

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0258b.1
(207 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
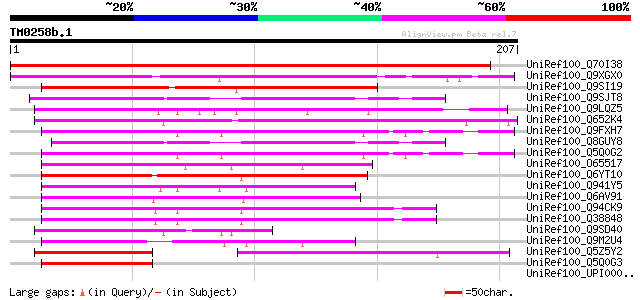
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q70I38 Hypothetical protein [Lotus japonicus] 413 e-114
UniRef100_Q9XGX0 Zinc finger protein SHI-like [Arabidopsis thali... 142 5e-33
UniRef100_Q9SI19 Hypothetical protein At2g18120 [Arabidopsis tha... 140 3e-32
UniRef100_Q9SJT8 Hypothetical protein At2g21400 [Arabidopsis tha... 136 4e-31
UniRef100_Q9LQZ5 F10A5.26 [Arabidopsis thaliana] 133 3e-30
UniRef100_Q652K4 Putative LRP1 [Oryza sativa] 126 3e-28
UniRef100_Q9FXH7 F6F9.16 protein [Arabidopsis thaliana] 125 9e-28
UniRef100_Q8GUY8 Hypothetical protein F3K23.16 [Arabidopsis thal... 123 3e-27
UniRef100_Q5Q0G2 Hypothetical protein [Arabidopsis thaliana] 123 3e-27
UniRef100_O65517 Hypothetical protein F23E13.150 [Arabidopsis th... 123 3e-27
UniRef100_Q6YT10 Putative lateral root primordia [Oryza sativa] 123 3e-27
UniRef100_Q941Y5 Putative LRP (Lateral root primordia) 1 [Oryza ... 115 9e-25
UniRef100_Q6AV91 Hypothetical protein OJ1286_E05.15 [Oryza sativa] 114 1e-24
UniRef100_Q94CK9 Lateral root primordia [Arabidopsis thaliana] 111 1e-23
UniRef100_Q38848 LRP1 [Arabidopsis thaliana] 110 2e-23
UniRef100_Q9SD40 Hypothetical protein F24M12.100 [Arabidopsis th... 92 1e-17
UniRef100_Q9M2U4 Hypothetical protein T12E18_120 [Arabidopsis th... 87 3e-16
UniRef100_Q5Z5Y2 Putative LRP1 [Oryza sativa] 80 4e-14
UniRef100_Q5Q0G3 Hypothetical protein [Arabidopsis thaliana] 78 1e-13
UniRef100_UPI000036462C UPI000036462C UniRef100 entry 41 0.022
>UniRef100_Q70I38 Hypothetical protein [Lotus japonicus]
Length = 246
Score = 413 bits (1062), Expect = e-114
Identities = 196/196 (100%), Positives = 196/196 (100%)
Query: 1 MSMMLGQEVKGSRCQECGNQAKKGCAYSRCRTCCNNKGFQCQTHVRSTWIPVDRRRHRLM 60
MSMMLGQEVKGSRCQECGNQAKKGCAYSRCRTCCNNKGFQCQTHVRSTWIPVDRRRHRLM
Sbjct: 1 MSMMLGQEVKGSRCQECGNQAKKGCAYSRCRTCCNNKGFQCQTHVRSTWIPVDRRRHRLM 60
Query: 61 EHQPPPTTNNPHHLHEDIPQSHNQNPFTSLEELKFPEAMSSMAVFSSVRVRSMDDSVNEM 120
EHQPPPTTNNPHHLHEDIPQSHNQNPFTSLEELKFPEAMSSMAVFSSVRVRSMDDSVNEM
Sbjct: 61 EHQPPPTTNNPHHLHEDIPQSHNQNPFTSLEELKFPEAMSSMAVFSSVRVRSMDDSVNEM 120
Query: 121 AYQTSVNIGGHRFSGILYDQGPEQQSLNASPLDQHQNLNLTTIHSHDGATMAPPSSSATA 180
AYQTSVNIGGHRFSGILYDQGPEQQSLNASPLDQHQNLNLTTIHSHDGATMAPPSSSATA
Sbjct: 121 AYQTSVNIGGHRFSGILYDQGPEQQSLNASPLDQHQNLNLTTIHSHDGATMAPPSSSATA 180
Query: 181 AHELFFPQPRSLASFR 196
AHELFFPQPRSLASFR
Sbjct: 181 AHELFFPQPRSLASFR 196
>UniRef100_Q9XGX0 Zinc finger protein SHI-like [Arabidopsis thaliana]
Length = 331
Score = 142 bits (358), Expect = 5e-33
Identities = 87/234 (37%), Positives = 123/234 (52%), Gaps = 38/234 (16%)
Query: 1 MSMMLGQEVKGSRCQECGNQAKKGCAYSRCRTCCNNKGFQCQTHVRSTWIPVDRRRHRLM 60
M M G G CQ+CGNQ+KK C++ RCRTCC ++G C THV+STW+P +RR R
Sbjct: 107 MMMRSGSGSGGPSCQDCGNQSKKDCSHMRCRTCCKSRGLDCPTHVKSTWVPAAKRRER-- 164
Query: 61 EHQPPPTTNNPHHLHEDIPQSHNQ----------------NPFTSLEELKFPEAMSSMAV 104
Q T P L +P+ + N + LE FP +SS AV
Sbjct: 165 -QQQLSTGQQPQQLGGSVPKRQRERIPARSTSMAYTRIPSNNTSGLEVGNFPPEVSSSAV 223
Query: 105 FSSVRVRSMDDSVNEMAYQTSVNIGGHRFSGILYDQGPEQQSLNASPLDQHQNLNLTTIH 164
F VRV S+DD E AY+T+V+IGGH F G+LYDQGP ++S + Q LNL T
Sbjct: 224 FRCVRVSSVDDEEEEYAYKTAVSIGGHVFKGVLYDQGPAERSSSGG---GSQPLNLIT-- 278
Query: 165 SHDGATMAPPSSS-------ATAAH-----ELFFPQPRSLASFRSGVPYFSHTK 206
+ A+ + P+ S +T+ H L +P P + +F +G +FS+++
Sbjct: 279 AGPSASSSSPNVSCNNGVVGSTSDHYIDPASLNYPTP--INTFMTGTHFFSNSR 330
>UniRef100_Q9SI19 Hypothetical protein At2g18120 [Arabidopsis thaliana]
Length = 222
Score = 140 bits (352), Expect = 3e-32
Identities = 68/144 (47%), Positives = 91/144 (62%), Gaps = 9/144 (6%)
Query: 14 CQECGNQAKKGCAYSRCRTCCNNKGFQCQTHVRSTWIPVDRRRHRLMEHQPPPTTNNPHH 73
CQECGNQAKKGC + RCRTCC + G C THVRSTWIP+ +RR R + Q P T+NP
Sbjct: 72 CQECGNQAKKGCTHGRCRTCCKSNGLHCPTHVRSTWIPIAKRRERQQQLQTP--TSNPTG 129
Query: 74 LHEDIPQSHNQNPFTSLE-------ELKFPEAMSSMAVFSSVRVRSMDDSVNEMAYQTSV 126
+ + + N +L+ E +FP+ +SS A+F VR+ DD + AYQT+V
Sbjct: 130 GSGRVGKYRDINQHATLDSSGLEMGETRFPDEVSSDALFRCVRMSGTDDGEGQYAYQTTV 189
Query: 127 NIGGHRFSGILYDQGPEQQSLNAS 150
I GH F GILY+QGPE +S+ ++
Sbjct: 190 GIAGHLFKGILYNQGPENKSMRST 213
>UniRef100_Q9SJT8 Hypothetical protein At2g21400 [Arabidopsis thaliana]
Length = 174
Score = 136 bits (342), Expect = 4e-31
Identities = 72/170 (42%), Positives = 98/170 (57%), Gaps = 23/170 (13%)
Query: 9 VKGSRCQECGNQAKKGCAYSRCRTCCNNKGFQCQTHVRSTWIPVDRRRHRLMEHQPPPTT 68
+ G +C++CGNQAKK C Y RCRTCC +K F CQTH++STW+P RR H + Q P +T
Sbjct: 4 IMGRKCEDCGNQAKKDCVYMRCRTCCKSKAFHCQTHIKSTWVPAYRRSHHKHQSQ-PLST 62
Query: 69 NNPHHLHEDIPQSHNQNPFTSLEELKFPEAMSSMAVFSSVRVRSMDDSVNEMAYQTSVNI 128
+ P + H FP +SS+A F V+V S+DD + AYQT+VNI
Sbjct: 63 SIPKGVQIHTTPGH------------FPAELSSLADFRCVKVSSIDDGKEQYAYQTTVNI 110
Query: 129 GGHRFSGILYDQGPEQQSLNASPLDQHQNLNLTTIHSHDGATMAPPSSSA 178
GGH F GIL+DQG L+ +D H N N S++ + PS+S+
Sbjct: 111 GGHVFRGILHDQG-----LHKVMVDHHYNKN-----SNNHQELLTPSTSS 150
>UniRef100_Q9LQZ5 F10A5.26 [Arabidopsis thaliana]
Length = 346
Score = 133 bits (334), Expect = 3e-30
Identities = 84/231 (36%), Positives = 116/231 (49%), Gaps = 48/231 (20%)
Query: 11 GSRCQECGNQAKKGCAYSRCRTCCNNKGFQCQTHVRSTWIPVDRRRHRL-----MEHQPP 65
G CQ+CGNQAKK C + RCRTCC ++GF CQTHV+STW+P +RR RL ++H
Sbjct: 122 GMNCQDCGNQAKKDCPHMRCRTCCKSRGFHCQTHVKSTWVPAAKRRERLAQLASLQHHSA 181
Query: 66 PT--TNNPHHLHE------DIPQSH--------------NQNPFTSLE-ELKFPEAMSSM 102
+ T N L E + + H N N + LE P ++S
Sbjct: 182 SSRETQNAKRLREASGGDNNDDKDHSGGGGSALANTRVVNANSNSGLEVSQHLPPEVNSP 241
Query: 103 AVFSSVRVRSMDDSVNEM--AYQTSVNIGGHRFSGILYDQGPEQQ--------SLNASPL 152
A+F VRV S+++ ++ AYQT+VNIGGH F GILYDQGPE Q + A+
Sbjct: 242 AIFRCVRVSSIEEDEDDQAYAYQTAVNIGGHIFKGILYDQGPEHQDNHHLNLLASTATTT 301
Query: 153 DQHQNLNLTTIHSHDGATMAPPSSSATAAHELFFPQPRSLASFRSGVPYFS 203
+ + T +++ M PSS P L SF +G P+F+
Sbjct: 302 NVEETATKTVTGNNNNGLMLDPSSL----------YPAQLNSFIAGTPFFT 342
>UniRef100_Q652K4 Putative LRP1 [Oryza sativa]
Length = 315
Score = 126 bits (317), Expect = 3e-28
Identities = 76/224 (33%), Positives = 105/224 (45%), Gaps = 29/224 (12%)
Query: 11 GSRCQECGNQAKKGCAYSRCRTCCNNKGFQCQTHVRSTWIPVDRRRHRLME--------- 61
G CQ+CGNQAKK C + RCRTCC ++GF C THV+STW+P +RR R +
Sbjct: 94 GISCQDCGNQAKKDCTHMRCRTCCKSRGFACATHVKSTWVPAAKRRERQQQLAALAASAA 153
Query: 62 ---------HQPPPTTNNPHHLHEDIPQSHNQNPFTSLEELKFPEAMSSMAVFSSVRVRS 112
P S +Q T E +FP +SS AVF VR+
Sbjct: 154 ATAGGAGPSRDPTKRPRARPSATTPTTSSGDQQMVTVAE--RFPREVSSEAVFRCVRLGP 211
Query: 113 MDDSVNEMAYQTSVNIGGHRFSGILYDQGPEQQSLNASPLDQHQNLNLTTIHSHDGATMA 172
+D + E+AYQT+V+IGGH F GIL+D GPE ++ + LT S A
Sbjct: 212 VDQAEAEVAYQTAVSIGGHVFKGILHDVGPEALAVAGGGGASEYHFRLTGDGSSPSTAAA 271
Query: 173 PPSSSATAAHELF--------FPQPRSLASFRSGVPYF-SHTKP 207
+ S + + +P P +F +G P+F H +P
Sbjct: 272 GEAGSGGGGNIIVSSAVVMDPYPTPGPYGAFPAGTPFFHGHPRP 315
>UniRef100_Q9FXH7 F6F9.16 protein [Arabidopsis thaliana]
Length = 345
Score = 125 bits (313), Expect = 9e-28
Identities = 84/235 (35%), Positives = 113/235 (47%), Gaps = 51/235 (21%)
Query: 14 CQECGNQAKKGCAYSRCRTCCNNKGFQCQTHVRSTWIPVDRRRHRLMEHQPPPT-----T 68
CQ+CGNQAKK C + RCRTCC ++GF CQTHV+STW+ +RR R + P
Sbjct: 119 CQDCGNQAKKDCPHMRCRTCCKSRGFDCQTHVKSTWVSAAKRRERQAQLAVLPAKRIRDA 178
Query: 69 NNPHHLHEDIPQSHNQN------------------PFTSLEELKFPEAMSSMAVFSSVRV 110
N+ +D ++ + LE P +SS AVF +RV
Sbjct: 179 NSRGGGDDDDDDKEDEKNDSCGGGSALACTRVVNASSSGLETSHLPPEISSPAVFRCMRV 238
Query: 111 RSMDDSVNEMAYQTSVNIGGHRFSGILYDQGPE------QQSLNASPLDQHQNLNL---- 160
S+DD E AYQT+V+IGGH F GILYDQGP SLN QH +LNL
Sbjct: 239 SSIDDEDEEYAYQTAVSIGGHVFKGILYDQGPSSDHHRYSSSLNGETSHQH-HLNLMDST 297
Query: 161 ---------TTIHSHDGATMAPPSSSATAAHELFFPQPRSLASFRSGVPYFSHTK 206
T +++++G+ PSS TA F A G P+F+ ++
Sbjct: 298 PSAATTNAVTAVNTNNGS--IDPSSLYTAVATPF------NAFVAGGTPFFASSR 344
>UniRef100_Q8GUY8 Hypothetical protein F3K23.16 [Arabidopsis thaliana]
Length = 162
Score = 123 bits (309), Expect = 3e-27
Identities = 69/161 (42%), Positives = 90/161 (55%), Gaps = 23/161 (14%)
Query: 18 GNQAKKGCAYSRCRTCCNNKGFQCQTHVRSTWIPVDRRRHRLMEHQPPPTTNNPHHLHED 77
GNQAKK C Y RCRTCC +K F CQTH+ STW+P RR H + Q P +T+ P +
Sbjct: 1 GNQAKKDCVYMRCRTCCKSKAFHCQTHIESTWVPAYRRSHHKHQSQ-PLSTSIPKGVQIH 59
Query: 78 IPQSHNQNPFTSLEELKFPEAMSSMAVFSSVRVRSMDDSVNEMAYQTSVNIGGHRFSGIL 137
H FP +SS+A F V+V S+DD + AYQT+VNIGGH F GIL
Sbjct: 60 TTPGH------------FPAELSSLADFRCVKVSSIDDGKEQYAYQTTVNIGGHVFRGIL 107
Query: 138 YDQGPEQQSLNASPLDQHQNLNLTTIHSHDGATMAPPSSSA 178
+DQG L+ +D H N N S++ + PS+S+
Sbjct: 108 HDQG-----LHKVMVDHHYNKN-----SNNHQELLTPSTSS 138
>UniRef100_Q5Q0G2 Hypothetical protein [Arabidopsis thaliana]
Length = 345
Score = 123 bits (308), Expect = 3e-27
Identities = 83/235 (35%), Positives = 112/235 (47%), Gaps = 51/235 (21%)
Query: 14 CQECGNQAKKGCAYSRCRTCCNNKGFQCQTHVRSTWIPVDRRRHRLMEHQPPPT-----T 68
CQ+CGNQAKK C + RCRTCC ++GF CQTHV+STW+ +RR R + P
Sbjct: 119 CQDCGNQAKKDCPHMRCRTCCKSRGFDCQTHVKSTWVSAAKRRERQAQLAVLPAKRIRDA 178
Query: 69 NNPHHLHEDIPQSHNQN------------------PFTSLEELKFPEAMSSMAVFSSVRV 110
N+ +D ++ + LE P +SS AVF +RV
Sbjct: 179 NSRGGGDDDDDDKEDEKNDSCGGGSALACTRVVNASSSGLETSHLPPEISSPAVFRCMRV 238
Query: 111 RSMDDSVNEMAYQTSVNIGGHRFSGILYDQGPE------QQSLNASPLDQHQNLNL---- 160
S+DD E AYQT+V+IGGH F G LYDQGP SLN QH +LNL
Sbjct: 239 SSIDDEDEEYAYQTAVSIGGHVFKGXLYDQGPSSDHHRYSSSLNGETSHQH-HLNLMDST 297
Query: 161 ---------TTIHSHDGATMAPPSSSATAAHELFFPQPRSLASFRSGVPYFSHTK 206
T +++++G+ PSS TA F A G P+F+ ++
Sbjct: 298 PSAATTNAVTAVNTNNGS--IDPSSLYTAVATPF------NAFVAGGTPFFASSR 344
>UniRef100_O65517 Hypothetical protein F23E13.150 [Arabidopsis thaliana]
Length = 322
Score = 123 bits (308), Expect = 3e-27
Identities = 65/158 (41%), Positives = 87/158 (54%), Gaps = 23/158 (14%)
Query: 14 CQECGNQAKKGCAYSRCRTCCNNKGFQCQTHVRSTWIPVDRRRHRLMEHQPPPTTNN--- 70
C++CGNQAKK C + RCRTCC ++GF C THVRSTWIPV RRR R + +
Sbjct: 94 CRDCGNQAKKDCTHMRCRTCCKSRGFDCSTHVRSTWIPVARRRERQQQLHMSTSGGGGGS 153
Query: 71 -------------PHHLHEDIPQSHNQNPFTS------LEELKFPEAMSSMAVFSSVRVR 111
H +P + + + S + E FP +SS A+F V++
Sbjct: 154 GSGGAGGGGSSIPKRHRDPTLPGTSSSSRLPSHSAGLEMGEASFPGEVSSDALFRCVKMS 213
Query: 112 SMDDSVN-EMAYQTSVNIGGHRFSGILYDQGPEQQSLN 148
+DD + + AYQT+VNIGGH F GILYDQGPE ++
Sbjct: 214 GVDDGGDGQYAYQTTVNIGGHLFKGILYDQGPESSYMS 251
>UniRef100_Q6YT10 Putative lateral root primordia [Oryza sativa]
Length = 324
Score = 123 bits (308), Expect = 3e-27
Identities = 61/140 (43%), Positives = 85/140 (60%), Gaps = 9/140 (6%)
Query: 14 CQECGNQAKKGCAYSRCRTCCNNKGFQCQTHVRSTWIPVDRRRHRLMEHQPPPTTNNPHH 73
CQ+CGNQAKK C + RCRTCC ++GF C THV+STW+P +RR R ++ +
Sbjct: 116 CQDCGNQAKKDCTHLRCRTCCKSRGFDCATHVKSTWVPAAKRRER--QNLLASAAESSKR 173
Query: 74 LHEDIPQSHNQNPFTSLEEL-------KFPEAMSSMAVFSSVRVRSMDDSVNEMAYQTSV 126
+ + + P TS E +FP +SS AVF VR+ ++++ E+AYQT+V
Sbjct: 174 PRDSAAAATSTTPTTSSGEQQQMMVGERFPREVSSEAVFRCVRLGPVEEADAEVAYQTTV 233
Query: 127 NIGGHRFSGILYDQGPEQQS 146
+IGGH F GIL+D GPE S
Sbjct: 234 SIGGHVFKGILHDVGPEHSS 253
>UniRef100_Q941Y5 Putative LRP (Lateral root primordia) 1 [Oryza sativa]
Length = 340
Score = 115 bits (287), Expect = 9e-25
Identities = 63/152 (41%), Positives = 79/152 (51%), Gaps = 24/152 (15%)
Query: 14 CQECGNQAKKGCAYSRCRTCCNNKGFQCQTHVRSTWIPVDRRRHRLM----EHQPPPT-- 67
CQ+CGNQAKK C + RCRTCC ++GF C THV+STW+P RRR R P T
Sbjct: 115 CQDCGNQAKKDCGHQRCRTCCKSRGFDCSTHVKSTWVPAARRRERQQLTGSASSSPATAS 174
Query: 68 ----TNNPHHLHEDIPQSHNQ----------NPFTSLEELKF----PEAMSSMAVFSSVR 109
+ P L SH + +S ++ F P + + AVF VR
Sbjct: 175 AAAASKKPRLLTSQTTTSHTSTSNATTPRSFDTTSSHQDASFRESLPRQVRAPAVFRCVR 234
Query: 110 VRSMDDSVNEMAYQTSVNIGGHRFSGILYDQG 141
V S+DD +E AYQ +V I GH F G LYDQG
Sbjct: 235 VTSIDDGEDEYAYQATVTINGHVFKGFLYDQG 266
>UniRef100_Q6AV91 Hypothetical protein OJ1286_E05.15 [Oryza sativa]
Length = 453
Score = 114 bits (286), Expect = 1e-24
Identities = 61/160 (38%), Positives = 80/160 (49%), Gaps = 30/160 (18%)
Query: 14 CQECGNQAKKGCAYSRCRTCCNNKGFQCQTHVRSTWIPVDRRRH---------------- 57
C +CGNQAKK C + RCRTCC ++GF C THVRSTW+P RRR
Sbjct: 232 CHDCGNQAKKDCVHHRCRTCCKSRGFDCPTHVRSTWVPAARRRERQQLAGAASSPPTSSA 291
Query: 58 ------------RLMEHQPPPTTNNPHHLHEDIPQSHNQNPFTSLEELK--FPEAMSSMA 103
RL+ Q TT+ + P+S + + + + P + + A
Sbjct: 292 FPAATTASAKKPRLLGSQTTTTTSRTSTSNATTPRSFDTSSSHQVASFRDALPRHVRAPA 351
Query: 104 VFSSVRVRSMDDSVNEMAYQTSVNIGGHRFSGILYDQGPE 143
VF VRV S+DD +E AYQ +V I GH F G LYDQG +
Sbjct: 352 VFRCVRVTSVDDGDDEFAYQAAVTINGHMFRGFLYDQGAD 391
>UniRef100_Q94CK9 Lateral root primordia [Arabidopsis thaliana]
Length = 320
Score = 111 bits (277), Expect = 1e-23
Identities = 68/191 (35%), Positives = 89/191 (45%), Gaps = 33/191 (17%)
Query: 14 CQECGNQAKKGCAYSRCRTCCNNKGFQCQTHVRSTWIPVDRRRHR-LMEHQPPPT----- 67
CQ+CGNQAKK C RCRTCC ++GF C THV+STW+ RRR R +M PT
Sbjct: 112 CQDCGNQAKKECKQRRCRTCCKSRGFDCSTHVKSTWVSAARRRERQVMPTGANPTAGSSL 171
Query: 68 -----TNNPHHLHEDIPQSHNQNPFTSLEEL-------------------KFPEAMSSMA 103
T P + Q TS +P + + A
Sbjct: 172 STSSGTKKPRIVGSQQQQQQQATSHTSTSNTPPQSFETSSSRQDGGGSREAWPGQVRAAA 231
Query: 104 VFSSVRVRSMDDSVNEMAYQTSVNIGGHRFSGILYDQGPEQQSLNASPLDQHQNLNLTTI 163
VF VRV +++D +E AYQ V IGGH F G LYDQG E + S D H +
Sbjct: 232 VFKCVRVTAVEDGDDEYAYQAVVKIGGHVFKGFLYDQGLEPKEGFPSMSDLHLG---GSA 288
Query: 164 HSHDGATMAPP 174
++H+G + + P
Sbjct: 289 NNHNGVSASAP 299
>UniRef100_Q38848 LRP1 [Arabidopsis thaliana]
Length = 320
Score = 110 bits (276), Expect = 2e-23
Identities = 68/191 (35%), Positives = 89/191 (45%), Gaps = 33/191 (17%)
Query: 14 CQECGNQAKKGCAYSRCRTCCNNKGFQCQTHVRSTWIPVDRRRHR-LMEHQPPPT----- 67
CQ+CGNQAKK C RCRTCC ++GF C THV+STW+ RRR R +M PT
Sbjct: 112 CQDCGNQAKKECKQRRCRTCCKSRGFDCSTHVKSTWVSAARRRERQVMPTGANPTAGSSL 171
Query: 68 -----TNNPHHLHEDIPQSHNQNPFTSLEEL-------------------KFPEAMSSMA 103
T P + Q TS +P + + A
Sbjct: 172 STSSGTKKPRIVGSQQQQQQQATSHTSTSNTPPQSFETSSSRQDGGGSREAWPGQVRAAA 231
Query: 104 VFSSVRVRSMDDSVNEMAYQTSVNIGGHRFSGILYDQGPEQQSLNASPLDQHQNLNLTTI 163
VF VRV +++D +E AYQ V IGGH F G LYDQG E + S D H +
Sbjct: 232 VFKCVRVTAVEDGDDEYAYQAVVKIGGHVFKGFLYDQGLEPKEGFPSMSDLHLG---GSA 288
Query: 164 HSHDGATMAPP 174
++H+G + + P
Sbjct: 289 NNHNGVSASVP 299
>UniRef100_Q9SD40 Hypothetical protein F24M12.100 [Arabidopsis thaliana]
Length = 252
Score = 91.7 bits (226), Expect = 1e-17
Identities = 49/112 (43%), Positives = 63/112 (55%), Gaps = 17/112 (15%)
Query: 11 GSRCQECGNQAKKGCAYSRCRTCCNNKGFQCQTHVRSTWIPVDRRRHRLME-HQPPPTTN 69
G CQ+CGNQAKK C++ RCRTCC ++GF+C THVRSTW+P +RR R + P T
Sbjct: 141 GVSCQDCGNQAKKDCSHMRCRTCCKSRGFECSTHVRSTWVPAAKRRERQQQLATVQPQTQ 200
Query: 70 NPHHLHEDIPQSHNQN-PFTS-------------LEELKFPEAMSSMAVFSS 107
P E +P+ H +N P TS LE FP +SS AV +
Sbjct: 201 LPR--GESVPKRHRENLPATSSSLVCTRIPSHSGLEVGNFPAEVSSSAVLGA 250
>UniRef100_Q9M2U4 Hypothetical protein T12E18_120 [Arabidopsis thaliana]
Length = 183
Score = 87.0 bits (214), Expect = 3e-16
Identities = 48/140 (34%), Positives = 73/140 (51%), Gaps = 22/140 (15%)
Query: 14 CQECGNQAKKGCAYSRCRTCCNNKGFQCQTHVRSTWIPVDRRRHRLMEHQPPPTTNNPHH 73
C++CGN+AKK C + RCRTCC ++G+ C THV+STWIP R ++++P
Sbjct: 41 CRDCGNRAKKECLFERCRTCCKSRGYNCVTHVKSTWIPSSATR----------SSSSPSE 90
Query: 74 LHEDIPQSHNQNP-------FTSLEELKF----PEAMSSMAVFSSVRVRSMDDSVN-EMA 121
+ + +P TS +E F P + + AVF RV ++ ++ E+
Sbjct: 91 RKKKLKIDKQSSPNVSLLPTTTSRQERGFREGLPGKIEAPAVFKRTRVTAISNNEQAEIG 150
Query: 122 YQTSVNIGGHRFSGILYDQG 141
YQ +V I GH F G L+ G
Sbjct: 151 YQATVTISGHIFKGFLHYYG 170
>UniRef100_Q5Z5Y2 Putative LRP1 [Oryza sativa]
Length = 384
Score = 79.7 bits (195), Expect = 4e-14
Identities = 29/48 (60%), Positives = 38/48 (78%)
Query: 11 GSRCQECGNQAKKGCAYSRCRTCCNNKGFQCQTHVRSTWIPVDRRRHR 58
G+ CQ+CGN AKK C++ RCRTCC ++GF C THV+STW+P +RR R
Sbjct: 130 GTSCQDCGNNAKKDCSHLRCRTCCRSRGFSCATHVKSTWVPAAKRRER 177
Score = 69.7 bits (169), Expect = 4e-11
Identities = 39/120 (32%), Positives = 62/120 (51%), Gaps = 9/120 (7%)
Query: 94 KFPEAMSSMAVFSSVRVRSMDDSVNEMAYQTSVNIGGHRFSGILYDQGPEQQSLNASPLD 153
+FP +S AVF VR+ ++D++ E+AYQT+V+IGGH F GIL D GP ++ P
Sbjct: 261 RFPPELSVEAVFRCVRIGAVDEADAELAYQTAVSIGGHTFKGILRDHGPADEAAGQLPPS 320
Query: 154 QHQNLNLTTIHSHDGATMAP---------PSSSATAAHELFFPQPRSLASFRSGVPYFSH 204
+ LT + + +++AT+A L P P + +F +G +F H
Sbjct: 321 SAEYHQLTGQGREESSPAGSSEGVGGGHGAATAATSAAVLMDPYPTPIGAFAAGTQFFPH 380
>UniRef100_Q5Q0G3 Hypothetical protein [Arabidopsis thaliana]
Length = 234
Score = 78.2 bits (191), Expect = 1e-13
Identities = 29/45 (64%), Positives = 36/45 (79%)
Query: 14 CQECGNQAKKGCAYSRCRTCCNNKGFQCQTHVRSTWIPVDRRRHR 58
CQ+CGNQAKK C + RCRTCC ++GF CQTHV+STW+ +RR R
Sbjct: 119 CQDCGNQAKKDCPHMRCRTCCKSRGFDCQTHVKSTWVSAAKRRER 163
>UniRef100_UPI000036462C UPI000036462C UniRef100 entry
Length = 449
Score = 40.8 bits (94), Expect = 0.022
Identities = 18/54 (33%), Positives = 27/54 (49%), Gaps = 4/54 (7%)
Query: 7 QEVKGSRCQECGNQAKKGCAYSRCRTCCNNKGFQ----CQTHVRSTWIPVDRRR 56
Q+ K +C++CGN C +S CR CC K F+ C +H D+R+
Sbjct: 379 QKPKYIKCEQCGNPKGNKCVFSLCRGCCKKKAFKEVADCPSHGLRFKTKADKRK 432
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.317 0.129 0.399
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 362,521,651
Number of Sequences: 2790947
Number of extensions: 14471203
Number of successful extensions: 39395
Number of sequences better than 10.0: 60
Number of HSP's better than 10.0 without gapping: 39
Number of HSP's successfully gapped in prelim test: 21
Number of HSP's that attempted gapping in prelim test: 39317
Number of HSP's gapped (non-prelim): 83
length of query: 207
length of database: 848,049,833
effective HSP length: 122
effective length of query: 85
effective length of database: 507,554,299
effective search space: 43142115415
effective search space used: 43142115415
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 72 (32.3 bits)
Lotus: description of TM0258b.1


