

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0226.6
(263 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
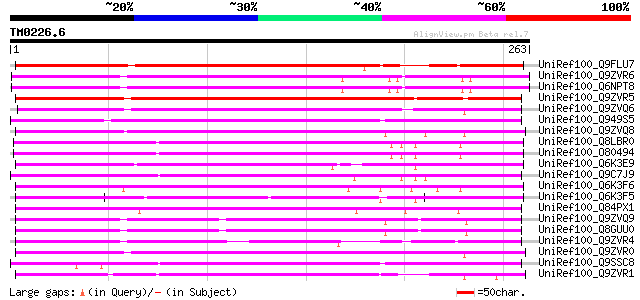
UniRef100_Q9FLU7 Phloem-specific lectin-like protein [Arabidopsi... 194 2e-48
UniRef100_Q9ZVR6 Putative phloem-specific lectin [Arabidopsis th... 193 3e-48
UniRef100_Q6NPT8 At2g02230 [Arabidopsis thaliana] 186 5e-46
UniRef100_Q9ZVR5 Lectin-like protein [Arabidopsis thaliana] 184 2e-45
UniRef100_Q9ZVQ6 Putative phloem-specific lectin [Arabidopsis th... 184 2e-45
UniRef100_Q949S5 Hypothetical protein At1g80110 [Arabidopsis tha... 177 3e-43
UniRef100_Q9ZVQ8 Putative phloem-specific lectin [Arabidopsis th... 174 3e-42
UniRef100_Q8LBR0 Phloem-specific lectin PP2-like protein [Arabid... 174 3e-42
UniRef100_O80494 T12M4.17 protein [Arabidopsis thaliana] 173 5e-42
UniRef100_Q6K3E9 F-box family protein-like [Oryza sativa] 171 1e-41
UniRef100_Q9C7J9 Hypothetical protein F14G9.15 [Arabidopsis thal... 167 3e-40
UniRef100_Q6K3F6 Phloem-specific lectin-like [Oryza sativa] 162 6e-39
UniRef100_Q6K3F5 F-box family protein-like [Oryza sativa] 159 7e-38
UniRef100_Q84PX1 Putative F-box protein [Oryza sativa] 151 1e-35
UniRef100_Q9ZVQ9 Hypothetical protein At2g02320 [Arabidopsis tha... 146 6e-34
UniRef100_Q8GUU0 Hypothetical protein T16F16.11 [Arabidopsis tha... 145 1e-33
UniRef100_Q9ZVR4 Putative phloem-specific lectin [Arabidopsis th... 144 2e-33
UniRef100_Q9ZVR0 Putative phloem-specific lectin [Arabidopsis th... 141 2e-32
UniRef100_Q9SSC8 F18B13.19 protein [Arabidopsis thaliana] 137 4e-31
UniRef100_Q9ZVR1 Putative phloem-specific lectin [Arabidopsis th... 135 8e-31
>UniRef100_Q9FLU7 Phloem-specific lectin-like protein [Arabidopsis thaliana]
Length = 251
Score = 194 bits (493), Expect = 2e-48
Identities = 110/262 (41%), Positives = 161/262 (60%), Gaps = 24/262 (9%)
Query: 4 LPEGCIAAILSRTTPVDAARLSLVSKIFRSAADSDAVWDRFLPSDSHSIVSQSLSRANAS 63
LPE CIA ++S T+P DA R+S VSK+ RSAADS+ W+RFLPSD + SLSR
Sbjct: 6 LPEECIATMISFTSPFDACRISAVSKLLRSAADSNTTWERFLPSDYRMYIDNSLSRF--- 62
Query: 64 SKKRFYLDLSDHPVVIDNGKKSFQLDKKSGKKCFMLGARALFIAWDNGDHYRCWTTLPES 123
S K+ +L + P++I++G+ SF ++K+SGKKC+ML AR L I W + + W ++P+S
Sbjct: 63 SNKQLFLRFCESPLLIEDGRTSFWMEKRSGKKCWMLSARKLDIVWVDSPEFWIWVSIPDS 122
Query: 124 RFPDVVVLRHLLWLEIHGMINTFSLSPNTQYGAFLVFKIIDAPNFHEVLVVLSVN----N 179
RF +V L + W EI G I+T LS T Y A+LVFK + +F + L V+
Sbjct: 123 RFEEVAGLLMVCWFEIRGKISTSLLSKATNYSAYLVFKEQEMGSFGFESLPLEVSFRSTR 182
Query: 180 IGGHNITKIAYLNPDSLEGEVHDRWDRVQGPSVRSDGWLEIEMGEFFISGLEDEEVEMSV 239
+N ++ +L + E R DGWLEIE+GE+++ G +DEE+EMSV
Sbjct: 183 TEVYNNRRV-FLKSGTQES--------------REDGWLEIELGEYYV-GFDDEEIEMSV 226
Query: 240 LQI-DGCFNVGLIIEGIEVRPK 260
L+ +G + G+I++GIE+RPK
Sbjct: 227 LETREGGWKGGIIVQGIEIRPK 248
>UniRef100_Q9ZVR6 Putative phloem-specific lectin [Arabidopsis thaliana]
Length = 317
Score = 193 bits (491), Expect = 3e-48
Identities = 118/289 (40%), Positives = 168/289 (57%), Gaps = 31/289 (10%)
Query: 2 ELLPEGCIAAILSRTTPVDAARLSLVSKIFRSAADSDAVWDRFLPSDSHSIVSQSLSRAN 61
+ LPE CI+ ++S T+P DA ++ VSK +SAA SD VW+ FLPS+ S+V QS AN
Sbjct: 33 DALPEDCISKVISHTSPRDACVVASVSKSVKSAAQSDLVWEMFLPSEYSSLVLQS---AN 89
Query: 62 ASSKKRFYLDLSDHPVVIDNGKKSFQLDKKSGKKCFMLGARALFIAWDNGDHYRCWTTLP 121
SKK +L L+D+ V+++NGKKSF ++K SGKKC+ML A L I W + Y W T+P
Sbjct: 90 HLSKKEIFLSLADNSVLVENGKKSFWVEKASGKKCYMLSAMELTIIWGDSPAYWKWITVP 149
Query: 122 ESRFPDVVVLRHLLWLEIHGMINTFSLSPNTQYGAFLVFKIIDAPN--FHEVLVVLSVNN 179
ES+F V LR++ W E+ G I+ LS T Y ++VFK + + F V V V
Sbjct: 150 ESKFEKVAELRNVCWFEVRGKISCGMLSKGTHYSVYVVFKTANGRSYGFDLVPVEAGVGF 209
Query: 180 IGGHNITKIAYL---NPDS----------------LEGEVHDRWDRVQGPSVRSDGWLEI 220
+G K Y N DS +EGE +R V GP R DGW E+
Sbjct: 210 VGKVATKKSVYFESGNADSRSATSHYSGISEEEEEVEGE-RERGMNVVGPKERVDGWSEV 268
Query: 221 EMGEFFIS--GLED---EEVEMSVLQI-DGCFNVGLIIEGIEVRPKNDN 263
E+G+F+I+ G D +E+E+S+++ +G + GLII+GIE+RP+ N
Sbjct: 269 ELGKFYINNGGCGDDGSDEIEISIMETQNGNWKSGLIIQGIEIRPERSN 317
>UniRef100_Q6NPT8 At2g02230 [Arabidopsis thaliana]
Length = 336
Score = 186 bits (472), Expect = 5e-46
Identities = 118/308 (38%), Positives = 168/308 (54%), Gaps = 50/308 (16%)
Query: 2 ELLPEGCIAAILSRTTPVDAARLSLVSKIFRSAADSDAVWDRFLPSDSHSIVSQSLSRAN 61
+ LPE CI+ ++S T+P DA ++ VSK +SAA SD VW+ FLPS+ S+V QS AN
Sbjct: 33 DALPEDCISKVISHTSPRDACVVASVSKSVKSAAQSDLVWEMFLPSEYSSLVLQS---AN 89
Query: 62 ASSKKRFYLDLSDHPVVIDNGKKSFQLDKKSGKKCFMLGARALFIAWDNGDHYRCWTTLP 121
SKK +L L+D+ V+++NGKKSF ++K SGKKC+ML A L I W + Y W T+P
Sbjct: 90 HLSKKEIFLSLADNSVLVENGKKSFWVEKASGKKCYMLSAMELTIIWGDSPAYWKWITVP 149
Query: 122 ESRFPDVVVLRHLLWLEIHGMINTFSLSPNTQYGAFLVFKIIDAPN--FHEVLVVLSVNN 179
ES+F V LR++ W E+ G I+ LS T Y ++VFK + + F V V V
Sbjct: 150 ESKFEKVAELRNVCWFEVRGKISCGMLSKGTHYSVYVVFKTANGRSYGFDLVPVEAGVGF 209
Query: 180 IGGHNITKIAYL---NPDS-----------------------------------LEGEVH 201
+G K Y N DS +EGE
Sbjct: 210 VGKVATKKSVYFESGNADSRSATSHYSGISYAMVSRAFRMRRPWMQVQREEEEEVEGE-R 268
Query: 202 DRWDRVQGPSVRSDGWLEIEMGEFFIS--GLED---EEVEMSVLQI-DGCFNVGLIIEGI 255
+R V GP R DGW E+E+G+F+I+ G D +E+E+S+++ +G + GLII+GI
Sbjct: 269 ERGMNVVGPKERVDGWSEVELGKFYINNGGCGDDGSDEIEISIMETQNGNWKSGLIIQGI 328
Query: 256 EVRPKNDN 263
E+RP+ N
Sbjct: 329 EIRPERSN 336
>UniRef100_Q9ZVR5 Lectin-like protein [Arabidopsis thaliana]
Length = 305
Score = 184 bits (467), Expect = 2e-45
Identities = 110/260 (42%), Positives = 161/260 (61%), Gaps = 10/260 (3%)
Query: 4 LPEGCIAAILSRTTPVDAARLSLVSKIFRSAADSDAVWDRFLPSDSHSIVSQSLSRANAS 63
LPE CI+ I+S T+P DA + VSK F SA +SD+VWD+FLPSD S+V S S
Sbjct: 45 LPEDCISNIISFTSPRDACVAASVSKTFESAVNSDSVWDKFLPSDYSSLVPPSRV---FS 101
Query: 64 SKKRFYLDLSDHPVVIDNGKKSFQLDKKSGKKCFMLGAR-ALFIAWDNGDHYRCWTTLPE 122
SKK Y + D+PV++++G KSF L+K++GKKCFML + +++I W + Y W ++PE
Sbjct: 102 SKKELYFAICDNPVLVEDGGKSFWLEKENGKKCFMLSPKKSMWITWVSTPQYWRWISIPE 161
Query: 123 SRFPDVVVLRHLLWLEIHGMINTFSLSPNTQYGAFLVFKIID-APNFHEVLVVLSVNNIG 181
+RF +V L ++ W E+ G +NT LSP T+Y A++VFK + PN +V V +V +G
Sbjct: 162 ARFEEVPELLNVCWFEVRGGMNTKELSPGTRYSAYIVFKTKNGCPNLGDVPVEATVGLVG 221
Query: 182 GHNITKIAYLNPDSLEGEVHDRWDRVQGPSVRSDGWLEIEMGEFF-ISGLEDEEVEMSVL 240
+ + Y S + + D V P+ R DGW+E E+G+FF SG + V+ S+L
Sbjct: 222 QESSQRHIYFVGPSDQRRDRETRD-VTRPTKRKDGWMEAELGQFFNESGC--DVVDTSIL 278
Query: 241 QIDGCF-NVGLIIEGIEVRP 259
+I + GLII+GIE RP
Sbjct: 279 EIKTPYWKRGLIIQGIEFRP 298
>UniRef100_Q9ZVQ6 Putative phloem-specific lectin [Arabidopsis thaliana]
Length = 272
Score = 184 bits (467), Expect = 2e-45
Identities = 107/259 (41%), Positives = 144/259 (55%), Gaps = 12/259 (4%)
Query: 5 PEGCIAAILSRTTPVDAARLSLVSKIFRSAADSDAVWDRFLPSDSHSIVSQSLSRANASS 64
PE CI+ I+S T P DA + VSK F S SD +W++FLP+D S++ S SS
Sbjct: 18 PEDCISYIISFTNPRDACVAATVSKTFESTVKSDIIWEKFLPADYESLIPPSRV---FSS 74
Query: 65 KKRFYLDLSDHPVVIDNGKKSFQLDKKSGKKCFMLGARALFIAWDNGDHYRCWTTLPESR 124
KK Y L + PV+ D+ KKS L+K SGK+C ML A L I W + Y W +PESR
Sbjct: 75 KKELYFSLCNDPVLFDDDKKSVWLEKASGKRCLMLSAMNLSIIWGDNPQYWQWIPIPESR 134
Query: 125 FPDVVVLRHLLWLEIHGMINTFSLSPNTQYGAFLVFKIID-APNFHEVLVVLSVNNIGGH 183
F V LR + W EI G NT LSP T+Y A++VFK +D F V + +V +G
Sbjct: 135 FEKVAKLRDVCWFEIRGRTNTRVLSPRTRYSAYIVFKGVDKCYGFQNVAIEAAVGVVGQE 194
Query: 184 NITKIAYLNPDSLEGEVHDRWDRVQGPSVRSDGWLEIEMGEFFISG--LEDEEVEMSVLQ 241
++ + G V P R DGW+EIE+GEFF G ++++E+EMS L+
Sbjct: 195 PSRRLICFSEAIRRGR-----RNVVKPKQREDGWMEIELGEFFNDGGIMDNDEIEMSALE 249
Query: 242 IDGC-FNVGLIIEGIEVRP 259
GLII+GIE+RP
Sbjct: 250 TKQLNRKCGLIIQGIEIRP 268
>UniRef100_Q949S5 Hypothetical protein At1g80110 [Arabidopsis thaliana]
Length = 257
Score = 177 bits (448), Expect = 3e-43
Identities = 105/260 (40%), Positives = 147/260 (56%), Gaps = 6/260 (2%)
Query: 1 MELLPEGCIAAILSRTTPVDAARLSLVSKIFRSAADSDAVWDRFLPSDSHSIVSQSLSRA 60
M LPE CIA ILS TTP+D RLS VS IFRSAA SD VW+ FLP+D + + A
Sbjct: 1 MNNLPEDCIAKILSLTTPLDVCRLSAVSSIFRSAAGSDDVWNHFLPAD---FPAGFAAPA 57
Query: 61 NASSKKRFYLDLSDHPVVIDNGKKSFQLDKKSGKKCFMLGARALFIAWDNGDHYRCWTTL 120
++K+ + L D+P++I+ SF L++KSG KC+M+ ARAL I W + Y W +L
Sbjct: 58 GLPTRKQLFFSLVDNPLLINGTLLSFSLERKSGNKCYMMAARALNIVWGHEQRYWHWISL 117
Query: 121 PESRFPDVVVLRHLLWLEIHGMINTFSLSPNTQYGAFLVFKIIDAPNFHEVLVVLSVNNI 180
P +RF +V L + WLEI G IN LS +T Y A+ VFK +P V S+
Sbjct: 118 PNTRFGEVAELIMVWWLEITGKINITLLSDDTLYAAYFVFKWNHSPYGFRQPVETSLVLA 177
Query: 181 GGHNITKIAYLNPDSLEGEVHDRWDRVQGPSVRSDGWLEIEMGEFFISGLEDEEVEMSVL 240
+ + + P + + Q P +R DGW E+E+G+FF + E+EMS+
Sbjct: 178 DTESTDNV--VQPSMISLMQDSGGEEGQSPVLRRDGWYEVELGQFFKRRGDLGEIEMSLK 235
Query: 241 QIDGCF-NVGLIIEGIEVRP 259
+ G + GLI+ GIE+RP
Sbjct: 236 ETKGPYEKKGLIVYGIEIRP 255
>UniRef100_Q9ZVQ8 Putative phloem-specific lectin [Arabidopsis thaliana]
Length = 305
Score = 174 bits (440), Expect = 3e-42
Identities = 106/266 (39%), Positives = 152/266 (56%), Gaps = 9/266 (3%)
Query: 4 LPEGCIAAILSRTTPVDAARLSLVSKIFRSAADSDAVWDRFLPSDSHSIVSQSLSRANAS 63
LPE C++ I+S T+P DA L+ VSK F SA SD VW++F+P + S++SQS +
Sbjct: 39 LPEECVSIIVSFTSPQDACVLASVSKTFASAVKSDIVWEKFIPPEYESLISQSRA-FKFL 97
Query: 64 SKKRFYLDLSDHPVVIDNGKKSFQLDKKSGKKCFMLGARALFIAWDNGDHYRCWTTLPES 123
SKK Y L D V+ID+GKKS ++K + K+C M+ A L IAW N W P++
Sbjct: 98 SKKELYFALCDKSVLIDDGKKSLWIEKANAKRCIMISAMNLAIAWGNSPQSWRWIPDPQA 157
Query: 124 RFPDVVVLRHLLWLEIHGMINTFSLSPNTQYGAFLVFKIIDAPNFHEVLVVLSVNNIGGH 183
RF V L + EI G IN+ +SP T+Y A++V+K ++ E + V V + G
Sbjct: 158 RFETVAELLEVCLFEIRGRINSRVISPKTRYSAYIVYKKLNICYGFENVAVEVVVGVVGQ 217
Query: 184 NITKIA--YLNPDSLEGEVHDRWDRVQG---PSVRSDGWLEIEMGEFFISG--LEDEEVE 236
++ + Y+ D E R DR + P R DGW+EI++GEFF G L D+E+E
Sbjct: 218 DLEESCRRYICFDETMDEQFRRRDRGKNLVKPERRKDGWMEIKIGEFFNEGGLLNDDEIE 277
Query: 237 MSVLQI-DGCFNVGLIIEGIEVRPKN 261
M L+ + GLII+GIE+RP N
Sbjct: 278 MVALEAKQRHWKRGLIIQGIEIRPTN 303
>UniRef100_Q8LBR0 Phloem-specific lectin PP2-like protein [Arabidopsis thaliana]
Length = 288
Score = 174 bits (440), Expect = 3e-42
Identities = 112/280 (40%), Positives = 154/280 (55%), Gaps = 23/280 (8%)
Query: 3 LLPEGCIAAILSRTTPVDAARLSLVSKIFRSAADSDAVWDRFLPSDSHSIVSQSLSRANA 62
+LPE C+A ILS TTP D + VS +FR A DSD VW++FLP+D ++S+S
Sbjct: 1 MLPEACVATILSFTTPADTISSAAVSSVFRVAGDSDFVWEKFLPTDYCHVISRSTDPHRI 60
Query: 63 -SSKKRFYLDLSDHPVVIDNGKKSFQLDKKSGKKCFMLGARALFIAWDNGDHYRCWTTLP 121
SSKK Y L + ++IDNG+K F+++K SGK ++L +R L I W + HY W+
Sbjct: 61 FSSKKELYRCLCE-SILIDNGRKIFKIEKLSGKISYILSSRDLSITWSDQRHYWSWSPRS 119
Query: 122 ESRFPDVVVLRHLLWLEIHGMINTFSLSPNTQYGAFLVFKIID-APNFHEVLVVLSVNNI 180
+SRF + V L WLEI G I T +LSPNT YGA+L+ K+ A V S+
Sbjct: 120 DSRFSEXVQLIMTDWLEIIGKIQTGALSPNTNYGAYLIMKVTSRAYGLDLVPAETSIKVG 179
Query: 181 GGHNITKIAYLN-----PDSLE----GEVHDRW-------DRVQGPSVRSDGWLEIEMGE 224
G K YL+ +E G+ R + P VR DGW+EIE+GE
Sbjct: 180 NGEKKIKSTYLSCLDNKKQQMERVFYGQREQRMATHEVVRSHRREPEVRDDGWMEIELGE 239
Query: 225 FFI---SGLEDEEVEMSVLQIDGC-FNVGLIIEGIEVRPK 260
F G +D+EV MS+ ++ G G+ I+GIEVRPK
Sbjct: 240 FETGSGEGDDDKEVVMSLTEVKGYQLKGGIAIDGIEVRPK 279
>UniRef100_O80494 T12M4.17 protein [Arabidopsis thaliana]
Length = 288
Score = 173 bits (438), Expect = 5e-42
Identities = 112/280 (40%), Positives = 154/280 (55%), Gaps = 23/280 (8%)
Query: 3 LLPEGCIAAILSRTTPVDAARLSLVSKIFRSAADSDAVWDRFLPSDSHSIVSQSLSRANA 62
+LPE C+A ILS TTP D + VS +FR A DSD VW++FLP+D ++S+S
Sbjct: 1 MLPEACVATILSFTTPADTISSAAVSSVFRVAGDSDFVWEKFLPTDYCHVISRSTDPHRI 60
Query: 63 -SSKKRFYLDLSDHPVVIDNGKKSFQLDKKSGKKCFMLGARALFIAWDNGDHYRCWTTLP 121
SSKK Y L + ++IDNG+K F+++K SGK ++L +R L I W + HY W+
Sbjct: 61 FSSKKELYRCLCE-SILIDNGRKIFKIEKLSGKISYILSSRDLSITWSDQRHYWSWSPRS 119
Query: 122 ESRFPDVVVLRHLLWLEIHGMINTFSLSPNTQYGAFLVFKIID-APNFHEVLVVLSVNNI 180
+SRF + V L WLEI G I T +LSPNT YGA+L+ K+ A V S+
Sbjct: 120 DSRFSEGVQLIMTDWLEIIGKIQTGALSPNTNYGAYLIMKVTSRAYGLDLVPAETSIKVG 179
Query: 181 GGHNITKIAYLN-----PDSLE----GEVHDRW-------DRVQGPSVRSDGWLEIEMGE 224
G K YL+ +E G+ R + P VR DGW+EIE+GE
Sbjct: 180 NGEKKIKSTYLSCLDNKKQQMERVFYGQREQRMATHEVVRSHRREPEVRDDGWMEIELGE 239
Query: 225 FFI---SGLEDEEVEMSVLQIDGC-FNVGLIIEGIEVRPK 260
F G +D+EV MS+ ++ G G+ I+GIEVRPK
Sbjct: 240 FETGSGEGDDDKEVVMSLTEVKGYQLKGGIAIDGIEVRPK 279
>UniRef100_Q6K3E9 F-box family protein-like [Oryza sativa]
Length = 297
Score = 171 bits (434), Expect = 1e-41
Identities = 113/288 (39%), Positives = 157/288 (54%), Gaps = 38/288 (13%)
Query: 4 LPEGCIAAILSRTTPVDAARLSLVSKIFRSAADSDAVWDRFLPSDSHSIVSQSLSRANAS 63
LPE ++A +SR +P DA + VS FR+AADSDAVW FLP + + LS A AS
Sbjct: 14 LPEELLSAAISRASPRDACHAAAVSPAFRAAADSDAVWASFLPRNLPDLADGELSPAPAS 73
Query: 64 SKKRFYLDLSDHPVVIDNGKKSFQLDKKSGKKCFMLGARALFIAWDNGDHYRCWTTLPES 123
KK +L LSD P ++ + S LD+++G KC+ML AR+L I W + HY W L +S
Sbjct: 74 -KKELFLRLSDGPYLLSDRLMSMWLDRETGAKCYMLSARSLVIIWGDTPHYWRWIPLTDS 132
Query: 124 RFPDVVVLRHLLWLEIHGMINTFSLSPNTQYGAFLVFKI------IDAPNFHEVLVVLSV 177
RF + L + WLEI G I++ LSPN+ Y A++VFKI +DAP F E V
Sbjct: 133 RFAEGAELIDVCWLEIRGRIHSKMLSPNSTYAAYMVFKIADEFYGLDAP-FQEASV---- 187
Query: 178 NNIGGHNITKIAYLNP-DSLEGEVHDRW-----------------------DRVQGPSVR 213
++GG TKI + DS + EV + + + V P R
Sbjct: 188 -SLGGRGSTKIVCVQSYDSEDEEVPENYWPMSIGPLLRRRARRRDRRLVLDEGVTVPQKR 246
Query: 214 SDGWLEIEMGEFFISGLEDEEVEMSVLQI-DGCFNVGLIIEGIEVRPK 260
+D W+E+EMGEF ED EV S+++ G + GLI++GIE+R K
Sbjct: 247 TDEWMELEMGEFINEEGEDGEVCFSLMETKGGNWKRGLIVQGIEIRLK 294
>UniRef100_Q9C7J9 Hypothetical protein F14G9.15 [Arabidopsis thaliana]
Length = 284
Score = 167 bits (422), Expect = 3e-40
Identities = 110/281 (39%), Positives = 155/281 (55%), Gaps = 23/281 (8%)
Query: 1 MELLPEGCIAAILSRTTPVDAARLSLVSKIFRSAADSDAVWDRFLPSDSHSIVSQSLSRA 60
M +LPE C+A IL+ T+P DA S VS +FR A DSD VW++FLPS S++SQS
Sbjct: 1 MMMLPEACVANILAFTSPADAFSSSEVSSVFRLAGDSDFVWEKFLPSHYKSLISQSTDHH 60
Query: 61 NA-SSKKRFYLDLSDHPVVIDNGKKSFQLDKKSGKKCFMLGARALFIAWDNGDHYRCWTT 119
SSKK Y L D ++IDN +K F+++K SGK ++L AR + I + + Y W+
Sbjct: 61 RIFSSKKEIYRCLCDS-LLIDNARKLFKINKFSGKISYILSARDISITYSDHASYCSWSN 119
Query: 120 LPESRFPDVVVLRHLLWLEIHGMINTFSLSPNTQYGAFLVFKIIDAPNFHEVLV----VL 175
+ +SRF + L LEI G I T LSPNT+YGA+L+ K+ + +++ V
Sbjct: 120 VSDSRFSESAELITTDRLEIKGKIQTTVLSPNTKYGAYLIMKVTNGAYGLDLVPAETSVK 179
Query: 176 SVNNIGGHNITKIAYLNPDSLE------GEVHDRW----DRVQG------PSVRSDGWLE 219
S N N T + L+ + G +R + V G P R DGWLE
Sbjct: 180 SKNGQNNKNTTYLCCLDEKKQQMKRLFYGNREERMAMTVEAVGGDGKRREPKARDDGWLE 239
Query: 220 IEMGEFFISGLEDEEVEMSVLQIDGC-FNVGLIIEGIEVRP 259
IE+GEF ED+EV MS+ ++ G G++I+GIEVRP
Sbjct: 240 IELGEFVTREGEDDEVNMSLTEVKGYQLKGGIVIDGIEVRP 280
>UniRef100_Q6K3F6 Phloem-specific lectin-like [Oryza sativa]
Length = 332
Score = 162 bits (411), Expect = 6e-39
Identities = 115/309 (37%), Positives = 154/309 (49%), Gaps = 52/309 (16%)
Query: 4 LPEGCIAAILSRTTPVDAARLSLVSKIFRSAADSDAVWDRFLPSDSHSIVSQS----LSR 59
LPE C+A +S TTP DA S VS FR+AADSDAVWD FLP D +I++++ +
Sbjct: 14 LPEECVAYAISMTTPGDACHSSAVSPAFRAAADSDAVWDSFLPPDHAAILARADDGIAAA 73
Query: 60 ANASSKKRFYLDLSDHPVVIDNGKKSFQLDKKSGKKCFMLGARALFIAW-DNGDHYRCWT 118
+SKK + L PV++D+ SF LD++SG KC ML ARAL IAW D+ +R
Sbjct: 74 GECASKKDLFARLCGRPVLLDDATMSFGLDRRSGAKCVMLSARALSIAWGDDPSRWRWTP 133
Query: 119 TLPESRFPDVVVLRHLLWLEIHGMINTFSLSPNTQYGAFLVFKIIDAPNFHE-------- 170
LP SRFP+V L + WLEI G + LSP T Y A+LV+ D E
Sbjct: 134 GLPGSRFPEVAELLDVCWLEITGKLQLSLLSPATTYAAYLVYSFADYTTGLECNIGMPTP 193
Query: 171 VLVVLSVNNIGGHNIT--------------KIAYLNPDSLEGEVHD-----RWDRVQGPS 211
+ V V+ GG KI + E +H R + G +
Sbjct: 194 MATVTVVSGAGGTTSRPPAAPATTTTTEQHKICLQHMGEEETIMHRQELVIRLRKAFGRT 253
Query: 212 VRSD------------------GWLEIEMGEFFI--SGLEDEEVEMSVLQIDGCFNVGLI 251
VR D GW E+E+GEF + +G ED VE+S + G + GLI
Sbjct: 254 VRFDPDMDIRCPRPRDGGGGGGGWREVELGEFAVPAAGGEDGVVEVSFKEETGRWKTGLI 313
Query: 252 IEGIEVRPK 260
++GIE+RPK
Sbjct: 314 VQGIELRPK 322
>UniRef100_Q6K3F5 F-box family protein-like [Oryza sativa]
Length = 479
Score = 159 bits (402), Expect = 7e-38
Identities = 100/264 (37%), Positives = 146/264 (54%), Gaps = 13/264 (4%)
Query: 4 LPEGCIAAILSRTTPVDAARLSLVSKIFRSAADSDAVWDRFLPSDSHSIVSQSLSRANAS 63
LPE + A LS T+P DA + V + FR+AADSDAVW RFLP D + LS S
Sbjct: 15 LPEELLVAALSLTSPRDACSAAAVCRDFRAAADSDAVWSRFLPRDLPRLADGELSPPPPS 74
Query: 64 SKKRFYLDLSDHPVVIDNGKKSFQLDKKSGKKCFMLGARALFIAWDNGDHYRCWTTLPES 123
+K F L LS P+++ + S L+++ G KC+ML ARAL I W + Y W L +S
Sbjct: 75 TKGLF-LRLSAAPLLLPHELTSMWLEREKGGKCYMLSARALQITWGDTPRYWRWIPLTDS 133
Query: 124 RFPDVVVLRHLLWLEIHGMINTFSLSPNTQYGAFLVFKIIDAPNFHEVLVVLSVNNIGGH 183
R +L + WLEIHG I + LS NT Y A+LV++I D + + +IGG
Sbjct: 134 RLEGAELL-SVCWLEIHGKILSKMLSRNTNYAAYLVYRIADRSYGLDFPFQEASVSIGGS 192
Query: 184 NITKIAYLNPDSLEGEVHDRW-------DRVQGPSVRSDGWLEIEMGEFFISGLEDEEVE 236
T+ S+E + R + ++ P RSDGW+E+++GE + +D EV
Sbjct: 193 TTTR----QVGSVERRLKRRCSHALVLAEDIEHPQKRSDGWMELKLGELYNEEGDDGEVC 248
Query: 237 MSVLQIDGCFNVGLIIEGIEVRPK 260
+S + +G + GL+++GIE+RPK
Sbjct: 249 ISFRETEGHWKRGLVVQGIEIRPK 272
Score = 63.5 bits (153), Expect = 5e-09
Identities = 51/164 (31%), Positives = 82/164 (49%), Gaps = 13/164 (7%)
Query: 49 SHSIVSQSLSRANASSKKRFYLDLSDHPVVIDNGKKSFQLDKKSGKKCFMLGARALFIAW 108
SH S SL + SSK+ +L +G S LD ++G KC+ML ARAL +A
Sbjct: 283 SHEKPSYSLLTTSRSSKEEIFLT---------DGLTSMWLDMETGFKCYMLSARALQLA- 332
Query: 109 DNGDHYRCWTTLPESRFPDVVVLRHLLWLEIHGMINTFSLSPNTQYGAFLVFKIIDAPNF 168
++ D +R + SRF +V+ L L I G I L NT Y A++VF ++D +F
Sbjct: 333 NSTDTWRLISLTGASRFSEVIELPACYELVICGKIPCKMLPGNTNYAAYIVFVVVD-DSF 391
Query: 169 HEVLVVLSVNNIGGHNIT--KIAYLNPDSLEGEVHDRWDRVQGP 210
++ + ++GG T ++ + + SL + H D ++ P
Sbjct: 392 GLATILDASVSVGGSLCTTRQVCFDSTSSLSADEHFVEDNIEVP 435
>UniRef100_Q84PX1 Putative F-box protein [Oryza sativa]
Length = 299
Score = 151 bits (382), Expect = 1e-35
Identities = 101/275 (36%), Positives = 141/275 (50%), Gaps = 19/275 (6%)
Query: 4 LPEGCIAAILSRTTPVDAARLSLVSKIFRSAADSDAVWDRFLPSDSHSIVSQSLSRANAS 63
LPE C+A +++ T+P DA RL+ VS FR+AA+SDAVWDRFLP D +I A A+
Sbjct: 23 LPEACLADVIALTSPRDACRLAAVSPSFRAAAESDAVWDRFLPPDYRAIAPLPPPPATAA 82
Query: 64 S------KKRFYLDLSDHPVVIDNGKKSFQLDKKSGKKCFMLGARALFIAWDNGDHYRCW 117
+ KK YL L D PV +D+G L+K+SG KCF L AR L + W++G+ W
Sbjct: 83 ASGGKRMKKGVYLGLCDKPVPVDDGSMMVWLEKESGAKCFALPARKLSLPWEDGEFSWRW 142
Query: 118 TTLPESRFPDVVVLRHLLWLEIHGMINTFSLSPNTQYGAFLVFKIIDAPNFHEVLVV--- 174
T P SRF +V L L+I+G + +L+P T Y A+LVF A H L
Sbjct: 143 TPHPLSRFEEVAQLVDCTCLDIYGRLPAAALTPATPYAAYLVFGTAAAAEGHRGLSFPDQ 202
Query: 175 -LSVNNIGGHNITKIAYLNPDSLEGE------VHDRWDRVQGPSVRSDGWLEIEMGEFF- 226
+V+ G L PD E + V+ P+ R DGW E+E+G
Sbjct: 203 ETTVSAAGRVVARHAVCLRPDDAEARKFRGVGLAGAGVPVRRPARRGDGWSEMELGRVAA 262
Query: 227 --ISGLEDEEVEMSVLQIDGCFNVGLIIEGIEVRP 259
++G E+V S + GL++E +E RP
Sbjct: 263 DEVAGAGGEDVVASFEVLGWYPKRGLVVECMEFRP 297
>UniRef100_Q9ZVQ9 Hypothetical protein At2g02320 [Arabidopsis thaliana]
Length = 307
Score = 146 bits (368), Expect = 6e-34
Identities = 99/263 (37%), Positives = 145/263 (54%), Gaps = 14/263 (5%)
Query: 4 LPEGCIAAILSRTTPVDAARLSLVSKIFRSAADSDAVWDRFLPSDSHSIVSQSLSRANAS 63
LPE CI+ I+S T+P DA +LVSK F SA SD VW++F+P + S++S+S + S
Sbjct: 43 LPEECISLIISFTSPRDACVFALVSKTFESAVQSDIVWEKFIPPEYESLLSRS---QHFS 99
Query: 64 SKKRFYLDLSDHPVVIDNGKKSFQLDKKSGKKCFMLGARALFIAWDNGDHYRCWTTLPES 123
SKK + L D V+I+ KK ++K +GK+C ML A AL + + H W T P S
Sbjct: 100 SKKELFFALCDESVLINVSKKDLWIEKATGKRCMMLSASALNL---STHHTWKWITNPVS 156
Query: 124 RFPDVV-VLRHLLWLEIHGMINTFSLSPNTQYGAFLVFKIIDAPNFHEVLVVLSVNNIGG 182
+ + V L W EI NT LSP T+Y ++VF D + + +V + G
Sbjct: 157 AWLETVPELLTTRWFEIRCRTNTRFLSPRTRYSVYIVFLKADICYGFAYVAMEAVVRMVG 216
Query: 183 HNITKIA---YLNPDSLEGEVHDRWDRVQGPSVRSDGWLEIEMGEFFISGL--EDEEVEM 237
H +++ +++E + R + V P R DGW+EIE+GEFF G ++E+EM
Sbjct: 217 HELSESCRRYVCFHEAMEWQFLTRKNLV-NPERREDGWMEIEIGEFFNEGAFRNNDEIEM 275
Query: 238 SVLQ-IDGCFNVGLIIEGIEVRP 259
SV + GLII+GIE+RP
Sbjct: 276 SVSETTQRNTKRGLIIQGIEIRP 298
>UniRef100_Q8GUU0 Hypothetical protein T16F16.11 [Arabidopsis thaliana]
Length = 307
Score = 145 bits (366), Expect = 1e-33
Identities = 99/263 (37%), Positives = 144/263 (54%), Gaps = 14/263 (5%)
Query: 4 LPEGCIAAILSRTTPVDAARLSLVSKIFRSAADSDAVWDRFLPSDSHSIVSQSLSRANAS 63
LPE CI+ I+S T+P DA +LVSK F SA SD VW++F+P + S++S+S + S
Sbjct: 43 LPEECISLIISFTSPRDACVFALVSKTFESAVQSDIVWEKFIPPEYESLLSRS---QHFS 99
Query: 64 SKKRFYLDLSDHPVVIDNGKKSFQLDKKSGKKCFMLGARALFIAWDNGDHYRCWTTLPES 123
SKK + L D V+I+ KK ++K +GK+C ML A AL + + H W T P S
Sbjct: 100 SKKELFFALCDESVLINVSKKDLWIEKATGKRCMMLSASALNL---STHHTWKWITNPVS 156
Query: 124 RFPDVV-VLRHLLWLEIHGMINTFSLSPNTQYGAFLVFKIIDAPNFHEVLVVLSVNNIGG 182
+ + V L W EI NT LSP T+Y ++VF D + + +V + G
Sbjct: 157 AWLETVPELLTTRWFEIRCRTNTRFLSPRTRYSVYIVFLKADICYGFAYVAMEAVVRMVG 216
Query: 183 HNITKIA---YLNPDSLEGEVHDRWDRVQGPSVRSDGWLEIEMGEFFISGL--EDEEVEM 237
H +++ +++E + R + V P R DGW+EIE+GEFF G ++E+EM
Sbjct: 217 HELSESCRRYVCFHEAMEWQFLTRKNLV-NPERREDGWMEIEIGEFFNEGAFRNNDEIEM 275
Query: 238 SVLQ-IDGCFNVGLIIEGIEVRP 259
SV + GLII GIE+RP
Sbjct: 276 SVSETTQRNSKRGLIIRGIEIRP 298
>UniRef100_Q9ZVR4 Putative phloem-specific lectin [Arabidopsis thaliana]
Length = 265
Score = 144 bits (364), Expect = 2e-33
Identities = 96/265 (36%), Positives = 135/265 (50%), Gaps = 47/265 (17%)
Query: 4 LPEGCIAAILSRTTPVDAARLSLVSKIFRSAADSDAVWDRFLPSDSHSIVSQSLSRANAS 63
LPE CI+ I+S TTP DA + VSK F SA SD+VW++FLP D S+V +S
Sbjct: 16 LPENCISNIISFTTPRDACFAASVSKAFESAVQSDSVWEKFLPLDYSSLVPES---RVFL 72
Query: 64 SKKRFYLDLSDHPVVIDNGKKSFQLDKKSGKKCFMLGARALFIAWDNGDHYRCWTTLPES 123
SKK L P++I+ GKKSF LDK SG+KC ML + + I+W N
Sbjct: 73 SKKELCFSLCRVPLLIEGGKKSFWLDKTSGEKCIMLSPKGMVISWVN-----------SP 121
Query: 124 RFPDVVVLRHLLWLEIHGMINTFSLSPNTQYGAFLVFKIIDA---------PNFHEVLVV 174
+F +V L + W E+ G ++T LSP T+Y ++VFK D P F
Sbjct: 122 QFEEVPELLYDSWFEVCGRLSTKYLSPRTRYSVYIVFKTNDLYPGVTLEPFPRF------ 175
Query: 175 LSVNNIGGHNITKIAYLNPDSLEGEVHDRWDRVQGPSVRSDGWLEIEMGEFFISGLEDEE 234
+ ++ P L+ E + V P R D W+E E+GEFF + +
Sbjct: 176 -------------VRFVGPTDLKYE----REYVTRPEKREDKWMEAELGEFF-NETSCGD 217
Query: 235 VEMSVLQIDGCFNVGLIIEGIEVRP 259
VE+SV+ + + GL+I+GIE RP
Sbjct: 218 VEVSVIDENSYWKSGLVIQGIEFRP 242
>UniRef100_Q9ZVR0 Putative phloem-specific lectin [Arabidopsis thaliana]
Length = 307
Score = 141 bits (355), Expect = 2e-32
Identities = 91/261 (34%), Positives = 137/261 (51%), Gaps = 6/261 (2%)
Query: 4 LPEGCIAAILSRTTPVDAARLSLVSKIFRSAADSDAVWDRFLPSDSHSIVSQSLSRANAS 63
LPE CI+ I+S T+P D + VSK F A D++W++FLPS+ S++ S
Sbjct: 48 LPEDCISNIISFTSPRDVCVSASVSKSFAHAVQCDSIWEKFLPSEYESLIPPWRV---FS 104
Query: 64 SKKRFYLDLSDHPVVIDNGKKSFQLDKKSGKKCFMLGARALFIAWDNGDHYRCWTTLPES 123
SKK Y L PV++++GKKSF L+ SGKKC +L A+ L+I N Y W L ES
Sbjct: 105 SKKDLYFTLCYDPVLVEDGKKSFWLETASGKKCVLLAAKELWITGGNNPEYWQWIELCES 164
Query: 124 RFPDVVVLRHLLWLEIHGMINTFSLSPNTQYGAFLVFKIIDAPNFHEVLVVLSVNNIGGH 183
F V L + ++ G ++T LS T Y ++V+KI D + L + G
Sbjct: 165 SFEKVPELLNNRSFQMGGSMSTQILSLGTHYSVYIVYKIKDERHGLRDLPIQVGVGFKGQ 224
Query: 184 NITKIAYLNPDSLEGEVHDRWDRVQGPSVRSDGWLEIEMGEFFISG--LEDEEVEMSVLQ 241
+ K +S + ++ R DGW+E E+G+FF G + +EVE+S++
Sbjct: 225 EMPKQFICFDESTDKTKEWPKKKLMKSKKRGDGWMEAEIGDFFNDGGLMGFDEVEVSIVD 284
Query: 242 IDG-CFNVGLIIEGIEVRPKN 261
+ G++IEGIE RPK+
Sbjct: 285 VTSPNLKCGVMIEGIEFRPKD 305
>UniRef100_Q9SSC8 F18B13.19 protein [Arabidopsis thaliana]
Length = 264
Score = 137 bits (344), Expect = 4e-31
Identities = 95/265 (35%), Positives = 138/265 (51%), Gaps = 9/265 (3%)
Query: 1 MELLPEGCIAAILSRTTPVDAARLSLVSKIFR---SAADSDAVWDRFL--PSDSHSIVSQ 55
M LPE CIA ILS TTP+D RLS VS I + S + R + P+ H +V
Sbjct: 1 MNNLPEDCIAKILSLTTPLDVCRLSAVSSISDPPPAPTMSGIISYRLISPPALRHLLVFP 60
Query: 56 SLSRANASSKKRFYLDLSDHPVVIDNGKKSFQLDKKSGKKCFMLGARALFIAWDNGDHYR 115
S +++ S +L + + + K+SF L++KSG KC+M+ ARAL I W + Y
Sbjct: 61 PGSNSSSPSSITLFLSTALF-CLTELYKQSFSLERKSGNKCYMMAARALNIVWGHEQRYW 119
Query: 116 CWTTLPESRFPDVVVLRHLLWLEIHGMINTFSLSPNTQYGAFLVFKIIDAPNFHEVLVVL 175
W +LP +RF +V L + WLEI G IN LS +T Y A+ VFK +P V
Sbjct: 120 HWISLPNTRFGEVAELIMVWWLEITGKINITLLSDDTLYAAYFVFKWNHSPYGFRQPVET 179
Query: 176 SVNNIGGHNITKIAYLNPDSLEGEVHDRWDRVQGPSVRSDGWLEIEMGEFFISGLEDEEV 235
S+ + + + P + + Q P +R DGW E+E+G+FF + E+
Sbjct: 180 SLVLADTESTDNV--VQPSMISLMQDSGGEEGQSPVLRRDGWYEVELGQFFKRRGDLGEI 237
Query: 236 EMSVLQIDGCF-NVGLIIEGIEVRP 259
EMS+ + G + GLI+ GIE+RP
Sbjct: 238 EMSLKETKGPYEKKGLIVYGIEIRP 262
>UniRef100_Q9ZVR1 Putative phloem-specific lectin [Arabidopsis thaliana]
Length = 284
Score = 135 bits (341), Expect = 8e-31
Identities = 97/265 (36%), Positives = 139/265 (51%), Gaps = 26/265 (9%)
Query: 4 LPEGCIAAILSRT-TPVDAARLSLVSKIFRSAADSDAVWDRFLPSDSHSIVSQSLSRANA 62
LP+ C+A I S T TP DA +LVSK F +SD+VW++FLP + VS
Sbjct: 37 LPDDCLAIISSFTSTPRDAFLAALVSKSFGLQFNSDSVWEKFLPPPDY--VSLLPKSRVF 94
Query: 63 SSKKRFYLDLSDHPVVIDNGKKSFQLDKKSGKKCFMLGARALFIAWDNGDHYRCWTTLPE 122
SSKK Y L D P NGK SF+LDK SGKKC ML A+ L I+ Y W ++PE
Sbjct: 95 SSKKELYFALCD-PFPNHNGKMSFRLDKASGKKCVMLSAKKLLISRVVNPKYWKWISIPE 153
Query: 123 SRFPDVVVLRHLLWLEIHGMINTFSLSPNTQYGAFLVFKIIDAPNFHEVLVVLSVNNIGG 182
SRF +V L ++ +I G++NT +SP T Y A++V+ N + + +
Sbjct: 154 SRFDEVPELLNIDSFDIRGVLNTRIISPGTHYSAYIVYTKTSHFNGFQTSPIQAGVGFQR 213
Query: 183 HNITKIAYLNPDSLEGEVHDRWDRVQGPSVRSDGWLEIEMGEFFISG--LEDEEVEMSVL 240
H ++K ++ DS + R DGW+E ++G+F+ G + +E+SV+
Sbjct: 214 HGMSK-TFIRFDSKK---------------RQDGWMEAKIGDFYNEGGLIGFNLIEVSVV 257
Query: 241 QIDGC----FNVGLIIEGIEVRPKN 261
+ GLIIEGIE RPK+
Sbjct: 258 DVARYPHMNMKSGLIIEGIEFRPKD 282
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.321 0.138 0.421
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 447,434,331
Number of Sequences: 2790947
Number of extensions: 18321155
Number of successful extensions: 35145
Number of sequences better than 10.0: 85
Number of HSP's better than 10.0 without gapping: 46
Number of HSP's successfully gapped in prelim test: 39
Number of HSP's that attempted gapping in prelim test: 34956
Number of HSP's gapped (non-prelim): 101
length of query: 263
length of database: 848,049,833
effective HSP length: 125
effective length of query: 138
effective length of database: 499,181,458
effective search space: 68887041204
effective search space used: 68887041204
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 73 (32.7 bits)
Lotus: description of TM0226.6


