

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0226.15
(219 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
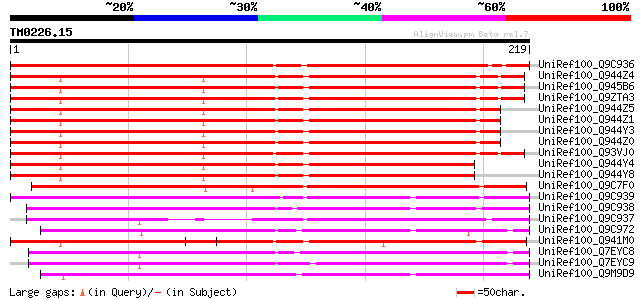
Sequences producing significant alignments: (bits) Value
UniRef100_Q9C936 Putative oxidoreductase; 38288-39393 [Arabidops... 249 4e-65
UniRef100_Q944Z4 2-oxoglutarate-dependent dioxygenase [Arabidops... 217 2e-55
UniRef100_Q945B6 AOP1.2 [Arabidopsis thaliana] 214 1e-54
UniRef100_Q9ZTA3 Putative oxidoreductase [Arabidopsis thaliana] 214 1e-54
UniRef100_Q944Z5 2-oxoglutarate-dependent dioxygenase [Arabidops... 214 2e-54
UniRef100_Q944Z1 2-oxoglutarate-dependent dioxygenase [Arabidops... 211 1e-53
UniRef100_Q944Y3 2-oxoglutarate-dependent dioxygenase [Arabidops... 211 1e-53
UniRef100_Q944Z0 2-oxoglutarate-dependent dioxygenase [Arabidops... 207 2e-52
UniRef100_Q93VJ0 AOP1 [Arabidopsis lyrata] 206 3e-52
UniRef100_Q944Y4 2-oxoglutarate-dependent dioxygenase [Arabidops... 206 5e-52
UniRef100_Q944Y8 2-oxoglutarate-dependent dioxygenase [Arabidops... 204 1e-51
UniRef100_Q9C7F0 Oxidoreductase, putative [Arabidopsis thaliana] 187 2e-46
UniRef100_Q9C939 Putative oxidoreductase; 32373-31266 [Arabidops... 161 1e-38
UniRef100_Q9C938 Putative oxidoreductase; 33116-34434 [Arabidops... 157 1e-37
UniRef100_Q9C937 Putative oxidoreductase; 36199-37309 [Arabidops... 147 1e-34
UniRef100_Q9C972 Putative oxidoreductase; 24302-25416 [Arabidops... 141 1e-32
UniRef100_Q941M0 2-oxoglutarate-dependent dioxygenase [Brassica ... 135 8e-31
UniRef100_Q7EYC8 Putative 2-oxoglutarate-dependent dioxygenase [... 134 2e-30
UniRef100_Q7EYC9 Putative 2-oxoglutarate-dependent dioxygenase [... 127 3e-28
UniRef100_Q9M9D9 T16N11.5 protein [Arabidopsis thaliana] 126 5e-28
>UniRef100_Q9C936 Putative oxidoreductase; 38288-39393 [Arabidopsis thaliana]
Length = 317
Score = 249 bits (636), Expect = 4e-65
Identities = 123/219 (56%), Positives = 163/219 (74%), Gaps = 5/219 (2%)
Query: 1 VKNTSVYEKVETLTKILWPNGNPSFRYALCSFSEHLSELDQIVRKMVLESLGVEKYLDEH 60
+ ++ + EKV+ T+ LWP GN SF + SFS+ LSELD +R+M++ES G++KY+DEH
Sbjct: 103 IDDSDIAEKVDAFTEKLWPQGNISFSTTIQSFSKKLSELDITIRRMIMESFGLDKYIDEH 162
Query: 61 LNSTNYFLKTMKYKGPQTSDTKVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKP 120
L+STNY L+ MKYKGP T +TKVGL+ HTD I+TIL+Q+ V+GLEV TKD + WI KP
Sbjct: 163 LHSTNYLLRVMKYKGPDTEETKVGLNAHTDKNIVTILYQNHVEGLEVQTKD-KNWIKVKP 221
Query: 121 SSESQSFIVMMNDCFHAWSNGRLNSPFHRVMMTGNEARYSAALFSVPKGGYIIKAPEELV 180
+ + SF VM+ D +A NGRL+SP+HRVMMTG E RYS LFS+PK G+I+ +P+ELV
Sbjct: 222 TQD--SFTVMIGDSLYALLNGRLHSPYHRVMMTGTETRYSLGLFSIPKAGHIVSSPDELV 279
Query: 181 DEEHPLLYKPFDLEEFLKHSFNKEKGKRDDQFALHSYCG 219
DEEHP L+KPFD EFL+ + E G+R Q AL +YCG
Sbjct: 280 DEEHPRLFKPFDHVEFLQFYYT-EAGQR-SQSALKTYCG 316
>UniRef100_Q944Z4 2-oxoglutarate-dependent dioxygenase [Arabidopsis thaliana]
Length = 320
Score = 217 bits (553), Expect = 2e-55
Identities = 119/222 (53%), Positives = 151/222 (67%), Gaps = 10/222 (4%)
Query: 1 VKNTSVYEKVETLTKILWPN-GNPSFRYALCSFSEHLSELDQIVRKMVLESLGVEKYLDE 59
+ + +V EKV T+ LWP+ GN S + FSE L ELD +VR+M++ES G+EKY+DE
Sbjct: 101 INDANVLEKVNDFTQQLWPDHGNKSISETIHLFSEQLVELDLMVRRMIMESFGIEKYIDE 160
Query: 60 HLNSTNYFLKTMKYKGPQTSD----TKVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKW 115
HLNST Y + MKY P D TK+GL +HTD I+TIL Q QVDGLEV TKD KW
Sbjct: 161 HLNSTYYLTRLMKYTSPPDDDDDEETKLGLRSHTDKNIITILHQYQVDGLEVKTKDD-KW 219
Query: 116 ISYKPSSESQSFIVMMNDCFHAWSNGRLNSPFHRVMMTGNEARYSAALFSVPKGGYIIKA 175
I KPS +S +VM+ D A NGRL+SP+HRV+MTG + RYS LFS+PK G II +
Sbjct: 220 IKVKPSQDS--VLVMVGDSLCALLNGRLHSPYHRVIMTGKKTRYSTGLFSIPKTGVIIDS 277
Query: 176 PEELVDEEHPLLYKPFDLEEFLKHSFNKEKGKRDDQFALHSY 217
PEELVD+EHP ++KPF+ +FL H F E G R Q ALH++
Sbjct: 278 PEELVDKEHPRIFKPFEYTDFL-HFFQTEAG-RIAQSALHAF 317
>UniRef100_Q945B6 AOP1.2 [Arabidopsis thaliana]
Length = 321
Score = 214 bits (546), Expect = 1e-54
Identities = 118/223 (52%), Positives = 150/223 (66%), Gaps = 11/223 (4%)
Query: 1 VKNTSVYEKVETLTKILWPN-GNPSFRYALCSFSEHLSELDQIVRKMVLESLGVEKYLDE 59
+ + +V EKV T+ LWP+ GN S + FSE L ELD +VR+M++ES G+E Y+DE
Sbjct: 101 IDDANVLEKVNDFTQQLWPDHGNKSISETIHLFSEQLVELDLMVRRMIMESFGIENYIDE 160
Query: 60 HLNSTNYFLKTMKYKGPQTSD-----TKVGLDTHTDTAILTILFQSQVDGLEVLTKDGRK 114
HLNST Y + MKY P D TK+GL +HTD I+TIL Q QVDGLEV TKD K
Sbjct: 161 HLNSTYYLTRLMKYTSPPDDDDDDEETKLGLRSHTDKNIITILHQYQVDGLEVKTKDD-K 219
Query: 115 WISYKPSSESQSFIVMMNDCFHAWSNGRLNSPFHRVMMTGNEARYSAALFSVPKGGYIIK 174
WI KPS +S +VM+ D A NGRL+SP+HRV+MTG + RYS LFS+PK G II
Sbjct: 220 WIKVKPSQDS--VLVMVGDSLCALLNGRLHSPYHRVIMTGKKTRYSTGLFSIPKTGVIID 277
Query: 175 APEELVDEEHPLLYKPFDLEEFLKHSFNKEKGKRDDQFALHSY 217
+PEELVD+EHP ++KPF+ +FL H F E G R Q ALH++
Sbjct: 278 SPEELVDKEHPRIFKPFEYTDFL-HFFQTEAG-RIAQSALHAF 318
>UniRef100_Q9ZTA3 Putative oxidoreductase [Arabidopsis thaliana]
Length = 322
Score = 214 bits (546), Expect = 1e-54
Identities = 118/224 (52%), Positives = 150/224 (66%), Gaps = 12/224 (5%)
Query: 1 VKNTSVYEKVETLTKILWPN-GNPSFRYALCSFSEHLSELDQIVRKMVLESLGVEKYLDE 59
+ + +V EKV T+ LWP+ GN S + FSE L ELD +VR+M++ES G+E Y+DE
Sbjct: 101 INDANVLEKVNDFTQQLWPDHGNKSISETIHLFSEQLVELDLMVRRMIMESFGIENYIDE 160
Query: 60 HLNSTNYFLKTMKYKGPQTSD------TKVGLDTHTDTAILTILFQSQVDGLEVLTKDGR 113
HLNST Y + MKY P D TK+GL +HTD I+TIL Q QVDGLEV TKD
Sbjct: 161 HLNSTYYLTRLMKYTSPPDDDDDDDEETKLGLRSHTDKNIITILHQYQVDGLEVKTKDD- 219
Query: 114 KWISYKPSSESQSFIVMMNDCFHAWSNGRLNSPFHRVMMTGNEARYSAALFSVPKGGYII 173
KWI KPS +S +VM+ D A NGRL+SP+HRV+MTG + RYS LFS+PK G II
Sbjct: 220 KWIKVKPSQDS--VLVMVGDSLCALLNGRLHSPYHRVIMTGKKTRYSTGLFSIPKTGVII 277
Query: 174 KAPEELVDEEHPLLYKPFDLEEFLKHSFNKEKGKRDDQFALHSY 217
+PEELVD+EHP ++KPF+ +FL H F E G R Q ALH++
Sbjct: 278 DSPEELVDKEHPRIFKPFEYTDFL-HFFQTEAG-RIAQSALHAF 319
>UniRef100_Q944Z5 2-oxoglutarate-dependent dioxygenase [Arabidopsis thaliana]
Length = 301
Score = 214 bits (544), Expect = 2e-54
Identities = 114/212 (53%), Positives = 145/212 (67%), Gaps = 9/212 (4%)
Query: 1 VKNTSVYEKVETLTKILWPN-GNPSFRYALCSFSEHLSELDQIVRKMVLESLGVEKYLDE 59
+ + +V EKV T+ LWP+ GN S + FSE L ELD +VR+M++ES G+EKY+DE
Sbjct: 94 INDANVLEKVNDFTQQLWPDHGNKSISETIHLFSEQLVELDLMVRRMIMESFGIEKYIDE 153
Query: 60 HLNSTNYFLKTMKYKGPQTSD----TKVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKW 115
HLNST Y + MKY P D TK+GL +HTD I+TIL Q QVDGLEV TKD KW
Sbjct: 154 HLNSTYYLTRLMKYTSPPDDDDDEETKLGLRSHTDKNIITILHQYQVDGLEVKTKDD-KW 212
Query: 116 ISYKPSSESQSFIVMMNDCFHAWSNGRLNSPFHRVMMTGNEARYSAALFSVPKGGYIIKA 175
I KPS +S +VM+ D A NGRL+SP+HRV+MTG + RYS LFS+PK G II +
Sbjct: 213 IKVKPSQDS--VLVMVGDSLCALLNGRLHSPYHRVIMTGKKTRYSTGLFSIPKTGVIIDS 270
Query: 176 PEELVDEEHPLLYKPFDLEEFLKHSFNKEKGK 207
PEELVD+EHP ++KPF+ +FL H F E G+
Sbjct: 271 PEELVDKEHPRIFKPFEYTDFL-HFFQTEAGR 301
>UniRef100_Q944Z1 2-oxoglutarate-dependent dioxygenase [Arabidopsis thaliana]
Length = 309
Score = 211 bits (537), Expect = 1e-53
Identities = 113/214 (52%), Positives = 144/214 (66%), Gaps = 11/214 (5%)
Query: 1 VKNTSVYEKVETLTKILWPN-GNPSFRYALCSFSEHLSELDQIVRKMVLESLGVEKYLDE 59
+ + +V EKV T+ LWP+ GN S + FSE L ELD +VR+M++ES G+E Y+DE
Sbjct: 95 INDANVLEKVNDFTQQLWPDHGNKSISETIHLFSEQLVELDLMVRRMIMESFGIENYIDE 154
Query: 60 HLNSTNYFLKTMKYKGPQTSD------TKVGLDTHTDTAILTILFQSQVDGLEVLTKDGR 113
HLNST Y + MKY P D TK+GL +HTD I+TIL Q QVDGLEV TKD
Sbjct: 155 HLNSTYYLTRLMKYTSPPDDDDDDDEETKLGLRSHTDKNIITILHQYQVDGLEVKTKDD- 213
Query: 114 KWISYKPSSESQSFIVMMNDCFHAWSNGRLNSPFHRVMMTGNEARYSAALFSVPKGGYII 173
KWI KPS +S +VM+ D A NGRL+SP+HRV+MTG + RYS LFS+PK G II
Sbjct: 214 KWIKVKPSQDS--VLVMVGDSLCALLNGRLHSPYHRVIMTGKKTRYSTGLFSIPKTGVII 271
Query: 174 KAPEELVDEEHPLLYKPFDLEEFLKHSFNKEKGK 207
+PEELVD+EHP ++KPF+ +FL H F E G+
Sbjct: 272 DSPEELVDKEHPRIFKPFEYTDFL-HFFQTEAGR 304
>UniRef100_Q944Y3 2-oxoglutarate-dependent dioxygenase [Arabidopsis thaliana]
Length = 301
Score = 211 bits (536), Expect = 1e-53
Identities = 113/215 (52%), Positives = 144/215 (66%), Gaps = 12/215 (5%)
Query: 1 VKNTSVYEKVETLTKILWPN-GNPSFRYALCSFSEHLSELDQIVRKMVLESLGVEKYLDE 59
+ + +V EKV T+ LWP+ GN S + FSE L ELD +VR+M++ES G+E Y+DE
Sbjct: 85 INDANVLEKVNDFTQQLWPDHGNKSISETIHRFSEKLVELDLMVRRMIMESFGIENYIDE 144
Query: 60 HLNSTNYFLKTMKYKGPQTSD-------TKVGLDTHTDTAILTILFQSQVDGLEVLTKDG 112
HLNST Y + MKY P D TK+GL +HTD I+TIL Q QVDGLEV TKD
Sbjct: 145 HLNSTYYLTRLMKYTSPPDDDDDDDDEETKLGLRSHTDKNIITILHQYQVDGLEVKTKDD 204
Query: 113 RKWISYKPSSESQSFIVMMNDCFHAWSNGRLNSPFHRVMMTGNEARYSAALFSVPKGGYI 172
KWI KPS +S +VM+ D A NGRL+SP+HRV+MTG + RYS LFS+PK G I
Sbjct: 205 -KWIKVKPSQDS--VLVMVGDSLCALLNGRLHSPYHRVIMTGKKTRYSTGLFSIPKTGVI 261
Query: 173 IKAPEELVDEEHPLLYKPFDLEEFLKHSFNKEKGK 207
I +PEELVD+EHP ++KPF+ +FL H F E G+
Sbjct: 262 IDSPEELVDKEHPRIFKPFEYTDFL-HFFQTEAGR 295
>UniRef100_Q944Z0 2-oxoglutarate-dependent dioxygenase [Arabidopsis halleri]
Length = 294
Score = 207 bits (526), Expect = 2e-52
Identities = 107/211 (50%), Positives = 146/211 (68%), Gaps = 8/211 (3%)
Query: 1 VKNTSVYEKVETLTKILWPN-GNPSFRYALCSFSEHLSELDQIVRKMVLESLGVEKYLDE 59
+ + +V EKV+ T+ LWP+ GN + + FSE L ELD +VR+M++ES G+EKY+DE
Sbjct: 88 IDDANVLEKVDDFTQQLWPDQGNKTISETIHRFSEQLVELDVMVRRMIMESFGIEKYIDE 147
Query: 60 HLNSTNYFLKTMKYKGPQTSD---TKVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWI 116
H+NS+NY ++ MKY P D TK+GL +HTD I+TIL Q QV GLEV TKD KWI
Sbjct: 148 HVNSSNYLMRFMKYTSPPDDDDEETKLGLRSHTDKNIITILHQYQVGGLEVKTKDD-KWI 206
Query: 117 SYKPSSESQSFIVMMNDCFHAWSNGRLNSPFHRVMMTGNEARYSAALFSVPKGGYIIKAP 176
+PS +S ++M+ D A NGRL SP+HRV+M + RYS LFS+PK G II +P
Sbjct: 207 KVQPSQDS--VLIMLGDSLCALLNGRLPSPYHRVIMMSKKTRYSTGLFSIPKTGVIIDSP 264
Query: 177 EELVDEEHPLLYKPFDLEEFLKHSFNKEKGK 207
EELVDEEHP ++KPF+ ++F+ H F+ E G+
Sbjct: 265 EELVDEEHPRIFKPFEYKDFI-HFFHTEAGR 294
>UniRef100_Q93VJ0 AOP1 [Arabidopsis lyrata]
Length = 318
Score = 206 bits (524), Expect = 3e-52
Identities = 114/221 (51%), Positives = 147/221 (65%), Gaps = 9/221 (4%)
Query: 1 VKNTSVYEKVETLTKILWPN-GNPSFRYALCSFSEHLSELDQIVRKMVLESLGVEKYLDE 59
+ + +V KV T+ LWP+ GN S + FS+ L ELD +VR+M++ES G+EKY+DE
Sbjct: 100 IDDANVLVKVNDFTQQLWPDHGNKSISETIHRFSDQLLELDVMVRRMIMESFGIEKYIDE 159
Query: 60 HLNSTNYFLKTMKYKGPQTSD---TKVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWI 116
HL STNY + MKY P +D T+VGL HTD I+TIL Q Q++GLEV TK+ WI
Sbjct: 160 HLKSTNYLFRMMKYTTPLDADVEETQVGLRAHTDKNIITILHQYQINGLEVKTKN-ENWI 218
Query: 117 SYKPSSESQSFIVMMNDCFHAWSNGRLNSPFHRVMMTGNEARYSAALFSVPKGGYIIKAP 176
KPS +S F+VM+ D A NGRL S +HRVMM GN+ RYS ALFS PK II +P
Sbjct: 219 KVKPSQDS--FVVMVGDSLCALLNGRLPSSYHRVMMMGNKTRYSTALFSAPKLETIIDSP 276
Query: 177 EELVDEEHPLLYKPFDLEEFLKHSFNKEKGKRDDQFALHSY 217
EELVDEEHP ++KPFD +F+ H F+ E G R Q LH++
Sbjct: 277 EELVDEEHPRMFKPFDYNDFV-HFFHTEAG-RKAQSTLHAF 315
>UniRef100_Q944Y4 2-oxoglutarate-dependent dioxygenase [Arabidopsis thaliana]
Length = 272
Score = 206 bits (523), Expect = 5e-52
Identities = 108/202 (53%), Positives = 138/202 (67%), Gaps = 9/202 (4%)
Query: 1 VKNTSVYEKVETLTKILWPN-GNPSFRYALCSFSEHLSELDQIVRKMVLESLGVEKYLDE 59
+ + +V EKV T+ LWP+ GN S + FSE L ELD +VR+M++ES G+E Y+DE
Sbjct: 74 INDANVLEKVNDFTQQLWPDHGNKSISETIHLFSEQLVELDLMVRRMIMESFGIENYIDE 133
Query: 60 HLNSTNYFLKTMKYKGPQTSD-----TKVGLDTHTDTAILTILFQSQVDGLEVLTKDGRK 114
HLNST Y + MKY P D TK+GL +HTD I+TIL Q QVDGLEV TKD K
Sbjct: 134 HLNSTYYLTRLMKYTSPPDDDDDDEETKLGLRSHTDKNIITILHQYQVDGLEVKTKDD-K 192
Query: 115 WISYKPSSESQSFIVMMNDCFHAWSNGRLNSPFHRVMMTGNEARYSAALFSVPKGGYIIK 174
WI KPS +S +VM+ D A NGRL+SP+HRV+MTG + RYS LFS+PK G II
Sbjct: 193 WIKVKPSQDS--VLVMVGDSLCALLNGRLHSPYHRVIMTGKKTRYSTGLFSIPKTGVIID 250
Query: 175 APEELVDEEHPLLYKPFDLEEF 196
+PEELVD+EHP ++KPF+ +F
Sbjct: 251 SPEELVDKEHPRIFKPFEYTDF 272
>UniRef100_Q944Y8 2-oxoglutarate-dependent dioxygenase [Arabidopsis thaliana]
Length = 267
Score = 204 bits (519), Expect = 1e-51
Identities = 108/206 (52%), Positives = 138/206 (66%), Gaps = 13/206 (6%)
Query: 1 VKNTSVYEKVETLTKILWPN-GNPSFRYALCSFSEHLSELDQIVRKMVLESLGVEKYLDE 59
+ + +V EKV T+ LWP+ GN S + FSE L ELD +VR+M++ES G+E Y+DE
Sbjct: 65 INDANVLEKVNDFTQQLWPDHGNKSISETIHRFSEKLVELDLMVRRMIMESFGIENYIDE 124
Query: 60 HLNSTNYFLKTMKYKGPQTSD---------TKVGLDTHTDTAILTILFQSQVDGLEVLTK 110
HLNST Y + MKY P D TK+GL +HTD I+TIL Q QVDGLEV TK
Sbjct: 125 HLNSTYYLTRLMKYTSPPDDDDDDDDDDEETKLGLRSHTDKNIITILHQYQVDGLEVKTK 184
Query: 111 DGRKWISYKPSSESQSFIVMMNDCFHAWSNGRLNSPFHRVMMTGNEARYSAALFSVPKGG 170
D KWI KPS +S +VM+ D A NGRL+SP+HRV+MTG + RYS LFS+PK G
Sbjct: 185 DD-KWIKVKPSQDS--VLVMVGDSLCALLNGRLHSPYHRVIMTGKKTRYSTGLFSIPKTG 241
Query: 171 YIIKAPEELVDEEHPLLYKPFDLEEF 196
II +PEELVD+EHP ++KPF+ +F
Sbjct: 242 VIIDSPEELVDKEHPRIFKPFEYTDF 267
>UniRef100_Q9C7F0 Oxidoreductase, putative [Arabidopsis thaliana]
Length = 322
Score = 187 bits (475), Expect = 2e-46
Identities = 107/212 (50%), Positives = 130/212 (60%), Gaps = 6/212 (2%)
Query: 10 VETLTKILWPNGNPSFRYALCSFSEHLSELDQIVRKMVLESLGVEKYLDEHLNSTNYFLK 69
V LT LWP GN + SF+E L EL+ VR M LES G+EKY++EHLN+ N +
Sbjct: 112 VNDLTHKLWPQGNIFVGKNVQSFAEKLIELNLTVRTMTLESFGLEKYMEEHLNAANKHFQ 171
Query: 70 TMKYKGPQTSDT--KVGLDTHTDTAILTILFQSQ-VDGLEVLTKDGRKWISYKPSSESQS 126
+KYKG +T K+G H D LTIL Q+ VDGLE+ TKDG +WI KPS S S
Sbjct: 172 LLKYKGISDDNTENKIGFYPHIDRHFLTILCQNDAVDGLEIKTKDGEEWIKVKPSQAS-S 230
Query: 127 FIVMMNDCFHAWSNGRLNSPFHRVMMTGNEARYSAALFSVPKGGYIIKAPEELVDEEHPL 186
FIVM H NG + P HRV++TG + RY AALF++PK G II APEE+VD+EHP
Sbjct: 231 FIVMAGASLHVLLNGGVFPPLHRVVITGKKDRYVAALFTIPKEGVIINAPEEMVDDEHPR 290
Query: 187 LYKPFDLEEFLKHSFNKEKGKRDDQFALHSYC 218
LYKPFD FLK F+ R D L YC
Sbjct: 291 LYKPFDFWGFLK--FSNLPNARKDISDLTDYC 320
>UniRef100_Q9C939 Putative oxidoreductase; 32373-31266 [Arabidopsis thaliana]
Length = 310
Score = 161 bits (407), Expect = 1e-38
Identities = 85/219 (38%), Positives = 129/219 (58%), Gaps = 6/219 (2%)
Query: 1 VKNTSVYEKVETLTKILWPNGNPSFRYALCSFSEHLSELDQIVRKMVLESLGVEKYLDEH 60
+ N + E + T ++WP GN F L ++E +ELDQ+V +MV +S VEKY D +
Sbjct: 97 IDNATSLEATRSFTGLMWPQGNEHFSECLYKYAEFAAELDQMVTRMVFQSYNVEKYYDPY 156
Query: 61 LNSTNYFLKTMKYKGPQTSDTKVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKP 120
+ ST Y L+ +K + P + +G THTD + TIL Q QV+GLE+ T++G + I+
Sbjct: 157 IESTTYLLRVLKNRAPNNENPTLGFVTHTDKSFTTILHQDQVNGLEMETREGER-ININL 215
Query: 121 SSESQSFIVMMNDCFHAWSNGRLNSPFHRVMMTGNEARYSAALFSVPKGGYIIKAPEELV 180
SS S F+V+ D AWSN R+ SP H+V+++G RYS +F+ G ++ PEEL+
Sbjct: 216 SSPS-LFMVVAGDALMAWSNDRVWSPRHQVLVSGETDRYSLGMFAFNNG--TLQVPEELI 272
Query: 181 DEEHPLLYKPFDLEEFLKHSFNKEKGKRDDQFALHSYCG 219
D +HPL+YKPFD L+ F + + + +YCG
Sbjct: 273 DHQHPLMYKPFDHLGLLR--FYRTDTGYKSECPIKAYCG 309
>UniRef100_Q9C938 Putative oxidoreductase; 33116-34434 [Arabidopsis thaliana]
Length = 314
Score = 157 bits (398), Expect = 1e-37
Identities = 86/212 (40%), Positives = 125/212 (58%), Gaps = 7/212 (3%)
Query: 8 EKVETLTKILWPNGNPSFRYALCSFSEHLSELDQIVRKMVLESLGVEKYLDEHLNSTNYF 67
E + T ++WP GN F + +FS ++ELD++V +M+ E+ GVEK+ + H+ S Y
Sbjct: 109 EIAQRFTHLMWPQGNDRFCNTVHTFSNAVAELDRLVVRMIFENYGVEKHYESHVGSKTYL 168
Query: 68 LKTMKYKGPQTSDTKVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSF 127
LK +KY P S + HTD L+IL Q+ V+GLEV +KDG +WIS + +S+
Sbjct: 169 LKFLKYLAPPESISMPAFPQHTDKTFLSILHQNDVNGLEVKSKDG-EWISLQ--LPPKSY 225
Query: 128 IVMMNDCFHAWSNGRLNSPFHRVMMTGNEARYSAALFSVPKGGYIIKAPEELVDEEHPLL 187
+VM D WSN R+ S HRV M G++ RY+ LFS ++ PEELVD++HPL+
Sbjct: 226 VVMAGDISMGWSNDRIRSCEHRVTMEGDKTRYTLGLFSFLTD--LVSIPEELVDDKHPLM 283
Query: 188 YKPFDLEEFLKHSFNKEKGKRDDQFALHSYCG 219
YKPFD + +F K R+ L +YCG
Sbjct: 284 YKPFDNIALI--NFYTTKEGREANSTLKAYCG 313
>UniRef100_Q9C937 Putative oxidoreductase; 36199-37309 [Arabidopsis thaliana]
Length = 289
Score = 147 bits (372), Expect = 1e-34
Identities = 91/213 (42%), Positives = 117/213 (54%), Gaps = 35/213 (16%)
Query: 8 EKVETLTKILWPNGNPSFRYALCSFSEHLSELDQIVRKMVLESLGV-EKYLDEHLNSTNY 66
+KV T LWP GN +F A+ SF+E +SELD + R+M++ES G+ E Y+ EHLNST
Sbjct: 110 DKVNAFTHKLWPQGNNNFSEAVMSFAEKVSELDFMTRRMIMESFGLDESYIKEHLNSTKC 169
Query: 67 FLKTMKYKGPQTSDTKVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQS 126
+D K DG+EV TKD + WI PS +S S
Sbjct: 170 L-----------NDVK--------------------DGIEVKTKDDKHWIKANPSQDS-S 197
Query: 127 FIVMMNDCFHAWSNGRLNSPFHRVMMTGNEARYSAALFSVPKGGYIIKAPEELVDEEHPL 186
FIV+ HA NGR+ + HRVM G R+SA LFSVPK +I APEELVD E+P
Sbjct: 198 FIVLGGAMLHALLNGRVLTGVHRVMRMGANIRFSAGLFSVPKTEDLIYAPEELVDAEYPR 257
Query: 187 LYKPFDLEEFLKHSFNKEKGKRDDQFALHSYCG 219
LYKP D E + +++ E R D AL +YCG
Sbjct: 258 LYKPVDFEAYFRYTI--EGPGRRDLSALRTYCG 288
>UniRef100_Q9C972 Putative oxidoreductase; 24302-25416 [Arabidopsis thaliana]
Length = 320
Score = 141 bits (356), Expect = 1e-32
Identities = 79/211 (37%), Positives = 119/211 (55%), Gaps = 12/211 (5%)
Query: 14 TKILWPNGNPSFRYALCSFSEHLSELDQIVRKMVLESLGVE--KYLDEHLNSTNYFLKTM 71
+K+LWP GN F ++ ++ELDQ V +M+ ES G++ K+ H ST Y L+ +
Sbjct: 114 SKLLWPQGNDPFCQTTHMYAMTMAELDQTVMRMLYESYGMDEKKHSVSHSESTRYLLRML 173
Query: 72 KYKGPQTSDTKVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMM 131
Y+ Q + G +HTD + ++IL Q+ V GL++ T G +W+ + PS F+V+
Sbjct: 174 SYRRQQNGEANTGFVSHTDKSFMSILHQNHVGGLQLKTMTG-QWVGFNPS--PTRFVVLS 230
Query: 132 NDCFHAWSNGRLNSPFHRVMMTGNEARYSAALFSVPKGGYIIKAPEELVDEEHPLLYKPF 191
AWSN R+ + +H+V+M+ +E RYS FS KG I+ PEELVD++HPL Y PF
Sbjct: 231 GMGLTAWSNDRIKACYHKVVMSADEIRYSLGFFSFHKG--TIRTPEELVDDQHPLRYNPF 288
Query: 192 D---LEEFLKHSFNKEKGKRDDQFALHSYCG 219
+ L F + N K +D L YCG
Sbjct: 289 EHDGLLRFYESYINSLKKSSED--LLQIYCG 317
>UniRef100_Q941M0 2-oxoglutarate-dependent dioxygenase [Brassica oleracea]
Length = 439
Score = 135 bits (340), Expect = 8e-31
Identities = 72/145 (49%), Positives = 95/145 (64%), Gaps = 6/145 (4%)
Query: 75 GPQTSDTKVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMMNDC 134
G + K+GL +HTD +LT+L+Q +++GLEVLTKD KWI KPS S F+VM D
Sbjct: 295 GADDEEKKLGLPSHTDKNLLTVLYQYEIEGLEVLTKD-EKWIRLKPSHNS--FVVMAGDS 351
Query: 135 FHAWSNGRLNSPFHRVMMTGNE-ARYSAALFSVPKGGYIIKAPEELVDEEHPLLYKPFDL 193
+A NGRL+ PFHRV +T + RYS ALFS P G YII+ P+ELVDE+HP L+KPF
Sbjct: 352 LYALMNGRLSRPFHRVRVTERKKTRYSIALFSTPNGDYIIEPPKELVDEKHPRLFKPFTY 411
Query: 194 EEFLKHSFNKEKGKRDDQFALHSYC 218
+ + SF + R + LH+YC
Sbjct: 412 VDLM--SFYHTEAGRRPRSTLHAYC 434
Score = 82.0 bits (201), Expect = 1e-14
Identities = 41/89 (46%), Positives = 58/89 (65%), Gaps = 2/89 (2%)
Query: 1 VKNTSVYEKVETLTKILWPN--GNPSFRYALCSFSEHLSELDQIVRKMVLESLGVEKYLD 58
+++++V EKV T++L P+ GN + + FSE L+ELD +VR+MV+ES G+E Y D
Sbjct: 96 IQDSNVLEKVNEFTQLLRPDCEGNKTMSETIQKFSEKLAELDVMVRRMVIESFGIENYFD 155
Query: 59 EHLNSTNYFLKTMKYKGPQTSDTKVGLDT 87
EHL STNY L+ MKY P D V + T
Sbjct: 156 EHLKSTNYRLRLMKYVAPPDVDANVAVST 184
>UniRef100_Q7EYC8 Putative 2-oxoglutarate-dependent dioxygenase [Oryza sativa]
Length = 312
Score = 134 bits (336), Expect = 2e-30
Identities = 73/212 (34%), Positives = 115/212 (53%), Gaps = 6/212 (2%)
Query: 9 KVETLTKILWPNGNPSFRYALCSFSEHLSELDQIVRKMVLESLGV-EKYLDEHLNSTNYF 67
+V +LWP GNP F + SF+ + +L++ V +M LE LGV E ++ HL + +Y
Sbjct: 101 RVREFAGLLWPEGNPEFCETIVSFATKMRDLERTVERMTLEGLGVGEDHIASHLAAQDYG 160
Query: 68 LKTMKYKGPQTSDTKVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSF 127
++ Y P + T + L H D ++ TI+ Q +V+GLEV DG W + P E +
Sbjct: 161 VRLSHYGPPPDASTAISLQAHRDDSMTTIIVQHEVEGLEVQAGDG-SWHAIPP--EPDTI 217
Query: 128 IVMMNDCFHAWSNGRLNSPFHRVMMTGNEARYSAALFSVPKGGYIIKAPEELVDEEHPLL 187
++ + F +NGR+ + HRV RY + S K G ++ A +ELVD EHPL
Sbjct: 218 AIVAGELFRVVTNGRVPASVHRVRTPSGRERYCVLVGSRSKDGAVLSAMDELVDGEHPLA 277
Query: 188 YKPFDLEEFLKHSFNKEKGKRDDQFALHSYCG 219
Y+P EEF++ +++E K D L ++CG
Sbjct: 278 YRPCKAEEFIQFRYSEEGRKFSD--PLKAFCG 307
>UniRef100_Q7EYC9 Putative 2-oxoglutarate-dependent dioxygenase [Oryza sativa]
Length = 311
Score = 127 bits (318), Expect = 3e-28
Identities = 68/212 (32%), Positives = 122/212 (57%), Gaps = 6/212 (2%)
Query: 9 KVETLTKILWPNGNPSFRYALCSFSEHLSELDQIVRKMVLESLGV-EKYLDEHLNSTNYF 67
+V +LWP+GNP F + SF++ ++EL++ V +M LE LGV E ++ HL++ +
Sbjct: 101 RVPEFAGLLWPDGNPEFCDTIVSFAKKMTELERAVERMTLEGLGVGEDHIASHLDAHDDA 160
Query: 68 LKTMKYKGPQTSDTKVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSF 127
++ +Y P + + + + H D ++TI+ Q +V+GLEV DG W + P ++ +F
Sbjct: 161 VRLSRYGPPPDAASAMSMGEHRDDTVITIIVQHEVEGLEVQASDG-SWHTIPPEPDTVAF 219
Query: 128 IVMMNDCFHAWSNGRLNSPFHRVMMTGNEARYSAALFSVPKGGYIIKAPEELVDEEHPLL 187
M + F +NGR+ + HRV + R A + KGG ++ A +ELVD +HPL
Sbjct: 220 --MAGELFTVVTNGRVPACVHRVRTPSHRERLVALFTTRCKGGTVVSAMDELVDGDHPLA 277
Query: 188 YKPFDLEEFLKHSFNKEKGKRDDQFALHSYCG 219
Y+P + +E+++ ++E G+ + L ++CG
Sbjct: 278 YRPCNEDEYVQFRHSEEGGRFSE--PLKAFCG 307
>UniRef100_Q9M9D9 T16N11.5 protein [Arabidopsis thaliana]
Length = 320
Score = 126 bits (316), Expect = 5e-28
Identities = 66/209 (31%), Positives = 116/209 (54%), Gaps = 7/209 (3%)
Query: 14 TKILWPNG---NPSFRYALCSFSEHLSELDQIVRKMVLESLGVEKYLDEHLNSTNYFLKT 70
++++WP+ N F + ++++ +EL+Q+V +M+ ES VEKY ++++ T Y L+
Sbjct: 114 SRLMWPDDHDDNDRFCETVHAYAKMQAELEQLVIRMLFESYNVEKYTEKYIGGTRYLLRL 173
Query: 71 MKYKGPQTSDTKVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVM 130
+KY+ + +HTD + ++IL Q+ + GL + ++ W + PS F+V+
Sbjct: 174 LKYRRLPNGEPNRKFISHTDKSFISILHQNHITGLMLKSEKEDVWYHFTPS--PTRFVVI 231
Query: 131 MNDCFHAWSNGRLNSPFHRVMMTGNEARYSAALFSVPKGGYIIKAPEELVDEEHPLLYKP 190
D AWSN R+ + +H+V M E RYS FS +G +I PEE+VD++HPL Y P
Sbjct: 232 AGDAIMAWSNDRIKACYHKVEMESVEMRYSLGFFSFQEG--MISTPEEMVDKDHPLAYNP 289
Query: 191 FDLEEFLKHSFNKEKGKRDDQFALHSYCG 219
F + L++ E + + +YCG
Sbjct: 290 FHHDGLLEYYETLEVHLKAHRTMTKAYCG 318
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.317 0.134 0.399
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 391,124,525
Number of Sequences: 2790947
Number of extensions: 16224905
Number of successful extensions: 40028
Number of sequences better than 10.0: 1099
Number of HSP's better than 10.0 without gapping: 569
Number of HSP's successfully gapped in prelim test: 530
Number of HSP's that attempted gapping in prelim test: 38241
Number of HSP's gapped (non-prelim): 1139
length of query: 219
length of database: 848,049,833
effective HSP length: 122
effective length of query: 97
effective length of database: 507,554,299
effective search space: 49232767003
effective search space used: 49232767003
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 72 (32.3 bits)
Lotus: description of TM0226.15


