

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0263.7
(787 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
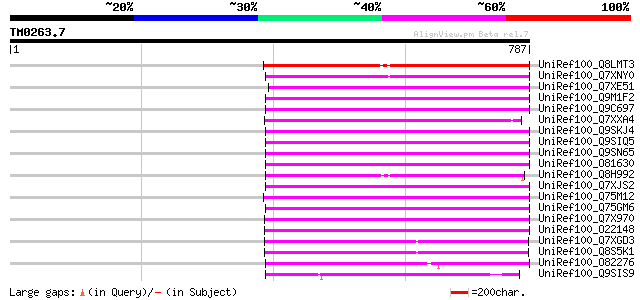
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q8LMT3 Putative retroelement [Oryza sativa] 302 3e-80
UniRef100_Q7XNY0 OSJNBb0015N08.11 protein [Oryza sativa] 292 3e-77
UniRef100_Q7XE51 Putative non-LTR retroelement reverse transcrip... 275 5e-72
UniRef100_Q9M1F2 Hypothetical protein F9K21.130 [Arabidopsis tha... 272 2e-71
UniRef100_Q9C697 Reverse transcriptase, putative; 16838-20266 [A... 272 2e-71
UniRef100_Q7XXA4 OSJNBa0019G23.12 protein [Oryza sativa] 271 4e-71
UniRef100_Q9SKJ4 Putative non-LTR retroelement reverse transcrip... 271 4e-71
UniRef100_Q9SIQ5 Putative non-LTR retroelement reverse transcrip... 271 4e-71
UniRef100_Q9SN65 Hypothetical protein F25I24.40 [Arabidopsis tha... 271 6e-71
UniRef100_O81630 F8M12.22 protein [Arabidopsis thaliana] 271 6e-71
UniRef100_Q8H992 Oryza sativa (japonica cultivar-group) retrotra... 263 1e-68
UniRef100_Q7XJS2 Putative non-LTR retroelement reverse transcrip... 262 3e-68
UniRef100_Q75M12 Hypothetical protein P0676G05.2 [Oryza sativa] 261 4e-68
UniRef100_Q75GM6 Putative non-LTR retroelement reverse transcrip... 261 4e-68
UniRef100_Q7X970 BZIP-like protein [Oryza sativa] 257 8e-67
UniRef100_O22148 Putative non-LTR retroelement reverse transcrip... 257 8e-67
UniRef100_Q7XGD3 Putative retroelement [Oryza sativa] 257 1e-66
UniRef100_Q8S5K1 Putative retroelement [Oryza sativa] 257 1e-66
UniRef100_O82276 Putative non-LTR retroelement reverse transcrip... 257 1e-66
UniRef100_Q9SIS9 Putative non-LTR retroelement reverse transcrip... 256 2e-66
>UniRef100_Q8LMT3 Putative retroelement [Oryza sativa]
Length = 764
Score = 302 bits (773), Expect = 3e-80
Identities = 166/404 (41%), Positives = 245/404 (60%), Gaps = 7/404 (1%)
Query: 385 LRGGNASFITLIPKVENPQGLGNYRPISLVGCVYKIVAKLLSMRLREVLSKVIDDRQTAF 444
+R N ITLIPKV++ + ++RPI L+ +K + K+L+ RL E+ KVI + QTAF
Sbjct: 28 MRRLNYGVITLIPKVKDANTIKSFRPICLLNVCFKFLTKILTRRLSEIAQKVIGESQTAF 87
Query: 445 IGGRFMLDSVVVANEVVEEAKRRKKECFVFKVDFEKAYDSVDWNFLFYMMSRIGFSHKWI 504
I GR LD VV+ +EV+ E K+ KK +FK+DFEKAYD V W+FLF ++ + GFS +WI
Sbjct: 88 IPGRQNLDGVVILHEVLHELKKEKKSGIIFKLDFEKAYDKVQWSFLFDVLHKKGFSDRWI 147
Query: 505 HWIKGCIQSASSSVLVNGSPTEEFNMGRGLRQGDPLAPFLFLIVAEGLNGLMRQAVQQKL 564
W+K +V +NG + F RG+RQGDPL+P LF +VA+ L+ ++ A Q
Sbjct: 148 QWVKMATIGGKMAVNINGEVKDFFKTYRGVRQGDPLSPLLFNLVADALSEMLNNAKQA-- 205
Query: 565 FTGYRVGGDEVEISLLQFADDTLFFGEASTQNILVVKSILRWFEAMSGLKVNFFKSKLAG 624
+ V G ++ LQ+ADDT+ F + +NI+ VK +L +EAMSGLK+N+ KS++
Sbjct: 206 -VPHLVPGG---LTHLQYADDTILFMTNTEENIVTVKFLLYCYEAMSGLKINYQKSEIMV 261
Query: 625 ISMVSRETQIFAAMLNCKVMGVPFTYLGIPVGANPRKAATWEPVIKKMQRKLSL*KHKSL 684
I ETQ A + NC+ +PFTYLGIP+ N A + K++++L+ K L
Sbjct: 262 IGGDEMETQRVADLFNCQAGKMPFTYLGIPISMNKLTNADLDIPPNKIEKRLATWKCGYL 321
Query: 685 SFGGRICLIKSVLSSLPLFFFSFFKAPKCIIDQCNKIQRKFLWGG-EEGRGVAWVKWEEI 743
S+GG+ LI S LSS+PL+ + P+ + ++ + I+ +F W G E+ R +KWE +
Sbjct: 322 SYGGKAILINSCLSSIPLYMMGVYLLPEGVHNKMDSIRARFFWEGLEKKRKYHMIKWEAL 381
Query: 744 CKKKEEGGLGVKDLESFNKALLVKWRWRLLREREWLWCQVLESK 787
C+ KE GGLG D N ALL KW +RL +E C +L +K
Sbjct: 382 CRPKEFGGLGFIDTRKMNIALLCKWIYRLESGKEDPCCVLLRNK 425
>UniRef100_Q7XNY0 OSJNBb0015N08.11 protein [Oryza sativa]
Length = 1026
Score = 292 bits (747), Expect = 3e-77
Identities = 150/400 (37%), Positives = 235/400 (58%), Gaps = 3/400 (0%)
Query: 389 NASFITLIPKVENPQGLGNYRPISLVGCVYKIVAKLLSMRLREVLSKVIDDRQTAFIGGR 448
N ITLIPK ++ + +RPI L+ +KI+ K+L RL ++ +I +QTAF+ R
Sbjct: 296 NFGVITLIPKTKDASQIQKFRPICLLNVSFKIITKVLMNRLNGTMAYIISKQQTAFLKNR 355
Query: 449 FMLDSVVVANEVVEEAKRRKKECFVFKVDFEKAYDSVDWNFLFYMMSRIGFSHKWIHWIK 508
F+++ VV+ +EV+ ++K+ +FKVDFEKAYD V+W F++ M+ GF +W WI
Sbjct: 356 FIMEGVVILHEVLNSIHQKKQSGILFKVDFEKAYDKVNWVFIYRMLKAKGFPDQWCDWIM 415
Query: 509 GCIQSASSSVLVNGSPTEEFNMGRGLRQGDPLAPFLFLIVAEGLNGLMRQAVQQKLFTGY 568
+ +V VN F +GLRQGDPL+P LF + AE L L+++A + L G
Sbjct: 416 KVVMGGKVAVKVNDQIGSFFKTHKGLRQGDPLSPLLFNLAAEALTLLVQRAEENSLIEGL 475
Query: 569 RVGGDEVEISLLQFADDTLFFGEASTQNILVVKSILRWFEAMSGLKVNFFKSKLAGISMV 628
GD +I++LQ+ADDT+F + +K IL FE +SGLK+NF KS++
Sbjct: 476 GTNGDN-KIAILQYADDTIFLINDKLDHAKNLKYILCLFEQLSGLKINFNKSEVFCFGEA 534
Query: 629 SRETQIFAAMLNCKVMGVPFTYLGIPVGANPRKAATWEPVIKKMQRKLSL*KHKSLSFGG 688
+ +++ + CKV +P YLGIP+ W+ KM+ KL + + S GG
Sbjct: 535 KEKQDLYSNIFTCKVGSLPLKYLGIPIDQKRILNKDWKLAENKMEHKLGCWQGRLQSIGG 594
Query: 689 RICLIKSVLSSLPLFFFSFFKAPKCIIDQCNKIQRKFLWGGEEG-RGVAWVKWEEICKKK 747
R+ L+ S LSS+P++ SF++ PK + ++ + +++FLW ++G R V W +C +
Sbjct: 595 RLILLNSTLSSVPMYMISFYRLPKGVQERIDYFRKRFLWQEDQGIRKYHLVNWPLVCSPR 654
Query: 748 EEGGLGVKDLESFNKALLVKWRWRLLREREWLWCQVLESK 787
++GGLGV DLE+ NKA+L KW WRL E W W +++ +K
Sbjct: 655 DQGGLGVLDLEAMNKAMLGKWIWRLENEEGW-WQEIIYAK 693
>UniRef100_Q7XE51 Putative non-LTR retroelement reverse transcriptase [Oryza sativa]
Length = 1652
Score = 275 bits (702), Expect = 5e-72
Identities = 152/396 (38%), Positives = 225/396 (56%), Gaps = 2/396 (0%)
Query: 393 ITLIPKVENPQGLGNYRPISLVGCVYKIVAKLLSMRLREVLSKVIDDRQTAFIGGRFMLD 452
ITL+PK ++ + YRPI L+ +KI K+++ R+ V KVI QT F+ GR +++
Sbjct: 1208 ITLLPKQKDASRIQQYRPICLLNVSFKIFTKVMANRIALVAQKVIKPSQTTFLSGRNIME 1267
Query: 453 SVVVANEVVEEAKRRKKECFVFKVDFEKAYDSVDWNFLFYMMSRIGFSHKWIHWIKGCIQ 512
VV+ +E + E ++KK + K+DFEKAYD VDW FL + GFS KW WI ++
Sbjct: 1268 GVVILHETLHELHKKKKNGVILKLDFEKAYDKVDWKFLQQSLRMKGFSSKWCDWIDSIVR 1327
Query: 513 SASSSVLVNGSPTEEFNMGRGLRQGDPLAPFLFLIVAEGLNGLMRQAVQQKLFTGYRVGG 572
S +V VN F +GLRQGDPL+P LF +V + L L+++A Q F G
Sbjct: 1328 GGSVAVKVNDEIGSYFQTRKGLRQGDPLSPILFNLVVDMLAILIQRAKDQGRFKGVVPHL 1387
Query: 573 DEVEISLLQFADDTLFFGEASTQNILVVKSILRWFEAMSGLKVNFFKSKLAGISMVSRET 632
+ +S+LQ+ADDT+ F + +K +L FE +S LK+NF+KS+L
Sbjct: 1388 VDNGLSILQYADDTILFMDHDLDEARDLKLVLSTFEKLSSLKINFYKSELFCYGKAKDVE 1447
Query: 633 QIFAAMLNCKVMGVPFTYLGIPVGANPRKAATWEPVIKKMQRKLSL*KHKSLSFGGRICL 692
+ + C PF YLGI + W+ V +++Q+KLS K K LS GGR+ L
Sbjct: 1448 HEYVKLFGCDTEDYPFKYLGIRMHHKRINNKDWQGVEERIQKKLSSWKGKFLSVGGRLVL 1507
Query: 693 IKSVLSSLPLFFFSFFKAPKCIIDQCNKIQRKFLWG-GEEGRGVAWVKWEEICKKKEEGG 751
I SVLS+L +F SFF+ PK I+ + + + +F W E + +W +CK KE GG
Sbjct: 1508 INSVLSNLAIFMLSFFEIPKGILKKLDYYRSRFFWQCDEHKKKYRLARWSVLCKPKECGG 1567
Query: 752 LGVKDLESFNKALLVKWRWRLLREREWLWCQVLESK 787
LG+++LE NK LL KW ++L+ E E +W +L +K
Sbjct: 1568 LGIQNLEIQNKCLLSKWLYKLINE-EGVWQDILRNK 1602
>UniRef100_Q9M1F2 Hypothetical protein F9K21.130 [Arabidopsis thaliana]
Length = 851
Score = 272 bits (696), Expect = 2e-71
Identities = 149/404 (36%), Positives = 231/404 (56%), Gaps = 5/404 (1%)
Query: 389 NASFITLIPKVENPQGLGNYRPISLVGCVYKIVAKLLSMRLREVLSKVIDDRQTAFIGGR 448
N + I +IPK++NPQ L +YRPI+L +YK+++K + RL+ L+ ++ D Q AFI GR
Sbjct: 106 NHTNICMIPKIQNPQTLSDYRPIALCNVLYKVISKCMVNRLKAHLNSIVSDSQAAFIPGR 165
Query: 449 FMLDSVVVANEVVEEAKRRK---KECFVFKVDFEKAYDSVDWNFLFYMMSRIGFSHKWIH 505
+ D+V++A+E++ K RK K K D KAYD V+W+FL M GF KWI
Sbjct: 166 IINDNVMIAHEIMHSLKVRKRVSKTYMAVKTDVSKAYDRVEWDFLETTMRLFGFCDKWIG 225
Query: 506 WIKGCIQSASSSVLVNGSPTEEFNMGRGLRQGDPLAPFLFLIVAEGLNGLMRQAVQQKLF 565
WI ++S SVL+NGSP + RG+RQGDPL+P+LF++ + L+ L++
Sbjct: 226 WIMAAVKSVHYSVLINGSPHGYISPTRGIRQGDPLSPYLFILCGDILSHLIKVKASSGDI 285
Query: 566 TGYRVGGDEVEISLLQFADDTLFFGEASTQNILVVKSILRWFEAMSGLKVNFFKSKLAGI 625
G R+G I+ LQFADD+LFF +A+ +N +K + +E SG K+N KS +
Sbjct: 286 RGVRIGNGAPAITHLQFADDSLFFCQANVRNCQALKDVFDVYEYYSGQKINVQKSLITFG 345
Query: 626 SMVSRETQI-FAAMLNCKVMGVPFTYLGIPVGANPRKAATWEPVIKKMQRKLSL*KHKSL 684
S V TQ +LN G YLG+P +K + +I +++ + + K L
Sbjct: 346 SRVYGSTQTRLKTLLNIPNQGGGGKYLGLPEQFGRKKKEMFNYIIDRVKERTASWSAKFL 405
Query: 685 SFGGRICLIKSVLSSLPLFFFSFFKAPKCIIDQCNKIQRKFLW-GGEEGRGVAWVKWEEI 743
S G+ L+KSV ++P++ S FK P+ I+ + + F W RG+ WV W+ +
Sbjct: 406 SPAGKEILLKSVALAMPVYAMSCFKLPQGIVSEIESLLMNFWWEKASNKRGIPWVAWKRL 465
Query: 744 CKKKEEGGLGVKDLESFNKALLVKWRWRLLREREWLWCQVLESK 787
K+EGGLG +DL FN ALL K WR+++ L+ +V++++
Sbjct: 466 QYSKKEGGLGFRDLAKFNDALLAKQAWRIIQYPNSLFARVMKAR 509
>UniRef100_Q9C697 Reverse transcriptase, putative; 16838-20266 [Arabidopsis thaliana]
Length = 1142
Score = 272 bits (696), Expect = 2e-71
Identities = 145/404 (35%), Positives = 229/404 (55%), Gaps = 5/404 (1%)
Query: 389 NASFITLIPKVENPQGLGNYRPISLVGCVYKIVAKLLSMRLREVLSKVIDDRQTAFIGGR 448
N + I LIPK E P + RPISL YK+++K+L RL+ VL +I + Q+AF+ GR
Sbjct: 269 NTTNICLIPKTERPTRMTELRPISLCNVGYKVISKILCQRLKTVLPNLISETQSAFVDGR 328
Query: 449 FMLDSVVVANEVVEEAKRR---KKECFVFKVDFEKAYDSVDWNFLFYMMSRIGFSHKWIH 505
+ D++++A E+ + K + K D KAYD V+WNF+ ++ ++GF KWI
Sbjct: 329 LISDNILIAQEMFHGLRTNSSCKDKFMAIKTDMSKAYDQVEWNFIEALLRKMGFCEKWIS 388
Query: 506 WIKGCIQSASSSVLVNGSPTEEFNMGRGLRQGDPLAPFLFLIVAEGLNGLMRQAVQQKLF 565
WI CI + VL+NG P RGLRQGDPL+P+LF++ E L +R+A +Q L
Sbjct: 389 WIMWCITTVQYKVLINGQPKGLIIPERGLRQGDPLSPYLFILCTEVLIANIRKAERQNLI 448
Query: 566 TGYRVGGDEVEISLLQFADDTLFFGEASTQNILVVKSILRWFEAMSGLKVNFFKSKLA-G 624
TG +V +S L FADD+LFF +A+ + ++ IL+ +E++SG ++NF KS + G
Sbjct: 449 TGIKVATPSPAVSHLLFADDSLFFCKANKEQCGIILEILKQYESVSGQQINFSKSSIQFG 508
Query: 625 ISMVSRETQIFAAMLNCKVMGVPFTYLGIPVGANPRKAATWEPVIKKMQRKLSL*KHKSL 684
+ +L +G +YLG+P K + V ++Q +++ K L
Sbjct: 509 HKVEDSIKADIKLILGIHNLGGMGSYLGLPESLGGSKTKVFSFVRDRLQSRINGWSAKFL 568
Query: 685 SFGGRICLIKSVLSSLPLFFFSFFKAPKCIIDQCNKIQRKFLWGGE-EGRGVAWVKWEEI 743
S GG+ +IKSV ++LP + S F+ PK I + KF W + RG+ W+ W+++
Sbjct: 569 SKGGKEVMIKSVAATLPRYVMSCFRLPKAITSKLTSAVAKFWWSSNGDSRGMHWMAWDKL 628
Query: 744 CKKKEEGGLGVKDLESFNKALLVKWRWRLLREREWLWCQVLESK 787
C K +GGLG ++++ FN ALL K WRL+ + L+ +V + +
Sbjct: 629 CSSKSDGGLGFRNVDDFNSALLAKQLWRLITAPDSLFAKVFKGR 672
>UniRef100_Q7XXA4 OSJNBa0019G23.12 protein [Oryza sativa]
Length = 1140
Score = 271 bits (694), Expect = 4e-71
Identities = 151/392 (38%), Positives = 218/392 (55%), Gaps = 5/392 (1%)
Query: 387 GGNASFITLIPKVENPQGLGNYRPISLVGCVYKIVAKLLSMRLREVLSKVIDDRQTAFIG 446
G N ITLIPKVE + YRPI L+ YKI K+ + R+ V +++ QTAF+
Sbjct: 719 GLNFGVITLIPKVEEANRIQQYRPICLLNVSYKIFTKVATNRISSVADNLVNPTQTAFMR 778
Query: 447 GRFMLDSVVVANEVVEEAKRRKKECFVFKVDFEKAYDSVDWNFLFYMMSRIGFSHKWIHW 506
GR +LD V + +E V E R+K +FK+DFEKAYD V W FL + GFS KWI W
Sbjct: 779 GRNILDGVAIIHETVHELHRKKLNGVIFKIDFEKAYDKVKWPFLLQTLRMKGFSPKWISW 838
Query: 507 IKGCIQSASSSVLVNGSPTEEFNMGRGLRQGDPLAPFLFLIVAEGLNGLMRQAVQQKLFT 566
IK I S +V VN F +GL+QGDPL+P LF + + L L+ +A Q
Sbjct: 839 IKSFIVGGSVAVKVNDDVGPFFQTKKGLQQGDPLSPILFNFIVDMLATLINRAKTQGQVD 898
Query: 567 GYRVGGDEVEISLLQFADDTLFFGEASTQNILVVKSILRWFEAMSGLKVNFFKSKLAGIS 626
G + +S+LQ+ DD + F + +KS+L FE +SGLK+NF KS+L
Sbjct: 899 GLIPHLIDGGLSILQYVDDIVLFMNHDLEKAQNMKSVLLAFEQLSGLKINFHKSELYCFG 958
Query: 627 MVSRETQIFAAMLNCKVMGVPFTYLGIPVGANPRKAATWEPVIKKMQRKLSL*KHKSLSF 686
+A + C+V PF YLGIP+ + A W+ V+++ ++KLS K K LS
Sbjct: 959 EALEYRDQYAQLFGCQVGNFPFRYLGIPIHCRKLRNAEWKEVVERFEKKLSSWKGKLLSL 1018
Query: 687 GGRICLIKSVLSSLPLFFFSFFKAPKCIIDQCNKIQRKFLWGGE-EGRGVAWVKWEEICK 745
GGR+ LI SVLSSLP++ SF P ++ + + ++ +F W G+ + KW+ IC+
Sbjct: 1019 GGRLTLINSVLSSLPMYMMSFLAIPSGVLKKLDYLRSRFYWQGDGHKKKYQLAKWDIICR 1078
Query: 746 KKEEGGLGVKDLESFNKALLVKW--RWRLLRE 775
K++G LG+ DLE L+ +W W +L +
Sbjct: 1079 PKDQGRLGIHDLEVI--TLVTRWLRTWAILHK 1108
>UniRef100_Q9SKJ4 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 1750
Score = 271 bits (694), Expect = 4e-71
Identities = 151/404 (37%), Positives = 229/404 (56%), Gaps = 5/404 (1%)
Query: 389 NASFITLIPKVENPQGLGNYRPISLVGCVYKIVAKLLSMRLREVLSKVIDDRQTAFIGGR 448
N + I +IPK+ NP L +YRPI+L +YK+++K L RL+ L+ ++ D Q AFI GR
Sbjct: 867 NHTNICMIPKITNPTTLSDYRPIALCNVLYKVISKCLVNRLKSHLNSIVSDSQAAFIPGR 926
Query: 449 FMLDSVVVANEVVEEAKRRK---KECFVFKVDFEKAYDSVDWNFLFYMMSRIGFSHKWIH 505
+ D+V++A+EV+ K RK K K D KAYD V+W+FL M GF +KWI
Sbjct: 927 IINDNVMIAHEVMHSLKVRKRVSKTYMAVKTDVSKAYDRVEWDFLETTMRLFGFCNKWIG 986
Query: 506 WIKGCIQSASSSVLVNGSPTEEFNMGRGLRQGDPLAPFLFLIVAEGLNGLMRQAVQQKLF 565
WI ++S SVL+NGSP RG+RQGDPL+P+LF++ + L+ L+
Sbjct: 987 WIMAAVKSVHYSVLINGSPHGYITPTRGIRQGDPLSPYLFILCGDILSHLINGRASSGDL 1046
Query: 566 TGYRVGGDEVEISLLQFADDTLFFGEASTQNILVVKSILRWFEAMSGLKVNFFKSKLAGI 625
G R+G I+ LQFADD+LFF +A+ +N +K + +E SG K+N KS +
Sbjct: 1047 RGVRIGNGAPAITHLQFADDSLFFCQANVRNCQALKDVFDVYEYYSGQKINVQKSMITFG 1106
Query: 626 SMVSRETQI-FAAMLNCKVMGVPFTYLGIPVGANPRKAATWEPVIKKMQRKLSL*KHKSL 684
S V TQ +L G YLG+P +K +E +I +++++ S + L
Sbjct: 1107 SRVYGSTQSRLKQILEIPNQGGGGKYLGLPEQFGRKKKEMFEYIIDRVKKRTSTWSARFL 1166
Query: 685 SFGGRICLIKSVLSSLPLFFFSFFKAPKCIIDQCNKIQRKFLW-GGEEGRGVAWVKWEEI 743
S G+ ++KSV ++P++ S FK PK I+ + + F W RG+ WV W+ +
Sbjct: 1167 SPAGKEIMLKSVALAMPVYAMSCFKLPKGIVSEIESLLMNFWWEKASNQRGIPWVAWKRL 1226
Query: 744 CKKKEEGGLGVKDLESFNKALLVKWRWRLLREREWLWCQVLESK 787
K+EGGLG +DL FN ALL K WRL++ L+ +V++++
Sbjct: 1227 QYSKKEGGLGFRDLAKFNDALLAKQAWRLIQYPNSLFARVMKAR 1270
>UniRef100_Q9SIQ5 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 1524
Score = 271 bits (694), Expect = 4e-71
Identities = 151/404 (37%), Positives = 229/404 (56%), Gaps = 5/404 (1%)
Query: 389 NASFITLIPKVENPQGLGNYRPISLVGCVYKIVAKLLSMRLREVLSKVIDDRQTAFIGGR 448
N + I +IPK+ NP L +YRPI+L +YK+++K L RL+ L+ ++ D Q AFI GR
Sbjct: 641 NHTNICMIPKITNPTTLSDYRPIALCNVLYKVISKCLVNRLKSHLNSIVSDSQAAFIPGR 700
Query: 449 FMLDSVVVANEVVEEAKRRK---KECFVFKVDFEKAYDSVDWNFLFYMMSRIGFSHKWIH 505
+ D+V++A+EV+ K RK K K D KAYD V+W+FL M GF +KWI
Sbjct: 701 IINDNVMIAHEVMHSLKVRKRVSKTYMAVKTDVSKAYDRVEWDFLETTMRLFGFCNKWIG 760
Query: 506 WIKGCIQSASSSVLVNGSPTEEFNMGRGLRQGDPLAPFLFLIVAEGLNGLMRQAVQQKLF 565
WI ++S SVL+NGSP RG+RQGDPL+P+LF++ + L+ L+
Sbjct: 761 WIMAAVKSVHYSVLINGSPHGYITPTRGIRQGDPLSPYLFILCGDILSHLINGRASSGDL 820
Query: 566 TGYRVGGDEVEISLLQFADDTLFFGEASTQNILVVKSILRWFEAMSGLKVNFFKSKLAGI 625
G R+G I+ LQFADD+LFF +A+ +N +K + +E SG K+N KS +
Sbjct: 821 RGVRIGNGAPAITHLQFADDSLFFCQANVRNCQALKDVFDVYEYYSGQKINVQKSMITFG 880
Query: 626 SMVSRETQI-FAAMLNCKVMGVPFTYLGIPVGANPRKAATWEPVIKKMQRKLSL*KHKSL 684
S V TQ +L G YLG+P +K +E +I +++++ S + L
Sbjct: 881 SRVYGSTQSKLKQILEIPNQGGGGKYLGLPEQFGRKKKEMFEYIIDRVKKRTSTWSARFL 940
Query: 685 SFGGRICLIKSVLSSLPLFFFSFFKAPKCIIDQCNKIQRKFLW-GGEEGRGVAWVKWEEI 743
S G+ ++KSV ++P++ S FK PK I+ + + F W RG+ WV W+ +
Sbjct: 941 SPAGKEIMLKSVALAMPVYAMSCFKLPKGIVSEIESLLMNFWWEKASNQRGIPWVAWKRL 1000
Query: 744 CKKKEEGGLGVKDLESFNKALLVKWRWRLLREREWLWCQVLESK 787
K+EGGLG +DL FN ALL K WRL++ L+ +V++++
Sbjct: 1001 QYSKKEGGLGFRDLAKFNDALLAKQAWRLIQYPNSLFARVMKAR 1044
>UniRef100_Q9SN65 Hypothetical protein F25I24.40 [Arabidopsis thaliana]
Length = 1294
Score = 271 bits (693), Expect = 6e-71
Identities = 149/404 (36%), Positives = 232/404 (56%), Gaps = 5/404 (1%)
Query: 389 NASFITLIPKVENPQGLGNYRPISLVGCVYKIVAKLLSMRLREVLSKVIDDRQTAFIGGR 448
N + I +IPK+ NP+ L +YRPI+L +YKI++K L RL+ L ++ D Q AFI GR
Sbjct: 847 NHTNICMIPKITNPETLSDYRPIALCNVLYKIISKCLVERLKGHLDAIVSDSQAAFIPGR 906
Query: 449 FMLDSVVVANEVVEEAKRRKK---ECFVFKVDFEKAYDSVDWNFLFYMMSRIGFSHKWIH 505
+ D+V++A+E++ K RK+ K D KAYD V+WNFL M GFS WI
Sbjct: 907 LVNDNVMIAHEMMHSLKTRKRVSQSYMAVKTDVSKAYDRVEWNFLETTMRLFGFSETWIK 966
Query: 506 WIKGCIQSASSSVLVNGSPTEEFNMGRGLRQGDPLAPFLFLIVAEGLNGLMRQAVQQKLF 565
WI G ++S + SVLVNG P RG+RQGDPL+P+LF++ A+ LN L++ V +
Sbjct: 967 WIMGAVKSVNYSVLVNGIPHGTIQPQRGIRQGDPLSPYLFILCADILNHLIKNRVAEGDI 1026
Query: 566 TGYRVGGDEVEISLLQFADDTLFFGEASTQNILVVKSILRWFEAMSGLKVNFFKSKLAGI 625
G R+G ++ LQFADD+LFF +++ +N +K + +E SG K+N KS +
Sbjct: 1027 RGIRIGNGVPGVTHLQFADDSLFFCQSNVRNCQALKDVFDVYEYYSGQKINMSKSMITFG 1086
Query: 626 SMVSRETQ-IFAAMLNCKVMGVPFTYLGIPVGANPRKAATWEPVIKKMQRKLSL*KHKSL 684
S V TQ +L + G YLG+P +K + +I++++++ S K L
Sbjct: 1087 SRVHGTTQNRLKNILGIQSHGGGGKYLGLPEQFGRKKRDMFNYIIERVKKRTSSWSAKYL 1146
Query: 685 SFGGRICLIKSVLSSLPLFFFSFFKAPKCIIDQCNKIQRKFLW-GGEEGRGVAWVKWEEI 743
S G+ ++KSV S+P++ S FK P I+ + + F W + R + W+ W+ +
Sbjct: 1147 SPAGKEIMLKSVAMSMPVYAMSCFKLPLNIVSEIEALLMNFWWEKNAKKREIPWIAWKRL 1206
Query: 744 CKKKEEGGLGVKDLESFNKALLVKWRWRLLREREWLWCQVLESK 787
K+EGGLG +DL FN ALL K WR++ L+ ++++++
Sbjct: 1207 QYSKKEGGLGFRDLAKFNDALLAKQVWRMINNPNSLFARIMKAR 1250
>UniRef100_O81630 F8M12.22 protein [Arabidopsis thaliana]
Length = 1662
Score = 271 bits (693), Expect = 6e-71
Identities = 149/404 (36%), Positives = 232/404 (56%), Gaps = 5/404 (1%)
Query: 389 NASFITLIPKVENPQGLGNYRPISLVGCVYKIVAKLLSMRLREVLSKVIDDRQTAFIGGR 448
N + I +IPK+ NP+ L +YRPI+L +YKI++K L RL+ L ++ D Q AFI GR
Sbjct: 867 NHTNICMIPKITNPETLSDYRPIALCNVLYKIISKCLVERLKGHLDAIVSDSQAAFIPGR 926
Query: 449 FMLDSVVVANEVVEEAKRRKK---ECFVFKVDFEKAYDSVDWNFLFYMMSRIGFSHKWIH 505
+ D+V++A+E++ K RK+ K D KAYD V+WNFL M GFS WI
Sbjct: 927 LVNDNVMIAHEMMHSLKTRKRVSQSYMAVKTDVSKAYDRVEWNFLETTMRLFGFSETWIK 986
Query: 506 WIKGCIQSASSSVLVNGSPTEEFNMGRGLRQGDPLAPFLFLIVAEGLNGLMRQAVQQKLF 565
WI G ++S + SVLVNG P RG+RQGDPL+P+LF++ A+ LN L++ V +
Sbjct: 987 WIMGAVKSVNYSVLVNGIPHGTIQPQRGIRQGDPLSPYLFILCADILNHLIKNRVAEGDI 1046
Query: 566 TGYRVGGDEVEISLLQFADDTLFFGEASTQNILVVKSILRWFEAMSGLKVNFFKSKLAGI 625
G R+G ++ LQFADD+LFF +++ +N +K + +E SG K+N KS +
Sbjct: 1047 RGIRIGNGVPGVTHLQFADDSLFFCQSNVRNCQALKDVFDVYEYYSGQKINMSKSMITFG 1106
Query: 626 SMVSRETQ-IFAAMLNCKVMGVPFTYLGIPVGANPRKAATWEPVIKKMQRKLSL*KHKSL 684
S V TQ +L + G YLG+P +K + +I++++++ S K L
Sbjct: 1107 SRVHGTTQNRLKNILGIQSHGGGGKYLGLPEQFGRKKRDMFNYIIERVKKRTSSWSAKYL 1166
Query: 685 SFGGRICLIKSVLSSLPLFFFSFFKAPKCIIDQCNKIQRKFLW-GGEEGRGVAWVKWEEI 743
S G+ ++KSV S+P++ S FK P I+ + + F W + R + W+ W+ +
Sbjct: 1167 SPAGKEIMLKSVAMSMPVYAMSCFKLPLNIVSEIEALLMNFWWEKNAKKREIPWIAWKRL 1226
Query: 744 CKKKEEGGLGVKDLESFNKALLVKWRWRLLREREWLWCQVLESK 787
K+EGGLG +DL FN ALL K WR++ L+ ++++++
Sbjct: 1227 QYSKKEGGLGFRDLAKFNDALLAKQVWRMINNPNSLFARIMKAR 1270
>UniRef100_Q8H992 Oryza sativa (japonica cultivar-group) retrotransposon Karma DNA,
complete sequence [Oryza sativa]
Length = 1197
Score = 263 bits (673), Expect = 1e-68
Identities = 141/398 (35%), Positives = 237/398 (59%), Gaps = 11/398 (2%)
Query: 389 NASFITLIPKVENPQGLGNYRPISLVGCVYKIVAKLLSMRLREVLSKVIDDRQTAFIGGR 448
N + I LIPK + YRPISL C KI++K+L+ RL++VL+++I QT F+ GR
Sbjct: 498 NRAHIVLIPKPGKENTVDGYRPISLQNCSVKILSKVLANRLQKVLTRMIHLDQTGFLKGR 557
Query: 449 FMLDSVVVANEVVEEAKRRKKECFVFKVDFEKAYDSVDWNFLFYMMSRIGFSHKWIHWIK 508
+ ++++ A E+++ RK + + K+DF KA+DSV W LF +++ GF + WI WIK
Sbjct: 558 CISENLIYATELIQACHARKCKTLIIKLDFAKAFDSVLWTSLFQILAVRGFPNNWISWIK 617
Query: 509 GCIQSASSSVLVNGSPTEEFNMGRGLRQGDPLAPFLFLIVAEGLNGLMRQAVQQKLFTGY 568
+Q++ S+VL+NG P + N +GLRQGDPL+P+LF++VA+ L L+ + Q + +
Sbjct: 618 SLLQTSKSAVLLNGIPGKWINCKKGLRQGDPLSPYLFILVADVLQRLLEKNFQIR----H 673
Query: 569 RVGGDEVEISLLQFADDTLFFGEASTQNILVVKSILRWFEAMSGLKVNFFKSKLAGISMV 628
+ D + +Q+ADDTL A ++L ++S L F +GL++NF KS + + +
Sbjct: 674 PIYQDR-PCATIQYADDTLVICRAEEDDVLALRSTLLQFSKATGLQINFAKSTMISLHID 732
Query: 629 SRETQIFAAMLNCKVMGVPFTYLGIPVGANPRKAATWEPVIKKMQRKLSL*KHKSLSFGG 688
+ + +L CK+ +P +YLG+P+ + +P++ K+ L+ + LS
Sbjct: 733 RSKESSLSELLQCKLESLPMSYLGLPLSLHKLTNNDLQPIVVKVDSFLTGWEASLLSQAE 792
Query: 689 RICLIKSVLSSLPLFFFSFFKAPKCIIDQCNKIQRKFLWGGEE--GRGVAWVKWEEICKK 746
R+ L+ +VLSS+P++ S FK P +I+ +K +R F W G++ V W+E+C
Sbjct: 793 RLILVNAVLSSVPVYAMSAFKLPPKVIEAIDKRRRAFFWTGDDTCSGAKCLVAWDEVCTA 852
Query: 747 KEEGGLGVKDLESFNKALLVKWRWRLLRER----EWLW 780
KE+GGLG+K L++ N+ALL+K + L + W+W
Sbjct: 853 KEKGGLGIKSLKTQNEALLLKRLFNLFSDNSSWTNWIW 890
>UniRef100_Q7XJS2 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 1344
Score = 262 bits (670), Expect = 3e-68
Identities = 144/404 (35%), Positives = 233/404 (57%), Gaps = 5/404 (1%)
Query: 389 NASFITLIPKVENPQGLGNYRPISLVGCVYKIVAKLLSMRLREVLSKVIDDRQTAFIGGR 448
N + I LIPK+ +PQ + + RPISL +YKI++K+L+ RL++ L ++ Q+AF+ R
Sbjct: 463 NHTHICLIPKITSPQRMSDLRPISLCSVLYKIISKILTQRLKKHLPAIVSTTQSAFVPQR 522
Query: 449 FMLDSVVVANEVVEEAK---RRKKECFVFKVDFEKAYDSVDWNFLFYMMSRIGFSHKWIH 505
+ D+++VA+E++ + R KE FK D KAYD V+W FL MM+ +GF++KWI
Sbjct: 523 LISDNILVAHEMIHSLRTNDRISKEHMAFKTDMSKAYDRVEWPFLETMMTALGFNNKWIS 582
Query: 506 WIKGCIQSASSSVLVNGSPTEEFNMGRGLRQGDPLAPFLFLIVAEGLNGLMRQAVQQKLF 565
WI C+ S S SVL+NG P RG+RQGDPL+P LF++ E L ++ +A Q
Sbjct: 583 WIMNCVTSVSYSVLINGQPYGHIIPTRGIRQGDPLSPALFVLCTEALIHILNKAEQAGKI 642
Query: 566 TGYRVGGDEVEISLLQFADDTLFFGEASTQNILVVKSILRWFEAMSGLKVNFFKSKLA-G 624
TG + +V ++ L FADDTL +A+ Q + L + +SG +N KS + G
Sbjct: 643 TGIQFQDKKVSVNHLLFADDTLLMCKATKQECEELMQCLSQYGQLSGQMINLNKSAITFG 702
Query: 625 ISMVSRETQIFAAMLNCKVMGVPFTYLGIPVGANPRKAATWEPVIKKMQRKLSL*KHKSL 684
++ + + + G YLG+P + K + + +K+Q +L+ K+L
Sbjct: 703 KNVDIQIKDWIKSRSGISLEGGTGKYLGLPECLSGSKRDLFGFIKEKLQSRLTGWYAKTL 762
Query: 685 SFGGRICLIKSVLSSLPLFFFSFFKAPKCIIDQCNKIQRKFLWGG-EEGRGVAWVKWEEI 743
S GG+ L+KS+ +LP++ S FK PK + + + F W ++ R + W+ W+ +
Sbjct: 763 SQGGKEVLLKSIALALPVYVMSCFKLPKNLCQKLTTVMMDFWWNSMQQKRKIHWLSWQRL 822
Query: 744 CKKKEEGGLGVKDLESFNKALLVKWRWRLLREREWLWCQVLESK 787
K++GG G KDL+ FN+ALL K WR+L+E+ L+ +V +S+
Sbjct: 823 TLPKDQGGFGFKDLQCFNQALLAKQAWRVLQEKGSLFSRVFQSR 866
>UniRef100_Q75M12 Hypothetical protein P0676G05.2 [Oryza sativa]
Length = 1936
Score = 261 bits (668), Expect = 4e-68
Identities = 150/407 (36%), Positives = 224/407 (54%), Gaps = 5/407 (1%)
Query: 386 RGGNASFITLIPKVENPQGLGNYRPISLVGCVYKIVAKLLSMRLREVLSKVIDDRQTAFI 445
+G N + I LIPK + P L +YRPISL VYK+V+K L RLR +L ++ Q+AFI
Sbjct: 1122 KGVNDTAIVLIPKKDQPIDLKDYRPISLCNVVYKVVSKCLVNRLRPILDDLVSKEQSAFI 1181
Query: 446 GGRFMLDSVVVANEVVEEAKRRKKE---CFVFKVDFEKAYDSVDWNFLFYMMSRIGFSHK 502
GR + D+ ++A E ++ KK +K+D KAYD VDW FL ++++GF+H+
Sbjct: 1182 QGRMITDNALLAFECFHSIQKNKKANSAACAYKLDLSKAYDRVDWRFLELALNKLGFAHR 1241
Query: 503 WIHWIKGCIQSASSSVLVNGSPTEEFNMGRGLRQGDPLAPFLFLIVAEGLNGLMRQAVQQ 562
W+ WI C+ + SV NG+ F RGLRQG+PL+PFLFL VA+GL+ L+++ V Q
Sbjct: 1242 WVSWIMLCVTTVRYSVKFNGTLLRSFAPTRGLRQGEPLSPFLFLFVADGLSLLLKEKVAQ 1301
Query: 563 KLFTGYRVGGDEVEISLLQFADDTLFFGEASTQNILVVKSILRWFEAMSGLKVNFFK-SK 621
T ++ IS L FADDTL F +A + VVK +L + +G +N K S
Sbjct: 1302 NSLTPLKICRQAPGISYLLFADDTLLFFKAEKKEAEVVKEVLTNYAQGTGQLINPAKCSI 1361
Query: 622 LAGISMVSRETQIFAAMLNCKVMGVPFTYLGIPVGANPRKAATWEPVIKKMQRKLSL*KH 681
L G + S ++ L + YLG P ++ + K+ +++
Sbjct: 1362 LFGEASPSSVSEDIRNTLQVERDNFEDRYLGFPTPEGRMHKGRFQSLQAKIAKRVIQWGE 1421
Query: 682 KSLSFGGRICLIKSVLSSLPLFFFSFFKAPKCIIDQCNKIQRKFLWGGEEG-RGVAWVKW 740
LS GG+ LIK+V+ ++P++ FK P + D+ K+ R F WG + G R W W
Sbjct: 1422 NFLSSGGKEILIKAVIQAIPVYVMGLFKFPDSVYDELTKMTRNFWWGADNGRRRTHWRAW 1481
Query: 741 EEICKKKEEGGLGVKDLESFNKALLVKWRWRLLREREWLWCQVLESK 787
+ + K K GGLG +D + FN+ALL + WRL+ L QVL++K
Sbjct: 1482 DSLTKAKINGGLGFRDYKLFNQALLTRQAWRLIEFPNSLCAQVLKAK 1528
>UniRef100_Q75GM6 Putative non-LTR retroelement reverse transcriptase [Oryza sativa]
Length = 1614
Score = 261 bits (668), Expect = 4e-68
Identities = 142/400 (35%), Positives = 225/400 (55%), Gaps = 2/400 (0%)
Query: 389 NASFITLIPKVENPQGLGNYRPISLVGCVYKIVAKLLSMRLREVLSKVIDDRQTAFIGGR 448
N I+LIPK NP + +RPI ++ +K ++K + RL E+ VI QTAFI GR
Sbjct: 1115 NFGVISLIPKNSNPTDIKQFRPICVLNDCFKFISKCVCNRLTEIARDVISPTQTAFIPGR 1174
Query: 449 FMLDSVVVANEVVEEAKRRKKECFVFKVDFEKAYDSVDWNFLFYMMSRIGFSHKWIHWIK 508
F+L+ V+ +EV+ E R+ E + K+DFEKAYD V W+FL +M R GF KW++WIK
Sbjct: 1175 FILEGCVIIHEVLHEMNRKNLEGIILKIDFEKAYDKVSWDFLIEVMVRKGFPSKWVNWIK 1234
Query: 509 GCIQSASSSVLVNGSPTEEFNMGRGLRQGDPLAPFLFLIVAEGLNGLMRQAVQQKLFTGY 568
C+ + +NG T+ F RGLRQGDPL+P LF ++++ L ++ A ++ + +G
Sbjct: 1235 TCVMGGRVCININGERTDFFRTFRGLRQGDPLSPLLFNLISDALAAMLDSAKREGVLSGL 1294
Query: 569 RVGGDEVEISLLQFADDTLFFGEASTQNILVVKSILRWFEAMSGLKVNFFKSKLAGISMV 628
I+ LQ+ADDT+ F + I+ K IL FE M+ LKVN+ KS++ + +
Sbjct: 1295 VPDIFPGGITHLQYADDTVLFVANDDKQIVATKFILYCFEEMAVLKVNYHKSEIFTLGLS 1354
Query: 629 SRETQIFAAMLNCKVMGVPFTYLGIPVGANPRKAATWEPVIKKMQRKLSL*KHKSLSFGG 688
+T A M NC V P YLG+P+G + ++ + +K++++L+ +LS G
Sbjct: 1355 DNDTNRVAMMFNCPVGQFPMKYLGLPIGPDKILNLGFDFLGQKLEKRLNS-WGNNLSHAG 1413
Query: 689 RICLIKSVLSSLPLFFFSFFKAPKCIIDQCNKIQRKFLWGGEEGRG-VAWVKWEEICKKK 747
R I + LSS+P + F++ P+ + + I+ ++ W +G VKWE++ K
Sbjct: 1414 RAVQINTCLSSIPSYAMCFYQLPEGVHQKFGSIRGRYYWARNRLKGKYHMVKWEDLAFPK 1473
Query: 748 EEGGLGVKDLESFNKALLVKWRWRLLREREWLWCQVLESK 787
+ GGLG + N ALL KW ++ E + L ++L K
Sbjct: 1474 DYGGLGFTETRRMNTALLAKWIMKIESEDDSLCIELLRRK 1513
>UniRef100_Q7X970 BZIP-like protein [Oryza sativa]
Length = 2367
Score = 257 bits (657), Expect = 8e-67
Identities = 148/406 (36%), Positives = 223/406 (54%), Gaps = 5/406 (1%)
Query: 387 GGNASFITLIPKVENPQGLGNYRPISLVGCVYKIVAKLLSMRLREVLSKVIDDRQTAFIG 446
G N + I LIPK E P L ++RPISL VYK+V+K L RLR +L ++ Q+AF+
Sbjct: 1217 GVNDTAIVLIPKKEQPVDLRDFRPISLCNVVYKVVSKCLVNRLRPILDDLVSVEQSAFVQ 1276
Query: 447 GRFMLDSVVVANEVVEEAKRRKKE---CFVFKVDFEKAYDSVDWNFLFYMMSRIGFSHKW 503
GR + D+ ++A E ++ KK +K+D KAYD VDW FL M+++GF+ +W
Sbjct: 1277 GRMITDNALLAFECFHAMQKNKKANHAACAYKLDLSKAYDRVDWRFLEMAMNKLGFARRW 1336
Query: 504 IHWIKGCIQSASSSVLVNGSPTEEFNMGRGLRQGDPLAPFLFLIVAEGLNGLMRQAVQQK 563
++WI C+ S V NG+ + F RGLRQGDPL PFLFL VA+GL+ L+++ V Q
Sbjct: 1337 VNWIMKCVTSVRYMVKFNGTLLQSFAPTRGLRQGDPLLPFLFLFVADGLSLLLKEKVAQN 1396
Query: 564 LFTGYRVGGDEVEISLLQFADDTLFFGEASTQNILVVKSILRWFEAMSGLKVNFFK-SKL 622
T ++V IS L FADDTL F +A + VVK +L + +G +N K S L
Sbjct: 1397 SLTPFKVCRAAPGISHLLFADDTLLFFKAHQREAEVVKEVLSSYAMGTGQLINPAKCSIL 1456
Query: 623 AGISMVSRETQIFAAMLNCKVMGVPFTYLGIPVGANPRKAATWEPVIKKMQRKLSL*KHK 682
G + ++ + +L + YLG P ++ + K+ +++
Sbjct: 1457 MGGASTPAVSEAISEILQVERDRFEDRYLGFPTPEGRMHKGRFQSLQAKIWKRVIQWGEN 1516
Query: 683 SLSFGGRICLIKSVLSSLPLFFFSFFKAPKCIIDQCNKIQRKFLWGGEEG-RGVAWVKWE 741
LS GG+ LIK+V+ ++P++ FK P+ +ID K+ + F W G R W W+
Sbjct: 1517 HLSTGGKEVLIKAVIQAIPVYVMGIFKLPESVIDDLTKLTKNFWWDSMNGQRKTHWKAWD 1576
Query: 742 EICKKKEEGGLGVKDLESFNKALLVKWRWRLLREREWLWCQVLESK 787
+ K K GGLG +D FN+ALL + WRL+ + L +VL++K
Sbjct: 1577 SLTKPKSLGGLGFRDYRLFNQALLARQAWRLITYPDSLCARVLKAK 1622
>UniRef100_O22148 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 1374
Score = 257 bits (657), Expect = 8e-67
Identities = 144/406 (35%), Positives = 229/406 (55%), Gaps = 5/406 (1%)
Query: 387 GGNASFITLIPKVENPQGLGNYRPISLVGCVYKIVAKLLSMRLREVLSKVIDDRQTAFIG 446
G N + I LIPK+ + + ++RPISL +YK++ KL++ RL+++L +I + Q AF+
Sbjct: 483 GMNKTNICLIPKILKAEKMTDFRPISLCNVIYKVIGKLMANRLKKILPSLISETQAAFVK 542
Query: 447 GRFMLDSVVVANEVVEEAKRRKK---ECFVFKVDFEKAYDSVDWNFLFYMMSRIGFSHKW 503
GR + D++++A+E++ K E K D KAYD V+W FL M +GF+ W
Sbjct: 543 GRLISDNILIAHELLHALSSNNKCSEEFIAIKTDISKAYDRVEWPFLEKAMRGLGFADHW 602
Query: 504 IHWIKGCIQSASSSVLVNGSPTEEFNMGRGLRQGDPLAPFLFLIVAEGLNGLMRQAVQQK 563
I I C++S VL+NG+P E RGLRQGDPL+P+LF+I E L +++ A Q+
Sbjct: 603 IRLIMECVKSVRYQVLINGTPHGEIIPSRGLRQGDPLSPYLFVICTEMLVKMLQSAEQKN 662
Query: 564 LFTGYRVGGDEVEISLLQFADDTLFFGEASTQNILVVKSILRWFEAMSGLKVNFFKSKLA 623
TG +V IS L FADD++F+ + + + + + I+ + SG +VN+ KS +
Sbjct: 663 QITGLKVARGAPPISHLLFADDSMFYCKVNDEALGQIIRIIEEYSLASGQRVNYLKSSIY 722
Query: 624 GISMVSRETQ-IFAAMLNCKVMGVPFTYLGIPVGANPRKAATWEPVIKKMQRKLSL*KHK 682
+S E + + L + G YLG+P K AT + ++ +K+ +
Sbjct: 723 FGKHISEERRCLVKRKLGIEREGGEGVYLGLPESFQGSKVATLSYLKDRLGKKVLGWQSN 782
Query: 683 SLSFGGRICLIKSVLSSLPLFFFSFFKAPKCIIDQCNKIQRKFLW-GGEEGRGVAWVKWE 741
LS GG+ L+K+V +LP + S FK PK I Q + +F W +EGRG+ W W
Sbjct: 783 FLSPGGKEILLKAVAMALPTYTMSCFKIPKTICQQIESVMAEFWWKNKKEGRGLHWKAWC 842
Query: 742 EICKKKEEGGLGVKDLESFNKALLVKWRWRLLREREWLWCQVLESK 787
+ + K GGLG K++E+FN ALL K WR++ E++ L +V +S+
Sbjct: 843 HLSRPKAVGGLGFKEIEAFNIALLGKQLWRMITEKDSLMAKVFKSR 888
>UniRef100_Q7XGD3 Putative retroelement [Oryza sativa]
Length = 1694
Score = 257 bits (656), Expect = 1e-66
Identities = 145/406 (35%), Positives = 223/406 (54%), Gaps = 8/406 (1%)
Query: 387 GGNASFITLIPKVENPQGLGNYRPISLVGCVYKIVAKLLSMRLREVLSKVIDDRQTAFIG 446
G N + I LIPK E PQ L ++RPISL VYKIV+K L RLR +L ++ Q+AF+
Sbjct: 1097 GVNETAIVLIPKTEQPQELKDFRPISLCNVVYKIVSKCLVNRLRPILDDLVSQNQSAFVP 1156
Query: 447 GRFMLDSVVVANEVVEEAKRRKKE---CFVFKVDFEKAYDSVDWNFLFYMMSRIGFSHKW 503
GR + D+ ++A E +R K +K+D KAYD VDW FL M ++GF+H W
Sbjct: 1157 GRLITDNALIAFEYFHHIQRNKNPENAYSAYKLDLSKAYDRVDWEFLEQAMVKLGFAHCW 1216
Query: 504 IHWIKGCIQSASSSVLVNGSPTEEFNMGRGLRQGDPLAPFLFLIVAEGLNGLMRQAVQQK 563
+ WI CI +V +NG+ F RGLRQGDPL+PFLFL VA+GL+ L+ + V Q
Sbjct: 1217 VKWIMACITLVRYAVKLNGTLLNTFAPSRGLRQGDPLSPFLFLFVADGLSFLLEEKVDQG 1276
Query: 564 LFTGYRVGGDEVEISLLQFADDTLFFGEASTQNILVVKSILRWFEAMSGLKVNFFKSKLA 623
+ + L FADDT F +A+ +V+K +L + + +G +N SK +
Sbjct: 1277 AMNPIHICRHAPAVLHLLFADDTPLFFKAANDQAVVMKEVLETYASCTGQLIN--PSKCS 1334
Query: 624 GISMVSRETQIFAAMLNCKVMGVPF--TYLGIPVGANPRKAATWEPVIKKMQRKLSL*KH 681
+ S T + + + F YLG P ++ + +++ +++ L
Sbjct: 1335 IMFGQSSPTAVQNQIKQTLQISNSFEDKYLGFPTPEGRMTKGKFQSLQERLWKRIMLWGE 1394
Query: 682 KSLSFGGRICLIKSVLSSLPLFFFSFFKAPKCIIDQCNKIQRKFLWGGEEG-RGVAWVKW 740
K LS GG+ LIK+VL ++P++ FK P+ + ++ K+ R F WG E+G R W W
Sbjct: 1395 KFLSTGGKEILIKAVLQAMPVYVMGLFKLPESVCEELTKLVRNFWWGAEKGMRRTHWKSW 1454
Query: 741 EEICKKKEEGGLGVKDLESFNKALLVKWRWRLLREREWLWCQVLES 786
+ + +K +GG+G +D FN+ALL + WRLL + L ++L++
Sbjct: 1455 DCLIAQKLKGGMGFRDFRLFNQALLARQAWRLLDRPDSLCARILKA 1500
>UniRef100_Q8S5K1 Putative retroelement [Oryza sativa]
Length = 1888
Score = 257 bits (656), Expect = 1e-66
Identities = 145/406 (35%), Positives = 223/406 (54%), Gaps = 8/406 (1%)
Query: 387 GGNASFITLIPKVENPQGLGNYRPISLVGCVYKIVAKLLSMRLREVLSKVIDDRQTAFIG 446
G N + I LIPK E PQ L ++RPISL VYKIV+K L RLR +L ++ Q+AF+
Sbjct: 1054 GVNETAIVLIPKTEQPQELKDFRPISLCNVVYKIVSKCLVNRLRPILDDLVSQNQSAFVP 1113
Query: 447 GRFMLDSVVVANEVVEEAKRRKKE---CFVFKVDFEKAYDSVDWNFLFYMMSRIGFSHKW 503
GR + D+ ++A E +R K +K+D KAYD VDW FL M ++GF+H W
Sbjct: 1114 GRLITDNALIAFEYFHHIQRNKNPENAYSAYKLDLSKAYDRVDWEFLEQAMVKLGFAHCW 1173
Query: 504 IHWIKGCIQSASSSVLVNGSPTEEFNMGRGLRQGDPLAPFLFLIVAEGLNGLMRQAVQQK 563
+ WI CI +V +NG+ F RGLRQGDPL+PFLFL VA+GL+ L+ + V Q
Sbjct: 1174 VKWIMACITLVRYAVKLNGTLLNTFAPSRGLRQGDPLSPFLFLFVADGLSFLLEEKVDQG 1233
Query: 564 LFTGYRVGGDEVEISLLQFADDTLFFGEASTQNILVVKSILRWFEAMSGLKVNFFKSKLA 623
+ + L FADDT F +A+ +V+K +L + + +G +N SK +
Sbjct: 1234 AMNPIHICRHAPAVLHLLFADDTPLFFKAANDQAVVMKEVLETYASCTGQLIN--PSKCS 1291
Query: 624 GISMVSRETQIFAAMLNCKVMGVPF--TYLGIPVGANPRKAATWEPVIKKMQRKLSL*KH 681
+ S T + + + F YLG P ++ + +++ +++ L
Sbjct: 1292 IMFGQSSPTAVQNQIKQTLQISNSFEDKYLGFPTPEGRMTKGKFQSLQERLWKRIMLWGE 1351
Query: 682 KSLSFGGRICLIKSVLSSLPLFFFSFFKAPKCIIDQCNKIQRKFLWGGEEG-RGVAWVKW 740
K LS GG+ LIK+VL ++P++ FK P+ + ++ K+ R F WG E+G R W W
Sbjct: 1352 KFLSTGGKEILIKAVLQAMPVYVMGLFKLPESVCEELTKLVRNFWWGAEKGMRRTHWKSW 1411
Query: 741 EEICKKKEEGGLGVKDLESFNKALLVKWRWRLLREREWLWCQVLES 786
+ + +K +GG+G +D FN+ALL + WRLL + L ++L++
Sbjct: 1412 DCLIAQKLKGGMGFRDFRLFNQALLARQAWRLLDRPDSLCARILKA 1457
>UniRef100_O82276 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 1231
Score = 257 bits (656), Expect = 1e-66
Identities = 150/407 (36%), Positives = 227/407 (54%), Gaps = 12/407 (2%)
Query: 389 NASFITLIPKVENPQGLGNYRPISLVGCVYKIVAKLLSMRLREVLSKVIDDRQTAFIGGR 448
N + + LI KV P+ + +RP+SL ++KI+ K++ RL+ V+SK+I Q +FI GR
Sbjct: 349 NDALLVLIAKVAKPERIQQFRPVSLCNVLFKIITKMMVTRLKNVISKLIGPAQASFIPGR 408
Query: 449 FMLDSVVVANEVVEEAKRRK--KECFVFKVDFEKAYDSVDWNFLFYMMSRIGFSHKWIHW 506
+D++V+ E V +R+K K + K+D EKAYD V W+FL + G S W
Sbjct: 409 LSIDNIVLVQEAVHSMRRKKGRKGWMLLKLDLEKAYDRVRWDFLQETLEAAGLSEGWTSR 468
Query: 507 IKGCIQSASSSVLVNGSPTEEFNMGRGLRQGDPLAPFLFLIVAEGLNGLMRQAVQQKLFT 566
I + S SVL NG T+ F RGLRQGDPL+P+LF++ E L L+ +V ++ +
Sbjct: 469 IMAGVTDPSMSVLWNGERTDSFVPARGLRQGDPLSPYLFVLCLERLCHLIEASVGKREWK 528
Query: 567 GYRVGGDEVEISLLQFADDTLFFGEASTQNILVVKSILRWFEAMSGLKVNFFKSKLAGIS 626
V ++S + FADD + F EAS I +++ +L F SG KV+ KSK+
Sbjct: 529 PIAVSCGGSKLSHVCFADDLILFAEASVAQIRIIRRVLERFCEASGQKVSLEKSKIFFSH 588
Query: 627 MVSRETQIFAAMLNCKVMGVPFT-----YLGIPVGANPRKAATWEPVIKKMQRKLSL*KH 681
VSRE + L + G+ T YLG+P+ T+ V++++ +L+ K
Sbjct: 589 NVSREME----QLISEESGIGCTKELGKYLGMPILQKRMNKETFGEVLERVSARLAGWKG 644
Query: 682 KSLSFGGRICLIKSVLSSLPLFFFSFFKAPKCIIDQCNKIQRKFLWGGE-EGRGVAWVKW 740
+SLS GRI L K+VLSS+P+ S P +D ++ R FLWG E + + W
Sbjct: 645 RSLSLAGRITLTKAVLSSIPVHVMSAILLPVSTLDTLDRYSRTFLWGSTMEKKKQHLLSW 704
Query: 741 EEICKKKEEGGLGVKDLESFNKALLVKWRWRLLREREWLWCQVLESK 787
+ICK K EGG+G++ NKAL+ K WRLL+++E LW +V+ K
Sbjct: 705 RKICKPKAEGGIGLRSARDMNKALVAKVGWRLLQDKESLWARVVRKK 751
>UniRef100_Q9SIS9 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 1715
Score = 256 bits (654), Expect = 2e-66
Identities = 148/390 (37%), Positives = 221/390 (55%), Gaps = 23/390 (5%)
Query: 389 NASFITLIPKVENPQGLGNYRPISLVGCVYKIVAKLLSMRLREVLSKVIDDRQTAFIGGR 448
N + I LIPK+ +P+ + +YRPISL YKI++K+L RL++ L VI D Q AF+ G+
Sbjct: 847 NQTQICLIPKIIDPKHMSDYRPISLCTASYKIISKILIKRLKQCLGDVISDSQAAFVPGQ 906
Query: 449 FMLDSVVVANEVVEEAKRRKKEC----FVFKVDFEKAYDSVDWNFLFYMMSRIGFSHKWI 504
+ D+V+VA+E++ K R+ EC K D KAYD V+WNFL +M ++GF+ +W+
Sbjct: 907 NISDNVLVAHELLHSLKSRR-ECQSGYVAVKTDISKAYDRVEWNFLEKVMIQLGFAPRWV 965
Query: 505 HWIKGCIQSASSSVLVNGSPTEEFNMGRGLRQGDPLAPFLFLIVAEGLNGLMRQAVQQKL 564
WI C+ S S VL+NGSP + RG+RQGDPL+P+LFL AE L+ ++R+A K
Sbjct: 966 KWIMTCVTSVSYEVLINGSPYGKIFPSRGIRQGDPLSPYLFLFCAEVLSNMLRKAEVNKQ 1025
Query: 565 FTGYRVGGDEVEISLLQFADDTLFFGEASTQNILVVKSILRWFEAMSGLKVNFFKSKLA- 623
G ++ D + IS L FADD+LFF AS QNI + I + +E SG K+N+ KS +
Sbjct: 1026 IHGMKITKDCLAISHLLFADDSLFFCRASNQNIEQLALIFKKYEEASGQKINYAKSSIIF 1085
Query: 624 GISMVSRETQIFAAMLNCKVMGVPFTYLGIPVGANPRKAATWEPVIKKMQRKLSL*KHKS 683
G + + Q +L + YLG+P RK +E ++ K++ + +
Sbjct: 1086 GQKIPTMRRQRLHRLLGIDNVRGGGKYLGLPEQLGRRKVELFEYIVTKVKERTEGWAYNY 1145
Query: 684 LSFGGRICLIKSVLSSLPLFFFSFFKAPKCIIDQCNKIQRKFLWGGEEGRGVAWVKWEEI 743
LS G+ +IK++ +LP++ + F P I ++ N + F WG
Sbjct: 1146 LSPAGKEIVIKAIAMALPVYSMNCFLLPTLICNEINSLITAFWWG--------------- 1190
Query: 744 CKKKEEGGLGVKDLESFNKALLVKWRWRLL 773
K+ EG LG KDL FN+ALL K WR+L
Sbjct: 1191 --KENEGDLGFKDLHQFNRALLAKQAWRIL 1218
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.341 0.150 0.502
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,202,124,475
Number of Sequences: 2790947
Number of extensions: 46643181
Number of successful extensions: 131529
Number of sequences better than 10.0: 1102
Number of HSP's better than 10.0 without gapping: 807
Number of HSP's successfully gapped in prelim test: 295
Number of HSP's that attempted gapping in prelim test: 129183
Number of HSP's gapped (non-prelim): 1392
length of query: 787
length of database: 848,049,833
effective HSP length: 135
effective length of query: 652
effective length of database: 471,271,988
effective search space: 307269336176
effective search space used: 307269336176
T: 11
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.9 bits)
S2: 79 (35.0 bits)
Lotus: description of TM0263.7


