

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0227.11
(425 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
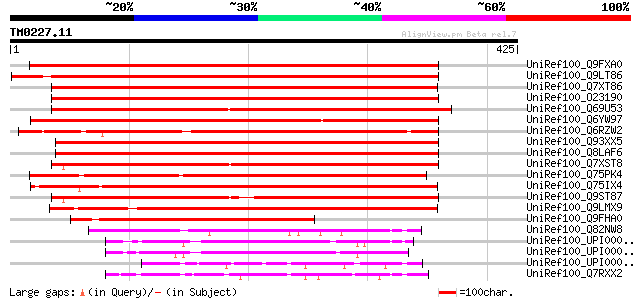
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9FXA0 F14J22.5 protein [Arabidopsis thaliana] 539 e-152
UniRef100_Q9LT86 MAP3K protein kinase-like protein [Arabidopsis ... 516 e-145
UniRef100_Q7XT86 OSJNBa0042L16.16 protein [Oryza sativa] 477 e-133
UniRef100_O23190 MAP3K-like protein kinase [Arabidopsis thaliana] 471 e-131
UniRef100_Q69U53 MAP3K-like protein [Oryza sativa] 458 e-127
UniRef100_Q6YW97 MAP3K protein kinase-like protein [Oryza sativa] 406 e-112
UniRef100_Q6RZW2 Hypothetical protein G9-3 [Musa acuminata] 404 e-111
UniRef100_Q93XX5 Hypothetical protein At5g67130; K21H1.9 [Arabid... 399 e-110
UniRef100_Q8LAF6 Hypothetical protein [Arabidopsis thaliana] 397 e-109
UniRef100_Q7XST8 OSJNBa0039K24.18 protein [Oryza sativa] 396 e-109
UniRef100_Q75PK4 Hypothetical protein LjNUF [Lotus japonicus] 396 e-109
UniRef100_Q75IX4 Expressed protein [Oryza sativa] 384 e-105
UniRef100_Q9ST87 CAA303710.1 protein [Oryza sativa] 374 e-102
UniRef100_Q9LMX9 F21F23.12 protein [Arabidopsis thaliana] 341 3e-92
UniRef100_Q9FHA0 Similarity to MAP3K-like protein kinase [Arabid... 260 6e-68
UniRef100_Q82NW8 Hypothetical protein [Streptomyces avermitilis] 106 1e-21
UniRef100_UPI00004301F4 UPI00004301F4 UniRef100 entry 76 2e-12
UniRef100_UPI00003C1EBF UPI00003C1EBF UniRef100 entry 73 2e-11
UniRef100_UPI0000235E3F UPI0000235E3F UniRef100 entry 70 1e-10
UniRef100_Q7RXX2 Hypothetical protein [Neurospora crassa] 70 1e-10
>UniRef100_Q9FXA0 F14J22.5 protein [Arabidopsis thaliana]
Length = 359
Score = 539 bits (1389), Expect = e-152
Identities = 246/343 (71%), Positives = 289/343 (83%)
Query: 17 ILVLLDSSLALKFKPLQQEGQICVANKNCNSGLHCETCVTNGNVRPRCTRTQPINPTSKV 76
I +LL SS L+ +EG+ C+ N NC++GLHCETC+ N + RPRC+RTQPINP +K
Sbjct: 11 IALLLQSSFLLEISSALKEGKTCITNSNCDAGLHCETCIANTDFRPRCSRTQPINPITKA 70
Query: 77 KGLPFNRYSWLTTHNSFALLGQKSMTGSVILSPTNQQDTITDQLNNGVRGLMLDLYDFEN 136
KGLPFN+YSWLTTHNSFA LG+ S TGS IL+PTNQQD+IT QLNNGVRG MLD+YDF+N
Sbjct: 71 KGLPFNKYSWLTTHNSFARLGEVSRTGSAILAPTNQQDSITSQLNNGVRGFMLDMYDFQN 130
Query: 137 DVWLCHSFGGQCYNYTAFQPAINVLKEIQVFLEANPSEIVTIIIEDYVTSPKGLTKVFDA 196
D+WLCHSF G C+N+TAFQPAIN+L+E QVFLE N E+VTIIIEDYV SPKGLTKVFDA
Sbjct: 131 DIWLCHSFDGTCFNFTAFQPAINILREFQVFLEKNKEEVVTIIIEDYVKSPKGLTKVFDA 190
Query: 197 AGLRKYWFPVSRMPKNGGDWPKVDDMVQKNQRLVVFTSKASKEASEGIAYEWRYLVENQY 256
AGLRK+ FPVSRMPKNGGDWP++DDMV+KNQRL+VFTS + KEA+EGIAY+W+Y+VENQY
Sbjct: 191 AGLRKFMFPVSRMPKNGGDWPRLDDMVRKNQRLLVFTSDSHKEATEGIAYQWKYMVENQY 250
Query: 257 GNGGMKAGSCPNRAESPSMNTTSRSLVLVNFFRDLPDVAQSCKDNSAPLLSMVSTCNQAA 316
GNGG+K G CPNRA+S M+ S+SLVLVN F D DV +CK NSA LL + TC QAA
Sbjct: 251 GNGGLKVGVCPNRAQSAPMSDKSKSLVLVNHFPDAADVIVACKQNSASLLESIKTCYQAA 310
Query: 317 GKRWPNFIAVDFYKRSDGGGAPEAVDVANGHLVCGCGNIATCK 359
G+RWPNFIAVDFYKRSDGGGAP+AVDVANG+L+CGC N A CK
Sbjct: 311 GQRWPNFIAVDFYKRSDGGGAPQAVDVANGNLICGCDNFAACK 353
>UniRef100_Q9LT86 MAP3K protein kinase-like protein [Arabidopsis thaliana]
Length = 413
Score = 516 bits (1329), Expect = e-145
Identities = 240/359 (66%), Positives = 289/359 (79%), Gaps = 7/359 (1%)
Query: 2 MMQKPTLAIATTLFT-ILVLLDSSLALKFKPLQQEGQICVANKNCNSGLHCETCVTNGNV 60
M Q+ T + L IL++L S ALK EG+ C+ +KNC+ GLHCE+C+ + +
Sbjct: 1 MFQRFTFFLTALLIPCILIILSPSYALK------EGETCIVSKNCDRGLHCESCLASDSF 54
Query: 61 RPRCTRTQPINPTSKVKGLPFNRYSWLTTHNSFALLGQKSMTGSVILSPTNQQDTITDQL 120
RPRC+R QPINPT+KVKGLP+N+YSWLTTHNSFA +G KS TGS+IL+P+NQQD+IT QL
Sbjct: 55 RPRCSRMQPINPTTKVKGLPYNKYSWLTTHNSFARMGAKSGTGSMILAPSNQQDSITSQL 114
Query: 121 NNGVRGLMLDLYDFENDVWLCHSFGGQCYNYTAFQPAINVLKEIQVFLEANPSEIVTIII 180
NGVRG MLDLYDF+ND+WLCHS+GG C+NYTAFQPA+N+LKE QVFL+ N +VT+I+
Sbjct: 115 LNGVRGFMLDLYDFQNDIWLCHSYGGNCFNYTAFQPAVNILKEFQVFLDKNKDVVVTLIL 174
Query: 181 EDYVTSPKGLTKVFDAAGLRKYWFPVSRMPKNGGDWPKVDDMVQKNQRLVVFTSKASKEA 240
EDYV SP GLT+VFDA+GLR + FPVSRMPKNG DWP +DDM+ +NQRL+VFTS KEA
Sbjct: 175 EDYVKSPNGLTRVFDASGLRNFMFPVSRMPKNGEDWPTLDDMICQNQRLLVFTSNPQKEA 234
Query: 241 SEGIAYEWRYLVENQYGNGGMKAGSCPNRAESPSMNTTSRSLVLVNFFRDLPDVAQSCKD 300
SEGIA+ WRY++ENQYG+GGMKAG C NR ES +M SRSL+LVN+F D DV SCK
Sbjct: 235 SEGIAFMWRYMIENQYGDGGMKAGVCTNRPESVAMGDRSRSLILVNYFPDTADVIGSCKQ 294
Query: 301 NSAPLLSMVSTCNQAAGKRWPNFIAVDFYKRSDGGGAPEAVDVANGHLVCGCGNIATCK 359
NSAPLL V C +A+GKRWPNFIAVDFYKRSDGGGAP+AVDVANGH VCGC +IA CK
Sbjct: 295 NSAPLLDTVKNCQEASGKRWPNFIAVDFYKRSDGGGAPKAVDVANGHAVCGCEDIAACK 353
>UniRef100_Q7XT86 OSJNBa0042L16.16 protein [Oryza sativa]
Length = 413
Score = 477 bits (1227), Expect = e-133
Identities = 216/323 (66%), Positives = 262/323 (80%)
Query: 36 GQICVANKNCNSGLHCETCVTNGNVRPRCTRTQPINPTSKVKGLPFNRYSWLTTHNSFAL 95
G+ C A++NC++GLHCETCV +GNVRPRCTR P++P +K + LPFNRY+WLTTHNSFA
Sbjct: 35 GETCAADRNCDAGLHCETCVADGNVRPRCTRVTPVDPQTKARDLPFNRYAWLTTHNSFAR 94
Query: 96 LGQKSMTGSVILSPTNQQDTITDQLNNGVRGLMLDLYDFENDVWLCHSFGGQCYNYTAFQ 155
LG +S TG+ I + NQQDTITDQLNNGVRGLMLD+YDF ND+WLCHSFGG C N+TAF
Sbjct: 95 LGTRSRTGTAIATAWNQQDTITDQLNNGVRGLMLDMYDFRNDIWLCHSFGGACQNFTAFV 154
Query: 156 PAINVLKEIQVFLEANPSEIVTIIIEDYVTSPKGLTKVFDAAGLRKYWFPVSRMPKNGGD 215
PA+ VL EI+ FL NPSE+VT+ +EDYV SP GLT+V +A+GL KY FP RMPK+GGD
Sbjct: 155 PAVEVLGEIERFLARNPSEVVTVFVEDYVESPMGLTRVLNASGLTKYVFPAWRMPKSGGD 214
Query: 216 WPKVDDMVQKNQRLVVFTSKASKEASEGIAYEWRYLVENQYGNGGMKAGSCPNRAESPSM 275
WP++ DMV+ N RL++FTSK++KEA+EGI YEW Y+VENQYG GM G CPNRAES +M
Sbjct: 215 WPRLSDMVRDNHRLLLFTSKSAKEAAEGIPYEWHYVVENQYGTKGMIKGRCPNRAESAAM 274
Query: 276 NTTSRSLVLVNFFRDLPDVAQSCKDNSAPLLSMVSTCNQAAGKRWPNFIAVDFYKRSDGG 335
N SRSLVLVN+FRDLP+ +CKDNSA LL M++TC+ + RW NFIAVDFYKRSD G
Sbjct: 275 NDLSRSLVLVNYFRDLPNFPVACKDNSAELLDMLTTCHDLSADRWANFIAVDFYKRSDRG 334
Query: 336 GAPEAVDVANGHLVCGCGNIATC 358
GA EA D ANG LVCGCG+++ C
Sbjct: 335 GAAEATDRANGGLVCGCGSVSAC 357
>UniRef100_O23190 MAP3K-like protein kinase [Arabidopsis thaliana]
Length = 799
Score = 471 bits (1211), Expect = e-131
Identities = 214/324 (66%), Positives = 261/324 (80%)
Query: 36 GQICVANKNCNSGLHCETCVTNGNVRPRCTRTQPINPTSKVKGLPFNRYSWLTTHNSFAL 95
G+ C + C++GL C++C NGN CTR QP+NPTSKV GLPFN+YSWLTTHNS+A+
Sbjct: 340 GETCSSTSECDAGLSCQSCPANGNTGSTCTRIQPLNPTSKVNGLPFNKYSWLTTHNSYAI 399
Query: 96 LGQKSMTGSVILSPTNQQDTITDQLNNGVRGLMLDLYDFENDVWLCHSFGGQCYNYTAFQ 155
G S TGS ++SP NQ+D+IT+QL NGVRG+MLD YDF+ND+WLCHS GG C+N+TAFQ
Sbjct: 400 TGANSATGSFLVSPKNQEDSITNQLKNGVRGIMLDTYDFQNDIWLCHSTGGTCFNFTAFQ 459
Query: 156 PAINVLKEIQVFLEANPSEIVTIIIEDYVTSPKGLTKVFDAAGLRKYWFPVSRMPKNGGD 215
PAIN LKEI FLE+N SEIVTII+EDYV S GLT VF+A+GL K+ P+SRMPK+G D
Sbjct: 460 PAINALKEINDFLESNLSEIVTIILEDYVKSQMGLTNVFNASGLSKFLLPISRMPKDGTD 519
Query: 216 WPKVDDMVQKNQRLVVFTSKASKEASEGIAYEWRYLVENQYGNGGMKAGSCPNRAESPSM 275
WP VDDMV++NQRLVVFTSK KEASEG+AY+W Y+VENQYGN GMK GSC +R+ES S+
Sbjct: 520 WPTVDDMVKQNQRLVVFTSKKDKEASEGLAYQWNYMVENQYGNDGMKDGSCSSRSESSSL 579
Query: 276 NTTSRSLVLVNFFRDLPDVAQSCKDNSAPLLSMVSTCNQAAGKRWPNFIAVDFYKRSDGG 335
+T SRSLV N+F P+ Q+C DNS+PL+ M+ TC++AAGKRWPNFIAVDFY+RSD G
Sbjct: 580 DTMSRSLVFQNYFETSPNSTQACADNSSPLIEMMRTCHEAAGKRWPNFIAVDFYQRSDSG 639
Query: 336 GAPEAVDVANGHLVCGCGNIATCK 359
GA EAVD ANG L CGC ++ CK
Sbjct: 640 GAAEAVDEANGRLTCGCDSLVYCK 663
>UniRef100_Q69U53 MAP3K-like protein [Oryza sativa]
Length = 411
Score = 458 bits (1179), Expect = e-127
Identities = 213/335 (63%), Positives = 257/335 (76%), Gaps = 1/335 (0%)
Query: 36 GQICVANKNCNSGLHCETCVTNGNVRPRCTRTQPINPTSKVKGLPFNRYSWLTTHNSFAL 95
G C + +C +GLHC C G CTR +PI+P + LPFN YSWLTTHNS+AL
Sbjct: 24 GDTCSSEGDCGAGLHCSDCGGGGGGDKTCTRAKPIDPLTHGTDLPFNNYSWLTTHNSYAL 83
Query: 96 LGQKSMTGSVILSPTNQQDTITDQLNNGVRGLMLDLYDFENDVWLCHSFGGQCYNYTAFQ 155
G S TGS +++ TNQ+DTIT QL NGVRGLMLD YDF NDVWLCHSF G+C+N+TAFQ
Sbjct: 84 AGSSSATGSALITQTNQEDTITAQLKNGVRGLMLDTYDFNNDVWLCHSFQGKCFNFTAFQ 143
Query: 156 PAINVLKEIQVFLEANPSEIVTIIIEDYVTSPKGLTKVFDAAGLRKYWFPVSRMPKNGGD 215
PAINVLKEI+ FL+ NPSE++TI +EDY T+ L KVF+A+GL KYWFPV++MPK+GGD
Sbjct: 144 PAINVLKEIRTFLDGNPSEVITIFLEDY-TASGSLPKVFNASGLMKYWFPVAKMPKSGGD 202
Query: 216 WPKVDDMVQKNQRLVVFTSKASKEASEGIAYEWRYLVENQYGNGGMKAGSCPNRAESPSM 275
WP + DM+ +N+RL+VFTSK SKEASEGIAYEW Y+VENQYGN GM G CPNRAESP+M
Sbjct: 203 WPLLKDMISQNERLLVFTSKKSKEASEGIAYEWSYVVENQYGNEGMVEGKCPNRAESPAM 262
Query: 276 NTTSRSLVLVNFFRDLPDVAQSCKDNSAPLLSMVSTCNQAAGKRWPNFIAVDFYKRSDGG 335
++ S+SLVL+NFF P C +NSAPL+SM+ TC+ +G RWPN+IAVDFY RSDGG
Sbjct: 263 DSKSQSLVLMNFFTTDPSQTGVCANNSAPLVSMLKTCHDLSGNRWPNYIAVDFYMRSDGG 322
Query: 336 GAPEAVDVANGHLVCGCGNIATCKVRLTPITFIPP 370
GAP A D+ANGHLVCGC NIA CK T T + P
Sbjct: 323 GAPLATDIANGHLVCGCDNIAYCKANSTFGTCVIP 357
>UniRef100_Q6YW97 MAP3K protein kinase-like protein [Oryza sativa]
Length = 412
Score = 406 bits (1044), Expect = e-112
Identities = 194/342 (56%), Positives = 243/342 (70%), Gaps = 1/342 (0%)
Query: 18 LVLLDSSLALKFKPLQQEGQICVANKNCNSGLHCETCVTNGNVRPRCTRTQPINPTSKVK 77
L+ L +L L Q G C + ++C +GL+C C G RP C R I PTS VK
Sbjct: 7 LLALHIALLLLLPCSCQVGDSCSSARDCGAGLYCGNCAATGKTRPSCIRDLAIQPTSIVK 66
Query: 78 GLPFNRYSWLTTHNSFALLGQKSMTGSVILSPTNQQDTITDQLNNGVRGLMLDLYDFEND 137
GLPFNRYSWL THNSF+++G+ S TG ++ NQ+DT+T+QL NGVRGLMLD+YDF +D
Sbjct: 67 GLPFNRYSWLVTHNSFSIIGEPSHTGVERVTFYNQEDTVTNQLRNGVRGLMLDMYDFNDD 126
Query: 138 VWLCHSFGGQCYNYTAFQPAINVLKEIQVFLEANPSEIVTIIIEDYVTSPKGLTKVFDAA 197
+WLCHS GQCYN+TAFQPAI+ LKE++ FL NP+EI+TI IEDYV S GL+K+F AA
Sbjct: 127 IWLCHSLQGQCYNFTAFQPAIDTLKEVEAFLSENPTEIITIFIEDYVHSTMGLSKLFTAA 186
Query: 198 GLRKYWFPVSRMPKNGGDWPKVDDMVQKNQRLVVFTSKASKEASEGIAYEWRYLVENQYG 257
L KYW+P+S MP NG DWP V DMV KN RL+VFTS +SKEASEGIAY+W YL+EN+ G
Sbjct: 187 DLTKYWYPISEMPTNGKDWPSVTDMVAKNHRLLVFTSDSSKEASEGIAYQWSYLLENESG 246
Query: 258 NGGMKAGSCPNRAESPSMNTTSRSLVLVNFFRDLPDVAQSCKDNSAPLLSMVSTCNQAAG 317
+ G+ GSCPNR ES +N+ S SL + N+F +P ++CK+NS L MV TC AAG
Sbjct: 247 DPGI-TGSCPNRKESQPLNSRSASLFMQNYFPTIPVENEACKENSVGLPQMVQTCYTAAG 305
Query: 318 KRWPNFIAVDFYKRSDGGGAPEAVDVANGHLVCGCGNIATCK 359
R PNFIAV++Y RSDGGG + D NG +CGC IA C+
Sbjct: 306 NRIPNFIAVNYYMRSDGGGVFDVQDRINGVTLCGCNTIAACQ 347
>UniRef100_Q6RZW2 Hypothetical protein G9-3 [Musa acuminata]
Length = 376
Score = 404 bits (1038), Expect = e-111
Identities = 207/384 (53%), Positives = 255/384 (65%), Gaps = 45/384 (11%)
Query: 8 LAIATTLFTILVLLDSSLALKFKPLQQEGQICVANKNCNSGLHCETCVTNGNVRPRCTRT 67
+A+ + F +L+ L +L + G+ C AN++C++GL C+ C + V C R
Sbjct: 1 MALPPSSFFLLLFLSVGSSL-LSSAAKLGEGCSANQDCDAGLRCDGCDGDLGV---CVRI 56
Query: 68 QPINPTSKV--------------------------------KGLPFNRYSWLTTHNSFAL 95
+P P SKV K LPFN+YSWLTTHNSFA
Sbjct: 57 RPYEPRSKVRIRHYPFSIRNLGLWVGWFRFRANLGLECAQGKDLPFNKYSWLTTHNSFAD 116
Query: 96 LGQKSMTGSVILSPTNQQDTITDQLNNGVRGLMLDLYDFENDVWLCHSFGGQCYNYTAFQ 155
G S TG+ +++ TNQ D IT QLNNGVRGLMLD+YDF ND+WLCHS Q
Sbjct: 117 AGAHSATGATLITFTNQHDNITSQLNNGVRGLMLDMYDFRNDIWLCHSTA------VYQQ 170
Query: 156 PAINVLKEIQVFLEANPSEIVTIIIEDYVTSPKGLTKVFDAAGLRKYWFPVSRMPKNGGD 215
PAINVLKEI+ FL ANPSE++TI IEDYV SP GL+KVF+A+GL KYWFPV +MPKNG D
Sbjct: 171 PAINVLKEIETFLAANPSEVITIFIEDYVKSPSGLSKVFNASGLMKYWFPVDQMPKNGSD 230
Query: 216 WPKVDDMVQKNQRLVVFTSKASKEASEGIAYEWRYLVENQYGNGGMKAGSCPNRAESPSM 275
WP + M+ +N RL+VFTS ASKEASEGIAYEW Y+VENQYG+ GM GSCP+RAES M
Sbjct: 231 WPLLSKMIDQNHRLLVFTSVASKEASEGIAYEWNYVVENQYGDEGMTPGSCPSRAESSPM 290
Query: 276 NTTSRSLVLVNFFRDLPDVAQSCKDNSAPLLSMVSTCNQAAGKRWPNFIAVDFYKRSDGG 335
+TT +SLVL+N+FR P + +C +NSAPLL M+ TC+ + RW NFIAVDFY + D
Sbjct: 291 STTLKSLVLMNYFRTNPSASSACHNNSAPLLDMLKTCHGLSANRWANFIAVDFYMKGD-- 348
Query: 336 GAPEAVDVANGHLVCGCGNIATCK 359
APEA DVANGH+VCGC NIA CK
Sbjct: 349 -APEAADVANGHMVCGCDNIAYCK 371
>UniRef100_Q93XX5 Hypothetical protein At5g67130; K21H1.9 [Arabidopsis thaliana]
Length = 426
Score = 399 bits (1025), Expect = e-110
Identities = 183/321 (57%), Positives = 237/321 (73%)
Query: 39 CVANKNCNSGLHCETCVTNGNVRPRCTRTQPINPTSKVKGLPFNRYSWLTTHNSFALLGQ 98
C + +C SGL+C C G +P CTR Q +PTS + GLPFN+Y+WL THN+F+
Sbjct: 40 CSSATDCVSGLYCGDCPAVGRSKPVCTRGQATSPTSIINGLPFNKYTWLMTHNAFSNANA 99
Query: 99 KSMTGSVILSPTNQQDTITDQLNNGVRGLMLDLYDFENDVWLCHSFGGQCYNYTAFQPAI 158
+ G ++ NQ+DTIT+QL NGVRGLMLD+YDF ND+WLCHS GQC+N+TAFQPAI
Sbjct: 100 PLLPGVERITFYNQEDTITNQLQNGVRGLMLDMYDFNNDIWLCHSLRGQCFNFTAFQPAI 159
Query: 159 NVLKEIQVFLEANPSEIVTIIIEDYVTSPKGLTKVFDAAGLRKYWFPVSRMPKNGGDWPK 218
N+L+E++ FL NP+EIVTIIIEDYV PKGL+ +F AGL KYWFPVS+MP+ G DWP
Sbjct: 160 NILREVEAFLSQNPTEIVTIIIEDYVHRPKGLSTLFANAGLDKYWFPVSKMPRKGEDWPT 219
Query: 219 VDDMVQKNQRLVVFTSKASKEASEGIAYEWRYLVENQYGNGGMKAGSCPNRAESPSMNTT 278
V DMVQ+N RL+VFTS A+KE EG+AY+WRY+VEN+ G+ G+K GSCPNR ES +N+
Sbjct: 220 VTDMVQENHRLLVFTSVAAKEDEEGVAYQWRYMVENESGDPGVKRGSCPNRKESQPLNSK 279
Query: 279 SRSLVLVNFFRDLPDVAQSCKDNSAPLLSMVSTCNQAAGKRWPNFIAVDFYKRSDGGGAP 338
S SL L+N+F P +CK++SAPL MV TC ++ G R PNF+AV+FY RSDGGG
Sbjct: 280 SSSLFLMNYFPTYPVEKDACKEHSAPLAEMVGTCLKSGGNRMPNFLAVNFYMRSDGGGVF 339
Query: 339 EAVDVANGHLVCGCGNIATCK 359
E +D NG ++CGC ++ C+
Sbjct: 340 EILDRMNGPVLCGCETLSACQ 360
>UniRef100_Q8LAF6 Hypothetical protein [Arabidopsis thaliana]
Length = 426
Score = 397 bits (1021), Expect = e-109
Identities = 182/321 (56%), Positives = 236/321 (72%)
Query: 39 CVANKNCNSGLHCETCVTNGNVRPRCTRTQPINPTSKVKGLPFNRYSWLTTHNSFALLGQ 98
C + +C SGL+C C G +P CTR Q +PTS + GLPFN+Y+WL THN+F+
Sbjct: 40 CSSATDCVSGLYCGDCPAVGRSKPVCTRGQATSPTSIINGLPFNKYTWLMTHNAFSNANA 99
Query: 99 KSMTGSVILSPTNQQDTITDQLNNGVRGLMLDLYDFENDVWLCHSFGGQCYNYTAFQPAI 158
+ G ++ NQ+DTIT+QL NGVRGLMLD+YDF ND+WLCHS GQC+N+T FQPAI
Sbjct: 100 PLLPGVERITFYNQEDTITNQLQNGVRGLMLDMYDFNNDIWLCHSLRGQCFNFTXFQPAI 159
Query: 159 NVLKEIQVFLEANPSEIVTIIIEDYVTSPKGLTKVFDAAGLRKYWFPVSRMPKNGGDWPK 218
N+L+E++ FL NP+EIVTIIIEDYV PKGL+ +F AGL KYWFPVS+MP+ G DWP
Sbjct: 160 NILREVEAFLSQNPTEIVTIIIEDYVHRPKGLSTLFANAGLDKYWFPVSKMPRKGEDWPT 219
Query: 219 VDDMVQKNQRLVVFTSKASKEASEGIAYEWRYLVENQYGNGGMKAGSCPNRAESPSMNTT 278
V DMVQ+N RL+VFTS A+KE EG+AY+WRY+VEN+ G+ G+K GSCPNR ES +N+
Sbjct: 220 VTDMVQENHRLLVFTSVAAKEDEEGVAYQWRYMVENESGDPGVKRGSCPNRKESQPLNSK 279
Query: 279 SRSLVLVNFFRDLPDVAQSCKDNSAPLLSMVSTCNQAAGKRWPNFIAVDFYKRSDGGGAP 338
S SL L+N+F P +CK++SAPL MV TC ++ G R PNF+AV+FY RSDGGG
Sbjct: 280 SSSLFLMNYFPTYPVEKDACKEHSAPLAEMVGTCLKSGGNRMPNFLAVNFYMRSDGGGVF 339
Query: 339 EAVDVANGHLVCGCGNIATCK 359
E +D NG ++CGC ++ C+
Sbjct: 340 EILDRMNGPVLCGCETLSACQ 360
>UniRef100_Q7XST8 OSJNBa0039K24.18 protein [Oryza sativa]
Length = 468
Score = 396 bits (1018), Expect = e-109
Identities = 183/327 (55%), Positives = 240/327 (72%), Gaps = 4/327 (1%)
Query: 36 GQICVANK--NCNSGLHCETCVT-NGNVRPRCTRTQPINPTSKVKGLPFNRYSWLTTHNS 92
G C A+ +C +G+ C TC G P C+RT P++P + L FNRY+WLTTHNS
Sbjct: 31 GDTCTASSASSCGAGMRCATCSPLPGMGPPVCSRTTPLDPKAHGTDLAFNRYTWLTTHNS 90
Query: 93 FALLGQKSMTGSVILSPTNQQDTITDQLNNGVRGLMLDLYDFENDVWLCHSFGGQCYNYT 152
FA++G S TG+ I++P NQ+DT+T QL NGVRGLMLD YDF+N+VWLCHSFGG+CYN+
Sbjct: 91 FAIVGSPSRTGTPIIAPPNQEDTVTAQLKNGVRGLMLDAYDFQNEVWLCHSFGGKCYNFA 150
Query: 153 AFQPAINVLKEIQVFLEANPSEIVTIIIEDYVTSPKGLTKVFDAAGLRKYWFPVSRMPKN 212
A+Q A++VLKEI FL+ANPSE++T+ +EDY P L KV +GL KY FP ++MPK
Sbjct: 151 AYQRAMDVLKEIGAFLDANPSEVITVFVEDYA-GPGSLGKVVGGSGLSKYLFPPAKMPKG 209
Query: 213 GGDWPKVDDMVQKNQRLVVFTSKASKEASEGIAYEWRYLVENQYGNGGMKAGSCPNRAES 272
GGDWP + DM+ +N RL++FTSK K+ S+G+AYEW Y++E QYGN G+ GSCP RAES
Sbjct: 210 GGDWPLLKDMIAQNHRLLMFTSKRGKDGSDGLAYEWDYVLETQYGNDGLVGGSCPKRAES 269
Query: 273 PSMNTTSRSLVLVNFFRDLPDVAQSCKDNSAPLLSMVSTCNQAAGKRWPNFIAVDFYKRS 332
+M++T +SL+L+NFF P + +C +NSAPL++ + C A+ KRWPNFIAVD+Y RS
Sbjct: 270 MAMDSTKQSLILMNFFSTNPSQSWACGNNSAPLVAKLKACYDASAKRWPNFIAVDYYMRS 329
Query: 333 DGGGAPEAVDVANGHLVCGCGNIATCK 359
GGGAP A DVANG CGC +IA CK
Sbjct: 330 KGGGAPLATDVANGRQQCGCDSIAYCK 356
>UniRef100_Q75PK4 Hypothetical protein LjNUF [Lotus japonicus]
Length = 337
Score = 396 bits (1018), Expect = e-109
Identities = 196/336 (58%), Positives = 252/336 (74%), Gaps = 10/336 (2%)
Query: 17 ILVLLDSSLALK-FKPLQQEGQICVANKN-CNSGLHCETCVTNGNVRPRCTRTQPINPTS 74
I L+ +SL + L Q + C + N C+ GL C C + RCTR + I+PTS
Sbjct: 7 IATLVSASLVFGCYYILVQIAETCSRDINDCDLGLQCLECHSQN----RCTRIRTISPTS 62
Query: 75 KVKGLPFNRYSWLTTHNSFALLGQKSMTGSVILSPTNQQDTITDQLNNGVRGLMLDLYDF 134
KV LPFN YSWLTTHNSFA G TGS +L+ TNQ+D+ITDQL NGVRGLMLD++D+
Sbjct: 63 KVMELPFNEYSWLTTHNSFAAKGVNWSTGSPVLAFTNQEDSITDQLKNGVRGLMLDMWDY 122
Query: 135 ENDVWLCHSFGGQCYNYTAFQPAINVLKEIQVFLEANPSEIVTIIIEDYVTSPKGLTKVF 194
E+ +WLC G C YT FQPA+NVLKE++VFL +P+EI+TI I+D+VTS G+ KVF
Sbjct: 123 EDTIWLCR---GPCTKYTTFQPALNVLKEVRVFLVTHPTEIITIFIDDHVTSGNGVNKVF 179
Query: 195 DAAGLRKYWFPVSRMPKNGGDWPKVDDMVQKNQRLVVFTSKASKEASEGIAYEWRYLVEN 254
D A LRK+WFPVS+MPKNG DWP V M++KN RL+VFTS AS+EASEGIAYEW Y+VE+
Sbjct: 180 DKARLRKFWFPVSKMPKNGSDWPTVKTMIRKNYRLIVFTSNASREASEGIAYEWNYVVES 239
Query: 255 QYGNGGMKAGSCPNRAESPSMNTTSRSLVLVNFFRDLPD-VAQSCKDNSAPLLSMVSTCN 313
Q+GN G+K GSC NR ES MN ++SLVL+N+FR++ + ++C+DNS+PL++M+ C
Sbjct: 240 QFGNVGIKGGSCQNRPESLPMNNATKSLVLMNYFRNVRNHDDEACRDNSSPLIAMMHVCF 299
Query: 314 QAAGKRWPNFIAVDFYKRSDGGGAPEAVDVANGHLV 349
+AAG RWPNFIAVDFYKR DGGGAPEA+D+AN +L+
Sbjct: 300 RAAGNRWPNFIAVDFYKRGDGGGAPEALDLANRNLI 335
>UniRef100_Q75IX4 Expressed protein [Oryza sativa]
Length = 360
Score = 384 bits (985), Expect = e-105
Identities = 189/343 (55%), Positives = 238/343 (69%), Gaps = 7/343 (2%)
Query: 18 LVLLDSSLALKFKPLQQEGQICVANKNCNSGLHCETCVTN--GNVRPRCTRTQPINPTSK 75
LVLL +AL G C +C G C C G+ R T P T+
Sbjct: 13 LVLL---VALSVAATANVGDSCSTAVDCGGGQWCFDCQPEFAGSSCVRSAATNPFQLTNN 69
Query: 76 VKGLPFNRYSWLTTHNSFALLGQKSMTGSVILSPTNQQDTITDQLNNGVRGLMLDLYDFE 135
LPFN+Y++LTTHNSFA++G+ S TG ++ NQ+DT+TDQLNNGVR LMLD YDF+
Sbjct: 70 --SLPFNKYAYLTTHNSFAIVGEPSHTGVPRITFDNQEDTVTDQLNNGVRALMLDTYDFK 127
Query: 136 NDVWLCHSFGGQCYNYTAFQPAINVLKEIQVFLEANPSEIVTIIIEDYVTSPKGLTKVFD 195
DVWLCHS GG+C ++TAF+PA++ KEI+ FL ANPSEIVT+I+EDYV +P GLT VF
Sbjct: 128 GDVWLCHSNGGKCNDFTAFEPALDTFKEIEAFLGANPSEIVTLILEDYVHAPNGLTNVFK 187
Query: 196 AAGLRKYWFPVSRMPKNGGDWPKVDDMVQKNQRLVVFTSKASKEASEGIAYEWRYLVENQ 255
A+GL KYWFPVS+MP+ G DWP V DMV NQRL+VFTS SK+A+EGIAY+W Y+VEN
Sbjct: 188 ASGLMKYWFPVSKMPQKGKDWPLVSDMVASNQRLLVFTSIRSKQATEGIAYQWNYMVENN 247
Query: 256 YGNGGMKAGSCPNRAESPSMNTTSRSLVLVNFFRDLPDVAQSCKDNSAPLLSMVSTCNQA 315
YG+ GM AG C NRAES +N ++SLVLVN+F +P +C +S L MV+TC A
Sbjct: 248 YGDDGMDAGKCSNRAESAPLNDKTKSLVLVNYFPSVPVKVTACLQHSKSLTDMVNTCYGA 307
Query: 316 AGKRWPNFIAVDFYKRSDGGGAPEAVDVANGHLVCGCGNIATC 358
AG RW N +AVD+YKRSDGGGA +A D+ NG L+CGC ++ C
Sbjct: 308 AGNRWANLLAVDYYKRSDGGGAFQATDLLNGRLLCGCQDVRAC 350
>UniRef100_Q9ST87 CAA303710.1 protein [Oryza sativa]
Length = 416
Score = 374 bits (960), Expect = e-102
Identities = 177/327 (54%), Positives = 231/327 (70%), Gaps = 16/327 (4%)
Query: 36 GQICVANK--NCNSGLHCETCVT-NGNVRPRCTRTQPINPTSKVKGLPFNRYSWLTTHNS 92
G C A+ +C +G+ C TC G P C+RT P++P + L FNRY+WLTTHNS
Sbjct: 31 GDTCTASSASSCGAGMRCATCSPLPGMGPPVCSRTTPLDPKAHGTDLAFNRYTWLTTHNS 90
Query: 93 FALLGQKSMTGSVILSPTNQQDTITDQLNNGVRGLMLDLYDFENDVWLCHSFGGQCYNYT 152
FA++G S TG+ I++P NQ+DT+T QL NGVRGLMLD YDF+N+VWLCHSFGG+CYN+
Sbjct: 91 FAIVGSPSRTGTPIIAPPNQEDTVTAQLKNGVRGLMLDAYDFQNEVWLCHSFGGKCYNFA 150
Query: 153 AFQPAINVLKEIQVFLEANPSEIVTIIIEDYVTSPKGLTKVFDAAGLRKYWFPVSRMPKN 212
A+Q A++VLKEI FL+ANPSE++T+ +EDY P L K RMPK
Sbjct: 151 AYQRAMDVLKEIGAFLDANPSEVITVFVEDYA-GPGSLGK------------SGGRMPKG 197
Query: 213 GGDWPKVDDMVQKNQRLVVFTSKASKEASEGIAYEWRYLVENQYGNGGMKAGSCPNRAES 272
GGDWP + DM+ +N RL++FTSK K+ S+G+AYEW Y++E QYGN G+ GSCP RAES
Sbjct: 198 GGDWPLLKDMIAQNHRLLMFTSKRGKDGSDGLAYEWDYVLETQYGNDGLVGGSCPKRAES 257
Query: 273 PSMNTTSRSLVLVNFFRDLPDVAQSCKDNSAPLLSMVSTCNQAAGKRWPNFIAVDFYKRS 332
+M++T +SL+L+NFF P + +C +NSAPL++ + C A+ KRWPNFIAVD+Y RS
Sbjct: 258 MAMDSTKQSLILMNFFSTNPSQSWACGNNSAPLVAKLKACYDASAKRWPNFIAVDYYMRS 317
Query: 333 DGGGAPEAVDVANGHLVCGCGNIATCK 359
GGGAP A DVANG CGC +IA CK
Sbjct: 318 KGGGAPLATDVANGRQQCGCDSIAYCK 344
>UniRef100_Q9LMX9 F21F23.12 protein [Arabidopsis thaliana]
Length = 346
Score = 341 bits (874), Expect = 3e-92
Identities = 168/326 (51%), Positives = 218/326 (66%), Gaps = 11/326 (3%)
Query: 34 QEGQICVANKNCNSGLHCETCVTNGNVRPRCTRTQPINPTSKVKG-LPFNRYSWLTTHNS 92
Q G C ++++CN GL C C G RC R+ + S V +PFN+Y++LTTHNS
Sbjct: 31 QLGDQCSSDEDCNVGLGCFKC---GIDVARCVRSNITDQFSIVNNSMPFNKYAFLTTHNS 87
Query: 93 FALLGQKSMTGSVILSPTNQQDTITDQLNNGVRGLMLDLYDFENDVWLCHSFGGQCYNYT 152
+A+ G+ L Q+DTI QLN+GVR LMLD YD+E DVW CHSF QC+ +T
Sbjct: 88 YAIEGKA-------LHVATQEDTIVQQLNSGVRALMLDTYDYEGDVWFCHSFDEQCFEFT 140
Query: 153 AFQPAINVLKEIQVFLEANPSEIVTIIIEDYVTSPKGLTKVFDAAGLRKYWFPVSRMPKN 212
F AI+ KEI FL ANPSEIVT+I+EDYV S GLTKVF +GL+K+WFPV MP
Sbjct: 141 KFNRAIDTFKEIFAFLTANPSEIVTLILEDYVKSQNGLTKVFTDSGLKKFWFPVQNMPIG 200
Query: 213 GGDWPKVDDMVQKNQRLVVFTSKASKEASEGIAYEWRYLVENQYGNGGMKAGSCPNRAES 272
G DWP V DMV N RL+VFTS SK+ +EGIAY+W Y+VENQYG+ G+K C NRA+S
Sbjct: 201 GQDWPLVKDMVANNHRLIVFTSAKSKQETEGIAYQWNYMVENQYGDDGVKPDECSNRADS 260
Query: 273 PSMNTTSRSLVLVNFFRDLPDVAQSCKDNSAPLLSMVSTCNQAAGKRWPNFIAVDFYKRS 332
+ +++LV VN F+ +P +C++NS LL M+ TC AAG RW NF+AV+FYKRS
Sbjct: 261 ALLTDKTKALVSVNHFKTVPVKILTCEENSEQLLDMIKTCYVAAGNRWANFVAVNFYKRS 320
Query: 333 DGGGAPEAVDVANGHLVCGCGNIATC 358
+GGG +A+D NG L+CG ++ C
Sbjct: 321 NGGGTFQAIDKLNGELLCGRDDVHAC 346
>UniRef100_Q9FHA0 Similarity to MAP3K-like protein kinase [Arabidopsis thaliana]
Length = 365
Score = 260 bits (664), Expect = 6e-68
Identities = 120/204 (58%), Positives = 153/204 (74%), Gaps = 5/204 (2%)
Query: 52 ETCVTNGNVRPRCTRTQPINPTSKVKGLPFNRYSWLTTHNSFALLGQKSMTGSVILSPTN 111
ET + V P +P ++ GLPFN+Y+WL THN+F+ + G ++ N
Sbjct: 38 ETVLPLEEVNPSAQEAKP-----QINGLPFNKYTWLMTHNAFSNANAPLLPGVERITFYN 92
Query: 112 QQDTITDQLNNGVRGLMLDLYDFENDVWLCHSFGGQCYNYTAFQPAINVLKEIQVFLEAN 171
Q+DTIT+QL NGVRGLMLD+YDF ND+WLCHS GQC+N+TAFQPAIN+L+E++ FL N
Sbjct: 93 QEDTITNQLQNGVRGLMLDMYDFNNDIWLCHSLRGQCFNFTAFQPAINILREVEAFLSQN 152
Query: 172 PSEIVTIIIEDYVTSPKGLTKVFDAAGLRKYWFPVSRMPKNGGDWPKVDDMVQKNQRLVV 231
P+EIVTIIIEDYV PKGL+ +F AGL KYWFPVS+MP+ G DWP V DMVQ+N RL+V
Sbjct: 153 PTEIVTIIIEDYVHRPKGLSTLFANAGLDKYWFPVSKMPRKGEDWPTVTDMVQENHRLLV 212
Query: 232 FTSKASKEASEGIAYEWRYLVENQ 255
FTS A+KE EG+AY+WRY+VEN+
Sbjct: 213 FTSVAAKEDEEGVAYQWRYMVENE 236
>UniRef100_Q82NW8 Hypothetical protein [Streptomyces avermitilis]
Length = 464
Score = 106 bits (264), Expect = 1e-21
Identities = 85/300 (28%), Positives = 136/300 (45%), Gaps = 31/300 (10%)
Query: 67 TQPINPTSKVKGLPFNRYSWLTTHNSFALLGQKSMTGSVILSPTNQQDTITDQLNNGVRG 126
T +NP ++ ++LT HN++A + NQ I QL +GVRG
Sbjct: 170 TPSVNPMPSPDQRTLDQVTFLTAHNAYANGVDGGFAPPFVNLVPNQTRGINQQLTDGVRG 229
Query: 127 LMLDLYDFENDVWLCHSFGGQCYNYTAFQPAINVLKEIQV---FLEANPSEIVTIIIEDY 183
M+D++ + LCH+ + T + + +IQ FL+ +P ++VT+ +EDY
Sbjct: 230 FMMDIHQTSDGAILCHN------SCTLVSKPVALWVDIQRMVDFLKQHPDQVVTVFLEDY 283
Query: 184 VTSPKGLTKVFDAAGLRKYWFPVSRMPKNGGDWPKVDDMVQKNQRLVVFT--SKASKEA- 240
V +++ +GL + R WP++ D++ N RL++FT S++S E+
Sbjct: 284 VDPGVLRSELARVSGLSDVLYRPDRTGVRQSGWPRMADLIAANHRLLIFTDHSRSSDESA 343
Query: 241 -----SEGIAYEWRYLVENQYGNG---GMKAGSCPNRAESPSMN-------TTSRSLVLV 285
S G+ Y+ + VEN + G G SC +R N + R L ++
Sbjct: 344 GLTRDSFGVMYQREWTVENYWSMGSGLGSSDWSCYSRWYGADTNIPLTYTESAFRPLFVM 403
Query: 286 NFFRDLPDVAQSCKDNSAPLLSMVSTCNQAAGKRWPNFIAVDFYKRSDGGGAPEAVDVAN 345
N FRD + + DN+ C AA K+ PNF+AVD Y D G AVD N
Sbjct: 404 NHFRDAAIASTATTDNTKLADRAQRFCRPAARKK-PNFLAVDRY---DLGNPTSAVDTLN 459
>UniRef100_UPI00004301F4 UPI00004301F4 UniRef100 entry
Length = 360
Score = 76.3 bits (186), Expect = 2e-12
Identities = 68/280 (24%), Positives = 123/280 (43%), Gaps = 47/280 (16%)
Query: 81 FNRYSWLTTHNSFALLGQKSMTGSVILSPTNQQDTITDQLNNGVRGLMLDLYDFENDVWL 140
++ +++ H+S+A+ GS + +Q +T QLN+G+R L + ++ + + L
Sbjct: 35 YSNVTFIGAHDSYAV-------GSSVAD--DQDKDVTSQLNDGIRTLQIQAHNASDGIHL 85
Query: 141 CHSF-----GGQCYNYTAFQPAINVLKEIQVFLEANPSEIVTIIIEDYVT-SPKGLTKVF 194
CHS GG +Y L + ++ NP++++TI+I + P + VF
Sbjct: 86 CHSSCSLLDGGLMSDY---------LSTVASWVNDNPNDVITIVIVNSDNLPPTSFSPVF 136
Query: 195 DAAGLRKYWFPVSRMPKNGGDWPKVDDMVQKNQRLVVFTS-KASKEASEGIAYEWRYLVE 253
++AGL + + P DWP + DM+ +V F +A + + E+ + E
Sbjct: 137 ESAGLSSKVYTPASQPTQLSDWPSLSDMIDAGTTVVAFMDYEADTSSVPYLLDEFAAMWE 196
Query: 254 NQYGNGGMKAGSCPNRAESPSMNTTSRSLVLVNFFRD-----------LPDVAQ----SC 298
+ YG + G NR S TS L+N F D +P+ + +
Sbjct: 197 DAYGVTTQEFGCAVNR----SSGDTSSQPFLINHFLDSTYSFSSIQVFVPNKDKLNETNA 252
Query: 299 KDNSAPLLSMVSTCNQAAGKRWPNFIAVDFYKRSDGGGAP 338
+ + + V+ C Q G+ PN I +DFY G +P
Sbjct: 253 ETGTGSIGYHVNNCRQLWGRN-PNHILLDFY--DSNGNSP 289
>UniRef100_UPI00003C1EBF UPI00003C1EBF UniRef100 entry
Length = 378
Score = 72.8 bits (177), Expect = 2e-11
Identities = 63/278 (22%), Positives = 120/278 (42%), Gaps = 40/278 (14%)
Query: 81 FNRYSWLTTHNSFALLGQKSMTGSVILSPTNQQDTITDQLNNGVRGLMLDLYDFEND--- 137
++ +++ HNS+A+ ++ G+ + NQ+ ++T QL +G+R L + + N
Sbjct: 54 YSNVTYIGAHNSYAV---GTLAGASV--GKNQEQSVTQQLTDGIRLLQVQAHKSSNSTSG 108
Query: 138 --VWLCHSF-----GGQCYNYTAFQPAINVLKEIQVFLEANPSEIVTIIIEDYVTSP-KG 189
+ LCHS GG NY L +++ ++++NP++++TI+I + P
Sbjct: 109 SGINLCHSSCQIENGGTLENY---------LSKVKTWVDSNPNDVITILIVNSDNQPVSS 159
Query: 190 LTKVFDAAGLRKYWFPVSRMPKNGGDWPKVDDMVQKNQRLVVFTSKASKEAS-EGIAYEW 248
F + GL + WP + ++ + LVVF ++ +S I +
Sbjct: 160 FGTAFQSTGLASKAYSPGTAALAKDSWPTLGSLIDSGKNLVVFIDNSADVSSVPYILPHF 219
Query: 249 RYLVENQYGNGGMKAGSCPNRAESPSMNTTSRSLVLVNFFRD-----------LPDVAQS 297
+ EN Y + +R S S S + L+N + D +P+ AQ
Sbjct: 220 QNTWENPYNQISVPFNCSVDRINSGS--EPSNLMYLINHYLDSSFNLFGTTVFVPNTAQL 277
Query: 298 CKDNS-APLLSMVSTCNQAAGKRWPNFIAVDFYKRSDG 334
NS + +++ C G +P ++ DFY DG
Sbjct: 278 NTTNSLSSIMTDAGNCASLHGTGYPTYVLTDFYDVGDG 315
>UniRef100_UPI0000235E3F UPI0000235E3F UniRef100 entry
Length = 381
Score = 70.1 bits (170), Expect = 1e-10
Identities = 81/263 (30%), Positives = 114/263 (42%), Gaps = 43/263 (16%)
Query: 111 NQQDTITDQLNNGVRGLMLDLYDFENDVWLCHSFGGQCYNYTAFQPAINVLKEIQVFLEA 170
NQ T QL+ GVR + ++D ++ LCHS C A + L EI+ +L++
Sbjct: 68 NQYYNTTVQLDAGVRLVSAQVHDSDSQWRLCHS---SCDLLDAGRLR-TWLSEIKSWLDS 123
Query: 171 NPSEIVTIII--EDYVTSPKGLTKVFDAAGLRKY-WFPVSRMPKNGGDWPKVDDMVQKNQ 227
NP+E+VT+++ D T+ L F+AA + Y + P S+ + WP + +++
Sbjct: 124 NPNEVVTVLLVNSDGATA-SDLAAEFEAADITDYAYTPTSQSAPS--SWPTLQELIDAGT 180
Query: 228 RLVVFTSKASKEASEGIAY---EWRYLVENQYGNGGMKAGSC-PNRAESPSMNTTS---- 279
RL+ F AS + + G Y E+ Y+ EN Y SC +R S N S
Sbjct: 181 RLMTFV--ASLDDNSGATYLMDEFTYIFENSYDVTSPSNFSCTADRPSSVKGNAASAISA 238
Query: 280 RSLVLVNFFRDLPDVAQSCKDNSAPLLSMVSTCN----------------QAAGKRWPNF 323
L L N F D AP S V T N Q A R P F
Sbjct: 239 NMLPLQNHFL----YQTILLDYQAPNESYVGTTNAPSGGEGNLGDAASTCQTAWGRQPAF 294
Query: 324 IAVDFYKRSDGGGAPEAVDVANG 346
I VDF+ D G A E VD NG
Sbjct: 295 ILVDFF---DKGPAIETVDKLNG 314
>UniRef100_Q7RXX2 Hypothetical protein [Neurospora crassa]
Length = 396
Score = 69.7 bits (169), Expect = 1e-10
Identities = 81/298 (27%), Positives = 125/298 (41%), Gaps = 46/298 (15%)
Query: 81 FNRYSWLTTHNSFALLGQKSMTGSVILSPTNQQDTITDQLNNGVRGLMLDLYDFENDVWL 140
+N + + H+S + L S + S+ NQ T L+ G+R L ++D + L
Sbjct: 79 YNNVTHMGAHDS-SFLRDASTSDSLA---GNQYFNATVALDAGIRLLQAQVHDVNGTLQL 134
Query: 141 CHSFGGQCYNYTAFQPAINVLKEIQVFLEANPSEIVTIIIEDYVTSPKGLTK----VFDA 196
CH+ C A P + L +I+ +++ NP+E+VTI++ V S L VF+
Sbjct: 135 CHT---SCSLLDA-GPLQDWLAKIKFWMDNNPNEVVTILL---VNSDNKLVSDYAAVFEG 187
Query: 197 AGLRKYWFPVSRMPKNGGDWPKVDDMVQKNQRLVVFTSKASKEASEGIAY---EWRYLVE 253
+G+ Y + +S WP + DM+ N+RLV F AS + S Y E+ ++ E
Sbjct: 188 SGISTYGYQLSNGSSASNTWPTLGDMITSNKRLVTFI--ASIDYSPTYPYLLSEFDHVFE 245
Query: 254 NQYG--------------NGGMKAGSCPNRAESPSMNTTSRSLVLVNFFRDLPDVAQSCK 299
Y G AG + P MN + SL+L +PD
Sbjct: 246 TAYNVLSLSGFNCTLDRPKGQGSAGDAISAGLMPLMNHFADSLLLQGV--QIPDETDIDI 303
Query: 300 DNSAPLLSM------VSTCNQAAGKRWPNFIAVDFYKRSDGGGAPEAVDVANGHLVCG 351
NS + TC + G + P FI VDF+ D G A + D NG G
Sbjct: 304 TNSPDTSTTGNLGLHADTCVKQWGVK-PTFILVDFF---DHGPAIDTADRLNGITATG 357
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.321 0.136 0.420
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 716,852,139
Number of Sequences: 2790947
Number of extensions: 30203689
Number of successful extensions: 64056
Number of sequences better than 10.0: 53
Number of HSP's better than 10.0 without gapping: 20
Number of HSP's successfully gapped in prelim test: 33
Number of HSP's that attempted gapping in prelim test: 63944
Number of HSP's gapped (non-prelim): 102
length of query: 425
length of database: 848,049,833
effective HSP length: 130
effective length of query: 295
effective length of database: 485,226,723
effective search space: 143141883285
effective search space used: 143141883285
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 76 (33.9 bits)
Lotus: description of TM0227.11


