKTHNGHFEYLVMLFGLTNAPATFQSLMNSVFRDFLRKFVLVFFDDILIYSSSIE 744
Query: 761 EHRDHLKLVLTVLLSNSFVANQSKCKFGCAQIDYLGHIISGAGVSVDPEKVQCIVEWPEP 820
EH++HL+LV V+ + A SK ++LGH IS + DP K+Q + EWP P
Sbjct: 745 EHKEHLRLVFEVMRLHKLFAKGSK--------EHLGHFISAREIETDPAKIQAVKEWPTP 796
Query: 821 KNVKGVRGFLGLTGYYRKFIKNYGKMAKPLTELTKKDNFTWGEEAKQAFHKLKTVMTSSP 880
VK VRGFLG GYYR+F++N+G +A PL LTK D F W EA+ AF LK V+ ++P
Sbjct: 797 TTVKQVRGFLGFAGYYRRFVRNFGVIAGPLHALTKTDGFCWSLEAQSAFDTLKAVLCNAP 856
Query: 881 VLALPDFDKSFEVECDAAGRGIGAVLMQQRQPIAYFSKALSNGNLTKSVYEKELMALVLC 940
VLALP FDK F VE DA G+GI AVLMQ+ P+AY S+ L L S+YEKEL+A +
Sbjct: 857 VLALPVFDKQFMVETDACGQGIRAVLMQKGHPLAYISRQLKGKQLHLSIYEKELLAFIFA 916
Query: 941 IQHWRHYLLGKAFTVYTDHKSLKHFLQQKISSPDQQCWLAKLLGYQFEVKYKPGQENKAA 1000
++ WRHYLL F + TD +SLK+ L+Q++++P QQ WL KLL + +E++Y+ G+EN A
Sbjct: 917 VRKWRHYLLPSHFIIKTDQRSLKYLLEQRLNTPVQQQWLPKLLEFDYEIQYRQGKENLVA 976
Query: 1001 DALSRRGDESDFFSLVSFPLWDDRKKLLEEIASDPYIKNLLQDVQQDPQSKPGFSVLQGV 1060
DALSR +S D K++ SD +K+++ +QQ P +K +S Q +
Sbjct: 977 DALSRVEGSEVLHMALSIVECDFLKEIQVAYESDGVLKDIISALQQHPDAKKHYSWSQDI 1036
Query: 1061 LLYQGRLVLSPTSPSIPWLLQEYHGSPTGGHSGFLRTYRRLAESLYWVGMQRTVRDFVRA 1120
L + ++V+ LLQ H S GG SG +++R+ YW GM + ++ F+R+
Sbjct: 1037 LRRKSKIVVPNDVEITNKLLQWLHCSGMGGRSGRDASHQRVKSLFYWKGMVKDIQAFIRS 1096
Query: 1121 CDTCQRQKYSAMSPGGLLQPFPVPNAVWEDLSLDFIIGLPKSKGYEAILVVVDRLSKYSH 1180
C TCQ+ K + GLLQP P+P+ +W D+S+DFI GLP S G I+VVVDRLSK +H
Sbjct: 1097 CGTCQQCKSDNAAYPGLLQPLPIPDKIWCDVSMDFIEGLPNSGGKSVIMVVVDRLSKAAH 1156
Query: 1181 FILLKHPYTAKSIAEVFVREIVRLHGIPNTVKSDRDPIFVSHFWSELFKLQGTKLKMSSS 1240
F+ L HPY+A ++A+ F+ + + HG P ++ SDRD +F S FW E FKLQG +L+MSS+
Sbjct: 1157 FVALAHPYSALTVAQAFLDNVYKHHGCPTSIVSDRDVLFTSDFWKEFFKLQGVELRMSSA 1216
Query: 1241 YHPETDGQTEVINRCLESYLRCFASDHPKTWSFWVPWAEFWYNSTFHCSIGQTPFEVVYG 1300
YHP++DGQTEV+NRCLE+YLRC P W+ W+P AE+WYN+ +H S TPFE+VYG
Sbjct: 1217 YHPQSDGQTEVVNRCLENYLRCMCHARPHLWNKWLPLAEYWYNTNYHSSSQMTPFELVYG 1276
Query: 1301 RKAPPIIRFLTNETKVVAVALELSERDEALKQLRTHLQRAQEQMASYANKKRRDLSFDIG 1360
+ P + +L ++KV VA L ER+ L L+ HL RAQ +M +A++ R + +FDIG
Sbjct: 1277 QAPPIHLPYLPGKSKVAVVARSLQERENMLLFLKFHLMRAQHRMKQFADQHRTERTFDIG 1336
Query: 1361 EWVFLKLRPHRQHSVVKRINQKLAARFYGPFQIEDKIGTVAYKLKLPVESRIHPVFHVSL 1420
++V++KL+P+RQ SVV R+NQKL+ +++GP++I +K G V
Sbjct: 1337 DFVYVKLQPYRQQSVVLRVNQKLSPKYFGPYKIIEKCGEV-------------------- 1376
Query: 1421 LKRAIGNYQVQGQLPTDLGIEEATDIYPEAILGTRIIRQGDSEVHQSLIKWKNRSIDDVT 1480
+GN QLP+ L + + PE IL +++++ L+KW +++ T
Sbjct: 1377 ---MVGNVTTSTQLPSVL--PDIFEKAPEYILERKLVKRQGRAATMVLVKWIGEPVEEAT 1431
Query: 1481 WE 1482
W+
Sbjct: 1432 WK 1433
>UniRef100_Q7XSK7 OSJNBa0059D20.8 protein [Oryza sativa]
Length = 1463
Score = 930 bits (2403), Expect = 0.0
Identities = 504/1177 (42%), Positives = 689/1177 (57%), Gaps = 53/1177 (4%)
Query: 341 RAMRVLILGDGETMDEDGEIVMLEEEAEEIEEEETVECKLMGVLGCMGEHQTQTMKLAGQ 400
R R L DG +D+ V +E +A + E + + G T++L
Sbjct: 293 RFCRRLFFVDGVEIDD----VAIEGDAAAAAGD--TEAPVFSLHAVAGVPIADTIQLQVT 346
Query: 401 VANVELLILIDSGASHNFISPKITSALGLVITPITTRSIKLGDGHKVVTRGICRGIKAKL 460
V + LL L+D G++H+FI + GL I + + +G +V G+ R +
Sbjct: 347 VGDASLLALLDGGSTHSFIGEEAARRAGLPIQSSPRMTAIVANGERVACPGVIRDAAFTI 406
Query: 461 GSIEITIDALVLELGGLDLVLGVSWLSTLGKVIMDWKLLTMQFVYGEQVVQLQGMRGKDS 520
D V+ L G D+VLG WL TLG ++ D+ +M F Q +G+
Sbjct: 407 NGSTFHTDLFVMPLAGFDVVLGTRWLGTLGPIVWDFTSRSMAFQRDGQRFAWKGVAST-- 464
Query: 521 YQSNLHSFLSDTHGREQMEWWGPQLQVVEATKKGHEIPQELRAVLADFHEVFTKKIQLPP 580
S TH R G L +L + +VF + LPP
Sbjct: 465 ---------STTHLRTLAAASGTLLD----------------ELLVAYEDVFGEPTGLPP 499
Query: 581 VRSRVHQITLNPEHGPINVRPYRYPHHQKEEIERQVTELLEAGIIRPSMSSFSSPVILVK 640
R R H I L P P+ VRPYRYP K+E+ERQ ++E G++R S S FSSPV+LVK
Sbjct: 500 PRGRDHAIVLKPSSAPVAVRPYRYPAAHKDELERQCAAMIEQGVVRRSDSPFSSPVLLVK 559
Query: 641 KKDKSWRMCVDYRALNKATIPDKYPIPIVDELLDELNGAVIFSKIDLKSGYHQIRVHEED 700
K D SWR CVDYRALN T+ D +PIP+VDEL +GA F+K+DL+SGYHQ+R+ ED
Sbjct: 560 KADGSWRFCVDYRALNALTVKDAFPIPVVDEL----HGARFFTKLDLRSGYHQVRMRPED 615
Query: 701 IPKTTFRTHNGHYEYLVMPFGLMNAPATFQAIMNDIFRPYLRKFVLVFFDDILIYSKSLL 760
+ KT FRTH+G YE+LVMPFGL NAPATFQA+MND+ RP+LR+FVLVFFDDILIYS++
Sbjct: 616 VHKTAFRTHDGLYEFLVMPFGLCNAPATFQALMNDVLRPFLRRFVLVFFDDILIYSETWT 675
Query: 761 EHRDHLKLVLTVLLSNSFVANQSKCKFGCAQIDYLGHIISGAGVSVDPEKVQCIVEWPEP 820
+H HL+ VL+VL + +SKC FG + YLGH+IS AGV++DP KVQ I EW P
Sbjct: 676 DHLRHLRTVLSVLRQHRLFVKRSKCTFGSPSVSYLGHVISEAGVAMDPAKVQAIHEWLVP 735
Query: 821 KNVKGVRGFLGLTGYYRKFIKNYGKMAKPLTELTKKDNFTWGEEAKQAFHKLKTVMTSSP 880
++ + VR FLGL GYYRKF+ NYG +A PLT LTKKD F+W E+ AF LK +TS+P
Sbjct: 736 RSARAVRSFLGLAGYYRKFVHNYGTIAAPLTALTKKDGFSWTEDTAAAFDALKAAVTSAP 795
Query: 881 VLALPDFDKSFEVECDAAGRGIGAVLMQQRQPIAYFSKALSNGNLTKSVYEKELMALVLC 940
VLA+PDF K F VE DA+ G GAVL+Q P+A+FS+ + + + YE+EL+ LV
Sbjct: 796 VLAMPDFAKPFTVEGDASTHGFGAVLVQDGHPVAFFSRPVVLRHRALAAYERELIGLVHA 855
Query: 941 IQHWRHYLLGKAFTVYTDHKSLKHFLQQKISSPDQQCWLAKLLGYQFEVKYKPGQENKAA 1000
++HWR YL G+ F V TDH SLK+ L Q++++ Q W+ KLLG+ F V+YKPG N A
Sbjct: 856 VRHWRPYLWGRRFVVKTDHYSLKYLLDQRLATIPQHHWVGKLLGFDFAVEYKPGAANTVA 915
Query: 1001 DALSRRGDESDFFSLVSFPLWDDRKKLLEEIASDPYIKNLLQDVQQDPQSKPGFSVLQGV 1060
DALSRR E +S P +D KL + DP + L +V ++ P ++++ +
Sbjct: 916 DALSRRDTEEGAILALSAPRFDFISKLHDAQRQDPALTALRDEVSAGTRTGP-WALVDDL 974
Query: 1061 LLYQGRLVLSPTSPSIPWLLQEYHGSPTGGHSGFLRTYRRLAESLYWVGMQRTVRDFVRA 1120
L Y L + P SP +++ H GH G RT RL + M++ V+D+VR+
Sbjct: 975 LQYNSWLYIPPASPLAREIIEATH---EDGHEGVKRTMHRLRREFHIPNMKQLVQDWVRS 1031
Query: 1121 CDTCQRQKYSAMSPGGLLQPFPVPNAVWEDLSLDFIIGLPKSKGYEAILVVVDRLSKYSH 1180
C CQR K +SP GLL P PVP VW D++LDFI LP+ +G IL VVDR SKY H
Sbjct: 1032 CAVCQRYKSEHLSPAGLLLPLPVPQGVWTDIALDFIEALPRVRGKSVILTVVDRFSKYCH 1091
Query: 1181 FILLKHPYTAKSIAEVFVREIVRLHGIPNTVKSDRDPIFVSHFWSELFKLQGTKLKMSSS 1240
FI L HPY+A+S+A+ F EIV LHG+P ++ SDRDPIF S FW EL +L GTKL M+++
Sbjct: 1092 FIPLAHPYSAESVAQAFFAEIVHLHGVPQSMVSDRDPIFTSTFWRELMRLMGTKLHMTTA 1151
Query: 1241 YHPETDGQTEVINRCLESYLRCFASDHPKTWSFWVPWAEFWYNSTFHCSIGQTPFEVVYG 1300
+HP++DGQ+E NR + YLRC D P+ W W+PWAEF +N+ + S+ TPF VVYG
Sbjct: 1152 FHPQSDGQSEAANRVIIMYLRCLTGDRPRQWLRWLPWAEFIFNTAYQSSLRDTPFRVVYG 1211
Query: 1301 RKAPPIIRFLTNETKVVAVALELSERDEALKQLRTHLQRAQEQMASYANKKRRDLSFDIG 1360
R P I + +T+V AVA + ER E L +R L++AQ + +K R +++ +G
Sbjct: 1212 RDPPSIRSYEAGDTRVAAVAKSMEERAEFLFDIRYRLEQAQAVQKLHYDKHHRHVAYQVG 1271
Query: 1361 EWVFLKLRPHRQHSVVKRINQKLAARFYGPFQIEDKIGTVAYKLKLPVESRIHPVFHVSL 1420
+W L+LR S+ + KL RFYGP++I + I VA +L+LP +R+H VFH+ L
Sbjct: 1272 DWALLRLRQRPTTSLPQSGTGKLKPRFYGPYRITELINDVAVRLELPAGARLHDVFHIGL 1331
Query: 1421 LKRAIGNYQVQGQLPTDLGIEE-ATDIYPEAILGTRIIRQGDSEVHQSLIKWKNRSIDDV 1479
LK+ G G P + A PE + R+ R V Q+L++WK S
Sbjct: 1332 LKKFHG--PPPGAPPALPPLHHGAIAPEPERAVRFRLAR----GVRQALVQWKGESPASA 1385
Query: 1480 TWEDNEVLAGQFPEFCLEDKAVSKEGGIERDSNTKWG 1516
TWED EVL ++P LED+ +EGG + WG
Sbjct: 1386 TWEDIEVLRAKYPALQLEDELSLEEGG-----DVMWG 1417
>UniRef100_Q7XJX3 OSJNBa0004L19.16 protein [Oryza sativa]
Length = 1586
Score = 899 bits (2322), Expect = 0.0
Identities = 471/1173 (40%), Positives = 689/1173 (58%), Gaps = 21/1173 (1%)
Query: 336 HKCP-ERAMRVLILGDGETMDE--DGEIV------MLEEEAEEIEEEETVECKLMGVLGC 386
H+C ++A+ L++ E DE +GE+ LE+E ++E E +
Sbjct: 419 HQCKVKKALNALLMEGEEGKDEGEEGELTGNQEDCKLEKEEAPPDDENQEELMFVSHNAV 478
Query: 387 MGEHQTQTMKLAGQVANVELLILIDSGASHNFISPKITSALGLVITPITTRSIKLGDGHK 446
G + T + Q+ + L+DSG++ F+ + + + + G +
Sbjct: 479 YGTTRPDTFSVIIQINGRRAVGLVDSGSTSTFMDQDYAVRNHCPLVSTDAKKVVVAGGGE 538
Query: 447 VVTRGICRGIKAKLGSIEITIDALVLELGGLDLVLGVSWLSTLGKVIMDWKLLTMQFVYG 506
+ + + ++ + ++ L G D++LG W+ + +D K + G
Sbjct: 539 LKSEVQVPELVYQIQGETFSNKFNIIPLKGYDVILGADWIYKYSPITLDLKKRELGITKG 598
Query: 507 EQVVQLQGMRGKDSYQSNLHSFLSDTHGREQMEWWGP----QLQVVEATKKGHEIPQELR 562
E+ V +Q D + H ++ + + G Q+ V+ + HEIP++++
Sbjct: 599 EKTVVIQ-----DFTRPGKHLWVDSKKVDQILRKGGLGCLFQITRVKEEETSHEIPEDIK 653
Query: 563 AVLADFHEVFTKKIQLPPVRSRVHQITLNPEHGPINVRPYRYPHHQKEEIERQVTELLEA 622
+L +F V LPP R+ H ITL P N+RPYR PH+QKE +E+ + EL+E+
Sbjct: 654 EILQEFPAVLKDPKGLPPRRNCDHVITLKSGAEPPNLRPYRVPHYQKEAMEKIIAELIES 713
Query: 623 GIIRPSMSSFSSPVILVKKKDKSWRMCVDYRALNKATIPDKYPIPIVDELLDELNGAVIF 682
I+ S +SSP ++V+KKD SWR+CVDYR LN T+ +K+P+PI+++LLDELNGA +F
Sbjct: 714 KEIQVSDIPYSSPAVMVRKKDGSWRLCVDYRQLNAQTVKNKFPMPIIEDLLDELNGAKVF 773
Query: 683 SKIDLKSGYHQIRVHEEDIPKTTFRTHNGHYEYLVMPFGLMNAPATFQAIMNDIFRPYLR 742
SK+DL+SGYHQIR+ +DIPKT FRTH GHYEY VMPFGL NAP TFQ++MN + P+LR
Sbjct: 774 SKLDLRSGYHQIRMATQDIPKTAFRTHLGHYEYQVMPFGLTNAPTTFQSLMNQVLAPFLR 833
Query: 743 KFVLVFFDDILIYSKSLLEHRDHLKLVLTVLLSNSFVANQSKCKFGCAQIDYLGHIISGA 802
K+VLVFFDDILIYSK EH++H++ V+ VL N V KC FG + YLGHIIS
Sbjct: 834 KYVLVFFDDILIYSKDWAEHKEHIRQVMKVLEENKLVVKLKKCAFGLPSVTYLGHIISQD 893
Query: 803 GVSVDPEKVQCIVEWPEPKNVKGVRGFLGLTGYYRKFIKNYGKMAKPLTELTKKDNFTWG 862
GV+ DP+KV+ I +P PK+V +R FLG+TGYYR+FIKNYG + +PL ++ KK+ F W
Sbjct: 894 GVATDPKKVEKIATYPTPKSVTDLRKFLGMTGYYRRFIKNYGIVCRPLHDMLKKEGFQWE 953
Query: 863 EEAKQAFHKLKTVMTSSPVLALPDFDKSFEVECDAAGRGIGAVLMQQRQPIAYFSKALSN 922
E +AF LKT M +SPVL+LPDF K F +E DA G GIGAVLMQ +P+AYFSK L
Sbjct: 954 REQTEAFETLKTHMCTSPVLSLPDFTKEFVIEADACGNGIGAVLMQSGRPLAYFSKTLGP 1013
Query: 923 GNLTKSVYEKELMALVLCIQHWRHYLLGKAFTVYTDHKSLKHFLQQKISSPDQQCWLAKL 982
+S+YEKE MA++ ++ WRHY+LG + TD +SLK + Q++ Q L KL
Sbjct: 1014 KAAAQSIYEKEAMAILEALKKWRHYVLGSRLIIKTDQQSLKFMMNQRLVEGIQHKLLLKL 1073
Query: 983 LGYQFEVKYKPGQENKAADALSRRGDESDFFSLVS-FPLWDDRKKLLEEIASDPYIKNLL 1041
+ Y + ++YK G+EN ADALSR ++ + P W + + D +L
Sbjct: 1074 MEYDYSIEYKAGKENLVADALSRIPPAEQCQAITTVIPEW--VRDIQRSYEGDVQAHKIL 1131
Query: 1042 QDVQQDPQSKPGFSVLQGVLLYQGRLVLSPTSPSIPWLLQEYHGSPTGGHSGFLRTYRRL 1101
+ + + +S G+L Y+GR+ + + L++ YH S GGHSG TY R+
Sbjct: 1132 SLIGTEGDTDGSYSQEAGLLRYKGRIYVGENTEIREELIRSYHSSAFGGHSGMRATYHRI 1191
Query: 1102 AESLYWVGMQRTVRDFVRACDTCQRQKYSAMSPGGLLQPFPVPNAVWEDLSLDFIIGLPK 1161
YW G+++ V F+R C CQ K + GLL P VP+ W +++DF+ GLPK
Sbjct: 1192 KSLFYWPGLKKAVEGFIRECPICQVTKAEHIHIPGLLDPLEVPDMAWAHITMDFVEGLPK 1251
Query: 1162 SKGYEAILVVVDRLSKYSHFILLKHPYTAKSIAEVFVREIVRLHGIPNTVKSDRDPIFVS 1221
S G + ILVVVDRL+KY+HFI + HPYT + + E+F+ I RLHG+P + +DRD IF S
Sbjct: 1252 SNGKDVILVVVDRLTKYAHFIAMAHPYTVEQVVELFMNNIHRLHGMPMAIITDRDRIFTS 1311
Query: 1222 HFWSELFKLQGTKLKMSSSYHPETDGQTEVINRCLESYLRCFASDHPKTWSFWVPWAEFW 1281
+ E+FK +LK S+SYHP+TDGQTE +N+CLESYLR P W W+ AE+W
Sbjct: 1312 QLFQEIFKSMKVRLKFSTSYHPQTDGQTERVNQCLESYLRSMTFQEPTRWHSWLALAEWW 1371
Query: 1282 YNSTFHCSIGQTPFEVVYGRKAPPIIRFLTNETKVVAVALELSERDEALKQLRTHLQRAQ 1341
YN+T+H SI TPF+ +YG P I F + + ++D +++L+ L AQ
Sbjct: 1372 YNTTYHTSIQMTPFQALYGYPPPQINEFSVPCNVSEEARVTIEQKDAIIQKLKYSLTEAQ 1431
Query: 1342 EQMASYANKKRRDLSFDIGEWVFLKLRPHRQHSVVKRINQKLAARFYGPFQIEDKIGTVA 1401
++ YA++ R + + +G+ V+LKL+P+RQ + R + KL ++FYGPF+I +K+G VA
Sbjct: 1432 RRIKHYADRNRSERTLAVGDMVYLKLQPYRQTAFGIRGSLKLRSKFYGPFKIMEKVGRVA 1491
Query: 1402 YKLKLPVESRIHPVFHVSLLKRAIGNYQVQGQLPTDLGIEEATDIYPEAILGTRIIRQGD 1461
YKL+LP S IHPVFHVS LK+ IG+ V +G + P A+L R+I +G
Sbjct: 1492 YKLQLPEGSNIHPVFHVSQLKKHIGSRAVPMANLPSVGPDGQIKTEPVAVLKRRMIPRGG 1551
Query: 1462 SEVHQSLIKWKNRSIDDVTWEDNEVLAGQFPEF 1494
V Q L+ W N S + TWED ++ FP F
Sbjct: 1552 VAVTQWLVLWHNLSPSEATWEDASMIQSMFPSF 1584
>UniRef100_Q8RUU3 Putative gag-pol polyprotein [Oryza sativa]
Length = 1338
Score = 893 bits (2307), Expect = 0.0
Identities = 467/1138 (41%), Positives = 674/1138 (59%), Gaps = 74/1138 (6%)
Query: 388 GEHQTQTMKLAGQVANVELLILIDSGASHNFISPKITSALGLVITPITTRSIKLGDGHKV 447
G + M + + + +L LIDSG H F+S + +G P ++ + +G KV
Sbjct: 255 GVRASDAMHITAHLGDTDLYTLIDSGLMHTFLSQDTAARVGRAPQPRMGLNVTVANGDKV 314
Query: 448 VTRGICRGIKAKLGSIEITIDALVLELGGLDLVLGVSWLSTLGKVIMDWKLLTMQFVYGE 507
G+ + ++ E D + D+ +LTM F +
Sbjct: 315 ACPGVFPDMPLQIAGEEFATD------------------------VYDFTVLTMSFWHR- 349
Query: 508 QVVQLQGMRGKDSYQSNLHSFLSDTHGREQMEWWGPQLQVVEATKKGHEIPQELRAVLAD 567
V L G+ G H R + P L ++L +
Sbjct: 350 --VTLHGLPG---------------HQRPRALACAPAAL--------------LDSLLEE 378
Query: 568 FHEVFTKKIQLPPVRSRVHQITLNPEHGPINVRPYRYPHHQKEEIERQVTELLEAGIIRP 627
F +VFT+ LPP R R H+I L P P+ VRPYRYP K+E+ERQ + E G+I
Sbjct: 379 FADVFTEPTGLPPARDRSHRIQLLPGTAPVAVRPYRYPVRHKDELERQCRVMEENGLIHR 438
Query: 628 SMSSFSSPVILVKKKDKSWRMCVDYRALNKATIPDKYPIPIVDELLDELNGAVIFSKIDL 687
S S+FSSPV+LVKK D SWR CV+YRALN+ T+ DKYPIP+VDELLDEL+GA IFSK+DL
Sbjct: 439 STSAFSSPVLLVKKADGSWRFCVNYRALNERTVKDKYPIPVVDELLDELHGAAIFSKLDL 498
Query: 688 KSGYHQIRVHEEDIPKTTFRTHNGHYEYLVMPFGLMNAPATFQAIMNDIFRPYLRKFVLV 747
+SGYHQ+R+H +DI KT FRTH+G YE+LV+PFGL NA ATFQ++MND+ RP+LR+FVLV
Sbjct: 499 RSGYHQVRMHPDDIDKTAFRTHDGLYEFLVIPFGLTNALATFQSLMNDVLRPFLRRFVLV 558
Query: 748 FFDDILIYSKSLLEHRDHLKLVLTVLLSNSFVANQSKCKFGCAQIDYLGHIISGAGVSVD 807
FFDDIL+YS + H HL+ V T L + +KC FG + YLGHIIS G ++D
Sbjct: 559 FFDDILVYSPTWTSHLQHLRTVFTALWAAQLFVKHTKCSFGDPSVAYLGHIISQHGFAMD 618
Query: 808 PEKVQCIVEWPEPKNVKGVRGFLGLTGYYRKFIKNYGKMAKPLTELTKKDNFTWGEEAKQ 867
K+Q + EWP P++ K +RGFLGL YYRKFI+++G +A PLT+L +KD+F W
Sbjct: 619 AAKIQAVAEWPRPRSPKELRGFLGLASYYRKFIQDFGSVAAPLTQLLRKDSFAWAPATDD 678
Query: 868 AFHKLKTVMTSSPVLALPDFDKSFEVECDAAGRGIGAVLMQQRQPIAYFSKALSNGNLTK 927
AF +LK +T++PVL+LPDF++ F VECDA+G G GAVL Q PIAYFS+ ++ +
Sbjct: 679 AFQRLKLALTTTPVLSLPDFNRPFVVECDASGTGFGAVLHQGEDPIAYFSRPIATRHHAL 738
Query: 928 SVYEKELMALVLCIQHWRHYLLGKAFTVYTDHKSLKHFLQQKISSPDQQCWLAKLLGYQF 987
+ YE+EL++LV ++HWR YL G+ F V TDH SLK L Q++S+ Q W++KLLG+ F
Sbjct: 739 AAYERELISLVQAVRHWRPYLWGRQFIVKTDHYSLKFLLDQRLSTIPQHHWVSKLLGFDF 798
Query: 988 EVKYKPGQENKAADALSRRGDESDFFSLVSFPLWDDRKKLLEEIASDPYIKNLLQDVQQD 1047
V+YKPG++N AADALS R ++S P +D K + +DP ++ L ++
Sbjct: 799 VVEYKPGKQNAAADALSCRAAPDSQAFVLSTPTFDLLKDIRTAGDTDPALQALRDEINSG 858
Query: 1048 PQSKPGFSVLQGVLLYQGRLVLSPTSPSIPWLLQEYHGSPTGGHSGFLRTYRRLAESLYW 1107
++ P ++V+ G++ Y+ R+ + P SP + ++ H GH G +T RL +
Sbjct: 859 TRTTP-WAVIDGLVTYKRRIYIPPGSPWVSVVVAAAHDD---GHEGIQKTLHRLRRDFHT 914
Query: 1108 VGMQRTVRDFVRACDTCQRQKYSAMSPGGLLQPFPVPNAVWEDLSLDFIIGLPKSKGYEA 1167
+R V D ++ C TCQR K + P GLL P PVP+A+W D+++DF+ GLP+ G
Sbjct: 915 PDDRRVVHDHIQGCLTCQRNKTDHLHPAGLLLPLPVPSAIWSDVAMDFVEGLPRVGGKSV 974
Query: 1168 ILVVVDRLSKYSHFILLKHPYTAKSIAEVFVREIVRLHGIPNTVKSDRDPIFVSHFWSEL 1227
IL VVDR SKY+H I L H YTA+++A F +IVRLHG+P ++ SDRDP+F S FW+ L
Sbjct: 975 ILTVVDRFSKYAHLIALAHSYTAETVARAFFVDIVRLHGVPESIVSDRDPVFTSAFWTAL 1034
Query: 1228 FKLQGTKLKMSSSYHPETDGQTEVINRCLESYLRCFASDHPKTWSFWVPWAEFWYNSTFH 1287
F TKL S+++HP++DGQ++ +N+ + LRC D + W W+PWAE+ YN++FH
Sbjct: 1035 FTATCTKLHRSTAFHPQSDGQSKAVNKAIAMCLRCMTGDRSRQWLRWLPWAEYIYNTSFH 1094
Query: 1288 CSIGQTPFEVVYGRKAPPIIRFLTNETKVVAVALELSERDEALKQLRTHLQRAQEQMASY 1347
++ TPF++VYGR P I + +E +V AVA + ERD L +R L++AQ+ Y
Sbjct: 1095 AALRDTPFKLVYGRDPPSIRAYDASELRVAAVAQSIEERDAFLADVRLRLEQAQQYAKRY 1154
Query: 1348 ANKKRRDLSFDIGEWVFLKLRPHRQHSVVKRINQKLAARFYGPFQIEDKIGTVAYKLKLP 1407
++K R++SF++G WV+L++R S+ + + KL RFYGP+++ I VAY+L LP
Sbjct: 1155 YDQKHREVSFEVGAWVWLRVRHRVPASLPEAVKGKLRPRFYGPYRVVAVINEVAYRLALP 1214
Query: 1408 VESRIHPVFHVSLLKRAIGNYQVQGQLPTDLGIEE--ATDIYPEAILGTRIIRQGDSEVH 1465
+R+H VFHV LLK +G V P L + A P +L R+ R V
Sbjct: 1215 PGTRLHDVFHVGLLKPFVG---VSPSAPLALPPIQHGAAQPVPRQVLRARLAR----GVR 1267
Query: 1466 QSLIKWKNRSIDDVTWEDNEVLAGQFPEFCLEDKAVSKEGGIERDSNTKWGMGLGKPK 1523
Q L++W+ +WED + ++P F L D+ + IE + WG+ + K
Sbjct: 1268 QLLVQWEGLPASATSWEDLDDFRNRYPSFQLADELL-----IEGGRDVMWGIPFRRRK 1320
>UniRef100_Q7XEK0 Hypothetical protein [Oryza sativa]
Length = 1611
Score = 888 bits (2295), Expect = 0.0
Identities = 477/1175 (40%), Positives = 709/1175 (59%), Gaps = 22/1175 (1%)
Query: 336 HKCP-ERAMRVLILGDGETMDEDGEIVMLEE-----EAEEIEEE-ETVECKLMGVL--GC 386
H+C ++A+ L++ + E+++ + + V EE + E++ E+ E V+ +LM +
Sbjct: 441 HQCKVKQAIHALLVENEESVEVEEDSVEEEEIKGEKQGEKLPEQTENVQEELMSISQSAV 500
Query: 387 MGEHQTQTMKLAGQVANVELLILIDSGASHNFISPKITSALGLVITPITTRSIKLGDGHK 446
G + T + +V + + L+DSG++ F+ K + R + + G +
Sbjct: 501 YGLTRPDTFSVMIKVNGKKAVGLVDSGSTTTFMDSKFAIKSQCTLENTKMRKVIVAGGGE 560
Query: 447 VVTRGICRGIKAKLGSIEITIDALVLELGGLDLVLGVSWLSTLGKVIMDWKLLTMQFVYG 506
+ + I G++ ++ T +L L D++LG W+ + +D + M+ G
Sbjct: 561 LKSELIVPGMEYEIQGESFTNSFNLLSLERYDIILGADWIFKYSPITLDLRKREMKITKG 620
Query: 507 EQVVQLQGMRGKDSYQSNLHSFLSDTHGREQMEWWGPQLQVVEATKKGHE---IPQELRA 563
+ +++Q Y + + + + G +Q+ T + + IP++++
Sbjct: 621 GRELEIQDFTKPGKYFQVSNKKMGKMIKKGAL---GCVIQINAITDQSNVEVGIPKDIQI 677
Query: 564 VLADFHEVFTKKIQLPPVRSRVHQITLNPEHGPINVRPYRYPHHQKEEIERQVTELLEAG 623
VL F +V + LPP RS H I L P N+RPYR PH QK +E +TEL
Sbjct: 678 VLQSFPKVLKEPKGLPPRRSCDHVINLKVGSEPPNLRPYRVPHFQKGAMEDIITELFRTQ 737
Query: 624 IIRPSMSSFSSPVILVKKKDKSWRMCVDYRALNKATIPDKYPIPIVDELLDELNGAVIFS 683
IR S S + SP ++V+KKD SWR+CVDYR LN TI +K+P+PI+++LLDEL+GA +FS
Sbjct: 738 EIRISDSPYVSPAVMVRKKDGSWRLCVDYRQLNAQTIKNKFPMPIIEDLLDELHGAKVFS 797
Query: 684 KIDLKSGYHQIRVHEEDIPKTTFRTHNGHYEYLVMPFGLMNAPATFQAIMNDIFRPYLRK 743
K+DL+SGYHQIR+ E DIPKT FRTH GHYEY VMPFGL NAPATFQA+MN + P+LRK
Sbjct: 798 KLDLRSGYHQIRMAEGDIPKTAFRTHLGHYEYNVMPFGLTNAPATFQALMNQVLAPFLRK 857
Query: 744 FVLVFFDDILIYSKSLLEHRDHLKLVLTVLLSNSFVANQSKCKFGCAQIDYLGHIISGAG 803
FVLVFF DILIYSK+ EH +H+KLV+ L +N V KC+FG ++ YLGHIIS G
Sbjct: 858 FVLVFFADILIYSKTQSEHLEHIKLVMQALSANQLVVRLKKCEFGLDRVSYLGHIISSEG 917
Query: 804 VSVDPEKVQCIVEWPEPKNVKGVRGFLGLTGYYRKFIKNYGKMAKPLTELTKKDNFTWGE 863
VS DP+K+ I PKNV VR FLG+ GYYR+FIK YG + +PL +L KKD F WG+
Sbjct: 918 VSTDPKKISDIKNRKPPKNVTEVREFLGMAGYYRRFIKGYGVICRPLHDLLKKDGFKWGD 977
Query: 864 EAKQAFHKLKTVMTSSPVLALPDFDKSFEVECDAAGRGIGAVLMQQRQPIAYFSKALSNG 923
++AF LK M +SPVLALPDF + F +E DA G GIGAVLMQ+ +P+AYFSKAL
Sbjct: 978 TQQEAFELLKEKMCNSPVLALPDFSQPFVIETDACGIGIGAVLMQKGRPLAYFSKALGPK 1037
Query: 924 NLTKSVYEKELMALVLCIQHWRHYLLGKAFTVYTDHKSLKHFLQQKISSPDQQCWLAKLL 983
+SVYEKE +A++ ++ WRHY+LG + + TD +SLK + Q++ Q L KL+
Sbjct: 1038 AAAQSVYEKEAIAILEALKKWRHYILGGSLIIKTDQQSLKFMMSQRLVEGIQHKLLLKLM 1097
Query: 984 GYQFEVKYKPGQENKAADALSRR---GDESDFFSLVSFPLW-DDRKKLLEEIASDPYIKN 1039
+ + ++YK G+EN ADALSR +E V P W D K+ EE D +
Sbjct: 1098 EFDYVIEYKSGKENLVADALSRSPNLKEEQCLPITVVVPEWVQDIKRSYEE---DIFAHK 1154
Query: 1040 LLQDVQQDPQSKPGFSVLQGVLLYQGRLVLSPTSPSIPWLLQEYHGSPTGGHSGFLRTYR 1099
+L ++ D + + + G+L Y+GR+ + T+ LL+ YH S GGHSG TY
Sbjct: 1155 ILSLIETDGDPERHYKLESGLLKYKGRIYVGETTEIRMLLLEAYHASYFGGHSGIRATYH 1214
Query: 1100 RLAESLYWVGMQRTVRDFVRACDTCQRQKYSAMSPGGLLQPFPVPNAVWEDLSLDFIIGL 1159
R+ + YW G+++ V ++R C TCQ K + GLL P VP+ W +++DFI GL
Sbjct: 1215 RIKQLFYWPGLKKQVEHYIRECPTCQITKAEHIHIPGLLNPLEVPDMAWTHITMDFIEGL 1274
Query: 1160 PKSKGYEAILVVVDRLSKYSHFILLKHPYTAKSIAEVFVREIVRLHGIPNTVKSDRDPIF 1219
PKS+G + ILVVVDRL+KY+HF+ L HPYT + + ++F+ I +LHG+P + +DRD +F
Sbjct: 1275 PKSQGKDVILVVVDRLTKYAHFLALSHPYTVEQVVQIFMDNIHKLHGMPMVIVTDRDRVF 1334
Query: 1220 VSHFWSELFKLQGTKLKMSSSYHPETDGQTEVINRCLESYLRCFASDHPKTWSFWVPWAE 1279
S+F+ E+FK Q KL+ S+++HP+TDGQTE +N+CLESYLR P+ W W+ AE
Sbjct: 1335 TSNFFQEIFKTQKVKLRFSTAHHPQTDGQTERVNQCLESYLRSMTFQEPQKWFSWLALAE 1394
Query: 1280 FWYNSTFHCSIGQTPFEVVYGRKAPPIIRFLTNETKVVAVALELSERDEALKQLRTHLQR 1339
+WYN+T+H SI TPF+ +YG P I F + L ++ L++L++ +
Sbjct: 1395 WWYNTTYHTSIQMTPFQALYGYPPPQITEFAIPCNMSEEARVTLEDKALILQKLKSSIGE 1454
Query: 1340 AQEQMASYANKKRRDLSFDIGEWVFLKLRPHRQHSVVKRINQKLAARFYGPFQIEDKIGT 1399
AQ ++ YA+K R + + ++G+ V+LKL+P+RQ ++ R + KL +++YGPF++ +K+G
Sbjct: 1455 AQRRIKFYADKGRSERTLELGDMVYLKLQPYRQVAMGIRGSLKLRSKYYGPFKVIEKMGA 1514
Query: 1400 VAYKLKLPVESRIHPVFHVSLLKRAIGNYQVQGQLPTDLGIEEATDIYPEAILGTRIIRQ 1459
VAYKL+LP + IHPVFHVS LK+ +G + +G + P A+L R+I +
Sbjct: 1515 VAYKLQLPDGAGIHPVFHVSQLKKHLGARAIPMPNLPAIGPDGQIKTEPAAVLQRRMIPR 1574
Query: 1460 GDSEVHQSLIKWKNRSIDDVTWEDNEVLAGQFPEF 1494
+ V Q LI W+N + + TWED + FP F
Sbjct: 1575 HNEPVTQWLILWENLTPAEATWEDASYIQAAFPNF 1609
>UniRef100_Q8SA91 Putative gag-pol polyprotein [Zea mays]
Length = 2396
Score = 881 bits (2276), Expect = 0.0
Identities = 484/1133 (42%), Positives = 675/1133 (58%), Gaps = 32/1133 (2%)
Query: 388 GEHQTQTMKLAGQVANVELLILIDSGASHNFISPKITSALGLVITPITTRSIKLGDGHKV 447
G QT+KL G + N LLILIDSG+SH F++ ++ L V + +T +++ +G V
Sbjct: 448 GSTGRQTLKLNGSIQNHPLLILIDSGSSHTFLNDQLRPHLQGVTSMASTLQVQVANGAMV 507
Query: 448 VTRGICRGIKAKLGSIEITIDALVLELGGLDLVLGVSWLSTLGKVIMDWKLLTMQFVYGE 507
+ ++ + T D L L D+V+G+ WL + + +DW + Y
Sbjct: 508 TCHYKLLQAQWQIQNCSFTSDVSFLPLPYYDMVVGMDWLESFSPMRVDWAQKWLIIPYQG 567
Query: 508 QVVQLQGMRGKDSYQSNLHSFLSDTHGREQMEWWGPQLQVVEATKKGHEIPQE---LRAV 564
V LQG N +DT +L +E+ P ++A+
Sbjct: 568 SSVLLQG---------NTAGVPADTV---------IELLFMESASSVSSSPDSHPAIQAL 609
Query: 565 LADFHEVFTKKIQLPPVRSRVHQITLNPEHGPINVRPYRYPHHQKEEIERQVTELLEAGI 624
L F VF + LPP R H I L P++VRPYRYP K++IE+QV E+L G+
Sbjct: 610 LQQFSSVFAEPQGLPPSRDCDHAIPLVEGAQPVSVRPYRYPPALKDKIEKQVQEMLHQGV 669
Query: 625 IRPSMSSFSSPVILVKKKDKSWRMCVDYRALNKATIPDKYPIPIVDELLDELNGAVIFSK 684
I+ S SSF+SPV+LVKKKD +WR CVDYR LN T+ KYP+P+ D+L+DEL + FSK
Sbjct: 670 IQKSNSSFASPVLLVKKKDMTWRFCVDYRYLNALTLKSKYPVPVFDQLIDELAHSKWFSK 729
Query: 685 IDLKSGYHQIRVHEEDIPKTTFRTHNGHYEYLVMPFGLMNAPATFQAIMNDIFRPYLRKF 744
+DL++GYHQI + + KT F+TH GHYE+ VM FGL AP TF + MN+ +P LRK
Sbjct: 730 LDLRAGYHQILLKPGEEYKTAFQTHVGHYEFRVMAFGLTGAPNTFLSAMNETLKPVLRKC 789
Query: 745 VLVFFDDILIYSKSLLEHRDHLKLVLTVLLSNSFVANQSKCKFGCAQIDYLGHIISGAGV 804
LVFFDDILIYSKS EH HL+ VL +LLS+++ SKC+F YLGHIIS GV
Sbjct: 790 ALVFFDDILIYSKSFEEHLLHLQKVLQLLLSDNWKVKLSKCEFAKTNTAYLGHIISEQGV 849
Query: 805 SVDPEKVQCIVEWPEPKNVKGVRGFLGLTGYYRKFIKNYGKMAKPLTELTKKDN-FTWGE 863
S P K+Q I W P + K +R FLGL G+YRKF+K++G +++PL +L KK F W
Sbjct: 850 STYPSKIQAISSWAVPTSAKELRCFLGLAGFYRKFVKHFGIISRPLFDLLKKHTLFVWTV 909
Query: 864 EAKQAFHKLKTVMTSSPVLALPDFDKSFEVECDAAGRGIGAVLMQQRQPIAYFSKALSNG 923
+ +AF LK + ++PVLALPDF + F + DA+ G+GAVLMQ P+A+ SKAL
Sbjct: 910 DHSKAFEVLKQALVTAPVLALPDFSQPFCIHTDASYYGVGAVLMQSGHPLAFLSKALGPK 969
Query: 924 NLTKSVYEKELMALVLCIQHWRHYLLGKAFTVYTDHKSLKHFLQQKISSPDQQCWLAKLL 983
N S YEKE MA++L I WR YL F +YTDH+SL +Q++ + QQ KL
Sbjct: 970 NQGLSTYEKEYMAIILAIAQWRSYLQLAEFIIYTDHRSLAQLNEQRLHTIWQQKMYTKLA 1029
Query: 984 GYQFEVKYKPGQENKAADALSRRGDESDFFSLVSFPLWDDRKKLLEEIASDPYIKNLLQD 1043
G Q+++ Y+ G +N AADALSR+ E +S + ++++E DP K LL
Sbjct: 1030 GLQYKIVYRKGVDNGAADALSRKVQEDSHCCAISHSVPTWLQEVVEGYDKDPTSKQLLAQ 1089
Query: 1044 VQQDPQSKPGFSVLQGVLLYQGRLVLSPTSPSIPWLLQEYHGSPTGGHSGFLRTYRRLAE 1103
+ + K FS+ QG++ ++ R+ L +LQ H + GGHSG TY ++ +
Sbjct: 1090 LILNSADKAPFSLHQGIIRHKNRIWLGGNLQLQQKVLQAMHDTAVGGHSGAPATYHKVKQ 1149
Query: 1104 SLYWVGMQRTVRDFVRACDTCQRQKYSAMSPGGLLQPFPVPNAVWEDLSLDFIIGLPKSK 1163
YW GM+ V +V++C CQ+ K GLLQP VP W +SLDFI GLP+S
Sbjct: 1150 MFYWPGMRADVLQYVQSCTVCQQSKPDRAKYPGLLQPLEVPPQAWHTISLDFIEGLPRSA 1209
Query: 1164 GYEAILVVVDRLSKYSHFILLKHPYTAKSIAEVFVREIVRLHGIPNTVKSDRDPIFVSHF 1223
Y ILVVVD+ SKY HF+ L HP+TA +A VF+ + +LHG+P + SDRD IF S F
Sbjct: 1210 HYNCILVVVDKFSKYGHFLPLLHPFTAAKVARVFLDNVYKLHGLPVNIISDRDRIFTSSF 1269
Query: 1224 WSELFKLQGTKLKMSSSYHPETDGQTEVINRCLESYLRCFASDHPKTWSFWVPWAEFWYN 1283
W +LF++ GT L MSSSYHP++DGQTE +N+CLE++LRC+ P WS W+ AE+WYN
Sbjct: 1270 WQQLFQITGTNLSMSSSYHPQSDGQTERLNQCLETFLRCYVHTCPSRWSAWLSVAEYWYN 1329
Query: 1284 STFHCSIGQTPFEVVYGRKAPPIIRFLTNETKVVAVALE--LSERDEALKQLRTHLQRAQ 1341
+T H ++G+TPFEV+YG P L +T V LE L ER+ K ++ HL RAQ
Sbjct: 1330 TTVHSTLGRTPFEVLYGH-TPRHFGILV-DTVVPQPELETWLKERELMTKVIKLHLHRAQ 1387
Query: 1342 EQMASYANKKRRDLSFDIGEWVFLKLRPHRQHSVVKRINQKLAARFYGPFQIEDKIGTVA 1401
++M A+K+R + F +G+WV+LKL+P+ Q SV R N KL+ +F+GPFQI D++G+VA
Sbjct: 1388 DRMKRQADKQRSERVFSVGDWVYLKLQPYIQSSVATRSNHKLSFKFFGPFQITDRLGSVA 1447
Query: 1402 YKLKLPVESRIHPVFHVSLLKRAIGNYQ-VQGQLPTDLGIEEATDIYPEAILGTRIIRQG 1460
Y+L LP S IHP+FHVS LKR IG Q QLP D+G + P IL R I +G
Sbjct: 1448 YRLALPASSSIHPIFHVSQLKRVIGRDQRASPQLPQDVGPIQV----PTRILQRRFIDRG 1503
Query: 1461 DSEVHQSLIKWKNRSIDDVTWEDNEVLAGQFPEFCLEDKAVSK-EGGIERDSN 1512
+ Q + W + D TWED E L +FP+ + D+A ++ +G + + S+
Sbjct: 1504 GELIAQVKVVWSGMTEDLATWEDVEALRARFPKALIWDQAGARGQGNVSKASS 1556
>UniRef100_Q60E20 Putative polyprotein [Oryza sativa]
Length = 1475
Score = 879 bits (2271), Expect = 0.0
Identities = 473/1165 (40%), Positives = 683/1165 (58%), Gaps = 33/1165 (2%)
Query: 350 DGETMDEDGEIVMLEEEAEEIEEEETVECK-------LMGVLGCMGEHQTQTMKLAGQVA 402
D ET + G+ EEE +E EE T E + + G + T + ++
Sbjct: 322 DKETSNTGGD----EEEKKETEESATSENESPTEELMYISQTAVQGTSRPDTFSVLIKIN 377
Query: 403 NVELLILIDSGASHNFISPKITSALGLVITPITTRSIKLGDGHKVVTRGICRGIKAKLGS 462
+ L+DSG++ F+ + T+ + + G ++ T + I ++
Sbjct: 378 GRTAVGLVDSGSTTTFMDQDYALRNYYPLKNTDTKKVVVAGGGELKTDVMVPDISYEIQG 437
Query: 463 IEITIDALVLELGGLDLVLGVSWLSTLGKVIMDWKLLTMQFVYGEQVVQLQGMRGKDSYQ 522
T +L L G D++LG W+ + +D K + G +V+ LQ D +
Sbjct: 438 ECFTNQFKLLPLKGYDIILGADWIYNYSPISLDLKQRILGITKGNKVILLQ-----DFTK 492
Query: 523 SNLHSFLSDTHGREQMEWWGP----QLQVVEAT--KKGHEIPQELRAVLADFHEVFTKKI 576
N H +S + ++ Q+ V+ T ++GH IP+++ ++ F V +
Sbjct: 493 PNKHFQISGKRLEKMLKKGALGMVIQVNVMSETVEEEGHVIPEDISDIIQQFPAVLKEPK 552
Query: 577 QLPPVRSRVHQITLNPEHGPINVRPYRYPHHQKEEIERQVTELLEAGIIRPSMSSFSSPV 636
LPP R H I L P N+RPYR PH+QKE +E + EL+E+ I+ S S +SSP
Sbjct: 553 GLPPKRECDHVINLQSGAVPPNIRPYRVPHYQKEAMENIINELIESKEIQTSDSPYSSPA 612
Query: 637 ILVKKKDKSWRMCVDYRALNKATIPDKYPIPIVDELLDELNGAVIFSKIDLKSGYHQIRV 696
++V+KKD SWRMCVDYR LN T+ +K+P+PI+++LLDELNGA IFSK+DL+SGYHQIR+
Sbjct: 613 VMVRKKDGSWRMCVDYRQLNAQTVKNKFPMPIIEDLLDELNGARIFSKLDLRSGYHQIRM 672
Query: 697 HEEDIPKTTFRTHNGHYEYLVMPFGLMNAPATFQAIMNDIFRPYLRKFVLVFFDDILIYS 756
E+D+ KT FRTH GHYEY VMPFGL N PATFQ++MN + P+LR+FVLVFFDDILIYS
Sbjct: 673 AEKDVHKTAFRTHLGHYEYQVMPFGLTNDPATFQSLMNHVLAPFLRRFVLVFFDDILIYS 732
Query: 757 KSLLEHRDHLKLVLTVLLSNSFVANQSKCKFGCAQIDYLGHIISGAGVSVDPEKVQCIVE 816
K+ EH +H+KLV+ L N V KC FG A + YLGH+IS GV+ DP+KV I
Sbjct: 733 KTRAEHLEHVKLVMQALQDNHLVIKLKKCAFGLASVSYLGHVISQDGVATDPKKVGKIKN 792
Query: 817 WPEPKNVKGVRGFLGLTGYYRKFIKNYGKMAKPLTELTKKDNFTWGEEAKQAFHKLKTVM 876
WP PK+V VR FLG+TGYYR+FI+ YG + +P+ ++ KK+ F WG + AF LK +
Sbjct: 793 WPTPKDVTDVRKFLGMTGYYRRFIQGYGTICRPIHDMLKKNGFQWGADQTTAFETLKHKL 852
Query: 877 TSSPVLALPDFDKSFEVECDAAGRGIGAVLMQQRQPIAYFSKALSNGNLTKSVYEKELMA 936
+SPVLALPDFD++F +E DA G GIGAVLMQ +PIA+FSKAL +S+YEKE MA
Sbjct: 853 RTSPVLALPDFDQAFTIEADACGVGIGAVLMQGGRPIAFFSKALGPKAAGQSIYEKEAMA 912
Query: 937 LVLCIQHWRHYLLGKAFTVYTDHKSLKHFLQQKISSPDQQCWLAKLLGYQFEVKYKPGQE 996
++ ++ WRHY+LG + TD +SLK + Q++ Q L KL+ Y + ++YK G+E
Sbjct: 913 ILEALKKWRHYVLGSKLIIKTDQQSLKFMMGQRLVEGIQHKLLLKLMEYDYTIEYKSGKE 972
Query: 997 NKAADALSRRGDESDFFS-----LVSFPLW--DDRKKLLEEIASDPYIKNLLQDVQQDPQ 1049
N ADALSR + V P W D ++ ++ + + L DP
Sbjct: 973 NLVADALSRLPQKEAVADRCHPMTVVIPEWIVDIQRSYENDVQAHKILS--LIGTAADPD 1030
Query: 1050 SKPGFSVLQGVLLYQGRLVLSPTSPSIPWLLQEYHGSPTGGHSGFLRTYRRLAESLYWVG 1109
+ + + G+L Y+GR+ + + L+ YH S GGHSG T+ R+ YW G
Sbjct: 1031 RE--YKLEAGLLKYKGRIYVGEATDIRRQLITTYHSSSFGGHSGMRATHHRIKMLFYWHG 1088
Query: 1110 MQRTVRDFVRACDTCQRQKYSAMSPGGLLQPFPVPNAVWEDLSLDFIIGLPKSKGYEAIL 1169
M+ V F+R C TCQ K + GLL P +P+ W +++DFI GLPKS+G + IL
Sbjct: 1089 MRGEVERFIRECPTCQITKSEHVHIPGLLNPLEIPDMAWTHITMDFIEGLPKSQGKDVIL 1148
Query: 1170 VVVDRLSKYSHFILLKHPYTAKSIAEVFVREIVRLHGIPNTVKSDRDPIFVSHFWSELFK 1229
VVVDRL+KY+HFI L HPY + + E F+ I +LHG+P + +DRD IF S + E+FK
Sbjct: 1149 VVVDRLTKYAHFIALAHPYDVEQVVEAFMNNIHKLHGMPMVIITDRDRIFTSSLFQEIFK 1208
Query: 1230 LQGTKLKMSSSYHPETDGQTEVINRCLESYLRCFASDHPKTWSFWVPWAEFWYNSTFHCS 1289
KL+ S++YHP+ DGQTE +N+CLESYLR P W W+ AE+WYN+TFH +
Sbjct: 1209 AMKVKLRFSTAYHPQMDGQTERVNQCLESYLRNMTFQEPHKWYSWLALAEWWYNTTFHTA 1268
Query: 1290 IGQTPFEVVYGRKAPPIIRFLTNETKVVAVALELSERDEALKQLRTHLQRAQEQMASYAN 1349
I TPF+ +YG P I F + + E++ L +L+ L AQ +M +A+
Sbjct: 1269 IQMTPFKAMYGYSPPQINEFSVPCNISEEARVTIEEKEAILNKLKNSLADAQHRMKYFAD 1328
Query: 1350 KKRRDLSFDIGEWVFLKLRPHRQHSVVKRINQKLAARFYGPFQIEDKIGTVAYKLKLPVE 1409
K R + + ++G+ V+LKL+P+RQ + R + KL ++FYGPF++ KIG +AYKL+LP +
Sbjct: 1329 KNRTERNLEVGDMVYLKLKPYRQSAFGIRGSLKLRSKFYGPFKVLQKIGQLAYKLQLPDD 1388
Query: 1410 SRIHPVFHVSLLKRAIGNYQVQGQLPTDLGIEEATDIYPEAILGTRIIRQGDSEVHQSLI 1469
++IHPVFHVS LK+ +G + + +G + P A+L R++ + V Q LI
Sbjct: 1389 AQIHPVFHVSQLKKHLGKHAIPMSNLPSVGPDGQIKTEPLAVLQRRMVPRKGVAVTQWLI 1448
Query: 1470 KWKNRSIDDVTWEDNEVLAGQFPEF 1494
W+N S + TWED V+ FP F
Sbjct: 1449 LWQNLSPAEATWEDASVIQAMFPSF 1473
>UniRef100_Q947Y5 Putative retroelement [Oryza sativa]
Length = 1923
Score = 863 bits (2231), Expect = 0.0
Identities = 453/970 (46%), Positives = 620/970 (63%), Gaps = 21/970 (2%)
Query: 557 IPQELRAVLADFHEVFTKKIQLPPVRSRVHQITLNPEHGPINVRPYRYPHHQKEEIERQV 616
IP ++ VL F EVF + ++PPVR+ H+I L P+N+RPYR+ K+EIERQV
Sbjct: 36 IPDSIKHVLDRFQEVFQEPTEMPPVRNCDHKIPLMEGASPVNLRPYRHTPALKDEIERQV 95
Query: 617 TELLEAGIIRPSMSSFSSPVILVKKKDKSWRMCVDYRALNKATIPDKYPIPIVDELLDEL 676
TE+L++G+I+ S S+FSSP +LVKKKD +WR+C+DY+ LN TI KYP+P++DELLDEL
Sbjct: 96 TEMLQSGVIQNSNSAFSSPALLVKKKDGTWRLCIDYKHLNAITIKGKYPLPVIDELLDEL 155
Query: 677 NGAVIFSKIDLKSGYHQIRVHEEDIPKTTFRTHNGHYEYLVMPFGLMNAPATFQAIMNDI 736
+GA FSK+DL++GYHQIR+ + KT F+TH+GHYEY VM FGL APATFQ MND
Sbjct: 156 SGAKYFSKLDLRAGYHQIRLQPGEEHKTAFQTHSGHYEYRVMSFGLTGAPATFQKAMNDT 215
Query: 737 FRPYLRKFVLVFFDDILIYSKSLLEHRDHLKLVLTVLLSNSFVANQSKCKFGCAQIDYLG 796
LRKF LVFFDDILIYS L H HL+ VL +L + + SKC F Q+ YLG
Sbjct: 216 LATVLRKFTLVFFDDILIYSPDLPSHIQHLEQVLQLLQAQQWKVKLSKCSFAQQQLAYLG 275
Query: 797 HIISGAGVSVDPEKVQCIVEWPEPKNVKGVRGFLGLTGYYRKFIKNYGKMAKPLTELTKK 856
HII GV+ DP K+ ++ W P++VK +RGFLGL GYYRKF++N+G + KPLT+L KK
Sbjct: 276 HIIGKDGVTTDPSKIADVLHWKIPQSVKQLRGFLGLAGYYRKFVRNFGTINKPLTQLLKK 335
Query: 857 D-NFTWGEEAKQAFHKLKTVMTSSPVLALPDFDKSFEVECDAAGRGIGAVLMQQRQPIAY 915
F W + +AF+ LK + S+PVLALPDF K+F VE DA GIGAVL Q R PIA+
Sbjct: 336 GVPFKWTAQMDEAFNALKQALVSAPVLALPDFSKTFTVETDACDMGIGAVLSQDRHPIAF 395
Query: 916 FSKALSNGNLTKSVYEKELMALVLCIQHWRHYLLGKAFTVYTDHKSLKHFLQQKISSPDQ 975
SKAL S YEKE +A++L + WR YL F + TDH +L H Q++ +P Q
Sbjct: 396 VSKALGPKTRGLSTYEKEYLAILLAVDQWRSYLQHDEFVILTDHHNLMHITDQRLHTPLQ 455
Query: 976 QCWLAKLLGYQFEVKYKPGQENKAADALSRRGDE-SDFFSLVS--FPLWDDRKKLLEEIA 1032
KL+G Q++V Y+ G N AADALSRR +E +D VS PLW +++
Sbjct: 456 HKAFTKLMGLQYKVCYRRGTSNAAADALSRRDEETNDQLWAVSECQPLW--LTAVVKGYE 513
Query: 1033 SDPYIKNLLQDVQQDPQSKPGFSVLQGVLLYQGRLVLSPTSPSIPWLLQEYHGSPTGGHS 1092
+D + LL ++ P ++ F ++QGVL Y+G++ + L+ H SP GGHS
Sbjct: 514 TDEQAQQLLTELALHPTAREHFHLVQGVLRYKGKIWIGHNLSLQQQLVTALHASPIGGHS 573
Query: 1093 GFLRTYRRLAESLYWVGMQRTVRDFVRACDTCQRQKYSAMSPGGLLQPFPVPNAVWEDLS 1152
GF TY+R+ W M++ ++ +V+ C CQ+ K GLLQP P+P W+ +S
Sbjct: 574 GFPVTYQRVKALFAWPQMKKMIQQWVKNCTICQQAKPDLAKYPGLLQPLPIPEGAWQVVS 633
Query: 1153 LDFIIGLPKSKGYEAILVVVDRLSKYSHFILLKHPYTAKSIAEVFVREIVRLHGIPNTVK 1212
LDFI GLPKS+ Y ILVVVD+ S+Y+HF+ L HP++A +A +++ I +LHG+P +
Sbjct: 634 LDFIEGLPKSERYNCILVVVDKFSRYAHFVPLSHPFSALDVAVSYMKNIYKLHGMPKVLI 693
Query: 1213 SDRDPIFVSHFWSELFKLQGTKLKMSSSYHPETDGQTEVINRCLESYLRCFASDHPKTWS 1272
SDRD IF S W LF G L +SS+YHP++DGQT+ +N+CLE +LRCF + P W
Sbjct: 694 SDRDKIFTSKLWEFLFLKSGIALHLSSAYHPQSDGQTKRVNQCLEMFLRCFTNAAPFKWV 753
Query: 1273 FWVPWAEFWYNSTFHCSIGQTPFEVVYGRKAPPIIRFLTNETKVVA-VALELSERDEALK 1331
W+ A +WYN+ FH ++ +TPFEV+YG P+ +T ET + +A+ L ER +
Sbjct: 754 TWLHLAGYWYNTCFHSALNKTPFEVLYGHN--PLQLGVTMETCAIPDLAVWLHERKLMAE 811
Query: 1332 QLRTHLQRAQEQMASYANKKRRDLSFDIGEWVFLKLRPHRQHSVVKRINQKLAARFYGPF 1391
L+ HL R Q++M +A+K R F +G+WVFLKL+P+ Q SV R KLA +FYGPF
Sbjct: 812 LLQQHLHRVQQKMKFHADKNRSFREFAVGDWVFLKLQPYVQKSVASRACHKLAFKFYGPF 871
Query: 1392 QIEDKIGTVAYKLKLPVESRIHPVFHVSLLKRAIG-NYQVQGQLPTDLGIEEATDIYPEA 1450
QI ++GTVAYKL+LP +S IHPVFHVS LK A G +QVQ +LP L + +YP
Sbjct: 872 QILARVGTVAYKLQLPDDSTIHPVFHVSQLKVAHGFKHQVQSRLPKFL----KSTVYPLQ 927
Query: 1451 ILGTRIIRQGDSEVHQSLIKWKNRSIDDVTWEDNEVLAGQFPEFCLEDKAVSKEGGI--- 1507
IL R+IR+G+ V Q L+ W + +D TWED E L +FP +A + G+
Sbjct: 928 ILDQRLIRKGNRTVSQVLVYWSDSVAEDATWEDREDLQQRFPMALAWGQATFQGEGVVKI 987
Query: 1508 ----ERDSNT 1513
E+++NT
Sbjct: 988 FPDGEQENNT 997
>UniRef100_Q8W150 Polyprotein [Oryza sativa]
Length = 933
Score = 836 bits (2159), Expect = 0.0
Identities = 434/882 (49%), Positives = 577/882 (65%), Gaps = 18/882 (2%)
Query: 619 LLEAGIIRPSMSSFSSPVILVKKKDKSWRMCVDYRALNKATIPDKYPIPIVDELLDELNG 678
+L+ GII+ S S FSSP +LVKKKD SWR+C+DYR LN T YP+PI+DELLDEL G
Sbjct: 1 MLQNGIIQHSSSPFSSPALLVKKKDGSWRVCIDYRQLNAITKKGTYPMPIIDELLDELAG 60
Query: 679 AVIFSKIDLKSGYHQIRVHEEDIPKTTFRTHNGHYEYLVMPFGLMNAPATFQAIMNDIFR 738
A IFSK+DL++GYHQIR+ E + KT F+TH+GHYEY VM FGL APATFQ MND R
Sbjct: 61 AKIFSKLDLRAGYHQIRMAEGEEFKTAFQTHSGHYEYKVMSFGLTGAPATFQGAMNDTLR 120
Query: 739 PYLRKFVLVFFDDILIYSKSLLEHRDHLKLVLTVLLSNSFVANQSKCKFGCAQIDYLGHI 798
P LRK LVFFDDILIYS + H DHLK VL +L ++ + SKC F QI YLGHI
Sbjct: 121 PLLRKCALVFFDDILIYSPDMNSHLDHLKQVLQLLDTHQWKVKLSKCDFAQTQISYLGHI 180
Query: 799 ISGAGVSVDPEKVQCIVEWPEPKNVKGVRGFLGLTGYYRKFIKNYGKMAKPLTELTKKD- 857
ISG GVS DP K+Q IV+W P +K +RGFLGL GYYRKF+K++G ++KPLT+L KKD
Sbjct: 181 ISGQGVSTDPSKIQSIVDWAVPTTLKKLRGFLGLAGYYRKFVKDFGTLSKPLTQLLKKDA 240
Query: 858 NFTWGEEAKQAFHKLKTVMTSSPVLALPDFDKSFEVECDAAGRGIGAVLMQQRQPIAYFS 917
F W E QAF LK +TS+PVLALP+F + F +E DA+ GIGAVL Q + P+A+ S
Sbjct: 241 PFVWSAEVNQAFQALKHALTSTPVLALPNFQQGFTIETDASDIGIGAVLSQNQHPVAFVS 300
Query: 918 KALSNGNLTKSVYEKELMALVLCIQHWRHYLLGKAFTVYTDHKSLKHFLQQKISSPDQQC 977
KAL S YEKE +A+++ + HWR YL + F + TDH SL H +Q++ +P QQ
Sbjct: 301 KALGPRTQGLSTYEKECLAIMMAVDHWRPYLQFQEFLIITDHHSLMHLTEQRLHTPWQQK 360
Query: 978 WLAKLLGYQFEVKYKPGQENKAADALSRRGDE--SDFFSL-VSFPLWDDRKKLLEEIASD 1034
KL G QF++ Y+ G+ N AADALSR E S+F + P+W + +L D
Sbjct: 361 AFTKLSGLQFQIVYRKGKHNAAADALSRHVPEETSEFLGISTCSPVW--LQDILHGYDQD 418
Query: 1035 PYIKNLLQDVQQDPQSKPGFSVLQGVLLYQGRLVLSPTSPSIPWLLQEYHGSPTGGHSGF 1094
P +LL + +P S P +S+ +G++ ++G++ + S ++ H SP GGHSGF
Sbjct: 419 PLALSLLTGLAVNPSSYPHYSLSKGLIKHKGKVWVGNNSNIQQQIISALHDSPLGGHSGF 478
Query: 1095 LRTYRRLAESLYWVGMQRTVRDFVRACDTCQRQKYSAMSPGGLLQPFPVPNAVWEDLSLD 1154
TY+R+ W M+ TV+ + +C C + K GLLQP PVP+ W+ +S+D
Sbjct: 479 PVTYKRIKSLFSWPHMKLTVQKQLASCAVCLQAKPDRSKYPGLLQPLPVPDGAWQIISMD 538
Query: 1155 FIIGLPKSKGYEAILVVVDRLSKYSHFILLKHPYTAKSIAEVFVREIVRLHGIPNTVKSD 1214
FI GLPKS + ILVVVD+ SKY+HF+ L HP++A +A+VF+ + +LHG+P + SD
Sbjct: 539 FIEGLPKSYHQDCILVVVDKFSKYAHFMPLSHPFSALDVAKVFMLNVYKLHGLPQIIISD 598
Query: 1215 RDPIFVSHFWSELFKLQGTKLKMSSSYHPETDGQTEVINRCLESYLRCFASDHPKTWSFW 1274
RD IF S W +LF GTKL +SS+YHP++DGQTE +N+CLE +LRCF P WS W
Sbjct: 599 RDKIFTSALWEQLFLRSGTKLHLSSAYHPQSDGQTERVNQCLEIFLRCFVHATPAKWSLW 658
Query: 1275 VPWAEFWYNSTFHCSIGQTPFEVVYGRKAPPIIRFLTNETKVVA-VALELSERDEALKQL 1333
+ AEFWYNS++H ++ +TPFEV+YG PP + + V++ + LSER + L
Sbjct: 659 LHLAEFWYNSSYHSALNKTPFEVLYG--YPPSHFGIRADACVISDLDSWLSERHLMTQLL 716
Query: 1334 RTHLQRAQEQMASYANKKRRDLSFDIGEWVFLKLRPHRQHSVVKRINQKLAARFYGPFQI 1393
R HL RAQ+ M + A+KKR SF +G+WVFLKL+P+ Q SV KR N KL+ R++GP+QI
Sbjct: 717 RQHLNRAQQVMKTQADKKRSFRSFQVGDWVFLKLQPYVQSSVAKRANHKLSFRYFGPYQI 776
Query: 1394 EDKIGTVAYKLKLPVESRIHPVFHVSLLKRAIG-NYQVQGQLPTDLGIEEATD--IYPEA 1450
K+G+VAYKL+LP +S +HPVFHVS LK + +Q QLP TD YP
Sbjct: 777 LSKVGSVAYKLQLPADSMVHPVFHVSQLKGTQNFKHSIQSQLP------NITDHIQYPVQ 830
Query: 1451 ILGTRIIRQGDSEVHQSLIKWKNRSIDDVTWEDNEVLAGQFP 1492
IL TRI ++G+ V Q L+ W N + TWED E L +FP
Sbjct: 831 ILDTRIQKKGNKVVRQILVCWSNLPAVEATWEDEEELKQRFP 872
>UniRef100_Q75LZ2 Putative gag-pol polyprotein [Oryza sativa]
Length = 1377
Score = 782 bits (2019), Expect = 0.0
Identities = 436/1074 (40%), Positives = 624/1074 (57%), Gaps = 110/1074 (10%)
Query: 364 EEEAEEIEEEETVECKLMGVLGCMGEHQTQTMKLAGQVANVELLILIDSGASHNFISPKI 423
E++A E+ M V G T++L G++ + +L+L+DSG+ +FIS ++
Sbjct: 406 EDQASISSEDSDSSLLAMSVHAVQGTDSQSTIRLQGKIQGLNILMLVDSGSLASFISDQL 465
Query: 424 TSALGLVITPITTRSIKLGDGHKVVTRGICRGIKAKLGSIEITIDALVLELGGLDLVLGV 483
L V S+K+ +G + + + + + V+ LGG D +LG+
Sbjct: 466 ADRLTGVQFLAHPLSVKVANGQVLHCQKEMPSAEWFVSGQKFLTTFKVVALGGYDAILGM 525
Query: 484 SWLSTLGKVIMDWKLLTMQFVYGEQVVQLQGMRGKDSYQSNLHSFLSDTHGREQMEWWGP 543
WL+ + +DW
Sbjct: 526 DWLTKYSPMQIDW----------------------------------------------- 538
Query: 544 QLQVVEATKKGHEIPQELRAVLADFHE---VFTKKIQLPPVRSRVHQITLNPEHGPINVR 600
QL+ + T GHE+ LR V +D + + ++ +Q +H I + +NVR
Sbjct: 539 QLKSIHLTVMGHEVL--LRGVQSDTSDCALISSQHLQ------HLHSI----DSVAVNVR 586
Query: 601 PYRYPHHQKEEIERQVTELLEAGIIRPSMSSFSSPVILVKKKDKSWRMCVDYRALNKATI 660
PYRY QK+EIE+QV ++L+ GII+PS S FSSPV+LVKKKD +WR CVDYR LN T+
Sbjct: 587 PYRYTPAQKDEIEKQVKDMLQKGIIQPSASPFSSPVLLVKKKDGTWRFCVDYRHLNAITV 646
Query: 661 PDKYPIPIVDELLDELNGAVIFSKIDLKSGYHQIRVHEEDIPKTTFRTHNGHYEYLVMPF 720
+KYP+PI+DEL+DEL GA FSK+DL+SGYHQIR+ + KT F+THNGH+E+ V+PF
Sbjct: 647 KNKYPLPIIDELMDELAGACWFSKLDLRSGYHQIRMAVGEEAKTAFKTHNGHFEFKVLPF 706
Query: 721 GLMNAPATFQAIMNDIFRPYLRKFVLVFFDDILIYSKSLLEHRDHLKLVLTVLLSNSFVA 780
GL +APATFQ +MN + LR+ VLVF DDIL+YS++L EH++HL+ V
Sbjct: 707 GLTSAPATFQGVMNTVLADQLRQNVLVFVDDILVYSRTLEEHKNHLRQV----------- 755
Query: 781 NQSKCKFGCAQIDYLGHIISGAGVSVDPEKVQCIVEWPEPKNVKGVRGFLGLTGYYRKFI 840
+ L HIIS GVS DPEK+Q + +WP P +VK VR FLGL GYYRKF+
Sbjct: 756 -----------FETLRHIISADGVSTDPEKIQVVRQWPVPVSVKDVRSFLGLAGYYRKFV 804
Query: 841 KNYGKMAKPLTE-LTKKDNFTWGEEAKQAFHKLKTVMTSSPVLALPDFDKSFEVECDAAG 899
+++G ++KPLT+ L K F W + ++AF LK + S+PVL +PDF K F VE DA+
Sbjct: 805 RHFGIISKPLTKLLCKGQPFIWTQHHQEAFDTLKQSLISAPVLVMPDFQKMFVVETDASD 864
Query: 900 RGIGAVLMQQRQPIAYFSKALSNGNLTKSVYEKELMALVLCIQHWRHYLLGKAFTVYTDH 959
RGIGAVLMQ + +A+ SKAL + S YEKE +A++L + HWR YL F + TDH
Sbjct: 865 RGIGAVLMQDQHSVAFLSKALGHRTQVLSTYEKESLAIILAVDHWRPYLQHDDFLIRTDH 924
Query: 960 KSLKHFLQQKISSPDQQCWLAKLLGYQFEVKYKPGQENKAADALSRR-GDESDFFSLVSF 1018
+SL Q +++ Q + KLLG ++ + YK G EN AADALS R D S +S
Sbjct: 925 RSLAFLDNQLLTTSWQYKAMTKLLGLRYRIVYKKGLENGAADALSHRSSDGLPILSALSV 984
Query: 1019 PLWDDRKKLLEEIASDPYIKNLLQDVQQDPQSKPGFSVLQGVLLYQGRLVLSPTSPSIPW 1078
L D + ++ SDP LL ++ F + G+L ++ R+ +
Sbjct: 985 GLPDWLQDVVSGYKSDPEALQLLHSFKEGTCHSAHFELQNGILYFKKRVWVGNNRQLQQQ 1044
Query: 1079 LLQEYHGSPTGGHSGFLRTYRRLAESLYWVGMQRTVRDFVRACDTCQRQKYSAMSPGGLL 1138
+L H + GGHSG ++DFV++C CQ+ K + GLL
Sbjct: 1045 VLANLHTAALGGHSG--------------------IQDFVQSCAVCQQAKPEHVPYPGLL 1084
Query: 1139 QPFPVPNAVWEDLSLDFIIGLPKSKGYEAILVVVDRLSKYSHFILLKHPYTAKSIAEVFV 1198
QP +P W+ +S+DFI GLP+S Y +ILV+VD+ +K++HF+ L HP+TA S+A++F+
Sbjct: 1085 QPVQIPEHAWQVISMDFIEGLPRSASYNSILVIVDKFTKFAHFLPLSHPFTAVSVAQLFM 1144
Query: 1199 REIVRLHGIPNTVKSDRDPIFVSHFWSELFKLQGTKLKMSSSYHPETDGQTEVINRCLES 1258
I +HG+P + SDRD +F S FW ELF+ GT+L+MSSSYHP+TDGQTE +N+CLE
Sbjct: 1145 DRIHSIHGLPQAIISDRDRVFTSIFWKELFRRSGTQLQMSSSYHPQTDGQTERVNQCLEM 1204
Query: 1259 YLRCFASDHPKTWSFWVPWAEFWYNSTFHCSIGQTPFEVVYGRKAPPIIRFLTNETKVVA 1318
+LRCF P WS W+ A+FWYN++FH ++ TPFE ++G K P L+ ++ V
Sbjct: 1205 FLRCFVHACPSRWSKWLSLAQFWYNTSFHSTLMLTPFEAMFGHK--PRHFGLSVDSVGVP 1262
Query: 1319 VALE--LSERDEALKQLRTHLQRAQEQMASYANKKRRDLSFDIGEWVFLKLRPHRQHSVV 1376
AL+ + +R + HL RAQ++M S A+ KR D F +G+WVF+KL+P+ Q SV+
Sbjct: 1263 SALDSWMQDRHNMQSLIYHHLVRAQQRMKSQADSKRTDRVFAVGDWVFMKLQPYVQQSVM 1322
Query: 1377 KRINQKLAARFYGPFQIEDKIGTVAYKLKLPVESRIHPVFHVSLLKRAIGNYQV 1430
R NQKLA +FYGPFQI ++G+VAYKL+LP S IHPV HVS LK+A+ +V
Sbjct: 1323 TRANQKLAFKFYGPFQILQRVGSVAYKLQLPATSLIHPVVHVSQLKKAVAASEV 1376
>UniRef100_Q8LLZ2 Putative retroelement [Oryza sativa]
Length = 1786
Score = 754 bits (1948), Expect = 0.0
Identities = 439/1129 (38%), Positives = 641/1129 (55%), Gaps = 75/1129 (6%)
Query: 401 VANVELLILIDSGASHNFISPKITSALGLVITPITTRSIKLGDGHKVVTRGICRGIKAKL 460
V ++ IL DSGASH+FIS L + + + G + + + ++
Sbjct: 683 VHSIPATILFDSGASHSFISVPFVGKNQLGVERLRNPLLITTPGGVMTAKYYSPAVPIEI 742
Query: 461 GSIEITIDALVLELGGLDLVLGVSWLSTLGKVIMDWKLLTMQFVYGEQVVQLQGMRGKDS 520
I D ++L+ LD++LG++WL+ V+ D T+ G +
Sbjct: 743 QGIPFPSDLILLDTKNLDVILGMNWLAQFQGVV-DCARRTVTLYRGPE------------ 789
Query: 521 YQSNLHSFLSDTHGREQMEWWGPQLQVVEATKKGHEIPQELRAVLADFHEVFTKKIQ-LP 579
+ + ++ P V + K H+I ++ +F +VF +++ +P
Sbjct: 790 ---------------QPVVFFAPPTSV--RSSKLHQIELSEIPIVREFGDVFPEELPGMP 832
Query: 580 PVRSRVHQITLNPEHGPINVRPYRYPHHQKEEIERQVTELLEAGIIRPSMSSFSSPVILV 639
P R +I L P P++ RPYR ++ E+++Q+ EL E G IRPS S + +PVI V
Sbjct: 833 PKREIEFRIDLAPGTTPLHKRPYRMAANELAEVKKQLEELKEKGYIRPSTSPWGAPVIFV 892
Query: 640 KKKDKSWRMCVDYRALNKATIPDKYPIPIVDELLDELNGAVIFSKIDLKSGYHQIRVHEE 699
+KKDK+ RMCVDYRALN+ TI +KYP+P +D+L D+L GA +FSKIDL+SGYHQ+R+ EE
Sbjct: 893 EKKDKTKRMCVDYRALNEVTIKNKYPLPRIDDLFDQLKGATVFSKIDLRSGYHQLRIREE 952
Query: 700 DIPKTTFRTHNGHYEYLVMPFGLMNAPATFQAIMNDIFRPYLRKFVLVFFDDILIYSKSL 759
DIPKT F T G YE+ VM FGL NAPA F +MN +F YL KFV+VF DDILIYS+S
Sbjct: 953 DIPKTAFTTRYGLYEFTVMSFGLTNAPAFFMNLMNKVFMEYLDKFVVVFIDDILIYSQSE 1012
Query: 760 LEHRDHLKLVLTVLLSNSFVANQSKCKFGCAQIDYLGHIISGAGVSVDPEKVQCIVEWPE 819
+H+ HL+LVL L + A SKC+F +++ +LGH+IS GV+VDPE V + EW +
Sbjct: 1013 EDHQHHLRLVLGKLREHQLYAKLSKCEFWLSEVKFLGHVISAKGVAVDPETVTAVTEWKQ 1072
Query: 820 PKNVKGVRGFLGLTGYYRKFIKNYGKMAKPLTELTKK-DNFTWGEEAKQAFHKLKTVMTS 878
PK V +R FLGL GYYR+FI+N+ K+A+P+T+L KK + F W + ++AF LK + S
Sbjct: 1073 PKTVTQIRSFLGLAGYYRRFIENFSKIARPMTQLLKKEEKFVWSPQCEKAFQTLKEKLVS 1132
Query: 879 SPVLALPDFDKSFEVECDAAGRGIGAVLMQQRQPIAYFSKALSNGNLTKSVYEKELMALV 938
SPVL LPD K F V CDA+ +G+G VLMQ +AY S+ L ++ EL A+V
Sbjct: 1133 SPVLILPDTRKDFMVYCDASRQGLGCVLMQDGHVVAYASRQLRPHEGNYPTHDLELAAVV 1192
Query: 939 LCIQHWRHYLLGKAFTVYTDHKSLKHFLQQKISSPDQQCWLAKLLGYQFEVKYKPGQENK 998
++ WRHYL+G +YTDHKSLK+ Q + Q+ WL + Y + Y PG+ N
Sbjct: 1193 HALKIWRHYLIGNRCEIYTDHKSLKYIFTQSDLNLRQRRWLELIKDYDVGIHYHPGKANV 1252
Query: 999 AADALSRRGD-----------------ESDFFSLVS---FPLWDDRKKLLEEI----ASD 1034
ADALSR+ E+ S+V + + LL++I +D
Sbjct: 1253 VADALSRKSHCNTLNVRGIPPELNQQMEALNLSIVGRGFLAALEAKPTLLDQIREAQKND 1312
Query: 1035 PYIKNLLQDVQQDPQSKPGFSVLQGVLLYQGRLVLSPTSPSIPWL-LQEYHGSPTGGHSG 1093
P + LL++++Q + GF+ + L+ G+ V P S + L LQE H SP H G
Sbjct: 1313 PDMHGLLKNMRQGKAA--GFTEDEHGTLWNGKRVCVPDSRELKQLILQEAHESPYSIHPG 1370
Query: 1094 FLRTYRRLAESLYWVGMQRTVRDFVRACDTCQRQKYSAMSPGGLLQPFPVPNAVWEDLSL 1153
+ Y L E +WV M+R + +FV CD CQR K P GLLQP VP W+++ +
Sbjct: 1371 STKMYLDLKEKYWWVSMKREIAEFVALCDVCQRVKAEHQRPAGLLQPLQVPEWKWDEIGM 1430
Query: 1154 DFIIGLPKSK-GYEAILVVVDRLSKYSHFILLKHPYTAKSIAEVFVREIVRLHGIPNTVK 1212
DFI GLPK++ GY++I V+VDRL+K + FI +K Y +AE++ IV LHGIP +
Sbjct: 1431 DFITGLPKTQGGYDSIWVIVDRLTKVARFIPVKTTYGGNKLAELYFARIVSLHGIPKKIV 1490
Query: 1213 SDRDPIFVSHFWSELFKLQGTKLKMSSSYHPETDGQTEVINRCLESYLRCFASDHPKTWS 1272
SDR F SHFW +L + GT+L S++YHP+TDGQTE +N+ LE LR D KTW
Sbjct: 1491 SDRGSQFTSHFWKKLQEELGTRLNFSTAYHPQTDGQTERLNQILEDMLRACVLDFGKTWD 1550
Query: 1273 FWVPWAEFWYNSTFHCSIGQTPFEVVYGRKA-PPIIRFLTNETKVVAVALELSERDEALK 1331
+P+AEF YN+++ SI P+E +YGRK P++ E++V + L E + ++
Sbjct: 1551 KSLPYAEFSYNNSYQASIQMAPYEALYGRKCRTPLLWDQVGESQVFGTDI-LREAEAKVR 1609
Query: 1332 QLRTHLQRAQEQMASYANKKRRDLSFDIGEWVFLKLRPHRQHSVVKRINQ--KLAARFYG 1389
+R +L+ AQ + SYA+ +RRDL F + ++V+L++ P R V R KLA R+ G
Sbjct: 1610 IIRDNLKVAQSRQKSYADNRRRDLEFAVDDFVYLRVTPLRG---VHRFQTKGKLAPRYVG 1666
Query: 1390 PFQIEDKIGTVAYKLKLPVE-SRIHPVFHVSLLKRAIGNYQVQGQLPTDLGIEEATDI-- 1446
PF+I + G VAY+L+LP +H VFHVS LKR + +V + IE D+
Sbjct: 1667 PFRIIARRGEVAYQLELPASLGNVHDVFHVSQLKRCL---RVPSEQADSEQIEVREDLTY 1723
Query: 1447 --YPEAILGTRIIRQGDSEVHQSLIKWKNRSIDDVTWEDNEVLAGQFPE 1493
P IL TR R + + ++W N + ++ TWE + L P+
Sbjct: 1724 VERPVKILDTRERRTRNRVIRFCKVQWSNHAEEEATWEREDELKAAHPD 1772
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.324 0.140 0.434
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,597,605,663
Number of Sequences: 2790947
Number of extensions: 112340985
Number of successful extensions: 341756
Number of sequences better than 10.0: 29731
Number of HSP's better than 10.0 without gapping: 3083
Number of HSP's successfully gapped in prelim test: 26648
Number of HSP's that attempted gapping in prelim test: 281178
Number of HSP's gapped (non-prelim): 50760
length of query: 1543
length of database: 848,049,833
effective HSP length: 141
effective length of query: 1402
effective length of database: 454,526,306
effective search space: 637245881012
effective search space used: 637245881012
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 82 (36.2 bits)
Lotus: description of TM0265.11


