

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0230.1
(420 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
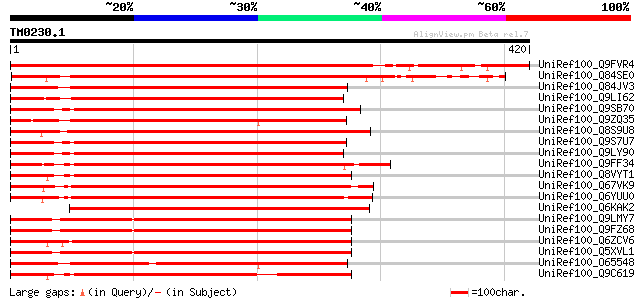
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9FVR4 Hypothetical protein F3C3.3 [Arabidopsis thaliana] 576 e-163
UniRef100_Q84SE0 Putative TPA: Cgi67 serine protease [Oryza sativa] 496 e-139
UniRef100_Q84JV3 Hypothetical protein At4g31020 [Arabidopsis tha... 348 2e-94
UniRef100_Q9LI62 Similarity to unknown protein [Arabidopsis thal... 348 2e-94
UniRef100_Q9SB70 Hypothetical protein F22K18.40 [Arabidopsis tha... 343 4e-93
UniRef100_Q9ZQ35 Hypothetical protein At2g24320 [Arabidopsis tha... 342 9e-93
UniRef100_Q8S9U8 P0519D04.38 protein [Oryza sativa] 337 3e-91
UniRef100_Q9S7U7 F28J7.1 protein [Arabidopsis thaliana] 336 8e-91
UniRef100_Q9LY90 Hypothetical protein F18O22_180 [Arabidopsis th... 333 4e-90
UniRef100_Q9FF34 Emb|CAB87778.1 [Arabidopsis thaliana] 332 9e-90
UniRef100_Q8VYT1 Hypothetical protein At1g66900 [Arabidopsis tha... 328 1e-88
UniRef100_Q67VK9 Cgi67 serine protease-like [Oryza sativa] 328 2e-88
UniRef100_Q6YUU0 Putative Cgi67 serine protease [Oryza sativa] 326 9e-88
UniRef100_Q6KAK2 Hypothetical protein OJ1004_E04.17 [Oryza sativa] 325 2e-87
UniRef100_Q9LMY7 F21F23.4 protein [Arabidopsis thaliana] 313 6e-84
UniRef100_Q9FZ68 F13B4.9 protein [Arabidopsis thaliana] 313 6e-84
UniRef100_Q6ZCV6 Hypothetical protein P0028A08.22 [Oryza sativa] 311 2e-83
UniRef100_Q5XVL1 Hypothetical protein [Arabidopsis thaliana] 310 6e-83
UniRef100_O65548 Hypothetical protein F6I18.70 [Arabidopsis thal... 305 1e-81
UniRef100_Q9C619 Hypothetical protein T4O24.3 [Arabidopsis thali... 295 2e-78
>UniRef100_Q9FVR4 Hypothetical protein F3C3.3 [Arabidopsis thaliana]
Length = 422
Score = 576 bits (1484), Expect = e-163
Identities = 286/431 (66%), Positives = 321/431 (74%), Gaps = 29/431 (6%)
Query: 1 MGCMVSQLAAKFAFFPPSPPTYQLKKRDDGKLTVVSAASPSVP-IPHADDSSLDVLLVDT 59
MGCM S LAAKFAFFPPSPPTY L K DGKL+ VS+AS S P A D SLDV +V T
Sbjct: 1 MGCMFSHLAAKFAFFPPSPPTYHLTKTPDGKLSAVSSASSSSSTFPSAGDPSLDVKVVKT 60
Query: 60 KHGNKIVAFYLKNPYARLTLLYSHGNAADLGQLYDLFVQLKVNLRVNLMGYDYSGYGAST 119
+ GNK+ AFYL+NP ARLTLLYSHGNAADLGQL+DLFVQLKVNLRVNLMGYDYSGYGAST
Sbjct: 61 RRGNKVTAFYLRNPNARLTLLYSHGNAADLGQLFDLFVQLKVNLRVNLMGYDYSGYGAST 120
Query: 120 GKPSESSTYADIEAIYECLESEYGVGQEDIILYGQSVGSGPTLHLAAKLPRLRGVVLHSG 179
GKPSE TYADIEA YECL+++YGVGQED+ILYGQSVGSGPTLHLA+KLPRLRGVVLHSG
Sbjct: 121 GKPSEYDTYADIEAAYECLQTDYGVGQEDLILYGQSVGSGPTLHLASKLPRLRGVVLHSG 180
Query: 180 ILSGLRVLCHVKFSFCFDIYKNINKIKKVKCPVLVIHGTEDDVVNWLHGNGLWKMARESY 239
ILSGLRVLCHVKF FC DIY N+NKIKKVKCPVLVIHGTEDDVVNWLHGN LWKMA+E Y
Sbjct: 181 ILSGLRVLCHVKFKFCCDIYSNVNKIKKVKCPVLVIHGTEDDVVNWLHGNRLWKMAKEPY 240
Query: 240 DPLWIKGGGHCNLELYPDYIRHLCKFIQEMESITTEKRLKKLRQSQGPQSNSSSCFSCSC 299
+PLWIKGGGHCNLE+YPDYIRHL +FIQ+ME+ TT+ RLK + Q + S+ C
Sbjct: 241 EPLWIKGGGHCNLEIYPDYIRHLYRFIQDMENTTTKSRLKTIWQEIRRRDESTGC----- 295
Query: 300 CRMKCCSNNCTGCSNCSCINPEC----CNCSCINPECFNCSCINCSC-SLK-CTQCCCLP 353
CCS C +CSC P C C+C C C +C C CSC +LK C CC P
Sbjct: 296 ----CCSGLCR--PSCSCPKPRCPKPSCSCGC---GCGDCGCFKCSCPTLKGCFSCCKKP 346
Query: 354 SCIKCCHLPKF--STCFGSSCCTKCSRPSCCFKC--CCNLRCARPSCCCMSCFCWRCCVG 409
SC+ C P F S+CFG C KCS C+KC C + C R SCCC CF W CC G
Sbjct: 347 SCVSSCCCPTFKCSSCFGKPKCPKCS----CWKCLKCPDTECCRSSCCCSGCFSWLCCCG 402
Query: 410 KHSGRDEKQSG 420
++ ++ G
Sbjct: 403 GGRRKEGERRG 413
>UniRef100_Q84SE0 Putative TPA: Cgi67 serine protease [Oryza sativa]
Length = 389
Score = 496 bits (1276), Expect = e-139
Identities = 257/418 (61%), Positives = 303/418 (72%), Gaps = 54/418 (12%)
Query: 2 GCMVSQLAAKFAFFPPSPPTYQLKKRD--DGKLTVVSAASPSVPIPHADDSSLDVLLVDT 59
GC VS LAA+FAFFPP P TY ++K + G +V++ P D+++DVLLVDT
Sbjct: 4 GCTVSSLAARFAFFPPEPATYAVRKDEACGGGGRLVASGVPR-------DAAVDVLLVDT 56
Query: 60 KHGNKIVAFYLKNPYARLTLLYSHGNAADLGQLYDLFVQLKVNLRVNLMGYDYSGYGAST 119
+ G+K+VAFYL+NP ARLT+LYSHGNAADLGQLYDLFVQLKVNL+VNLMGYDYSGYGAST
Sbjct: 57 RKGSKVVAFYLRNPAARLTVLYSHGNAADLGQLYDLFVQLKVNLKVNLMGYDYSGYGAST 116
Query: 120 GKPSESSTYADIEAIYECLESEYGVGQEDIILYGQSVGSGPTLHLAAKLPRLRGVVLHSG 179
GKPSE +TYADIEA+Y+CLE+EYG+ QED+ILYGQSVGSGPTLHLA++LPRLRGVVLHS
Sbjct: 117 GKPSEENTYADIEAVYQCLETEYGISQEDLILYGQSVGSGPTLHLASRLPRLRGVVLHSA 176
Query: 180 ILSGLRVLCHVKFSFCFDIYKNINKIKKVKCPVLVIHGTEDDVVNWLHGNGLWKMARESY 239
ILSGLRV+CHV F+FCFDIYKN+ KIKKVK PVLVIHGT+DDVVNW HGN LWK+ARE Y
Sbjct: 177 ILSGLRVVCHVNFTFCFDIYKNVKKIKKVKSPVLVIHGTDDDVVNWSHGNELWKLAREPY 236
Query: 240 DPLWIKGGGHCNLELYPDYIRHLCKFIQEMESITTEKRLKKLRQSQGP---------QSN 290
DPLWIKGGGHCNLELYPD+IRHL KFI+EME+ITT+ RLKK+RQS P +
Sbjct: 237 DPLWIKGGGHCNLELYPDFIRHLSKFIREMENITTKTRLKKIRQSLQPAPKKVHHRASTG 296
Query: 291 SSSCFSCSCC---RMKCCSNNCTGCSNCSCINPECCN--CSCINPECFNCSCINCSCSLK 345
+++ F+ +CC R++ S C GC N SC CC+ SC + F CS SCS +
Sbjct: 297 TTTTFTTNCCCRIRVRKPSCRCPGC-NLSC---GCCSGLMSCFSSRLFKCSTCFSSCSCR 352
Query: 346 CTQCCCLPSCIKCCHLPKFSTCFGSSCCTKCSRPSCCFKC--CCNLRCARPSCCCMSC 401
C KC TCF SC SCC C C +C CCC SC
Sbjct: 353 --------GCFKC------PTCFSFSC-------SCCRSCLKCPTFKC----CCCGSC 385
>UniRef100_Q84JV3 Hypothetical protein At4g31020 [Arabidopsis thaliana]
Length = 294
Score = 348 bits (893), Expect = 2e-94
Identities = 166/274 (60%), Positives = 213/274 (77%), Gaps = 10/274 (3%)
Query: 1 MGCMVSQLAAKFAFFPPSPPTYQLKKRDD-GKLTVVSAASPSVPIPHADDSSLDVLLVDT 59
MG + S +AAKFAFFPP P TY + K D+ GKL ++ D +++V + T
Sbjct: 1 MGNVTSNVAAKFAFFPPEPATYGVTKDDETGKLVFAGVSA---------DKNVEVHQLTT 51
Query: 60 KHGNKIVAFYLKNPYARLTLLYSHGNAADLGQLYDLFVQLKVNLRVNLMGYDYSGYGAST 119
K GNK+VA + ++P+AR TLLYSHGNAADLGQ+ +LF++L+ +LRVN+M YDYSGYGAST
Sbjct: 52 KSGNKVVATFWRHPFARFTLLYSHGNAADLGQMVELFIELRAHLRVNIMSYDYSGYGAST 111
Query: 120 GKPSESSTYADIEAIYECLESEYGVGQEDIILYGQSVGSGPTLHLAAKLPRLRGVVLHSG 179
GKPSE +TY DIEA+Y CL S+YG+ QE+IILYGQSVGSGPTLH+A++L RLRGVVLHS
Sbjct: 112 GKPSEFNTYYDIEAVYSCLRSDYGIKQEEIILYGQSVGSGPTLHMASRLKRLRGVVLHSA 171
Query: 180 ILSGLRVLCHVKFSFCFDIYKNINKIKKVKCPVLVIHGTEDDVVNWLHGNGLWKMARESY 239
ILSG+RVL VK + FDI+KNI+KI+ V VLVIHGT D++V+ HG LW++A+E Y
Sbjct: 172 ILSGIRVLYPVKMTLWFDIFKNIDKIRHVNSQVLVIHGTNDEIVDLSHGKRLWELAKEKY 231
Query: 240 DPLWIKGGGHCNLELYPDYIRHLCKFIQEMESIT 273
DPLW+KGGGHCNLE YP+YI+HL KF+ ME ++
Sbjct: 232 DPLWVKGGGHCNLETYPEYIKHLKKFVNAMEKLS 265
>UniRef100_Q9LI62 Similarity to unknown protein [Arabidopsis thaliana]
Length = 399
Score = 348 bits (892), Expect = 2e-94
Identities = 165/270 (61%), Positives = 212/270 (78%), Gaps = 9/270 (3%)
Query: 1 MGCMVSQLAAKFAFFPPSPPTYQLKKRDDGKLTVVSAASPSVPIPHADDSSLDVLLVDTK 60
MG + S +AAKFAFFPP+PP+Y ++ + GKL ++ + +++VL + TK
Sbjct: 1 MGAVTSSMAAKFAFFPPNPPSYGVEVVE-GKLRLIGVENVK--------ENVEVLKLKTK 51
Query: 61 HGNKIVAFYLKNPYARLTLLYSHGNAADLGQLYDLFVQLKVNLRVNLMGYDYSGYGASTG 120
GN++VA Y+KNP A LTLLYSHGNAADLGQ+++LF +L ++LRVNL+GYDYSGYG S+G
Sbjct: 52 RGNQVVAAYIKNPTASLTLLYSHGNAADLGQMFELFSELSLHLRVNLIGYDYSGYGRSSG 111
Query: 121 KPSESSTYADIEAIYECLESEYGVGQEDIILYGQSVGSGPTLHLAAKLPRLRGVVLHSGI 180
KPSE +TY+DIEA+Y CLE +YGV ++D+ILYGQSVGSGPTL LA++LP LR VVLHS I
Sbjct: 112 KPSEQNTYSDIEAVYRCLEEKYGVKEQDVILYGQSVGSGPTLELASRLPNLRAVVLHSAI 171
Query: 181 LSGLRVLCHVKFSFCFDIYKNINKIKKVKCPVLVIHGTEDDVVNWLHGNGLWKMARESYD 240
SGLRV+ VK ++ FDIYKN+ KI VKCPVLVIHGT DDVVNW HG L+++ +E Y+
Sbjct: 172 ASGLRVMYPVKRTYWFDIYKNVEKISFVKCPVLVIHGTSDDVVNWSHGKQLFELCKEKYE 231
Query: 241 PLWIKGGGHCNLELYPDYIRHLCKFIQEME 270
PLWIKGG HC+LELYP YI+HL KF+ +E
Sbjct: 232 PLWIKGGNHCDLELYPQYIKHLRKFVSAIE 261
>UniRef100_Q9SB70 Hypothetical protein F22K18.40 [Arabidopsis thaliana]
Length = 365
Score = 343 bits (881), Expect = 4e-93
Identities = 164/284 (57%), Positives = 219/284 (76%), Gaps = 8/284 (2%)
Query: 1 MGCMVSQLAAKFAFFPPSPPTYQLKKRDDGKLTVVSAASPSVPIPHADDSSLDVLLVDTK 60
MG + S +AAK AFFPP+PP+Y+L + + +L ++S P PH ++ +D+L + T+
Sbjct: 1 MGGVTSSMAAKLAFFPPNPPSYKLVRDETTELFLMS------PFPHREN--VDILRLPTR 52
Query: 61 HGNKIVAFYLKNPYARLTLLYSHGNAADLGQLYDLFVQLKVNLRVNLMGYDYSGYGASTG 120
G +IVA Y++ P A TLLYSHGNAAD+GQ+Y+LF++L ++LRVNLMGYDYSGYG S+G
Sbjct: 53 RGTEIVAMYIRYPMAVTTLLYSHGNAADIGQMYELFIELSIHLRVNLMGYDYSGYGQSSG 112
Query: 121 KPSESSTYADIEAIYECLESEYGVGQEDIILYGQSVGSGPTLHLAAKLPRLRGVVLHSGI 180
KP+E +TYADIEA Y+CLE YG QE+IILYGQSVGSGPT+ LAA+LPRLR +LHS I
Sbjct: 113 KPTEQNTYADIEAAYKCLEENYGAKQENIILYGQSVGSGPTVDLAARLPRLRASILHSPI 172
Query: 181 LSGLRVLCHVKFSFCFDIYKNINKIKKVKCPVLVIHGTEDDVVNWLHGNGLWKMARESYD 240
LSGLRV+ VK ++ FDIYKNI+KI V+CPVLVIHGT DDVV++ HG LW++ +E Y+
Sbjct: 173 LSGLRVMYPVKRTYWFDIYKNIDKITLVRCPVLVIHGTADDVVDFSHGKQLWELCQEKYE 232
Query: 241 PLWIKGGGHCNLELYPDYIRHLCKFIQEMESITTEKRLKKLRQS 284
PLW+KGG HC+LEL+P+YI HL KF+ +E +++ R+S
Sbjct: 233 PLWLKGGNHCDLELFPEYIGHLKKFVSAVEKSASKRNSSFSRRS 276
>UniRef100_Q9ZQ35 Hypothetical protein At2g24320 [Arabidopsis thaliana]
Length = 316
Score = 342 bits (878), Expect = 9e-93
Identities = 166/295 (56%), Positives = 216/295 (72%), Gaps = 32/295 (10%)
Query: 1 MGCMVSQLAAKFAFFPPSPPTYQLKKRDDGKLTVVSAASPSVPIPHADDSSLDVLLVDTK 60
MG + S +AAKFAFFPP PPTY + K ++ + + +P + S+DV + TK
Sbjct: 1 MGNVTSNMAAKFAFFPP-PPTYDVGKDEETGKLMFTGITP--------EKSMDVHQLTTK 51
Query: 61 HGNKIVAFYLKNPYARLTLLYSHGNAADLGQLYDLFVQLKVNLRVNLMGYDYSGYGASTG 120
GNK++A + K+P++R TLLYSHGNAADLGQ+ DLF++L+ +LRVN+M YDYSGYGASTG
Sbjct: 52 SGNKVIATFWKHPFSRFTLLYSHGNAADLGQMVDLFIELRAHLRVNIMSYDYSGYGASTG 111
Query: 121 KPSESSTYADIEAIYECLESEYGVGQEDIILYGQSVGSGPTLHLAAKLPRLRGVVLHSGI 180
KP+E +TY DIEA+Y CL +EYG+ QE++ILYGQSVGSGPTLHLA+++ RLRG+VLHS I
Sbjct: 112 KPTELNTYYDIEAVYNCLRTEYGIMQEEMILYGQSVGSGPTLHLASRVKRLRGIVLHSAI 171
Query: 181 LSGLRVLCHVKFSFCFDIYK-----------------------NINKIKKVKCPVLVIHG 217
LSGLRVL VK +F FD+YK NI+KI+ V CPVLVIHG
Sbjct: 172 LSGLRVLYPVKMTFWFDMYKVSLISLVSGYYYRVSLSNSGILQNIDKIRHVTCPVLVIHG 231
Query: 218 TEDDVVNWLHGNGLWKMARESYDPLWIKGGGHCNLELYPDYIRHLCKFIQEMESI 272
T+DD+VN HG LW++A++ YDPLW+KGGGHCNLE YP+YI+H+ KF+ ME +
Sbjct: 232 TKDDIVNMSHGKRLWELAKDKYDPLWVKGGGHCNLETYPEYIKHMRKFMNAMEKL 286
>UniRef100_Q8S9U8 P0519D04.38 protein [Oryza sativa]
Length = 546
Score = 337 bits (865), Expect = 3e-91
Identities = 169/313 (53%), Positives = 216/313 (68%), Gaps = 26/313 (8%)
Query: 1 MGCMVSQLAAKFAFFPPSPPTYQL---------------------KKRDDGKLTVVSAAS 39
MG + S +AA+FAFFPP+PP+Y + ++RD G S A+
Sbjct: 183 MGGVTSTIAARFAFFPPTPPSYTVVADAATGRLAIPEISRPPARRRRRDGGGDASASGAA 242
Query: 40 PSVPIPHADDSSLDVLLVDTKHGNKIVAFYLKNPYARLTLLYSHGNAADLGQLYDLFVQL 99
P+ D+ +V+ + T+ GN+IV ++++ A TLLYSHGNAADLGQ+Y LFV+L
Sbjct: 243 PA-----EDEDGTEVVRLRTRRGNEIVGVHVRHERASATLLYSHGNAADLGQMYGLFVEL 297
Query: 100 KVNLRVNLMGYDYSGYGASTGKPSESSTYADIEAIYECLESEYGVGQEDIILYGQSVGSG 159
LR+NL GYDYSGYG STGKP+E +TYADIEA Y CL+ +YGV EDIILYGQSVGSG
Sbjct: 298 SRRLRINLFGYDYSGYGRSTGKPTECNTYADIEAAYNCLKEKYGVADEDIILYGQSVGSG 357
Query: 160 PTLHLAAKLPRLRGVVLHSGILSGLRVLCHVKFSFCFDIYKNINKIKKVKCPVLVIHGTE 219
PT+ LA++LP LRGVVLHS ILSGLRVL VK ++ FDIYKNI+KI V CPVLVIHGT
Sbjct: 358 PTIDLASRLPNLRGVVLHSPILSGLRVLYPVKRTYWFDIYKNIDKIGLVNCPVLVIHGTS 417
Query: 220 DDVVNWLHGNGLWKMARESYDPLWIKGGGHCNLELYPDYIRHLCKFIQEMESITTEKRLK 279
DDVV+ HG LW++ + Y PLW+ GGGHCNLELYPDYI+HL KF+ + +++ LK
Sbjct: 418 DDVVDCSHGKQLWELCKVKYSPLWLTGGGHCNLELYPDYIKHLKKFVSSLGKKSSKPDLK 477
Query: 280 KLRQSQGPQSNSS 292
++ +G S S
Sbjct: 478 EITMKEGASSKDS 490
>UniRef100_Q9S7U7 F28J7.1 protein [Arabidopsis thaliana]
Length = 361
Score = 336 bits (861), Expect = 8e-91
Identities = 160/272 (58%), Positives = 212/272 (77%), Gaps = 8/272 (2%)
Query: 1 MGCMVSQLAAKFAFFPPSPPTYQLKKRDDGKLTVVSAASPSVPIPHADDSSLDVLLVDTK 60
MG + S +AAKFAFFPPSPP+Y++ + L ++S P PH ++ ++++ + T+
Sbjct: 1 MGGVTSSVAAKFAFFPPSPPSYKVVTDELTGLLLLS------PFPHREN--VEIVKLRTR 52
Query: 61 HGNKIVAFYLKNPYARLTLLYSHGNAADLGQLYDLFVQLKVNLRVNLMGYDYSGYGASTG 120
G +IV Y+++P A TLLYSHGNAADLGQ+Y+LF++L ++L+VNLMGYDYSGYG STG
Sbjct: 53 RGTEIVGMYVRHPMATSTLLYSHGNAADLGQMYELFIELSIHLKVNLMGYDYSGYGQSTG 112
Query: 121 KPSESSTYADIEAIYECLESEYGVGQEDIILYGQSVGSGPTLHLAAKLPRLRGVVLHSGI 180
KPSE +TYADIEA+Y+CLE +G QE +ILYGQSVGSGPTL LA++LP+LR VVLHS I
Sbjct: 113 KPSEHNTYADIEAVYKCLEETFGSKQEGVILYGQSVGSGPTLDLASRLPQLRAVVLHSPI 172
Query: 181 LSGLRVLCHVKFSFCFDIYKNINKIKKVKCPVLVIHGTEDDVVNWLHGNGLWKMARESYD 240
LSGLRV+ VK ++ FDIYKNI+KI V CPVL+IHGT D+VV+ HG LW++ ++ Y+
Sbjct: 173 LSGLRVMYSVKKTYWFDIYKNIDKIPYVDCPVLIIHGTSDEVVDCSHGKQLWELCKDKYE 232
Query: 241 PLWIKGGGHCNLELYPDYIRHLCKFIQEMESI 272
PLW+KGG HC+LE YP+YIRHL KFI +E +
Sbjct: 233 PLWVKGGNHCDLEHYPEYIRHLKKFIATVERL 264
>UniRef100_Q9LY90 Hypothetical protein F18O22_180 [Arabidopsis thaliana]
Length = 369
Score = 333 bits (855), Expect = 4e-90
Identities = 163/270 (60%), Positives = 210/270 (77%), Gaps = 8/270 (2%)
Query: 1 MGCMVSQLAAKFAFFPPSPPTYQLKKRDDGKLTVVSAASPSVPIPHADDSSLDVLLVDTK 60
MG + S +AAKFAFFPPSP +Y+L + L +++ P PH ++ +++L + T+
Sbjct: 1 MGGVTSSVAAKFAFFPPSPSSYKLVYDELTGLLLMN------PFPHREN--VEILKLPTR 52
Query: 61 HGNKIVAFYLKNPYARLTLLYSHGNAADLGQLYDLFVQLKVNLRVNLMGYDYSGYGASTG 120
G +IVA Y+++P A TLLYSHGNAADLGQ+Y+LF++L ++L+VNLMGYDYSGYG STG
Sbjct: 53 RGTEIVAMYVRHPMATSTLLYSHGNAADLGQMYELFIELSIHLKVNLMGYDYSGYGQSTG 112
Query: 121 KPSESSTYADIEAIYECLESEYGVGQEDIILYGQSVGSGPTLHLAAKLPRLRGVVLHSGI 180
KPSE TYADIEA Y+CLE YG QEDIILYGQSVGSGPTL LAA+LP+LR VLHS I
Sbjct: 113 KPSEHHTYADIEAAYKCLEETYGAKQEDIILYGQSVGSGPTLDLAARLPQLRAAVLHSPI 172
Query: 181 LSGLRVLCHVKFSFCFDIYKNINKIKKVKCPVLVIHGTEDDVVNWLHGNGLWKMARESYD 240
LSGLRV+ VK ++ FDI+KNI+KI V CPVLVIHGT D+VV+ HG LW++++E Y+
Sbjct: 173 LSGLRVMYPVKKTYWFDIFKNIDKIPLVNCPVLVIHGTCDEVVDCSHGKQLWELSKEKYE 232
Query: 241 PLWIKGGGHCNLELYPDYIRHLCKFIQEME 270
PLW++GG HC+LE YP+YI+HL KFI +E
Sbjct: 233 PLWLEGGNHCDLEHYPEYIKHLKKFITTVE 262
>UniRef100_Q9FF34 Emb|CAB87778.1 [Arabidopsis thaliana]
Length = 336
Score = 332 bits (852), Expect = 9e-90
Identities = 169/317 (53%), Positives = 222/317 (69%), Gaps = 22/317 (6%)
Query: 1 MGCMVSQLAAKFAFFPPSPPTYQLKKRDDGKLTVVSAASPSVPIPHADDSSLDVLLVDTK 60
MG + S +AAKFAFFPPSPP+Y D +L + +P DD +DVL + T+
Sbjct: 1 MGGVTSSIAAKFAFFPPSPPSYGFVS-DVDRLYITE-------VPRRDD--VDVLKLKTR 50
Query: 61 HGNKIVAFYLKNPYARLTLLYSHGNAADLGQLYDLFVQLKVNLRVNLMGYDYSGYGASTG 120
GN+IVA Y+K+P A TLLYSHGNAADLGQ+++LF++L LR+NLMGYDYSGYG STG
Sbjct: 51 RGNEIVAIYIKHPKANGTLLYSHGNAADLGQMFELFIELSNRLRLNLMGYDYSGYGQSTG 110
Query: 121 KPSESSTYADIEAIYECLESEYGVGQEDIILYGQSVGSGPTLHLAAKLPRLRGVVLHSGI 180
K SE +TYADI+A Y CL+ YGV + +ILYGQSVGSGPT+ LA++ P LRGVVLHS I
Sbjct: 111 KASECNTYADIDAAYTCLKEHYGVKDDQLILYGQSVGSGPTIDLASRTPNLRGVVLHSPI 170
Query: 181 LSGLRVLCHVKFSFCFDIYKNINKIKKVKCPVLVIHGTEDDVVNWLHGNGLWKMARESYD 240
LSG+RVL VK ++ FDIYKNI+KI V CPVLVIHGT D+VV+ HG LW++++E Y+
Sbjct: 171 LSGMRVLYPVKRTYWFDIYKNIDKIGAVTCPVLVIHGTADEVVDCSHGKQLWELSKEKYE 230
Query: 241 PLWIKGGGHCNLELYPDYIRHLCKFIQEM---------ESITTEKRLKKLRQSQGPQSNS 291
PLW+ GGGHCNLELYP++I+HL K++ + ++ TT+ K QS+ ++
Sbjct: 231 PLWVSGGGHCNLELYPEFIKHLKKYVISISKGPRTGSNKTATTDAAKK---QSKPAENGR 287
Query: 292 SSCFSCSCCRMKCCSNN 308
+ F CC + N+
Sbjct: 288 ADTFQLGCCLPEVSRNS 304
>UniRef100_Q8VYT1 Hypothetical protein At1g66900 [Arabidopsis thaliana]
Length = 272
Score = 328 bits (842), Expect = 1e-88
Identities = 163/278 (58%), Positives = 210/278 (74%), Gaps = 11/278 (3%)
Query: 1 MGCMVSQLAAKFAFFPPSPPTYQLKKRDD--GKLTVVSAASPSVPIPHADDSSLDVLLVD 58
MG + S +AAKFAFFPP+PP+Y++ D G+L + IP DD +D+L +
Sbjct: 1 MGGVTSSIAAKFAFFPPTPPSYEVIADDSCGGRLYIPE-------IPRRDD--VDILKLR 51
Query: 59 TKHGNKIVAFYLKNPYARLTLLYSHGNAADLGQLYDLFVQLKVNLRVNLMGYDYSGYGAS 118
T+ GN+IVA Y+K+ A TLLYSHGNAADLGQ+++LFV+L LRVNLMGYDYSGYG S
Sbjct: 52 TRCGNEIVAVYVKHSKANGTLLYSHGNAADLGQMFELFVELSNRLRVNLMGYDYSGYGQS 111
Query: 119 TGKPSESSTYADIEAIYECLESEYGVGQEDIILYGQSVGSGPTLHLAAKLPRLRGVVLHS 178
TG+ SE +TYADIEA Y+CL+ +YGV + +I+YGQSVGSGPT+ LA++ P LRGVVL
Sbjct: 112 TGQASECNTYADIEASYKCLKEKYGVKDDQLIVYGQSVGSGPTVDLASRTPNLRGVVLQC 171
Query: 179 GILSGLRVLCHVKFSFCFDIYKNINKIKKVKCPVLVIHGTEDDVVNWLHGNGLWKMARES 238
ILSG+RVL VK ++ FDIYKNI+KI V CPVLVIHGT D+VV+W HG LW++++E
Sbjct: 172 PILSGMRVLYPVKCTYWFDIYKNIDKIGSVTCPVLVIHGTADEVVDWSHGKRLWELSKEK 231
Query: 239 YDPLWIKGGGHCNLELYPDYIRHLCKFIQEMESITTEK 276
Y+PLWI GGGHC+LELYPD+IRHL KF+ + + E+
Sbjct: 232 YEPLWISGGGHCDLELYPDFIRHLKKFVVSLGNKQAEQ 269
>UniRef100_Q67VK9 Cgi67 serine protease-like [Oryza sativa]
Length = 389
Score = 328 bits (841), Expect = 2e-88
Identities = 164/303 (54%), Positives = 213/303 (70%), Gaps = 22/303 (7%)
Query: 1 MGCMVSQLAAKFAFFPPSPPTYQLKK---------RDDGKLTVVSAASPSVPIPHADDSS 51
MG + S +AA+FAFFPP+PP+Y ++ +DG + +S +P +
Sbjct: 1 MGAVASTVAARFAFFPPAPPSYGVEPPPSPSPAAAAEDGAVVELSG------VPRR--AG 52
Query: 52 LDVLLVDTKHGNKIVAFYLKNPYARLTLLYSHGNAADLGQLYDLFVQLKVNLRVNLMGYD 111
++ + T G ++VA Y++ P ARLTLLYSHGNAADLGQ+Y+LFV+L +L VNLMGYD
Sbjct: 53 VEARRLPTGRGTEVVAMYVRQPGARLTLLYSHGNAADLGQMYELFVELSSHLNVNLMGYD 112
Query: 112 YSGYGASTGKPSESSTYADIEAIYECLESEYGVGQEDIILYGQSVGSGPTLHLAAKLPRL 171
YSGYG S+GKPSE +TY+DIEA Y CL YG +E+IILYGQSVGSGPTL LA++LP L
Sbjct: 113 YSGYGQSSGKPSEQNTYSDIEAAYRCLVETYGATEENIILYGQSVGSGPTLDLASRLPHL 172
Query: 172 RGVVLHSGILSGLRVLCHVKFSFCFDIYKNINKIKKVKCPVLVIHGTEDDVVNWLHGNGL 231
R VVLHS ILSGLRV+ VK ++ FDIYKNI+K+ VKCPVLVIHGT D+VV+ HG L
Sbjct: 173 RAVVLHSPILSGLRVMYPVKHTYWFDIYKNIDKVPLVKCPVLVIHGTADEVVDCSHGRAL 232
Query: 232 WKMARESYDPLWIKGGGHCNLELYPDYIRHLCKFIQEMESITTEKRLKKLRQSQGPQSNS 291
W++++ Y+PLW+KGG HCNLELYP+YI+HL KF+ +E + K +S G S
Sbjct: 233 WELSKIKYEPLWVKGGNHCNLELYPEYIKHLKKFVMAIEKLPPTK-----DESSGSSGPS 287
Query: 292 SSC 294
C
Sbjct: 288 DPC 290
>UniRef100_Q6YUU0 Putative Cgi67 serine protease [Oryza sativa]
Length = 389
Score = 326 bits (835), Expect = 9e-88
Identities = 165/307 (53%), Positives = 216/307 (69%), Gaps = 21/307 (6%)
Query: 1 MGCMVSQLAAKFAFFPPSPPTYQLK--------------KRDDGKLTVVSAASPSVPIPH 46
MG + S +AA+FAFFPPSPP+Y + ++D G VV + +P
Sbjct: 1 MGAVTSTVAARFAFFPPSPPSYGAEAPPPPAAAGAGVGVEKDGGGGGVVVELTD---VPR 57
Query: 47 ADDSSLDVLLVDTKHGNKIVAFYLKNPYARLTLLYSHGNAADLGQLYDLFVQLKVNLRVN 106
+++ + TK G ++VA Y++ ARLTLLYSHGNAADLGQ+++LFV+L +L VN
Sbjct: 58 R--GNVEARRLRTKRGTEVVAMYVRQAGARLTLLYSHGNAADLGQMFELFVELSAHLNVN 115
Query: 107 LMGYDYSGYGASTGKPSESSTYADIEAIYECLESEYGVGQEDIILYGQSVGSGPTLHLAA 166
LMGYDYSGYG S+GKPSE +TYADIEA+Y CL YG +E+IILYGQSVGSGPTL LA+
Sbjct: 116 LMGYDYSGYGQSSGKPSEHNTYADIEAVYRCLVETYGASEENIILYGQSVGSGPTLDLAS 175
Query: 167 KLPRLRGVVLHSGILSGLRVLCHVKFSFCFDIYKNINKIKKVKCPVLVIHGTEDDVVNWL 226
+LP LR VVLHS ILSGLRV+ VK ++ FDIYKNI+KI V+CPVLVIHGT D+VV+
Sbjct: 176 RLPHLRAVVLHSPILSGLRVMYPVKHTYWFDIYKNIDKIPLVRCPVLVIHGTADEVVDCS 235
Query: 227 HGNGLWKMARESYDPLWIKGGGHCNLELYPDYIRHLCKFIQEMESITTEKRLKKLRQSQG 286
HG LW++++ Y+PLW+KGG HCNLELYP+YI+HL KF+ +E + + +S G
Sbjct: 236 HGRALWELSKVKYEPLWVKGGNHCNLELYPEYIKHLKKFVGAIEK--SPPLYDESPESSG 293
Query: 287 PQSNSSS 293
P N+ +
Sbjct: 294 PSDNTQT 300
>UniRef100_Q6KAK2 Hypothetical protein OJ1004_E04.17 [Oryza sativa]
Length = 264
Score = 325 bits (832), Expect = 2e-87
Identities = 151/243 (62%), Positives = 195/243 (80%)
Query: 49 DSSLDVLLVDTKHGNKIVAFYLKNPYARLTLLYSHGNAADLGQLYDLFVQLKVNLRVNLM 108
D+ ++V + TK G ++VA + ++P ARLTLLYSHGNAADLGQ+ LF++L+ +LRVN+M
Sbjct: 7 DAGVEVHALPTKGGTRVVAAFWRHPSARLTLLYSHGNAADLGQMLGLFLELRAHLRVNIM 66
Query: 109 GYDYSGYGASTGKPSESSTYADIEAIYECLESEYGVGQEDIILYGQSVGSGPTLHLAAKL 168
YDYSGYGASTGKPSE +TY DIEA+Y+CL YG+ ED+ILYGQSVGSGPTLHLA++L
Sbjct: 67 SYDYSGYGASTGKPSEYNTYCDIEAVYDCLTKVYGIEPEDLILYGQSVGSGPTLHLASRL 126
Query: 169 PRLRGVVLHSGILSGLRVLCHVKFSFCFDIYKNINKIKKVKCPVLVIHGTEDDVVNWLHG 228
+LRGVVLHS ILSG+RVL VK + FDI+KNI+KIK+V CPVLVIHGT DD+V++ HG
Sbjct: 127 EKLRGVVLHSAILSGIRVLYPVKVTLWFDIFKNIDKIKQVDCPVLVIHGTADDIVDFSHG 186
Query: 229 NGLWKMARESYDPLWIKGGGHCNLELYPDYIRHLCKFIQEMESITTEKRLKKLRQSQGPQ 288
LW++A+E Y+PLW+KGGGHCNLE YP+YIRHL KFI ME ++ +K K + +
Sbjct: 187 KRLWELAKEKYEPLWVKGGGHCNLETYPEYIRHLRKFINAMEKLSKDKTAKAPQLAPSSS 246
Query: 289 SNS 291
+N+
Sbjct: 247 NNN 249
>UniRef100_Q9LMY7 F21F23.4 protein [Arabidopsis thaliana]
Length = 341
Score = 313 bits (802), Expect = 6e-84
Identities = 157/278 (56%), Positives = 201/278 (71%), Gaps = 9/278 (3%)
Query: 1 MGCMVSQLAAKFAFFPPSPPTYQLKKRDD-GKLTVVSAASPSVPIPH-ADDSSLDVLLVD 58
MG S +AAK AFFPP+PP+Y + + GK+ + S +PH D +++V+ +
Sbjct: 1 MGIATSTMAAKLAFFPPNPPSYTVVTEESTGKMMI------STNLPHYLRDENIEVVKIR 54
Query: 59 TKHGNKIVAFYLKNPYARLTLLYSHGNAADLGQLYDLFVQLKVNLRVNLMGYDYSGYGAS 118
TK GN+IVA Y+KNP A+LT+L+SHGNA+DL Q++ + +L + L VNLMGYDYSGYG S
Sbjct: 55 TKRGNEIVAMYVKNPTAKLTVLFSHGNASDLAQIFYILAEL-IQLNVNLMGYDYSGYGQS 113
Query: 119 TGKPSESSTYADIEAIYECLESEYGVGQEDIILYGQSVGSGPTLHLAAKLPRLRGVVLHS 178
+GKPSE TYADIEA Y L YG E IILYGQSVGSGP+L LA++LPRLR +VLHS
Sbjct: 114 SGKPSEQDTYADIEAAYNWLRQTYGTKDERIILYGQSVGSGPSLELASRLPRLRALVLHS 173
Query: 179 GILSGLRVLCHVKFSFCFDIYKNINKIKKVKCPVLVIHGTEDDVVNWLHGNGLWKMARES 238
LSGLRV+ VK SF FDIYKNI+KI V+CPVLVIHGT+DDVVN HG LW + +E
Sbjct: 174 PFLSGLRVMYPVKHSFPFDIYKNIDKIHLVECPVLVIHGTDDDVVNISHGKHLWGLCKEK 233
Query: 239 YDPLWIKGGGHCNLELYPDYIRHLCKFIQEMESITTEK 276
Y+PLW+KG GH ++E+ P+Y+ HL KFI +E + K
Sbjct: 234 YEPLWLKGRGHSDIEMSPEYLPHLRKFISAIEKLPVPK 271
>UniRef100_Q9FZ68 F13B4.9 protein [Arabidopsis thaliana]
Length = 358
Score = 313 bits (802), Expect = 6e-84
Identities = 157/278 (56%), Positives = 201/278 (71%), Gaps = 9/278 (3%)
Query: 1 MGCMVSQLAAKFAFFPPSPPTYQLKKRDD-GKLTVVSAASPSVPIPH-ADDSSLDVLLVD 58
MG S +AAK AFFPP+PP+Y + + GK+ + S +PH D +++V+ +
Sbjct: 1 MGIATSTMAAKLAFFPPNPPSYTVVTEESTGKMMI------STNLPHYLRDENIEVVKIR 54
Query: 59 TKHGNKIVAFYLKNPYARLTLLYSHGNAADLGQLYDLFVQLKVNLRVNLMGYDYSGYGAS 118
TK GN+IVA Y+KNP A+LT+L+SHGNA+DL Q++ + +L + L VNLMGYDYSGYG S
Sbjct: 55 TKRGNEIVAMYVKNPTAKLTVLFSHGNASDLAQIFYILAEL-IQLNVNLMGYDYSGYGQS 113
Query: 119 TGKPSESSTYADIEAIYECLESEYGVGQEDIILYGQSVGSGPTLHLAAKLPRLRGVVLHS 178
+GKPSE TYADIEA Y L YG E IILYGQSVGSGP+L LA++LPRLR +VLHS
Sbjct: 114 SGKPSEQDTYADIEAAYNWLRQTYGTKDERIILYGQSVGSGPSLELASRLPRLRALVLHS 173
Query: 179 GILSGLRVLCHVKFSFCFDIYKNINKIKKVKCPVLVIHGTEDDVVNWLHGNGLWKMARES 238
LSGLRV+ VK SF FDIYKNI+KI V+CPVLVIHGT+DDVVN HG LW + +E
Sbjct: 174 PFLSGLRVMYPVKHSFPFDIYKNIDKIHLVECPVLVIHGTDDDVVNISHGKHLWGLCKEK 233
Query: 239 YDPLWIKGGGHCNLELYPDYIRHLCKFIQEMESITTEK 276
Y+PLW+KG GH ++E+ P+Y+ HL KFI +E + K
Sbjct: 234 YEPLWLKGRGHSDIEMSPEYLPHLRKFISAIEKLPVPK 271
>UniRef100_Q6ZCV6 Hypothetical protein P0028A08.22 [Oryza sativa]
Length = 414
Score = 311 bits (797), Expect = 2e-83
Identities = 156/284 (54%), Positives = 203/284 (70%), Gaps = 10/284 (3%)
Query: 1 MGCMVSQLAAKFAFFPPSPPTYQLKKRDD----GKLTVVSAASPS----VPIPHADDSSL 52
MG + S +AAKFAFFPP PP Y + ++ G +AA+ +P + +
Sbjct: 1 MGGVTSSVAAKFAFFPPDPPAYGVADEEEPPPPGSAAPAAAATARRVSLTGVPWRE--GV 58
Query: 53 DVLLVDTKHGNKIVAFYLKNPYARLTLLYSHGNAADLGQLYDLFVQLKVNLRVNLMGYDY 112
+ V T+ G +I+A Y++ P ARLT+LYSHGNAAD+G++Y+LFV+ L VNLMGYDY
Sbjct: 59 EARRVRTRRGTEIIAVYVRCPKARLTVLYSHGNAADIGKMYELFVEFSARLHVNLMGYDY 118
Query: 113 SGYGASTGKPSESSTYADIEAIYECLESEYGVGQEDIILYGQSVGSGPTLHLAAKLPRLR 172
SGYG S+GK SE++T+ADIEA Y+CL YG +EDIILYGQSVGSGPT+ LAA+L R+R
Sbjct: 119 SGYGRSSGKASEANTFADIEAAYKCLVEVYGTREEDIILYGQSVGSGPTVDLAAQLHRIR 178
Query: 173 GVVLHSGILSGLRVLCHVKFSFCFDIYKNINKIKKVKCPVLVIHGTEDDVVNWLHGNGLW 232
VVLHS ILSGLRV+ VK ++ FDIYKNI K+ VK PVLVIHGT DD+V+ HG LW
Sbjct: 179 AVVLHSPILSGLRVMYSVKKTYWFDIYKNIEKMPLVKSPVLVIHGTNDDIVDCSHGKQLW 238
Query: 233 KMARESYDPLWIKGGGHCNLELYPDYIRHLCKFIQEMESITTEK 276
++ + Y+PLWI+GG HCNL+ +P YIRHL KFI +E++ EK
Sbjct: 239 ELCQNKYEPLWIEGGDHCNLQTFPVYIRHLKKFISTIENMPLEK 282
>UniRef100_Q5XVL1 Hypothetical protein [Arabidopsis thaliana]
Length = 358
Score = 310 bits (793), Expect = 6e-83
Identities = 156/278 (56%), Positives = 200/278 (71%), Gaps = 9/278 (3%)
Query: 1 MGCMVSQLAAKFAFFPPSPPTYQLKKRDD-GKLTVVSAASPSVPIPH-ADDSSLDVLLVD 58
MG S +AAK AFFPP+PP+Y + + GK+ + S +PH D +++V+ +
Sbjct: 1 MGIATSTMAAKLAFFPPNPPSYTVVTEESTGKMMI------STNLPHYLRDENIEVVKIR 54
Query: 59 TKHGNKIVAFYLKNPYARLTLLYSHGNAADLGQLYDLFVQLKVNLRVNLMGYDYSGYGAS 118
TK GN+IVA Y+KNP A+LT+L+S GNA+DL Q++ + +L + L VNLMGYDYSGYG S
Sbjct: 55 TKRGNEIVAMYVKNPTAKLTVLFSXGNASDLAQIFYILAEL-IQLNVNLMGYDYSGYGQS 113
Query: 119 TGKPSESSTYADIEAIYECLESEYGVGQEDIILYGQSVGSGPTLHLAAKLPRLRGVVLHS 178
+GKPSE TYADIEA Y L YG E IILYGQSVGSGP+L LA++LPRLR +VLHS
Sbjct: 114 SGKPSEQDTYADIEAAYNWLRQTYGTKDERIILYGQSVGSGPSLELASRLPRLRALVLHS 173
Query: 179 GILSGLRVLCHVKFSFCFDIYKNINKIKKVKCPVLVIHGTEDDVVNWLHGNGLWKMARES 238
LSGLRV+ VK SF FDIYKNI+KI V+CPVLVIHGT+DDVVN HG LW + +E
Sbjct: 174 PFLSGLRVMYPVKHSFPFDIYKNIDKIHLVECPVLVIHGTDDDVVNISHGKHLWGLCKEK 233
Query: 239 YDPLWIKGGGHCNLELYPDYIRHLCKFIQEMESITTEK 276
Y+PLW+KG GH ++E+ P+Y+ HL KFI +E + K
Sbjct: 234 YEPLWLKGRGHSDIEMSPEYLPHLRKFISAIEKLPVPK 271
>UniRef100_O65548 Hypothetical protein F6I18.70 [Arabidopsis thaliana]
Length = 307
Score = 305 bits (782), Expect = 1e-81
Identities = 156/292 (53%), Positives = 203/292 (69%), Gaps = 33/292 (11%)
Query: 1 MGCMVSQLAAKFAFFPPSPPTYQLKKRDD-GKLTVVSAASPSVPIPHADDSSLDVLLVDT 59
MG + S +AAKFAFFPP P TY + K D+ GKL ++ D +++V + T
Sbjct: 1 MGNVTSNVAAKFAFFPPEPATYGVTKDDETGKLVFAGVSA---------DKNVEVHQLTT 51
Query: 60 KHGNKIVAFYLKNPYARLTLLYSHGNAADLGQLYDLFVQLKVNLRVNLMGYDYSGYGAST 119
K GNK+VA + ++P+AR TLLYSHGNAADLGQ+ +LF++L+ +LRVN+M Y T
Sbjct: 52 KSGNKVVATFWRHPFARFTLLYSHGNAADLGQMVELFIELRAHLRVNIMRYILK-----T 106
Query: 120 GKPSESSTYADIEAIYECLESEYGVGQEDIILYGQSVGSGPTLHLAAKLPRLRGVVLHSG 179
PSE +TY DIEA+Y CL S+YG+ QE+IILYGQSVGSGPTLH+A++L RLRGVVLHS
Sbjct: 107 LMPSEFNTYYDIEAVYSCLRSDYGIKQEEIILYGQSVGSGPTLHMASRLKRLRGVVLHSA 166
Query: 180 ILSGLRVLCHVKFSFCFDIYK------------------NINKIKKVKCPVLVIHGTEDD 221
ILSG+RVL VK + FDI+K NI+KI+ V VLVIHGT D+
Sbjct: 167 ILSGIRVLYPVKMTLWFDIFKVRKAHTKDLLLVGLHIYSNIDKIRHVNSQVLVIHGTNDE 226
Query: 222 VVNWLHGNGLWKMARESYDPLWIKGGGHCNLELYPDYIRHLCKFIQEMESIT 273
+V+ HG LW++A+E YDPLW+KGGGHCNLE YP+YI+HL KF+ ME ++
Sbjct: 227 IVDLSHGKRLWELAKEKYDPLWVKGGGHCNLETYPEYIKHLKKFVNAMEKLS 278
>UniRef100_Q9C619 Hypothetical protein T4O24.3 [Arabidopsis thaliana]
Length = 256
Score = 295 bits (754), Expect = 2e-78
Identities = 151/278 (54%), Positives = 197/278 (70%), Gaps = 27/278 (9%)
Query: 1 MGCMVSQLAAKFAFFPPSPPTYQLKKRDD--GKLTVVSAASPSVPIPHADDSSLDVLLVD 58
MG + S +AAKFAFFPP+PP+Y++ D G+L + IP DD +D+L +
Sbjct: 1 MGGVTSSIAAKFAFFPPTPPSYEVIADDSCGGRLYIPE-------IPRRDD--VDILKLR 51
Query: 59 TKHGNKIVAFYLKNPYARLTLLYSHGNAADLGQLYDLFVQLKVNLRVNLMGYDYSGYGAS 118
T+ GN+IVA Y+K+ A TLLYSHGNAADLGQ+++LFV+L LRVNLMGYDYSGYG S
Sbjct: 52 TRCGNEIVAVYVKHSKANGTLLYSHGNAADLGQMFELFVELSNRLRVNLMGYDYSGYGQS 111
Query: 119 TGKPSESSTYADIEAIYECLESEYGVGQEDIILYGQSVGSGPTLHLAAKLPRLRGVVLHS 178
TG+ SE +TYADIEA Y+CL+ +YGV + +I+YGQSVGSGPT+ LA++ P LRGVVL
Sbjct: 112 TGQASECNTYADIEASYKCLKEKYGVKDDQLIVYGQSVGSGPTVDLASRTPNLRGVVLQC 171
Query: 179 GILSGLRVLCHVKFSFCFDIYKNINKIKKVKCPVLVIHGTEDDVVNWLHGNGLWKMARES 238
ILSG+RVL VK ++ FDIYK GT D+VV+W HG LW++++E
Sbjct: 172 PILSGMRVLYPVKCTYWFDIYK----------------GTADEVVDWSHGKRLWELSKEK 215
Query: 239 YDPLWIKGGGHCNLELYPDYIRHLCKFIQEMESITTEK 276
Y+PLWI GGGHC+LELYPD+IRHL KF+ + + E+
Sbjct: 216 YEPLWISGGGHCDLELYPDFIRHLKKFVVSLGNKQAEQ 253
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.325 0.139 0.477
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 743,188,812
Number of Sequences: 2790947
Number of extensions: 33673700
Number of successful extensions: 205392
Number of sequences better than 10.0: 2995
Number of HSP's better than 10.0 without gapping: 817
Number of HSP's successfully gapped in prelim test: 2319
Number of HSP's that attempted gapping in prelim test: 148715
Number of HSP's gapped (non-prelim): 16342
length of query: 420
length of database: 848,049,833
effective HSP length: 130
effective length of query: 290
effective length of database: 485,226,723
effective search space: 140715749670
effective search space used: 140715749670
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 76 (33.9 bits)
Lotus: description of TM0230.1


