

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0243.12
(1626 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
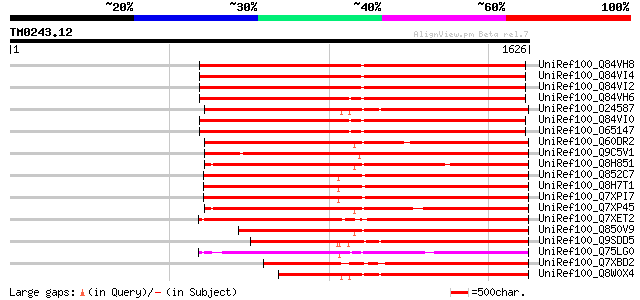
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q84VH8 Gag-pol polyprotein [Glycine max] 989 0.0
UniRef100_Q84VI4 Gag-pol polyprotein [Glycine max] 987 0.0
UniRef100_Q84VI2 Gag-pol polyprotein [Glycine max] 983 0.0
UniRef100_Q84VH6 Gag-pol polyprotein [Glycine max] 975 0.0
UniRef100_O24587 Pol protein [Zea mays] 972 0.0
UniRef100_Q84VI0 Gag-pol polyprotein [Glycine max] 965 0.0
UniRef100_O65147 Gag-pol polyprotein [Glycine max] 965 0.0
UniRef100_Q60DR2 Putative polyprotein [Oryza sativa] 946 0.0
UniRef100_Q9C5V1 Gag/pol polyprotein [Arabidopsis thaliana] 946 0.0
UniRef100_Q8H851 Putative Zea mays retrotransposon Opie-2 [Oryza... 945 0.0
UniRef100_Q852C7 Putative gag-pol polyprotein [Oryza sativa] 938 0.0
UniRef100_Q8H7T1 Putative Zea mays retrotransposon Opie-2 [Oryza... 935 0.0
UniRef100_Q7XPI7 OSJNBb0004A17.2 protein [Oryza sativa] 935 0.0
UniRef100_Q7XP45 OSJNBa0063G07.6 protein [Oryza sativa] 927 0.0
UniRef100_Q7XET2 Hypothetical protein [Oryza sativa] 918 0.0
UniRef100_Q850V9 Putative polyprotein [Oryza sativa] 889 0.0
UniRef100_Q9SDD5 Similar to copia-type pol polyprotein [Oryza sa... 860 0.0
UniRef100_Q75LG0 Putative integrase [Oryza sativa] 825 0.0
UniRef100_Q7XBD2 Retrotransposon Opie-2 [Zea mays] 777 0.0
UniRef100_Q8W0X4 Putative pol protein [Zea mays] 772 0.0
>UniRef100_Q84VH8 Gag-pol polyprotein [Glycine max]
Length = 1576
Score = 989 bits (2558), Expect = 0.0
Identities = 498/1020 (48%), Positives = 680/1020 (65%), Gaps = 7/1020 (0%)
Query: 596 MLQISLIAPLKHQSWYLDSGCSRHMTGEKRMFRELKLKPGGEVGFGGNEKGKIIGTGTIC 655
++ SL A K + WYLDSGCSRHMTG K ++ V FG KGKIIG G +
Sbjct: 549 VVHTSLRASAK-EDWYLDSGCSRHMTGVKEFLLNIEPCSTSYVTFGDGSKGKIIGMGKLV 607
Query: 656 VDSSPCIDNVLLVDGLTHNLLSISQLADKGYDVIFNQKSCRAVSQIDGSVLFNSKRKNNI 715
D P ++ VLLV GLT NL+SISQL D+G++V F + C ++ ++ S+ K+N
Sbjct: 608 HDGLPSLNKVLLVKGLTANLISISQLCDEGFNVNFTKSECLVTNEKSEVLMKGSRSKDNC 667
Query: 716 YKIRLSELEAQNVKCLLSVNEEQWVWHRRLGHASMRKISQLSKLNLVRGLPNLKFASDAL 775
Y E + CL S +E +WH+R GH +R + ++ VRG+PNLK +
Sbjct: 668 YLWTPQETSYSST-CLSSKEDEVRIWHQRFGHLHLRGMKKIIDKGAVRGIPNLKIEEGRI 726
Query: 776 CEACQKGKFTKVPFKAKNVVSTSRPLELLHIDLFGPVKTESIGGKRYGMVIVDDYSRWTW 835
C CQ GK K+ + +TSR LELLH+DL GP++ ES+GGKRY V+VDD+SR+TW
Sbjct: 727 CGECQIGKQVKMSHQKLRHQTTSRVLELLHMDLMGPMQVESLGGKRYAYVVVDDFSRFTW 786
Query: 836 VKFLTRKDESHAVFSTFIAQVQNEKACRIVRVRSDLGGEFENDKFESLFDSYGIAHDFSC 895
V F+ K E+ VF ++Q EK C I R+RSD G EFEN +F S GI H+FS
Sbjct: 787 VNFIREKSETFEVFKELSLRLQREKDCVIKRIRSDHGREFENSRFTEFCTSEGITHEFSA 846
Query: 896 PRTPQQNGVVERKNRTLQEMARTMLQETGMAKHFWAETVNTACYIQNRISVRPILNKTPY 955
TPQQNG+VERKNRTLQE AR ML + + WAE +NTACYI NR+++R T Y
Sbjct: 847 AITPQQNGIVERKNRTLQEAARVMLHAKELPYNLWAEAMNTACYIHNRVTLRRGTPTTLY 906
Query: 956 ELWKNIKPNISYFHPFGCVCYVLNTKDRLHKFDAKSSKCLLLGYSDRSKGFRFYNTDAKT 1015
E+WK KP++ +FH FG CY+L +++ K D KS + LGYS S+ +R +N+ +T
Sbjct: 907 EIWKGRKPSVKHFHIFGSPCYILADREQRRKMDPKSDAGIFLGYSTNSRAYRVFNSRTRT 966
Query: 1016 IEESIHVRFDDKLDSDQSKLVEKFADLSINVSDKGKAPEEVEPEEDEPEEEAGPSNSQTL 1075
+ ESI+V DD + + + E L NV+D K+ E E D +E+ +
Sbjct: 967 VMESINVVVDDLSPARKKDVEEDVRTLGDNVADAAKSGENAE-NSDSATDESNINQPDKR 1025
Query: 1076 KKSRITAAHPKELILGNKDEPVRTRSAFRPSEETLLSLKGLVSLIEPKSIDEALQDKEWI 1135
+RI HPKELI+G+ + V TRS E ++S VS IEPK++ EAL D+ WI
Sbjct: 1026 SSTRIQKMHPKELIIGDPNRGVTTRSR----EVEIVSNSCFVSKIEPKNVKEALTDEFWI 1081
Query: 1136 LAMEEELNQFSKNDVWSLVKKPESVHVIGTKWVFRNKLNEKGDVVRNKARLVAQGYSQQE 1195
AM+EEL QF +N+VW LV +PE +VIGTKW+F+NK NE+G + RNKARLVAQGY+Q E
Sbjct: 1082 NAMQEELEQFKRNEVWELVPRPEGTNVIGTKWIFKNKTNEEGVITRNKARLVAQGYTQIE 1141
Query: 1196 GIDYTETFAPVARLEAIRLLISFSVNHNIVLHQMDVKSAFLNGYISEEVYVHQPPGFEDE 1255
G+D+ ETFAPVARLE+IRLL+ + L+QMDVKSAFLNGY++EEVYV QP GF D
Sbjct: 1142 GVDFDETFAPVARLESIRLLLGVACILKFKLYQMDVKSAFLNGYLNEEVYVEQPKGFADP 1201
Query: 1256 KKPDHVFKLKKSLYGLKQAPRAWYERLSSFLLENEFVRGKVDTTLFCKTYKDDILIVQIY 1315
PDHV++LKK+LYGLKQAPRAWYERL+ FL + + +G +D TLF K ++++I QIY
Sbjct: 1202 THPDHVYRLKKALYGLKQAPRAWYERLTEFLTQQGYRKGGIDKTLFVKQDAENLMIAQIY 1261
Query: 1316 VDDIIFGSANQSLCKEFSEMMQAEFEMSMMGELKYFLGIQVDQTPEGTYIHQSKYTKELL 1375
VDDI+FG + + + F + MQ+EFEMS++GEL YFLG+QV Q + ++ QS+Y K ++
Sbjct: 1262 VDDIVFGGMSNEMLRHFVQQMQSEFEMSLVGELTYFLGLQVKQMEDSIFLSQSRYAKNIV 1321
Query: 1376 KKFNMLESTVAKTPMHPTCILEKEDKSGKVCQKLYRGMIGSLLYLTASRPDILFSVHLCA 1435
KKF M ++ +TP L K++ V Q LYR MIGSLLYLTASRPDI ++V +CA
Sbjct: 1322 KKFGMENASHKRTPAPTHLKLSKDEAGTSVDQSLYRSMIGSLLYLTASRPDITYAVGVCA 1381
Query: 1436 RFQSDPRETHLTAVKRILRYLKGTTNLGLMYKKTSEYKLSGYCDADYAGDRTERKSTSGN 1495
R+Q++P+ +HLT VKRIL+Y+ GT++ G+MY S L GYCDAD+AG +RKSTSG
Sbjct: 1382 RYQANPKISHLTQVKRILKYVNGTSDYGIMYCHCSNPMLVGYCDADWAGSADDRKSTSGG 1441
Query: 1496 CQFLGSNLVSWASKRQSTIALSTAEAEYISAAICSTQMLWMKHQLEDYQILESNIPIYCD 1555
C +LG+NL+SW SK+Q+ ++LSTAEAEYI+A +Q++WMK L++Y + + + +YCD
Sbjct: 1442 CFYLGNNLISWFSKKQNCVSLSTAEAEYIAAGSSCSQLVWMKQMLKEYNVEQDVMTLYCD 1501
Query: 1556 NTAAISLSKNPILHSRAKHIEVKYHFIRDYVQKGVLLLKFVDTDHQWADIFTKSLAEDRF 1615
N +AI++SKNP+ HSR KHI++++H+IRD V V+ LK VDT+ Q ADIFTK+L ++F
Sbjct: 1502 NMSAINISKNPVQHSRTKHIDIRHHYIRDLVDDKVITLKHVDTEEQIADIFTKALDANQF 1561
>UniRef100_Q84VI4 Gag-pol polyprotein [Glycine max]
Length = 1574
Score = 987 bits (2551), Expect = 0.0
Identities = 497/1020 (48%), Positives = 679/1020 (65%), Gaps = 7/1020 (0%)
Query: 596 MLQISLIAPLKHQSWYLDSGCSRHMTGEKRMFRELKLKPGGEVGFGGNEKGKIIGTGTIC 655
++ SL A K + WYLDSGCSRHMTG K ++ V FG KGKIIG G +
Sbjct: 547 VVHTSLRASAK-EDWYLDSGCSRHMTGVKEFLLNIEPCSTSYVTFGDGSKGKIIGMGKLV 605
Query: 656 VDSSPCIDNVLLVDGLTHNLLSISQLADKGYDVIFNQKSCRAVSQIDGSVLFNSKRKNNI 715
D P ++ VLLV GLT NL+SISQL D+G++V F + C ++ ++ S+ K+N
Sbjct: 606 HDGLPSLNKVLLVKGLTANLISISQLCDEGFNVNFTKSECLVTNEKSEVLMKGSRSKDNC 665
Query: 716 YKIRLSELEAQNVKCLLSVNEEQWVWHRRLGHASMRKISQLSKLNLVRGLPNLKFASDAL 775
Y E + CL S +E +WH+R GH +R + ++ VRG+PNLK +
Sbjct: 666 YLWTPQETSYSST-CLSSKEDEVRIWHQRFGHLHLRGMKKIIDKGAVRGIPNLKIEEGRI 724
Query: 776 CEACQKGKFTKVPFKAKNVVSTSRPLELLHIDLFGPVKTESIGGKRYGMVIVDDYSRWTW 835
C CQ GK K+ + +TSR LELLH+DL GP++ ES+GGKRY V+VDD+SR+TW
Sbjct: 725 CGECQIGKQVKMSHQKLQHQTTSRVLELLHMDLMGPMQVESLGGKRYAYVVVDDFSRFTW 784
Query: 836 VKFLTRKDESHAVFSTFIAQVQNEKACRIVRVRSDLGGEFENDKFESLFDSYGIAHDFSC 895
VKF+ K E+ VF ++Q EK C I R+RSD G EFEN + S GI H+FS
Sbjct: 785 VKFIREKSETFEVFKELSLRLQREKDCVIKRIRSDHGREFENSRLTEFCTSEGITHEFSA 844
Query: 896 PRTPQQNGVVERKNRTLQEMARTMLQETGMAKHFWAETVNTACYIQNRISVRPILNKTPY 955
TPQQNG+VERKNRTLQE AR ML + + WAE +NTACYI NR+++R T Y
Sbjct: 845 AITPQQNGIVERKNRTLQEAARVMLHAKELPYNLWAEAMNTACYIHNRVTLRRGTPTTLY 904
Query: 956 ELWKNIKPNISYFHPFGCVCYVLNTKDRLHKFDAKSSKCLLLGYSDRSKGFRFYNTDAKT 1015
E+WK KP++ +FH FG CY+L +++ K D KS + LGYS S+ +R +N+ +T
Sbjct: 905 EIWKGRKPSVKHFHIFGSPCYILADREQRRKMDPKSDAGIFLGYSTNSRAYRVFNSRTRT 964
Query: 1016 IEESIHVRFDDKLDSDQSKLVEKFADLSINVSDKGKAPEEVEPEEDEPEEEAGPSNSQTL 1075
+ ESI+V DD + + + E NV+D K+ E E D +E+ +
Sbjct: 965 VMESINVVVDDLSPARKKDVEEDVRTSGDNVADAAKSGENAE-NSDSATDESNINQPDKR 1023
Query: 1076 KKSRITAAHPKELILGNKDEPVRTRSAFRPSEETLLSLKGLVSLIEPKSIDEALQDKEWI 1135
+RI HPKELI+G+ + V TRS E ++S VS IEPK++ EAL D+ WI
Sbjct: 1024 SSTRIQKMHPKELIIGDPNRGVTTRSR----EVEIVSNSCFVSKIEPKNVKEALTDEFWI 1079
Query: 1136 LAMEEELNQFSKNDVWSLVKKPESVHVIGTKWVFRNKLNEKGDVVRNKARLVAQGYSQQE 1195
AM+EEL QF +N+VW LV +PE +VIGTKW+F+NK NE+G + RNKARLVAQGY+Q E
Sbjct: 1080 NAMQEELEQFKRNEVWELVPRPEGTNVIGTKWIFKNKTNEEGVITRNKARLVAQGYTQIE 1139
Query: 1196 GIDYTETFAPVARLEAIRLLISFSVNHNIVLHQMDVKSAFLNGYISEEVYVHQPPGFEDE 1255
G+D+ ETFAPVARLE+IRLL+ + L+QMDVKSAFLNGY++EEVYV QP GF D
Sbjct: 1140 GVDFDETFAPVARLESIRLLLGVACILKFKLYQMDVKSAFLNGYLNEEVYVEQPKGFADP 1199
Query: 1256 KKPDHVFKLKKSLYGLKQAPRAWYERLSSFLLENEFVRGKVDTTLFCKTYKDDILIVQIY 1315
PDHV++LKK+LYGLKQAPRAWYERL+ FL + + +G +D TLF K ++++I QIY
Sbjct: 1200 THPDHVYRLKKALYGLKQAPRAWYERLTEFLTQQGYRKGGIDKTLFVKQDAENLMIAQIY 1259
Query: 1316 VDDIIFGSANQSLCKEFSEMMQAEFEMSMMGELKYFLGIQVDQTPEGTYIHQSKYTKELL 1375
VDDI+FG + + + F + MQ+EFEMS++GEL YFLG+QV Q + ++ QS+Y K ++
Sbjct: 1260 VDDIVFGGMSNEMLRHFVQQMQSEFEMSLVGELTYFLGLQVKQMEDSIFLSQSRYAKNIV 1319
Query: 1376 KKFNMLESTVAKTPMHPTCILEKEDKSGKVCQKLYRGMIGSLLYLTASRPDILFSVHLCA 1435
KKF M ++ +TP L K++ V Q LYR MIGSLLYLTASRPDI ++V +CA
Sbjct: 1320 KKFGMENASHKRTPAPTHLKLSKDEAGTSVDQSLYRSMIGSLLYLTASRPDITYAVGVCA 1379
Query: 1436 RFQSDPRETHLTAVKRILRYLKGTTNLGLMYKKTSEYKLSGYCDADYAGDRTERKSTSGN 1495
R+Q++P+ +HLT VKRIL+Y+ GT++ G+MY S L GYCDAD+AG +RKSTSG
Sbjct: 1380 RYQANPKISHLTQVKRILKYVNGTSDYGIMYCHCSNPMLVGYCDADWAGSADDRKSTSGG 1439
Query: 1496 CQFLGSNLVSWASKRQSTIALSTAEAEYISAAICSTQMLWMKHQLEDYQILESNIPIYCD 1555
C +LG+NL+SW SK+Q+ ++LSTAEAEYI+A +Q++WMK L++Y + + + +YCD
Sbjct: 1440 CFYLGNNLISWFSKKQNCVSLSTAEAEYIAAGSSCSQLVWMKQMLKEYNVEQDVMTLYCD 1499
Query: 1556 NTAAISLSKNPILHSRAKHIEVKYHFIRDYVQKGVLLLKFVDTDHQWADIFTKSLAEDRF 1615
N +AI++SKNP+ HSR KHI++++H+IRD V V+ LK VDT+ Q ADIFTK+L ++F
Sbjct: 1500 NMSAINISKNPVQHSRTKHIDIRHHYIRDLVDDKVITLKHVDTEEQIADIFTKALDANQF 1559
>UniRef100_Q84VI2 Gag-pol polyprotein [Glycine max]
Length = 1576
Score = 983 bits (2540), Expect = 0.0
Identities = 495/1020 (48%), Positives = 678/1020 (65%), Gaps = 7/1020 (0%)
Query: 596 MLQISLIAPLKHQSWYLDSGCSRHMTGEKRMFRELKLKPGGEVGFGGNEKGKIIGTGTIC 655
++ SL A K + WYLDSGCSRHMTG K ++ V FG KGKIIG G +
Sbjct: 549 VVHTSLRASAK-EDWYLDSGCSRHMTGVKEFLLNIEPCSTSYVTFGDGSKGKIIGMGKLV 607
Query: 656 VDSSPCIDNVLLVDGLTHNLLSISQLADKGYDVIFNQKSCRAVSQIDGSVLFNSKRKNNI 715
D P ++ VLLV GLT NL+SISQL D+G++V F + C ++ ++ S+ K+N
Sbjct: 608 HDGLPSLNKVLLVKGLTANLISISQLCDEGFNVNFTKSECLVTNEKSEVLMKGSRSKDNC 667
Query: 716 YKIRLSELEAQNVKCLLSVNEEQWVWHRRLGHASMRKISQLSKLNLVRGLPNLKFASDAL 775
Y E + CL S +E +WH+R GH +R + ++ + VRG+PNLK +
Sbjct: 668 YLWTPQETSYSST-CLSSKEDEVRIWHQRFGHLHLRGMKKILDKSAVRGIPNLKIEEGRI 726
Query: 776 CEACQKGKFTKVPFKAKNVVSTSRPLELLHIDLFGPVKTESIGGKRYGMVIVDDYSRWTW 835
C CQ GK K+ + +TSR LELLH+DL GP++ ES+GGKRY V+VDD+SR+TW
Sbjct: 727 CGECQIGKQVKMSHQKLQHQTTSRVLELLHMDLMGPMQVESLGGKRYAYVVVDDFSRFTW 786
Query: 836 VKFLTRKDESHAVFSTFIAQVQNEKACRIVRVRSDLGGEFENDKFESLFDSYGIAHDFSC 895
V F+ K + VF ++Q EK C I R+RSD G EFEN +F S GI H+FS
Sbjct: 787 VNFIREKSGTFEVFKKLSLRLQREKDCVIKRIRSDHGREFENSRFTEFCTSEGITHEFSA 846
Query: 896 PRTPQQNGVVERKNRTLQEMARTMLQETGMAKHFWAETVNTACYIQNRISVRPILNKTPY 955
TPQQNG+VERKNRTLQE AR ML + + WAE +NTACYI NR+++R T Y
Sbjct: 847 AITPQQNGIVERKNRTLQEAARVMLHAKELPYNLWAEAMNTACYIHNRVTLRRGTPTTLY 906
Query: 956 ELWKNIKPNISYFHPFGCVCYVLNTKDRLHKFDAKSSKCLLLGYSDRSKGFRFYNTDAKT 1015
E+WK KP++ +FH FG CY+L +++ K D KS + LGYS S+ +R +N+ +T
Sbjct: 907 EIWKGRKPSVKHFHIFGSPCYILADREQRRKMDPKSDAGIFLGYSTNSRAYRVFNSRTRT 966
Query: 1016 IEESIHVRFDDKLDSDQSKLVEKFADLSINVSDKGKAPEEVEPEEDEPEEEAGPSNSQTL 1075
+ ESI+V DD + + + E NV+D K+ E E D +E+ +
Sbjct: 967 VMESINVVVDDLSPARKKDVEEDVRTSGDNVADAAKSGENAE-NSDSATDESNINQPDKR 1025
Query: 1076 KKSRITAAHPKELILGNKDEPVRTRSAFRPSEETLLSLKGLVSLIEPKSIDEALQDKEWI 1135
+RI HPKELI+G+ + V TRS E ++S VS IEPK++ EAL D+ WI
Sbjct: 1026 SSTRIQKMHPKELIIGDPNRGVTTRSR----EVEIVSNSCFVSKIEPKNVKEALTDEFWI 1081
Query: 1136 LAMEEELNQFSKNDVWSLVKKPESVHVIGTKWVFRNKLNEKGDVVRNKARLVAQGYSQQE 1195
AM+EEL QF +N+VW LV +PE +VIGTKW+F+NK NE+G + RNKARLVAQGY+Q E
Sbjct: 1082 NAMQEELEQFKRNEVWELVPRPEGTNVIGTKWIFKNKTNEEGVITRNKARLVAQGYTQIE 1141
Query: 1196 GIDYTETFAPVARLEAIRLLISFSVNHNIVLHQMDVKSAFLNGYISEEVYVHQPPGFEDE 1255
G+D+ ETFAPVARLE+IRLL+ + L+QMDVKSAFLNGY++EEVYV QP GF D
Sbjct: 1142 GVDFDETFAPVARLESIRLLLGVACILKFKLYQMDVKSAFLNGYLNEEVYVEQPKGFADP 1201
Query: 1256 KKPDHVFKLKKSLYGLKQAPRAWYERLSSFLLENEFVRGKVDTTLFCKTYKDDILIVQIY 1315
PDHV++LKK+LYGLKQAPRAWYERL+ FL + + +G +D TLF K ++++I QIY
Sbjct: 1202 THPDHVYRLKKALYGLKQAPRAWYERLTEFLTQQGYRKGGIDKTLFVKQDAENLMIAQIY 1261
Query: 1316 VDDIIFGSANQSLCKEFSEMMQAEFEMSMMGELKYFLGIQVDQTPEGTYIHQSKYTKELL 1375
VDDI+FG + + + F + MQ+EFEMS++GEL YFLG+QV Q + ++ QS+Y K ++
Sbjct: 1262 VDDIVFGGMSNEMLRHFVQQMQSEFEMSLVGELTYFLGLQVKQMEDSIFLSQSRYAKNIV 1321
Query: 1376 KKFNMLESTVAKTPMHPTCILEKEDKSGKVCQKLYRGMIGSLLYLTASRPDILFSVHLCA 1435
KKF M ++ +TP L K++ V QK YR MIGSLLYLTASRPDI ++V +CA
Sbjct: 1322 KKFGMENASHKRTPAPTHLKLSKDEAGTSVDQKPYRSMIGSLLYLTASRPDITYAVGVCA 1381
Query: 1436 RFQSDPRETHLTAVKRILRYLKGTTNLGLMYKKTSEYKLSGYCDADYAGDRTERKSTSGN 1495
R+Q++P+ +HL VKRIL+Y+ GT++ G+MY S L GYCDAD+AG +RKSTSG
Sbjct: 1382 RYQANPKISHLNQVKRILKYVNGTSDYGIMYCHCSSSMLVGYCDADWAGSADDRKSTSGG 1441
Query: 1496 CQFLGSNLVSWASKRQSTIALSTAEAEYISAAICSTQMLWMKHQLEDYQILESNIPIYCD 1555
C +LG+NL+SW SK+Q+ ++LSTAEAEYI+A +Q++WMK L++Y + + + +YCD
Sbjct: 1442 CFYLGNNLISWFSKKQNCVSLSTAEAEYIAAGSSCSQLVWMKQMLKEYNVEQDVMTLYCD 1501
Query: 1556 NTAAISLSKNPILHSRAKHIEVKYHFIRDYVQKGVLLLKFVDTDHQWADIFTKSLAEDRF 1615
N +AI++SKNP+ HSR KHI++++H+IRD V V+ LK VDT+ Q ADIFTK+L ++F
Sbjct: 1502 NMSAINISKNPVQHSRTKHIDIRHHYIRDLVDDKVITLKHVDTEEQIADIFTKALDANQF 1561
>UniRef100_Q84VH6 Gag-pol polyprotein [Glycine max]
Length = 1577
Score = 975 bits (2521), Expect = 0.0
Identities = 494/1022 (48%), Positives = 678/1022 (66%), Gaps = 11/1022 (1%)
Query: 596 MLQISLIAPLKHQSWYLDSGCSRHMTGEKRMFRELKLKPGGEVGFGGNEKGKIIGTGTIC 655
++ SL A K + WYLDSGCSRHMTG K ++ V FG KGKI G G +
Sbjct: 550 VVHTSLRASAK-EDWYLDSGCSRHMTGVKEFLVNIEPCSTSYVTFGDGSKGKITGMGKLV 608
Query: 656 VDSSPCIDNVLLVDGLTHNLLSISQLADKGYDVIFNQKSCRAVSQIDGSVLFNSKRKNNI 715
+ P ++ VLLV GLT NL+SISQL D+G++V F + C ++ ++ S+ K+N
Sbjct: 609 HEGLPSLNKVLLVKGLTVNLISISQLCDEGFNVNFTKSECLVTNEKSEVLMKGSRSKDNC 668
Query: 716 YKIRLSELEAQNVKCLLSVNEEQWVWHRRLGHASMRKISQLSKLNLVRGLPNLKFASDAL 775
Y E + + CL S +E +WH+R GH +R + ++ VRG+PNLK +
Sbjct: 669 YLWTPQE-SSHSSTCLFSKEDEVKIWHQRFGHLHLRGMKKIIDKGAVRGIPNLKIEEGRI 727
Query: 776 CEACQKGKFTKVPFKAKNVVSTSRPLELLHIDLFGPVKTESIGGKRYGMVIVDDYSRWTW 835
C CQ GK K+ + +TSR LELLH+DL GP++ ES+GGKRY V+VDD+SR+TW
Sbjct: 728 CGECQIGKQVKMSHQKLQHQTTSRVLELLHMDLMGPMQVESLGGKRYAYVVVDDFSRFTW 787
Query: 836 VKFLTRKDESHAVFSTFIAQVQNEKACRIVRVRSDLGGEFENDKFESLFDSYGIAHDFSC 895
V F+ K ++ VF ++Q EK C I R+RSD G EFEN KF S GI H+FS
Sbjct: 788 VNFIREKSDTFEVFKELSLRLQREKDCVIKRIRSDHGREFENSKFTEFCTSEGITHEFSA 847
Query: 896 PRTPQQNGVVERKNRTLQEMARTMLQETGMAKHFWAETVNTACYIQNRISVRPILNKTPY 955
TPQQNG+VERKNRTLQE AR ML + + WAE +NTACYI NR+++R T Y
Sbjct: 848 AITPQQNGIVERKNRTLQEAARVMLHAKELPYNLWAEAMNTACYIHNRVTLRRGTPTTLY 907
Query: 956 ELWKNIKPNISYFHPFGCVCYVLNTKDRLHKFDAKSSKCLLLGYSDRSKGFRFYNTDAKT 1015
E+WK KP + +FH FG CY+L +++ K D KS + LGYS S+ +R +N+ +T
Sbjct: 908 EIWKGRKPTVKHFHIFGSPCYILADREQRRKMDPKSDAGIFLGYSTNSRAYRVFNSRTRT 967
Query: 1016 IEESIHVRFDDKLDSDQSKLVEKFADLSINVSDKGKAPEEVEPEEDEPEEEAGPSNSQTL 1075
+ ESI+V DD + + + E NV+D K+ E E + +E P+ +Q
Sbjct: 968 VMESINVVVDDLTPARKKDVEEDVRTSGDNVADTAKSAENAENSDSATDE---PNINQPD 1024
Query: 1076 KKS--RITAAHPKELILGNKDEPVRTRSAFRPSEETLLSLKGLVSLIEPKSIDEALQDKE 1133
K+ RI HPKELI+G+ + V TRS E ++S VS IEPK++ EAL D+
Sbjct: 1025 KRPSIRIQKMHPKELIIGDPNRGVTTRSR----EIEIVSNSCFVSKIEPKNVKEALTDEF 1080
Query: 1134 WILAMEEELNQFSKNDVWSLVKKPESVHVIGTKWVFRNKLNEKGDVVRNKARLVAQGYSQ 1193
WI AM+EEL QF +N+VW LV +PE +VIGTKW+F+NK NE+G + RNKARLVAQGY+Q
Sbjct: 1081 WINAMQEELEQFKRNEVWELVPRPEGTNVIGTKWIFKNKTNEEGVITRNKARLVAQGYTQ 1140
Query: 1194 QEGIDYTETFAPVARLEAIRLLISFSVNHNIVLHQMDVKSAFLNGYISEEVYVHQPPGFE 1253
EG+D+ ETFAPVARLE+IRLL+ + L+QMDVKSAFLNGY++EE YV QP GF
Sbjct: 1141 IEGVDFDETFAPVARLESIRLLLGVACILKFKLYQMDVKSAFLNGYLNEEAYVEQPKGFV 1200
Query: 1254 DEKKPDHVFKLKKSLYGLKQAPRAWYERLSSFLLENEFVRGKVDTTLFCKTYKDDILIVQ 1313
D PDHV++LKK+LYGLKQAPRAWYERL+ FL + + +G +D TLF K ++++I Q
Sbjct: 1201 DPTHPDHVYRLKKALYGLKQAPRAWYERLTEFLTQQGYRKGGIDKTLFVKQDAENLMIAQ 1260
Query: 1314 IYVDDIIFGSANQSLCKEFSEMMQAEFEMSMMGELKYFLGIQVDQTPEGTYIHQSKYTKE 1373
IYVDDI+FG + + + F + MQ+EFEMS++GEL YFLG+QV Q + ++ QSKY K
Sbjct: 1261 IYVDDIVFGGMSNEMLRHFVQQMQSEFEMSLVGELTYFLGLQVKQMEDSIFLSQSKYAKN 1320
Query: 1374 LLKKFNMLESTVAKTPMHPTCILEKEDKSGKVCQKLYRGMIGSLLYLTASRPDILFSVHL 1433
++KKF M ++ +TP L K++ V Q LYR MIGSLLYLTASRPDI ++V +
Sbjct: 1321 IVKKFGMENASHKRTPAPTHLKLSKDEAGTSVDQSLYRSMIGSLLYLTASRPDITYAVGV 1380
Query: 1434 CARFQSDPRETHLTAVKRILRYLKGTTNLGLMYKKTSEYKLSGYCDADYAGDRTERKSTS 1493
CAR+Q++P+ +HL VKRIL+Y+ GT++ G+MY S L GYCDAD+AG +RKSTS
Sbjct: 1381 CARYQANPKISHLNQVKRILKYVNGTSDYGIMYCHCSGSMLVGYCDADWAGSADDRKSTS 1440
Query: 1494 GNCQFLGSNLVSWASKRQSTIALSTAEAEYISAAICSTQMLWMKHQLEDYQILESNIPIY 1553
G C +LG+NL+SW SK+Q+ ++LSTAEAEYI+A +Q++WMK L++Y + + + +Y
Sbjct: 1441 GGCFYLGNNLISWFSKKQNCVSLSTAEAEYIAAGSSCSQLVWMKQMLKEYNVEQDVMTLY 1500
Query: 1554 CDNTAAISLSKNPILHSRAKHIEVKYHFIRDYVQKGVLLLKFVDTDHQWADIFTKSLAED 1613
CDN +AI++SKNP+ HSR KHI++++H+IR+ V V+ L+ VDT+ Q ADIFTK+L
Sbjct: 1501 CDNMSAINISKNPVQHSRTKHIDIRHHYIRELVDDKVITLEHVDTEEQIADIFTKALDAK 1560
Query: 1614 RF 1615
+F
Sbjct: 1561 QF 1562
>UniRef100_O24587 Pol protein [Zea mays]
Length = 1068
Score = 972 bits (2513), Expect = 0.0
Identities = 515/1058 (48%), Positives = 674/1058 (63%), Gaps = 48/1058 (4%)
Query: 609 SWYLDSGCSRHMTGEKRMFRE-LKLKPGGE-VGFGGNEKGKIIGTGTICVDSSPCIDNVL 666
SW +DSGC+ HMTGEK+MF +K K + + FG +GK+ G G I + + I NV
Sbjct: 10 SWIIDSGCTNHMTGEKKMFTSYVKNKDSQDSIIFGDGNQGKVKGLGKIAISNEHSISNVF 69
Query: 667 LVDGLTHNLLSISQLADKGYDVIFNQKSCRAVSQIDGSVLFNSKRKNNIYKIRLSELEAQ 726
LV+ L +NLLS+SQL + GY+ +F + DGS+ F +Y + ++ EA
Sbjct: 70 LVESLGYNLLSVSQLCNMGYNCLFTNIDVSVFRRCDGSLAFKGVLDGKLYLVDFAKEEAG 129
Query: 727 NVKCLLSVNEEQWVWHRRLGHASMRKISQLSKLNLVRGLPNLKFASDALCEACQKGKFTK 786
CL++ W+WHRRL H M+ + +L K V GL N++F D C ACQ GK
Sbjct: 130 LDACLIAKTSMGWLWHRRLAHVGMKNLHKLLKGEHVIGLTNVQFEKDRPCAACQAGKQVG 189
Query: 787 VPFKAKNVVSTSRPLELLHIDLFGPVKTESIGGKRYGMVIVDDYSRWTWVKFLTRKDESH 846
KNV++TSRPLE+LH+DLFGPV SIGG +YG+VIVDD+SR+TWV FL K E+
Sbjct: 190 GSHHTKNVMTTSRPLEMLHMDLFGPVAYLSIGGSKYGLVIVDDFSRFTWVFFLQEKSETQ 249
Query: 847 AVFSTFIAQVQNEKACRIVRVRSDLGGEFENDKFESLFDSYGIAHDFSCPRTPQQNGVVE 906
F+ + QNE ++ ++RSD G EF+N + E + GI H+FS P TPQQNGVVE
Sbjct: 250 GTLKRFLRRAQNEFELKVKKIRSDNGSEFKNLQVEEFLEEEGIKHEFSAPYTPQQNGVVE 309
Query: 907 RKNRTLQEMARTMLQETGMAKHFWAETVNTACYIQNRISVRPILNKTPYELWKNIKPNIS 966
RKNRTL +MARTML E + FW E VNTAC+ NR+ + IL T YEL KPN+S
Sbjct: 310 RKNRTLIDMARTMLGEFKTPECFWTEAVNTACHAINRVYLHRILKNTSYELLTGNKPNVS 369
Query: 967 YFHPFGCVCYVLNTKDRLHKFDAKSSKCLLLGYSDRSKGFRFYNTDAKTIEESIHVRFDD 1026
YF FG CY+L K R KF K+ + LLGY +K +R +N + +E S V FD+
Sbjct: 370 YFRVFGSKCYILVKKGRNSKFAPKAVEGFLLGYDSNTKAYRVFNKSSGLVEVSGDVVFDE 429
Query: 1027 KLDSDQSKLV-------EKFADLSINVSDKGKAPEEVEPEEDEP---------------- 1063
S + ++V E +I G+ + + E ++P
Sbjct: 430 TNGSPREQVVDCDDVDEEDIPTAAIRTMAIGEVRPQEQDEREQPSPSTMVHPPTQDDEQV 489
Query: 1064 -----------------EEEAGPSNSQTLKKSRITAAHPKELILGNKDEPVRTRSAFRPS 1106
EEEA P+ T ++ I HP + ILG+ + V TRS
Sbjct: 490 HQQEVCDQGGAQDDHVLEEEAQPA-PPTQVRAMIQRDHPVDQILGDISKGVTTRSRL--- 545
Query: 1107 EETLLSLKGLVSLIEPKSIDEALQDKEWILAMEEELNQFSKNDVWSLVKKPESVHVIGTK 1166
VS IEP ++EAL D +W+LAM+EELN F +N+VW+LV +P+ +V+GTK
Sbjct: 546 -VNFCEHNSFVSSIEPFRVEEALLDPDWVLAMQEELNNFKRNEVWTLVPRPKQ-NVVGTK 603
Query: 1167 WVFRNKLNEKGDVVRNKARLVAQGYSQQEGIDYTETFAPVARLEAIRLLISFSVNHNIVL 1226
WVFRNK +E+G V RNKARLVA+GY+Q G+D+ ETFAPVARLE+IR+L++++ +H+ L
Sbjct: 604 WVFRNKQDERGVVTRNKARLVAKGYAQVAGLDFEETFAPVARLESIRILLAYAAHHSFRL 663
Query: 1227 HQMDVKSAFLNGYISEEVYVHQPPGFEDEKKPDHVFKLKKSLYGLKQAPRAWYERLSSFL 1286
+QMDVKSAFLNG I EEVYV QPPGFEDE+ PDHV KL K+LYGLKQAPRAWYE L FL
Sbjct: 664 YQMDVKSAFLNGPIKEEVYVEQPPGFEDERYPDHVCKLSKALYGLKQAPRAWYECLRDFL 723
Query: 1287 LENEFVRGKVDTTLFCKTYKDDILIVQIYVDDIIFGSANQSLCKEFSEMMQAEFEMSMMG 1346
+ N F GK D TLF KT D+ + QIYVDDIIFGS NQ C+EFS +M +FEMSMMG
Sbjct: 724 IANAFKVGKADPTLFTKTCDGDLFVCQIYVDDIIFGSTNQKSCEEFSRVMTQKFEMSMMG 783
Query: 1347 ELKYFLGIQVDQTPEGTYIHQSKYTKELLKKFNMLESTVAKTPMHPTCILEKEDKSGKVC 1406
EL YFLG QV Q +GT+I Q+KYT++LLK+F M ++ AKTPM + V
Sbjct: 784 ELNYFLGFQVKQLKDGTFISQTKYTQDLLKRFGMKDAKPAKTPMGTDGHTDLNKGGKSVD 843
Query: 1407 QKLYRGMIGSLLYLTASRPDILFSVHLCARFQSDPRETHLTAVKRILRYLKGTTNLGLMY 1466
QK YR MIGSLLYL ASRPDI+ SV +CARFQSDP+E HL AVKRILRYL T GL Y
Sbjct: 844 QKAYRSMIGSLLYLCASRPDIMLSVCMCARFQSDPKECHLVAVKRILRYLVATPCFGLWY 903
Query: 1467 KKTSEYKLSGYCDADYAGDRTERKSTSGNCQFLGSNLVSWASKRQSTIALSTAEAEYISA 1526
K S + L GY D+DYAG + +RKSTSG CQFLG +LVSW SK+Q+++ALSTAEAEY++A
Sbjct: 904 PKGSTFDLVGYSDSDYAGCKVDRKSTSGTCQFLGRSLVSWNSKKQTSVALSTAEAEYVAA 963
Query: 1527 AICSTQMLWMKHQLEDYQILESNIPIYCDNTAAISLSKNPILHSRAKHIEVKYHFIRDYV 1586
C Q+LWM+ L D+ S +P+ CDN +AI +++NP+ HSR KHI++++HF+RD+
Sbjct: 964 GQCCAQLLWMRQTLRDFGYNLSKVPLLCDNESAIRMAENPVEHSRTKHIDIRHHFLRDHQ 1023
Query: 1587 QKGVLLLKFVDTDHQWADIFTKSLAEDRFNFILKNLNM 1624
QKG + + V T++Q ADIFTK L E F + LN+
Sbjct: 1024 QKGDIEVFHVSTENQLADIFTKPLDEKTFCRLRSELNV 1061
>UniRef100_Q84VI0 Gag-pol polyprotein [Glycine max]
Length = 1576
Score = 965 bits (2495), Expect = 0.0
Identities = 490/1022 (47%), Positives = 672/1022 (64%), Gaps = 11/1022 (1%)
Query: 596 MLQISLIAPLKHQSWYLDSGCSRHMTGEKRMFRELKLKPGGEVGFGGNEKGKIIGTGTIC 655
++ SL A K + WYLDSGCSRHMTG K ++ V FG KGKI G G +
Sbjct: 549 VVHTSLRASAK-EDWYLDSGCSRHMTGVKEFLVNIEPCSTSYVTFGDGSKGKITGMGKLV 607
Query: 656 VDSSPCIDNVLLVDGLTHNLLSISQLADKGYDVIFNQKSCRAVSQIDGSVLFNSKRKNNI 715
D P ++ VLLV GLT NL+SISQL D+G++V F + C ++ ++ S+ K+N
Sbjct: 608 HDGLPSLNKVLLVKGLTANLISISQLCDEGFNVNFTKSECLVTNEKSEVLMKGSRSKDNC 667
Query: 716 YKIRLSELEAQNVKCLLSVNEEQWVWHRRLGHASMRKISQLSKLNLVRGLPNLKFASDAL 775
Y E + CL S +E +WH+R GH +R + ++ VRG+PNLK +
Sbjct: 668 YLWTPQETSYSST-CLSSKEDEVKIWHQRFGHLHLRGMKKIIDKGAVRGIPNLKIEEGRI 726
Query: 776 CEACQKGKFTKVPFKAKNVVSTSRPLELLHIDLFGPVKTESIGGKRYGMVIVDDYSRWTW 835
C CQ GK K+ + +TS LELLH+DL GP++ ES+GGKRY V+VDD+SR+TW
Sbjct: 727 CGECQIGKQVKMSHQKLQHQTTSMVLELLHMDLMGPMQVESLGGKRYAYVVVDDFSRFTW 786
Query: 836 VKFLTRKDESHAVFSTFIAQVQNEKACRIVRVRSDLGGEFENDKFESLFDSYGIAHDFSC 895
V F+ K ++ VF ++Q EK C I R+RSD G EFEN KF S GI H+FS
Sbjct: 787 VNFIREKSDTFEVFKELSLRLQREKDCVIKRIRSDHGREFENSKFTEFCTSEGITHEFSA 846
Query: 896 PRTPQQNGVVERKNRTLQEMARTMLQETGMAKHFWAETVNTACYIQNRISVRPILNKTPY 955
TPQQNG+VERKNRTLQE R ML + + WAE +NTACYI NR+++R T Y
Sbjct: 847 AITPQQNGIVERKNRTLQEATRVMLHAKELPYNLWAEAMNTACYIHNRVTLRRGTPTTLY 906
Query: 956 ELWKNIKPNISYFHPFGCVCYVLNTKDRLHKFDAKSSKCLLLGYSDRSKGFRFYNTDAKT 1015
E+WK KP + +FH FG CY+L +++ K D KS + LGYS S+ +R +N+ +T
Sbjct: 907 EIWKGRKPTVKHFHIFGSPCYILADREQRRKMDPKSDAGIFLGYSTNSRAYRVFNSRTRT 966
Query: 1016 IEESIHVRFDDKLDSDQSKLVEKFADLSINVSDKGKAPEEVEPEEDEPEEEAGPSNSQTL 1075
+ ESI+V DD + + + E NV+D K+ E E + +E P+ +Q
Sbjct: 967 VMESINVVVDDLTPARKKDVEEDVRTSEDNVADTAKSAENAEKSDSTTDE---PNINQPD 1023
Query: 1076 KKS--RITAAHPKELILGNKDEPVRTRSAFRPSEETLLSLKGLVSLIEPKSIDEALQDKE 1133
K RI PKELI+G+ + V TRS E ++S VS IEPK++ EAL D+
Sbjct: 1024 KSPFIRIQKMQPKELIIGDPNRGVTTRSR----EIEIVSNSCFVSKIEPKNVKEALTDEF 1079
Query: 1134 WILAMEEELNQFSKNDVWSLVKKPESVHVIGTKWVFRNKLNEKGDVVRNKARLVAQGYSQ 1193
WI AM+EEL QF +N+VW LV +PE +VIGTKW+F+NK NE+G + RNKARLVAQGY+Q
Sbjct: 1080 WINAMQEELEQFKRNEVWELVPRPEGTNVIGTKWIFKNKTNEEGVITRNKARLVAQGYTQ 1139
Query: 1194 QEGIDYTETFAPVARLEAIRLLISFSVNHNIVLHQMDVKSAFLNGYISEEVYVHQPPGFE 1253
EG+D+ ETFAPVARLE+IRLL+ + L+QMDVKSAFLNGY++EE YV QP GF
Sbjct: 1140 IEGVDFDETFAPVARLESIRLLLGVACILKFKLYQMDVKSAFLNGYLNEEAYVEQPKGFV 1199
Query: 1254 DEKKPDHVFKLKKSLYGLKQAPRAWYERLSSFLLENEFVRGKVDTTLFCKTYKDDILIVQ 1313
D DHV++LKK+LYGLKQAPRAWYERL+ FL + + +G +D TLF K ++++I Q
Sbjct: 1200 DPTHLDHVYRLKKALYGLKQAPRAWYERLTEFLTQQGYRKGGIDKTLFVKQDAENLMIAQ 1259
Query: 1314 IYVDDIIFGSANQSLCKEFSEMMQAEFEMSMMGELKYFLGIQVDQTPEGTYIHQSKYTKE 1373
IYVDDI+FG + + + F MQ+EFEMS++GEL YFLG+QV Q + ++ QSKY K
Sbjct: 1260 IYVDDIVFGGMSNEMLRHFVPQMQSEFEMSLVGELHYFLGLQVKQMEDSIFLSQSKYAKN 1319
Query: 1374 LLKKFNMLESTVAKTPMHPTCILEKEDKSGKVCQKLYRGMIGSLLYLTASRPDILFSVHL 1433
++KKF M ++ +TP L K++ V Q LYR MIGSLLYLTASRPDI F+V +
Sbjct: 1320 IVKKFGMENASHKRTPAPTHLKLSKDEAGTSVDQNLYRSMIGSLLYLTASRPDITFAVGV 1379
Query: 1434 CARFQSDPRETHLTAVKRILRYLKGTTNLGLMYKKTSEYKLSGYCDADYAGDRTERKSTS 1493
CAR+Q++P+ +HL VKRIL+Y+ GT++ G+MY S+ L GYCDAD+AG +RK TS
Sbjct: 1380 CARYQANPKISHLNQVKRILKYVNGTSDYGIMYCHCSDSMLVGYCDADWAGSADDRKCTS 1439
Query: 1494 GNCQFLGSNLVSWASKRQSTIALSTAEAEYISAAICSTQMLWMKHQLEDYQILESNIPIY 1553
G C +LG+NL+SW SK+Q+ ++LSTAEAEYI+A +Q++WMK L++Y + + + +Y
Sbjct: 1440 GGCFYLGTNLISWFSKKQNCVSLSTAEAEYIAAGSSCSQLVWMKQMLKEYNVEQDVMTLY 1499
Query: 1554 CDNTAAISLSKNPILHSRAKHIEVKYHFIRDYVQKGVLLLKFVDTDHQWADIFTKSLAED 1613
CDN +AI++SKNP+ H+R KHI++++H+IRD V ++ L+ VDT+ Q ADIFTK+L +
Sbjct: 1500 CDNMSAINISKNPVQHNRTKHIDIRHHYIRDLVDDKIITLEHVDTEEQVADIFTKALDAN 1559
Query: 1614 RF 1615
+F
Sbjct: 1560 QF 1561
>UniRef100_O65147 Gag-pol polyprotein [Glycine max]
Length = 1550
Score = 965 bits (2494), Expect = 0.0
Identities = 491/1022 (48%), Positives = 673/1022 (65%), Gaps = 11/1022 (1%)
Query: 596 MLQISLIAPLKHQSWYLDSGCSRHMTGEKRMFRELKLKPGGEVGFGGNEKGKIIGTGTIC 655
++ SL A K + WYLDSGCSRHMTG K ++ V FG KGKI G G +
Sbjct: 523 VVHTSLRASAK-EDWYLDSGCSRHMTGVKEFLVNIEPCSTSYVTFGDGSKGKITGMGKLV 581
Query: 656 VDSSPCIDNVLLVDGLTHNLLSISQLADKGYDVIFNQKSCRAVSQIDGSVLFNSKRKNNI 715
D P ++ VLLV GLT NL+SISQL D+G++V F + C ++ ++ S+ K+N
Sbjct: 582 HDGLPSLNKVLLVKGLTANLISISQLCDEGFNVNFTKSECLVTNEKSEVLMKGSRSKDNC 641
Query: 716 YKIRLSELEAQNVKCLLSVNEEQWVWHRRLGHASMRKISQLSKLNLVRGLPNLKFASDAL 775
Y E + CL S +E +WH+R GH +R + ++ VRG+PNLK +
Sbjct: 642 YLWTPQETSYSST-CLFSKEDEVKIWHQRFGHLHLRGMKKIIDKGAVRGIPNLKIEEGRI 700
Query: 776 CEACQKGKFTKVPFKAKNVVSTSRPLELLHIDLFGPVKTESIGGKRYGMVIVDDYSRWTW 835
C CQ GK K+ + +TSR LELLH+DL GP++ ES+G KRY V+VDD+SR+TW
Sbjct: 701 CGECQIGKQVKMSNQKLQHQTTSRVLELLHMDLMGPMQVESLGRKRYAYVVVDDFSRFTW 760
Query: 836 VKFLTRKDESHAVFSTFIAQVQNEKACRIVRVRSDLGGEFENDKFESLFDSYGIAHDFSC 895
V F+ K ++ VF ++Q EK C I R+RSD G EFEN KF S GI H+FS
Sbjct: 761 VNFIREKSDTFEVFKELSLRLQREKDCVIKRIRSDHGREFENSKFTEFCTSEGITHEFSA 820
Query: 896 PRTPQQNGVVERKNRTLQEMARTMLQETGMAKHFWAETVNTACYIQNRISVRPILNKTPY 955
TPQQNG+VERKNRTLQE AR ML + + WAE +NTACYI NR+++R T Y
Sbjct: 821 AITPQQNGIVERKNRTLQEAARVMLHAKELPYNLWAEAMNTACYIHNRVTLRRGTPTTLY 880
Query: 956 ELWKNIKPNISYFHPFGCVCYVLNTKDRLHKFDAKSSKCLLLGYSDRSKGFRFYNTDAKT 1015
E+WK KP + +FH G CY+L +++ K D KS + LGYS S+ +R +N+ +T
Sbjct: 881 EIWKGRKPTVKHFHICGSPCYILADREQRRKMDPKSDAGIFLGYSTNSRAYRVFNSRTRT 940
Query: 1016 IEESIHVRFDDKLDSDQSKLVEKFADLSINVSDKGKAPEEVEPEEDEPEEEAGPSNSQTL 1075
+ ESI+V DD + + + E NV+D K+ E E + +E P+ +Q
Sbjct: 941 VMESINVVVDDLTPARKKDVEEDVRTSGDNVADTAKSAENAENSDSATDE---PNINQPD 997
Query: 1076 KKS--RITAAHPKELILGNKDEPVRTRSAFRPSEETLLSLKGLVSLIEPKSIDEALQDKE 1133
K+ RI HPKELI+G+ + V TRS E ++S VS IEPK++ EAL D+
Sbjct: 998 KRPSIRIQKMHPKELIIGDPNRGVTTRSR----EIEIISNSCFVSKIEPKNVKEALTDEF 1053
Query: 1134 WILAMEEELNQFSKNDVWSLVKKPESVHVIGTKWVFRNKLNEKGDVVRNKARLVAQGYSQ 1193
WI AM+EEL QF +N+VW LV +PE +VIGTKW+F+NK NE+G + RNKARLVAQGY+Q
Sbjct: 1054 WINAMQEELEQFKRNEVWELVPRPEGTNVIGTKWIFKNKTNEEGVITRNKARLVAQGYTQ 1113
Query: 1194 QEGIDYTETFAPVARLEAIRLLISFSVNHNIVLHQMDVKSAFLNGYISEEVYVHQPPGFE 1253
EG+D+ ETFAP ARLE+IRLL+ + L+QMDVKSAFLNGY++EE YV QP GF
Sbjct: 1114 IEGVDFDETFAPGARLESIRLLLGVACILKFKLYQMDVKSAFLNGYLNEEAYVEQPKGFV 1173
Query: 1254 DEKKPDHVFKLKKSLYGLKQAPRAWYERLSSFLLENEFVRGKVDTTLFCKTYKDDILIVQ 1313
D PDHV++LKK+LYGLKQAPRAWYERL+ FL + + +G +D TLF K ++++I Q
Sbjct: 1174 DPTHPDHVYRLKKALYGLKQAPRAWYERLTEFLTQQGYRKGGIDKTLFVKQDAENLMIAQ 1233
Query: 1314 IYVDDIIFGSANQSLCKEFSEMMQAEFEMSMMGELKYFLGIQVDQTPEGTYIHQSKYTKE 1373
IYVDDI+FG + + + F + MQ+EFEMS++GEL YFLG+QV Q + ++ QSKY K
Sbjct: 1234 IYVDDIVFGGMSNEMLRHFVQQMQSEFEMSLVGELTYFLGLQVKQMEDSIFLSQSKYAKN 1293
Query: 1374 LLKKFNMLESTVAKTPMHPTCILEKEDKSGKVCQKLYRGMIGSLLYLTASRPDILFSVHL 1433
++KKF M ++ +TP L K++ V Q LYR MIGSLLYLTASRPDI ++V
Sbjct: 1294 IVKKFGMENASHKRTPAPTHLKLSKDEAGTSVDQSLYRSMIGSLLYLTASRPDITYAVGG 1353
Query: 1434 CARFQSDPRETHLTAVKRILRYLKGTTNLGLMYKKTSEYKLSGYCDADYAGDRTERKSTS 1493
CAR+Q++P+ +HL VKRIL+Y+ GT++ G+MY S+ L GYCDAD+AG +RKST
Sbjct: 1354 CARYQANPKISHLNQVKRILKYVNGTSDYGIMYCHCSDSMLVGYCDADWAGSVDDRKSTF 1413
Query: 1494 GNCQFLGSNLVSWASKRQSTIALSTAEAEYISAAICSTQMLWMKHQLEDYQILESNIPIY 1553
G C +LG+N +SW SK+Q+ ++LSTAEAEYI+A +Q++WMK L++Y + + + +Y
Sbjct: 1414 GGCFYLGTNFISWFSKKQNCVSLSTAEAEYIAAGSSCSQLVWMKQMLKEYNVEQDVMTLY 1473
Query: 1554 CDNTAAISLSKNPILHSRAKHIEVKYHFIRDYVQKGVLLLKFVDTDHQWADIFTKSLAED 1613
CDN +AI++SKNP+ HSR KHI++++H+IRD V V+ L+ VDT+ Q ADIFTK+L +
Sbjct: 1474 CDNLSAINISKNPVQHSRTKHIDIRHHYIRDLVDDKVITLEHVDTEEQIADIFTKALDAN 1533
Query: 1614 RF 1615
+F
Sbjct: 1534 QF 1535
>UniRef100_Q60DR2 Putative polyprotein [Oryza sativa]
Length = 1577
Score = 946 bits (2445), Expect = 0.0
Identities = 498/1042 (47%), Positives = 668/1042 (63%), Gaps = 47/1042 (4%)
Query: 610 WYLDSGCSRHMTGEKRMFR--ELKLKPGGEVGFGGNEKGKIIGTGTICVDSSPCIDNVLL 667
W LDSGC++HMTG++ MF E+ +V FG N KGK+IG G I + + IDNV L
Sbjct: 549 WVLDSGCTQHMTGDRAMFTTFEVGRNEQEKVTFGDNSKGKVIGLGKIAISNDLSIDNVSL 608
Query: 668 VDGLTHNLLSISQLADKGYDVIFNQKSCRAVSQIDGSVLFNSKRKNNIYKIRLSELEAQN 727
V L NLLS++Q+ D F + S +D S +F R N+Y + + EA
Sbjct: 609 VKSLNFNLLSVAQICDLSLSCAFFPQEVIVSSLLDKSCVFKGFRYGNLYLVDFNSSEANL 668
Query: 728 VKCLLSVNEEQWVWHRRLGHASMRKISQLSKLNLVRGLPNLKFASDALCEACQKGKFTKV 787
CL++ W+WHRRL H M ++S+ SK +LV GL ++KF D LC ACQ GK
Sbjct: 669 KTCLVAKTSLGWLWHRRLAHVGMNQLSKFSKRDLVMGLKDVKFEKDKLCSACQAGKQVAC 728
Query: 788 PFKAKNVVSTSRPLELLHIDLFGPVKTESIGGKRYGMVIVDDYSRWTWVKFLTRKDESHA 847
K+++STS+PLELLH+DLF P +SIGG + +VIVDDYSR+TWV FL K
Sbjct: 729 SHPTKSIMSTSKPLELLHMDLFDPTTYKSIGGNSHCLVIVDDYSRYTWVFFLHDKSIVAD 788
Query: 848 VFSTFIAQVQNEKACRIVRVRSDLGGEFENDKFESLFDSYGIAHDFSCPRTPQQNGVVER 907
+F F + QNE +C +V++RS++G EF+N E D GI H+ +PQQNGVVER
Sbjct: 789 LFKKFAKRAQNEFSCTLVKIRSNIGSEFKNTNIEDYCDDLGIKHELFATYSPQQNGVVER 848
Query: 908 KNRTLQEMARTMLQETGMAKHFWAETVNTACYIQNRISVRPILNKTPYELWKNIKPNISY 967
KNRTL EMARTML E G++ FWAE +NTAC+ NR+ + +L KT YE+ KPNI+Y
Sbjct: 849 KNRTLIEMARTMLDEYGVSDSFWAEAINTACHATNRLYLHRVLKKTSYEVIVGRKPNIAY 908
Query: 968 FHPFGCVCYVLNTKDRLHKFDAKSSKCLLLGYSDRSKGFRFYNTDAKTIEESIHVRFDDK 1027
F FGC CY+ RL KF+++ + LLGY+ +SK +R YN + +EE+ V+FD+
Sbjct: 909 FRVFGCKCYIHRKGVRLTKFESRCDEGFLLGYASKSKAYRVYNKNKGIVEETADVQFDET 968
Query: 1028 LDSDQS-KLVEKFADLSI-----NVSDKGKAPEEVEPE-----EDEPEEEAGPSNSQTLK 1076
S + + ++ D + N+S P EVE + +DEP A PS +Q
Sbjct: 969 NGSQEGHENLDDVGDEGLMRVMKNMSIGDVKPIEVEDKPSTSTQDEPSTSAMPSQAQVEV 1028
Query: 1077 KSR--------------ITAAHPKELILGNKDEPVRTRSAFRPSEETLLSLKGLVSLIEP 1122
+ ++ HP + +LG+ + V+TRS ++ VS +E
Sbjct: 1029 EEEKAQEPPMPPRIHTALSKDHPIDQVLGDISKGVQTRSRVT----SICEHYSFVSCLER 1084
Query: 1123 KSIDEALQDKEWILAMEEELNQFSKNDVWSLVKKPESVHVIGTKWVFRNKLNEKGDVVRN 1182
K +DEAL D +W+ AM EEL F++N VW+LV++P +VIGTKWVFRNK +E G VVRN
Sbjct: 1085 KHVDEALCDPDWMNAMHEELKNFARNKVWTLVERPRDHNVIGTKWVFRNKQDENGLVVRN 1144
Query: 1183 KARLVAQGYSQQEGIDYTETFAPVARLEAIRLLISFSVNHNIVLHQMDVKSAFLNGYISE 1242
KARLVAQG++Q EG+D+ ETFAPVARLEAI +L++F+ +I L QMDVKSAFLN
Sbjct: 1145 KARLVAQGFTQVEGLDFGETFAPVARLEAICILLAFASCFDIKLFQMDVKSAFLN----- 1199
Query: 1243 EVYVHQPPGFEDEKKPDHVFKLKKSLYGLKQAPRAWYERLSSFLLENEFVRGKVDTTLFC 1302
D K P+HV+KL K+LYGL+QAPRAWYERL FLL +F GKVD TLF
Sbjct: 1200 -----------DTKYPNHVYKLSKALYGLRQAPRAWYERLRDFLLSKDFKIGKVDITLFT 1248
Query: 1303 KTYKDDILIVQIYVDDIIFGSANQSLCKEFSEMMQAEFEMSMMGELKYFLGIQVDQTPEG 1362
K DD + QIYVDDIIFGS N+ CKEF +MM EFEMSM+GEL +FLG+Q+ Q G
Sbjct: 1249 KIIGDDFFVYQIYVDDIIFGSTNEVFCKEFGDMMSREFEMSMIGELSFFLGLQIKQLKNG 1308
Query: 1363 TYIHQSKYTKELLKKFNMLESTVAKTPMHPTCILEKEDKSGKVCQKLYRGMIGSLLYLTA 1422
T++ Q+KY K+LLK+F + ++ KTPM L+ ++ V KLYR MIGSLLYLT
Sbjct: 1309 TFVSQTKYIKDLLKRFGLEDAKPIKTPMATNGHLDLDEGGKPVDLKLYRSMIGSLLYLTV 1368
Query: 1423 SRPDILFSVHLCARFQSDPRETHLTAVKRILRYLKGTTNLGLMYKKTSEYKLSGYCDADY 1482
SRPDI+FSV +CARFQ+ P+E HL AVKRILRYLK ++ +GL Y K +++KL GY D DY
Sbjct: 1369 SRPDIMFSVCMCARFQAAPKECHLVAVKRILRYLKHSSTIGLWYPKGAKFKLVGYSDPDY 1428
Query: 1483 AGDRTERKSTSGNCQFLGSNLVSWASKRQSTIALSTAEAEYISAAICSTQMLWMKHQLED 1542
AG + +RKSTS +CQ LG +LVSW+SK+Q+++ALSTAE EY+SA C Q+LWMK L D
Sbjct: 1429 AGCKVDRKSTSSSCQMLGRSLVSWSSKKQNSVALSTAETEYVSAGSCCAQLLWMKQTLLD 1488
Query: 1543 YQILESNIPIYCDNTAAISLSKNPILHSRAKHIEVKYHFIRDYVQKGVLLLKFVDTDHQW 1602
Y I + P+ CDN AI ++ NP+ HSR KHI++++HF+RD+V K +++ + T+ Q
Sbjct: 1489 YGISFTKTPLLCDNDGAIKIANNPVQHSRTKHIDIRHHFLRDHVAKCDIVISHIRTEDQL 1548
Query: 1603 ADIFTKSLAEDRFNFILKNLNM 1624
ADIFTK L E RF + LN+
Sbjct: 1549 ADIFTKPLDETRFCKLRNELNI 1570
>UniRef100_Q9C5V1 Gag/pol polyprotein [Arabidopsis thaliana]
Length = 1643
Score = 946 bits (2445), Expect = 0.0
Identities = 482/1035 (46%), Positives = 668/1035 (63%), Gaps = 28/1035 (2%)
Query: 610 WYLDSGCSRHMTGEKRMFRELKLKPGGEVGFGGNEKGKIIGTGTICVDSSPCIDNVLLVD 669
WY DSG SRHMTG + V FGG KG+I G G + P + NV V+
Sbjct: 614 WYFDSGASRHMTGSQANLNNYSSVKESNVMFGGGAKGRIKGKGDLTETEKPHLTNVYFVE 673
Query: 670 GLTHNLLSISQLADKGYDVIFNQKSCRAVSQIDGSVLFNSKRKNNIYKIRLSELEAQNVK 729
GLT NL+S+SQL D+G V FN+ C A ++ + + L + NN Y ++
Sbjct: 674 GLTANLISVSQLCDEGLTVSFNKVKCWATNERNQNTLTGVRTGNNCYMWEEPKI------ 727
Query: 730 CLLSVNEEQWVWHRRLGHASMRKISQLSKLNLVRGLPNLKFASDALCEACQKGKFTKVPF 789
CL + E+ +WH+RLGH + R +S+L +VRG+P LK +C AC +GK +V
Sbjct: 728 CLRAEKEDPVLWHQRLGHMNARSMSKLVNKEMVRGVPELKHIEKIVCGACNQGKQIRVQH 787
Query: 790 KAKNVVSTSRPLELLHIDLFGPVKTESIGGKRYGMVIVDDYSRWTWVKFLTRKDESHAVF 849
K + T++ L+L+H+DL GP++TESI GKRY V+VDD+SR+ WV+F+ K E+ F
Sbjct: 788 KRVEGIQTTQVLDLIHMDLMGPMQTESIAGKRYVFVLVDDFSRYAWVRFIREKSETANSF 847
Query: 850 STFIAQVQNEKACRIVRVRSDLGGEFENDKFESLFDSYGIAHDFSCPRTPQQNGVVERKN 909
Q++NEK I ++RSD GGEF N+ F S +S GI H +S PRTPQ NGVVERKN
Sbjct: 848 KILALQLKNEKKMGIKQIRSDRGGEFMNEAFNSFCESQGIFHQYSAPRTPQSNGVVERKN 907
Query: 910 RTLQEMARTMLQETGMAKHFWAETVNTACYIQNRISVRPILNKTPYELWKNIKPNISYFH 969
RTLQEMAR M+ G+ + FWAE ++TACY+ NR+ VR +KTPYE+WK KPN+SYF
Sbjct: 908 RTLQEMARAMIHGNGVPEKFWAEAISTACYVINRVYVRLGSDKTPYEIWKGKKPNLSYFR 967
Query: 970 PFGCVCYVLNTKDRLHKFDAKSSKCLLLGYSDRSKGFRFYNTDAKTIEESIHVRFDDKLD 1029
FGCVCY++N KD+L KFD++S + LGY+ S +R +N IEES++V FDD
Sbjct: 968 VFGCVCYIMNDKDQLGKFDSRSEEGFFLGYATNSLAYRVWNKQRGKIEESMNVVFDDGSM 1027
Query: 1030 SDQSKLVEKFADLSINVSDKGKAPEEVEPEEDEPEEEAGPSNSQTLKKSRITAAHPKELI 1089
+ +V + ++S+ ++ ++G + + + +++ H + I
Sbjct: 1028 PELQIIVRNRNEPQTSISNNHGEERNDNQFDNGDINKSGEESDEEVPPAQVHRDHASKDI 1087
Query: 1090 LGNKD-------------------EPVRTRSAFRPSEETLLSLKGLVSLIEPKSIDEALQ 1130
+G+ + R ++F EE + S VS++EPK++ EAL+
Sbjct: 1088 IGDPSGERVTRGVKQDYRQLAGIKQKHRVMASFACFEEIMFSC--FVSIVEPKNVKEALE 1145
Query: 1131 DKEWILAMEEELNQFSKNDVWSLVKKPESVHVIGTKWVFRNKLNEKGDVVRNKARLVAQG 1190
D WILAMEEEL +FS++ VW LV +P V+VIGTKW+F+NK +E G++ RNKARLVAQG
Sbjct: 1146 DHFWILAMEEELEEFSRHQVWDLVPRPPQVNVIGTKWIFKNKFDEVGNITRNKARLVAQG 1205
Query: 1191 YSQQEGIDYTETFAPVARLEAIRLLISFSVNHNIVLHQMDVKSAFLNGYISEEVYVHQPP 1250
Y+Q EG+D+ ETFAPVARLE IR L+ + LHQMDVK AFLNG I EEVYV QP
Sbjct: 1206 YTQVEGLDFDETFAPVARLECIRFLLGTACGMGFKLHQMDVKCAFLNGIIEEEVYVEQPK 1265
Query: 1251 GFEDEKKPDHVFKLKKSLYGLKQAPRAWYERLSSFLLENEFVRGKVDTTLFCKTYKDDIL 1310
GFE+ + P++V+KLKK+LYGLKQAPRAWYERL++FL+ + RG VD TLF K I+
Sbjct: 1266 GFENLEFPEYVYKLKKALYGLKQAPRAWYERLTTFLIVQGYTRGSVDKTLFVKNDVHGII 1325
Query: 1311 IVQIYVDDIIFGSANQSLCKEFSEMMQAEFEMSMMGELKYFLGIQVDQTPEGTYIHQSKY 1370
I+QIYVDDI+FG + L K F + M EF MSM+GELKYFLG+Q++QT EG I QS Y
Sbjct: 1326 IIQIYVDDIVFGGTSDKLVKTFVKTMTTEFRMSMVGELKYFLGLQINQTDEGITISQSTY 1385
Query: 1371 TKELLKKFNMLESTVAKTPMHPTCILEKEDKSGKVCQKLYRGMIGSLLYLTASRPDILFS 1430
+ L+K+F M S A TPM T L K++K KV +KLYRGMIGSLLYLTA+RPD+ S
Sbjct: 1386 AQNLVKRFGMCSSKPAPTPMSTTTKLFKDEKGVKVDEKLYRGMIGSLLYLTATRPDLCLS 1445
Query: 1431 VHLCARFQSDPRETHLTAVKRILRYLKGTTNLGLMYKKTSEYKLSGYCDADYAGDRTERK 1490
V LCAR+QS+P+ +HL AVKRI++Y+ GT N GL Y + + L GYCDAD+ G+ +R+
Sbjct: 1446 VGLCARYQSNPKASHLLAVKRIIKYVSGTINYGLNYTRDTSLVLVGYCDADWGGNLDDRR 1505
Query: 1491 STSGNCQFLGSNLVSWASKRQSTIALSTAEAEYISAAICSTQMLWMKHQLEDY-QILESN 1549
ST+G FLGSNL+SW SK+Q+ ++LS+ ++EYI+ C TQ+LWM+ DY
Sbjct: 1506 STTGGVFFLGSNLISWHSKKQNCVSLSSTQSEYIALGSCCTQLLWMRQMGLDYGMTFPDP 1565
Query: 1550 IPIYCDNTAAISLSKNPILHSRAKHIEVKYHFIRDYVQKGVLLLKFVDTDHQWADIFTKS 1609
+ + CDN +AI++SKNP+ HS KHI +++HF+R+ V++ + ++ V T+ Q DIFTK
Sbjct: 1566 LLVKCDNESAIAISKNPVQHSVTKHIAIRHHFVRELVEEKQITVEHVPTEIQLVDIFTKP 1625
Query: 1610 LAEDRFNFILKNLNM 1624
L + F + K+L +
Sbjct: 1626 LDLNTFVNLQKSLGI 1640
>UniRef100_Q8H851 Putative Zea mays retrotransposon Opie-2 [Oryza sativa]
Length = 2011
Score = 945 bits (2443), Expect = 0.0
Identities = 503/1044 (48%), Positives = 670/1044 (63%), Gaps = 49/1044 (4%)
Query: 610 WYLDSGCSRHMTGEKRMFRELKLKPGG----EVGFGGNEKGKIIGTGTICVDSSPCIDNV 665
W LDSGC++HMTG++ MF ++ GG +V FG N KGK+IG G I + + IDNV
Sbjct: 558 WVLDSGCTQHMTGDRAMFTTFEV--GGNEQEKVTFGDNSKGKVIGLGKIAISNDLSIDNV 615
Query: 666 LLVDGLTHNLLSISQLADKGYDVIFNQKSCRAVSQIDGSVLFNSKRKNNIYKIRLSELEA 725
LV L NLLS++Q+ D F + S +D S +F R N+Y + EA
Sbjct: 616 SLVKSLNFNLLSVAQICDLVLSCAFFPQEVIVSSLLDKSCVFKGFRYGNLYLGDFNSSEA 675
Query: 726 QNVKCLLSVNEEQWVWHRRLGHASMRKISQLSKLNLVRGLPNLKFASDALCEACQKGKFT 785
CL++ W+WHRRL H M ++S+LSK +LV GL ++KF D LC ACQ GK
Sbjct: 676 NLKTCLVAKTSLGWLWHRRLAHVGMNQLSKLSKRDLVVGLKDVKFEKDKLCSACQAGKQV 735
Query: 786 KVPFKAKNVVSTSRPLELLHIDLFGPVKTESIGGKRYGMVIVDDYSRWTWVKFLTRKDES 845
K+++STSRPLELLH++ FGP +SIGG + +VIVDDYSR+TW+ FL K
Sbjct: 736 ACSHPTKSIMSTSRPLELLHMEFFGPTTYKSIGGNSHCLVIVDDYSRYTWMFFLHDKSIV 795
Query: 846 HAVFSTFIAQVQNEKACRIVRVRSDLGGEFENDKFESLFDSYGIAHDFSCPRTPQQNGVV 905
+F F + QNE C +V++RSD G +F+N E D GI H+ S +PQQNGVV
Sbjct: 796 AELFKKFAKRGQNEFNCTLVKIRSDNGSKFKNTNIEDYCDDLGIKHELSATYSPQQNGVV 855
Query: 906 ERKNRTLQEMARTMLQETGMAKHFWAETVNTACYIQNRISVRPILNKTPYELWKNIKPNI 965
E KNRTL EMARTML E G++ FWAE +NTAC+ NR+ + +L KT YEL KPN+
Sbjct: 856 EMKNRTLIEMARTMLDEYGVSDSFWAEAINTACHATNRLYLHRLLKKTSYELIVGRKPNV 915
Query: 966 SYFHPFGCVCYVLNTKDRLHKFDAKSSKCLLLGYSDRSKGFRFYNTDAKTIEESIHVRFD 1025
+YF FGC CY+ RL KF+++ + LLGY+ SK +R YN + +EE+ V+FD
Sbjct: 916 AYFRVFGCKCYIYRKGVRLTKFESRCDEGFLLGYASNSKAYRVYNKNKGIVEETADVQFD 975
Query: 1026 DKLDSDQS-KLVEKFADLSI-----NVSDKGKAPEEVEPE-----EDEPEEEAGPSNSQT 1074
+ S + + ++ D + N+S P EVE + +DEP A PS +Q
Sbjct: 976 ETNGSQEGHENLDDVGDEGLMRAMKNMSIGDVKPIEVEDKPSTSTQDEPSTSATPSQAQV 1035
Query: 1075 LKK--------------SRITAAHPKELILGNKDEPVRTRSAFRPSEETLLSLKGLVSLI 1120
+ + ++ HP + +LG+ + V+TRS ++ VS +
Sbjct: 1036 EVEEEKAQDLPMPPRIHTALSKDHPIDQVLGDISKGVQTRSRV----ASICEHYSFVSCL 1091
Query: 1121 EPKSIDEALQDKEWILAMEEELNQFSKNDVWSLVKKPESVHVIGTKWVFRNKLNEKGDVV 1180
EPK +DEAL D +WI AM +ELN F++N VW+LV++ +VIGTKWVFRNK +E G VV
Sbjct: 1092 EPKHVDEALCDPDWINAMHKELNNFARNKVWTLVERLRDHNVIGTKWVFRNKQDENGLVV 1151
Query: 1181 RNKARLVAQGYSQQEGIDYTETFAPVARLEAIRLLISFSVNHNIVLHQMDVKSAFLNGYI 1240
RNKAR VAQG++Q EG+D+ ETFAPV RLEAI +L++F+ NI L QMDVKSAFLNG I
Sbjct: 1152 RNKARFVAQGFTQVEGLDFGETFAPVTRLEAICILLAFASCFNIKLFQMDVKSAFLNGEI 1211
Query: 1241 SEEVYVHQPPGFEDEKKPDHVFKLKKSLYGLKQAPRAWYERLSSFLLENEFVRGKVDTTL 1300
+E V+V QPPGFED K P+HV+KL K+LYGLKQAPRAWYERL FLL +F GKVDTTL
Sbjct: 1212 AELVFVEQPPGFEDPKYPNHVYKLSKALYGLKQAPRAWYERLRDFLLSKDFKIGKVDTTL 1271
Query: 1301 FCKTYKDDILIVQIYVDDIIFGSANQSLCKEFSEMMQAEFEMSMMGELKYFLGIQVDQTP 1360
F K DD + QIYVDDIIFG N+ CKEF +MM EFEMSM+GEL +F G+Q+ Q
Sbjct: 1272 FTKIIGDDFFVCQIYVDDIIFGCTNEVFCKEFGDMMSREFEMSMIGELSFFHGLQIKQLK 1331
Query: 1361 EGTYIHQSKYTKELLKKFNMLESTVAKTPMHPTCILEKEDKSGKVCQKLYRGMIGSLLYL 1420
+GT F + ++ KTPM L+ ++ V KLYR MIGSLLYL
Sbjct: 1332 DGT--------------FGLEDAKPIKTPMATNGHLDLDEGGKPVDLKLYRSMIGSLLYL 1377
Query: 1421 TASRPDILFSVHLCARFQSDPRETHLTAVKRILRYLKGTTNLGLMYKKTSEYKLSGYCDA 1480
TASRPDI+FSV +CARFQ+ P+E HL AVKRILRYLK ++ +GL Y K +++KL GY D+
Sbjct: 1378 TASRPDIMFSVCMCARFQAAPKECHLVAVKRILRYLKHSSTIGLWYPKGAKFKLVGYSDS 1437
Query: 1481 DYAGDRTERKSTSGNCQFLGSNLVSWASKRQSTIALSTAEAEYISAAICSTQMLWMKHQL 1540
DYAG + +RKSTSG+CQ LG +LVSW+SK+Q+ +AL AEAEY+SA C Q+LWMK L
Sbjct: 1438 DYAGCKVDRKSTSGSCQMLGRSLVSWSSKKQNFVALFIAEAEYVSAGSCCAQLLWMKQIL 1497
Query: 1541 EDYQILESNIPIYCDNTAAISLSKNPILHSRAKHIEVKYHFIRDYVQKGVLLLKFVDTDH 1600
DY I + P+ C+N +AI ++ NP+ HSR KHI++++HF+RD+V K +++ + T+
Sbjct: 1498 LDYGISFTKTPLLCENDSAIKIANNPVQHSRTKHIDIRHHFLRDHVAKCDIVISHIRTED 1557
Query: 1601 QWADIFTKSLAEDRFNFILKNLNM 1624
Q ADIFTK L E RF + LN+
Sbjct: 1558 QLADIFTKPLDETRFCKLRNELNL 1581
>UniRef100_Q852C7 Putative gag-pol polyprotein [Oryza sativa]
Length = 1969
Score = 938 bits (2425), Expect = 0.0
Identities = 492/1048 (46%), Positives = 661/1048 (62%), Gaps = 38/1048 (3%)
Query: 606 KHQSWYLDSGCSRHMTGEKRMFRELKLKPGGE-VGFGGNEKGKIIGTGTICVDSSPCIDN 664
+ SW +DSGCSRHMTGE + F L G E + FG G+++ GTI V+ + +
Sbjct: 750 RSNSWLVDSGCSRHMTGEAKWFTSLTRASGDETITFGDASSGRVMAKGTIKVNDKFMLKD 809
Query: 665 VLLVDGLTHNLLSISQLADKGYDVIFNQKSCRAVSQIDGSVLFNSKRKNNIYKIRLSELE 724
V LV L +NLLS+SQL D+ +V F + R + + V F+ R ++
Sbjct: 810 VALVSKLKYNLLSVSQLCDENLEVRFKKDRSRVLDASESPV-FDISRVGRVFFANFDSSA 868
Query: 725 AQNVKCLL-SVNEEQWVWHRRLGHASMRKISQLSKLNLVRGLPNLKFASDALCEACQKGK 783
+CL+ S N + + WHRRLGH +S++S ++L+RGLP LK D +C C+ GK
Sbjct: 869 PGPSRCLIASENRDLFFWHRRLGHIGFDHLSRISGMDLIRGLPKLKVQKDLVCAPCRHGK 928
Query: 784 FTKVPFKAKNVVSTSRPLELLHIDLFGPVKTESIGGKRYGMVIVDDYSRWTWVKFLTRKD 843
T K +V T P +LLH+D GP + +S+GGK Y +V+VDD+SR++WV FL K+
Sbjct: 929 MTSSSHKPVTMVMTDGPGQLLHMDTVGPARVQSVGGKWYVLVVVDDFSRYSWVYFLESKE 988
Query: 844 ESHAVFSTFIAQVQNEKACRIVRVRSDLGGEFENDKFESLFDSYGIAHDFSCPRTPQQNG 903
E+ F + + E + +RSD G EF+N FES DS G+ H FS P PQQNG
Sbjct: 989 ETFGFFQSLARSLALEFPGALRAIRSDNGSEFKNSAFESFCDSSGVEHQFSSPYVPQQNG 1048
Query: 904 VVERKNRTLQEMARTMLQETGMAKHFWAETVNTACYIQNRISVRPILNKTPYELWKNIKP 963
VVERKNRTL EMARTML E + FW E ++ AC+I NR+ +R IL+KTPYEL +P
Sbjct: 1049 VVERKNRTLVEMARTMLDEFTTPRKFWTEAISAACFISNRVFLRTILHKTPYELRFGRRP 1108
Query: 964 NISYFHPFGCVCYVLNTKDRLHKFDAKSSKCLLLGYSDRSKGFRFYNTDAKTIEESIHVR 1023
+S+ FGC C+VL + + L KF+++S + LGY+ S+ +R Y I E+ V
Sbjct: 1109 KVSHLRVFGCKCFVLKSGN-LDKFESRSLDGIFLGYATHSRAYRVYVLSTNKIVETCEVT 1167
Query: 1024 FDDKL---------------------DSDQSKLVEKFADLSINVSDKGKAPEEVEPEEDE 1062
FD+ D D + D + V + G +P P D
Sbjct: 1168 FDEASPGARPEISGVPDESIFVDEDSDDDDDDSIPPPLDSTPPVQETG-SPSTTSPSGDA 1226
Query: 1063 PE------EEAGPSNSQTLKKSRITAAHPKELILGNKDEPVRTRSAFRPSEETLLSLKGL 1116
P EE S I HP + ++G E V ++ L
Sbjct: 1227 PTTSSSAAEEIDGGTSGPTAPRHIQNRHPPDSMIGGLGERVTRNRSYE------LVNSAF 1280
Query: 1117 VSLIEPKSIDEALQDKEWILAMEEELNQFSKNDVWSLVKKPESVHVIGTKWVFRNKLNEK 1176
V+ EPK++ AL D+ W+ AM EEL F +N VWSLV+ P +VIGTKWVF+NKL E
Sbjct: 1281 VASFEPKNVCHALSDENWVNAMHEELENFERNKVWSLVEPPLGFNVIGTKWVFKNKLGED 1340
Query: 1177 GDVVRNKARLVAQGYSQQEGIDYTETFAPVARLEAIRLLISFSVNHNIVLHQMDVKSAFL 1236
G +VRNKARLVAQG++Q EG+D+ ETFAPVARLEAIR+L++F+ + L QMDVKSAFL
Sbjct: 1341 GSIVRNKARLVAQGFTQVEGLDFEETFAPVARLEAIRILLAFAASKGFKLFQMDVKSAFL 1400
Query: 1237 NGYISEEVYVHQPPGFEDEKKPDHVFKLKKSLYGLKQAPRAWYERLSSFLLENEFVRGKV 1296
NG I EEVYV QPPGFE+ K P+HVFKL+K+LYGLKQAPRAWYERL +FLL+N F G V
Sbjct: 1401 NGVIEEEVYVKQPPGFENPKFPNHVFKLEKALYGLKQAPRAWYERLKTFLLQNGFEMGAV 1460
Query: 1297 DTTLFCKTYKDDILIVQIYVDDIIFGSANQSLCKEFSEMMQAEFEMSMMGELKYFLGIQV 1356
D TLF D L+VQIYVDDIIFG ++ +L +FS++M EFEMSMMGEL +FLG+Q+
Sbjct: 1461 DKTLFTLHSGIDFLLVQIYVDDIIFGGSSHALVAQFSDVMSREFEMSMMGELTFFLGLQI 1520
Query: 1357 DQTPEGTYIHQSKYTKELLKKFNMLESTVAKTPMHPTCILEKEDKSGKVCQKLYRGMIGS 1416
QT EG ++HQ+KY+KELLKKF+M + TPM T L ++ +V Q+ YR MIGS
Sbjct: 1521 KQTKEGIFVHQTKYSKELLKKFDMADCKPIATPMATTSSLGPDEDGEEVDQREYRSMIGS 1580
Query: 1417 LLYLTASRPDILFSVHLCARFQSDPRETHLTAVKRILRYLKGTTNLGLMYKKTSEYKLSG 1476
LLYLTASRPDI FSV LCARFQ+ PR +H AVKRI RY+K T G+ Y +S +
Sbjct: 1581 LLYLTASRPDIHFSVCLCARFQASPRTSHRQAVKRIFRYIKSTLEYGIWYSCSSALSVRA 1640
Query: 1477 YCDADYAGDRTERKSTSGNCQFLGSNLVSWASKRQSTIALSTAEAEYISAAICSTQMLWM 1536
+ DAD+AG + +RKSTSG C FLG++LVSW+S++QS++A STAEAEY++AA +Q+LWM
Sbjct: 1641 FSDADFAGCKIDRKSTSGTCHFLGTSLVSWSSRKQSSVAQSTAEAEYVAAASACSQVLWM 1700
Query: 1537 KHQLEDYQILESNIPIYCDNTAAISLSKNPILHSRAKHIEVKYHFIRDYVQKGVLLLKFV 1596
L+DY + S +P+ CDNT+AI+++KNP+ HSR KHIE++YHF+RD V+KG ++L+FV
Sbjct: 1701 ISTLKDYGLSFSGVPLLCDNTSAINIAKNPVQHSRTKHIEIRYHFLRDNVEKGTIVLEFV 1760
Query: 1597 DTDHQWADIFTKSLAEDRFNFILKNLNM 1624
+++ Q ADIFTK L RF F+ L +
Sbjct: 1761 ESEKQLADIFTKPLDRSRFEFLRSELGV 1788
>UniRef100_Q8H7T1 Putative Zea mays retrotransposon Opie-2 [Oryza sativa]
Length = 2145
Score = 935 bits (2416), Expect = 0.0
Identities = 490/1048 (46%), Positives = 660/1048 (62%), Gaps = 38/1048 (3%)
Query: 606 KHQSWYLDSGCSRHMTGEKRMFRELKLKPGGE-VGFGGNEKGKIIGTGTICVDSSPCIDN 664
+ SW +DSGCSRHMTGE + F L E + FG G+++ GTI V+ + +
Sbjct: 750 RSNSWLVDSGCSRHMTGEAKWFTSLTRASSDETITFGDASSGRVMAKGTIKVNDKFMLKD 809
Query: 665 VLLVDGLTHNLLSISQLADKGYDVIFNQKSCRAVSQIDGSVLFNSKRKNNIYKIRLSELE 724
V LV L +NLLS+SQL D+ +V F + R + + V F+ R ++
Sbjct: 810 VALVSKLKYNLLSVSQLCDENLEVRFKKDRSRVLDASESPV-FDISRVGRVFFANFDSSA 868
Query: 725 AQNVKCLL-SVNEEQWVWHRRLGHASMRKISQLSKLNLVRGLPNLKFASDALCEACQKGK 783
+CL+ S N + + WHRRLGH +S++S ++L+RGLP LK D +C C+ GK
Sbjct: 869 PGPSRCLIASENRDLFFWHRRLGHIGFDHLSRISGMDLIRGLPKLKVPKDLVCAPCRHGK 928
Query: 784 FTKVPFKAKNVVSTSRPLELLHIDLFGPVKTESIGGKRYGMVIVDDYSRWTWVKFLTRKD 843
T K +V T P +LLH+D GP + +S+GGK Y +V+VDD+SR++WV FL K+
Sbjct: 929 MTSSSHKPVTMVMTDGPGQLLHMDTVGPARVQSVGGKWYVLVVVDDFSRYSWVYFLESKE 988
Query: 844 ESHAVFSTFIAQVQNEKACRIVRVRSDLGGEFENDKFESLFDSYGIAHDFSCPRTPQQNG 903
E+ F + + E + +RSD G EF+N FES DS G+ H FS P PQQNG
Sbjct: 989 ETFGFFQSLARSLALEFPGALRAIRSDNGSEFKNSAFESFCDSSGVEHQFSSPYVPQQNG 1048
Query: 904 VVERKNRTLQEMARTMLQETGMAKHFWAETVNTACYIQNRISVRPILNKTPYELWKNIKP 963
VVERKNRTL EMARTML E + FW E ++ AC+I NR+ +R IL+KTPYEL +P
Sbjct: 1049 VVERKNRTLVEMARTMLDEFTTPRKFWTEAISAACFISNRVFLRTILHKTPYELRFGRRP 1108
Query: 964 NISYFHPFGCVCYVLNTKDRLHKFDAKSSKCLLLGYSDRSKGFRFYNTDAKTIEESIHVR 1023
+S+ FGC C+VL + + L KF+++S + LGY+ S+ +R Y I E+ V
Sbjct: 1109 KVSHLRVFGCKCFVLKSGN-LDKFESRSLDGIFLGYATHSRAYRVYVLSTNKIVETCEVT 1167
Query: 1024 FDDKL---------------------DSDQSKLVEKFADLSINVSDKGKAPEEVEPEEDE 1062
FD+ D D + D + V + G +P P D
Sbjct: 1168 FDEASPGARPEISGVPDESIFVDEDSDDDDDDSIPPPLDSTPPVQETG-SPSTTSPSGDA 1226
Query: 1063 PE------EEAGPSNSQTLKKSRITAAHPKELILGNKDEPVRTRSAFRPSEETLLSLKGL 1116
P EE S I HP + ++G E V ++ L
Sbjct: 1227 PTTSSSAAEEIDGGTSGPTAPRHIQNRHPPDSMIGGLGERVTRNRSYE------LVNSAF 1280
Query: 1117 VSLIEPKSIDEALQDKEWILAMEEELNQFSKNDVWSLVKKPESVHVIGTKWVFRNKLNEK 1176
V+ EPK++ AL D+ W+ AM EEL F +N VWSLV+ P +VIGTKWVF+NKL E
Sbjct: 1281 VASFEPKNVCHALSDENWVNAMHEELENFERNKVWSLVEPPLGFNVIGTKWVFKNKLGED 1340
Query: 1177 GDVVRNKARLVAQGYSQQEGIDYTETFAPVARLEAIRLLISFSVNHNIVLHQMDVKSAFL 1236
G +VRNKARLVAQG++Q EG+D+ ETFAPVARLEAIR+L++F+ + L QMDVKSAFL
Sbjct: 1341 GSIVRNKARLVAQGFTQVEGLDFEETFAPVARLEAIRILLAFAASKGFKLFQMDVKSAFL 1400
Query: 1237 NGYISEEVYVHQPPGFEDEKKPDHVFKLKKSLYGLKQAPRAWYERLSSFLLENEFVRGKV 1296
NG I EEVYV QPPGFE+ K P+HVFKL+K+LYGLKQAPRAWYERL +FLL+N F G V
Sbjct: 1401 NGVIEEEVYVKQPPGFENPKFPNHVFKLEKALYGLKQAPRAWYERLKTFLLQNGFEMGAV 1460
Query: 1297 DTTLFCKTYKDDILIVQIYVDDIIFGSANQSLCKEFSEMMQAEFEMSMMGELKYFLGIQV 1356
D TLF D L+VQIYVDDIIFG ++ +L +FS++M EFEMSMMGEL +FLG+Q+
Sbjct: 1461 DKTLFTLHSGIDFLLVQIYVDDIIFGGSSHALVAQFSDVMSREFEMSMMGELTFFLGLQI 1520
Query: 1357 DQTPEGTYIHQSKYTKELLKKFNMLESTVAKTPMHPTCILEKEDKSGKVCQKLYRGMIGS 1416
QT EG ++HQ+KY+KELLKKF+M + TPM T L ++ +V Q+ YR MIGS
Sbjct: 1521 KQTKEGIFVHQTKYSKELLKKFDMADCKPIATPMATTSSLGPDEDGEEVDQREYRSMIGS 1580
Query: 1417 LLYLTASRPDILFSVHLCARFQSDPRETHLTAVKRILRYLKGTTNLGLMYKKTSEYKLSG 1476
LLYLTASRPDI FSV LCARFQ+ PR +H AVKR+ RY+K T G+ Y +S +
Sbjct: 1581 LLYLTASRPDIHFSVCLCARFQASPRTSHRQAVKRMFRYIKSTLEYGIWYSCSSALSVRA 1640
Query: 1477 YCDADYAGDRTERKSTSGNCQFLGSNLVSWASKRQSTIALSTAEAEYISAAICSTQMLWM 1536
+ DAD+AG + +RKSTSG C FLG++LVSW+S++QS++A STAEAEY++AA +Q+LWM
Sbjct: 1641 FSDADFAGCKIDRKSTSGTCHFLGTSLVSWSSRKQSSVAQSTAEAEYVAAASACSQVLWM 1700
Query: 1537 KHQLEDYQILESNIPIYCDNTAAISLSKNPILHSRAKHIEVKYHFIRDYVQKGVLLLKFV 1596
L+DY + S +P+ CDNT+AI+++KNP+ HSR KHIE++YHF+RD V+KG ++L+FV
Sbjct: 1701 ISTLKDYGLSFSGVPLLCDNTSAINIAKNPVQHSRTKHIEIRYHFLRDNVEKGTIVLEFV 1760
Query: 1597 DTDHQWADIFTKSLAEDRFNFILKNLNM 1624
+++ Q ADIFTK L RF F+ L +
Sbjct: 1761 ESEKQLADIFTKPLDRSRFEFLRSELGV 1788
>UniRef100_Q7XPI7 OSJNBb0004A17.2 protein [Oryza sativa]
Length = 1877
Score = 935 bits (2416), Expect = 0.0
Identities = 490/1048 (46%), Positives = 660/1048 (62%), Gaps = 38/1048 (3%)
Query: 606 KHQSWYLDSGCSRHMTGEKRMFRELKLKPGGE-VGFGGNEKGKIIGTGTICVDSSPCIDN 664
+ SW +DSGCSRHMTGE + F L E + FG G+++ GTI V+ + +
Sbjct: 832 RSNSWLVDSGCSRHMTGEAKWFTSLTRASSDETITFGDASSGRVMAKGTIKVNDKFMLKD 891
Query: 665 VLLVDGLTHNLLSISQLADKGYDVIFNQKSCRAVSQIDGSVLFNSKRKNNIYKIRLSELE 724
V LV L +NLLS+SQL D+ +V F + R + + V F+ R ++
Sbjct: 892 VALVSKLKYNLLSVSQLCDENLEVRFKKDRSRVLDASESPV-FDISRVGRVFFANFDSSA 950
Query: 725 AQNVKCLL-SVNEEQWVWHRRLGHASMRKISQLSKLNLVRGLPNLKFASDALCEACQKGK 783
+CL+ S N + + WHRRLGH +S++S ++L+RGLP LK D +C C+ GK
Sbjct: 951 PGPSRCLIASENRDLFFWHRRLGHIGFDHLSRISGMDLIRGLPKLKVPKDLVCAPCRHGK 1010
Query: 784 FTKVPFKAKNVVSTSRPLELLHIDLFGPVKTESIGGKRYGMVIVDDYSRWTWVKFLTRKD 843
T K +V T P +LLH+D GP + +S+GGK Y +V+VDD+SR++WV FL K+
Sbjct: 1011 MTSSSHKPVTMVMTDGPGQLLHMDTVGPARVQSVGGKWYVLVVVDDFSRYSWVYFLESKE 1070
Query: 844 ESHAVFSTFIAQVQNEKACRIVRVRSDLGGEFENDKFESLFDSYGIAHDFSCPRTPQQNG 903
E+ F + + E + +RSD G EF+N FES DS G+ H FS P PQQNG
Sbjct: 1071 ETFGFFQSLARSLALEFPGALRAIRSDNGSEFKNSAFESFCDSSGVEHQFSSPYVPQQNG 1130
Query: 904 VVERKNRTLQEMARTMLQETGMAKHFWAETVNTACYIQNRISVRPILNKTPYELWKNIKP 963
VVERKNRTL EMARTML E + FW E ++ AC+I NR+ +R IL+KTPYEL +P
Sbjct: 1131 VVERKNRTLVEMARTMLDEFTTPRKFWTEAISAACFISNRVFLRTILHKTPYELRFGRRP 1190
Query: 964 NISYFHPFGCVCYVLNTKDRLHKFDAKSSKCLLLGYSDRSKGFRFYNTDAKTIEESIHVR 1023
+S+ FGC C+VL + + L KF+++S + LGY+ S+ +R Y I E+ V
Sbjct: 1191 KVSHLRVFGCKCFVLKSGN-LDKFESRSLDGIFLGYATHSRAYRVYVLSTNKIVETCEVT 1249
Query: 1024 FDDKL---------------------DSDQSKLVEKFADLSINVSDKGKAPEEVEPEEDE 1062
FD+ D D + D + V + G +P P D
Sbjct: 1250 FDEASPGARPEISGVPDESIFVDEDSDDDDDDSIPPPLDSTPPVQETG-SPSTTSPSGDA 1308
Query: 1063 PE------EEAGPSNSQTLKKSRITAAHPKELILGNKDEPVRTRSAFRPSEETLLSLKGL 1116
P EE S I HP + ++G E V ++ L
Sbjct: 1309 PTTSSSAAEEIDGGTSGPTAPRHIQNRHPPDSMIGGLGERVTRNRSYE------LVNSAF 1362
Query: 1117 VSLIEPKSIDEALQDKEWILAMEEELNQFSKNDVWSLVKKPESVHVIGTKWVFRNKLNEK 1176
V+ EPK++ AL D+ W+ AM EEL F +N VWSLV+ P +VIGTKWVF+NKL E
Sbjct: 1363 VASFEPKNVCHALSDENWVNAMHEELENFERNKVWSLVEPPLGFNVIGTKWVFKNKLGED 1422
Query: 1177 GDVVRNKARLVAQGYSQQEGIDYTETFAPVARLEAIRLLISFSVNHNIVLHQMDVKSAFL 1236
G +VRNKARLVAQG++Q EG+D+ ETFAPVARLEAIR+L++F+ + L QMDVKSAFL
Sbjct: 1423 GSIVRNKARLVAQGFTQVEGLDFEETFAPVARLEAIRILLAFAASKGFKLFQMDVKSAFL 1482
Query: 1237 NGYISEEVYVHQPPGFEDEKKPDHVFKLKKSLYGLKQAPRAWYERLSSFLLENEFVRGKV 1296
NG I EEVYV QPPGFE+ K P+HVFKL+K+LYGLKQAPRAWYERL +FLL+N F G V
Sbjct: 1483 NGVIEEEVYVKQPPGFENPKFPNHVFKLEKALYGLKQAPRAWYERLKTFLLQNGFEMGAV 1542
Query: 1297 DTTLFCKTYKDDILIVQIYVDDIIFGSANQSLCKEFSEMMQAEFEMSMMGELKYFLGIQV 1356
D TLF D L+VQIYVDDIIFG ++ +L +FS++M EFEMSMMGEL +FLG+Q+
Sbjct: 1543 DKTLFTLHSGIDFLLVQIYVDDIIFGGSSHALVAQFSDVMSREFEMSMMGELTFFLGLQI 1602
Query: 1357 DQTPEGTYIHQSKYTKELLKKFNMLESTVAKTPMHPTCILEKEDKSGKVCQKLYRGMIGS 1416
QT EG ++HQ+KY+KELLKKF+M + TPM T L ++ +V Q+ YR MIGS
Sbjct: 1603 KQTKEGIFVHQTKYSKELLKKFDMADCKPIATPMATTSSLGPDEDGEEVDQREYRSMIGS 1662
Query: 1417 LLYLTASRPDILFSVHLCARFQSDPRETHLTAVKRILRYLKGTTNLGLMYKKTSEYKLSG 1476
LLYLTASRPDI FSV LCARFQ+ PR +H AVKR+ RY+K T G+ Y +S +
Sbjct: 1663 LLYLTASRPDIHFSVCLCARFQASPRTSHRQAVKRMFRYIKSTLEYGIWYSCSSALSVRA 1722
Query: 1477 YCDADYAGDRTERKSTSGNCQFLGSNLVSWASKRQSTIALSTAEAEYISAAICSTQMLWM 1536
+ DAD+AG + +RKSTSG C FLG++LVSW+S++QS++A STAEAEY++AA +Q+LWM
Sbjct: 1723 FSDADFAGCKIDRKSTSGTCHFLGTSLVSWSSRKQSSVAQSTAEAEYVAAASACSQVLWM 1782
Query: 1537 KHQLEDYQILESNIPIYCDNTAAISLSKNPILHSRAKHIEVKYHFIRDYVQKGVLLLKFV 1596
L+DY + S +P+ CDNT+AI+++KNP+ HSR KHIE++YHF+RD V+KG ++L+FV
Sbjct: 1783 ISTLKDYGLSFSGVPLLCDNTSAINIAKNPVQHSRTKHIEIRYHFLRDNVEKGTIVLEFV 1842
Query: 1597 DTDHQWADIFTKSLAEDRFNFILKNLNM 1624
+++ Q ADIFTK L RF F+ L +
Sbjct: 1843 ESEKQLADIFTKPLDRSRFEFLRSELGV 1870
>UniRef100_Q7XP45 OSJNBa0063G07.6 protein [Oryza sativa]
Length = 1539
Score = 927 bits (2396), Expect = 0.0
Identities = 495/1044 (47%), Positives = 664/1044 (63%), Gaps = 64/1044 (6%)
Query: 610 WYLDSGCSRHMTGEKRMFRELKLKPGG----EVGFGGNEKGKIIGTGTICVDSSPCIDNV 665
W LDSGC++HMTG++ MF ++ GG +V F N K K+IG G I + + IDNV
Sbjct: 524 WVLDSGCTQHMTGDRAMFTTFEV--GGNEQEKVTFVDNSKRKVIGLGKIAISNDLSIDNV 581
Query: 666 LLVDGLTHNLLSISQLADKGYDVIFNQKSCRAVSQIDGSVLFNSKRKNNIYKIRLSELEA 725
V L NLLS++Q+ D G F + S +D S +F R N+Y + + EA
Sbjct: 582 SFVKSLNFNLLSVAQICDLGLSCAFFPQEVIVSSLLDKSCVFKGFRYGNLYFVDFNSSEA 641
Query: 726 QNVKCLLSVNEEQWVWHRRLGHASMRKISQLSKLNLVRGLPNLKFASDALCEACQKGKFT 785
CL++ W+WHRRL H M ++S+LSK +LV GL ++KF D LC ACQ GK
Sbjct: 642 NLKTCLVAKTSLGWLWHRRLAHVGMNQLSKLSKRDLVVGLKDVKFEKDKLCSACQAGKQV 701
Query: 786 KVPFKAKNVVSTSRPLELLHIDLFGPVKTESIGGKRYGMVIVDDYSRWTWVKFLTRKDES 845
K+++STSRPLELLH+DLFGP +SIGG + +VIVDDYSR+TWV FL K
Sbjct: 702 ACSHPTKSIMSTSRPLELLHMDLFGPTTYKSIGGNSHCLVIVDDYSRYTWVFFLHDKSIV 761
Query: 846 HAVFSTFIAQVQNEKACRIVRVRSDLGGEFENDKFESLFDSYGIAHDFSCPRTPQQNGVV 905
+F + QNE +C +V++RSD G EF+N E D GI H+ S +PQQNGVV
Sbjct: 762 AELFKKIAKRAQNEFSCTLVKIRSDNGSEFKNTNIEDYCDDLGIKHELSATYSPQQNGVV 821
Query: 906 ERKNRTLQEMARTMLQETGMAKHFWAETVNTACYIQNRISVRPILNKTPYELWKNIKPNI 965
ERKNRTL EMARTML E G++ FWAE +NTAC+ NR + +L T YEL KPN+
Sbjct: 822 ERKNRTLIEMARTMLDEYGVSDSFWAEAINTACHATNRFYLHRLLKNTSYELIVGRKPNV 881
Query: 966 SYFHPFGCVCYVLNTKDRLHKFDAKSSKCLLLGYSDRSKGFRFYNTDAKTIEESIHVRFD 1025
+YF FGC CY+ RL KF+++ + LLGY+ SK +R YN + T+EE+ V+FD
Sbjct: 882 AYFRVFGCKCYIYRKGVRLTKFESRCDEGFLLGYASNSKAYRVYNKNKGTVEETADVQFD 941
Query: 1026 DKLDSDQS-KLVEKFADLSI-----NVSDKGKAPEEVEPE-----EDEPEEEAGPSNSQT 1074
+ S + + ++ D + N+S P EVE + +DEP A PS +Q
Sbjct: 942 ETNGSQEGHENLDDVGDEGLIRAMKNMSFGDVKPIEVEDKPSTSTQDEPSTFATPSQAQV 1001
Query: 1075 LKK--------------SRITAAHPKELILGNKDEPVRTRSAFRPSEETLLSLKGLVSLI 1120
+ + ++ HP + +LG+ + V+T S ++ VS +
Sbjct: 1002 EVEEEKAQDPPIPPRIHTTLSKDHPIDQVLGDISKGVQTLSRV----ASICEHYSFVSCL 1057
Query: 1121 EPKSIDEALQDKEWILAMEEELNQFSKNDVWSLVKKPESVHVIGTKWVFRNKLNEKGDVV 1180
EPK +DEAL D +W+ AM EELN F++N VW+LV++P +VIGTKWVFRNK +E G VV
Sbjct: 1058 EPKHVDEALCDPDWMNAMHEELNNFARNKVWTLVERPRDHNVIGTKWVFRNKQDENGLVV 1117
Query: 1181 RNKARLVAQGYSQQEGIDYTETFAPVARLEAIRLLISFSVNHNIVLHQMDVKSAFLNGYI 1240
RNKARLVAQG++Q EG+D+ ETFAPVARLEAI +L++F+ +I L QMDVKSAFLNG I
Sbjct: 1118 RNKARLVAQGFTQVEGLDFGETFAPVARLEAICILLAFASWFDIKLFQMDVKSAFLNGEI 1177
Query: 1241 SEEVYVHQPPGFEDEKKPDHVFKLKKSLYGLKQAPRAWYERLSSFLLENEFVRGKVDTTL 1300
+E V+V QPPGFED K P+H FK+ GKVDTTL
Sbjct: 1178 AELVFVEQPPGFEDPKYPNHDFKI-----------------------------GKVDTTL 1208
Query: 1301 FCKTYKDDILIVQIYVDDIIFGSANQSLCKEFSEMMQAEFEMSMMGELKYFLGIQVDQTP 1360
F K DD + QIYVDDIIFGS N+ CKEF +MM EFEMSM+GEL +FLG+Q+ Q
Sbjct: 1209 FTKIIGDDFFVCQIYVDDIIFGSTNEVFCKEFGDMMSREFEMSMIGELSFFLGLQIKQLK 1268
Query: 1361 EGTYIHQSKYTKELLKKFNMLESTVAKTPMHPTCILEKEDKSGKVCQKLYRGMIGSLLYL 1420
+GT++ Q+KY K+LLK+F + ++ KTPM L+ ++ V KLYR MIGSLLYL
Sbjct: 1269 DGTFVSQTKYIKDLLKRFGLEDAKPIKTPMATNGHLDLDEGGKPVDLKLYRSMIGSLLYL 1328
Query: 1421 TASRPDILFSVHLCARFQSDPRETHLTAVKRILRYLKGTTNLGLMYKKTSEYKLSGYCDA 1480
TASRPDI+FSV +CA FQ+ P+E HL AVKRILRYLK ++ +GL Y K +++KL GY D+
Sbjct: 1329 TASRPDIMFSVCMCAWFQAAPKECHLVAVKRILRYLKYSSTIGLWYPKGAKFKLVGYSDS 1388
Query: 1481 DYAGDRTERKSTSGNCQFLGSNLVSWASKRQSTIALSTAEAEYISAAICSTQMLWMKHQL 1540
DYAG + +R STSG+CQ LG +LVSW+SK+Q+++ALSTAEAEY+SA C Q+LWMK L
Sbjct: 1389 DYAGCKVDRNSTSGSCQMLGRSLVSWSSKKQNSVALSTAEAEYVSAGSCCAQLLWMKQTL 1448
Query: 1541 EDYQILESNIPIYCDNTAAISLSKNPILHSRAKHIEVKYHFIRDYVQKGVLLLKFVDTDH 1600
DY I + P+ CDN +AI ++ NP+ HSR KHI++++HF+RD+V K +++ + T+
Sbjct: 1449 LDYGISFTKTPLLCDNDSAIKIANNPVQHSRTKHIDIRHHFLRDHVAKCDIVISHIRTED 1508
Query: 1601 QWADIFTKSLAEDRFNFILKNLNM 1624
Q ADIFTK L E RF + LN+
Sbjct: 1509 QLADIFTKPLDETRFCKLRNELNV 1532
>UniRef100_Q7XET2 Hypothetical protein [Oryza sativa]
Length = 1419
Score = 918 bits (2372), Expect = 0.0
Identities = 480/1036 (46%), Positives = 663/1036 (63%), Gaps = 36/1036 (3%)
Query: 591 RTRLSMLQISLIAPLKHQSWYLDSGCSRHMTGEKRMFRELKLKPGGE-VGFGGNEKGKII 649
++++ +L+++L+A K W +DSG SRHMTG+K F LK E + F ++
Sbjct: 411 QSKVLVLEVALVAR-KENVWIVDSGDSRHMTGDKNWFSSLKKASKTESIVFADASTSAVV 469
Query: 650 GTGTICVDSSPCIDNVLLVDGLTHNLLSISQLADKGYDVIFNQKSCRAVSQIDGSVLFNS 709
TG + V+ + NV LV L +NLLS+SQ+ D+ ++V F + + SVL N
Sbjct: 470 ATGLVKVNEKFELKNVALVKDLKYNLLSVSQIVDENFEVHFKKTGSKVFDSCGDSVL-NI 528
Query: 710 KRKNNIYKIRLSELEAQNVKCLLS-VNEEQWVWHRRLGHASMRKISQLSKLNLVRGLPNL 768
R ++K + + CL++ +++ WHRRLGH +++LS L+LVRGL L
Sbjct: 529 SRYERVFKADFENSVSLVITCLVAKFDKDVMFWHRRLGHVGFDHLTRLSGLDLVRGLSKL 588
Query: 769 KFASDALCEACQKGKFTKVPFKAKNVVSTSRPLELLHIDLFGPVKTESIGGKRYGMVIVD 828
K D +C C+ K V T P +LLH+D GP + +S+GGK Y +VIVD
Sbjct: 589 KKDLDLICTPCRHAKMVSTSHAPIVSVMTDAPGQLLHMDTVGPARVQSVGGKWYVLVIVD 648
Query: 829 DYSRWTWVKFLTRKDESHAVFSTFIAQVQNEKACRIVRVRSDLGGEFENDKFESLFDSYG 888
D+SR++WV F+T KDE+ F +++ E + R+RSD GGEF+N FE + G
Sbjct: 649 DFSRYSWVFFMTTKDEAFQHFRGLFLRLELEFPGSLKRIRSDNGGEFKNASFEQFCNERG 708
Query: 889 IAHDFSCPRTPQQNGVVERKNRTLQEMARTMLQETGMAKHFWAETVNTACYIQNRISVRP 948
+ H+FS PR PQQN VVERKNR L EMARTML E + FWAE +NTACYI NR+ +R
Sbjct: 709 LEHEFSSPRVPQQNSVVERKNRVLVEMARTMLDEYKTTRKFWAEAINTACYISNRVFLRS 768
Query: 949 ILNKTPYELWKNIKPNISYFHPFGCVCYVLNTKDRLHKFDAKSSKCLLLGYSDRSKGFRF 1008
L KT YEL +P +S+ FGC C+VL + + L KF+A+S+ L LGY ++G+R
Sbjct: 769 KLGKTSYELRFGHQPKVSHLRVFGCKCFVLKSGN-LDKFEARSTDGLFLGYPAHTRGYRV 827
Query: 1009 YNTDAKTIEESIHVRFDDKLDSDQSKLVEKFADLSINVSDKGKAPEEVEPEEDEPEEEAG 1068
I E+ V FD+ + + + + +G+ E E D +++
Sbjct: 828 LILGTNKIVETCEVSFDEASPGTRPDIAGTLSQV------QGEDGRIFEDESDYDDDDEV 881
Query: 1069 PSNSQTLKKSRITAAHPKELILGNKDEPVRTRSAFRPSEETLLSLKGLVSLIEPKSIDEA 1128
S + +S++T V SAF V+ EPK + A
Sbjct: 882 GSAGERTTRSKVTT------------HDVCANSAF-------------VASFEPKDVSHA 916
Query: 1129 LQDKEWILAMEEELNQFSKNDVWSLVKKPESVHVIGTKWVFRNKLNEKGDVVRNKARLVA 1188
L D+ WI AM EEL F +N VW+LV+ P +VIGTKWVF+NK NE G +VRNKARLVA
Sbjct: 917 LTDESWINAMHEELENFERNKVWTLVEPPSGHNVIGTKWVFKNKQNEDGLIVRNKARLVA 976
Query: 1189 QGYSQQEGIDYTETFAPVARLEAIRLLISFSVNHNIVLHQMDVKSAFLNGYISEEVYVHQ 1248
QG++Q EG+D+ ETFAPVAR+EAIRLL++F+ + L+QMDVKSAFLNG+I EEVYV Q
Sbjct: 977 QGFTQVEGLDFDETFAPVARIEAIRLLLAFAASKGFKLYQMDVKSAFLNGFIQEEVYVKQ 1036
Query: 1249 PPGFEDEKKPDHVFKLKKSLYGLKQAPRAWYERLSSFLLENEFVRGKVDTTLFCKTYKDD 1308
PPGFE+ P+HVFKL K+LYGLKQAPRAWY+RL +FLL F GKVD TLF + D+
Sbjct: 1037 PPGFENPDFPNHVFKLSKALYGLKQAPRAWYDRLKNFLLAKGFTMGKVDKTLFVLKHGDN 1096
Query: 1309 ILIVQIYVDDIIFGSANQSLCKEFSEMMQAEFEMSMMGELKYFLGIQVDQTPEGTYIHQS 1368
L VQIYVDDIIFG + ++ +F+E M+ EFEMSMMGEL YFLG+Q+ QTP+GT++HQ+
Sbjct: 1097 QLFVQIYVDDIIFGCSTHAVVVDFAETMRREFEMSMMGELSYFLGLQIKQTPQGTFVHQT 1156
Query: 1369 KYTKELLKKFNMLESTVAKTPMHPTCILEKEDKSGKVCQKLYRGMIGSLLYLTASRPDIL 1428
KYTK+LL++F M TP+ T +L+ ++ V QK YR MIGSLLYLTASRPDI
Sbjct: 1157 KYTKDLLRRFKMENCKPISTPIGSTAVLDPDEDGEAVDQKEYRSMIGSLLYLTASRPDIQ 1216
Query: 1429 FSVHLCARFQSDPRETHLTAVKRILRYLKGTTNLGLMYKKTSEYKLSGYCDADYAGDRTE 1488
F+V LCARFQ+ PR +H AVKRI+RYL T G+ Y +S LSGY DAD+ G R +
Sbjct: 1217 FAVCLCARFQASPRASHRQAVKRIMRYLNHTLEFGIWYSTSSSICLSGYSDADFGGCRID 1276
Query: 1489 RKSTSGNCQFLGSNLVSWASKRQSTIALSTAEAEYISAAICSTQMLWMKHQLEDYQILES 1548
RKSTSG C FLG++L++W+S++QS++A STAE+EY++AA C +Q+LW+ L+DY I
Sbjct: 1277 RKSTSGTCHFLGTSLIAWSSRKQSSVAQSTAESEYVAAASCCSQILWLLSTLKDYGITFE 1336
Query: 1549 NIPIYCDNTAAISLSKNPILHSRAKHIEVKYHFIRDYVQKGVLLLKFVDTDHQWADIFTK 1608
+P++CDNT+AI+++KNP+ HSR KHI++++HF+RD+V+KG + L+F+DT Q ADIFTK
Sbjct: 1337 KVPLFCDNTSAINIAKNPVQHSRTKHIDIRFHFLRDHVEKGDVELQFLDTKLQIADIFTK 1396
Query: 1609 SLAEDRFNFILKNLNM 1624
L +RF F+ L +
Sbjct: 1397 PLDSNRFAFLRGELGV 1412
>UniRef100_Q850V9 Putative polyprotein [Oryza sativa]
Length = 1128
Score = 889 bits (2298), Expect = 0.0
Identities = 464/935 (49%), Positives = 624/935 (66%), Gaps = 31/935 (3%)
Query: 717 KIRLSELEAQNVKCLLSVNEEQWVWHRRLGHASMRKISQLSKLNLVRGLPNLKFASDALC 776
K+ + +NV L++ W+WHRRL H M ++S+LSK +LV GL ++KF D LC
Sbjct: 191 KVTFGDNSKRNVIGLVAKTSFGWLWHRRLAHVGMNQLSKLSKRDLVVGLKDVKFEKDKLC 250
Query: 777 EACQKGKFTKVPFKAKNVVSTSRPLELLHIDLFGPVKTESIGGKRYGMVIVDDYSRWTWV 836
ACQ K K+++STSRPLELLH+DLFGP +SIGG + +VIVDDYS +TWV
Sbjct: 251 SACQASKQVACSHPTKSIMSTSRPLELLHMDLFGPTTYKSIGGNSHCLVIVDDYSCYTWV 310
Query: 837 KFLTRKDESHAVFSTFIAQVQNEKACRIVRVRSDLGGEFENDKFESLFDSYGIAHDFSCP 896
FL K +F F + QNE +C +V++RSD G +F+N E D I H+ S
Sbjct: 311 FFLHDKCIVAELFKKFAKRAQNEFSCTLVKIRSDNGSKFKNTNIEDYCDDLSIKHELSAT 370
Query: 897 RTPQQNGVVERKNRTLQEMARTMLQETGMAKHFWAETVNTACYIQNRISVRPILNKTPYE 956
+PQQNGVVERKNRTL EMARTML E G++ FWAE +NTAC+ NR+ + +L KT YE
Sbjct: 371 YSPQQNGVVERKNRTLIEMARTMLDEYGVSDSFWAEAINTACHATNRLYLHRLLKKTSYE 430
Query: 957 LWKNIKPNISYFHPFGCVCYVLNTKDRLHKFDAKSSKCLLLGYSDRSKGFRFYNTDAKTI 1016
L KPN++YF FGC CY+ RL KF+++ + LLGY+ SK +R YN + +
Sbjct: 431 LIVGRKPNVAYFRVFGCKCYIYRKGVRLTKFESRCDEGFLLGYASNSKAYRVYNKNKGIV 490
Query: 1017 EESIHVRFDDKLDSDQS-KLVEKFADLSI-----NVSDKGKAPEEVEPE-----EDEPEE 1065
EE+ V+FD+ S + + ++ D + N+S P EVE + +DEP
Sbjct: 491 EETADVQFDETNGSQEGHENLDDVGDEGLMRAMKNMSIGDVKPIEVEDKPSTSTQDEPST 550
Query: 1066 EAGPSNSQT-LKKSR-------------ITAAHPKELILGNKDEPVRTRSAFRPSEETLL 1111
A PS +Q ++K + ++ HP + +LG+ + V+TRS ++
Sbjct: 551 SASPSQAQVEVEKEKAQDPPMPPRIYTALSKDHPIDQVLGDISKGVQTRSPVA----SIC 606
Query: 1112 SLKGLVSLIEPKSIDEALQDKEWILAMEEELNQFSKNDVWSLVKKPESVHVIGTKWVFRN 1171
VS +EPK +DEAL D +W+ A+ EELN F++N VW+LV++P +VIGTKWVFRN
Sbjct: 607 EHYSFVSCLEPKHVDEALYDPDWMNAIHEELNNFARNKVWTLVERPRDHNVIGTKWVFRN 666
Query: 1172 KLNEKGDVVRNKARLVAQGYSQQEGIDYTETFAPVARLEAIRLLISFSVNHNIVLHQMDV 1231
K +E VVRNKARLVAQG++Q E +D+ ETF PVARLEAIR+L++F+ +I L QMDV
Sbjct: 667 KQDENRLVVRNKARLVAQGFTQVEDLDFGETFGPVARLEAIRILLAFASCFDIKLFQMDV 726
Query: 1232 KSAFLNGYISEEVYVHQPPGFEDEKKPDHVFKLKKSLYGLKQAPRAWYERLSSFLLENEF 1291
KSAFLNG I+E V+V QPPGF+D K P+HV+KL K+LYGLKQAPRAWYERL FLL +F
Sbjct: 727 KSAFLNGEIAELVFVEQPPGFDDPKYPNHVYKLSKALYGLKQAPRAWYERLRDFLLSKDF 786
Query: 1292 VRGKVDTTLFCKTYKDDILIVQIYVDDIIFGSANQSLCKEFSEMMQAEFEMSMMGELKYF 1351
GKVDTTLF K DD + QIYVDDIIFGS N+ CKEF +MM EFEMSM+ EL +F
Sbjct: 787 KIGKVDTTLFTKIIGDDFFVCQIYVDDIIFGSTNEVFCKEFGDMMSREFEMSMIEELSFF 846
Query: 1352 LGIQVDQTPEGTYIHQSKYTKELLKKFNMLESTVAKTPMHPTCILEKEDKSGKVCQKLYR 1411
LG+Q+ Q +GT++ Q+KY K+LLK+F + ++ KTPM L+ ++ V KLYR
Sbjct: 847 LGLQIKQLKDGTFVSQTKYIKDLLKRFGLEDAKPIKTPMATNWHLDLDEGGKPVDLKLYR 906
Query: 1412 GMIGSLLYLTASRPDILFSVHLCARFQSDPRETHLTAVKRILRYLKGTTNLGLMYKKTSE 1471
MIGSLLYLTASRPDI+FSV + ARFQ+ P+E HL AVKRILRYLK ++ + L Y K ++
Sbjct: 907 SMIGSLLYLTASRPDIMFSVCMYARFQAAPKECHLVAVKRILRYLKHSSTISLWYPKGAK 966
Query: 1472 YKLSGYCDADYAGDRTERKSTSGNCQFLGSNLVSWASKRQSTIALSTAEAEYISAAICST 1531
+KL GY D+DYAG + +RKSTSG+CQ LG +LVSW+SK+Q+++ALSTAEAEYISA C
Sbjct: 967 FKLVGYSDSDYAGYKVDRKSTSGSCQMLGRSLVSWSSKKQNSVALSTAEAEYISAGSCCA 1026
Query: 1532 QMLWMKHQLEDYQI--LESNIPIYCDNTAAISLSKNPILHSRAKHIEVKYHFIRDYVQKG 1589
Q+LWMK L DY I E+ P+ C+N + I ++ NP+ H R KHI++++HF+ D+V K
Sbjct: 1027 QLLWMKQILLDYGISFTETQTPLLCNNDSTIKIANNPVQHFRTKHIDIRHHFLTDHVAKC 1086
Query: 1590 VLLLKFVDTDHQWADIFTKSLAEDRFNFILKNLNM 1624
+++ + T+ Q ADIFTK L E RF + LN+
Sbjct: 1087 DIVISHIRTEDQLADIFTKPLDETRFCKLRNELNV 1121
Score = 38.1 bits (87), Expect = 2.3
Identities = 19/43 (44%), Positives = 27/43 (62%), Gaps = 2/43 (4%)
Query: 610 WYLDSGCSRHMTGEKRMFR--ELKLKPGGEVGFGGNEKGKIIG 650
W LDS C++ MTG++ MF E++ K +V FG N K +IG
Sbjct: 162 WVLDSVCTQRMTGDRAMFTTFEVEGKEQEKVTFGDNSKRNVIG 204
>UniRef100_Q9SDD5 Similar to copia-type pol polyprotein [Oryza sativa]
Length = 940
Score = 860 bits (2221), Expect = 0.0
Identities = 461/905 (50%), Positives = 601/905 (65%), Gaps = 39/905 (4%)
Query: 754 SQLSKLNLVRGLPNLKFASDALCEACQKGKFTKVPFKAKNVVSTSRPLELLHIDLFGPVK 813
SQ SK + GL N++F D +C ACQ GK KNV++T+RPLELLH+DLFGP+
Sbjct: 34 SQTSKARHILGLTNIQFEKDRVCSACQAGKQIGAHHPVKNVMTTTRPLELLHMDLFGPIA 93
Query: 814 TESIGGKRYGMVIVDDYSRWTWVKFLTRKDESHAVFSTFIAQVQNEKACRIVRVRSDLGG 873
SIGG +YG+VIVDD+S +TWV FL K E+ A+F F + QNE +I +RSD
Sbjct: 94 YLSIGGNKYGLVIVDDFSCFTWVFFLHDKSETQAIFKKFARRAQNEFDLKIKNIRSDNVK 153
Query: 874 EFENDKFESLFDSYGIAHDFSCPRTPQQNGVVERKNRTLQEMARTMLQETGMAKHFWAET 933
EF+N ES D GI H+FS P +PQQNGV ERKNRTL E+ARTML E + FWAE
Sbjct: 154 EFKNTCIESFLDEEGIKHEFSAPYSPQQNGVAERKNRTLIEIARTMLDEYKTSDRFWAEA 213
Query: 934 VNTACYIQNRISVRPILNKTPYELWKNIKPNISYFHPFGCVCYVLNTKDRLHKFDAKSSK 993
VNT C+ NR+ + +L KTPYEL KPN+SYF FG CY+LN K R KF K
Sbjct: 214 VNTVCHDINRLYLHRLLKKTPYELLTGNKPNVSYFRVFGSKCYILNKKARSSKFAPKVDG 273
Query: 994 CLLLGYSDRSKGFRFYNTDAKTIEESIHVRFDD-------KLDS-------DQSKLVEKF 1039
LLGY +R +N + +E + V FD+ ++DS D + +++
Sbjct: 274 GFLLGYGSNECAYRVFNKTSGIVEIAPDVTFDETNGSQVEQVDSHVLGEEEDPREAIKRL 333
Query: 1040 A--DLSINVSDKGKAPE-EVEPEE------------DEPEE-EAGPSNSQTLKKSRITAA 1083
A D+ +G + +VEP D+ EE E P +S L RI +
Sbjct: 334 ALGDVRPREPQQGASSSTQVEPPTSTQANDPSTSSLDQGEEGEQVPPSSINLAHPRIHQS 393
Query: 1084 ----HPKELILGNKDEPVRTRSAFRPSEETLLSLKGLVSLIEPKSIDEALQDKEWILAME 1139
HP + ILG+ ++ V TRS E VS +EP ++EAL D +W++AM+
Sbjct: 394 IQRDHPTDNILGDINKGVSTRSHIANFCEHY----SFVSSLEPLRVEEALNDPDWVMAMQ 449
Query: 1140 EELNQFSKNDVWSLVKKPESVHVIGTKWVFRNKLNEKGDVVRNKARLVAQGYSQQEGIDY 1199
EELN F++N+VW+LV++ +VIGTKW+FRNK +E G V+RNKARLVAQG++Q EG+D+
Sbjct: 450 EELNNFTRNEVWTLVERSRQ-NVIGTKWIFRNKQDEAGVVIRNKARLVAQGFTQIEGLDF 508
Query: 1200 TETFAPVARLEAIRLLISFSVNHNIVLHQMDVKSAFLNGYISEEVYVHQPPGFEDEKKPD 1259
ETFAPVARLE+IR+L++F+ N N L+QMDVKSAFLNG I+E VYV QPPGF+D K P+
Sbjct: 509 GETFAPVARLESIRILLTFATNLNFKLYQMDVKSAFLNGLINELVYVEQPPGFKDPKYPN 568
Query: 1260 HVFKLKKSLYGLKQAPRAWYERLSSFLLENEFVRGKVDTTLFCKTYKDDILIVQIYVDDI 1319
HV+KL K+LY LKQAPRAWYE L +FL++N F GK D+TLF K + +DI + QIYVDDI
Sbjct: 569 HVYKLHKALYELKQAPRAWYECLRNFLVKNGFEIGKADSTLFTKRHDNDIFVCQIYVDDI 628
Query: 1320 IFGSANQSLCKEFSEMMQAEFEMSMMGELKYFLGIQVDQTPEGTYIHQSKYTKELLKKFN 1379
IFGS N+S +EFS MM FEMSMMGELK+FLG+Q+ Q EGT+I Q+KY K++LKKF
Sbjct: 629 IFGSTNKSFSEEFSRMMTKRFEMSMMGELKFFLGLQIKQLKEGTFICQTKYLKDMLKKFG 688
Query: 1380 MLESTVAKTPMHPTCILEKEDKSGKVCQKLYRGMIGSLLYLTASRPDILFSVHLCARFQS 1439
M + TPM L+ ++ V QK+YR +IGSLLYL ASRPDI+ SV +CARFQ+
Sbjct: 689 MENAKPIHTPMPSNGHLDLNEQGKDVDQKVYRSIIGSLLYLCASRPDIMLSVCMCARFQA 748
Query: 1440 DPRETHLTAVKRILRYLKGTTNLGLMYKKTSEYKLSGYCDADYAGDRTERKSTSGNCQFL 1499
P+E HL AVKRILRYL T NLGL Y K + + L GY DADYAG + +RKSTSG CQFL
Sbjct: 749 APKECHLVAVKRILRYLVHTPNLGLWYPKGARFDLIGYADADYAGCKVDRKSTSGTCQFL 808
Query: 1500 GSNLVSWASKRQSTIALSTAEAEYISAAICSTQMLWMKHQLEDYQILESNIPIYCDNTAA 1559
G +LVSW+SK+Q+++ALSTAEAEYIS C Q++WMK L DY + S IP+ CDN +A
Sbjct: 809 GRSLVSWSSKKQNSVALSTAEAEYISTGSCCAQLIWMKQTLRDYGLNVSKIPLLCDNESA 868
Query: 1560 ISLSKNPILHSRAKHIEVKYHFIRDYVQKGVLLLKFVDTDHQWADIFTKSLAEDRFNFIL 1619
I ++ NP+ HSR KHI++++HF+RD+ +G + ++ V D Q ADIFTK L E RF +
Sbjct: 869 IKIANNPVQHSRTKHIDIRHHFLRDHSTRGDIDIQHVRIDKQLADIFTKPLDEARFCELR 928
Query: 1620 KNLNM 1624
LN+
Sbjct: 929 SELNI 933
>UniRef100_Q75LG0 Putative integrase [Oryza sativa]
Length = 1507
Score = 825 bits (2130), Expect = 0.0
Identities = 451/1061 (42%), Positives = 638/1061 (59%), Gaps = 92/1061 (8%)
Query: 591 RTRLSMLQISLIAPLKHQSWYLDSGCSRHMTGEKRMFRELKLKPGGEVGFGGNEKGKIIG 650
++++ +L+++L+A K W +DSGC RHMTG ++ + +LK
Sbjct: 505 QSKVLVLEVALVAR-KENVWIVDSGCLRHMTGMVKVNEKFELK----------------- 546
Query: 651 TGTICVDSSPCIDNVLLVDGLTHNLLSISQLADKGYDVIFNQKSCRAVSQIDGSVLFNSK 710
NV LV+ L +NLLS+ Q+ + ++V F + + SVL N
Sbjct: 547 -------------NVALVEDLNYNLLSVLQIVYENFEVHFKKTRSKVFDSCGDSVL-NIS 592
Query: 711 RKNNIYKIRLSELEAQNVKCLLS-VNEEQWVWHRRLGHASMRKISQLSKLNLVRGLPNLK 769
R ++K + + CL++ +++ WHRRLGH +++LS L+LVRGLP LK
Sbjct: 593 RYGRVFKADFENPVSPVITCLVAKFDKDVMFWHRRLGHVGFDHLTRLSGLDLVRGLPKLK 652
Query: 770 FASDALCEACQKGKFTKVPFKAKNVVSTSRPLELLHIDLFGPVKTESIGGKRYGMVIVDD 829
D C C K V T P +LLH+D GP + +S+GGK Y +VI+DD
Sbjct: 653 KDLDLDCAPCHHAKMVASSHAPIVSVMTDAPRQLLHMDTVGPARVQSVGGKWYVLVIIDD 712
Query: 830 YSRWTWVKFLTRKDESHAVFSTFIAQVQNEKACRIVRVRSDLGGEFENDKFESLFDSYGI 889
+SR++WV F+ KDE+ F +++ E + R+RSD G E++N FE + G+
Sbjct: 713 FSRYSWVFFMATKDEAFQHFRGLFLRLELEFPGSLKRIRSDNGSEYKNASFEQFCNERGL 772
Query: 890 AHDFSCPRTPQQNGVVERKNRTLQEMARTMLQETGMAKHFWAETVNTACYIQNRISVRPI 949
H+FS PR PQQNGVVERKN L EM RTML E + FWAE +NTA YI NR+ +R
Sbjct: 773 EHEFSSPRVPQQNGVVERKNHVLVEMVRTMLDEYKTPRKFWAEAINTAYYISNRVFLRSK 832
Query: 950 LNKTPYELWKNIKPNISYFHPFGCVCYVLNTKDRLHKFDAKSSKCLLLGYSDRSKGFRFY 1009
L K+ YEL +P +S+ FGC C+VL + + L KF+A+S+ L LGY ++G+R
Sbjct: 833 LGKSSYELRFGHQPKVSHLRVFGCKCFVLKSGN-LDKFEARSTDGLFLGYPAHTRGYRTL 891
Query: 1010 NT----DAKTIEESIHVRFDDKLDS----------------------DQSKLVEKFADLS 1043
+ D + E+ DD++ S ++S A+ S
Sbjct: 892 SQVQGEDGRIFEDESDDNDDDEVGSAGQTGRQAGQTAGTPPVRPAHEERSDRPGLSAEGS 951
Query: 1044 INVSDKGKAPEEVEPEEDEPEEEAGPSNSQTLKKSRITAAHPKELILGNKDEPVRTRSAF 1103
++ G P E+ E S S+ + I HP E I+GN E TRS
Sbjct: 952 VDAVRDG--PLEITTSTSTDTEHG--STSEVVAPLHIQQRHPPEQIIGNIGERT-TRS-- 1004
Query: 1104 RPSEETLLSLKGLVSLIEPKSIDEALQDKEWILAMEEELNQFSKNDVWSLVKKPESVHVI 1163
+ + + + V+ EPK + AL D+ WI AM EEL F +N VW+LV+ P ++I
Sbjct: 1005 KVTTHDVCANYAFVASFEPKDVSHALTDESWINAMHEELENFERNKVWTLVEPPSGHNII 1064
Query: 1164 GTKWVFRNKLNEKGDVVRNKARLVAQGYSQQEGIDYTETFAPVARLEAIRLLISFSVNHN 1223
GTKWVF+NK NE +VRNKARLVAQG++Q EG+D+ ETFAPVAR+EAIRL ++F+ +
Sbjct: 1065 GTKWVFKNKQNEDDLIVRNKARLVAQGFTQVEGLDFDETFAPVARIEAIRLFLAFASSKG 1124
Query: 1224 IVLHQMDVKSAFLNGYISEEVYVHQPPGFEDEKKPDHVFKLKKSLYGLKQAPRAWYERLS 1283
L+QMDVKSAFLNG+I EEVYV QPPGFE+ P++VFKL K+LYGLKQAPRAWY+RL
Sbjct: 1125 FKLYQMDVKSAFLNGFIQEEVYVKQPPGFENPDFPNYVFKLSKALYGLKQAPRAWYDRLK 1184
Query: 1284 SFLLENEFVRGKVDTTLFCKTYKDDILIVQIYVDDIIFGSANQSLCKEFSEMMQAEFEMS 1343
+FLL F GKVD TLF L +F+E M+ EFEMS
Sbjct: 1185 NFLLAKGFTMGKVDKTLFV-------------------------LKHDFAETMRREFEMS 1219
Query: 1344 MMGELKYFLGIQVDQTPEGTYIHQSKYTKELLKKFNMLESTVAKTPMHPTCILEKEDKSG 1403
MMGEL YFLG+Q+ QTP+GT++HQ+KYTK+LL++F M TP+ T +L+ ++
Sbjct: 1220 MMGELSYFLGLQIKQTPQGTFVHQTKYTKDLLRRFKMENCKPISTPIDSTAVLDPDEDGE 1279
Query: 1404 KVCQKLYRGMIGSLLYLTASRPDILFSVHLCARFQSDPRETHLTAVKRILRYLKGTTNLG 1463
V QK YR MIGSLLYLTASRP+I F+V LCARFQ+ PR +H AVKRI+RYL T G
Sbjct: 1280 AVDQKEYRSMIGSLLYLTASRPEIQFAVCLCARFQASPRASHRQAVKRIMRYLNHTLEFG 1339
Query: 1464 LMYKKTSEYKLSGYCDADYAGDRTERKSTSGNCQFLGSNLVSWASKRQSTIALSTAEAEY 1523
+ Y +S LSGY DA++ G R +RKSTSG C FLG++L++W+S++QS++A STAE+EY
Sbjct: 1340 IWYSTSSSLCLSGYSDANFGGCRIDRKSTSGTCHFLGTSLIAWSSRKQSSVAQSTAESEY 1399
Query: 1524 ISAAICSTQMLWMKHQLEDYQILESNIPIYCDNTAAISLSKNPILHSRAKHIEVKYHFIR 1583
++AA C +Q+LW+ L++Y + +P++CDNT+AI+++KN + HSR KH+++++HF+R
Sbjct: 1400 VAAASCCSQILWLLSTLKNYGLTFEKVPLFCDNTSAINIAKNLVQHSRTKHVDIRFHFLR 1459
Query: 1584 DYVQKGVLLLKFVDTDHQWADIFTKSLAEDRFNFILKNLNM 1624
D+V+KG + L+F+DT Q ADIFTK L + F F+ L +
Sbjct: 1460 DHVEKGDVELQFLDTKLQIADIFTKPLDSNCFTFLCGELGI 1500
>UniRef100_Q7XBD2 Retrotransposon Opie-2 [Zea mays]
Length = 1512
Score = 777 bits (2007), Expect = 0.0
Identities = 410/831 (49%), Positives = 540/831 (64%), Gaps = 56/831 (6%)
Query: 794 VVSTSRPLELLHIDLFGPVKTESIGGKRYGMVIVDDYSRWTWVKFLTRKDESHAVFSTFI 853
+V PLELLH+DLFGPV SIGG +YG+VIVDD+SR+TWV FL K E+ F+
Sbjct: 731 IVFLQIPLELLHMDLFGPVAYLSIGGSKYGLVIVDDFSRFTWVFFLQDKSETQGTLKRFL 790
Query: 854 AQVQNEKACRIVRVRSDLGGEFENDKFESLFDSYGIAHDFSCPRTPQQNGVVERKNRTLQ 913
+ QNE ++ ++RSD G EF+N + E + GI H+FS P TPQQNGVVERKNRTL
Sbjct: 791 RRAQNEFELKVKKIRSDNGSEFKNLQVEEFLEEEGIKHEFSAPYTPQQNGVVERKNRTLI 850
Query: 914 EMARTMLQETGMAKHFWAETVNTACYIQNRISVRPILNKTPYELWKNIKPNISYFHPFGC 973
+MARTML E + FW+E VNTAC+ NR+ + +L KT YEL KPN+SYF FG
Sbjct: 851 DMARTMLGEFKTPECFWSEAVNTACHAINRVYLHRLLKKTSYELLTGNKPNVSYFRVFGS 910
Query: 974 VCYVLNTKDRLHKFDAKSSKCLLLGYSDRSKGFRFYNTDAKTIEESIHVRFDDKLDSDQS 1033
CY+L K R KF K+ + LLGY +K +R +N + +E S V FD+ S +
Sbjct: 911 KCYILVKKGRNSKFAPKAVEGFLLGYDSNTKAYRVFNKSSGLVEVSSDVVFDETNGSPRE 970
Query: 1034 KLVEKFADLSINVSDKGKAPEEVEPEEDEPEEEAGPSNSQTLKKSRITAAHPKELILGNK 1093
++V+ D+ +EE P+ + ++ I P+E +
Sbjct: 971 QVVDC----------------------DDVDEEDVPTAA--IRTMAIGEVRPQEQ--DER 1004
Query: 1094 DEPVRTRSAFRPSEETLLSLKGLVSLIEPKSIDEALQDKEWILAMEEELNQFSKNDVWSL 1153
D+P + + P+++ +P ++EAL D +W+LAM+EELN F +N+VWSL
Sbjct: 1005 DQPSSSTTVHPPTQDDE----------QPFRVEEALLDLDWVLAMQEELNNFKRNEVWSL 1054
Query: 1154 VKKPESVHVIGTKWVFRNKLNEKGDVVRNKARLVAQGYSQQEGIDYTETFAPVARLEAIR 1213
+E G V RNKARLVA+GY+Q G+D+ ETFAPVARLE+IR
Sbjct: 1055 --------------------DEHGVVTRNKARLVAKGYAQVAGLDFEETFAPVARLESIR 1094
Query: 1214 LLISFSVNHNIVLHQMDVKSAFLNGYISEEVYVHQPPGFEDEKKPDHVFKLKKSLYGLKQ 1273
+L++++ +H+ L+QMDVKSAFLNG I EEVYV QPPGFEDE+ PDHV KL K+LYGLKQ
Sbjct: 1095 ILLAYAAHHSFRLYQMDVKSAFLNGPIKEEVYVEQPPGFEDERYPDHVCKLAKALYGLKQ 1154
Query: 1274 APRAWYERLSSFLLENEFVRGKVDTTLFCKTYKDDILIVQIYVDDIIFGSANQSLCKEFS 1333
APRAWYE L FL+ N F GK D TLF KT D+ + QI+VDDIIFGS NQ C+EFS
Sbjct: 1155 APRAWYECLRDFLIANAFKVGKADPTLFTKTCNGDLFVCQIFVDDIIFGSTNQKSCEEFS 1214
Query: 1334 EMMQAEFEMSMMGELKYFLGIQVDQTPEGTYIHQSKYTKELLKKFNMLESTVAKTPMHPT 1393
+M +FEMSMMG+L YFLG QV Q +GT+I Q KYT++LLK+F M ++ AKTPM
Sbjct: 1215 RVMTQKFEMSMMGKLNYFLGFQVKQLKDGTFISQMKYTQDLLKRFGMKDAKPAKTPMGTD 1274
Query: 1394 CILEKEDKSGKVCQKLYRGMIGSLLYLTASRPDILFSVHLCARFQSDPRETHLTAVKRIL 1453
+ V QK YR MIGSLLYL ASRPDI+ SV +CARFQS+P+E HL AVKRIL
Sbjct: 1275 GHTDLNKGGKSVDQKAYRSMIGSLLYLCASRPDIMLSVCMCARFQSEPKECHLVAVKRIL 1334
Query: 1454 RYLKGTTNLGLMYKKTSEYKLSGYCDADYAGDRTERKSTSGNCQFLGSNLVSWASKRQST 1513
RYL T GL Y K S + L GY D+D AG + +RKSTSG CQFLG +LVSW SK+Q++
Sbjct: 1335 RYLVATPCFGLWYPKGSTFDLVGYSDSDNAGCKVDRKSTSGTCQFLGRSLVSWNSKKQTS 1394
Query: 1514 IALSTAEAEYISAAICSTQMLWMKHQLEDYQILESNIPIYCDNTAAISLSKNPILHSRAK 1573
+ALSTAEAEY++A C Q+LWM+ L D+ S +P+ CDN +AI +++NP+ H+R K
Sbjct: 1395 VALSTAEAEYVAAGQCCAQLLWMRQTLRDFGYNLSKVPLLCDNESAIRMAENPVEHNRTK 1454
Query: 1574 HIEVKYHFIRDYVQKGVLLLKFVDTDHQWADIFTKSLAEDRFNFILKNLNM 1624
HI++++HF+RD+ QKG + + V T++Q ADIFTK L E F + LN+
Sbjct: 1455 HIDIRHHFLRDHQQKGDIEVFHVSTENQLADIFTKPLDEKTFCRLRSELNV 1505
Score = 45.4 bits (106), Expect = 0.014
Identities = 23/49 (46%), Positives = 31/49 (62%), Gaps = 2/49 (4%)
Query: 609 SWYLDSGCSRHMTGEKRMFRE-LKLKPGGE-VGFGGNEKGKIIGTGTIC 655
SW +DSGC+ HMTGEK+MF +K K + + FG +GK+ TIC
Sbjct: 671 SWIIDSGCTNHMTGEKKMFTSYVKNKDSQDSIIFGDGNQGKLSLLATIC 719
>UniRef100_Q8W0X4 Putative pol protein [Zea mays]
Length = 1657
Score = 772 bits (1994), Expect = 0.0
Identities = 411/823 (49%), Positives = 531/823 (63%), Gaps = 46/823 (5%)
Query: 842 KDESHAVFSTFIAQVQNEKACRIVRVRSDLGGEFENDKFESLFDSYGIAHDFSCPRTPQQ 901
K E+ F+ + QNE ++ ++RSD G EF+N + E + GI H+FS P TPQQ
Sbjct: 834 KSETQGTLKRFLRRAQNEFELKVKKIRSDNGSEFKNLQVEEFLEEEGIKHEFSAPYTPQQ 893
Query: 902 NGVVERKNRTLQEMARTMLQETGMAKHFWAETVNTACYIQNRISVRPILNKTPYELWKNI 961
NGVVERKNRTL +MARTML E + FW E VNTAC+ NR+ + +L KT YEL +
Sbjct: 894 NGVVERKNRTLIDMARTMLGEFKTPECFWTEAVNTACHAINRVYLHRLLKKTSYELLTDN 953
Query: 962 KPNISYFHPFGCVCYVLNTKDRLHKFDAKSSKCLLLGYSDRSKGFRFYNTDAKTIEESIH 1021
KPN+SYF FG CY+L K R KF K+ + LLGY +K +R +N + +E S
Sbjct: 954 KPNVSYFRVFGSKCYILVKKGRNSKFAPKAVEGFLLGYDSNTKAYRVFNKSSGLVEVSSD 1013
Query: 1022 VRFDDKLDSDQSKLV-------EKFADLSINVSDKGKAPEEVEPEEDEP----------- 1063
V FD+ S + ++V E +I G+ + + E D+P
Sbjct: 1014 VVFDETNGSPREQVVDCDDVDEEDVPTAAIRTMAIGEVRPQEQDERDQPSSSTMVHPPTE 1073
Query: 1064 ----------------------EEEAGPSNSQTLKKSRITAAHPKELILGNKDEPVRTRS 1101
EEEA P+ T ++ I HP + ILG+ + V TRS
Sbjct: 1074 DDEQVPQVEALDQGGAQDDQVMEEEAQPA-PPTQVRAMIQRDHPVDQILGDISKGVTTRS 1132
Query: 1102 AFRPSEETLLSLKGLVSLIEPKSIDEALQDKEWILAMEEELNQFSKNDVWSLVKKPESVH 1161
VS IEP ++EAL D +W+LAM+EELN F +N+VW+LV +P+ +
Sbjct: 1133 RL----VNFCEHYSFVSSIEPFRVEEALLDPDWVLAMQEELNNFKRNEVWTLVPRPKQ-N 1187
Query: 1162 VIGTKWVFRNKLNEKGDVVRNKARLVAQGYSQQEGIDYTETFAPVARLEAIRLLISFSVN 1221
V+GTKWVFRNK +E G V RNKARLVA+GY+Q G+D+ ETFAPVARLE+IR+L++++ +
Sbjct: 1188 VVGTKWVFRNKQDEHGVVTRNKARLVAKGYAQVAGLDFEETFAPVARLESIRILLAYAAH 1247
Query: 1222 HNIVLHQMDVKSAFLNGYISEEVYVHQPPGFEDEKKPDHVFKLKKSLYGLKQAPRAWYER 1281
H+ L+QMDVKSAFLNG I EEVYV QPPGFEDE+ PDHV KL K+LYGLKQAPRAWYE
Sbjct: 1248 HSFRLYQMDVKSAFLNGPIKEEVYVEQPPGFEDERYPDHVCKLSKALYGLKQAPRAWYEC 1307
Query: 1282 LSSFLLENEFVRGKVDTTLFCKTYKDDILIVQIYVDDIIFGSANQSLCKEFSEMMQAEFE 1341
L FLL N F GK D TLF KT D+ + QIYVDDIIFGS NQ C+EFS +M +FE
Sbjct: 1308 LRDFLLANAFKVGKADPTLFTKTCDGDLFVCQIYVDDIIFGSTNQKSCEEFSRVMTQKFE 1367
Query: 1342 MSMMGELKYFLGIQVDQTPEGTYIHQSKYTKELLKKFNMLESTVAKTPMHPTCILEKEDK 1401
MSMMGEL YFLG QV Q +GT+I Q+KYT++LLK+F M ++ AKTPM +
Sbjct: 1368 MSMMGELNYFLGFQVKQLKDGTFISQTKYTQDLLKRFGMKDAKPAKTPMGTDGHTDLNKG 1427
Query: 1402 SGKVCQKLYRGMIGSLLYLTASRPDILFSVHLCARFQSDPRETHLTAVKRILRYLKGTTN 1461
V QK YR MIGSLLYL ASRPDI+ SV +CARFQSDP+E HL AVKRILRYL T
Sbjct: 1428 GKSVDQKAYRSMIGSLLYLCASRPDIMLSVCMCARFQSDPKECHLVAVKRILRYLVATPC 1487
Query: 1462 LGLMYKKTSEYKLSGYCDADYAGDRTERKSTSGNCQFLGSNLVSWASKRQSTIALSTAEA 1521
GL Y K S + L GY D+DYAG + +RKSTSG CQFLG +LVSW SK+Q+++ALSTAEA
Sbjct: 1488 FGLWYPKGSTFDLVGYSDSDYAGCKVDRKSTSGTCQFLGRSLVSWNSKKQTSVALSTAEA 1547
Query: 1522 EYISAAICSTQMLWMKHQLEDYQILESNIPIYCDNTAAISLSKNPILHSRAKHIEVKYHF 1581
EY++A C Q+LWM+ L D+ S +P+ CDN + I +++NP+ HSR KHI++++HF
Sbjct: 1548 EYVAAGQCCAQLLWMRQTLRDFGYNLSKVPLLCDNESDIRMAENPVEHSRTKHIDIRHHF 1607
Query: 1582 IRDYVQKGVLLLKFVDTDHQWADIFTKSLAEDRFNFILKNLNM 1624
+RD+ QKG + + V T++Q ADIFTK L E F + LN+
Sbjct: 1608 LRDHQQKGDIEVFHVSTENQLADIFTKPLDEKTFCRLRSELNV 1650
Score = 40.4 bits (93), Expect = 0.46
Identities = 20/41 (48%), Positives = 27/41 (65%), Gaps = 2/41 (4%)
Query: 609 SWYLDSGCSRHMTGEKRMFRE-LKLKPGGE-VGFGGNEKGK 647
SW +DSGC+ HMTGEK+MF +K K + + FG +GK
Sbjct: 792 SWIIDSGCTNHMTGEKKMFTSYVKNKDSQDSIIFGDGNQGK 832
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.332 0.142 0.433
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,440,507,465
Number of Sequences: 2790947
Number of extensions: 97681516
Number of successful extensions: 331504
Number of sequences better than 10.0: 3151
Number of HSP's better than 10.0 without gapping: 1953
Number of HSP's successfully gapped in prelim test: 1201
Number of HSP's that attempted gapping in prelim test: 322998
Number of HSP's gapped (non-prelim): 5535
length of query: 1626
length of database: 848,049,833
effective HSP length: 141
effective length of query: 1485
effective length of database: 454,526,306
effective search space: 674971564410
effective search space used: 674971564410
T: 11
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (22.0 bits)
S2: 82 (36.2 bits)
Lotus: description of TM0243.12


