

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0253.9
(677 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
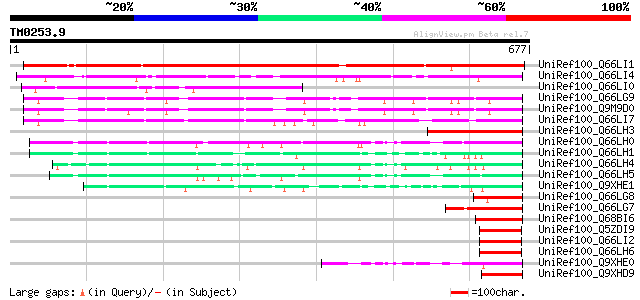
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q66LI1 CENP-C [Medicago truncatula] 687 0.0
UniRef100_Q66LI4 CENP-C [Beta vulgaris subsp. vulgaris] 243 2e-62
UniRef100_Q66LI0 CENP-C [Solanum tuberosum] 231 4e-59
UniRef100_Q66LG9 CENP-C [Arabidopsis thaliana] 221 7e-56
UniRef100_Q9M9D0 T16N11.16 protein [Arabidopsis thaliana] 215 4e-54
UniRef100_Q66LI7 CENP-C [Arabidopsis arenosa] 207 1e-51
UniRef100_Q66LH3 CENP-C [Glycine max] 167 9e-40
UniRef100_Q66LH0 CENP-C1 [Saccharum officinarum] 97 2e-18
UniRef100_Q66LH1 CENP-C2 [Saccharum officinarum] 96 5e-18
UniRef100_Q66LH4 CENP-C [Sorghum propinquum] 92 7e-17
UniRef100_Q66LH5 CENP-C [Sorghum bicolor] 89 3e-16
UniRef100_Q9XHE1 CENPCA protein [Zea mays] 87 2e-15
UniRef100_Q66LG8 CENP-C [Lycopersicon esculentum] 73 3e-11
UniRef100_Q66LG7 CENP-C [Triticum aestivum] 72 5e-11
UniRef100_Q68BI6 Centromere protein C homolog [Arabidopsis thali... 72 7e-11
UniRef100_Q5ZDI9 CENPCA protein-like [Oryza sativa] 70 2e-10
UniRef100_Q66LI2 CENP-C [Oryza sativa] 70 2e-10
UniRef100_Q66LH6 CENP-C [Oryza sativa] 70 2e-10
UniRef100_Q9XHE0 CENPCB protein [Zea mays] 62 4e-08
UniRef100_Q9XHD9 CENPCC protein [Zea mays] 56 4e-06
>UniRef100_Q66LI1 CENP-C [Medicago truncatula]
Length = 697
Score = 687 bits (1772), Expect = 0.0
Identities = 390/663 (58%), Positives = 476/663 (70%), Gaps = 22/663 (3%)
Query: 18 DPLSNYRGLSLFPSTSSALPLSVA-DDLDAIHNHLRSMALRSPAKLVEQTKAILEENPGL 76
DP++NY GLSLF ST S P S DLDAI+N+LRSM L SP +L EQ ++ILE N G
Sbjct: 10 DPIANYSGLSLFRSTFSLQPSSNPFHDLDAINNNLRSMDLGSPTRLAEQGQSILENNLG- 68
Query: 77 FNSQDDERNDVVDVDVDVAEDGQDFPRKRRPGLGLKRAR--FSLKPSTSVSLESLLPNLD 134
FN+++ + DV + DV E+G++FPRKRRPGLGL RAR FSLKP+ S+E LLP+LD
Sbjct: 69 FNTENLTQ-DVENDDVFAVEEGEEFPRKRRPGLGLNRARPRFSLKPTKKPSVEDLLPSLD 127
Query: 135 IENLKDPVEFFKAHERMENARREIQKQMG-VSVEQNQDDVSTRPRQRRPGLRGNDQRSIK 193
I++ KDP EFF AHER ENARRE+QKQ+G VS E NQD ST+PR RRPGL G ++ +K
Sbjct: 128 IKDHKDPEEFFLAHERRENARRELQKQLGIVSSEPNQD--STKPRDRRPGLPGFNRGPVK 185
Query: 194 YRHRYSTEISDKNDHAMASQEAFGSDSLDPDTEKIENGEACLTSLENEVTDSSATEGNKI 253
YRHR+S E D N ++SQE F SD+LD + + G+A TSL+NEV S A E NK
Sbjct: 186 YRHRFSQETLDNNVDVLSSQEVFESDNLDLVGDNTDTGDASPTSLDNEVAGSPAVEENKG 245
Query: 254 CEILDGLLRCNSEDLEGDGAMDLLQERLQIKPVVLEKVSVPDFPDNQPIDLKSLRGSLSK 313
+IL GLL CNSE+LEGDGAM+LLQERL IKP+V EK+SVPDFPD QPIDLK LR + SK
Sbjct: 246 NDILQGLLTCNSEELEGDGAMNLLQERLNIKPIVFEKLSVPDFPDIQPIDLKFLRENSSK 305
Query: 314 PRKALSNIDNLLNRRKNKTPLRQDAGSPAEQLASPTPPRSPFAPLSSLLNHISRLKPSVD 373
PRKALSNIDNLLNR KTPLR+D G +QL SPTPPRSPFA LS L I R KPSVD
Sbjct: 306 PRKALSNIDNLLNRIDIKTPLRRDVGYTEKQLGSPTPPRSPFASLSKLQKQILRSKPSVD 365
Query: 374 PFSAHDIDHLSTTKYSPVHKMNQELNVVGSAKPSNELNDHITKDAVAIGETNAVPDTLRN 433
PFSAH+IDH+S SP +NQE+N+VGS+KP++EL+ + +D +A GETN + DT
Sbjct: 366 PFSAHEIDHISKRNSSPTDTINQEVNIVGSSKPADELSAPVIEDVIAAGETNTILDT--- 422
Query: 434 CASASENSKEDNSGKSSNKLDAPLIEDILAVSESCLIEDAVMNSTSTSQMPMEDNSMEPE 493
SE SKE+ S KSS +++APLIED + VSE+ +++ V+N TST M DNS EPE
Sbjct: 423 ----SEKSKEEVSRKSSEQVNAPLIEDKVGVSETSSVDNPVINCTSTPLKSMVDNSREPE 478
Query: 494 FDANVDRNEPHADMDVDNGGSGMGETVMDDTVAKPNIEVNEQCQFEGKILTENTDAFTAS 553
F+ANVD NEP DMDVD G SGMG+ MDD V + N+E + Q E L EN FTAS
Sbjct: 479 FNANVDSNEPPVDMDVDIGSSGMGKRAMDDIVGRQNVE-PQPYQSEDN-LPENMHEFTAS 536
Query: 554 MPTDDTDIDVVNPLADQSSPG---VIQANSIDKRTTCTNEGSEQCLQEERDGSRAPVE-Q 609
+PTDD ++++V PLADQS+ QANS+DKR+ +N+G E LQE GS APV Q
Sbjct: 537 LPTDDANLNLVIPLADQSNASNQDEHQANSMDKRSGRSNDGPELSLQENTVGSVAPVNGQ 596
Query: 610 KRVKSRSQKDTK-SKRLAKRQSLAAAGTSWTSGIRRSTRIRTRPLEYWKGERLVYGRIHQ 668
KRVK +QK +K KR R SLA AGTSW SG+RRS R RTRPLEYWKGER+VYGR+H+
Sbjct: 597 KRVKVCAQKVSKGKKREQHRMSLADAGTSWESGVRRSKRFRTRPLEYWKGERMVYGRVHE 656
Query: 669 SKS 671
S S
Sbjct: 657 SLS 659
>UniRef100_Q66LI4 CENP-C [Beta vulgaris subsp. vulgaris]
Length = 710
Score = 243 bits (619), Expect = 2e-62
Identities = 217/706 (30%), Positives = 345/706 (48%), Gaps = 86/706 (12%)
Query: 9 TVAEQQQPLDPLSNYRGLSLFPSTSSALPLSVADDLD----------AIHNHLRSMALRS 58
T E +DPL++Y LSLFP T S+L S + +D +I HL++ L S
Sbjct: 5 TETEGSDLVDPLADYSSLSLFPRTFSSLSTSSSSSIDLRKPNSPILNSILTHLKAKVLSS 64
Query: 59 PAKLVEQTKAILEENPGLFNSQDDERNDVVDVDVDVAEDGQDFPRKRRPGLGLKRARFSL 118
P K+++Q K ILE++ + E +AE+ + PR+RRP LGLKRA+FS
Sbjct: 65 PDKMLKQAKPILEDSLNFLKTDKTEA---------IAEN-EKVPRERRPALGLKRAKFSA 114
Query: 119 KPSTSVSLESLLPNLDIENLKDPVEFFKAHERMENARREIQKQMG---VSVEQNQDDVST 175
KP S SL ++D++ L DP E F A ERMENA++E+Q+ G ++QN+ ++
Sbjct: 115 KPMPSQPDASLEFSIDVDKLSDPEELFSAFERMENAKKEVQRLRGEPLFDLDQNRASLAR 174
Query: 176 RPRQRRPGLRGNDQRSIKYRHR-YSTE-ISDKNDHAMASQEAFGSDSLDPDTEKI-ENGE 232
RPR RP L G S Y HR YS++ ++D ++ SQE + L P + + +
Sbjct: 175 RPR--RPSLLG----SSTYTHRPYSSKSMADVDETLFPSQETIYDEILSPIRDDVLPHAN 228
Query: 233 ACLTSLENEVTDSSATEGNKICEILDGLLRCNSEDLEGDGAMDLLQERLQIKPVVLEKVS 292
S ++DS + +K+ E D LL N E L+ D +LL+++LQIKPV + K +
Sbjct: 229 VVNHSPSVILSDSKSRTTSKVSEF-DELLSSNYEGLDEDEVENLLRDKLQIKPVEIGKAA 287
Query: 293 VPDFPDNQPIDLK-SLRGSLSKPRKALSNIDNLLNRRK--NKTPLRQDAGSPAEQLASPT 349
+P N + L+ S+ SL+ P+ ++N+++ R K D SPA+
Sbjct: 288 LP----NWRVQLRESMTSSLALPK-----VNNMVSHRSAARKAVRSSDKVSPAKS----- 333
Query: 350 PPRSPFAPLSSLLNHISRLKPSVDPFSAHDIDHLSTTKYSPVHKMNQELNVVGSAKPSN- 408
+SP A +S L HI++ PS DPFS I+ P + + G + +
Sbjct: 334 --QSPLASISLLNRHIAKHTPSKDPFSPIHINSSPAEATIPAREPENRADQAGESDEIDA 391
Query: 409 ELNDHITKDAVAIGETN---AVPDTLRNC-----ASASENSKEDNSGKSS---NKLD--- 454
N + A GE N ++ D+ + S+ N KED G+ NK D
Sbjct: 392 RKNSCAFSTSPAHGEENIAVSIQDSFVSMEKDVDTSSEPNEKEDLIGQLDQIGNKKDLCE 451
Query: 455 ---APLIED-ILAVSESCLIEDAVMNSTSTSQMPMEDNSMEPEFDANVDRNEPHADMDVD 510
+P+ E+ +AVS + A N T++ +D+ P + V R +D++
Sbjct: 452 FGRSPVHEEESVAVSGQDSLALASKNVVPTNEPEKQDDEDVPAAELPV-RQLLFEGLDME 510
Query: 511 NGGSGMGETVMDDTVAKPNIEVNEQCQFEGKILTENTDAFTASMPTDDTDIDVVNPLADQ 570
SG GE V + +E C+ + EN ++ ++ ++ V P ++
Sbjct: 511 EPTSGGGEGVHLGEL----VEKGGTCEPSTR---ENCSVEELNVSVEELNVSVEEPTTER 563
Query: 571 SSPGVIQANSIDKRTTCTNEGS-EQCLQEERDGSRAPVEQ----KRVKSRSQKDTKSKR- 624
+ ++ + T + G E+ +Q E+ S A V K+ K + + +SKR
Sbjct: 564 VASSKRKSKELGISTAPSEAGVFEEGMQAEKSQSDALVSSEGRSKKRKVQGAQPRRSKRD 623
Query: 625 -LAKRQSLAAAGTSWTSGIRRSTRIRTRPLEYWKGERLVYGRIHQS 669
+R SL AGT W +G+RRSTRI+ RPL+YWKGER +YGR+H+S
Sbjct: 624 LQQRRTSLYCAGTKWEAGVRRSTRIKMRPLQYWKGERFLYGRVHES 669
>UniRef100_Q66LI0 CENP-C [Solanum tuberosum]
Length = 384
Score = 231 bits (590), Expect = 4e-59
Identities = 153/384 (39%), Positives = 220/384 (56%), Gaps = 30/384 (7%)
Query: 16 PLDPLSNYRGLSLFPST---SSALPLSV-ADDLDAIHNHLRSMALRSPAKLVEQTKAILE 71
P+DPL + GLSL P+T S+ +SV DL+ IHN ++SM + P L+E+ + I++
Sbjct: 10 PVDPLHSLAGLSLLPTTVRVSTDASVSVNPKDLELIHNFMKSMETKGPG-LLEEAREIVD 68
Query: 72 ENPGLFNSQDDERNDVVDVDVDVAEDGQDFPRKRRPGLGLKRARFSLKP-STSVSLESLL 130
L N++ +D D+A G++ ++RRPGLG KRARFSLKP STS S+
Sbjct: 69 NGAELLNTKFTSFILSKGIDGDLAMKGKEKLQERRPGLGRKRARFSLKPPSTSQPTVSVA 128
Query: 131 PNLDIENLKDPVEFFKAHERMENARREIQKQMGVSVEQNQDDVSTRP---RQRRPGLRGN 187
P LDI+ L DPVEFF E++E+A +EI++Q G S+ + DV+ P R+RRPG+ G
Sbjct: 129 PRLDIDQLSDPVEFFSVAEKLEDAEKEIERQKGSSI--HDPDVNNPPANARRRRPGILG- 185
Query: 188 DQRSIKYRHRYSTEISDKNDHAMASQEAFGSDSLDPDTEKIENGEACLTSL--------E 239
+S+KY+HR+S+ + +D ++SQE D L +E+G L E
Sbjct: 186 --KSVKYKHRFSSTQPENDDAFISSQETLEDDIL------VEHGSQLPEELHGLNVELQE 237
Query: 240 NEVTDSSATEGNKICEILDGLLRCNSEDLEGDGAMDLLQERLQIKPVVLEKVSVPDFPDN 299
E+T S N+I +ILD LL + EDL+ D A+ LQERLQI P+ L + +P+FP
Sbjct: 238 AELTGSVKKTENRINKILDELLSGSDEDLDRDMAVSKLQERLQINPIELGTLCIPEFPVT 297
Query: 300 QPIDLKSLRGSLSKPRKALSNIDNLLNRRKNKTPL--RQDAGSPAEQLASPTPPRSPFAP 357
+D K+ + KPRK + L+ TP +Q SPA +LASPTPP+SPF
Sbjct: 298 GKLDGKAFGERIQKPRKFSLEVRELVKSATEGTPSSHKQHEESPASKLASPTPPKSPFGS 357
Query: 358 LSSLLNHISRLKPSVDPFSAHDID 381
LS L + + P DPFS +ID
Sbjct: 358 LSLLKKKLMQSNPLRDPFSPLNID 381
>UniRef100_Q66LG9 CENP-C [Arabidopsis thaliana]
Length = 705
Score = 221 bits (562), Expect = 7e-56
Identities = 213/712 (29%), Positives = 334/712 (45%), Gaps = 123/712 (17%)
Query: 18 DPLSNYRGLSLFPSTSSAL-----PLSVADDLDAIHNHLRSMALRSPAKLVEQTKAILEE 72
DPL Y GLSLFP T +L P ++DL H L+SM ++ EQ KAILE
Sbjct: 15 DPLQAYSGLSLFPRTLKSLSNPLPPSYQSEDLQQTHTLLQSMPFEIQSEHQEQAKAILE- 73
Query: 73 NPGLFNSQDDERNDVVDVDVDVAEDGQDFPRKRRPGLGLKRARFSLKPSTSVSLESLLPN 132
DVDVDV + R+RRPGL KR FSL +TS + P+
Sbjct: 74 ----------------DVDVDVQLNPIPNKRERRPGLDRKRKSFSLHLTTSQP-PPVAPS 116
Query: 133 LDIENLKDPVEFFKAHERMENARREIQKQMGVSVEQNQDDVSTRPRQRRPGLRGNDQRSI 192
D +FF A+++ E A RE QKQ G SV Q++ +R R RRPG+ G +R
Sbjct: 117 FDPSKYPRSEDFFAAYDKFELANREWQKQTGSSVIDIQENPPSR-RPRRPGIPGRKRRPF 175
Query: 193 K--YRHRYSTEI----SDKNDHAMASQEAFGSDSLDPDTEKIENGEACLTSLENEVTDSS 246
K + Y T++ + + + +AS+++ S + A +T+++ EV DS+
Sbjct: 176 KESFTDSYFTDVINLEASEKEIPIASEQSLESATA-----------AHVTTVDREVDDST 224
Query: 247 ATEGNKICEILDGLLRCNSEDLEGDGAMDLLQERLQIKPVVLEKVSVPDFPDNQPIDLKS 306
+ +L LL C+ E+LEGDGA+ LL+ERLQIK +EK S+P+F D + ++LK+
Sbjct: 225 VDTDKDLNNVLKDLLACSREELEGDGAIKLLEERLQIKSFNIEKFSIPEFQDVRKMNLKA 284
Query: 307 LRGSLSKPRKALSNIDNLLNRRKNKTPLRQDAGSPAEQLASPTPPRSPFAPLSSLLNHIS 366
GS RK+LS+I N+L + N+ +R+++ SP+ Q + H S
Sbjct: 285 -SGSNPPNRKSLSDIQNIL-KGTNRVAVRKNSHSPSPQ----------------TIKHFS 326
Query: 367 RLKPSVDPFSAHDIDHL-------STTKYSPVHK--MNQELNVVGSAKPSNELNDHITKD 417
P VD FS DI +L S P+ K N VG+ ++ ND + K
Sbjct: 327 SPNPPVDQFSFPDIHNLLPGDQQPSEVNVQPIAKDIPNTSPTNVGTVDVASPFNDSVVKR 386
Query: 418 AVAIGETNAVPDTLRNCASASENSKEDNSGKSSNKLDAPLIEDILAVSESCLIEDAVMNS 477
+ GE ++ + + S S++ N D +++ I S + L ++ M +
Sbjct: 387 S---GEDDS---HIHSGIHRSHLSRDGNP-------DICVMDSISNRSSAMLQKNVDMRT 433
Query: 478 TSTS-QMPMED-----NSMEPEFDANVDRNEPHADMDVDNGGSGMGE--TVMDDTV---- 525
+PM + N+ + E DA ++ + + + + TV +D++
Sbjct: 434 KGKEVDVPMSESGANRNTGDRENDAEINEETDNLERLAECASKEVTRPFTVEEDSIPYQQ 493
Query: 526 ---AKPNIEVNEQCQFEGKILTENTDAFTASMPTDDTDIDVVNPLADQSSPGV-----IQ 577
+K EQ G L E+ + ++ + + L +++P V Q
Sbjct: 494 GASSKSPNRAPEQYNTMGGSL-EHAEHNQGLHEEENVNTGSASGLQVENAPEVHKYSHKQ 552
Query: 578 ANSIDKR------------TTCTNEGSEQCLQEERDGSRAPVEQKRVKSRSQKDTKSKR- 624
N KR T G ++ ++ SRA +Q + KS +++ K K+
Sbjct: 553 TNKRRKRGSSDSNVKKRSKTVHGETGGDKQMKTLPHESRAK-KQTKGKSNEREEKKPKKT 611
Query: 625 -------LAKRQSLAAAGTSWTSGIRRSTRIRTRPLEYWKGERLVYGRIHQS 669
+ R+SLAAAGT G+RRSTRI++RPLEYW+GER +YGRIH+S
Sbjct: 612 LTHEGKLFSCRKSLAAAGTKIEGGVRRSTRIKSRPLEYWRGERFLYGRIHES 663
>UniRef100_Q9M9D0 T16N11.16 protein [Arabidopsis thaliana]
Length = 710
Score = 215 bits (547), Expect = 4e-54
Identities = 213/717 (29%), Positives = 334/717 (45%), Gaps = 128/717 (17%)
Query: 18 DPLSNYRGLSLFPSTSSAL-----PLSVADDLDAIHNHLRSMALRSPAKLVEQTKAILEE 72
DPL Y GLSLFP T +L P ++DL H L+SM ++ EQ KAILE
Sbjct: 15 DPLQAYSGLSLFPRTLKSLSNPLPPSYQSEDLQQTHTLLQSMPFEIQSEHQEQAKAILE- 73
Query: 73 NPGLFNSQDDERNDVVDVDVDVAEDGQDFPRKRRPGLGLKRARFSLKPSTSVSLESLLPN 132
DVDVDV + R+RRPGL KR FSL +TS + P+
Sbjct: 74 ----------------DVDVDVQLNPIPNKRERRPGLDRKRKSFSLHLTTSQP-PPVAPS 116
Query: 133 LDIENLKDPVEFFKAHERMEN-----ARREIQKQMGVSVEQNQDDVSTRPRQRRPGLRGN 187
D +FF A+++ E A RE QKQ G SV Q++ +R R RRPG+ G
Sbjct: 117 FDPSKYPRSEDFFAAYDKFEPGFETVANREWQKQTGSSVIDIQENPPSR-RPRRPGIPGR 175
Query: 188 DQRSIK--YRHRYSTEI----SDKNDHAMASQEAFGSDSLDPDTEKIENGEACLTSLENE 241
+R K + Y T++ + + + +AS+++ S + A +T+++ E
Sbjct: 176 KRRPFKESFTDSYFTDVINLEASEKEIPIASEQSLESATA-----------AHVTTVDRE 224
Query: 242 VTDSSATEGNKICEILDGLLRCNSEDLEGDGAMDLLQERLQIKPVVLEKVSVPDFPDNQP 301
V DS+ + +L LL C+ E+LEGDGA+ LL+ERLQIK +EK S+P+F D +
Sbjct: 225 VDDSTVDTDKDLNNVLKDLLACSREELEGDGAIKLLEERLQIKSFNIEKFSIPEFQDVRK 284
Query: 302 IDLKSLRGSLSKPRKALSNIDNLLNRRKNKTPLRQDAGSPAEQLASPTPPRSPFAPLSSL 361
++LK+ GS RK+LS+I N+L + N+ +R+++ SP+ Q
Sbjct: 285 MNLKA-SGSNPPNRKSLSDIQNIL-KGTNRVAVRKNSHSPSPQ----------------T 326
Query: 362 LNHISRLKPSVDPFSAHDIDHL-------STTKYSPVHK--MNQELNVVGSAKPSNELND 412
+ H S P VD FS DI +L S P+ K N VG+ ++ ND
Sbjct: 327 IKHFSSPNPPVDQFSFPDIHNLLPGDQQPSEVNVQPIAKDIPNTSPTNVGTVDVASPFND 386
Query: 413 HITKDAVAIGETNAVPDTLRNCASASENSKEDNSGKSSNKLDAPLIEDILAVSESCLIED 472
+ K + GE ++ + + S S++ N D +++ I S + L ++
Sbjct: 387 SVVKRS---GEDDS---HIHSGIHRSHLSRDGNP-------DICVMDSISNRSSAMLQKN 433
Query: 473 AVMNSTSTS-QMPMED-----NSMEPEFDANVDRNEPHADMDVDNGGSGMGE--TVMDDT 524
M + +PM + N+ + E DA ++ + + + + TV +D+
Sbjct: 434 VDMRTKGKEVDVPMSESGANRNTGDRENDAEINEETDNLERLAECASKEVTRPFTVEEDS 493
Query: 525 V-------AKPNIEVNEQCQFEGKILTENTDAFTASMPTDDTDIDVVNPLADQSSPGV-- 575
+ +K EQ G L E+ + ++ + + L +++P V
Sbjct: 494 IPYQQGASSKSPNRAPEQYNTMGGSL-EHAEHNQGLHEEENVNTGSASGLQVENAPEVHK 552
Query: 576 ---IQANSIDKR------------TTCTNEGSEQCLQEERDGSRAPVEQKRVKSRSQKDT 620
Q N KR T G ++ ++ SRA +Q + KS +++
Sbjct: 553 YSHKQTNKRRKRGSSDSNVKKRSKTVHGETGGDKQMKTLPHESRAK-KQTKGKSNEREEK 611
Query: 621 KSKR--------LAKRQSLAAAGTSWTSGIRRSTRIRTRPLEYWKGERLVYGRIHQS 669
K K+ + R+SLAAAGT G+RRSTRI++RPLEYW+GER +YGRIH+S
Sbjct: 612 KPKKTLTHEGKLFSCRKSLAAAGTKIEGGVRRSTRIKSRPLEYWRGERFLYGRIHES 668
>UniRef100_Q66LI7 CENP-C [Arabidopsis arenosa]
Length = 710
Score = 207 bits (526), Expect = 1e-51
Identities = 196/717 (27%), Positives = 311/717 (43%), Gaps = 127/717 (17%)
Query: 18 DPLSNYRGLSLFPSTSSAL--------PLSVADDLDAIHNHLRSMALRSPAKLVEQTKAI 69
DPL Y GLSLFP T +L PL ++DL H LRSM + EQ KAI
Sbjct: 14 DPLQAYSGLSLFPRTLKSLSNPLPPPPPLHQSEDLQQTHTLLRSMPFEVQNEHQEQAKAI 73
Query: 70 LEENPGLFNSQDDERNDVVDVDVDVAEDGQDFPRKRRPGLGLKRARFSLKPSTSVSLESL 129
LE D++VDV + R+RRPGL KR FSL TS +
Sbjct: 74 LE-----------------DMNVDVKLNPIPNKRERRPGLDRKRKSFSLN-LTSSQPPPV 115
Query: 130 LPNLDIENLKDPVEFFKAHERMENARREIQKQMGVSVEQNQDDVSTRPRQRRPGLRGNDQ 189
P+ D P +FF +++ E A RE QKQ G SV Q + +R R RRPG+ G +
Sbjct: 116 APSFDGSKYPRPEDFFAEYDKFELANREWQKQTGSSVIDTQQNPPSR-RPRRPGIPGRKR 174
Query: 190 RSIKYRHRYSTEISDKNDHAMASQEAFGSDSLDPDTEKIENGEACLTSLENEVTDSSATE 249
R ++ S N A + S+ T A +T+++ EV DS+
Sbjct: 175 RPFEHTFTDSYFTDAINLEASDKENPIASEQCLERTTA-----AHVTTVDREVDDSTVDT 229
Query: 250 GNKICEILDGLLRCNSEDLEGDGAMDLLQERLQIKPVVLEKVSVPDFPDNQPIDLKSLRG 309
+ IL LL + ++LEGD A+ LL++ LQI+ + +EK S+P+F D + ++LK+ G
Sbjct: 230 DKDLNNILKKLLASSRDELEGDAAVKLLEDHLQIESLNVEKFSIPEFQDVRKMNLKA-SG 288
Query: 310 SLSKPRKALSNIDNLLNRRKNKTPLRQDAGSPAEQ----LASPTPPRSPFA--------- 356
S RK+LS+I N+L + ++ R+++ SP+ Q +SP PP F+
Sbjct: 289 SNPSNRKSLSDIQNIL-KGTHRVAGRKNSHSPSPQTRKHFSSPNPPVDQFSFPDIHNLLP 347
Query: 357 --------PLSSLLNHISRLKP--------------SVDPFSAHDIDHLSTTKYSPVHKM 394
+ L I+ P SV+ S D H+ +S +H+
Sbjct: 348 GDQQPSEVDVQPLAKDIANTSPSNVGTVDVASPFNNSVEKRSDEDDSHI----HSGIHRS 403
Query: 395 ------NQELNVVGSAKPSNELNDHITKDAVAIGETNAVPDTLRNCASASENSKEDNSGK 448
N ++ V+ S N + D G+ VP SE+ N+G+
Sbjct: 404 HLRPDGNVDICVMDSISNRNSAMLEVNVDMRTTGKEVDVP--------MSESGANRNTGQ 455
Query: 449 SSNKLD-------APLIEDI--------LAVSESCLIEDAVMNSTSTSQMPMEDNSMEPE 493
N +D ++ + V E + +S S ++ P + N+M+
Sbjct: 456 RENDIDINEETGHLEMLAEYASKEATRPFTVEEDSIPYQQGTSSNSPNRAPEQYNTMDGP 515
Query: 494 FDANVDRNEPHADMDVD-NGGSGMGETVMDDTVAKPNIEVNEQCQFEGKILTENTDAFTA 552
+ H + +V+ + SG+ E + + + + N+
Sbjct: 516 SEHAEHNQGLHEEENVNTDSASGLQENALQEVHNSSHKQTNK----------------LR 559
Query: 553 SMPTDDTDIDVVNPLADQSSPGVIQANSIDKRTTCTNEGSEQCLQEERDGSRAPVEQKRV 612
+ D+++ + + G Q ++ + + + + E E+K
Sbjct: 560 KRGSSDSNVKKRSKTVHGETGGDPQMKTLPHESGVKKQTKRKSNERE--------EKKPK 611
Query: 613 KSRSQKDTKSKRLAKRQSLAAAGTSWTSGIRRSTRIRTRPLEYWKGERLVYGRIHQS 669
+R + K ++R+SLAAAGT G+RRSTRI++RPLEYWKGER +YGRIH+S
Sbjct: 612 NTRKTLTREGKLFSRRKSLAAAGTKMEGGVRRSTRIKSRPLEYWKGERFLYGRIHES 668
>UniRef100_Q66LH3 CENP-C [Glycine max]
Length = 170
Score = 167 bits (423), Expect = 9e-40
Identities = 82/126 (65%), Positives = 101/126 (80%), Gaps = 1/126 (0%)
Query: 545 ENTDAFTASMPTDDTDIDVVNPLADQSSPGVIQANSIDKRTTCTNEGSEQCLQEERDGSR 604
EN FTAS+PTD +++V+NPLADQ +P +ANS+DKR +++ E CLQE+ DGS
Sbjct: 5 ENMQTFTASIPTDGMNLNVLNPLADQPNPAGYEANSVDKRARRSDDDPEHCLQEKTDGSM 64
Query: 605 APVE-QKRVKSRSQKDTKSKRLAKRQSLAAAGTSWTSGIRRSTRIRTRPLEYWKGERLVY 663
PV QKRVKS QK +K+K+L+ R SLAAAGTSWTSG+RRSTRIRTRPLEYWKGERLVY
Sbjct: 65 VPVNGQKRVKSLVQKVSKNKKLSNRHSLAAAGTSWTSGLRRSTRIRTRPLEYWKGERLVY 124
Query: 664 GRIHQS 669
GR+H+S
Sbjct: 125 GRVHKS 130
>UniRef100_Q66LH0 CENP-C1 [Saccharum officinarum]
Length = 709
Score = 96.7 bits (239), Expect = 2e-18
Identities = 165/699 (23%), Positives = 295/699 (41%), Gaps = 105/699 (15%)
Query: 27 SLFPSTSSALPLSVADDLDAIHNHLR-SMALRSPAKLVEQTKAILEENPGLFNSQDDERN 85
+L P+ +SA S + DA+ + + +L+ +L++Q K +L+E+ + D+
Sbjct: 20 TLGPAPASATGSSPSKARDALLEAIAVARSLKGSEELLKQAKMVLKEHGDIQALYHDD-- 77
Query: 86 DVVDVDVDVAEDGQDFPRKRRPGLGLKRARFSLKPSTSVSLESLLPNLDIENLKDPVEFF 145
V +G + RRP L KRARF++K + S + ++ + N+ DPVE+F
Sbjct: 78 ---GVKARPPANGSKEQQGRRPALNRKRARFTMKDTASKPMP-VVDQSKLTNISDPVEYF 133
Query: 146 KAHERMENARREIQKQMGVSVEQ--NQDDVSTRPRQRRPGLRGNDQ-RSIKYRHRYSTEI 202
+R+E A+ EIQ+ G + ++ N D V RQ PG RG S K T+
Sbjct: 134 MTLDRLEEAQEEIQRLHGGAEKRVLNFDPVDEPKRQ--PGFRGRKSVCSFKVIEDADTQD 191
Query: 203 SDKNDHAMASQEAFGSDSLDPDTEKIENGEACLTSLENEV----TDSSATEGNKICEILD 258
+ + + S D + E C+ S E DS A + + D
Sbjct: 192 PIEVPASQTATTTGSQFSQDVMHVVADKNEQCVPSSSGEAISGKEDSLAEKDGR-----D 246
Query: 259 GL--LRCNSEDLEGDGAMDLLQERLQIKPVVLEKVSVPD-FPDNQPIDLKS-LRGS---- 310
L L + + L+ + +++ L K + E+VS+ + P +P+ + +GS
Sbjct: 247 DLTYLLTSMQHLDESEEEEFIRKTLGFKEIRKERVSLHNSIPGVRPVRSNTEQKGSMRVH 306
Query: 311 -----LSKPRK-ALSNIDNLLNRR---KNKTPLRQDAGSPAEQLASPTPPRS----PFAP 357
L +PR+ +S ++ L R K + GSP + P+ P
Sbjct: 307 PPESPLPQPRQDRISELEKHLFPRGAANAKCTDDESEGSPDIVMGEPSLVHDSSDVPMTD 366
Query: 358 LSSLLNHISRLKPSVDPFSAHDIDHLSTTKYSPVHKMNQEL--NVVGSAKPS--NELNDH 413
+ + I R P+V +A I L P H ++ + VG + + + N+
Sbjct: 367 ENFTGSEIDRETPNVGARAAGHI--LDPEPCLPDHAYERQSRGSSVGLCRDTEVTKENEA 424
Query: 414 ITKDAVAIGETNAVPD--TLRNCASASENSKEDNSGKSSNKL-------------DAPL- 457
+ +++ E + D T+ S +E S G S+ +L D L
Sbjct: 425 CRRSNISVEEDDVPIDYPTIGRSTSETETSSHHLEGSSTEELVSKPGRHAAPDGIDRTLH 484
Query: 458 -----IEDILAVSESCLIEDAVMNSTSTSQMPMEDNSMEPEFDANVDRNEPHADMDVDNG 512
I+ + V E +++D S+ + +MP+ED ++P N+P +G
Sbjct: 485 AAEDSIQHLEVVKEGGVLQD---KSSQSLEMPLED--IDPV-------NQPQM-----HG 527
Query: 513 GSGMGETVMDDTVAKPNIEVNEQCQFEGKILTENTDAFTASMPTDDTDIDVVNPLADQSS 572
GS + ++ EGK+ ++ +AD+SS
Sbjct: 528 GSTKKLAPDLCNALSLTKQKKQKAAQEGKMKKQSKRG---------------KKVADESS 572
Query: 573 PGV-IQANSIDKRTTCTNEGSEQCLQEERDGSRAP-VEQKRVKSRSQKDTKSKRLAKRQS 630
+ I ++D N+ E ++++R S P + + +Q+ K+K+L +R+
Sbjct: 573 HALEIPQANLDSENQPHND--EVNIEQQRVLSTTPSPNHAKGQKGAQRTNKTKKLNQRKI 630
Query: 631 LAAAGTSWTSGIRRSTRIRTRPLEYWKGERLVYGRIHQS 669
L AG + S +RRSTR R+RPLE+W GERL+YG I+ +
Sbjct: 631 LGDAGLAQPSVVRRSTRTRSRPLEHWLGERLLYGPINDT 669
>UniRef100_Q66LH1 CENP-C2 [Saccharum officinarum]
Length = 709
Score = 95.5 bits (236), Expect = 5e-18
Identities = 170/701 (24%), Positives = 282/701 (39%), Gaps = 109/701 (15%)
Query: 27 SLFPSTSSALPLSVADDLDAIHNHLR-SMALRSPAKLVEQTKAILEENPGLFNSQDDERN 85
+L P+ +SA S + DA+ + + +L+ +L++Q K +L+E+ + D+
Sbjct: 20 TLGPAHASATGPSPSKAGDALLEAIAVARSLKGSEELLKQAKMVLKEHGDIQALYHDD-- 77
Query: 86 DVVDVDVDVAEDGQDFPRKRRPGLGLKRARFSLKPSTSVSLESLLPNLDIENLKDPVEFF 145
V +G + RRP L KRARF++K + S + ++ + N+ DPVE+F
Sbjct: 78 ---GVKARPPANGSKEQQGRRPALNRKRARFTMKDTASKPMP-VVDQSKLMNISDPVEYF 133
Query: 146 KAHERMENARREIQKQMGVSVEQ--NQDDVSTRPRQRRPGLRGNDQR-SIKYRHRYSTEI 202
+R+E A+ EIQ+ G + ++ N D V RQ PG G S K T+
Sbjct: 134 MTLDRLEEAQEEIQRLHGGAEKRVLNFDPVDEPKRQ--PGFHGRKSVCSFKVIEDADTQD 191
Query: 203 SDKNDHAMASQEAFGSDSLDPDTEKIENGEACLTSLENEVTDSSATEGNKICEILDGL-- 260
+ + + S D + E C+ S E S + ++ D L
Sbjct: 192 PIEVPASQTATTTGSQFSQDVMHVVADKNEQCVPSSSGEAI-SGKEDSLAEKDVRDDLTY 250
Query: 261 LRCNSEDLEGDGAMDLLQERLQIKPVVLEKVSVPDFPDNQPIDLKSLRGSLSKPRKALSN 320
L + + L+ + +++ L K + E+VS L S+ R SN
Sbjct: 251 LLTSMQHLDESEEEEFIRKTLGFKEIRKERVS--------------LHNSIPGVRPVRSN 296
Query: 321 IDNLLNRRKN--KTPLRQDAGSPAEQLASPTPPRSPFAPLSSLLNHISRLKPSV---DPF 375
+ + R + ++PL Q +L PR A + + S P + +P
Sbjct: 297 TEQKGSMRVHPPESPLPQPRQDRISELEKHLFPRG--AANAKCTDDESEGSPDIVMGEPL 354
Query: 376 SAHDI-DHLSTTKYSPVHKMNQELNVVGSAKPSNELNDH--ITKDAVAIGETNAVPDTLR 432
HD D L T + ++++E VG+ + L+ + A + R
Sbjct: 355 LVHDSSDVLMTDENFTGSEIDRETPNVGARAAGHILDPEPCLLDHAYERQSRGSSVGLCR 414
Query: 433 NCASASENSKEDNSGKSSNKLDAPLIEDILAVSESCLIEDAVMNSTSTSQMPMEDNSMEP 492
+ A EN S S + D P+ D + S T TS +E +S E
Sbjct: 415 DTEVAKENEACRRSNISVEEDDVPI--DYPTIGRST-------RETETSSHHLEGSSTEE 465
Query: 493 EFDANVDRNEPHADMDVDNGGSGMGETVMDDTVAKPNIEVNEQCQFEGKILTENTDAFTA 552
V + HA D G+ T+ + ++EV + EG +L + + +
Sbjct: 466 L----VSKPGRHAAPD------GIDRTLHAAEDSIQHLEVVK----EGGVLQDKSSQ-SL 510
Query: 553 SMPTDDTDIDVVNP------LADQSSPGVIQANSI--DKRTTCTNEG------------- 591
MP +D ID VN + +P + A S+ K+ EG
Sbjct: 511 EMPLED--IDPVNQPQMHGGSTKKLAPDLCNALSLTKQKKQKAAQEGKMKKQSKRGKKVA 568
Query: 592 --SEQCL---QEERDGSRAP------VEQKRVKS------------RSQKDTKSKRLAKR 628
S L Q D P +EQ+RV S +Q+ K+K+L +R
Sbjct: 569 DESSHALEIPQANLDSENQPHNDDVNIEQQRVLSITPSPNHAKGQKGAQRTNKTKKLNQR 628
Query: 629 QSLAAAGTSWTSGIRRSTRIRTRPLEYWKGERLVYGRIHQS 669
+ L AG + S +RRSTR R+RPLE+W GERL+YG I+ +
Sbjct: 629 KILGDAGLAQPSVVRRSTRTRSRPLEHWLGERLLYGPINDT 669
>UniRef100_Q66LH4 CENP-C [Sorghum propinquum]
Length = 758
Score = 91.7 bits (226), Expect = 7e-17
Identities = 168/706 (23%), Positives = 286/706 (39%), Gaps = 126/706 (17%)
Query: 57 RSPA---KLVEQTKAILEENPGLFNSQDDERNDVVDVDVDVAEDGQDFPRKRRPGLGLKR 113
RSP +LV+Q K +L+E+ + Q +D V V V + + + RR L KR
Sbjct: 46 RSPKGSEELVKQAKMVLKEHGDI---QPLYHDDGVQVRPPVNDSKEQ--QGRRLALNRKR 100
Query: 114 ARFSLKPSTSVSLESLLPNLDIENLKDPVEFFKAHERMENARREIQKQMGVSVEQNQDDV 173
ARF++K + S ++ + N+ DPVE+F +R+E A+ EIQ+ G + +
Sbjct: 101 ARFTMKDTASKPTP-VVDQSKLTNISDPVEYFMTLDRLEEAQEEIQRLNGAAEKCVLKFD 159
Query: 174 STRPRQRRPGLRGNDQ-RSIKYRHRYSTEISDKNDHAMASQEAFGSDSLDPDTEKIENGE 232
++R+PGL G RS K T+ + + + S D + E
Sbjct: 160 PVDQQKRQPGLCGRKSIRSFKIIGDADTQDPIEVPASQTTTLTGSQVSQDAMHAVADKNE 219
Query: 233 ACLTSLENEVT----DSSATEGNK--ICEILDGLLRCNSEDLEGDGAMDLLQERLQIKPV 286
C+ S E DS A + + +L+ L + + EG + + L I
Sbjct: 220 QCVRSSSGEAISGKEDSLAQKDGRDDFTYLLNSLQHLDESEEEG-----FILKTLGIGEP 274
Query: 287 VLEKVSVPDFPDNQPIDLKSLR------GSL-SKPRKALSNIDNLLNRRKNKTPLRQDAG 339
E+VS F ++ P ++SLR GS+ + P ++L + L +R G
Sbjct: 275 RKERVS---FRNSIP-GVRSLRSNNEQKGSMRAHPLESLL-LQPLQDRISELEKHLFPGG 329
Query: 340 SPAEQLASPTPPRSPFAPLS--SLLNHISRLKPSVDPFSAHDIDH-------------LS 384
+ + SP + SLL+ S + + + F+ +ID L
Sbjct: 330 AAGAKCTDDESEGSPDIAMGEPSLLHDFSHVPMADENFTGSEIDRETPNLGARAADHTLD 389
Query: 385 TTKYSPVHKMNQELNVVGSAKPSN--ELNDHITKDAVAIGETNAVPD--TLRNCASASEN 440
++ + VG + + + N+ + +++ E + D T+ S +E
Sbjct: 390 PEPSDHAYERQPGGSSVGLCRDTEVAKENEACRRSNISMEEDDVPIDYPTIGRSTSETEA 449
Query: 441 SKEDNSGKSSNKLDA--------PLIEDILAVSESCLIEDAVMNSTSTSQMPMEDNSMEP 492
S G+S+ +L I+ + V E +++D S+ + +MP+ED ++P
Sbjct: 450 SSHHLEGRSTEELGINRTLHTAEDSIQHLEVVKEGGVLQD---KSSQSLEMPLED--IDP 504
Query: 493 EFDANVDRNEPHADMDVDNGGS----------GMGETVMDDTVAKPNIEVNEQCQFEGKI 542
N+P +GGS + T A ++ +Q + K+
Sbjct: 505 V-------NQPQM-----HGGSTKKLAPDLCNALSLTKQKKQQAAQEGKMKKQSKRGKKV 552
Query: 543 LTENTDAFTASMPT---------DDTDIDVVNPLADQSSP-------GVIQANSIDKRTT 586
E++ A S DD +I+ L+ SP G +AN K
Sbjct: 553 ADESSHAPEISQANLDSENQPHNDDVNIEQQKVLSSTLSPNHAKGQKGAQRANKTKKLNQ 612
Query: 587 CTNEGSEQCL-----QEERDGSRAP------VEQKRVKSRS------------QKDTKSK 623
G E L Q D P +EQ+ V S + Q+ K+K
Sbjct: 613 RKILGDESSLALEISQANLDSENQPHNDDLNIEQQTVLSSTLSPNHDKGQKGAQRTNKTK 672
Query: 624 RLAKRQSLAAAGTSWTSGIRRSTRIRTRPLEYWKGERLVYGRIHQS 669
+L +R+ L AG + SG+RRSTR R+RPLE+W GERL+YG I+ +
Sbjct: 673 KLNQRKILGDAGLAQPSGVRRSTRTRSRPLEHWLGERLLYGPINDT 718
>UniRef100_Q66LH5 CENP-C [Sorghum bicolor]
Length = 694
Score = 89.4 bits (220), Expect = 3e-16
Identities = 149/657 (22%), Positives = 267/657 (39%), Gaps = 87/657 (13%)
Query: 53 SMALRSPAKLVEQTKAILEENPGLFNSQDDERNDVVDVDVDVAEDGQDFPRKRRPGLGLK 112
S +L+ +LV+Q K +L+E+ + D+ V V +G + RP L K
Sbjct: 45 SRSLKGSEELVKQAKMVLKEHGDIQPLYHDD-----GVQVRPPVNGSKEQQGGRPALNRK 99
Query: 113 RARFSLKPSTSVSLESLLPNLDIENLKDPVEFFKAHERMENARREIQKQMGVSVEQNQDD 172
RARF++K + S ++ + N+ DPVE+F +++E A+ EIQ+ G + +
Sbjct: 100 RARFTMKDTASKPTP-VVDQSKLTNISDPVEYFMTLDQLEEAQEEIQRLNGAAEKCVLKF 158
Query: 173 VSTRPRQRRPGLRGNDQ-RSIKYRHRYSTEISDKNDHAMASQEAFGSDSLDPDTEKIENG 231
+R+PGL G RS K T+ + + + S D +
Sbjct: 159 DPVDQPKRQPGLCGRKSIRSFKIIGDADTQDPIEVPASQTTTLTGSQVSQDVVHAVADKN 218
Query: 232 EACLTSLENEVT---DSSATEGN---KICEILDGLLRCNSEDLEGD-----GAMDLLQER 280
E C+ S E + S T+ + +L+ L + + EG G + +ER
Sbjct: 219 EQCVRSSSAEAISGKEDSLTQKDGRDDFTYLLNSLQHLDESEEEGFILKTLGIGEPRKER 278
Query: 281 LQIKPVVL----------EKVSVPDFPDNQPIDLKSLRGSLSKPRKALSNIDNLLNRRKN 330
+ + + +K S+ P P+ L+ L+ +S+ K L
Sbjct: 279 VSFRNSIPGVRSIRSNNEQKGSMRAHPLESPL-LQPLQDRISELEKHLFPG----GAADA 333
Query: 331 KTPLRQDAGSPAEQLASPTPPRS----PFAPLSSLLNHISRLKPSVDPFSAHDIDHLSTT 386
K + GSP + P+ P + + I R P++ A DH+
Sbjct: 334 KCTDDESEGSPDIAMGEPSLVHDSSDVPMVDENFTGSEIDRETPNL---GARAADHILDP 390
Query: 387 KYSP-VHKMNQELNVVGSAKPSN--ELNDHITKDAVAIGETNAVPD--TLRNCASASENS 441
+ S ++ + VG + + + N+ + +++ E + D T+ S +E S
Sbjct: 391 EPSDHAYERQPGGSSVGLCRDTEVAKENEACRRSNISMEEDDVPIDYPTIGRSTSETEAS 450
Query: 442 KEDNSGKSSNKLDA--------PLIEDILAVSESCLIEDAVMNSTSTSQMPMEDNSMEPE 493
G+S+ +L I+ + V E +++D S+ + +MP+ED ++P
Sbjct: 451 SHHLEGRSTEELGINRTLHTAEDSIQHLEVVKEGGVLQD---KSSQSLEMPLED--IDPV 505
Query: 494 FDANVDRNEPHADMDVDNGGSGMGETVMDDTVAKPNIEVNEQCQFEGKILTENTDAFTAS 553
N+P +GGS + +Q EGK+ ++
Sbjct: 506 -------NQPQM-----HGGSTKKLAPDLCNALSLTKQKKQQAAQEGKMKKQSKRG---- 549
Query: 554 MPTDDTDIDVVNPLADQSSPGV-IQANSIDKRTTCTNEGSEQCLQEERDGSRAPVEQKRV 612
+AD+SS + I ++D N+ + +P K
Sbjct: 550 -----------KKVADESSLALEISQANLDSENQPHNDDQNIEQHTVLSSTLSPNHDKGQ 598
Query: 613 KSRSQKDTKSKRLAKRQSLAAAGTSWTSGIRRSTRIRTRPLEYWKGERLVYGRIHQS 669
K +Q+ K+K+L +R+ L AG + SG+RRSTR R+RPLE+W GERL+YG I+ +
Sbjct: 599 KG-AQRTNKTKKLNQRKILGDAGLAQPSGVRRSTRTRSRPLEHWLGERLLYGPINDT 654
>UniRef100_Q9XHE1 CENPCA protein [Zea mays]
Length = 701
Score = 86.7 bits (213), Expect = 2e-15
Identities = 144/633 (22%), Positives = 243/633 (37%), Gaps = 113/633 (17%)
Query: 97 DGQDFPRKRRPGLGLKRARFSLKPSTSVSLESLLPNLDIENLKDPVEFFKAHERMENARR 156
+G + RRP L KRARF++K + S + ++ + N+ DP+ FF +R+E A
Sbjct: 82 NGSKEQQGRRPALDRKRARFAMKDTGSKPVP-VVDQSKLSNISDPITFFMTLDRLEEAEE 140
Query: 157 EIQKQMGVSVEQNQDDVSTRPRQRRPGLRGNDQ-RSIKYRHRYSTEISDKNDHAMASQEA 215
EI++ G + ++ + R+PGLRG RS K T+ D N+ +
Sbjct: 141 EIKRLNGEAEKRTLNFDPVDEPIRQPGLRGRKSVRSFKVIEDVGTQ--DPNEAPASQTAT 198
Query: 216 FGSDSLDPDTEKI---ENGEACLTSLENEVTDSSATEGNKICEILDGLLRCNSEDLEGDG 272
L D +NG + + +++ + K + + +DL+
Sbjct: 199 MTGSQLSQDVMHAVAGKNGRSVSSRSSEAISEKEVSLAEKDGRDDLTYILTSIQDLDESE 258
Query: 273 AMDLLQERLQIKPVVLEKVSVPDFPDNQPIDLKSLRGSLS-----KPRKALSNIDNLLNR 327
+ +++ L IK E+VS+ N + ++SLR + K S++
Sbjct: 259 EEEFIRKTLGIKETRKERVSL----RNSILGVRSLRSNTEQKGSMKVHPPESHLPQTRQD 314
Query: 328 RKNKTPLRQDAGSPAEQLASPTPPRSPFAP-----LSSLLNHISRLKPSVDPFSAHDID- 381
R ++ G A A T S +P SL+++ S + + + F+A D+D
Sbjct: 315 RISELEKHLFPGGAAN--AKCTDDESEGSPDIVMGEPSLVHYSSDVLMTDENFTASDVDR 372
Query: 382 ---HLSTTKYSPVHKMNQELNVVGSAKPSNELNDHITKDAVAIGETNAVPDTLRNCASAS 438
+L PV P L DH + DT A
Sbjct: 373 ETPNLGARAADPV------------LDPEPNLPDHAYERQPGGSSLGLYRDT--EVGVAK 418
Query: 439 ENSKEDNSGKSSNKLDAPLIEDILAVSESCLIEDAVMNSTS-TSQMPMEDNSMEPEFDAN 497
EN + S S + D P+ D + + S N T +S P+E +S E
Sbjct: 419 ENETCNRSNISVEEDDVPI--DYITIGRS-------TNETEVSSSHPLEGSSTE----EL 465
Query: 498 VDRNEPHADMDVDNGGSGMGETVMDDTVAKPNIEVNEQCQFEGKILTENTDAFTASMPTD 557
V + HA D G+ T A+ +I+ E + EG +L + + MP +
Sbjct: 466 VTKPGRHAAPD------GICRT---SHAAEDSIQHLEAVK-EGGVLQDKSSQL-LEMPQE 514
Query: 558 DTDIDVVNPLADQSSPGVIQANSIDKRTTCTNEGSEQCLQEER-----------DGSRAP 606
D +NPL G S + T + +Q +QE + G
Sbjct: 515 D-----INPLNQAQMHGGSTKKSAPDLSP-TKQKKQQAVQERKRKQQSKRGKKVAGESLE 568
Query: 607 VEQKRVKSR------------------------------SQKDTKSKRLAKRQSLAAAGT 636
+ Q +S +Q+ ++K+ +R+ L A
Sbjct: 569 IPQTNFESENQPHNDDVNIEKQTITSSTLSPNGAKGQKGAQRRNQTKKRNQRKILGDADL 628
Query: 637 SWTSGIRRSTRIRTRPLEYWKGERLVYGRIHQS 669
+ G+R+S+R R+RPLEYW GERL+YG IH +
Sbjct: 629 ACQPGVRKSSRTRSRPLEYWLGERLLYGPIHDN 661
>UniRef100_Q66LG8 CENP-C [Lycopersicon esculentum]
Length = 138
Score = 72.8 bits (177), Expect = 3e-11
Identities = 36/71 (50%), Positives = 48/71 (66%), Gaps = 7/71 (9%)
Query: 605 APVEQKRVKSRSQKDTK-------SKRLAKRQSLAAAGTSWTSGIRRSTRIRTRPLEYWK 657
+P K+ K + QK + K L+ R SLA AGTS+ SG+RRS R++TRPLEYWK
Sbjct: 31 SPQHHKQAKDKQQKAKELAVGRRERKHLSSRPSLADAGTSFESGVRRSKRMKTRPLEYWK 90
Query: 658 GERLVYGRIHQ 668
GERL+YGR+ +
Sbjct: 91 GERLLYGRVDE 101
>UniRef100_Q66LG7 CENP-C [Triticum aestivum]
Length = 234
Score = 72.0 bits (175), Expect = 5e-11
Identities = 38/103 (36%), Positives = 63/103 (60%), Gaps = 4/103 (3%)
Query: 569 DQSSPGVIQANSIDKRTTCTNEGSEQCLQEERDGSRAP--VEQKRVKSRSQKDTKSKRLA 626
+ S+P I + D T + + + ++++ +R P + +++ Q+ K K +
Sbjct: 94 EASNPLGIPPENCDTETQPSMQDTN--IEQQTVDTRVPRSPNKGKLQKEGQRRKKRKEVN 151
Query: 627 KRQSLAAAGTSWTSGIRRSTRIRTRPLEYWKGERLVYGRIHQS 669
+R+SL A G +W SG+RRSTRIR+RPL+ W GERL+YGRIH +
Sbjct: 152 RRKSLTAFGLAWQSGVRRSTRIRSRPLQDWLGERLLYGRIHDT 194
>UniRef100_Q68BI6 Centromere protein C homolog [Arabidopsis thaliana]
Length = 318
Score = 71.6 bits (174), Expect = 7e-11
Identities = 33/62 (53%), Positives = 45/62 (72%)
Query: 608 EQKRVKSRSQKDTKSKRLAKRQSLAAAGTSWTSGIRRSTRIRTRPLEYWKGERLVYGRIH 667
E++ K + + K + R+SLAAAGT G+RRSTRI++RPLEYW+GER +YGRIH
Sbjct: 215 EREEKKPKKTLTHEGKLFSCRKSLAAAGTKIEGGVRRSTRIKSRPLEYWRGERFLYGRIH 274
Query: 668 QS 669
+S
Sbjct: 275 ES 276
>UniRef100_Q5ZDI9 CENPCA protein-like [Oryza sativa]
Length = 618
Score = 70.5 bits (171), Expect = 2e-10
Identities = 30/55 (54%), Positives = 43/55 (77%)
Query: 613 KSRSQKDTKSKRLAKRQSLAAAGTSWTSGIRRSTRIRTRPLEYWKGERLVYGRIH 667
K+ +K K + L +R+SLA AG +W +G+RRSTRIR++PL++W GER +YGRIH
Sbjct: 522 KNEVRKRNKKQDLNRRKSLADAGLTWQAGVRRSTRIRSKPLQHWLGERFIYGRIH 576
>UniRef100_Q66LI2 CENP-C [Oryza sativa]
Length = 490
Score = 70.5 bits (171), Expect = 2e-10
Identities = 30/55 (54%), Positives = 43/55 (77%)
Query: 613 KSRSQKDTKSKRLAKRQSLAAAGTSWTSGIRRSTRIRTRPLEYWKGERLVYGRIH 667
K+ +K K + L +R+SLA AG +W +G+RRSTRIR++PL++W GER +YGRIH
Sbjct: 394 KNEVRKRNKKQDLNRRKSLADAGLTWQAGVRRSTRIRSKPLQHWLGERFIYGRIH 448
>UniRef100_Q66LH6 CENP-C [Oryza sativa]
Length = 117
Score = 70.5 bits (171), Expect = 2e-10
Identities = 30/55 (54%), Positives = 43/55 (77%)
Query: 613 KSRSQKDTKSKRLAKRQSLAAAGTSWTSGIRRSTRIRTRPLEYWKGERLVYGRIH 667
K+ +K K + L +R+SLA AG +W +G+RRSTRIR++PL++W GER +YGRIH
Sbjct: 21 KNEVRKRNKKQDLNRRKSLADAGLTWQAGVRRSTRIRSKPLQHWLGERFIYGRIH 75
>UniRef100_Q9XHE0 CENPCB protein [Zea mays]
Length = 451
Score = 62.4 bits (150), Expect = 4e-08
Identities = 70/271 (25%), Positives = 118/271 (42%), Gaps = 71/271 (26%)
Query: 407 SNELNDHITKDAVAIGETNAVPDTLRNCASASENSKEDNSGKSSNKLDAPLIEDILAVSE 466
S++L +T++ V+ +A PD + + A+E+S I+ + V +
Sbjct: 204 SHDLEGSLTEELVSEPGRHAAPDDIYRTSHAAEDS----------------IQHLEVVKQ 247
Query: 467 SCLIEDAVMNSTSTSQMPMEDNSMEPEFDANVDRNEPHADMDVDNGGSGMGETVMDDTVA 526
+++D S+ + +MP+ED ++P + R + H GGS T +
Sbjct: 248 GGVLQD---KSSQSLEMPLED--IDPLY-----RPQMH-------GGSTKKSTPVLCNAL 290
Query: 527 KPNIEVNEQCQFEGKILTENTDAFTASMPTDDTDIDVVNPLADQSSPGVIQANSIDKRTT 586
P IE +Q EGK + V ++D+SS +
Sbjct: 291 SP-IEQKQQAAQEGKRKQQAKR---------------VKKVSDESSHAL----------- 323
Query: 587 CTNEGSEQCLQEERDGSRAPV--EQKRVKSRS------QKDTKSKRLAKRQSLAAAGTSW 638
E S+ + E + V EQ V S + Q+ K+K+L +R+ L +
Sbjct: 324 ---EISQANFEPENEPHNVDVNMEQPTVMSSAKGQKGAQRRNKTKKLNQRKILGDVDLAR 380
Query: 639 TSGIRRSTRIRTRPLEYWKGERLVYGRIHQS 669
SG+RRS+RIR+RPLEYW GERL+YG + +
Sbjct: 381 QSGVRRSSRIRSRPLEYWLGERLLYGPVQDT 411
>UniRef100_Q9XHD9 CENPCC protein [Zea mays]
Length = 440
Score = 55.8 bits (133), Expect = 4e-06
Identities = 25/54 (46%), Positives = 38/54 (70%)
Query: 616 SQKDTKSKRLAKRQSLAAAGTSWTSGIRRSTRIRTRPLEYWKGERLVYGRIHQS 669
+Q+ ++K+ +R+ L A + G+R+S+R R+RPLEYW GERL+YG IH S
Sbjct: 371 AQRRNQTKKRNQRKILGDADLACQPGVRKSSRTRSRPLEYWLGERLLYGPIHDS 424
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.309 0.128 0.354
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,128,082,480
Number of Sequences: 2790947
Number of extensions: 49090120
Number of successful extensions: 128756
Number of sequences better than 10.0: 417
Number of HSP's better than 10.0 without gapping: 44
Number of HSP's successfully gapped in prelim test: 378
Number of HSP's that attempted gapping in prelim test: 128277
Number of HSP's gapped (non-prelim): 778
length of query: 677
length of database: 848,049,833
effective HSP length: 134
effective length of query: 543
effective length of database: 474,062,935
effective search space: 257416173705
effective search space used: 257416173705
T: 11
A: 40
X1: 16 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.7 bits)
S2: 78 (34.7 bits)
Lotus: description of TM0253.9


