

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0226.7
(399 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
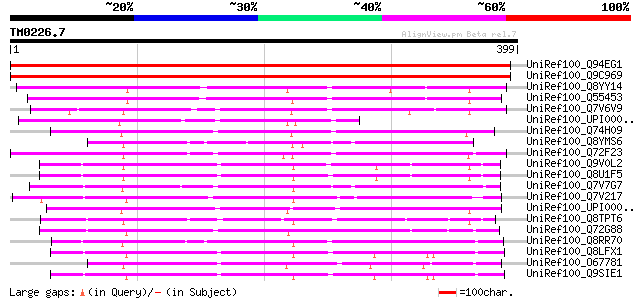
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q94EG1 Putative aspartate aminotransferase [Oryza sativa] 618 e-175
UniRef100_Q9C969 Putative aspartate aminotransferase; 38163-3625... 585 e-166
UniRef100_Q8YY14 Alr1039 protein [Anabaena sp.] 220 5e-56
UniRef100_Q55453 Aspartate aminotransferase [Synechocystis sp.] 199 1e-49
UniRef100_Q7V6V9 Aminotransferases class-I [Prochlorococcus mari... 189 2e-46
UniRef100_UPI0000349D32 UPI0000349D32 UniRef100 entry 166 8e-40
UniRef100_Q74H09 Aminotransferase, classes I and II [Geobacter s... 147 4e-34
UniRef100_Q8YMS6 Aspartate aminotransferase [Anabaena sp.] 147 5e-34
UniRef100_Q72F23 Aromatic aminotransferase [Desulfovibrio vulgaris] 144 6e-33
UniRef100_Q9V0L2 Aspartate aminotransferase [Pyrococcus abyssi] 143 7e-33
UniRef100_Q8U1F5 Aspartate transaminase [Pyrococcus furiosus] 143 1e-32
UniRef100_Q7V7G7 Aminotransferases class-I [Prochlorococcus mari... 142 2e-32
UniRef100_Q7V217 Aminotransferases class-I [Prochlorococcus mari... 141 3e-32
UniRef100_UPI000033BD5F UPI000033BD5F UniRef100 entry 140 5e-32
UniRef100_Q8TPT6 Aspartate aminotransferase [Methanosarcina acet... 140 6e-32
UniRef100_Q72G88 Aspartate aminotransferase [Thermus thermophilus] 140 8e-32
UniRef100_Q8RR70 Aspartate aminotransferase [Phormidium lapideum] 140 8e-32
UniRef100_Q8LFX1 Putative aspartate aminotransferase [Arabidopsi... 140 8e-32
UniRef100_O67781 Aspartate aminotransferase [Aquifex aeolicus] 140 8e-32
UniRef100_Q9SIE1 Putative aspartate aminotransferase [Arabidopsi... 139 1e-31
>UniRef100_Q94EG1 Putative aspartate aminotransferase [Oryza sativa]
Length = 394
Score = 618 bits (1593), Expect = e-175
Identities = 285/394 (72%), Positives = 355/394 (89%)
Query: 1 MGSYANLARRALETEMPIMVQMQEVLRGAKNCVSLAQGVVYWQPPKPALDKVKELVWEPS 60
MGS+ LARRA+ETE P+MV+MQE+LRG K+ +SLAQGVVYWQPP+ A++K+KE+VWEPS
Sbjct: 1 MGSFGRLARRAVETEAPVMVKMQELLRGNKDVMSLAQGVVYWQPPEAAMNKIKEIVWEPS 60
Query: 61 VSRYGADEGLPELRAALVKKLRQENNLHKSSVMVTAGANQAFVNLVLTLCDAGDSVVMFA 120
+S+YG+D+GLPELR AL++KLR+EN L KSS+MVT+GANQAFVN+VLTLCDAGD+VVMFA
Sbjct: 61 ISKYGSDDGLPELREALLEKLRRENKLTKSSIMVTSGANQAFVNVVLTLCDAGDAVVMFA 120
Query: 121 PYYFNAYMSFQMTGITNIHVGPGSPDTLYPDVDWLERILTESKPVPKLVTVVNPGNPTGT 180
PYYFN+YMSFQMTG+T+I VG +P+TL+PDVDWLE++L E+ P+PKLV+VVNPGNP+G
Sbjct: 121 PYYFNSYMSFQMTGVTDILVGASNPETLHPDVDWLEKVLQENNPIPKLVSVVNPGNPSGA 180
Query: 181 YIPESLLQRIANLCKNAGSWLLVDNTYEYFMFDGLKHSCVEGNHIVNIFSFSKAYGMMGW 240
+IP+ +L+RI+ LC+NAG+WL+VDNTYEYFM+DG++H C+EGNHIVN+FSFSKAYGMMGW
Sbjct: 181 FIPKPMLERISELCRNAGAWLVVDNTYEYFMYDGMEHYCLEGNHIVNLFSFSKAYGMMGW 240
Query: 241 RVGYIAYPSEVEGLSAQLLKVQDNIPICASIISQHLALHSLEMGPEWVRERVQTLAKNRE 300
RVGYIA+P+E +GL AQLLKVQDNIPICASII Q LAL++LE GPEW+RERV+ L KNRE
Sbjct: 241 RVGYIAHPNEADGLHAQLLKVQDNIPICASIIGQRLALYALEAGPEWIRERVRDLVKNRE 300
Query: 301 IVSEALSPLGEGSVKGGEGAIYLFAKLPKGGYDDFDVVRWLANRHGIAVIPGSACGAAGN 360
++ EA+SPLG+ SVKGGEGAIYL+AKLP+ DDF+VVRWLAN+HG+AVIPGSA G G
Sbjct: 301 LLMEAMSPLGKDSVKGGEGAIYLWAKLPEKCSDDFEVVRWLANKHGVAVIPGSASGGPGY 360
Query: 361 LRISFGGLTESDCRAAAERLKKGLEELVANGLVQ 394
+R+SFGGL ESD R AAERL++GL+ELV G+VQ
Sbjct: 361 IRVSFGGLKESDTRLAAERLRRGLQELVTEGMVQ 394
>UniRef100_Q9C969 Putative aspartate aminotransferase; 38163-36256 [Arabidopsis
thaliana]
Length = 394
Score = 585 bits (1508), Expect = e-166
Identities = 274/394 (69%), Positives = 335/394 (84%)
Query: 1 MGSYANLARRALETEMPIMVQMQEVLRGAKNCVSLAQGVVYWQPPKPALDKVKELVWEPS 60
MGS+ L+RR L T+MP+M Q++ ++ N +SLAQGVV+WQPP+ AL+KVKELVW+P
Sbjct: 1 MGSFGMLSRRTLGTDMPVMAQIRSLMAELTNPMSLAQGVVHWQPPQKALEKVKELVWDPI 60
Query: 61 VSRYGADEGLPELRAALVKKLRQENNLHKSSVMVTAGANQAFVNLVLTLCDAGDSVVMFA 120
+S YG DEGLPELR AL+KKLR+EN L S VMVTAGANQAFVNLV+TLCDAGDSVVMF
Sbjct: 61 ISSYGPDEGLPELRQALLKKLREENKLTNSQVMVTAGANQAFVNLVITLCDAGDSVVMFE 120
Query: 121 PYYFNAYMSFQMTGITNIHVGPGSPDTLYPDVDWLERILTESKPVPKLVTVVNPGNPTGT 180
PYYFN+YM+FQMTG+TNI VGPG DTLYPD DWLER L+ESKP PK+VTVVNPGNP+GT
Sbjct: 121 PYYFNSYMAFQMTGVTNIIVGPGQSDTLYPDADWLERTLSESKPTPKVVTVVNPGNPSGT 180
Query: 181 YIPESLLQRIANLCKNAGSWLLVDNTYEYFMFDGLKHSCVEGNHIVNIFSFSKAYGMMGW 240
Y+PE LL+RIA +CK+AG WL+VDNTYEYFM+DGLKH CVEG+HIVN+FSFSK YGMMGW
Sbjct: 181 YVPEPLLKRIAQICKDAGCWLIVDNTYEYFMYDGLKHCCVEGDHIVNVFSFSKTYGMMGW 240
Query: 241 RVGYIAYPSEVEGLSAQLLKVQDNIPICASIISQHLALHSLEMGPEWVRERVQTLAKNRE 300
R+GYIAY ++G + +L+K+QDNIPICA+IISQ LA+++LE G W+ ERV++L KNR+
Sbjct: 241 RLGYIAYSERLDGFATELVKIQDNIPICAAIISQRLAVYALEEGSGWITERVKSLVKNRD 300
Query: 301 IVSEALSPLGEGSVKGGEGAIYLFAKLPKGGYDDFDVVRWLANRHGIAVIPGSACGAAGN 360
IV EAL PLG+ +VKGGEGAIYL+AKLP+G DDF VVRWLA+RHG+ VIPG A G+ G
Sbjct: 301 IVKEALEPLGKENVKGGEGAIYLWAKLPEGHRDDFKVVRWLAHRHGVVVIPGCASGSPGY 360
Query: 361 LRISFGGLTESDCRAAAERLKKGLEELVANGLVQ 394
LR+SFGGL E + RAAA RL+KG+EEL+ +G+V+
Sbjct: 361 LRVSFGGLQEVEMRAAAARLRKGIEELLHHGMVE 394
>UniRef100_Q8YY14 Alr1039 protein [Anabaena sp.]
Length = 398
Score = 220 bits (561), Expect = 5e-56
Identities = 132/404 (32%), Positives = 214/404 (52%), Gaps = 27/404 (6%)
Query: 6 NLARRALETEMPIMVQMQEVLRGAKNCVSLAQGVVYWQPPKPALDKVKELVWEPSVSRYG 65
N R + PI+ + E+++ + +SL QGVV + PP A++ + + +P+ + Y
Sbjct: 3 NFTSRMQAVQSPIIPVVGELIQNSPGTISLGQGVVSYSPPPEAIELLPRFLADPANNLYK 62
Query: 66 ADEGLPELRAALVKKLRQENNLHKSS---VMVTAGANQAFVNLVLTLCDAGDSVVMFAPY 122
A EG+P L AL +KL NN+ ++ ++VTAG+N AF+N +L + GD +++ PY
Sbjct: 63 AVEGIPPLLNALTEKLSTFNNIEITTDNCIVVTAGSNMAFMNAILAITSPGDEIILNTPY 122
Query: 123 YFNAYMSFQMTGITNIHVGPGSPDTLYPDVDWLERILTESKPVPKLVTVVNPGNPTGTYI 182
YFN M+ M G + V L P E I P + V ++P NPTG
Sbjct: 123 YFNHEMAITMAGCRAVLVETDENYQLCP-----EAIAQAITPKTRAVVTISPNNPTGVVY 177
Query: 183 PESLLQRIANLCKNAGSWLLVDNTYEYFMFDGLKH-----SCVEGNHIVNIFSFSKAYGM 237
E LL+ + +C N G + + D YEYF +DG+KH + ++++S SKAYG
Sbjct: 178 CEDLLRNVNQICANYGIYHISDEAYEYFTYDGVKHVSPASFAGSSEYTISLYSLSKAYGF 237
Query: 238 MGWRVGYIAYPSEVEGLSAQLLKVQDNIPICASIISQHLALHSLEMGPEWVRERVQTLAK 297
WR+GY+ P L + KVQD I IC ++SQ+ AL +L+ PE++++ + LA+
Sbjct: 238 ASWRIGYMVIPKH---LLVAIKKVQDTILICPPVVSQYAALGALQAKPEYLQDHIGALAQ 294
Query: 298 N-------REIVSEALSPL-GEGSVKGGEGAIYLFAKLPKGGYDDFDVVRWLANRHGIAV 349
R+IV + L L G ++ +GA Y+F K+ D F +V+ L + +AV
Sbjct: 295 PAVGIAQVRQIVFDYLKQLQGLCNITPADGAFYVFLKV-HTQIDAFALVKQLIQEYKVAV 353
Query: 350 IPGSACGAAGN--LRISFGGLTESDCRAAAERLKKGLEELVANG 391
IPG+ G LR+++G L + + ERL +GL+ ++ G
Sbjct: 354 IPGTTFGMENGCYLRVAYGALQKDTVKEGIERLVQGLKTILDTG 397
>UniRef100_Q55453 Aspartate aminotransferase [Synechocystis sp.]
Length = 389
Score = 199 bits (505), Expect = 1e-49
Identities = 122/384 (31%), Positives = 204/384 (52%), Gaps = 20/384 (5%)
Query: 15 EMPIMVQMQEVLRGAKNCVSLAQGVVYWQPPKPALDKVKELVWEPSVSRYGADEGLPELR 74
+ P++ + + + + +SL QGV ++ PP V+E + + + +Y G+P L
Sbjct: 14 QSPVIPVVGQWIVDSPGTISLGQGVAFYPPPNEVAVAVRESLEQTPLHQYQPVAGIPSLI 73
Query: 75 AALVKKLRQENNLHKSS---VMVTAGANQAFVNLVLTLCDAGDSVVMFAPYYFNAYMSFQ 131
+AL +KLR++N+++ SS V+VTAGAN F+N VL + + GD +++ PYYFN M+ +
Sbjct: 74 SALTEKLRRDNDINLSSDQAVVVTAGANMGFLNAVLAITEVGDEIILNTPYYFNHEMAVR 133
Query: 132 MTGITNIHVGPGSPDTLYPDVDWLERILTESKPVPKLVTVVNPGNPTGTYIPESLLQRIA 191
+ G + V L L+ I P + V ++P NPTG PE+ L+ +
Sbjct: 134 IAGCQPVLVPTDDQYQLQ-----LDLIAQAIAPRTRAVVTISPNNPTGAIYPEADLRAVN 188
Query: 192 NLCKNAGSWLLVDNTYEYFMFDGLKHSCVE-----GNHIVNIFSFSKAYGMMGWRVGYIA 246
LC+ G + + D Y+YF +D E G H ++++SFSKAYGM GWRVGY+
Sbjct: 189 QLCQERGIYHIHDEAYDYFAYDQTPIFSPEAMGDSGGHTISLYSFSKAYGMAGWRVGYMV 248
Query: 247 YPSEVEGLSAQLLKVQDNIPICASIISQHLALHSLEMGPEWVRERVQTLAKNREIVSEAL 306
P E L + K+QD IC ++SQ+ AL L +G + + + +A R+ + E L
Sbjct: 249 IPLE---LLLAVKKIQDTNLICPVVVSQYAALACLRVGKNYSAQFLPEMAACRQQLLETL 305
Query: 307 SPLGE-GSVKGGEGAIYLFAKLPKGGYDDFDVVRWLANRHGIAVIPGSACGAAGN--LRI 363
L + + +GA Y ++ D ++V+ L + +AV+PGS G LRI
Sbjct: 306 GQLSDYCRLVVPQGAFYCLLEV-NSPLTDLELVKRLIDEFKVAVLPGSTFGVDSGCYLRI 364
Query: 364 SFGGLTESDCRAAAERLKKGLEEL 387
++G L ++ A RL++GL+ +
Sbjct: 365 AYGALRQATASVAIARLEQGLKSI 388
>UniRef100_Q7V6V9 Aminotransferases class-I [Prochlorococcus marinus]
Length = 404
Score = 189 bits (479), Expect = 2e-46
Identities = 130/391 (33%), Positives = 198/391 (50%), Gaps = 28/391 (7%)
Query: 17 PIMVQMQEVLRGAKNCVSLAQGVVYWQPP---KPALDKVKELVWEPSVSRYGADEGLPEL 73
P++ + E++ +SLAQG+V W PP K A++ L E S++RYG G P+L
Sbjct: 22 PVIPALNELVAKTPGTLSLAQGMVNWPPPIAVKLAMNNAL-LNQESSLNRYGPARGDPDL 80
Query: 74 RAALVKKLRQENNLH--KSSVMVTAGANQAFVNLVLTLCDAGDSVVMFAPYYFNAYMSFQ 131
+ +KL +N L +S VMVTAG+N AF + LCD GD V++ PYYFN +M+ Q
Sbjct: 81 LELIKQKLMMQNGLDLAESMVMVTAGSNMAFHAIAQVLCDPGDEVILPLPYYFNHFMAIQ 140
Query: 132 MTGITNIHVGPGSPDTLYPDVDWLERILTESKPVPKLVTVVNPGNPTGTYIPESLLQRIA 191
+ G + V G L P+ +E +T+ + + ++P NP+G P++LL I
Sbjct: 141 LAGGVPVPVNAG----LIPNPGLIEAAITKR---TRAIVTISPNNPSGIVFPQTLLAAIN 193
Query: 192 NLCKNAGSWLLVDNTYEYFMFDGLKHSCV-----EGNHIVNIFSFSKAYGMMGWRVGYIA 246
+C G + D YE F+F + H GNH V+++SFSKAYGM GWR+GY++
Sbjct: 194 RICAQHGLLHISDEAYEDFVFGDVPHWSPGSLPGAGNHTVSLYSFSKAYGMAGWRLGYMS 253
Query: 247 YPSEVEGLSAQLLKVQDNIPICASIISQHLALHSLEMGPEWVRERVQTLAKNREIVSEAL 306
P S L KVQD + IC Q A+ +L G +WVR+ V L +++ +
Sbjct: 254 VPIV---WSKALEKVQDTVLICPPRFCQRAAIAALLDGSDWVRQNVSQLMSRYQLLLKRF 310
Query: 307 SPLGEGS---VKGGEGAIYLFAKLPKGGYDDFDVVRWLANRHGIAVIPGSACGAAGN--- 360
+ + + GA Y ++ G D ++R L + +A I G + G
Sbjct: 311 AASNDRPWRFLHEPNGAFYCLLEVDCGCNGD-TLMRQLVRDYRVATIGGCSFGFKNESCV 369
Query: 361 LRISFGGLTESDCRAAAERLKKGLEELVANG 391
LRIS G L ++ A RL+ G+ V G
Sbjct: 370 LRISVGMLEGAELIDAFNRLEAGMLNAVQQG 400
>UniRef100_UPI0000349D32 UPI0000349D32 UniRef100 entry
Length = 271
Score = 166 bits (421), Expect = 8e-40
Identities = 94/275 (34%), Positives = 153/275 (55%), Gaps = 15/275 (5%)
Query: 8 ARRALETEMPIMVQMQEVLRGAKNCVSLAQGVVYWQPPKPALDKVKELVWEPSVSRYGAD 67
++R + PI+ + +++ +SL QG+V++ PP +++ +P+ +YG
Sbjct: 5 SKRMASVQQPIIPVIGDLILANPGTISLGQGIVHYPPPTEVESTLRDFWSDPTAHQYGPV 64
Query: 68 EGLPELRAALVKKLRQEN--NLHKSSVMVTAGANQAFVNLVLTLCDAGDSVVMFAPYYFN 125
EGL LR+AL +KL QEN L V+VTAG+N F+N++L + D GD +++ P+YFN
Sbjct: 65 EGLDPLRSALAEKLEQENAIRLEGRGVVVTAGSNMGFLNVLLAISDPGDEIILPEPFYFN 124
Query: 126 AYMSFQMTGITNIHVGPGSPDTLYPDVDWLERILTESKPVPKLVTVVNPGNPTGTYIPES 185
M+ ++ I V + D + +D +E +T+ + V V+P NPTG PE+
Sbjct: 125 QEMAVRIADC--IPVTVPTDDAGHLQIDAIESAITKR---TRAVVTVSPNNPTGIVYPEA 179
Query: 186 LLQRIANLCKNAGSWLLVDNTYEYFMFDGLKH---SCVEGN--HIVNIFSFSKAYGMMGW 240
L+ + C+ G + + D YEYF++D +H +E + H + ++S SKAYG W
Sbjct: 180 DLRAVNEACRARGIYHISDEAYEYFLYDDAEHFSPGTIEQSAPHTICLYSLSKAYGFASW 239
Query: 241 RVGYIAYPSEVEGLSAQLLKVQDNIPICASIISQH 275
R+GY P E+ G L KVQD ICA+ ISQ+
Sbjct: 240 RIGYAVIPEELFG---ALRKVQDTNVICATAISQY 271
>UniRef100_Q74H09 Aminotransferase, classes I and II [Geobacter sulfurreducens]
Length = 391
Score = 147 bits (372), Expect = 4e-34
Identities = 98/358 (27%), Positives = 168/358 (46%), Gaps = 15/358 (4%)
Query: 33 VSLAQGVVYWQPPKPALDKVKELVWEPSVSRYGADEGLPELRAALVKKLRQENN--LHKS 90
V L Q V + P + D + L+ +P VS+Y DEGLPE+R + + + ++
Sbjct: 36 VDLCQAVPDYPPARQLTDYLAALLDDPLVSKYSPDEGLPEVREGVCARYGRVYGAAMNPD 95
Query: 91 SVMVTAGANQAFVNLVLTLCDAGDSVVMFAPYYFNAYMSFQMTGITNIHVGPGSPDTLYP 150
+ +T GA+QAF ++TLC AGD V++ P YF+ M+ + G+ +++ P
Sbjct: 96 QLCLTIGASQAFWLAMVTLCRAGDEVIVPLPAYFDHPMALDILGVRPVYLPFDEERGGVP 155
Query: 151 DVDWLERILTESKPVPKLVTVVNPGNPTGTYIPESLLQRIANLCKNAGSWLLVDNTYEYF 210
D +ER++T P + + +V P NPTG P +Q + + + G L++D TY F
Sbjct: 156 DPAAVERLIT---PRTRAILLVTPSNPTGVVTPPETIQELHGVARRRGIALVLDETYADF 212
Query: 211 MFDGLKHSCV-----EGNHIVNIFSFSKAYGMMGWRVGYIAYPSEVEGLSAQLLKVQDNI 265
+ G + + G+H++++ SF K Y + G+R G +A E G LK QD +
Sbjct: 213 IPGGERPHDLFLDPRWGDHLIHLMSFGKTYALTGYRAGCLAASKEFIG---HALKAQDTM 269
Query: 266 PICASIISQHLALHSLEMGPEWVRERVQTLAKNREIVSEALSPLGEGSVKGGEGAIYLFA 325
+C I+Q+ L+ + WV E + + ++ + G G + +
Sbjct: 270 AVCQPRITQYAVLYGVSHLDGWVEENRLMMTRRHDLFRSLFTRPGNPFSLVASGTFFAWV 329
Query: 326 KLPKGGYDDFDVVRWLANRHGIAVIPGSACGAA--GNLRISFGGLTESDCRAAAERLK 381
+ P + R LA GI +PG G LR++FG + + A ER +
Sbjct: 330 RHPLQEGTGREAARRLAVEAGIICLPGEVFGPGLEPYLRLAFGNIRDEAIPGAVERFR 387
>UniRef100_Q8YMS6 Aspartate aminotransferase [Anabaena sp.]
Length = 388
Score = 147 bits (371), Expect = 5e-34
Identities = 100/313 (31%), Positives = 164/313 (51%), Gaps = 16/313 (5%)
Query: 62 SRYGADEGLPELRAALVKKLRQENNLH--KSSVMVTAGANQAFVNLVLTLCDAGDSVVMF 119
++YGA G P+LR A+ +KL+++N+L +V+VT G + NL++ L D GD V++
Sbjct: 61 TKYGAAAGEPKLREAIARKLQKDNHLDYKPENVIVTNGGKHSLYNLIVALIDPGDEVIIP 120
Query: 120 APYYFNAYMSFQMTGITNIHVGPGSPDTLYPDVDWLERILTESKPVPKLVTVVNPGNPTG 179
APY+ + + G ++ V P T Y E++ P KL + +P NPTG
Sbjct: 121 APYWLSYPEMVTLVGGKSVIV-PTDASTGYKITP--EQLRKAITPKTKLFVLNSPSNPTG 177
Query: 180 -TYIPESLLQRIANLCKNAGSWLLVDNTYEYFMFDGLKHSCVE--GNHIVNIF----SFS 232
Y PE + + +A + +A +++ D YE ++DG +H + G I N F+
Sbjct: 178 MVYTPEEI-KALAQVVVDADIYVVSDEIYEKILYDGAQHISIGSLGKEIFNRTLISNGFA 236
Query: 233 KAYGMMGWRVGYIAYPSEVEGLSAQLLKVQDNIPICASIISQHLALHSLEMGPEWVRERV 292
KAY M GWR+GY+A P ++ +A ++ +C +Q+ A+ +LE + V E
Sbjct: 237 KAYSMTGWRLGYLAGPVDIIK-AASSIQGHSTSNVCT--FAQYGAIAALEDSQDCVEEMR 293
Query: 293 QTLAKNREIVSEALSPLGEGSVKGGEGAIYLFAKLPKGGYDDFDVVRWLANRHGIAVIPG 352
Q AK R+++ + L+ + S +GA YLF + K G + L H +AVIPG
Sbjct: 294 QAFAKRRQVMLDRLNAIPGLSTAKPDGAFYLFPDISKTGLKSLEFCDALIEEHKVAVIPG 353
Query: 353 SACGAAGNLRISF 365
A GA N+R+S+
Sbjct: 354 IAFGADDNIRLSY 366
>UniRef100_Q72F23 Aromatic aminotransferase [Desulfovibrio vulgaris]
Length = 399
Score = 144 bits (362), Expect = 6e-33
Identities = 118/403 (29%), Positives = 185/403 (45%), Gaps = 20/403 (4%)
Query: 1 MGSYANLARRALETEMPIMVQMQEVLRGAKNCVSLAQGVVYWQPPKPALDKV-KELVWEP 59
M ++RR + + M + CVSL QGV ++ P+ ++ V + L +
Sbjct: 1 MSDVFRVSRRVQQIRISATKLMPMLAARVGGCVSLGQGVPSFRTPEHIVEAVCRALRDKA 60
Query: 60 SVSRYGADEGLPELRAALVKKLRQENNLH---KSSVMVTAGANQAFVNLVLTLCDAGDSV 116
RY G+P LR A+ L S V VT GA +A + +LT+ D GD V
Sbjct: 61 DAGRYTLQPGMPALREAIAADLAARKGYMVNPDSEVGVTVGAMEALLMALLTVVDRGDEV 120
Query: 117 VMFAPYYFNAYMSFQMTGITNIHVGPGSPDTLYPDVDWLERILTESKPVPKLVTVVNPGN 176
++ +P Y + M +HV P D DVD + +T P + V V NPGN
Sbjct: 121 IIPSPGYASHAEQVLMAEGVPVHV-PLRADDWGLDVDAIRAAVT---PRTRAVIVCNPGN 176
Query: 177 PTGTYIPESLLQRIANLCKNAGSWLLVDNTYEYFMFDG---LKHSCVE--GNHIVNIFSF 231
PTGT ++ ++ + L L+ D TY+Y ++ G L + + H++ + SF
Sbjct: 177 PTGTVYDDADVRALCELALERNIMLISDETYDYMVYGGGEPLSPASLPEMRRHVIVVNSF 236
Query: 232 SKAYGMMGWRVGYIAYPSEVEGLSAQLLKVQDNIPICASIISQHLALHSLEMGPEWVRER 291
SK Y + GWRVGY A + G +LLKV D ICA +SQ+ AL +L + V +
Sbjct: 237 SKKYALTGWRVGYCAADAAWMG---ELLKVHDAAAICAPAVSQYAALAALTGPQDCVDDM 293
Query: 292 VQTLAKNREIVSEALSPLG-EGSVKGGEGAIYLFAKLPKGGYDDFDVVRWLANRHGIAVI 350
L+ R + L + GA Y+ A+ V R L + +
Sbjct: 294 RAALSARRNLACARLDAMAPHFDYVQPRGAFYIMARYTFTDAPSDMVARRLLEEGRVITV 353
Query: 351 PGSACGAAG--NLRISFGGLTESDCRAAAERLKKGLEELVANG 391
PG++ G G +LR+SF G+ E++ A +R+ + +VA+G
Sbjct: 354 PGASFGPTGERHLRLSF-GMEEAELDEAFDRMAAWTQRVVASG 395
>UniRef100_Q9V0L2 Aspartate aminotransferase [Pyrococcus abyssi]
Length = 389
Score = 143 bits (361), Expect = 7e-33
Identities = 104/375 (27%), Positives = 186/375 (48%), Gaps = 21/375 (5%)
Query: 24 EVLRGAKNCVSLAQGVVYWQPPKPALDKVKELVWEPSVSRYGADEGLPELRAALVKKLRQ 83
++ G K+ +SL G + P+ + KE + + ++ YG + GLPELR A+ +KL++
Sbjct: 20 DIAAGMKDVISLGIGEPDFDTPQHIKEYAKEAL-DMGLTHYGPNIGLPELREAIAEKLKK 78
Query: 84 ENNLH---KSSVMVTAGANQAFVNLVLTLCDAGDSVVMFAPYYFNAYMSFQMTGITNIHV 140
+NN+ +MV GANQAF+ + G+ V++ P + + + + G + V
Sbjct: 79 QNNIEADPNKEIMVLVGANQAFLMGLSAFLKDGEEVLIPTPAFVSYAPAVILAGGKPVEV 138
Query: 141 GPGSPDTLYPDVDWLERILTESKPVPKLVTVVNPGNPTGTYIPESLLQRIANLCKNAGSW 200
+ +VD L++ +TE K + + +P NPTG+ + + L+ IA+
Sbjct: 139 PTYEENEFRLNVDELKKYVTEKT---KALIINSPCNPTGSVLKKKDLEEIADFAVEHDLI 195
Query: 201 LLVDNTYEYFMFDGLKHSCVEG-----NHIVNIFSFSKAYGMMGWRVGYIAYPSEVEGLS 255
++ D YE+F++D +KH + + + FSK + M GWR+G++A PS +
Sbjct: 196 VISDEVYEHFIYDDVKHYSIASLDGMFERTITVNGFSKTFAMTGWRLGFVAAPS---WII 252
Query: 256 AQLLKVQDNIPICASIISQHLALHSLEMGPEW--VRERVQTLAKNREIVSEALSPLGEGS 313
+++K Q C Q+ A +L W V E + + R++V + L+ +G +
Sbjct: 253 EKMVKFQMYNATCPVTFIQYAAAKALRDERSWKAVEEMRKEYDRRRKLVWKRLNEMGLPT 312
Query: 314 VKGGEGAIYLFAKLPKGGYDDFDVVRWLANRHGIAVIPGSACGAAGN--LRISFGGLTES 371
VK +GA Y+F ++ G + + +AV+PGSA G AG +RIS+ E
Sbjct: 313 VK-PKGAFYIFPRIKDTGLTSKEFSELMLMEAKVAVVPGSAFGKAGEGYVRISYATAYEK 371
Query: 372 DCRAAAERLKKGLEE 386
A +R++K L E
Sbjct: 372 -LEEAMDRMEKVLRE 385
>UniRef100_Q8U1F5 Aspartate transaminase [Pyrococcus furiosus]
Length = 389
Score = 143 bits (360), Expect = 1e-32
Identities = 103/375 (27%), Positives = 184/375 (48%), Gaps = 21/375 (5%)
Query: 24 EVLRGAKNCVSLAQGVVYWQPPKPALDKVKELVWEPSVSRYGADEGLPELRAALVKKLRQ 83
++ G K+ +SL G + P + KE + + ++ YG + GLPELR A+ +KL++
Sbjct: 20 DLAAGMKDVISLGIGEPDFDTPAHIKEYAKEGL-DKGLTHYGPNVGLPELREAIAEKLKK 78
Query: 84 ENNLH---KSSVMVTAGANQAFVNLVLTLCDAGDSVVMFAPYYFNAYMSFQMTGITNIHV 140
+N + S +MV GANQAF+ + T G+ V++ +P + + + + G + V
Sbjct: 79 QNGIEADPNSEIMVLVGANQAFLMSLATFLKDGEEVLIPSPMFVSYAPAVILAGGKPVEV 138
Query: 141 GPGSPDTLYPDVDWLERILTESKPVPKLVTVVNPGNPTGTYIPESLLQRIANLCKNAGSW 200
+ +VD L++ ++E + + + +P NPTG + + L+ IA+
Sbjct: 139 PTYEDNEFRINVDDLKKHVSEKT---RALIINSPNNPTGAVLTKKDLEEIADFANEHDLM 195
Query: 201 LLVDNTYEYFMFDGLKHSCVEG-----NHIVNIFSFSKAYGMMGWRVGYIAYPSEVEGLS 255
++ D YE+F++DG KH + + + FSK + M GWR+G++ PS V
Sbjct: 196 IISDEVYEHFIYDGAKHYSIAALDGMFGRTITVNGFSKTFAMTGWRLGFVVAPSWV---I 252
Query: 256 AQLLKVQDNIPICASIISQHLALHSLEMGPEW--VRERVQTLAKNREIVSEALSPLGEGS 313
+++K Q C Q+ A +L W V E + + R +V + L+ +G +
Sbjct: 253 EKMVKFQMYNATCPVTFIQYAAAKALRDERSWKAVEEMRKEYERRRNLVWKRLNEMGLPT 312
Query: 314 VKGGEGAIYLFAKLPKGGYDDFDVVRWLANRHGIAVIPGSACGAAGN--LRISFGGLTES 371
V+ +GA Y+F ++ G + + +AV+PGSA G AG +RIS+ E
Sbjct: 313 VE-PKGAFYIFPRIKDTGLSSKEFSELMLMEAKVAVVPGSAFGRAGEGYIRISYATAYEK 371
Query: 372 DCRAAAERLKKGLEE 386
A R++K L+E
Sbjct: 372 -LEEAMNRMEKVLKE 385
>UniRef100_Q7V7G7 Aminotransferases class-I [Prochlorococcus marinus]
Length = 392
Score = 142 bits (358), Expect = 2e-32
Identities = 106/381 (27%), Positives = 192/381 (49%), Gaps = 19/381 (4%)
Query: 16 MPIMVQMQEVLRGAKNCVSLAQGVVYWQPPKPALDKVKELVWEPSVSRYGADEGLPELRA 75
+ I + + + + ++ SL+ G + P+ +D + + + ++RYG G PELR
Sbjct: 20 LAISARAKALQQEGRDICSLSAGEPDFNTPEFIIDATVKALRD-GITRYGPAAGDPELRE 78
Query: 76 ALVKKLRQENNL--HKSSVMVTAGANQAFVNLVLTLCDAGDSVVMFAPYYFNAYMSFQMT 133
A+ KL +EN + + V+VT G QA NL + + GD V++ APY+ + ++
Sbjct: 79 AIATKLSKENTVPTNAEQVLVTNGGKQAIFNLFQVILNPGDEVLIPAPYWLSYPEMARLA 138
Query: 134 GITNIHVGPGSPDTLYP-DVDWLERILTESKPVPKLVTVVNPGNPTGTYIPESLLQRIAN 192
G + P +P+ + D++ LE + P +L+ + +PGNPTG + L+ +A+
Sbjct: 139 G-AKVTTLPSTPENGFCLDLNNLEASIG---PKTRLLLLNSPGNPTGRVMARKELETLAD 194
Query: 193 LCKNAGSWLLV-DNTYEYFMFDGLKHSCVEG------NHIVNIFSFSKAYGMMGWRVGYI 245
L ++ L++ D YE+ + +G +H N + F+K + M GWR+GY+
Sbjct: 195 LLRDHPQILVMSDEIYEFILEEGQQHYSFSAIAPDLSNRTFIVNGFAKGWAMTGWRLGYL 254
Query: 246 AYPSEVEGLSAQLLKVQDNIPICASIISQHLALHSLEMGPEWVRERVQTLAKNREIVSEA 305
A P++ +A L+ Q +C+ +Q AL +L+ E V++ V++ RE+++
Sbjct: 255 AGPADAVK-AATALQSQSTSNVCS--FAQRGALAALQGSRECVKKMVKSYNTRRELLTSG 311
Query: 306 LSPLGEGSVKGGEGAIYLFAKLPKGGYDDFDVVRWLANRHGIAVIPGSACGAAGNLRISF 365
L L S+ +GA Y F KLP D HG+A++PG+A G +R++
Sbjct: 312 LLSLEGMSLVPPKGAFYAFPKLPPESLDSVSFCEQALENHGLAMVPGAAFGDDSCVRLTC 371
Query: 366 GGLTESDCRAAAERLKKGLEE 386
E+ C ERL+K L++
Sbjct: 372 AVSPETIC-DGLERLRKALKQ 391
>UniRef100_Q7V217 Aminotransferases class-I [Prochlorococcus marinus subsp. pastoris]
Length = 393
Score = 141 bits (356), Expect = 3e-32
Identities = 109/398 (27%), Positives = 188/398 (46%), Gaps = 28/398 (7%)
Query: 3 SYANLARRALETEMPIMVQMQ----EVLRGAKNCVSLAQGVVYWQPPKPALDKVKELVWE 58
S NL++RAL E + +Q+ ++ + K+ +L+ G + P L E +++
Sbjct: 2 SEVNLSKRALSIEPSLTLQISAKANQLAKEGKDICNLSAGEPDFDAPNEILKATSEAIFD 61
Query: 59 PSVSRYGADEGLPELRAALVKKLRQENNLHKS--SVMVTAGANQAFVNLVLTLCDAGDSV 116
++YG G ELR A+ +KL+ +NNL+ +VMVT GA Q+ NL L + GD V
Sbjct: 62 -GYTKYGPASGDLELRKAIAEKLQTQNNLNVEYENVMVTNGAKQSIYNLFQVLLNDGDEV 120
Query: 117 VMFAPYYFNAYMSFQMTGITNIHVGPGSPDTLYPDVDWLERILTESKPVPKLVTVVNPGN 176
++ APY+ + ++ G + + + D ++ L+ ++ P K + + +P N
Sbjct: 121 IIPAPYWLSYPQMVRLAGGKPVFLNSSAEDGFKINIQDLK---SKISPKTKFIIINSPNN 177
Query: 177 PTGTYIPESLLQRIANLCK-NAGSWLLVDNTYEYFMFDGLKHSCVEG------NHIVNIF 229
PTG +P+ L +IA L + N +L D YE + KH + I I
Sbjct: 178 PTGRVMPKEELLQIAELVRENKNINILSDEIYELILKKEFKHLSLASLATDLKERIFIIN 237
Query: 230 SFSKAYGMMGWRVGYIAYPSEVEGLSAQLLKVQDNIPICASIISQHLALHSLEMGPEWVR 289
F+K + M GWRVGY+ +V S+ L + Q +C+ + Q AL +L++ E+
Sbjct: 238 GFAKGWAMTGWRVGYLVGQKDVIKASSAL-QSQSTSNVCSFV--QRGALEALKINREFFL 294
Query: 290 ERVQTLAKNREIVSEALSPLGEGSVKGGEGAIYLFAKLPKGGYDDFDVVRWLANRHGIAV 349
E RE++ + L + + GA Y F +LP + + + N +G+ V
Sbjct: 295 EINSHYDLRREVLYKGLKNIEGLFISPPNGAFYAFPRLPNNSMTSVEFCKRILNDYGLVV 354
Query: 350 IPGSACGAAGNLRISFGGLTESDCRAAAERLKKGLEEL 387
+PG G +RIS C ++ E++ GL L
Sbjct: 355 VPGKPFGDDQCIRIS--------CASSKEKILDGLTRL 384
>UniRef100_UPI000033BD5F UPI000033BD5F UniRef100 entry
Length = 400
Score = 140 bits (354), Expect = 5e-32
Identities = 101/367 (27%), Positives = 171/367 (46%), Gaps = 16/367 (4%)
Query: 30 KNCVSLAQGVVYWQPPKPALDKVKELVWEPSVSRYGADEGLPELRAALVKKLRQENN--L 87
K ++LAQ V + P + + + ++Y A G+P LR +L + + +
Sbjct: 33 KPLLNLAQAVPSYPPASSMSKHLADKLNLFETAQYTAIAGIPPLRNSLASHMECSHGGRI 92
Query: 88 HKSSVMVTAGANQAFVNLVLTLCDAGDSVVMFAPYYFNAYMSFQMTGITNIHVGPGSPDT 147
+ +V+++AG NQAF ++ + AGD V++ PYYFN M +M GI + + +
Sbjct: 93 NPENVLISAGCNQAFCLTIMAVAKAGDEVILSVPYYFNHMMWLEMLGIKPVLIPFRKDLS 152
Query: 148 LYPDVDWLERILTESKPVPKLVTVVNPGNPTGTYIPESLLQRIANLCKNAGSWLLVDNTY 207
P + +E ++TE K + +V+P NPTG P ++L + L + L++D TY
Sbjct: 153 SVPSPEDVEGLITEK---TKAIVLVSPNNPTGAIYPSNILSQFFELAHTSKIALVLDETY 209
Query: 208 EYFMFDGLKH-----SCVEGNHIVNIFSFSKAYGMMGWRVGYIAYPSEVEGLSAQLLKVQ 262
+ F+ H SC E V ++SFSK + + G+RVG I + + KV
Sbjct: 210 KDFLTTMPPHTIFHSSCWE-ETFVQLYSFSKVFSLTGYRVGSIIGGQSIIDAAT---KVM 265
Query: 263 DNIPICASIISQHLALHSLEMGPEWVRERVQTLAKNREIVSEALSPLGEGSVKGGEGAIY 322
D + ICA I+Q AL+ L +WV + + + V++ S GA +
Sbjct: 266 DTLAICAPRIAQDAALYGLNNLEKWVANKRTGIQERLNYVNDIFSSDQTSFELISSGAYF 325
Query: 323 LFAKLPKGGYDDFDVVRWLANRHGIAVIPGSACGAAGN--LRISFGGLTESDCRAAAERL 380
+ K P DV R LA + I +PGS G LR++F T + +RL
Sbjct: 326 AYVKHPFDKIPARDVARHLAEKENILCLPGSFFGPGQEQFLRLAFANATVEELSLLPKRL 385
Query: 381 KKGLEEL 387
+ ++ +
Sbjct: 386 RSSVDSI 392
>UniRef100_Q8TPT6 Aspartate aminotransferase [Methanosarcina acetivorans]
Length = 380
Score = 140 bits (353), Expect = 6e-32
Identities = 109/367 (29%), Positives = 182/367 (48%), Gaps = 22/367 (5%)
Query: 25 VLRGAKNCVSLAQGVVYWQPPKPALDKVKELVWEPSVSRYGADEGLPELRAALVKKLRQE 84
+++ + ++ + G + PK D + ++E + Y G+PELRAA+ +KL+ E
Sbjct: 24 MIKEGTDVINFSLGEPDFDTPKNICDAAAKAMYEGK-THYAPSAGIPELRAAIAEKLKTE 82
Query: 85 NNLH--KSSVMVTAGANQAFVNLVLTLCDAGDSVVMFAPYYFNAYMSFQMTGITNIHVGP 142
N+L + V+VT GA QA +++ D GD ++F P + + +G + V P
Sbjct: 83 NHLEVTEKDVLVTPGAKQAIFEIMMGALDDGDRALLFDPAWVTYDACIRFSGANTVWV-P 141
Query: 143 GSPDTLYPDVDWLERILTESKPVPKLVTVVNPGNPTGTYIPESLLQRIANLCKNAGSWLL 202
P+ + ++ E I ++K L+ V +PGNPTG + LQ IA+L + ++
Sbjct: 142 TVPERGFLPDNFAEYINDKTK----LIVVNSPGNPTGGVFGKKTLQCIADLAIDHDLLVV 197
Query: 203 VDNTYEYFMFDGLKHSCVEG-----NHIVNIFSFSKAYGMMGWRVGYIAYPSEVEGLSAQ 257
D YE ++D +H + + + + FSKAY M GWR+GY+ P E+ L
Sbjct: 198 SDEIYEKIIYDR-EHISIGSFDGMQDRTITVNGFSKAYAMTGWRLGYLTAPPEIFKL--- 253
Query: 258 LLKVQDNIPICASIISQHLALHSLEMGPEWVRERVQTLAKNREIVSEALSPLGEGSVKGG 317
L K+Q + A+ Q+ L +L+ + V+ V R+I+ + L+ +G K
Sbjct: 254 LQKIQSHSVSSATTFVQYGGLEALQGPQDGVKAMVDRFKMRRDILIDGLNKIGI-ECKKP 312
Query: 318 EGAIYLFAKLPKGGYDDFDVVRWLANRHGIAVIPGSACGAAGN--LRISFGGLTESDCRA 375
+GA Y FA + + G R L H +AV PG A GA+G +RIS+ + R
Sbjct: 313 DGAFYAFANVSEYGNGTEVAERLLKEAH-VAVTPGIAFGASGEDFIRISYATSIDR-IRE 370
Query: 376 AAERLKK 382
A ERL+K
Sbjct: 371 ALERLEK 377
>UniRef100_Q72G88 Aspartate aminotransferase [Thermus thermophilus]
Length = 385
Score = 140 bits (352), Expect = 8e-32
Identities = 102/366 (27%), Positives = 172/366 (46%), Gaps = 11/366 (3%)
Query: 24 EVLRGAKNCVSLAQGVVYWQPPKPALDKVKELVWEPSVSRYGADEGLPELRAALVKKLRQ 83
E+ R + V+L G + P+ + + + + ++Y G+PELR AL +K R+
Sbjct: 25 ELRRQGVDLVALTAGEPDFDTPEHVKEAARRALAQGK-TKYAPPAGIPELREALAEKFRR 83
Query: 84 ENNLHKS--SVMVTAGANQAFVNLVLTLCDAGDSVVMFAPYYFNAYMSFQMTGITNIHVG 141
EN L + +VT G QA NL + D GD V++ +PY+ + + G + V
Sbjct: 84 ENGLSVTPEETIVTVGGKQALFNLFQAILDPGDEVIVLSPYWVSYPEMVRFAGGVVVEVE 143
Query: 142 PGSPDTLYPDVDWLERILTESKPVPKLVTVVNPGNPTGTYIPESLLQRIANLCKNAGSWL 201
+ PD + + R +T P K + V +P NPTG P+ +L+ +A L +L
Sbjct: 144 TLPEEGFVPDPERVRRAIT---PRTKALVVNSPNNPTGAVYPKEVLEALAKLAVEHDFYL 200
Query: 202 LVDNTYEYFMFDGLKHS--CVEGNHIVNIFSFSKAYGMMGWRVGYIAYPSEVEGLSAQLL 259
+ D YE+ +++G S V H + + +KA+ M GWR+GY P EV A +
Sbjct: 201 VSDEIYEHLLYEGEHFSPGRVAPEHTLTVNGAAKAFAMTGWRIGYACGPKEVIKAMASVS 260
Query: 260 KVQDNIPICASIISQHLALHSLEMGPEWVRERVQTLAKNREIVSEALSPLGEGSVKGGEG 319
P + + AL + E +V + + R+++ E L+ LG +V+ G
Sbjct: 261 SQSTTSPDTIAQWATLEALTNQEASRAFVEMAREAYRRRRDLLLEGLTALGLKAVR-PSG 319
Query: 320 AIYLFAKLPKGGYDDFDVVRWLANRHGIAVIPGSACGAAGNLRISFGGLTESDCRAAAER 379
A Y+ D+ L G+AV+PG+ A G++R+S+ +E + R A ER
Sbjct: 320 AFYVLMDTSPIAPDEVRAAERLLEA-GVAVVPGTDFAAFGHVRLSY-ATSEENLRKALER 377
Query: 380 LKKGLE 385
+ LE
Sbjct: 378 FARVLE 383
>UniRef100_Q8RR70 Aspartate aminotransferase [Phormidium lapideum]
Length = 388
Score = 140 bits (352), Expect = 8e-32
Identities = 102/364 (28%), Positives = 175/364 (48%), Gaps = 18/364 (4%)
Query: 34 SLAQGVVYWQPPKPALDKVKELVWEPSVSRYGADEGLPELRAALVKKLRQENNL--HKSS 91
S + G + PK ++ K + E +RYG G P LR A+ +KL+++N L +
Sbjct: 34 SFSAGEPDFNTPKHIVEAAKAAL-EQGKTRYGPAAGEPRLREAIAQKLQRDNGLCYGADN 92
Query: 92 VMVTAGANQAFVNLVLTLCDAGDSVVMFAPYYFNAYMSFQMTGITNIHVGPGSPDTLYPD 151
++VT G Q+ NL+L + + GD V++ AP++ + ++ T + + P + +T +
Sbjct: 93 ILVTNGGKQSIFNLMLAMIEPGDEVIIPAPFWVSYPEMVKLAEGTPV-ILPTTVETQFKV 151
Query: 152 VDWLERILTESKPVPKLVTVVNPGNPTGTYIPESLLQRIANLCKNAGSWLLVDNTYEYFM 211
E+I P KL+ P NPTG ++ IA + AG W+L D YE +
Sbjct: 152 SP--EQIRQAITPKTKLLVFNTPSNPTGMVYTPDEVRAIAQVAVEAGLWVLSDEIYEKIL 209
Query: 212 FDGLKHSCVEG------NHIVNIFSFSKAYGMMGWRVGYIAYPSEVEGLSAQLLKVQDNI 265
+D +H + V F+K Y M GWRVG++A P L K+Q +
Sbjct: 210 YDDAQHLSIGAASPEAYERSVVCSGFAKTYAMTGWRVGFLAGPVP---LVKAATKIQGHS 266
Query: 266 PICASIISQHLALHSLEMGPEWVRERVQTLAKNREIVSEALSPLGEGSVKGGEGAIYLFA 325
+Q+ A+ + E + V+E + A+ R + +AL+ + +GA Y+F
Sbjct: 267 TSNVCTFAQYGAIAAYENSQDCVQEMLAAFAERRRYMLDALNAMPGLECPKPDGAFYMFP 326
Query: 326 KLPKGGYDDFDVVRWLANRHGIAVIPGSACGAAGNLRISFGGLTESD-CRAAAERLKKGL 384
+ K G D L ++H +A +PG+A GA +R+S+ T+ D + ERL+K L
Sbjct: 327 SIAKTGRSSLDFCSELLDQHQVATVPGAAFGADDCIRLSYA--TDLDTIKRGMERLEKFL 384
Query: 385 EELV 388
++
Sbjct: 385 HGIL 388
>UniRef100_Q8LFX1 Putative aspartate aminotransferase [Arabidopsis thaliana]
Length = 475
Score = 140 bits (352), Expect = 8e-32
Identities = 108/379 (28%), Positives = 184/379 (48%), Gaps = 31/379 (8%)
Query: 33 VSLAQGVVYWQPPKPALDKVKELVWEPSVSRYGADEGLPELRAALVKKLRQENNLHKS-- 90
+ LA G + PK + + E +RY + G+ ELR A+ +KL++EN L +
Sbjct: 102 IRLAAGEPDFDTPKVVAEAGINAIRE-GFTRYTLNAGITELREAICRKLKEENGLSYAPD 160
Query: 91 SVMVTAGANQAFVNLVLTLCDAGDSVVMFAPYYFNAYMSFQMTGITNIHVGPGSPDTLYP 150
++V+ GA Q+ + VL +C GD V++ APY+ + ++ T + + +
Sbjct: 161 QILVSNGAKQSLLQAVLAVCSPGDEVIIPAPYWVSYTEQARLADATPVVIPTKISNNFLL 220
Query: 151 DVDWLERILTESKPVPKLVTVVNPGNPTGTYIPESLLQRIAN-LCKNAGSWLLVDNTYEY 209
D LE LTE +L+ + +P NPTG+ P+SLL+ IA + K+ +L D YE+
Sbjct: 221 DPKDLESKLTEKS---RLLILCSPSNPTGSVYPKSLLEEIARIIAKHPRLLVLSDEIYEH 277
Query: 210 FMFDGLKHSCVEG-----NHIVNIFSFSKAYGMMGWRVGYIAYPSEVEGLSAQLLKVQDN 264
++ H+ + + FSKA+ M GWR+GY+A P + A K+Q
Sbjct: 278 IIYAPATHTSFASLPDMYERTLTVNGFSKAFAMTGWRLGYLAGPKHI---VAACSKLQGQ 334
Query: 265 IPICASIISQHLALHSLEMGP---EWVRERVQTLAKNREIVSEALSPLGEGSVKGGEGAI 321
+ AS I+Q + +L +G E V E V+ + R+ + ++L + + +GA
Sbjct: 335 VSSGASSIAQKAGVAALGLGKAGGETVAEMVKAYRERRDFLVKSLGDIKGVKISEPQGAF 394
Query: 322 YLFAKL------PKGGY----DDFDVVRWLANRHGIAVIPGSACGAAGNLRISFGGLTES 371
YLF G+ D + + ++ +A++PG A G +RIS+ T
Sbjct: 395 YLFIDFSAYYGSEAEGFGLINDSSSLALYFLDKFQVAMVPGDAFGDXSCIRISYA--TSL 452
Query: 372 D-CRAAAERLKKGLEELVA 389
D +AA E+++K LE L A
Sbjct: 453 DVLQAAVEKIRKALEPLRA 471
>UniRef100_O67781 Aspartate aminotransferase [Aquifex aeolicus]
Length = 394
Score = 140 bits (352), Expect = 8e-32
Identities = 102/340 (30%), Positives = 163/340 (47%), Gaps = 22/340 (6%)
Query: 62 SRYGADEGLPELRAALVKKLRQENNLH--KSSVMVTAGANQAFVNLVLTLCDAGDSVVMF 119
++Y G+PELR A+ +KL +EN + S ++V+AGA + + + D GD V++
Sbjct: 63 TKYAPSAGIPELREAIAEKLLKENKVEYKPSEIVVSAGAKMVLFLIFMAILDEGDEVLLP 122
Query: 120 APYYFNAYMSFQMTGITNIHVGPGSPDTLYPDVDWLERILTESKPVPKLVTVVNPGNPTG 179
+PY+ + G + V ++ ++ +TE K + + +P NPTG
Sbjct: 123 SPYWVTYPEQIRFFGGVPVEVPLKKEKGFQLSLEDVKEKVTERT---KAIVINSPNNPTG 179
Query: 180 TYIPESLLQRIANLCKNAGSWLLVDNTYEYFMFDGLK------HSCVEGNHIVNIFSFSK 233
E L++IA C G +++ D YEYF++ K S N + +FSK
Sbjct: 180 AVYEEEELKKIAEFCVERGIFIISDECYEYFVYGDAKFVSPASFSDEVKNITFTVNAFSK 239
Query: 234 AYGMMGWRVGYIAYPSEVEGLSAQLLKVQDNIPICASIISQHLALHSLE--MGPEWVRER 291
+Y M GWR+GY+A P E + A L + +Q+ AL +L+ ++V E
Sbjct: 240 SYSMTGWRIGYVACPEEYAKVIASL---NSQSVSNVTTFAQYGALEALKNPKSKDFVNEM 296
Query: 292 VQTLAKNREIVSEALSPLGEGSVKGGEGAIYLF----AKLPKGGYDDFDVVRWLANRHGI 347
+ R+ E LS + V EGA Y+F A K G D + +L + +
Sbjct: 297 RNAFERRRDTAVEELSKIPGMDVVKPEGAFYIFPDFSAYAEKLG-GDVKLSEFLLEKAKV 355
Query: 348 AVIPGSACGAAGNLRISFGGLTESDCRAAAERLKKGLEEL 387
AV+PGSA GA G LR+S+ L+E R+KK LEE+
Sbjct: 356 AVVPGSAFGAPGFLRLSY-ALSEERLVEGIRRIKKALEEI 394
>UniRef100_Q9SIE1 Putative aspartate aminotransferase [Arabidopsis thaliana]
Length = 475
Score = 139 bits (350), Expect = 1e-31
Identities = 108/379 (28%), Positives = 184/379 (48%), Gaps = 31/379 (8%)
Query: 33 VSLAQGVVYWQPPKPALDKVKELVWEPSVSRYGADEGLPELRAALVKKLRQENNLHKS-- 90
+ LA G + PK + + E +RY + G+ ELR A+ +KL++EN L +
Sbjct: 102 IRLAAGEPDFDTPKVVAEAGINAIRE-GFTRYTLNAGITELREAICRKLKEENGLSYAPD 160
Query: 91 SVMVTAGANQAFVNLVLTLCDAGDSVVMFAPYYFNAYMSFQMTGITNIHVGPGSPDTLYP 150
++V+ GA Q+ + VL +C GD V++ APY+ + ++ T + + +
Sbjct: 161 QILVSNGAKQSLLQAVLAVCSPGDEVIIPAPYWVSYTEQARLADATPVVIPTKISNNFLL 220
Query: 151 DVDWLERILTESKPVPKLVTVVNPGNPTGTYIPESLLQRIAN-LCKNAGSWLLVDNTYEY 209
D LE LTE +L+ + +P NPTG+ P+SLL+ IA + K+ +L D YE+
Sbjct: 221 DPKDLESKLTEKS---RLLILCSPSNPTGSVYPKSLLEEIARIIAKHPRLLVLSDEIYEH 277
Query: 210 FMFDGLKHSCVEG-----NHIVNIFSFSKAYGMMGWRVGYIAYPSEVEGLSAQLLKVQDN 264
++ H+ + + FSKA+ M GWR+GY+A P + A K+Q
Sbjct: 278 IIYAPATHTSFASLPDMYERTLTVNGFSKAFAMTGWRLGYLAGPKHI---VAACSKLQGQ 334
Query: 265 IPICASIISQHLALHSLEMGP---EWVRERVQTLAKNREIVSEALSPLGEGSVKGGEGAI 321
+ AS I+Q + +L +G E V E V+ + R+ + ++L + + +GA
Sbjct: 335 VSSGASSIAQKAGVAALGLGKAGGETVAEMVKAYRERRDFLVKSLGDIKGVKISEPQGAF 394
Query: 322 YLFAKL------PKGGY----DDFDVVRWLANRHGIAVIPGSACGAAGNLRISFGGLTES 371
YLF G+ D + + ++ +A++PG A G +RIS+ T
Sbjct: 395 YLFIDFSAYYGSEAEGFGLINDSSSLALYFLDKFQVAMVPGDAFGDDSCIRISYA--TSL 452
Query: 372 D-CRAAAERLKKGLEELVA 389
D +AA E+++K LE L A
Sbjct: 453 DVLQAAVEKIRKALEPLRA 471
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.319 0.136 0.411
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 690,800,340
Number of Sequences: 2790947
Number of extensions: 29584158
Number of successful extensions: 77027
Number of sequences better than 10.0: 3813
Number of HSP's better than 10.0 without gapping: 903
Number of HSP's successfully gapped in prelim test: 2910
Number of HSP's that attempted gapping in prelim test: 70933
Number of HSP's gapped (non-prelim): 3946
length of query: 399
length of database: 848,049,833
effective HSP length: 129
effective length of query: 270
effective length of database: 488,017,670
effective search space: 131764770900
effective search space used: 131764770900
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 76 (33.9 bits)
Lotus: description of TM0226.7


