

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0252a.5
(150 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
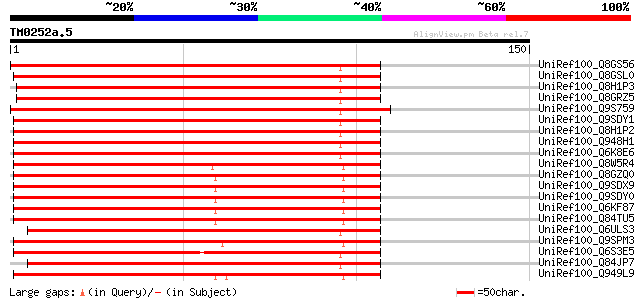
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q8GS56 Phosphoenolpyruvate carboxylase kinase [Lotus j... 188 2e-47
UniRef100_Q8GSL0 Phosphoenolpyruvate carboxylase kinase [Glycine... 165 2e-40
UniRef100_Q8H1P3 Phosphoenolpyruvate carboxylase kinase [Glycine... 161 3e-39
UniRef100_Q8GRZ5 Phosphoenolpyruvate carboxylase kinase [Glycine... 161 3e-39
UniRef100_Q9S759 Phosphoenolpyruvate carboxylase kinase [Kalanch... 152 1e-36
UniRef100_Q9SDY1 Phosphoenolpyruvate carboxylase kinase [Glycine... 147 5e-35
UniRef100_Q8H1P2 Phosphoenolpyruvate carboxylase kinase [Glycine... 147 5e-35
UniRef100_Q948H1 Putative phosphoenolpyruvate kinase [Oryza sativa] 144 4e-34
UniRef100_Q6K8E6 Putative phosphoenolpyruvate carboxylase kinase... 144 4e-34
UniRef100_Q8W5R4 Phosphoenolpyruvate carboxylase kinase [Flaveri... 140 6e-33
UniRef100_Q8GZQ0 PEP carboxylase kinase [Solanum tuberosum] 139 1e-32
UniRef100_Q9SDX9 Phosphoenolpyruvate carboxylase kinase 1 [Lycop... 139 1e-32
UniRef100_Q9SDY0 Phosphoenolpyruvate carboxylase kinase [Lycoper... 139 1e-32
UniRef100_Q6KF87 PEPC kinase 1b [Solanum tuberosum] 139 1e-32
UniRef100_Q84TU5 Phosphoenolpyruvate carboxylase kinase 1 [Solan... 136 8e-32
UniRef100_Q6ULS3 Phosphoenolpyruvate carboxylase kinase 4 [Glyci... 134 3e-31
UniRef100_Q9SPM3 Phosphoenolpyruvate carboxylase-kinase [Mesembr... 134 4e-31
UniRef100_Q6S3E5 Phosphoenolpyruvate carboxylase kinase 1 [Clusi... 132 1e-30
UniRef100_Q84JP7 Phosphoenolpyruvate carboxylase kinase 2 [Lycop... 131 3e-30
UniRef100_Q949L9 Putative phosphoenolpyruvate carboxylase kinase... 131 3e-30
>UniRef100_Q8GS56 Phosphoenolpyruvate carboxylase kinase [Lotus japonicus]
Length = 277
Score = 188 bits (478), Expect = 2e-47
Identities = 96/126 (76%), Positives = 97/126 (76%), Gaps = 19/126 (15%)
Query: 1 GGPIPEVEVASLMKQLLEAITHCHRLGVAHHDVKPDNVLFGAGGDLKLADFGSAEWFGDG 60
G IPEVE A LMKQLLEA+ HCHRLGVAH DVKPDNVLFG GGDLKLADFGSAEWFGDG
Sbjct: 104 GTSIPEVEAAGLMKQLLEAVAHCHRLGVAHRDVKPDNVLFGGGGDLKLADFGSAEWFGDG 163
Query: 61 RRMSGVVGTPYYVAPEVLMGWENGEKVDVWSCGV-------------------IFEAVIW 101
RRMSGVVGTPYYVAPEVLMG E GEKVDVWSCGV IFEAVI
Sbjct: 164 RRMSGVVGTPYYVAPEVLMGREYGEKVDVWSCGVILYIMLSGTPPFYGDSAAEIFEAVIR 223
Query: 102 GNLRFP 107
GNLRFP
Sbjct: 224 GNLRFP 229
>UniRef100_Q8GSL0 Phosphoenolpyruvate carboxylase kinase [Glycine max]
Length = 274
Score = 165 bits (418), Expect = 2e-40
Identities = 82/125 (65%), Positives = 91/125 (72%), Gaps = 19/125 (15%)
Query: 2 GPIPEVEVASLMKQLLEAITHCHRLGVAHHDVKPDNVLFGAGGDLKLADFGSAEWFGDGR 61
GPI E + A+LMK LLEA+ HCHRLGVAH D+KPDN+LF + +LKLADFGSAEWFGDGR
Sbjct: 102 GPIQESQAAALMKNLLEAVAHCHRLGVAHRDIKPDNILFDSADNLKLADFGSAEWFGDGR 161
Query: 62 RMSGVVGTPYYVAPEVLMGWENGEKVDVWSCGV-------------------IFEAVIWG 102
MSGVVGTPYYVAPEVL+G E EKVDVWSCGV IFEAV+
Sbjct: 162 SMSGVVGTPYYVAPEVLLGREYDEKVDVWSCGVILYIMLAGIPPFYGDSAAEIFEAVVRA 221
Query: 103 NLRFP 107
NLRFP
Sbjct: 222 NLRFP 226
>UniRef100_Q8H1P3 Phosphoenolpyruvate carboxylase kinase [Glycine max]
Length = 274
Score = 161 bits (407), Expect = 3e-39
Identities = 80/124 (64%), Positives = 89/124 (71%), Gaps = 19/124 (15%)
Query: 3 PIPEVEVASLMKQLLEAITHCHRLGVAHHDVKPDNVLFGAGGDLKLADFGSAEWFGDGRR 62
P E + ASL+K LLEA+ HCHRLGVAH D+KPDN+LF + +LKLADFGSAEWFGDGR
Sbjct: 103 PFSESQAASLIKNLLEAVAHCHRLGVAHRDIKPDNILFDSADNLKLADFGSAEWFGDGRS 162
Query: 63 MSGVVGTPYYVAPEVLMGWENGEKVDVWSCGV-------------------IFEAVIWGN 103
MSGVVGTPYYVAPEVL+G E EKVDVWSCGV IFEAV+ N
Sbjct: 163 MSGVVGTPYYVAPEVLLGREYDEKVDVWSCGVILYIMLAGIPPFYGDSAAEIFEAVVRAN 222
Query: 104 LRFP 107
LRFP
Sbjct: 223 LRFP 226
>UniRef100_Q8GRZ5 Phosphoenolpyruvate carboxylase kinase [Glycine max]
Length = 274
Score = 161 bits (407), Expect = 3e-39
Identities = 80/124 (64%), Positives = 89/124 (71%), Gaps = 19/124 (15%)
Query: 3 PIPEVEVASLMKQLLEAITHCHRLGVAHHDVKPDNVLFGAGGDLKLADFGSAEWFGDGRR 62
P E + ASL+K LLEA+ HCHRLGVAH D+KPDN+LF + +LKLADFGSAEWFGDGR
Sbjct: 103 PFSESQAASLIKNLLEAVAHCHRLGVAHRDIKPDNILFDSADNLKLADFGSAEWFGDGRS 162
Query: 63 MSGVVGTPYYVAPEVLMGWENGEKVDVWSCGV-------------------IFEAVIWGN 103
MSGVVGTPYYVAPEVL+G E EKVDVWSCGV IFEAV+ N
Sbjct: 163 MSGVVGTPYYVAPEVLLGREYDEKVDVWSCGVILYIMLAGIPPFYGDSAAEIFEAVVKAN 222
Query: 104 LRFP 107
LRFP
Sbjct: 223 LRFP 226
>UniRef100_Q9S759 Phosphoenolpyruvate carboxylase kinase [Kalanchoe fedtschenkoi]
Length = 274
Score = 152 bits (384), Expect = 1e-36
Identities = 78/129 (60%), Positives = 86/129 (66%), Gaps = 19/129 (14%)
Query: 1 GGPIPEVEVASLMKQLLEAITHCHRLGVAHHDVKPDNVLFGAGGDLKLADFGSAEWFGDG 60
GG +PE E + + L+E I HCHR GV H DVKP+NVLF + G LKLADFGSAEWFGDG
Sbjct: 102 GGAVPEQEASVIAASLMEGIAHCHRRGVCHRDVKPENVLFDSVGRLKLADFGSAEWFGDG 161
Query: 61 RRMSGVVGTPYYVAPEVLMGWENGEKVDVWSCGV-------------------IFEAVIW 101
R MSGVVGTPYYVAPEVL G E EKVDVWS GV IFEAV+
Sbjct: 162 RGMSGVVGTPYYVAPEVLQGREYNEKVDVWSAGVILYTMLAGFPPFYGETAQDIFEAVMR 221
Query: 102 GNLRFPLES 110
GNLRFP +
Sbjct: 222 GNLRFPFRA 230
>UniRef100_Q9SDY1 Phosphoenolpyruvate carboxylase kinase [Glycine max]
Length = 274
Score = 147 bits (371), Expect = 5e-35
Identities = 73/125 (58%), Positives = 85/125 (67%), Gaps = 19/125 (15%)
Query: 2 GPIPEVEVASLMKQLLEAITHCHRLGVAHHDVKPDNVLFGAGGDLKLADFGSAEWFGDGR 61
GP+ E ASL+KQLLEA+ HCH G+AH D+KP+N+LF G LKL+DFGSAEW G+G
Sbjct: 97 GPLTEPHAASLLKQLLEAVAHCHAQGLAHRDIKPENILFDEGNKLKLSDFGSAEWLGEGS 156
Query: 62 RMSGVVGTPYYVAPEVLMGWENGEKVDVWSCGV-------------------IFEAVIWG 102
MSGVVGTPYYVAPEV+MG E EKVDVWS GV IFE+V+
Sbjct: 157 SMSGVVGTPYYVAPEVIMGREYDEKVDVWSSGVILYAMLAGFPPFYGESAPEIFESVLRA 216
Query: 103 NLRFP 107
NLRFP
Sbjct: 217 NLRFP 221
>UniRef100_Q8H1P2 Phosphoenolpyruvate carboxylase kinase [Glycine max]
Length = 282
Score = 147 bits (371), Expect = 5e-35
Identities = 73/125 (58%), Positives = 85/125 (67%), Gaps = 19/125 (15%)
Query: 2 GPIPEVEVASLMKQLLEAITHCHRLGVAHHDVKPDNVLFGAGGDLKLADFGSAEWFGDGR 61
GP+ E ASL+KQLLEA+ HCH G+AH D+KP+N+LF G LKL+DFGSAEW G+G
Sbjct: 105 GPLTEPHAASLLKQLLEAVAHCHAQGLAHRDIKPENILFDEGNKLKLSDFGSAEWLGEGS 164
Query: 62 RMSGVVGTPYYVAPEVLMGWENGEKVDVWSCGV-------------------IFEAVIWG 102
MSGVVGTPYYVAPEV+MG E EKVDVWS GV IFE+V+
Sbjct: 165 SMSGVVGTPYYVAPEVIMGREYDEKVDVWSSGVILYAMLAGFPPFYGESAPEIFESVLRA 224
Query: 103 NLRFP 107
NLRFP
Sbjct: 225 NLRFP 229
>UniRef100_Q948H1 Putative phosphoenolpyruvate kinase [Oryza sativa]
Length = 369
Score = 144 bits (363), Expect = 4e-34
Identities = 73/125 (58%), Positives = 83/125 (66%), Gaps = 19/125 (15%)
Query: 2 GPIPEVEVASLMKQLLEAITHCHRLGVAHHDVKPDNVLFGAGGDLKLADFGSAEWFGDGR 61
G +PE E A L+ QL A+ CHR GVAH DVKPDN+LF GG LKL DFGSA WFGDGR
Sbjct: 117 GRLPEHEAADLVAQLASALASCHRRGVAHRDVKPDNLLFDGGGVLKLGDFGSAGWFGDGR 176
Query: 62 RMSGVVGTPYYVAPEVLMGWENGEKVDVWSCGV-------------------IFEAVIWG 102
M+G+VGTPYYVAPEV+ G E GEKVDVWS GV +F+ V+ G
Sbjct: 177 PMTGLVGTPYYVAPEVVAGREYGEKVDVWSAGVVLYMMLSGTLPFYGATAAEVFQCVLRG 236
Query: 103 NLRFP 107
NLRFP
Sbjct: 237 NLRFP 241
>UniRef100_Q6K8E6 Putative phosphoenolpyruvate carboxylase kinase [Oryza sativa]
Length = 289
Score = 144 bits (363), Expect = 4e-34
Identities = 73/125 (58%), Positives = 83/125 (66%), Gaps = 19/125 (15%)
Query: 2 GPIPEVEVASLMKQLLEAITHCHRLGVAHHDVKPDNVLFGAGGDLKLADFGSAEWFGDGR 61
G +PE E A L+ QL A+ CHR GVAH DVKPDN+LF GG LKL DFGSA WFGDGR
Sbjct: 117 GRLPEHEAADLVAQLASALASCHRRGVAHRDVKPDNLLFDGGGVLKLGDFGSAGWFGDGR 176
Query: 62 RMSGVVGTPYYVAPEVLMGWENGEKVDVWSCGV-------------------IFEAVIWG 102
M+G+VGTPYYVAPEV+ G E GEKVDVWS GV +F+ V+ G
Sbjct: 177 PMTGLVGTPYYVAPEVVAGREYGEKVDVWSAGVVLYMMLSGTLPFYGATAAEVFQCVLRG 236
Query: 103 NLRFP 107
NLRFP
Sbjct: 237 NLRFP 241
>UniRef100_Q8W5R4 Phosphoenolpyruvate carboxylase kinase [Flaveria trinervia]
Length = 281
Score = 140 bits (353), Expect = 6e-33
Identities = 74/127 (58%), Positives = 84/127 (65%), Gaps = 21/127 (16%)
Query: 2 GPIPEVEVASLMKQLLEAITHCHRLGVAHHDVKPDNVLFGAGGDLKLADFGSAEWF--GD 59
G E E A++ L+ AI++CHRLG+AH D+KPDNVLF + G LKLADFGSAEWF D
Sbjct: 104 GVFSETEAATVFAPLMSAISYCHRLGIAHRDLKPDNVLFDSRGGLKLADFGSAEWFAMND 163
Query: 60 GRRMSGVVGTPYYVAPEVLMGWENGEKVDVWSCGVI-------------------FEAVI 100
R MSG+VGTPYYVAPEVL G E EKVDVWS GVI FEAV+
Sbjct: 164 RRTMSGIVGTPYYVAPEVLSGMEYNEKVDVWSAGVILYIMLAGVPPFHGDSPADTFEAVL 223
Query: 101 WGNLRFP 107
GNLRFP
Sbjct: 224 RGNLRFP 230
>UniRef100_Q8GZQ0 PEP carboxylase kinase [Solanum tuberosum]
Length = 279
Score = 139 bits (351), Expect = 1e-32
Identities = 73/127 (57%), Positives = 86/127 (67%), Gaps = 21/127 (16%)
Query: 2 GPIPEVEVASLMKQLLEAITHCHRLGVAHHDVKPDNVLFGAGGDLKLADFGSAEWFG--D 59
G + E A+++ QL+ AI++CH +GVAH D+KPDNVLF + LKLADFGSAEWF +
Sbjct: 102 GSLSESAAAAILTQLVSAISYCHHMGVAHRDIKPDNVLFDSENRLKLADFGSAEWFAGCE 161
Query: 60 GRRMSGVVGTPYYVAPEVLMGWENGEKVDVWSCGVI-------------------FEAVI 100
GR MSGVVGTPYYVAPEVLMG E EKVDVWS GVI F+AV+
Sbjct: 162 GRMMSGVVGTPYYVAPEVLMGKEYNEKVDVWSAGVILYIMLSGVPPFYGETPTETFQAVL 221
Query: 101 WGNLRFP 107
GNLRFP
Sbjct: 222 RGNLRFP 228
>UniRef100_Q9SDX9 Phosphoenolpyruvate carboxylase kinase 1 [Lycopersicon esculentum]
Length = 279
Score = 139 bits (351), Expect = 1e-32
Identities = 73/127 (57%), Positives = 86/127 (67%), Gaps = 21/127 (16%)
Query: 2 GPIPEVEVASLMKQLLEAITHCHRLGVAHHDVKPDNVLFGAGGDLKLADFGSAEWFG--D 59
G + E A+++ QL+ AI++CH +GVAH D+KPDNVLF + LKLADFGSAEWF +
Sbjct: 102 GSLSESAAAAILTQLVSAISYCHHMGVAHRDIKPDNVLFDSENRLKLADFGSAEWFAGCE 161
Query: 60 GRRMSGVVGTPYYVAPEVLMGWENGEKVDVWSCGVI-------------------FEAVI 100
GR MSGVVGTPYYVAPEVLMG E EKVDVWS GVI F+AV+
Sbjct: 162 GRMMSGVVGTPYYVAPEVLMGKEYNEKVDVWSAGVILYIMLSGVPPFYGETPTETFQAVL 221
Query: 101 WGNLRFP 107
GNLRFP
Sbjct: 222 RGNLRFP 228
>UniRef100_Q9SDY0 Phosphoenolpyruvate carboxylase kinase [Lycopersicon esculentum]
Length = 276
Score = 139 bits (350), Expect = 1e-32
Identities = 73/127 (57%), Positives = 86/127 (67%), Gaps = 21/127 (16%)
Query: 2 GPIPEVEVASLMKQLLEAITHCHRLGVAHHDVKPDNVLFGAGGDLKLADFGSAEWFG--D 59
G + E A+++ QL+ AI++CH +GVAH D+KPDNVLF + LKLADFGSAEWF +
Sbjct: 99 GSLSESAAAAILTQLVSAISYCHHMGVAHRDIKPDNVLFDSENRLKLADFGSAEWFAGCE 158
Query: 60 GRRMSGVVGTPYYVAPEVLMGWENGEKVDVWSCGVI-------------------FEAVI 100
GR MSGVVGTPYYVAPEVLMG E EKVDVWS GVI F+AV+
Sbjct: 159 GRLMSGVVGTPYYVAPEVLMGKEYNEKVDVWSAGVILYIMLSGVPPFYGETPTETFQAVL 218
Query: 101 WGNLRFP 107
GNLRFP
Sbjct: 219 RGNLRFP 225
>UniRef100_Q6KF87 PEPC kinase 1b [Solanum tuberosum]
Length = 277
Score = 139 bits (350), Expect = 1e-32
Identities = 73/127 (57%), Positives = 86/127 (67%), Gaps = 21/127 (16%)
Query: 2 GPIPEVEVASLMKQLLEAITHCHRLGVAHHDVKPDNVLFGAGGDLKLADFGSAEWFG--D 59
G + E A+++ QL+ AI++CH +GVAH D+KPDNVLF + LKLADFGSAEWF +
Sbjct: 102 GSLSESAAAAILTQLVSAISYCHHMGVAHRDIKPDNVLFDSENRLKLADFGSAEWFAGCE 161
Query: 60 GRRMSGVVGTPYYVAPEVLMGWENGEKVDVWSCGVI-------------------FEAVI 100
GR MSGVVGTPYYVAPEVLMG E EKVDVWS GVI F+AV+
Sbjct: 162 GRMMSGVVGTPYYVAPEVLMGKEYNEKVDVWSVGVILYIMLSGVPPFYGETPTETFQAVL 221
Query: 101 WGNLRFP 107
GNLRFP
Sbjct: 222 RGNLRFP 228
>UniRef100_Q84TU5 Phosphoenolpyruvate carboxylase kinase 1 [Solanum tuberosum]
Length = 266
Score = 136 bits (343), Expect = 8e-32
Identities = 72/127 (56%), Positives = 85/127 (66%), Gaps = 21/127 (16%)
Query: 2 GPIPEVEVASLMKQLLEAITHCHRLGVAHHDVKPDNVLFGAGGDLKLADFGSAEWFG--D 59
G + E A+++ QL+ AI++CH +GVAH D+KPDNVLF + LKLADFGSAEWF +
Sbjct: 95 GSLSESAAAAILTQLVSAISYCHHMGVAHRDIKPDNVLFDSENRLKLADFGSAEWFAGCE 154
Query: 60 GRRMSGVVGTPYYVAPEVLMGWENGEKVDVWSCGVI-------------------FEAVI 100
GR MSGVVGTPYYVAPEVLMG E EKVDVWS VI F+AV+
Sbjct: 155 GRMMSGVVGTPYYVAPEVLMGKEYNEKVDVWSARVILYIMLSGVPPFYGETPTETFQAVL 214
Query: 101 WGNLRFP 107
GNLRFP
Sbjct: 215 RGNLRFP 221
>UniRef100_Q6ULS3 Phosphoenolpyruvate carboxylase kinase 4 [Glycine max]
Length = 270
Score = 134 bits (338), Expect = 3e-31
Identities = 71/121 (58%), Positives = 81/121 (66%), Gaps = 19/121 (15%)
Query: 6 EVEVASLMKQLLEAITHCHRLGVAHHDVKPDNVLFGAGGDLKLADFGSAEWFGDGRRMSG 65
E E AS+M QL++A+ HCHRLGVAH DVKPDN+LF LKLADFGSA+ F +G MSG
Sbjct: 105 EPEAASVMWQLMQAVAHCHRLGVAHRDVKPDNILFDEENRLKLADFGSADTFKEGEPMSG 164
Query: 66 VVGTPYYVAPEVLMGWENGEKVDVWSCGV-------------------IFEAVIWGNLRF 106
VVGTP+YVAPEVL G + EKVDVWS GV IFEAV+ NLRF
Sbjct: 165 VVGTPHYVAPEVLAGRDYNEKVDVWSAGVVLYQMLAGFLPFRGDSPVEIFEAVLRANLRF 224
Query: 107 P 107
P
Sbjct: 225 P 225
>UniRef100_Q9SPM3 Phosphoenolpyruvate carboxylase-kinase [Mesembryanthemum
crystallinum]
Length = 279
Score = 134 bits (337), Expect = 4e-31
Identities = 71/126 (56%), Positives = 80/126 (63%), Gaps = 20/126 (15%)
Query: 2 GPIPEVEVASLMKQLLEAITHCHRLGVAHHDVKPDNVLFGAGGDLKLADFGSAEWFGDG- 60
GP E + A++ QL EA+ HCHR VAH D+KPDN+LF + LKL DFGSAEWFG G
Sbjct: 108 GPFSEPDAAAIFCQLAEALAHCHRNYVAHRDIKPDNILFDSRNRLKLCDFGSAEWFGAGD 167
Query: 61 RRMSGVVGTPYYVAPEVLMGWENGEKVDVWSCGVI-------------------FEAVIW 101
R M GVVGTPYYVAPEVL G + EK DVWS GVI FEAV+
Sbjct: 168 REMRGVVGTPYYVAPEVLSGKDYNEKADVWSAGVILYIMLGGVPPFYGETVEETFEAVLR 227
Query: 102 GNLRFP 107
GNLRFP
Sbjct: 228 GNLRFP 233
>UniRef100_Q6S3E5 Phosphoenolpyruvate carboxylase kinase 1 [Clusia minor]
Length = 258
Score = 132 bits (333), Expect = 1e-30
Identities = 69/125 (55%), Positives = 83/125 (66%), Gaps = 20/125 (16%)
Query: 2 GPIPEVEVASLMKQLLEAITHCHRLGVAHHDVKPDNVLFGAGGDLKLADFGSAEWFGDGR 61
GPIPE + + QL++A++HCH+ GV H D+KPDN+LF + LKLADFGSAE G G
Sbjct: 93 GPIPEAQSKAQFVQLMKAVSHCHKSGVVHRDIKPDNILFDSRESLKLADFGSAE-CGKGS 151
Query: 62 RMSGVVGTPYYVAPEVLMGWENGEKVDVWSCGV-------------------IFEAVIWG 102
++GVVGTPYYVAPE+L G E GEKVDVWS GV IFEAV+ G
Sbjct: 152 DLNGVVGTPYYVAPEILEGREYGEKVDVWSAGVVLYVMLAGFPPFYGETVVEIFEAVLRG 211
Query: 103 NLRFP 107
NLRFP
Sbjct: 212 NLRFP 216
>UniRef100_Q84JP7 Phosphoenolpyruvate carboxylase kinase 2 [Lycopersicon esculentum]
Length = 278
Score = 131 bits (330), Expect = 3e-30
Identities = 67/121 (55%), Positives = 79/121 (64%), Gaps = 19/121 (15%)
Query: 6 EVEVASLMKQLLEAITHCHRLGVAHHDVKPDNVLFGAGGDLKLADFGSAEWFGDGRRMSG 65
E + +M L++AI HCHRLGVAH D+KPDN+LF +LKLADFGSAE F +G+ MSG
Sbjct: 108 ESDAVDVMVPLMKAIAHCHRLGVAHRDIKPDNILFTDSNELKLADFGSAECFHEGQLMSG 167
Query: 66 VVGTPYYVAPEVLMGWENGEKVDVWSCGV-------------------IFEAVIWGNLRF 106
VVGTPYYVAPEVL G EK+D+WS GV IFEAV+ NLRF
Sbjct: 168 VVGTPYYVAPEVLAGRNYSEKIDIWSAGVILYIMLAGVPPFFGDSASEIFEAVLRANLRF 227
Query: 107 P 107
P
Sbjct: 228 P 228
>UniRef100_Q949L9 Putative phosphoenolpyruvate carboxylase kinase [Beta vulgaris]
Length = 282
Score = 131 bits (329), Expect = 3e-30
Identities = 68/128 (53%), Positives = 82/128 (63%), Gaps = 22/128 (17%)
Query: 2 GPIPEVEVASLMKQLLEAITHCHRLGVAHHDVKPDNVLFGAGGDLKLADFGSAEWFG--D 59
GP E + S+ QL++A+ HCHR + H DVKPDN+LF + +KL DFGSAEWFG +
Sbjct: 104 GPFSETDACSVFTQLIDALAHCHRNYICHRDVKPDNILFDSRNRVKLCDFGSAEWFGGVE 163
Query: 60 GR-RMSGVVGTPYYVAPEVLMGWENGEKVDVWSCGVI-------------------FEAV 99
GR RM+G+VGTPYYVAPEVL G E EK+DVWS GVI F AV
Sbjct: 164 GRERMNGIVGTPYYVAPEVLAGREYNEKIDVWSAGVILYIMLGGVPPFYGDTVEDTFAAV 223
Query: 100 IWGNLRFP 107
+ GNLRFP
Sbjct: 224 LRGNLRFP 231
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.334 0.150 0.505
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 250,348,766
Number of Sequences: 2790947
Number of extensions: 10325553
Number of successful extensions: 66161
Number of sequences better than 10.0: 16849
Number of HSP's better than 10.0 without gapping: 12066
Number of HSP's successfully gapped in prelim test: 4784
Number of HSP's that attempted gapping in prelim test: 38645
Number of HSP's gapped (non-prelim): 18054
length of query: 150
length of database: 848,049,833
effective HSP length: 126
effective length of query: 24
effective length of database: 496,390,511
effective search space: 11913372264
effective search space used: 11913372264
T: 11
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.5 bits)
S2: 67 (30.4 bits)
Lotus: description of TM0252a.5


