

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0239.11
(307 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
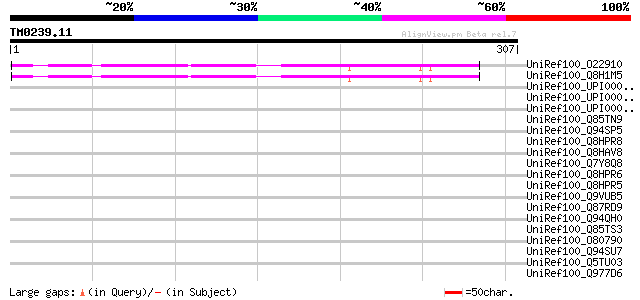
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_O22910 Hypothetical protein At2g47360 [Arabidopsis tha... 213 5e-54
UniRef100_Q8H1M5 Hypothetical protein At2g47360/T8I13.20 [Arabid... 211 3e-53
UniRef100_UPI0000368B65 UPI0000368B65 UniRef100 entry 42 0.027
UniRef100_UPI00003AF2F5 UPI00003AF2F5 UniRef100 entry 39 0.23
UniRef100_UPI00003AF2F4 UPI00003AF2F4 UniRef100 entry 39 0.23
UniRef100_Q85TN9 NADH subunit 2 [Jenynsia maculata] 38 0.30
UniRef100_Q94SP5 NADH-ubiquinone oxidoreductase chain 2 [Beryx s... 36 1.5
UniRef100_Q8HPR8 NADH dehydrogenase subunit 2 [Etheostoma obeyense] 36 1.5
UniRef100_Q8HAV8 NADH dehydrogenase subunit 2 [Etheostoma virgatum] 35 1.9
UniRef100_Q7Y8Q8 NADH dehydrogenase subunit 2 [Ophiodon elongatus] 35 1.9
UniRef100_Q8HPR6 NADH dehydrogenase subunit 2 [Etheostoma virgatum] 35 2.5
UniRef100_Q8HPR5 NADH dehydrogenase subunit 2 [Etheostoma virgatum] 35 2.5
UniRef100_Q9VUB5 CG9007-PA [Drosophila melanogaster] 35 2.5
UniRef100_Q87RD9 Ferrous iron transport protein B [Vibrio paraha... 35 3.3
UniRef100_Q94QH0 NADH-ubiquinone oxidoreductase chain 2 [Arctosc... 35 3.3
UniRef100_Q85TS3 NADH subunit 2 [Aphanius iberus] 35 3.3
UniRef100_O80790 Putative non-LTR retroelement reverse transcrip... 34 4.3
UniRef100_Q94SU7 NADH-ubiquinone oxidoreductase chain 2 [Poromit... 34 4.3
UniRef100_Q5TU03 ENSANGP00000026920 [Anopheles gambiae str. PEST] 34 4.3
UniRef100_Q977D6 Hypothetical protein ST0010 [Sulfolobus tokodaii] 34 4.3
>UniRef100_O22910 Hypothetical protein At2g47360 [Arabidopsis thaliana]
Length = 303
Score = 213 bits (542), Expect = 5e-54
Identities = 124/298 (41%), Positives = 175/298 (58%), Gaps = 45/298 (15%)
Query: 2 GRWQQKAKEFFSNHGGAPTPKATFHLLTITLLSLLLPLSFFLLASLSAAKHYLQTFTLFL 61
G+W+++A++ F L+T TLLSLLLPLSF LL+ LS+A +F L
Sbjct: 6 GKWRKEAEQLVVK---------PFRLVTTTLLSLLLPLSFLLLSRLSSA-----SFLFSL 51
Query: 62 HSPQQQQQPFSFLFTLALQINPCILYVLVSIVSIATLINGLMGKITLLNDSSTSPVLQPS 121
Q Q + F+F+L L+ NP I+Y +VS +S+ TL+ GL KIT D S P
Sbjct: 52 TKSQPQTESSFFVFSLFLRANPAIVYAVVSSISVYTLVLGLTTKITA-TDPKHSIAFYPH 110
Query: 122 LYIAWILLCAFQFCVGLGIEASIAAGIFEFDNSSSSSSSSFGESAAERSLLSKVFFLLGL 181
+ IAW+ L Q VG+G+E +I+ G+ +ER+ LS++ F GL
Sbjct: 111 VSIAWLTLFLVQISVGIGLETTISNGLI---------------IGSERNFLSRLVFFFGL 155
Query: 182 HETTQVWYRVVVRPVVDDTVFGG----AKKEKWIERVAVAASLGVLWWWRLREEVESLVV 237
HE +WYRV+VRPVVD+T+ GG ++E +ERVA+A S G LWWW+LR+EVE+LV
Sbjct: 156 HEVMLLWYRVIVRPVVDNTLLGGEDGQRREETVVERVALAVSCGTLWWWKLRDEVEALVG 215
Query: 238 MAEAKKSEQF-----GELELK------DFIGWWLYYVTVTIGMVRIVKGLMWMFMVAL 284
+AEAK++ G + + DF+ WWLYY+ VTIGMVRI+KG +W M+ L
Sbjct: 216 VAEAKRALLLLLPIDGNVNVSFDVGTVDFVNWWLYYMVVTIGMVRIIKGSLWFGMILL 273
>UniRef100_Q8H1M5 Hypothetical protein At2g47360/T8I13.20 [Arabidopsis thaliana]
Length = 303
Score = 211 bits (536), Expect = 3e-53
Identities = 123/298 (41%), Positives = 173/298 (57%), Gaps = 45/298 (15%)
Query: 2 GRWQQKAKEFFSNHGGAPTPKATFHLLTITLLSLLLPLSFFLLASLSAAKHYLQTFTLFL 61
G+W+++A++ F L+T TLLSLLLP SF LL+ LS+A +F L
Sbjct: 6 GKWRKEAEQLVVK---------PFRLVTTTLLSLLLPXSFLLLSRLSSA-----SFLFSL 51
Query: 62 HSPQQQQQPFSFLFTLALQINPCILYVLVSIVSIATLINGLMGKITLLNDSSTSPVLQPS 121
Q Q + F+F L L+ NP I+Y +VS +S+ TL+ GL KIT D S P
Sbjct: 52 TKSQPQTESSFFVFXLFLRANPAIVYAVVSSISVYTLVLGLTTKITA-TDPKHSIAFYPH 110
Query: 122 LYIAWILLCAFQFCVGLGIEASIAAGIFEFDNSSSSSSSSFGESAAERSLLSKVFFLLGL 181
+ IAW+ L Q VG+G+E +I+ G+ +ER+ LS++ F GL
Sbjct: 111 VSIAWLTLFLVQISVGIGLETTISNGLI---------------IGSERNFLSRLVFFFGL 155
Query: 182 HETTQVWYRVVVRPVVDDTVFGG----AKKEKWIERVAVAASLGVLWWWRLREEVESLVV 237
HE +WYRV+VRPVVD+T+ GG ++E +ERVA+A S G LWWW+LR+EVE+LV
Sbjct: 156 HEVMLLWYRVIVRPVVDNTLLGGEDGQRREETVVERVALAVSCGTLWWWKLRDEVEALVG 215
Query: 238 MAEAKKSEQF-----GELELK------DFIGWWLYYVTVTIGMVRIVKGLMWMFMVAL 284
+AEAK++ G + + DF+ WWLYY+ VTIGMVRI+KG +W M+ L
Sbjct: 216 VAEAKRALLLLLPIDGNVNVSFDVGTVDFVNWWLYYMVVTIGMVRIIKGSLWFGMILL 273
>UniRef100_UPI0000368B65 UPI0000368B65 UniRef100 entry
Length = 1123
Score = 41.6 bits (96), Expect = 0.027
Identities = 42/158 (26%), Positives = 71/158 (44%), Gaps = 34/158 (21%)
Query: 27 LLTITLLSLLLPLSFFLLASLSAAKHYLQTFTLFLHSPQQQQQPFSFLFTLALQINPCIL 86
L T + + + L F LLA L L+T +H+ +Q++ P ++ALQ+NPC+
Sbjct: 121 LWTTDMAACVKELVFHLLAEL------LRT----VHTLEQRRHPAGLSSSIALQLNPCL- 169
Query: 87 YVLVSIVSIATLINGLMGKITLLNDSSTS----------PVLQPSLYIAWILLCAFQFCV 136
++ L ++ L D T +++ L +A + L
Sbjct: 170 ----------AMLMALQSELHKLYDEETQNWGRFSTYFHALMEGCLAVAEVTLPTNMSVT 219
Query: 137 GLGIEASIAAGIFEFDNSSSSSSSSFGESAAERSLLSK 174
G+ ++ A + +SSSSSSSS G++ SLLSK
Sbjct: 220 ASGVTSATAPNL---SDSSSSSSSSPGQTPQSPSLLSK 254
>UniRef100_UPI00003AF2F5 UPI00003AF2F5 UniRef100 entry
Length = 1247
Score = 38.5 bits (88), Expect = 0.23
Identities = 37/135 (27%), Positives = 60/135 (44%), Gaps = 12/135 (8%)
Query: 52 HYLQTFTLFLHSPQQQQQPFSFLFTLALQINPCILYVLVSIVSIATLI-----NGLMGKI 106
H L +H+ +Q++ P ++ALQ+NPC+ ++ + L N + G
Sbjct: 536 HLLAELLRKIHNLEQKKSPAGLSSSIALQLNPCLAMLMALQSELHKLYDEETQNWVSGNA 595
Query: 107 TLLNDSSTSPVLQPSLYI-AWILLCAFQFCVGLGIEASIAAGIF------EFDNSSSSSS 159
+ + + S Y A + +C V L I S+ A + +SSSSSS
Sbjct: 596 CGGSGVGAADQGRFSTYFHALMEVCLAVAEVTLPINMSVTANVISSTSAPNLSDSSSSSS 655
Query: 160 SSFGESAAERSLLSK 174
SS G++ SLLSK
Sbjct: 656 SSPGQTPQSPSLLSK 670
>UniRef100_UPI00003AF2F4 UPI00003AF2F4 UniRef100 entry
Length = 1405
Score = 38.5 bits (88), Expect = 0.23
Identities = 37/135 (27%), Positives = 60/135 (44%), Gaps = 12/135 (8%)
Query: 52 HYLQTFTLFLHSPQQQQQPFSFLFTLALQINPCILYVLVSIVSIATLI-----NGLMGKI 106
H L +H+ +Q++ P ++ALQ+NPC+ ++ + L N + G
Sbjct: 694 HLLAELLRKIHNLEQKKSPAGLSSSIALQLNPCLAMLMALQSELHKLYDEETQNWVSGNA 753
Query: 107 TLLNDSSTSPVLQPSLYI-AWILLCAFQFCVGLGIEASIAAGIF------EFDNSSSSSS 159
+ + + S Y A + +C V L I S+ A + +SSSSSS
Sbjct: 754 CGGSGVGAADQGRFSTYFHALMEVCLAVAEVTLPINMSVTANVISSTSAPNLSDSSSSSS 813
Query: 160 SSFGESAAERSLLSK 174
SS G++ SLLSK
Sbjct: 814 SSPGQTPQSPSLLSK 828
>UniRef100_Q85TN9 NADH subunit 2 [Jenynsia maculata]
Length = 348
Score = 38.1 bits (87), Expect = 0.30
Identities = 49/170 (28%), Positives = 72/170 (41%), Gaps = 24/170 (14%)
Query: 26 HLLTITLLSLLLPLSFFLLASLSAAKHYLQTFTL---FLHSPQQQQQPFSFLFTLALQIN 82
H LT TL++ L L L S LQ L + S Q+ PFS L LQI
Sbjct: 91 HPLTTTLVTFALALKIGLAPLHSWMPEVLQGLDLTTGLILSTWQKLAPFS----LILQIQ 146
Query: 83 PCILYVLVSIVSIATLINGLMGKITLLNDSSTSPVLQPS--LYIAWILLCAFQFCVGLGI 140
P +L+ + +TLI G G LN + +L S ++ W++L QF L +
Sbjct: 147 PTNHNILIILGLTSTLIGGWGG----LNQTQLRKILAYSSIAHLGWMVLIV-QFSPALTL 201
Query: 141 EASIAAGI----------FEFDNSSSSSSSSFGESAAERSLLSKVFFLLG 180
A + + F + SS S+S+ +S A S+L + LG
Sbjct: 202 LALLTYFVMTFSLFLTFKLNFSTNISSLSTSWAKSPALTSMLPLMLLSLG 251
>UniRef100_Q94SP5 NADH-ubiquinone oxidoreductase chain 2 [Beryx splendens]
Length = 348
Score = 35.8 bits (81), Expect = 1.5
Identities = 46/170 (27%), Positives = 78/170 (45%), Gaps = 24/170 (14%)
Query: 26 HLLTITLLSLLLPLSFFLLASLSAAKHYLQTFTL---FLHSPQQQQQPFSFLFTLALQIN 82
H L IT+++L L L L S LQ L + S Q+ PF+ L LQI
Sbjct: 91 HPLPITIITLALALKIGLAPVHSWLPEVLQGLDLTTGLILSTWQKLAPFALL----LQIQ 146
Query: 83 PCILYVLVSIVSIATLINGLMGKITLLNDSSTSPVLQPS--LYIAWILLCAFQFCVGLGI 140
P ++++ ++TL+ G G LN + +L S ++ W++L QF L +
Sbjct: 147 PANPTTMIALGLMSTLVGGWGG----LNQTQLRKILAYSSIAHLGWMIL-IIQFNPSLTL 201
Query: 141 EASIAAGI--------FEFDNSS--SSSSSSFGESAAERSLLSKVFFLLG 180
A + I F+ +NS+ ++ ++S+ +S A +L+ V LG
Sbjct: 202 LALLTYFIMTFSTFLVFKLNNSTNINTLATSWAKSPALTALMPLVLLSLG 251
>UniRef100_Q8HPR8 NADH dehydrogenase subunit 2 [Etheostoma obeyense]
Length = 348
Score = 35.8 bits (81), Expect = 1.5
Identities = 47/170 (27%), Positives = 77/170 (44%), Gaps = 24/170 (14%)
Query: 26 HLLTITLLSLLLPLSFFLLASLSAAKHYLQTFTL---FLHSPQQQQQPFSFLFTLALQIN 82
H L ITL++L L L L S LQ L + S Q+ PF+ L LQI
Sbjct: 91 HPLPITLITLALALKIGLAPVHSWLPEVLQGLDLSTGLILSTWQKLAPFALL----LQIQ 146
Query: 83 PCILYVLVSIVSIATLINGLMGKITLLNDSSTSPVLQPS--LYIAWILLCAFQFCVGLGI 140
P VLV++ +TLI G G LN + +L S ++ W++L QF L +
Sbjct: 147 PANSTVLVALGLASTLIGGWGG----LNQTQLRKILAYSSIAHLGWMIL-VLQFSPSLTL 201
Query: 141 EASI--------AAGIFEFDNSS--SSSSSSFGESAAERSLLSKVFFLLG 180
+ + +F+ +NS+ ++ ++S+ ++ A +L + LG
Sbjct: 202 LTLLTYFVMTFSSFLVFKLNNSTNINTLATSWAKAPALTALTPLILLSLG 251
>UniRef100_Q8HAV8 NADH dehydrogenase subunit 2 [Etheostoma virgatum]
Length = 348
Score = 35.4 bits (80), Expect = 1.9
Identities = 47/170 (27%), Positives = 76/170 (44%), Gaps = 24/170 (14%)
Query: 26 HLLTITLLSLLLPLSFFLLASLSAAKHYLQTFTL---FLHSPQQQQQPFSFLFTLALQIN 82
H L ITL++L L L L S LQ L + S Q+ PF+ L LQI
Sbjct: 91 HPLPITLITLALALKIGLAPVHSWLPEVLQGLDLTTGLILSTWQKLAPFALL----LQIQ 146
Query: 83 PCILYVLVSIVSIATLINGLMGKITLLNDSSTSPVLQPS--LYIAWILLCAFQFCVGLGI 140
P +L+++ +TLI G G LN + VL S ++ W++L QF L +
Sbjct: 147 PANSVILMALGLASTLIGGWGG----LNQTQLRKVLAYSSIAHLGWMIL-VLQFSPSLTL 201
Query: 141 EASIAAG--------IFEFDNSSSSS--SSSFGESAAERSLLSKVFFLLG 180
A + +F+ + S+S + ++S+ ++ A +L + LG
Sbjct: 202 LALLTYFVMTLSTFLVFKLNKSTSINMLATSWAKAPAITALAPLILLSLG 251
>UniRef100_Q7Y8Q8 NADH dehydrogenase subunit 2 [Ophiodon elongatus]
Length = 348
Score = 35.4 bits (80), Expect = 1.9
Identities = 48/170 (28%), Positives = 75/170 (43%), Gaps = 24/170 (14%)
Query: 26 HLLTITLLSLLLPLSFFLLASLSAAKHYLQTFTL---FLHSPQQQQQPFSFLFTLALQIN 82
H L ITL++L L L L S LQ L + S Q+ PF+ L LQI
Sbjct: 91 HPLPITLITLALALKVGLAPLHSWLPEVLQGLDLTTGLILSTWQKLAPFALL----LQIQ 146
Query: 83 PCILYVLVSIVSIATLINGLMGKITLLNDSSTSPVLQPS--LYIAWILLCAFQFCVGLGI 140
P +L+++ ++TL+ G G LN + +L S ++ W++L QF L +
Sbjct: 147 PANSTLLIALGLMSTLVGGWGG----LNQTQLRKILAYSSIAHLGWMVL-VMQFSPSLTL 201
Query: 141 EASIAAGIFEFD-------NSS---SSSSSSFGESAAERSLLSKVFFLLG 180
+ I F NSS ++ ++S+ ++ A SL V LG
Sbjct: 202 LTLLTYLIMTFSTFLVFKLNSSTNINALATSWAKAPALTSLTPLVLLSLG 251
>UniRef100_Q8HPR6 NADH dehydrogenase subunit 2 [Etheostoma virgatum]
Length = 348
Score = 35.0 bits (79), Expect = 2.5
Identities = 45/170 (26%), Positives = 76/170 (44%), Gaps = 24/170 (14%)
Query: 26 HLLTITLLSLLLPLSFFLLASLSAAKHYLQTFTL---FLHSPQQQQQPFSFLFTLALQIN 82
H L ITL++L L L L S LQ L + S Q+ PF+ L LQI
Sbjct: 91 HPLPITLITLALALKIGLAPVHSWLPEVLQGLDLTTGLILSTWQKLAPFALL----LQIQ 146
Query: 83 PCILYVLVSIVSIATLINGLMGKITLLNDSSTSPVLQPS--LYIAWILLCAFQFCVGLGI 140
P +L++ +TL+ G G LN + +L S ++ W++L QF L +
Sbjct: 147 PANSNILIAFGLTSTLVGGWGG----LNQTQLRKILAYSSIAHLGWMIL-VLQFSPSLTL 201
Query: 141 EASI--------AAGIFEFDNSSSSS--SSSFGESAAERSLLSKVFFLLG 180
+ + +F+ +NS+S + ++S+ ++ A +L + LG
Sbjct: 202 LTLLTYLVMTFSSFLVFKLNNSTSINMLATSWDKAPALTALAPMILLSLG 251
>UniRef100_Q8HPR5 NADH dehydrogenase subunit 2 [Etheostoma virgatum]
Length = 348
Score = 35.0 bits (79), Expect = 2.5
Identities = 45/170 (26%), Positives = 76/170 (44%), Gaps = 24/170 (14%)
Query: 26 HLLTITLLSLLLPLSFFLLASLSAAKHYLQTFTL---FLHSPQQQQQPFSFLFTLALQIN 82
H L ITL++L L L L S LQ L + S Q+ PF+ L LQI
Sbjct: 91 HPLPITLITLALALKIGLAPVHSWLPEVLQGLDLTTGLILSTWQKLAPFALL----LQIQ 146
Query: 83 PCILYVLVSIVSIATLINGLMGKITLLNDSSTSPVLQPS--LYIAWILLCAFQFCVGLGI 140
P +L++ +TL+ G G LN + +L S ++ W++L QF L +
Sbjct: 147 PANSNILIAFGLASTLVGGWGG----LNQTQLRKILAYSSIAHLGWMIL-VLQFSPSLTL 201
Query: 141 EASI--------AAGIFEFDNSSSSS--SSSFGESAAERSLLSKVFFLLG 180
+ + +F+ +NS+S + ++S+ ++ A +L + LG
Sbjct: 202 LTLLTYLVMTFSSFLVFKLNNSTSINMLATSWAKAPALTALAPLILLSLG 251
>UniRef100_Q9VUB5 CG9007-PA [Drosophila melanogaster]
Length = 3146
Score = 35.0 bits (79), Expect = 2.5
Identities = 25/66 (37%), Positives = 34/66 (50%), Gaps = 1/66 (1%)
Query: 131 AFQFCVGLGIEASIAAGIFEFDNSSSSSSSSFGESAAERSLLSKVFFLLGLHETTQVWYR 190
A Q +GLG+ A++AAG NS + S+SS GES ++ L S L L T V
Sbjct: 1350 AIQLTMGLGVGATVAAGASVLPNSRNRSTSSSGES-SQMGLNSPQLGQLNLGFKTSVTAT 1408
Query: 191 VVVRPV 196
+ PV
Sbjct: 1409 SLTAPV 1414
>UniRef100_Q87RD9 Ferrous iron transport protein B [Vibrio parahaemolyticus]
Length = 758
Score = 34.7 bits (78), Expect = 3.3
Identities = 35/116 (30%), Positives = 49/116 (42%), Gaps = 20/116 (17%)
Query: 85 ILYVLVSIVSIATLINGLMGKITLLNDSSTS----------PVLQPSLYIAWILLCAFQF 134
I++ L + +A + GL K TL SS S P LQ + W L +F
Sbjct: 451 IVFALYLLGIVAAIFTGLFLKHTLYPGSSDSLVMEMPDYELPTLQNVVIKTWQKLK--RF 508
Query: 135 CVGLGIEASIAAGIFEFDNSSSSSSSSFGESAAERSLLSK-------VFFLLGLHE 183
+G G + I F NS + S FG +E S+LSK VF +G+ E
Sbjct: 509 VLGAGKTIVVVVAILSFLNSLGTDGS-FGNEDSENSVLSKAAQVVTPVFAPIGIQE 563
>UniRef100_Q94QH0 NADH-ubiquinone oxidoreductase chain 2 [Arctoscopus japonicus]
Length = 348
Score = 34.7 bits (78), Expect = 3.3
Identities = 47/170 (27%), Positives = 74/170 (42%), Gaps = 24/170 (14%)
Query: 26 HLLTITLLSLLLPLSFFLLASLSAAKHYLQTFTL---FLHSPQQQQQPFSFLFTLALQIN 82
H L +TL++L L L L S LQ L + S Q+ PF+ L +QI
Sbjct: 91 HPLPVTLVTLALALKLGLAPIHSWLPEVLQGLDLTTGLILSTWQKLAPFALL----MQIQ 146
Query: 83 PCILYVLVSIVSIATLINGLMGKITLLNDSSTSPVLQPS--LYIAWILLCAFQFCVGLG- 139
P +L+ + +TL+ G G LN + +L S ++ W+LL QF L
Sbjct: 147 PANSTLLIVLGLASTLVGGWGG----LNQTQLRKILAYSSIAHLGWMLL-VLQFSPSLTF 201
Query: 140 -------IEASIAAGIFEFDNSSS--SSSSSFGESAAERSLLSKVFFLLG 180
+ S +F+ NS+S + ++S+ ++ A SL V LG
Sbjct: 202 ITLLTYILMTSSTFLVFKLTNSTSINALATSWAKAPAFTSLTPLVLLSLG 251
>UniRef100_Q85TS3 NADH subunit 2 [Aphanius iberus]
Length = 348
Score = 34.7 bits (78), Expect = 3.3
Identities = 46/170 (27%), Positives = 72/170 (42%), Gaps = 24/170 (14%)
Query: 26 HLLTITLLSLLLPLSFFLLASLSAAKHYLQTFTL---FLHSPQQQQQPFSFLFTLALQIN 82
H L TL++L L L L + LQ L + S Q+ PF L LQ+
Sbjct: 91 HPLPTTLVTLALALKIGLAPLHAWMPEVLQGLDLTTGLILSTWQKLAPFC----LILQLQ 146
Query: 83 PCILYVLVSIVSIATLINGLMGKITLLNDSSTSPVLQPS--LYIAWILLCAFQFCVGLGI 140
VL+ + ++TL+ G G LN + +L S ++ W++L QF L +
Sbjct: 147 HTNHQVLLILGLLSTLVGGWGG----LNQTQLRKILAYSSIAHLGWMIL-VLQFSPSLTL 201
Query: 141 EASIAAGIFEFD----------NSSSSSSSSFGESAAERSLLSKVFFLLG 180
+ I F + +S SSS+ ++ A S+L + FLLG
Sbjct: 202 VTLLTYFIMTFSLFLAFKLNHSTNINSLSSSWAKAPALTSILPLILFLLG 251
>UniRef100_O80790 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 970
Score = 34.3 bits (77), Expect = 4.3
Identities = 23/67 (34%), Positives = 37/67 (54%), Gaps = 6/67 (8%)
Query: 218 ASLGVLWWWRLREEVESLVVMAEAKKSEQFGELE----LKDFIGWWLYYVTVTIGMVRIV 273
A+L L WWRL + E+L +KK + GE++ LK W + +VT+G+ +V
Sbjct: 508 ATLDSLDWWRLLHDKETLWARVLSKK-YKVGEMQNSNWLKPKSSWSSTWRSVTVGLREVV 566
Query: 274 -KGLMWM 279
KG+ W+
Sbjct: 567 AKGVRWV 573
>UniRef100_Q94SU7 NADH-ubiquinone oxidoreductase chain 2 [Poromitra oscitans]
Length = 348
Score = 34.3 bits (77), Expect = 4.3
Identities = 45/170 (26%), Positives = 78/170 (45%), Gaps = 24/170 (14%)
Query: 26 HLLTITLLSLLLPLSFFLLASLSAAKHYLQTFTL---FLHSPQQQQQPFSFLFTLALQIN 82
H + IT+++L L L L S LQ L + S Q+ PF+ L +QI
Sbjct: 91 HPIPITIITLALALKIGLAPVHSWLPEVLQGLDLTTGLILSTWQKLAPFA----LIMQIQ 146
Query: 83 PCILYVLVSIVSIATLINGLMGKITLLNDSSTSPVLQPS--LYIAWILLCAFQFCVGLGI 140
P ++++ ++TL+ G G LN + +L S ++ W++L QF L +
Sbjct: 147 PYNPTTMIALGLMSTLVGGWGG----LNQTQLRKILAYSSIAHLGWMVL-IIQFSPSLTL 201
Query: 141 EASIAAGI--------FEFDNSS--SSSSSSFGESAAERSLLSKVFFLLG 180
A + I F+ +NS+ ++ ++S+ +S A SL+ V LG
Sbjct: 202 LALLTYFIMTFSTFLAFKLNNSTNINTLATSWAKSPALTSLVPLVLLSLG 251
>UniRef100_Q5TU03 ENSANGP00000026920 [Anopheles gambiae str. PEST]
Length = 288
Score = 34.3 bits (77), Expect = 4.3
Identities = 35/150 (23%), Positives = 62/150 (41%), Gaps = 28/150 (18%)
Query: 27 LLTITLLSLLLPLSFFLLASLSAAKHYLQTFTLFLHSPQQQQQPFSFLFTLALQINPCIL 86
LL LL LLL LS LL +L Y S + Q +P SF+ TL +
Sbjct: 76 LLPFALLGLLLCLS--LLLALRTGNSY--------RSDRNQSRPVSFIDTLIWMVG---- 121
Query: 87 YVLVSIVSIATLINGLMGKITLLNDSSTSPVLQPSLYIAWILLCAFQFCVG--LGIEASI 144
+ G I N SS+ ++ +LY++ I+ C++ + L ++AS+
Sbjct: 122 ------------VLAQQGSIIRPNSSSSMVIILVALYMSAIIYCSYLTKIASLLSVDASV 169
Query: 145 AAGIFEFDNSSSSSSSSFGESAAERSLLSK 174
+ ++ G + A+ +++ K
Sbjct: 170 NLDLQTVLSAGQYQIGFVGNNTADETVIQK 199
>UniRef100_Q977D6 Hypothetical protein ST0010 [Sulfolobus tokodaii]
Length = 130
Score = 34.3 bits (77), Expect = 4.3
Identities = 25/79 (31%), Positives = 39/79 (48%), Gaps = 3/79 (3%)
Query: 109 LNDSSTSPVLQPSLYIAWILLCAFQFCVGLGIEASIAAGIFEFDNSSSSSSSSFGESAAE 168
+++ + P L P L + IL C+FQ C G GI A + A + F S + S SSF S +
Sbjct: 26 VDNQTLLPFLTPELLL--ILYCSFQ-CTGYGIFALLNASMNGFIFSKNLSPSSFISSCID 82
Query: 169 RSLLSKVFFLLGLHETTQV 187
+ S F L + ++
Sbjct: 83 TGISSSPIFFSPLSKNLKL 101
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.325 0.137 0.417
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 478,992,990
Number of Sequences: 2790947
Number of extensions: 17829906
Number of successful extensions: 74072
Number of sequences better than 10.0: 44
Number of HSP's better than 10.0 without gapping: 3
Number of HSP's successfully gapped in prelim test: 41
Number of HSP's that attempted gapping in prelim test: 74052
Number of HSP's gapped (non-prelim): 47
length of query: 307
length of database: 848,049,833
effective HSP length: 127
effective length of query: 180
effective length of database: 493,599,564
effective search space: 88847921520
effective search space used: 88847921520
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 74 (33.1 bits)
Lotus: description of TM0239.11


