

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0228a.2
(510 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
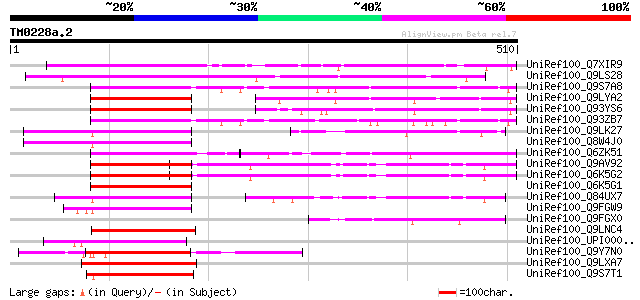
UniRef100_Q7XIR9 Putative RING3 protein [Oryza sativa] 286 1e-75
UniRef100_Q9LS28 Similarity to kinase [Arabidopsis thaliana] 241 3e-62
UniRef100_Q9S7A8 F28J7.10 protein [Arabidopsis thaliana] 224 3e-57
UniRef100_Q9LYA2 Kinase-like protein [Arabidopsis thaliana] 140 1e-31
UniRef100_Q93YS6 Putative kinase [Arabidopsis thaliana] 140 1e-31
UniRef100_Q93ZB7 AT3g01770/F28J7_10 [Arabidopsis thaliana] 135 3e-30
UniRef100_Q9LK27 Gb|AAF01563.1 [Arabidopsis thaliana] 132 3e-29
UniRef100_Q8W4J0 Hypothetical protein At3g27260; K17E12.8 [Arabi... 130 1e-28
UniRef100_Q6ZK51 Putative bromodomain-containing protein [Oryza ... 130 1e-28
UniRef100_Q9AV92 Kinase-like protein [Oryza sativa] 123 1e-26
UniRef100_Q6K5G2 Putative global transcription factor group E [O... 123 1e-26
UniRef100_Q6K5G1 Putative global transcription factor group E [O... 123 1e-26
UniRef100_Q84UX7 Global transcription factor group E [Zea mays] 122 2e-26
UniRef100_Q9FGW9 Similarity to kinase [Arabidopsis thaliana] 120 7e-26
UniRef100_Q9FGX0 Gb|AAC55944.1 [Arabidopsis thaliana] 120 9e-26
UniRef100_Q9LNC4 F9P14.9 protein [Arabidopsis thaliana] 116 1e-24
UniRef100_UPI000049A4F5 UPI000049A4F5 UniRef100 entry 111 6e-23
UniRef100_Q9Y7N0 Hypothetical bromodomain protein C1450.02 [Schi... 106 1e-21
UniRef100_Q9LXA7 Bromodomain protein-like [Arabidopsis thaliana] 103 9e-21
UniRef100_Q9S7T1 Hypothetical protein T18K17.19 [Arabidopsis tha... 103 9e-21
>UniRef100_Q7XIR9 Putative RING3 protein [Oryza sativa]
Length = 484
Score = 286 bits (732), Expect = 1e-75
Identities = 198/495 (40%), Positives = 279/495 (56%), Gaps = 54/495 (10%)
Query: 38 QTDENGRCNSAE--KLFKPDSIKRGPPEGFEG-QKEKRQKIDRKGSTQCAAILKCLMSYP 94
+++++G + E KL K D + G + K K ++ S QC +ILK LM +
Sbjct: 17 ESEDSGTDSEVEGSKLSKKDGVTSVYTCGHQPTSKNKVDPMNTSKSRQCGSILKKLMDHK 76
Query: 95 YSWVFNKPVDPVALNIPDYFAIIAKPMDLGTISSKLKKNVYSCVEEFAADVRLTFSNAMT 154
W+FN PVDPV IPDYF +I PMDLGT+ KL YS EFAADVRLTFSNAM
Sbjct: 77 SGWIFNTPVDPVVYGIPDYFDVIRNPMDLGTVKRKLTSKQYSNPYEFAADVRLTFSNAMK 136
Query: 155 YNPPHNDVHMMAKELNQIFDSKWKDMDKKWKCEDELGKSVTGTIKETARKSCNVMHPRQK 214
YNPP NDVH +A +LN+IFDS+WK +++KWK + + + + + K+ + P K
Sbjct: 137 YNPPGNDVHGIADQLNKIFDSEWKLLERKWKDRNLVQEQPSLKVL----KAQPAVTP--K 190
Query: 215 DTFPKKSQVSEHKRVLKISSLAARDAKVEVPKLSQVHCKSIEKDLIKGSE-ERDRIAPGA 273
PK + V K + A +KV++ K ++GSE + P
Sbjct: 191 PVLPKGVTAGTNSAVSKTLATAL-SSKVKI------------KFSVRGSELTSSKDTPLQ 237
Query: 274 VAFKQKRQIKSTSPLERKSDPDSDGAVSSLDSEHVYPGSQHATPATDASSGEVWST---- 329
++ I + P + D + S + G++ P DAS+ + S+
Sbjct: 238 AVGRRDGTINQSLPCTK--DNAKTPRIQSSEDRSESTGNE-LRPCDDASTSPLASSRQEE 294
Query: 330 ---PDFPVQLSPKKALRAAMLKSRFADTILKAQQKTLLEHGDKGDPLKMQLEKERLERIQ 386
P+ P LSP KALRAAMLKSRFA TI+KAQQK LL+HG K DP K+QLEKERLE+ Q
Sbjct: 295 EYLPEEP--LSPSKALRAAMLKSRFAGTIVKAQQKALLDHGKKIDPAKLQLEKERLEKRQ 352
Query: 387 REERARIEAQIKTAEAAARARAEEELRQQREKEREAARAAIEEMKRTVEIEHNLEIVKEL 446
+EE+ARIEAQ+K AEAAA+ + +EE+R +RE+ER AAR A+ MK+TV+I+ N + +K+L
Sbjct: 353 QEEKARIEAQVKAAEAAAQLKLDEEMRMKREQERRAARLALHMMKKTVDID-NSDFLKDL 411
Query: 447 ETLSGCTLSYKALGGRNDYKVALETLDKLQFE----NPLEQLGLFIKDELADEDEEVING 502
E LS K K+ ++ +D + +PLE+LGLF+K +L +E E +
Sbjct: 412 ENLS------KKWELNPPGKLIVDFVDGIDLPPGLGSPLERLGLFMKKDLEEEVEHEMED 465
Query: 503 T--------VEEGEI 509
+ VEEGEI
Sbjct: 466 SVSPSTEIDVEEGEI 480
>UniRef100_Q9LS28 Similarity to kinase [Arabidopsis thaliana]
Length = 506
Score = 241 bits (616), Expect = 3e-62
Identities = 170/495 (34%), Positives = 259/495 (51%), Gaps = 42/495 (8%)
Query: 17 FSTKRIEVDPGQKCEFGQRGLQTDENGRCNSAEKL----------FKPDSIKRGPPEGF- 65
F R+ P K +FG +G +S++K+ S KRG P+
Sbjct: 8 FVLVRMVAIPNIKIKFGPQGSVRTFQTLSDSSKKIEHVVTEDLSQSSEKSKKRGGPKELD 67
Query: 66 EGQKEKRQKIDRKGSTQCAAILKCLMSYPYSWVFNKPVDPVALNIPDYFAIIAKPMDLGT 125
E Q +K+Q++D S+QC A+L+ LM + W+F +PVDPV + IPDYF +I KPMDLGT
Sbjct: 68 EVQPKKKQRLDCDWSSQCLALLRFLMEHRGGWLFKEPVDPVKMEIPDYFNVIQKPMDLGT 127
Query: 126 ISSKLKKNVYSCVEEFAADVRLTFSNAMTYNPPHNDVHMMAKELNQIFDSKWKDMDKKWK 185
+ SKL KNVYS +EFAADVRLTF+NAM YNP N+VH +AKE+N+IF+ +W+ + KK
Sbjct: 128 VKSKLLKNVYSNADEFAADVRLTFANAMHYNPLWNEVHTIAKEINEIFEVRWESLMKKKV 187
Query: 186 CEDELGKSVTGTIKETARKSCNVMHPRQKDTFPKKSQVSEHKRVLKISSLAARDAKVEVP 245
+ G ++ + C+ K + SL+++ KV+
Sbjct: 188 LRLSWNEVREGYKRQPVERDCSRRSSTGTSASSGVGLTKPAKENSEKGSLSSKPVKVQSK 247
Query: 246 K---------LSQVHCKSIEKDLIKGSEERDRIAPGAVAFKQKRQIKSTSPLERKS-DPD 295
K L+ C I +K V Q+K+ S + DP
Sbjct: 248 KNTPAVTPKALATCKCGRIICICLKSCSSFG----SDVCSLTDCQLKNISGAQASELDPQ 303
Query: 296 SDGAVSSLDSEHVYPGSQHATPATDASSGEVWSTPDFPV--QLSPKKALRAAMLKSRFAD 353
S+G+ +S + SQ P+ G T FP + P+KALRAA+LK+++A
Sbjct: 304 SNGSDTSKKERNGSLKSQLDKPSNSDLLGNELKTA-FPALPPVPPEKALRAAILKAQYAG 362
Query: 354 TILKAQQKTLLEHGDKGDPLKMQLEKERLERIQREERARIEAQIKTAEAAARARAEEELR 413
TI+KA+ + +L +K D +++Q+EKE++ER QREE+ARIEA+++ A+ A R RA++EL+
Sbjct: 363 TIIKAKHRIVLGQNNKADLIRIQIEKEQMERAQREEKARIEAEMRAAKVAERMRAQDELK 422
Query: 414 QQREKEREAARAAIEEMKRTVEIEHN--LEIVKELETLSGCTLSYKA--------LGGRN 463
Q+RE + R I +MK+ + E N ++ K+ + GC KA L +N
Sbjct: 423 QKRESQ----RLEIAKMKKGFDFERNNHSKLKKKFVKVCGCFSLTKARLLLEELGLVLKN 478
Query: 464 DYKVALETLDKLQFE 478
DY LE + +F+
Sbjct: 479 DYCPELEVIGSEKFD 493
>UniRef100_Q9S7A8 F28J7.10 protein [Arabidopsis thaliana]
Length = 601
Score = 224 bits (572), Expect = 3e-57
Identities = 174/483 (36%), Positives = 244/483 (50%), Gaps = 69/483 (14%)
Query: 82 QCAAILKCLMSYPYSWVFNKPVDPVALNIPDYFAIIAKPMDLGTISSKLKKNVYSCVEEF 141
QC ++LK LMS + W+FN PVD V LNIPDYF II PMDLGT+ SKL YS EF
Sbjct: 132 QCESLLKRLMSQQHCWLFNTPVDVVKLNIPDYFTIIKHPMDLGTVKSKLTSGTYSSPSEF 191
Query: 142 AADVRLTFSNAMTYNPPHNDVHMMAKELNQIFDSKWKDMDKKWKCEDELGKSVTGTIKET 201
+ADVRLTF NAMTYNP N+V+ A L++ F+ +WK ++KK KS +
Sbjct: 192 SADVRLTFRNAMTYNPSDNNVYRFADTLSKFFEVRWKTIEKK----SSGTKSEPSNLATL 247
Query: 202 ARKSCNVMHP----RQKDTFPKKSQVSEHKRVL----------KISSLAARDAKVEVPKL 247
A K + P R+ + + S + KRV+ + SL + V++
Sbjct: 248 AHKDIAIPEPVAKKRKMNAVKRNSLLEPAKRVMTDEDRVKLGRDLGSLT--EFPVQIINF 305
Query: 248 SQVHCKSIEKDLIKGSEERDRIAPGAVAFKQKRQIKSTSP--LERKSDPDSDGAV-SSLD 304
+ H E + ++ I ++ Q++ L DS+G S L+
Sbjct: 306 LRDHSSKEE----RSGDDEIEIDINDLSHDALFQLRDLFDEFLRENQKKDSNGEPWSELE 361
Query: 305 SEHV------YPGSQHATPAT-------------DASSGEV-----------WSTPDFPV 334
E V +P S +T T DAS G++ ++
Sbjct: 362 DEDVDIGNYEHPISHISTVRTEKADSVGGLNQMEDASRGKLSLIEGADGHQDGNSAPKEK 421
Query: 335 QLSPKKALRAAMLKSRFADTILKAQQKTLLEHGDKGDPLKMQLEKERLERIQREERARIE 394
+L P+K RAA+LK+RFAD ILKAQ+ T L +K DP +Q EKE LE +++E+AR++
Sbjct: 422 ELPPEKRYRAALLKNRFADIILKAQEIT-LNQNEKRDPETLQREKEELELQKKKEKARLQ 480
Query: 395 AQIKTAEAAARARAEEELRQQREKEREAARAAIEEMKRTVEIEHNLEIVKELETLSGCTL 454
A+ K AE A R +E +++ E EREAAR A+ EM+++VEI N +K+LE L T+
Sbjct: 481 AEAKEAEEARRKAEAQEAKRKLELEREAARQALLEMEKSVEINENTRFLKDLELLK--TV 538
Query: 455 SYKALGGRNDYKVALETLDKLQF--ENPLEQLGLFIKDELADEDEEVI------NGTVEE 506
+ L D + L F NPLEQLGLF+K E DEDE + VEE
Sbjct: 539 NTDQLRNLRDVGSESDGLAVFGFGGSNPLEQLGLFMKHE-EDEDESDMLAFPDPGNEVEE 597
Query: 507 GEI 509
GEI
Sbjct: 598 GEI 600
>UniRef100_Q9LYA2 Kinase-like protein [Arabidopsis thaliana]
Length = 703
Score = 140 bits (352), Expect = 1e-31
Identities = 65/102 (63%), Positives = 77/102 (74%)
Query: 82 QCAAILKCLMSYPYSWVFNKPVDPVALNIPDYFAIIAKPMDLGTISSKLKKNVYSCVEEF 141
QC A+LK LMS+ Y WVFN PVD V LNI DYF +I PMDLGT+ +KL YSC EF
Sbjct: 140 QCEALLKRLMSHQYGWVFNTPVDVVKLNILDYFNVIEHPMDLGTVKNKLTSGTYSCPSEF 199
Query: 142 AADVRLTFSNAMTYNPPHNDVHMMAKELNQIFDSKWKDMDKK 183
AADVRLTFSNAMTYNPP NDV++MA L + F+ +WK ++KK
Sbjct: 200 AADVRLTFSNAMTYNPPGNDVYVMADTLRKFFEVRWKTLEKK 241
Score = 107 bits (266), Expect = 1e-21
Identities = 92/303 (30%), Positives = 147/303 (48%), Gaps = 46/303 (15%)
Query: 248 SQVHCKSIEKDLIKGSEERDRIA-----PGAVAFKQKRQIKSTSPLERKSD-PDSDGAVS 301
S + +IEKDL+ G+ + A+ + S S + S P + +
Sbjct: 405 SSISPVTIEKDLVLGNSNDYPLGCTTDCSSFDAYNLGNSLGSVSGDPKMSSLPRASKGLG 464
Query: 302 SLDSEHVYPGSQHATPATDASSGEV---------------------WSTPDFPVQLSPKK 340
++D E + G+ A+P +S G + ++ QL P+K
Sbjct: 465 TIDLEPMLDGATSASPTRGSSVGGLDQLESASPEKISSVEADCQQDGNSAQNEKQLPPEK 524
Query: 341 ALRAAMLKSRFADTILKAQQKTLLEHGDKGDPLKMQLEKERLERIQREERARIEAQIKTA 400
+ RAA+LK+RFAD ILKA++K L D DP K+Q E+E LE +++E+AR++A+ K A
Sbjct: 525 SYRAAILKNRFADIILKAREKP-LNQNDTRDPEKLQREREELELQKKKEKARLQAEAKAA 583
Query: 401 EAAAR-------ARAEEELRQQREKEREAARAAIEEMKRTVEIEHNLEIVKELETLSGCT 453
E A R A A E +++ E EREAAR A+ E +VE+ N + +++LE L
Sbjct: 584 EEARRKAEAQAAAEAAAEAKRKLELEREAARQALME---SVELNENAKFLEDLELLKTVD 640
Query: 454 LSY--KALGGRNDYKVALETLDKLQFENPLEQLGLFIKDELADEDEEVING-----TVEE 506
+ + + V L + NPLEQLGLF+K + +E+ + + +EE
Sbjct: 641 TDHLTNTIEEEDGPDVGLRSF-SFGGSNPLEQLGLFMKQDEDEEEADPLTSPAPEIDIEE 699
Query: 507 GEI 509
GEI
Sbjct: 700 GEI 702
>UniRef100_Q93YS6 Putative kinase [Arabidopsis thaliana]
Length = 688
Score = 140 bits (352), Expect = 1e-31
Identities = 65/102 (63%), Positives = 77/102 (74%)
Query: 82 QCAAILKCLMSYPYSWVFNKPVDPVALNIPDYFAIIAKPMDLGTISSKLKKNVYSCVEEF 141
QC A+LK LMS+ Y WVFN PVD V LNI DYF +I PMDLGT+ +KL YSC EF
Sbjct: 140 QCEALLKRLMSHQYGWVFNTPVDVVKLNILDYFNVIEHPMDLGTVKNKLTSGTYSCPSEF 199
Query: 142 AADVRLTFSNAMTYNPPHNDVHMMAKELNQIFDSKWKDMDKK 183
AADVRLTFSNAMTYNPP NDV++MA L + F+ +WK ++KK
Sbjct: 200 AADVRLTFSNAMTYNPPGNDVYVMADTLRKFFEVRWKTLEKK 241
Score = 119 bits (297), Expect = 3e-25
Identities = 96/290 (33%), Positives = 155/290 (53%), Gaps = 35/290 (12%)
Query: 248 SQVHCKSIEKDLIKGSEERDRIAPGAVAFKQKRQIKSTSPLERKS------DPDSDGAVS 301
S + +IEKDL+ G+ + + G+V+ K S+ P K +P DGA S
Sbjct: 405 SSISPVTIEKDLVLGNSNGNSL--GSVSGDPKM---SSLPRASKGLGTIDLEPMLDGATS 459
Query: 302 SLDSEHVYPGS----QHATP----ATDASSGEVWSTPDFPVQLSPKKALRAAMLKSRFAD 353
+ + G + A+P + +A + ++ QL P+K+ RAA+LK+RFAD
Sbjct: 460 ASPTRGSSVGGLDQLESASPEKISSVEADCQQDGNSAQNEKQLPPEKSYRAAILKNRFAD 519
Query: 354 TILKAQQKTLLEHGDKGDPLKMQLEKERLERIQREERARIEAQIKTAEAAAR-------A 406
ILKA++K L ++ D DP K+Q E+E LE +++E+AR++A+ K AE A R A
Sbjct: 520 IILKAREKPLNQN-DTRDPEKLQREREELELQKKKEKARLQAEAKAAEEARRKAEAQAAA 578
Query: 407 RAEEELRQQREKEREAARAAIEEMKRTVEIEHNLEIVKELETLSGCTLSY--KALGGRND 464
A E +++ E EREAAR A+ EM+++VE+ N + +++LE L + + +
Sbjct: 579 EAAAEAKRKLELEREAARQALMEMEQSVELNENAKFLEDLELLKTVDTDHLTNTIEEEDG 638
Query: 465 YKVALETLDKLQFENPLEQLGLFIKDELADEDEEVING-----TVEEGEI 509
V L + NPLEQLGLF+K + +E+ + + +EEGEI
Sbjct: 639 PDVGLRSF-SFGGSNPLEQLGLFMKQDEDEEEADPLTSPAPEIDIEEGEI 687
>UniRef100_Q93ZB7 AT3g01770/F28J7_10 [Arabidopsis thaliana]
Length = 620
Score = 135 bits (340), Expect = 3e-30
Identities = 143/505 (28%), Positives = 209/505 (41%), Gaps = 94/505 (18%)
Query: 82 QCAAILKCLMSYPYSWVFNKPVDPVALNIPDYFAIIAKPMDLGTISSKLKKNVYSCVEEF 141
QC ++LK LMS + W+FN PVD V LNIPDYF II PMDLGT+ SKL YS EF
Sbjct: 132 QCESLLKRLMSQQHCWLFNTPVDVVKLNIPDYFTIIKHPMDLGTVKSKLTSGTYSSPSEF 191
Query: 142 AADVRLTFSNAMTYNPPHNDVHMMAKELNQIFDSKWKDMDKKWKCEDELGKSVTGTIKET 201
+ADVRLTF NAMTYNP N+V+ A L++ F+ +WK ++KK KS +
Sbjct: 192 SADVRLTFRNAMTYNPSDNNVYRFADTLSKFFEVRWKTIEKK----SSGTKSEPSNLATL 247
Query: 202 ARKSCNVMHP----RQKDTFPKKSQVSEHKRVL----------KISSLAARDAKVEVPKL 247
A K + P R+ + + S + KRV+ + SL + V++
Sbjct: 248 AHKDIAIPEPVAKKRKMNAVKRNSLLEPAKRVMTDEDRVKLGRDLGSLT--EFPVQIINF 305
Query: 248 SQVHCKSIEKDLIKGSEERDRIAPGAVAFKQKRQIKSTSP--LERKSDPDSDGAVSSLDS 305
+ H E + ++ I ++ Q++ L DS+G L+
Sbjct: 306 LRDHSSKEE----RSGDDEIEIDINDLSHDALFQLRDLFDEFLRENQKKDSNGEPCVLEL 361
Query: 306 EHVYPGSQHATPATDASSGEVWSTPDFPVQLSPKKALRAAMLKSRFADTILKAQQK---- 361
H GS T G D + + +++ D++ Q
Sbjct: 362 LH---GSGPGNSLTQHCDGSELEDEDVDIGNYEHPISHISTVRTE-KDSVGGLNQMEDAS 417
Query: 362 ----TLLEHGD-----KGDPLKMQLEKERLERIQREERARIEAQIKTAEAAA---RARAE 409
+L+E D P + +L E+ R + + +K E R
Sbjct: 418 RGKLSLIEGADGHQDGNSAPKEKELPPEKRYRAALLKNRFADIILKAQEITLNQNEKRDP 477
Query: 410 EELRQQRE-----KEREAAR--------------AAIEEMKRTVEIE------------- 437
E L++++E K++E AR A +E KR +E+E
Sbjct: 478 ETLQREKEELELQKKKEKARLQAEAKEAEEARRKAEAQEAKRKLELEREAARQALLEMEK 537
Query: 438 -----HNLEIVKELETLSGCTLSYKALGGRNDYKVALETLDKLQF--ENPLEQLGLFIKD 490
N +K+LE L T++ L D + L F NPLEQLGLF+K
Sbjct: 538 SVEINENTRFLKDLELLK--TVNTDQLRNLRDVGSESDGLAVFGFGGSNPLEQLGLFMKH 595
Query: 491 ELADEDEEVI------NGTVEEGEI 509
E DEDE + VEEGEI
Sbjct: 596 E-EDEDESDMLAFPDPGNEVEEGEI 619
>UniRef100_Q9LK27 Gb|AAF01563.1 [Arabidopsis thaliana]
Length = 818
Score = 132 bits (331), Expect = 3e-29
Identities = 70/174 (40%), Positives = 97/174 (55%), Gaps = 5/174 (2%)
Query: 15 IKFSTKRIEVDPGQKCEFGQRGLQTDENGRCNSAEKLFKPDSIKRGPPEGFEGQKEKRQK 74
+ ++ R+ GQK + + S +K+ + RG G G+ E ++
Sbjct: 107 VSSTSDRVGFSTGQKISSRVSNSKKPSDFAVGSGKKVRHQNGTSRGWNRGTSGKFESSKE 166
Query: 75 IDRKGST-----QCAAILKCLMSYPYSWVFNKPVDPVALNIPDYFAIIAKPMDLGTISSK 129
QC +L+ L S+P+SWVF PVD V LNIPDY I PMDLGT+
Sbjct: 167 TMTSTPNITLMKQCDTLLRKLWSHPHSWVFQAPVDVVKLNIPDYLTTIKHPMDLGTVKKN 226
Query: 130 LKKNVYSCVEEFAADVRLTFSNAMTYNPPHNDVHMMAKELNQIFDSKWKDMDKK 183
L VYS EFAADVRLTF+NAMTYNPP +DVH+M L+++F+++WK + KK
Sbjct: 227 LASGVYSSPHEFAADVRLTFTNAMTYNPPGHDVHIMGDILSKLFEARWKTIKKK 280
Score = 112 bits (281), Expect = 2e-23
Identities = 85/228 (37%), Positives = 129/228 (56%), Gaps = 36/228 (15%)
Query: 283 KSTSPLERKSDPDSDGAVSSLDSEHVYPGSQHATPATDASSGEVWSTPDFPVQLSPKKAL 342
+ST LE+ D S +SS +S+ + G+ TPA S +K
Sbjct: 500 QSTGALEQM-DICSQQKLSSDESDGQHEGNILETPA------------------SSEKRY 540
Query: 343 RAAMLKSRFADTILKAQQKTLLEHGDKGDPLKMQLEKERLERIQREERARIEAQI----- 397
RAA+LK+RFAD ILKA++K L ++G KGDP +++ E+E L +++E+AR++A+
Sbjct: 541 RAALLKNRFADIILKAREKPLPQNGIKGDPERLRKEREELVLQKKKEKARLQAEAEAAED 600
Query: 398 --KTAEAAARARAEEELRQQREKEREAARAAIEEMKRTVEIEHNLEIVKELETLSGCTLS 455
+ AEA A A A E +++RE EREAAR A+ +M++TVEI N +++LE LS + +
Sbjct: 601 ARRQAEAEAAAEAAAEAKRKRELEREAARQALLKMEKTVEINENSRFLEDLEMLS--SSA 658
Query: 456 YKALGGRNDYKVALETLD-----KLQFENPLEQLGLFIKDELADEDEE 498
+ L + LD L+ NPLEQLGL++K D+DEE
Sbjct: 659 PEQLPSSAEETSPERPLDALGSFNLRGSNPLEQLGLYMKQ---DDDEE 703
>UniRef100_Q8W4J0 Hypothetical protein At3g27260; K17E12.8 [Arabidopsis thaliana]
Length = 503
Score = 130 bits (326), Expect = 1e-28
Identities = 69/174 (39%), Positives = 97/174 (55%), Gaps = 5/174 (2%)
Query: 15 IKFSTKRIEVDPGQKCEFGQRGLQTDENGRCNSAEKLFKPDSIKRGPPEGFEGQKEKRQK 74
+ ++ R+ GQK + + S +K+ + RG G G+ E ++
Sbjct: 107 VSSTSDRVGFSTGQKISSRVSNSKKPSDFAVGSGKKVRHQNGTSRGWNRGTSGKFESSKE 166
Query: 75 IDRKGST-----QCAAILKCLMSYPYSWVFNKPVDPVALNIPDYFAIIAKPMDLGTISSK 129
QC +L+ L S+P+SWVF PVD V LNIPDY I PMDLGT+
Sbjct: 167 TMTSTPNITLMKQCDTLLRKLWSHPHSWVFQAPVDVVKLNIPDYLTTIKHPMDLGTVKKN 226
Query: 130 LKKNVYSCVEEFAADVRLTFSNAMTYNPPHNDVHMMAKELNQIFDSKWKDMDKK 183
L VYS EFAADVRLTF++AMTYNPP +DVH+M L+++F+++WK + KK
Sbjct: 227 LASGVYSSPHEFAADVRLTFTDAMTYNPPGHDVHIMGDILSKLFEARWKTIKKK 280
>UniRef100_Q6ZK51 Putative bromodomain-containing protein [Oryza sativa]
Length = 791
Score = 130 bits (326), Expect = 1e-28
Identities = 109/294 (37%), Positives = 147/294 (49%), Gaps = 32/294 (10%)
Query: 233 SSLAARDAKVEVPKLSQVHCKSIEKDL----IKGSEERDRIAPGAVAFKQKRQIKSTSPL 288
S A R +K S +S D GS+ + P K Q S L
Sbjct: 512 SETAERSSKHSTSSSSSSDSESSSSDSDSSSSSGSDLDVNVPPSTSGAKDNTQ--SAVRL 569
Query: 289 ERKSDPDSDGAVSSLDSEHVYPGSQHATPATDASSGEVWSTPDFPVQLSPKKALRAAMLK 348
++++DP L S ++ S P + GE S P P K RAA+L
Sbjct: 570 DQENDP--------LSSTNLPQQSSDPVPISAEDEGENVSEKQVP----PAKQYRAAVLL 617
Query: 349 SRFADTILKAQQKTLLEHGDKGDPLKMQLEKERLERIQREERARIEAQIKTAE------- 401
+RFADTI KA++KTL + K DP K+Q + E LER++REERARI+A+ K AE
Sbjct: 618 NRFADTIFKAREKTL-DQVAKKDPEKLQHDMEELERLRREERARIQAEAKAAEDARKRAE 676
Query: 402 AAARARAEEELRQQREKEREAARAAIEEMKRTVEIEHNLEIVKELETLSGCTL--SYKAL 459
AAA A A E ++QRE+EREAAR A+++M++TV+I +K+LE L T + +
Sbjct: 677 AAAAAEAAAEAKRQREREREAARKALQQMEKTVDINEGNLFLKDLEMLGTVTSGEQFPSS 736
Query: 460 GGRNDYKVALETLDKLQFENPLEQLGLFIK--DELADEDEEVINGT--VEEGEI 509
G E L NPLEQLGL++K DE +E E T VEEGEI
Sbjct: 737 VGETSPTHTPEGLGFQLGSNPLEQLGLYMKNDDEEDEEGESADEPTIDVEEGEI 790
Score = 118 bits (296), Expect = 4e-25
Identities = 69/150 (46%), Positives = 89/150 (59%), Gaps = 10/150 (6%)
Query: 82 QCAAILKCLMSYPYSWVFNKPVDPVALNIPDYFAIIAKPMDLGTISSKLKKNVYSCVEEF 141
QC +LK L + ++ F PVD V LNIPDYF II KPMDLGTI KL +YS +F
Sbjct: 165 QCGNLLKNLFKHQWAGPFLAPVDVVQLNIPDYFDIIKKPMDLGTIEKKLNAGMYSTPWDF 224
Query: 142 AADVRLTFSNAMTYNPPHNDVHMMAKELNQIFDSKWKDMDKKWKCEDELGKSVTGTIKET 201
AADVRLTF NA+TYNP NDV++M K L IF+++WK ++KK D+ K +
Sbjct: 225 AADVRLTFDNAVTYNPVGNDVNLMGKTLKCIFETRWKFIEKKLPSLDD---------KFS 275
Query: 202 ARKSCNVMHPRQKDTFPKKSQVSEHKRVLK 231
R+ + +KDT +K SE K K
Sbjct: 276 VRREPSQKGAVKKDTI-EKDYPSEKKHSTK 304
>UniRef100_Q9AV92 Kinase-like protein [Oryza sativa]
Length = 714
Score = 123 bits (309), Expect = 1e-26
Identities = 62/102 (60%), Positives = 76/102 (73%)
Query: 82 QCAAILKCLMSYPYSWVFNKPVDPVALNIPDYFAIIAKPMDLGTISSKLKKNVYSCVEEF 141
QC AILK LM+ S +F+ PVD V LNIPDYF II KPMDLGTI +KL Y+ EF
Sbjct: 170 QCDAILKKLMTQKCSNIFDSPVDAVKLNIPDYFQIIKKPMDLGTIRNKLDSGSYTSPSEF 229
Query: 142 AADVRLTFSNAMTYNPPHNDVHMMAKELNQIFDSKWKDMDKK 183
AADVRLTFSNAMTYNP + VH A +LN++F+S+W+ ++KK
Sbjct: 230 AADVRLTFSNAMTYNPRGHVVHDYAIQLNKMFESRWRTIEKK 271
Score = 114 bits (286), Expect = 5e-24
Identities = 122/369 (33%), Positives = 176/369 (47%), Gaps = 45/369 (12%)
Query: 161 DVHMMAKELNQIFDSKWKDMDKKWKCEDELGKSVTGTIKETARKSCNVMHPRQKDTFPKK 220
D+H ++ +L +F+ K K +DK + E +S + + ++ NV T P K
Sbjct: 370 DIHAVSDDL--LFELK-KHVDKYLQ---EREQSQQAKSEPSENEAANVSGLSHSSTNPCK 423
Query: 221 S--QVSEHKRVLKISS--LAARDA-----KVEVPKLSQVHCKSIEKDLIKGSEERDRIAP 271
V E + +S L +DA K P S S D GS+
Sbjct: 424 GGDPVEEDVDICGNASPILIEKDAHNNPNKCGSPSSSSSDSGSSSSDSESGSDSESEQEK 483
Query: 272 GAVAFKQKRQIKSTSPLER-KSDPDSD-GAVSSLDSEHVYPGSQHATPATDASSGEVWST 329
G K K +S +E+ KSD S A+ D + + PA + + S
Sbjct: 484 GGSPGKPKGSKRSEQLVEQEKSDVISPVDAIRPADDVELREQDNESKPAPEGEN----SK 539
Query: 330 PDFPVQLSPKKALRAAMLKSRFADTILKAQQKTLLEHGDKGDPLKMQLEKERLERIQREE 389
PD Q+SP K LR A L+SR+AD I+KAQ L + GDK +E LE++Q+EE
Sbjct: 540 PDR--QVSPDKLLRTAFLRSRYADVIVKAQG-ILSQGGDK---------QEELEKLQKEE 587
Query: 390 RARIEAQIKTAEAAARARAEEELRQQREKEREAARAAIEEMKRTVEIEHNLEIVKELETL 449
+AR+ A+ A A RA AE E +++R+ ERE AR A++EM+RTVEI NL + K+LE L
Sbjct: 588 KARLLAEGNAAMEARRAEAEAEAKRKRDLEREKARQALQEMERTVEINDNLHL-KDLEML 646
Query: 450 SGCTLSYKALGGRNDYKVALETLDKLQ-----FENPLEQLGLFIK--DELADEDEEVING 502
T + + D + D + NPLEQLGLF+K +E +ED +
Sbjct: 647 GTATTEH--IVSSVDETSPEHSQDGMPSFLPGSGNPLEQLGLFMKADEEEEEEDPSCVPS 704
Query: 503 T--VEEGEI 509
T EEGEI
Sbjct: 705 TKDAEEGEI 713
>UniRef100_Q6K5G2 Putative global transcription factor group E [Oryza sativa]
Length = 714
Score = 123 bits (309), Expect = 1e-26
Identities = 62/102 (60%), Positives = 76/102 (73%)
Query: 82 QCAAILKCLMSYPYSWVFNKPVDPVALNIPDYFAIIAKPMDLGTISSKLKKNVYSCVEEF 141
QC AILK LM+ S +F+ PVD V LNIPDYF II KPMDLGTI +KL Y+ EF
Sbjct: 170 QCDAILKKLMTQKCSNIFDSPVDAVKLNIPDYFQIIKKPMDLGTIRNKLDSGSYTSPSEF 229
Query: 142 AADVRLTFSNAMTYNPPHNDVHMMAKELNQIFDSKWKDMDKK 183
AADVRLTFSNAMTYNP + VH A +LN++F+S+W+ ++KK
Sbjct: 230 AADVRLTFSNAMTYNPRGHVVHDYAIQLNKMFESRWRTIEKK 271
Score = 116 bits (291), Expect = 1e-24
Identities = 123/369 (33%), Positives = 178/369 (47%), Gaps = 45/369 (12%)
Query: 161 DVHMMAKELNQIFDSKWKDMDKKWKCEDELGKSVTGTIKETARKSCNVMHPRQKDTFPKK 220
D+H ++ +L +F+ K K +DK + E +S + + ++ NV T P K
Sbjct: 370 DIHAVSDDL--LFELK-KHVDKYLQ---EREQSQQAKSEPSENEAANVSGLSHSSTNPCK 423
Query: 221 S--QVSEHKRVLKISS--LAARDA-----KVEVPKLSQVHCKSIEKDLIKGSEERDRIAP 271
V E + +S L +DA K P S S D GS+
Sbjct: 424 GGDPVEEDVDICGNASPILIEKDAHNNPNKCGSPSSSSSDSGSSSSDSESGSDSESEQEK 483
Query: 272 GAVAFKQKRQIKSTSPLER-KSDPDSD-GAVSSLDSEHVYPGSQHATPATDASSGEVWST 329
G K K +S +E+ KSD S A+ D + + PA + + S
Sbjct: 484 GGSPGKPKGSKRSEQLVEQEKSDVISPVDAIRPADDVELREQDNESKPAPEGEN----SK 539
Query: 330 PDFPVQLSPKKALRAAMLKSRFADTILKAQQKTLLEHGDKGDPLKMQLEKERLERIQREE 389
PD Q+SP K LRAA+L+SR+AD I+KAQ L + GDK +E LE++Q+EE
Sbjct: 540 PDR--QVSPDKLLRAALLRSRYADVIVKAQG-ILSQGGDK---------QEELEKLQKEE 587
Query: 390 RARIEAQIKTAEAAARARAEEELRQQREKEREAARAAIEEMKRTVEIEHNLEIVKELETL 449
+AR+ A+ A A RA AE E +++R+ ERE AR A++EM+RTVEI NL + K+LE L
Sbjct: 588 KARLLAEGNAAMEARRAEAEAEAKRKRDLEREKARQALQEMERTVEINDNLHL-KDLEML 646
Query: 450 SGCTLSYKALGGRNDYKVALETLDKLQ-----FENPLEQLGLFIK--DELADEDEEVING 502
T + + D + D + NPLEQLGLF+K +E +ED +
Sbjct: 647 GTATTEH--IVSSVDETSPEHSQDGMPSFLPGSGNPLEQLGLFMKADEEEEEEDPSSVPS 704
Query: 503 T--VEEGEI 509
T EEGEI
Sbjct: 705 TKDAEEGEI 713
>UniRef100_Q6K5G1 Putative global transcription factor group E [Oryza sativa]
Length = 480
Score = 123 bits (309), Expect = 1e-26
Identities = 62/102 (60%), Positives = 76/102 (73%)
Query: 82 QCAAILKCLMSYPYSWVFNKPVDPVALNIPDYFAIIAKPMDLGTISSKLKKNVYSCVEEF 141
QC AILK LM+ S +F+ PVD V LNIPDYF II KPMDLGTI +KL Y+ EF
Sbjct: 170 QCDAILKKLMTQKCSNIFDSPVDAVKLNIPDYFQIIKKPMDLGTIRNKLDSGSYTSPSEF 229
Query: 142 AADVRLTFSNAMTYNPPHNDVHMMAKELNQIFDSKWKDMDKK 183
AADVRLTFSNAMTYNP + VH A +LN++F+S+W+ ++KK
Sbjct: 230 AADVRLTFSNAMTYNPRGHVVHDYAIQLNKMFESRWRTIEKK 271
>UniRef100_Q84UX7 Global transcription factor group E [Zea mays]
Length = 696
Score = 122 bits (307), Expect = 2e-26
Identities = 66/149 (44%), Positives = 89/149 (59%), Gaps = 11/149 (7%)
Query: 46 NSAEKLFKPDSIKRGPPEGFEGQKEKRQKIDRKGST-----------QCAAILKCLMSYP 94
++A + KP +RG G K + + T QC AILK LM+
Sbjct: 124 SAAPRAKKPQKSQRGGTNVIRGAKGRFLPTKPRPETSTVLSEAAAFKQCEAILKKLMTQK 183
Query: 95 YSWVFNKPVDPVALNIPDYFAIIAKPMDLGTISSKLKKNVYSCVEEFAADVRLTFSNAMT 154
YS +FN PVD V L IPDYF I+ PMDLGT+ KL+ Y+ +FAADVRLTF+NAM
Sbjct: 184 YSHIFNVPVDIVKLQIPDYFDIVKTPMDLGTVKKKLESGSYTSPSDFAADVRLTFNNAMA 243
Query: 155 YNPPHNDVHMMAKELNQIFDSKWKDMDKK 183
YNP + VH MA +LN++F+S+W+ ++KK
Sbjct: 244 YNPRGHAVHDMAIQLNKMFESRWRPIEKK 272
Score = 106 bits (265), Expect = 1e-21
Identities = 93/274 (33%), Positives = 141/274 (50%), Gaps = 37/274 (13%)
Query: 238 RDAKVEVPKLSQVHCKSIEKDLIKGSE---ERDRI-APGAVAFKQKRQIK----STSPLE 289
R++K P S +S D GS+ E +++ +PG +A K+ + S +
Sbjct: 443 RNSKCGSPSSSSSDSESSSSDSDSGSDSESESEKVGSPGKLAKGTKKPDQLVEQERSDVI 502
Query: 290 RKSDPDSDGAVSSLDSEHVYPGSQHATPATDASSGEVWSTPDFPVQLSPKKALRAAMLKS 349
+D + A+ L E + PA + E PD Q+SP + LRAA+L+S
Sbjct: 503 SPADANRPAAIVGLHGE-----DSESKPAPGGENSE----PD--TQVSPDRLLRAALLRS 551
Query: 350 RFADTILKAQQKTLLEHGDKGDPLKMQLEKERLERIQREERARIEAQIKTAEAAARARAE 409
R+AD I+KA+ L + GDK +E LE++Q+EE+ R+ A+ A A RA AE
Sbjct: 552 RYADVIVKARG-ILSQGGDK---------QEELEKLQKEEKERLLAEGTAAMEACRAEAE 601
Query: 410 EELRQQREKEREAARAAIEEMKRTVEIEHNLEIVKELETLSGCTLSYKALGGRNDYKVAL 469
E +++R ERE AR A++EM+RTVEI NL + K+LE L T + + D
Sbjct: 602 AEKKRKRNFEREKARQALQEMERTVEINDNLHL-KDLELLGTATTEH--IVSSVDETSPE 658
Query: 470 ETLDKLQ-----FENPLEQLGLFIKDELADEDEE 498
+ D + NPLEQLGLFIK + ++DE+
Sbjct: 659 RSHDGMAGYHPGAVNPLEQLGLFIKVDDEEDDEQ 692
>UniRef100_Q9FGW9 Similarity to kinase [Arabidopsis thaliana]
Length = 477
Score = 120 bits (302), Expect = 7e-26
Identities = 67/154 (43%), Positives = 88/154 (56%), Gaps = 25/154 (16%)
Query: 55 DSIKRGPPEGFE----GQKEKRQKI------DRKGST---------------QCAAILKC 89
D +R PPE F Q +KR + ++KG + +C +L
Sbjct: 112 DGPRRPPPENFATFVGSQGKKRPPVRSDKQRNKKGPSRLNVPTSYTVASVMKECETLLNR 171
Query: 90 LMSYPYSWVFNKPVDPVALNIPDYFAIIAKPMDLGTISSKLKKNVYSCVEEFAADVRLTF 149
L S+ W F PVDPV LNIPDYF +I PMDLGTI S+L K YS +FAADVRLTF
Sbjct: 172 LWSHKSGWPFRTPVDPVMLNIPDYFNVIKHPMDLGTIRSRLCKGEYSSPLDFAADVRLTF 231
Query: 150 SNAMTYNPPHNDVHMMAKELNQIFDSKWKDMDKK 183
SN++ YNPP N H MA+ +++ F+S WK ++KK
Sbjct: 232 SNSIAYNPPGNQFHTMAQGISKYFESGWKSIEKK 265
>UniRef100_Q9FGX0 Gb|AAC55944.1 [Arabidopsis thaliana]
Length = 569
Score = 120 bits (301), Expect = 9e-26
Identities = 78/210 (37%), Positives = 125/210 (59%), Gaps = 25/210 (11%)
Query: 301 SSLDSEHVYPGSQHATPATDASSGEVWSTPDFPVQLSPKKALRAAMLKSRFADTILKAQQ 360
+++D+ + P + A P S PD SP K RAA LK+RFADTI+KA++
Sbjct: 38 TTMDAVVLVPDEETAPPERQIS-------PD-----SPDKRYRAAFLKNRFADTIMKARE 85
Query: 361 KTLLEHGDKGDPLKMQLEKERLERIQREERARIEAQIKTAEAA---ARARAEEELRQQRE 417
K + G+KGDP K+++E+E E+ REE+ R++A+ K AE A A+A A E+ R++RE
Sbjct: 86 KAFTK-GEKGDPEKLRIEREEFEKRLREEKERLQAEAKAAEEARRKAKAEAAEKARRERE 144
Query: 418 KEREAARAAIEEMKRTVEIEHNLEIVKELETLSG-------CTLSYKALGGR-NDYKVAL 469
+EREAAR A+++M++TVEI + +++L+ L S + + + ++ + L
Sbjct: 145 QEREAARQALQKMEKTVEINEGIRFMEDLQMLRATGTEGDQLPTSMEVMSPKFSEDMLGL 204
Query: 470 ETLDKLQFENPLEQLGLFIK-DELADEDEE 498
+ NPLE LGL++K DE DE+E+
Sbjct: 205 GSFKMESNSNPLEHLGLYMKMDEDEDEEED 234
>UniRef100_Q9LNC4 F9P14.9 protein [Arabidopsis thaliana]
Length = 766
Score = 116 bits (291), Expect = 1e-24
Identities = 56/105 (53%), Positives = 70/105 (66%)
Query: 83 CAAILKCLMSYPYSWVFNKPVDPVALNIPDYFAIIAKPMDLGTISSKLKKNVYSCVEEFA 142
C+A+L+ LM + + WVFN PVD L + DY+ II PMDLGTI S L KN+Y EFA
Sbjct: 425 CSALLERLMKHKHGWVFNAPVDVKGLGLLDYYTIIEHPMDLGTIKSALMKNLYKSPREFA 484
Query: 143 ADVRLTFSNAMTYNPPHNDVHMMAKELNQIFDSKWKDMDKKWKCE 187
DVRLTF NAMTYNP DVH+MA L QIF+ +W ++ + E
Sbjct: 485 EDVRLTFHNAMTYNPEGQDVHLMAVTLLQIFEERWAVIEADYNRE 529
>UniRef100_UPI000049A4F5 UPI000049A4F5 UniRef100 entry
Length = 265
Score = 111 bits (277), Expect = 6e-23
Identities = 60/155 (38%), Positives = 93/155 (59%), Gaps = 11/155 (7%)
Query: 35 RGLQTDENGRCNSAEKLFKPDSIKRGPPE--------GFEGQKE---KRQKIDRKGSTQC 83
+G+ +E + E+ K I+ G G+E +++ K + ++ C
Sbjct: 10 KGVSKEEEKKGQKKEREKKESKIEEGSSHRPMTRSMSGYEIEEKPVIKNEPLNSFEKELC 69
Query: 84 AAILKCLMSYPYSWVFNKPVDPVALNIPDYFAIIAKPMDLGTISSKLKKNVYSCVEEFAA 143
+++K LM S VF +PVDP NIP+YF II PMDLGT+ K+KKN+Y ++EF+
Sbjct: 70 MSVMKQLMKVSESEVFMEPVDPEIWNIPNYFDIIKTPMDLGTVIKKIKKNMYYSIDEFSN 129
Query: 144 DVRLTFSNAMTYNPPHNDVHMMAKELNQIFDSKWK 178
DVRLTF+NAMT+NPP N VH A++L +IF++ ++
Sbjct: 130 DVRLTFTNAMTFNPPGNYVHSYAEKLYKIFENYYR 164
>UniRef100_Q9Y7N0 Hypothetical bromodomain protein C1450.02 [Schizosaccharomyces
pombe]
Length = 578
Score = 106 bits (265), Expect = 1e-21
Identities = 53/112 (47%), Positives = 72/112 (63%), Gaps = 6/112 (5%)
Query: 77 RKGSTQ---CAAILKCLMSYPY---SWVFNKPVDPVALNIPDYFAIIAKPMDLGTISSKL 130
RK ++Q C+ +LK L Y ++ F +PVDPVA + PDYF +I +PMDL TI SKL
Sbjct: 251 RKNNSQMRFCSTVLKELYKRQYESFAFPFYQPVDPVACDCPDYFDVIKEPMDLSTIQSKL 310
Query: 131 KKNVYSCVEEFAADVRLTFSNAMTYNPPHNDVHMMAKELNQIFDSKWKDMDK 182
KN YS +EEF +D+ L F+N TYNPP VH+M ++L +F KW+ K
Sbjct: 311 NKNEYSTLEEFESDILLMFNNCFTYNPPGTPVHVMGRQLENVFKEKWEARPK 362
Score = 85.9 bits (211), Expect = 3e-15
Identities = 83/303 (27%), Positives = 124/303 (40%), Gaps = 40/303 (13%)
Query: 10 TRKLKIKFSTKRIEVDPGQKCEFGQRGLQTDENGRCNSAEKLF-KPDSIKRGPPEGFEGQ 68
+R+ ++K TK + G G +Q+ + + E L K DS G G Q
Sbjct: 5 SRENEVKAETKDEIANDGSPQLNGDNNIQSSDGHNDENEESLSRKRDS--SGATVGDLKQ 62
Query: 69 KEKR-----------QKIDRKG-----STQCAAILKCLMSYPYSWVFNKPVDPVALNIPD 112
+EK +KI G C AI++ L S F PVDP+ NIPD
Sbjct: 63 EEKESMPKKEPEPTVKKIRGSGMPPPQQKYCLAIVRQLKRTKNSAPFKVPVDPIKQNIPD 122
Query: 113 YFAIIAKPMDLGTISSKLKKNVYSCVEEFAADVRLTFSNAMTYNPPHNDVHMMAKELNQI 172
Y I+ PMDLGTI KL YS +EF D+ L FSN YN + V M K L ++
Sbjct: 123 YPTIVKNPMDLGTIEKKLTSYEYSVPQEFIDDMNLMFSNCFLYNGTESPVGSMGKALQEV 182
Query: 173 FDSKWKDMDKKWKCEDELGKSVTGTIKETARKSCNVMHPRQKDTFPKKSQVSEHKRVLKI 232
F+ + K + + + +K++ +KS + PR +R +
Sbjct: 183 FERQLKQL-------PDAEQPAAAPVKKSKQKSASTAPPRT-------------RRNSSV 222
Query: 233 SSLAARDAKVEVPKLSQVHCKSIEKDLIKGSEERDRIAPGAVAFKQKRQIKSTS-PLERK 291
SS +A A PK + K + + + R + KRQ +S + P +
Sbjct: 223 SSTSASVAASTAPKAASPAVLPEGKPRRRKNNSQMRFCSTVLKELYKRQYESFAFPFYQP 282
Query: 292 SDP 294
DP
Sbjct: 283 VDP 285
>UniRef100_Q9LXA7 Bromodomain protein-like [Arabidopsis thaliana]
Length = 678
Score = 103 bits (258), Expect = 9e-21
Identities = 51/116 (43%), Positives = 75/116 (63%)
Query: 73 QKIDRKGSTQCAAILKCLMSYPYSWVFNKPVDPVALNIPDYFAIIAKPMDLGTISSKLKK 132
+K+ + T C IL LM + +SWVF PVD V L + DY I+ KPMDLGT+ L+K
Sbjct: 243 EKVLKSMMTTCGQILVKLMKHKWSWVFLNPVDVVGLGLHDYHRIVDKPMDLGTVKMNLEK 302
Query: 133 NVYSCVEEFAADVRLTFSNAMTYNPPHNDVHMMAKELNQIFDSKWKDMDKKWKCED 188
+Y +FA+DVRLTF+NAM+YNP DV++MA++L FD + K+++ ++
Sbjct: 303 GLYRSPIDFASDVRLTFTNAMSYNPKGQDVYLMAEKLLSQFDVWFNPTLKRFEAQE 358
>UniRef100_Q9S7T1 Hypothetical protein T18K17.19 [Arabidopsis thaliana]
Length = 461
Score = 103 bits (258), Expect = 9e-21
Identities = 51/112 (45%), Positives = 73/112 (64%), Gaps = 4/112 (3%)
Query: 78 KGSTQ----CAAILKCLMSYPYSWVFNKPVDPVALNIPDYFAIIAKPMDLGTISSKLKKN 133
KG+ Q C +L LM + W+FN PVD V L + DY II +PMDLGT+ ++L K+
Sbjct: 114 KGTVQILKSCNNLLTKLMKHKSGWIFNTPVDVVTLGLHDYHNIIKEPMDLGTVKTRLSKS 173
Query: 134 VYSCVEEFAADVRLTFSNAMTYNPPHNDVHMMAKELNQIFDSKWKDMDKKWK 185
+Y EFA DVRLTF+NAM YNP +DV+ MA+ L +F+ KW ++ +++
Sbjct: 174 LYKSPLEFAEDVRLTFNNAMLYNPVGHDVYHMAEILLNLFEEKWVPLETQYE 225
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.313 0.130 0.364
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 835,307,940
Number of Sequences: 2790947
Number of extensions: 34954016
Number of successful extensions: 189300
Number of sequences better than 10.0: 4458
Number of HSP's better than 10.0 without gapping: 1557
Number of HSP's successfully gapped in prelim test: 3087
Number of HSP's that attempted gapping in prelim test: 155919
Number of HSP's gapped (non-prelim): 18525
length of query: 510
length of database: 848,049,833
effective HSP length: 132
effective length of query: 378
effective length of database: 479,644,829
effective search space: 181305745362
effective search space used: 181305745362
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 77 (34.3 bits)
Lotus: description of TM0228a.2


