

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
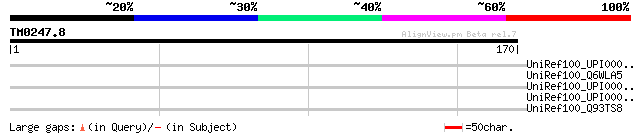
Query= TM0247.8
(170 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_UPI00002ECBC4 UPI00002ECBC4 UniRef100 entry 35 1.00
UniRef100_Q6WLA5 CAD [Leptopeza sp. NCSU-99071981] 33 2.2
UniRef100_UPI000042E2D4 UPI000042E2D4 UniRef100 entry 33 3.8
UniRef100_UPI0000309804 UPI0000309804 UniRef100 entry 32 8.4
UniRef100_Q93TS8 Dissimilatory sulfite reductase alpha subunit [... 32 8.4
>UniRef100_UPI00002ECBC4 UPI00002ECBC4 UniRef100 entry
Length = 304
Score = 34.7 bits (78), Expect = 1.00
Identities = 24/73 (32%), Positives = 36/73 (48%), Gaps = 6/73 (8%)
Query: 8 ISQTTSPTSLDTSPRVLAKLMSLKAYPKVKSILRQNTIIIKRVQPLTMTLRVINTLTKSH 67
+S + P + P L+ L + + P S +T+ QPLT+TL +TLT+SH
Sbjct: 37 VSPSRPPGLSASQPLSLSTLRTSDSQPL--SYSASHTLTCSHAQPLTLTLTHAHTLTRSH 94
Query: 68 ASVKFHALTRSHA 80
A H L SH+
Sbjct: 95 A----HMLNLSHS 103
>UniRef100_Q6WLA5 CAD [Leptopeza sp. NCSU-99071981]
Length = 1289
Score = 33.5 bits (75), Expect = 2.2
Identities = 21/69 (30%), Positives = 36/69 (51%), Gaps = 2/69 (2%)
Query: 94 SGVSSIHKVSRLSEASRLIPWMANKLLNERRSTNILRTYQTPQHLDPTTLLVFPKLWFDF 153
+ + + + V +L E +++ PW NK+ N N+L T +L+ TTLL KL F
Sbjct: 765 AAIKANYTVEKLYELTKIDPWFLNKMKNIIDYLNLLEV--TGNNLNRTTLLEAKKLGFSD 822
Query: 154 QCLPSRLLS 162
+ + S + S
Sbjct: 823 RQIASAIKS 831
>UniRef100_UPI000042E2D4 UPI000042E2D4 UniRef100 entry
Length = 2094
Score = 32.7 bits (73), Expect = 3.8
Identities = 30/120 (25%), Positives = 54/120 (45%), Gaps = 11/120 (9%)
Query: 6 AVISQTTSPTSLDTSPRVLAKLMSLKAYPKVKSILRQNTIIIKRVQPLTMTLRVINTLTK 65
A +S T+PT D R AK +++ P T+ +K ++ L ++ +
Sbjct: 1753 ADLSDVTAPTRPDVPERP-AKATAIEKGP-------MGTVAVKVIEQLNDEVQRLKDALA 1804
Query: 66 SHASVKFHALTRSHASVEFHAFHKVSRLSGVSSIHKVSRLSEASRLIPWMANKLLNERRS 125
++A TR+ +VEFHA + L S+ S++S+ RLI + + +R S
Sbjct: 1805 ANAEYHQQLRTRNAGTVEFHA---LVNLVEQQSVIPFSQISQVQRLIDELYPTVTKQRDS 1861
>UniRef100_UPI0000309804 UPI0000309804 UniRef100 entry
Length = 286
Score = 31.6 bits (70), Expect = 8.4
Identities = 19/67 (28%), Positives = 33/67 (48%)
Query: 5 NAVISQTTSPTSLDTSPRVLAKLMSLKAYPKVKSILRQNTIIIKRVQPLTMTLRVINTLT 64
N +I + S+D + ++KL+S A ++ +I R T+ + P + L + TL
Sbjct: 123 NMLIIYNDNGISIDPNVGAISKLLSRFASSRIYNIFRDETLELAEKAPFSERLGLKATLQ 182
Query: 65 KSHASVK 71
K H S K
Sbjct: 183 KLHDSAK 189
>UniRef100_Q93TS8 Dissimilatory sulfite reductase alpha subunit [sulfate-reducing
bacterium AK-01]
Length = 351
Score = 31.6 bits (70), Expect = 8.4
Identities = 26/108 (24%), Positives = 44/108 (40%), Gaps = 1/108 (0%)
Query: 4 CNAVISQTTSPTSLDTSPRVLAKLMSLKAYPKVKSILRQNTIIIKRVQPLTMTLRVINTL 63
C A +S+T+S +SL +P + + P + + +I R PLT+ + T
Sbjct: 143 CTARLSRTSSSSSLTAAPTAAWLPLPVPTCPSSEPGATTSALIRMR-SPLTLAANIRPTP 201
Query: 64 TKSHASVKFHALTRSHASVEFHAFHKVSRLSGVSSIHKVSRLSEASRL 111
+ + H +R +S H H+V S S+ AS L
Sbjct: 202 ALTKTATGEHLTSRRKSSTCAHRLHEVQAASWKSTTKSAPAACIASTL 249
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.322 0.131 0.375
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 231,793,699
Number of Sequences: 2790947
Number of extensions: 7454512
Number of successful extensions: 19805
Number of sequences better than 10.0: 5
Number of HSP's better than 10.0 without gapping: 1
Number of HSP's successfully gapped in prelim test: 4
Number of HSP's that attempted gapping in prelim test: 19803
Number of HSP's gapped (non-prelim): 6
length of query: 170
length of database: 848,049,833
effective HSP length: 118
effective length of query: 52
effective length of database: 518,718,087
effective search space: 26973340524
effective search space used: 26973340524
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 70 (31.6 bits)
Lotus: description of TM0247.8


