

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0226.11
(231 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
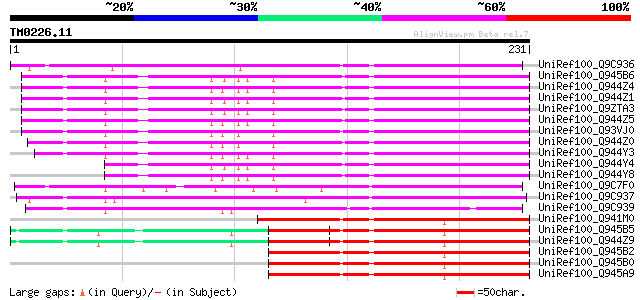
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9C936 Putative oxidoreductase; 38288-39393 [Arabidops... 231 9e-60
UniRef100_Q945B6 AOP1.2 [Arabidopsis thaliana] 180 2e-44
UniRef100_Q944Z4 2-oxoglutarate-dependent dioxygenase [Arabidops... 177 2e-43
UniRef100_Q944Z1 2-oxoglutarate-dependent dioxygenase [Arabidops... 176 6e-43
UniRef100_Q9ZTA3 Putative oxidoreductase [Arabidopsis thaliana] 176 6e-43
UniRef100_Q944Z5 2-oxoglutarate-dependent dioxygenase [Arabidops... 173 3e-42
UniRef100_Q93VJ0 AOP1 [Arabidopsis lyrata] 173 4e-42
UniRef100_Q944Z0 2-oxoglutarate-dependent dioxygenase [Arabidops... 171 2e-41
UniRef100_Q944Y3 2-oxoglutarate-dependent dioxygenase [Arabidops... 170 3e-41
UniRef100_Q944Y4 2-oxoglutarate-dependent dioxygenase [Arabidops... 167 2e-40
UniRef100_Q944Y8 2-oxoglutarate-dependent dioxygenase [Arabidops... 166 3e-40
UniRef100_Q9C7F0 Oxidoreductase, putative [Arabidopsis thaliana] 155 8e-37
UniRef100_Q9C937 Putative oxidoreductase; 36199-37309 [Arabidops... 146 5e-34
UniRef100_Q9C939 Putative oxidoreductase; 32373-31266 [Arabidops... 144 2e-33
UniRef100_Q941M0 2-oxoglutarate-dependent dioxygenase [Brassica ... 120 2e-26
UniRef100_Q945B5 AOP2 [Arabidopsis thaliana] 113 3e-24
UniRef100_Q944Z9 2-oxoglutarate-dependent dioxygenase [Arabidops... 113 3e-24
UniRef100_Q945B2 2-oxoglutarate-dependent dioxygenase [Arabidops... 113 3e-24
UniRef100_Q945B0 2-oxoglutarate-dependent dioxygenase [Arabidops... 113 3e-24
UniRef100_Q945A9 2-oxoglutarate-dependent dioxygenase [Arabidops... 113 3e-24
>UniRef100_Q9C936 Putative oxidoreductase; 38288-39393 [Arabidopsis thaliana]
Length = 317
Score = 231 bits (590), Expect = 9e-60
Identities = 131/295 (44%), Positives = 167/295 (56%), Gaps = 71/295 (24%)
Query: 1 MGSETTL---KLPVIDFTNLNNLGGTSPNWEAVKSKVHKALVEYMQL------------- 44
MGSET L +LPVIDF+N NL P W+ ++ V KAL +Y
Sbjct: 1 MGSETPLLPLRLPVIDFSN-KNLKPGEPEWDLTRADVQKALQDYGYFEASFDRIPFELRK 59
Query: 45 -------------------NVSKKPYFGYAGQFPNIPLFKSMGIDDANVYEKVESLTKIL 85
NVSKKP+ GY GQ+P +PL++SMGIDD+++ EKV++ T+ L
Sbjct: 60 SVFGALEELFDLPLQTKLRNVSKKPFHGYVGQYPMVPLYESMGIDDSDIAEKVDAFTEKL 119
Query: 86 WPNGNPSFRYASQPIT--------------------------------YFLKTMKYKGPQ 113
WP GN SF Q + Y L+ MKYKGP
Sbjct: 120 WPQGNISFSTTIQSFSKKLSELDITIRRMIMESFGLDKYIDEHLHSTNYLLRVMKYKGPD 179
Query: 114 TSDTKVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMMNDCFHA 173
T +TKVGL+ HTD I+TIL+Q+ V+GLEV TKD + WI KP+ +S F VM+ D +A
Sbjct: 180 TEETKVGLNAHTDKNIVTILYQNHVEGLEVQTKD-KNWIKVKPTQDS--FTVMIGDSLYA 236
Query: 174 WSNGRLHSPFHRVMMTGNEARYSAALFFVPKGGCIIKAPEELVDEEHPLLYKPFD 228
NGRLHSP+HRVMMTG E RYS LF +PK G I+ +P+ELVDEEHP L+KPFD
Sbjct: 237 LLNGRLHSPYHRVMMTGTETRYSLGLFSIPKAGHIVSSPDELVDEEHPRLFKPFD 291
>UniRef100_Q945B6 AOP1.2 [Arabidopsis thaliana]
Length = 321
Score = 180 bits (457), Expect = 2e-44
Identities = 122/296 (41%), Positives = 156/296 (52%), Gaps = 78/296 (26%)
Query: 6 TLKLPVIDFTNLNNLGGTSPNWEAVKSKVHKALVEY------------------------ 41
+ +LPVIDF++ N G+S W+ VK+ V KAL +Y
Sbjct: 11 SFQLPVIDFSDQNLKPGSS-KWDEVKADVLKALEDYGCFEAFFDKLSVELNRSVFEAMED 69
Query: 42 --------MQLNVSKKPYFGYAGQFPNIPLFKSMGIDDANVYEKVESLTKILWPN-GNPS 92
Q NVS KP+ GY L++S+GIDDANV EKV T+ LWP+ GN S
Sbjct: 70 LFELPIPTKQRNVSSKPFHGYLCH----NLYESLGIDDANVLEKVNDFTQQLWPDHGNKS 125
Query: 93 FR-----YASQPI--------------------------TYFL-KTMKYKGPQTSD---- 116
++ Q + TY+L + MKY P D
Sbjct: 126 ISETIHLFSEQLVELDLMVRRMIMESFGIENYIDEHLNSTYYLTRLMKYTSPPDDDDDDE 185
Query: 117 -TKVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMMNDCFHAWS 175
TK+GL +HTD I+TIL Q QVDGLEV TKD KWI KPS +S +VM+ D A
Sbjct: 186 ETKLGLRSHTDKNIITILHQYQVDGLEVKTKDD-KWIKVKPSQDS--VLVMVGDSLCALL 242
Query: 176 NGRLHSPFHRVMMTGNEARYSAALFFVPKGGCIIKAPEELVDEEHPLLYKPFDRED 231
NGRLHSP+HRV+MTG + RYS LF +PK G II +PEELVD+EHP ++KPF+ D
Sbjct: 243 NGRLHSPYHRVIMTGKKTRYSTGLFSIPKTGVIIDSPEELVDKEHPRIFKPFEYTD 298
>UniRef100_Q944Z4 2-oxoglutarate-dependent dioxygenase [Arabidopsis thaliana]
Length = 320
Score = 177 bits (449), Expect = 2e-43
Identities = 120/295 (40%), Positives = 155/295 (51%), Gaps = 77/295 (26%)
Query: 6 TLKLPVIDFTNLNNLGGTSPNWEAVKSKVHKALVEY------------------------ 41
+ +LPVIDF++ N G+S W+ V + V KAL +Y
Sbjct: 11 SFQLPVIDFSDQNLKPGSS-KWDEVTADVLKALEDYGCFEASFDKLSVELNRSVFEAMED 69
Query: 42 --------MQLNVSKKPYFGYAGQFPNIPLFKSMGIDDANVYEKVESLTKILWPN-GNPS 92
Q NVS KP+ GY L++S+GI+DANV EKV T+ LWP+ GN S
Sbjct: 70 LFELPIPTKQRNVSSKPFHGYLCH----NLYESLGINDANVLEKVNDFTQQLWPDHGNKS 125
Query: 93 F-----------------------------RYASQPI--TYFL-KTMKYKGPQTSD---- 116
+Y + + TY+L + MKY P D
Sbjct: 126 ISETIHLFSEQLVELDLMVRRMIMESFGIEKYIDEHLNSTYYLTRLMKYTSPPDDDDDEE 185
Query: 117 TKVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMMNDCFHAWSN 176
TK+GL +HTD I+TIL Q QVDGLEV TKD KWI KPS +S +VM+ D A N
Sbjct: 186 TKLGLRSHTDKNIITILHQYQVDGLEVKTKDD-KWIKVKPSQDS--VLVMVGDSLCALLN 242
Query: 177 GRLHSPFHRVMMTGNEARYSAALFFVPKGGCIIKAPEELVDEEHPLLYKPFDRED 231
GRLHSP+HRV+MTG + RYS LF +PK G II +PEELVD+EHP ++KPF+ D
Sbjct: 243 GRLHSPYHRVIMTGKKTRYSTGLFSIPKTGVIIDSPEELVDKEHPRIFKPFEYTD 297
>UniRef100_Q944Z1 2-oxoglutarate-dependent dioxygenase [Arabidopsis thaliana]
Length = 309
Score = 176 bits (445), Expect = 6e-43
Identities = 120/297 (40%), Positives = 155/297 (51%), Gaps = 79/297 (26%)
Query: 6 TLKLPVIDFTNLNNLGGTSPNWEAVKSKVHKALVEY------------------------ 41
+ +LPVIDF++ N G+S W+ V + V KAL +Y
Sbjct: 5 SFQLPVIDFSDQNLKPGSS-KWDEVTADVLKALEDYGCFEASFDKLSVELNRSVFEAMED 63
Query: 42 --------MQLNVSKKPYFGYAGQFPNIPLFKSMGIDDANVYEKVESLTKILWPN-GNPS 92
Q NVS KP+ GY L++S+GI+DANV EKV T+ LWP+ GN S
Sbjct: 64 LFELPIPTKQRNVSSKPFHGYLCH----NLYESLGINDANVLEKVNDFTQQLWPDHGNKS 119
Query: 93 FR-----YASQPI--------------------------TYFL-KTMKYKGPQTSD---- 116
++ Q + TY+L + MKY P D
Sbjct: 120 ISETIHLFSEQLVELDLMVRRMIMESFGIENYIDEHLNSTYYLTRLMKYTSPPDDDDDDD 179
Query: 117 --TKVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMMNDCFHAW 174
TK+GL +HTD I+TIL Q QVDGLEV TKD KWI KPS +S +VM+ D A
Sbjct: 180 EETKLGLRSHTDKNIITILHQYQVDGLEVKTKDD-KWIKVKPSQDS--VLVMVGDSLCAL 236
Query: 175 SNGRLHSPFHRVMMTGNEARYSAALFFVPKGGCIIKAPEELVDEEHPLLYKPFDRED 231
NGRLHSP+HRV+MTG + RYS LF +PK G II +PEELVD+EHP ++KPF+ D
Sbjct: 237 LNGRLHSPYHRVIMTGKKTRYSTGLFSIPKTGVIIDSPEELVDKEHPRIFKPFEYTD 293
>UniRef100_Q9ZTA3 Putative oxidoreductase [Arabidopsis thaliana]
Length = 322
Score = 176 bits (445), Expect = 6e-43
Identities = 120/297 (40%), Positives = 155/297 (51%), Gaps = 79/297 (26%)
Query: 6 TLKLPVIDFTNLNNLGGTSPNWEAVKSKVHKALVEY------------------------ 41
+ +LPVIDF++ N G+S W+ V + V KAL +Y
Sbjct: 11 SFQLPVIDFSDQNLKPGSS-KWDEVTADVLKALEDYGCFEASFDKLSVELNRSVFEAMED 69
Query: 42 --------MQLNVSKKPYFGYAGQFPNIPLFKSMGIDDANVYEKVESLTKILWPN-GNPS 92
Q NVS KP+ GY L++S+GI+DANV EKV T+ LWP+ GN S
Sbjct: 70 LFELPIPTKQRNVSSKPFHGYLCH----NLYESLGINDANVLEKVNDFTQQLWPDHGNKS 125
Query: 93 FR-----YASQPI--------------------------TYFL-KTMKYKGPQTSD---- 116
++ Q + TY+L + MKY P D
Sbjct: 126 ISETIHLFSEQLVELDLMVRRMIMESFGIENYIDEHLNSTYYLTRLMKYTSPPDDDDDDD 185
Query: 117 --TKVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMMNDCFHAW 174
TK+GL +HTD I+TIL Q QVDGLEV TKD KWI KPS +S +VM+ D A
Sbjct: 186 EETKLGLRSHTDKNIITILHQYQVDGLEVKTKDD-KWIKVKPSQDS--VLVMVGDSLCAL 242
Query: 175 SNGRLHSPFHRVMMTGNEARYSAALFFVPKGGCIIKAPEELVDEEHPLLYKPFDRED 231
NGRLHSP+HRV+MTG + RYS LF +PK G II +PEELVD+EHP ++KPF+ D
Sbjct: 243 LNGRLHSPYHRVIMTGKKTRYSTGLFSIPKTGVIIDSPEELVDKEHPRIFKPFEYTD 299
>UniRef100_Q944Z5 2-oxoglutarate-dependent dioxygenase [Arabidopsis thaliana]
Length = 301
Score = 173 bits (439), Expect = 3e-42
Identities = 119/295 (40%), Positives = 154/295 (51%), Gaps = 77/295 (26%)
Query: 6 TLKLPVIDFTNLNNLGGTSPNWEAVKSKVHKALVEY------------------------ 41
+ +LPVIDF++ N G+S W+ V + V KAL +Y
Sbjct: 4 SFQLPVIDFSDQNLKPGSS-KWDEVTADVLKALEDYGCFEASFDKLSVELNRSVFEAMED 62
Query: 42 --------MQLNVSKKPYFGYAGQFPNIPLFKSMGIDDANVYEKVESLTKILWPN-GNPS 92
Q NVS K + GY L++S+GI+DANV EKV T+ LWP+ GN S
Sbjct: 63 LFELPIPTKQRNVSSKLFHGYLCH----NLYESLGINDANVLEKVNDFTQQLWPDHGNKS 118
Query: 93 F-----------------------------RYASQPI--TYFL-KTMKYKGPQTSD---- 116
+Y + + TY+L + MKY P D
Sbjct: 119 ISETIHLFSEQLVELDLMVRRMIMESFGIEKYIDEHLNSTYYLTRLMKYTSPPDDDDDEE 178
Query: 117 TKVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMMNDCFHAWSN 176
TK+GL +HTD I+TIL Q QVDGLEV TKD KWI KPS +S +VM+ D A N
Sbjct: 179 TKLGLRSHTDKNIITILHQYQVDGLEVKTKDD-KWIKVKPSQDS--VLVMVGDSLCALLN 235
Query: 177 GRLHSPFHRVMMTGNEARYSAALFFVPKGGCIIKAPEELVDEEHPLLYKPFDRED 231
GRLHSP+HRV+MTG + RYS LF +PK G II +PEELVD+EHP ++KPF+ D
Sbjct: 236 GRLHSPYHRVIMTGKKTRYSTGLFSIPKTGVIIDSPEELVDKEHPRIFKPFEYTD 290
>UniRef100_Q93VJ0 AOP1 [Arabidopsis lyrata]
Length = 318
Score = 173 bits (438), Expect = 4e-42
Identities = 120/294 (40%), Positives = 149/294 (49%), Gaps = 76/294 (25%)
Query: 6 TLKLPVIDFTNLNNLGGTSPNWEAVKSKVHKALVEY------------------------ 41
+LKLPVIDF+N G+S W+ VK+ V KAL +Y
Sbjct: 10 SLKLPVIDFSNQTLKPGSS-KWDEVKADVRKALEDYGCFEASFDKVSVELKTSVFEAMEE 68
Query: 42 --------MQLNVSKKPYFGYAGQFPNIPLFKSMGIDDANVYEKVESLTKILWPN-GNPS 92
Q NVS KP+ GY L++S GIDDANV KV T+ LWP+ GN S
Sbjct: 69 LFELPIQTKQRNVSSKPFHGYICH----NLYQSXGIDDANVLVKVNDFTQQLWPDHGNKS 124
Query: 93 F-----RYASQPI---------------------------TYFLKTMKYKGPQTSD---T 117
R++ Q + Y + MKY P +D T
Sbjct: 125 ISETIHRFSDQLLELDVMVRRMIMESFGIEKYIDEHLKSTNYLFRMMKYTTPLDADVEET 184
Query: 118 KVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMMNDCFHAWSNG 177
+VGL HTD I+TIL Q Q++GLEV TK+ WI KPS +S F+VM+ D A NG
Sbjct: 185 QVGLRAHTDKNIITILHQYQINGLEVKTKN-ENWIKVKPSQDS--FVVMVGDSLCALLNG 241
Query: 178 RLHSPFHRVMMTGNEARYSAALFFVPKGGCIIKAPEELVDEEHPLLYKPFDRED 231
RL S +HRVMM GN+ RYS ALF PK II +PEELVDEEHP ++KPFD D
Sbjct: 242 RLPSSYHRVMMMGNKTRYSTALFSAPKLETIIDSPEELVDEEHPRMFKPFDYND 295
>UniRef100_Q944Z0 2-oxoglutarate-dependent dioxygenase [Arabidopsis halleri]
Length = 294
Score = 171 bits (432), Expect = 2e-41
Identities = 114/291 (39%), Positives = 150/291 (51%), Gaps = 76/291 (26%)
Query: 9 LPVIDFTNLNNLGGTSPNWEAVKSKVHKALVEY--------------------------- 41
LPVIDF++ G+S W+ VK+ V KAL +Y
Sbjct: 1 LPVIDFSDQTLKPGSS-KWDEVKADVRKALEDYGCFEASFDKVSVELKKSVFEAMEELFE 59
Query: 42 -----MQLNVSKKPYFGYAGQFPNIPLFKSMGIDDANVYEKVESLTKILWPN-GNPSF-- 93
Q NVS KP+ GY L++S+GIDDANV EKV+ T+ LWP+ GN +
Sbjct: 60 LPIQTKQRNVSSKPFHGYLCH----NLYQSLGIDDANVLEKVDDFTQQLWPDQGNKTISE 115
Query: 94 ---RYASQPI---------------------------TYFLKTMKYKGPQTSD---TKVG 120
R++ Q + Y ++ MKY P D TK+G
Sbjct: 116 TIHRFSEQLVELDVMVRRMIMESFGIEKYIDEHVNSSNYLMRFMKYTSPPDDDDEETKLG 175
Query: 121 LDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMMNDCFHAWSNGRLH 180
L +HTD I+TIL Q QV GLEV TKD KWI +PS +S ++M+ D A NGRL
Sbjct: 176 LRSHTDKNIITILHQYQVGGLEVKTKDD-KWIKVQPSQDS--VLIMLGDSLCALLNGRLP 232
Query: 181 SPFHRVMMTGNEARYSAALFFVPKGGCIIKAPEELVDEEHPLLYKPFDRED 231
SP+HRV+M + RYS LF +PK G II +PEELVDEEHP ++KPF+ +D
Sbjct: 233 SPYHRVIMMSKKTRYSTGLFSIPKTGVIIDSPEELVDEEHPRIFKPFEYKD 283
>UniRef100_Q944Y3 2-oxoglutarate-dependent dioxygenase [Arabidopsis thaliana]
Length = 301
Score = 170 bits (430), Expect = 3e-41
Identities = 117/292 (40%), Positives = 150/292 (51%), Gaps = 80/292 (27%)
Query: 12 IDFTNLNNLGGTSPNWEAVKSKVHKALVEY------------------------------ 41
IDF++ G+S W+ VK+ V KAL Y
Sbjct: 1 IDFSDQTLKPGSS-KWDEVKADVRKALENYGCFEASFDKLSVELNRSVFEAMEDLFELPI 59
Query: 42 --MQLNVSKKPYFGYAGQFPNIPLFKSMGIDDANVYEKVESLTKILWPN-GNPSF----- 93
Q NVS KP+ GY L++S+GI+DANV EKV T+ LWP+ GN S
Sbjct: 60 PTKQRNVSSKPFHGYLCH----NLYESLGINDANVLEKVNDFTQQLWPDHGNKSISETIH 115
Query: 94 RYASQPI--------------------------TYFL-KTMKYKGPQTSD-------TKV 119
R++ + + TY+L + MKY P D TK+
Sbjct: 116 RFSEKLVELDLMVRRMIMESFGIENYIDEHLNSTYYLTRLMKYTSPPDDDDDDDDEETKL 175
Query: 120 GLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMMNDCFHAWSNGRL 179
GL +HTD I+TIL Q QVDGLEV TKD KWI KPS +S +VM+ D A NGRL
Sbjct: 176 GLRSHTDKNIITILHQYQVDGLEVKTKDD-KWIKVKPSQDS--VLVMVGDSLCALLNGRL 232
Query: 180 HSPFHRVMMTGNEARYSAALFFVPKGGCIIKAPEELVDEEHPLLYKPFDRED 231
HSP+HRV+MTG + RYS LF +PK G II +PEELVD+EHP ++KPF+ D
Sbjct: 233 HSPYHRVIMTGKKTRYSTGLFSIPKTGVIIDSPEELVDKEHPRIFKPFEYTD 284
>UniRef100_Q944Y4 2-oxoglutarate-dependent dioxygenase [Arabidopsis thaliana]
Length = 272
Score = 167 bits (423), Expect = 2e-40
Identities = 104/227 (45%), Positives = 131/227 (56%), Gaps = 45/227 (19%)
Query: 43 QLNVSKKPYFGYAGQFPNIPLFKSMGIDDANVYEKVESLTKILWPN-GNPSFR-----YA 96
Q NVS KP+ GY L++S+GI+DANV EKV T+ LWP+ GN S ++
Sbjct: 52 QRNVSSKPFHGYLCH----NLYESLGINDANVLEKVNDFTQQLWPDHGNKSISETIHLFS 107
Query: 97 SQPI--------------------------TYFL-KTMKYKGPQTSD-----TKVGLDTH 124
Q + TY+L + MKY P D TK+GL +H
Sbjct: 108 EQLVELDLMVRRMIMESFGIENYIDEHLNSTYYLTRLMKYTSPPDDDDDDEETKLGLRSH 167
Query: 125 TDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMMNDCFHAWSNGRLHSPFH 184
TD I+TIL Q QVDGLEV TKD KWI KPS +S +VM+ D A NGRLHSP+H
Sbjct: 168 TDKNIITILHQYQVDGLEVKTKDD-KWIKVKPSQDS--VLVMVGDSLCALLNGRLHSPYH 224
Query: 185 RVMMTGNEARYSAALFFVPKGGCIIKAPEELVDEEHPLLYKPFDRED 231
RV+MTG + RYS LF +PK G II +PEELVD+EHP ++KPF+ D
Sbjct: 225 RVIMTGKKTRYSTGLFSIPKTGVIIDSPEELVDKEHPRIFKPFEYTD 271
>UniRef100_Q944Y8 2-oxoglutarate-dependent dioxygenase [Arabidopsis thaliana]
Length = 267
Score = 166 bits (421), Expect = 3e-40
Identities = 104/231 (45%), Positives = 132/231 (57%), Gaps = 49/231 (21%)
Query: 43 QLNVSKKPYFGYAGQFPNIPLFKSMGIDDANVYEKVESLTKILWPN-GNPSF-----RYA 96
Q NVS KP+ GY L++S+GI+DANV EKV T+ LWP+ GN S R++
Sbjct: 43 QRNVSSKPFHGYLCH----NLYESLGINDANVLEKVNDFTQQLWPDHGNKSISETIHRFS 98
Query: 97 SQPI--------------------------TYFL-KTMKYKGPQTSD---------TKVG 120
+ + TY+L + MKY P D TK+G
Sbjct: 99 EKLVELDLMVRRMIMESFGIENYIDEHLNSTYYLTRLMKYTSPPDDDDDDDDDDEETKLG 158
Query: 121 LDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMMNDCFHAWSNGRLH 180
L +HTD I+TIL Q QVDGLEV TKD KWI KPS +S +VM+ D A NGRLH
Sbjct: 159 LRSHTDKNIITILHQYQVDGLEVKTKDD-KWIKVKPSQDS--VLVMVGDSLCALLNGRLH 215
Query: 181 SPFHRVMMTGNEARYSAALFFVPKGGCIIKAPEELVDEEHPLLYKPFDRED 231
SP+HRV+MTG + RYS LF +PK G II +PEELVD+EHP ++KPF+ D
Sbjct: 216 SPYHRVIMTGKKTRYSTGLFSIPKTGVIIDSPEELVDKEHPRIFKPFEYTD 266
>UniRef100_Q9C7F0 Oxidoreductase, putative [Arabidopsis thaliana]
Length = 322
Score = 155 bits (392), Expect = 8e-37
Identities = 112/295 (37%), Positives = 142/295 (47%), Gaps = 74/295 (25%)
Query: 3 SETTLKLPVIDFTNLNNLGGTSPNWEAVKSKVHKALVEY--------------------- 41
S+ L LP+IDF+N +L +P W+ V+S+V KAL EY
Sbjct: 7 SQVPLSLPIIDFSN-PDLKPETPEWDLVRSQVRKALEEYGCFEALFDGASMELRKALFES 65
Query: 42 --------MQLNVSKKPYFGYAGQF--PNIPLFKSMG---IDDANVYEKVESLTKILWPN 88
++ +S K Y G P +P+ + MG ID+ NV V LT LWP
Sbjct: 66 SKEVFDLPLETKLSTKTDVHYEGYLTIPRVPIQEGMGFYGIDNPNV---VNDLTHKLWPQ 122
Query: 89 GN-------PSFRYASQPITYFLKTM-------------------------KYKGPQTSD 116
GN SF + ++TM KYKG +
Sbjct: 123 GNIFVGKNVQSFAEKLIELNLTVRTMTLESFGLEKYMEEHLNAANKHFQLLKYKGISDDN 182
Query: 117 T--KVGLDTHTDTAILTILFQSQ-VDGLEVLTKDGRKWISYKPSSESQSFIVMMNDCFHA 173
T K+G H D LTIL Q+ VDGLE+ TKDG +WI KPS S SFIVM H
Sbjct: 183 TENKIGFYPHIDRHFLTILCQNDAVDGLEIKTKDGEEWIKVKPSQAS-SFIVMAGASLHV 241
Query: 174 WSNGRLHSPFHRVMMTGNEARYSAALFFVPKGGCIIKAPEELVDEEHPLLYKPFD 228
NG + P HRV++TG + RY AALF +PK G II APEE+VD+EHP LYKPFD
Sbjct: 242 LLNGGVFPPLHRVVITGKKDRYVAALFTIPKEGVIINAPEEMVDDEHPRLYKPFD 296
>UniRef100_Q9C937 Putative oxidoreductase; 36199-37309 [Arabidopsis thaliana]
Length = 289
Score = 146 bits (368), Expect = 5e-34
Identities = 102/263 (38%), Positives = 132/263 (49%), Gaps = 38/263 (14%)
Query: 4 ETTL--KLPVIDFTNLNNLGGTSPNWEAVKSKVHKALVEY-----------MQLN----- 45
ETTL +LPVIDFT+ + GT W++V++ V +AL EY ++L
Sbjct: 5 ETTLALQLPVIDFTSRDLKPGTI-QWDSVRADVRRALEEYGCFEALFDKVPLELRKAVFD 63
Query: 46 ----------------VSKKPYFGYAGQFPNIPLFKSMGIDDANVYEKVESLTKILWPNG 89
VSK+ Y GY GQ P +PLF+ MG+D A +KV + T LWP G
Sbjct: 64 ASEEVFQLPLETKKRVVSKRKYRGYVGQIPTLPLFEVMGVDFAENPDKVNAFTHKLWPQG 123
Query: 90 NPSFRYASQPITYFLKTMKYKGPQTSDTKVGLDTHTDTAIL--TILFQSQVDGLEVLTKD 147
N +F A + + + + GLD L T DG+EV TKD
Sbjct: 124 NNNFSEAVMSFAEKVSELDFMTRRMIMESFGLDESYIKEHLNSTKCLNDVKDGIEVKTKD 183
Query: 148 GRKWISYKPSSESQSFIVMMNDCFHAWSNGRLHSPFHRVMMTGNEARYSAALFFVPKGGC 207
+ WI PS +S SFIV+ HA NGR+ + HRVM G R+SA LF VPK
Sbjct: 184 DKHWIKANPSQDS-SFIVLGGAMLHALLNGRVLTGVHRVMRMGANIRFSAGLFSVPKTED 242
Query: 208 IIKAPEELVDEEHPLLYKPFDRE 230
+I APEELVD E+P LYKP D E
Sbjct: 243 LIYAPEELVDAEYPRLYKPVDFE 265
>UniRef100_Q9C939 Putative oxidoreductase; 32373-31266 [Arabidopsis thaliana]
Length = 310
Score = 144 bits (363), Expect = 2e-33
Identities = 93/286 (32%), Positives = 134/286 (46%), Gaps = 70/286 (24%)
Query: 8 KLPVIDFTNLNNLGGTSPNWEAVKSKVHKALVEY-------------------------- 41
K+P +DF+ + GT WE+ + + +AL EY
Sbjct: 4 KIPTLDFSREDLKPGTK-YWESTRENIRQALEEYGCFIIDLKDKTPLDLLDRVFGSLVDL 62
Query: 42 -------MQLNVSKKPYFGYAGQFPNIPLFKSMGIDDANVYEKVESLTKILWPNGNPSF- 93
N KP GY GQ P +PL +S+GID+A E S T ++WP GN F
Sbjct: 63 FDLPTQTKMKNKYDKPLNGYVGQIPALPLHESLGIDNATSLEATRSFTGLMWPQGNEHFS 122
Query: 94 ----RYAS---------------------------QPITYFLKTMKYKGPQTSDTKVGLD 122
+YA + TY L+ +K + P + +G
Sbjct: 123 ECLYKYAEFAAELDQMVTRMVFQSYNVEKYYDPYIESTTYLLRVLKNRAPNNENPTLGFV 182
Query: 123 THTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMMNDCFHAWSNGRLHSP 182
THTD + TIL Q QV+GLE+ T++G + I+ SS S F+V+ D AWSN R+ SP
Sbjct: 183 THTDKSFTTILHQDQVNGLEMETREGER-ININLSSPS-LFMVVAGDALMAWSNDRVWSP 240
Query: 183 FHRVMMTGNEARYSAALFFVPKGGCIIKAPEELVDEEHPLLYKPFD 228
H+V+++G RYS +F G ++ PEEL+D +HPL+YKPFD
Sbjct: 241 RHQVLVSGETDRYSLGMFAFNNG--TLQVPEELIDHQHPLMYKPFD 284
>UniRef100_Q941M0 2-oxoglutarate-dependent dioxygenase [Brassica oleracea]
Length = 439
Score = 120 bits (302), Expect = 2e-26
Identities = 64/122 (52%), Positives = 81/122 (65%), Gaps = 4/122 (3%)
Query: 111 GPQTSDTKVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMMNDC 170
G + K+GL +HTD +LT+L+Q +++GLEVLTKD KWI KPS S F+VM D
Sbjct: 295 GADDEEKKLGLPSHTDKNLLTVLYQYEIEGLEVLTKD-EKWIRLKPSHNS--FVVMAGDS 351
Query: 171 FHAWSNGRLHSPFHRVMMTGNE-ARYSAALFFVPKGGCIIKAPEELVDEEHPLLYKPFDR 229
+A NGRL PFHRV +T + RYS ALF P G II+ P+ELVDE+HP L+KPF
Sbjct: 352 LYALMNGRLSRPFHRVRVTERKKTRYSIALFSTPNGDYIIEPPKELVDEKHPRLFKPFTY 411
Query: 230 ED 231
D
Sbjct: 412 VD 413
Score = 45.1 bits (105), Expect = 0.001
Identities = 39/120 (32%), Positives = 54/120 (44%), Gaps = 37/120 (30%)
Query: 1 MGSETTLKLPVIDFTNLNNLGGTSPNWEAVKSKVHKALVEY------------------- 41
MG++T +LPVI ++ L S W V+S V KAL +Y
Sbjct: 1 MGADTP-QLPVIYLSD-QTLKPGSEKWVEVRSDVRKALEDYGAFEVSYDRVSEELKKSVL 58
Query: 42 -------------MQLNVSKKPYFGYAGQFPNIPLFKSMGIDDANVYEKVESLTKILWPN 88
Q NVS KPY GY+ + L +S+GI D+NV EKV T++L P+
Sbjct: 59 DAMIELFELPVEAKQRNVSPKPYTGYS---THNGLSESLGIQDSNVLEKVNEFTQLLRPD 115
>UniRef100_Q945B5 AOP2 [Arabidopsis thaliana]
Length = 432
Score = 113 bits (283), Expect = 3e-24
Identities = 58/117 (49%), Positives = 80/117 (67%), Gaps = 4/117 (3%)
Query: 116 DTKVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMMNDCFHAWS 175
+ K+GL HTD + T+LFQ +++GLEV TKD KWI KPS + FIV+ D A
Sbjct: 293 EKKLGLPCHTDKNLFTVLFQHEIEGLEVKTKD-EKWIRVKPSPNT--FIVIAGDSLCALM 349
Query: 176 NGRLHSPFHRVMMTGNE-ARYSAALFFVPKGGCIIKAPEELVDEEHPLLYKPFDRED 231
NGR+ +P+HRV +T + RY+AA+F PK +I+AP+ELVDE+HP L++PFD D
Sbjct: 350 NGRIRAPYHRVRVTEKKRTRYTAAIFTCPKPDYVIEAPKELVDEKHPRLFRPFDYRD 406
Score = 47.4 bits (111), Expect = 3e-04
Identities = 51/208 (24%), Positives = 77/208 (36%), Gaps = 76/208 (36%)
Query: 1 MGSETTLKLPVIDFTNLNNLGGTSPNWEAVKSKVHKAL---------------------- 38
MGS +L+LP+I+ + G+S W V+S V KAL
Sbjct: 1 MGS-CSLQLPLINLADKTLEPGSS-KWAEVRSDVRKALEDFGCFEASYDKVSLELQESIM 58
Query: 39 ----------VEYMQLNVSKKPYFGYAGQFPNIPLFKSMGIDDANVYEKVESLTKILWPN 88
VE Q NV KPY GY + L +S+GI +AN+ E V T+ LWP+
Sbjct: 59 KTMEELFALPVETKQRNVCPKPYVGYLN---HNNLSESLGISNANILENVNEFTQQLWPD 115
Query: 89 GNPSFRYAS----------------------------------QPITYFLKTMKYKGPQT 114
G+ + + + Y ++ MKY P
Sbjct: 116 GDGNENISKTIQLFAEKLVEIDVMVRRMVMESFGIEKYIDDHLKSTEYRMRLMKYIAPPE 175
Query: 115 SDTKVGLDTHTDTAILTILFQSQVDGLE 142
D +D + D +L + +DG+E
Sbjct: 176 GDANTTVDDYAD-----LLAKLNIDGVE 198
>UniRef100_Q944Z9 2-oxoglutarate-dependent dioxygenase [Arabidopsis thaliana]
Length = 432
Score = 113 bits (283), Expect = 3e-24
Identities = 58/117 (49%), Positives = 80/117 (67%), Gaps = 4/117 (3%)
Query: 116 DTKVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMMNDCFHAWS 175
+ K+GL HTD + T+LFQ +++GLEV TKD KWI KPS + FIV+ D A
Sbjct: 293 EKKLGLPCHTDKNLFTVLFQHEIEGLEVKTKD-EKWIRVKPSPNT--FIVIAGDSLCALM 349
Query: 176 NGRLHSPFHRVMMTGNE-ARYSAALFFVPKGGCIIKAPEELVDEEHPLLYKPFDRED 231
NGR+ +P+HRV +T + RY+AA+F PK +I+AP+ELVDE+HP L++PFD D
Sbjct: 350 NGRIRAPYHRVRVTEKKRTRYTAAIFTCPKPDYVIEAPKELVDEKHPRLFRPFDYRD 406
Score = 47.0 bits (110), Expect = 4e-04
Identities = 50/208 (24%), Positives = 77/208 (36%), Gaps = 76/208 (36%)
Query: 1 MGSETTLKLPVIDFTNLNNLGGTSPNWEAVKSKVHKAL---------------------- 38
MGS +L+LP+I+ + G+S W V+S V KAL
Sbjct: 1 MGS-CSLQLPLINLADKTLEPGSS-KWAEVRSDVRKALQDFGCFEASYDKVSLELQESIM 58
Query: 39 ----------VEYMQLNVSKKPYFGYAGQFPNIPLFKSMGIDDANVYEKVESLTKILWPN 88
VE Q NV KPY GY + L +S+GI +AN+ E + T+ LWP+
Sbjct: 59 KTMEELFALPVETKQRNVCPKPYVGYLN---HNNLSESLGISNANILENINEFTQQLWPH 115
Query: 89 GNPSFRYAS----------------------------------QPITYFLKTMKYKGPQT 114
G+ + + + Y ++ MKY P
Sbjct: 116 GDGNENISKTIQLFAEKLVEIDVMVRRMVMESFGIEKYIDDHLKSTEYRMRLMKYIAPPE 175
Query: 115 SDTKVGLDTHTDTAILTILFQSQVDGLE 142
D +D + D +L + +DG+E
Sbjct: 176 GDANTTVDDYAD-----LLAKLNIDGVE 198
>UniRef100_Q945B2 2-oxoglutarate-dependent dioxygenase [Arabidopsis thaliana]
Length = 414
Score = 113 bits (283), Expect = 3e-24
Identities = 58/117 (49%), Positives = 80/117 (67%), Gaps = 4/117 (3%)
Query: 116 DTKVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMMNDCFHAWS 175
+ K+GL HTD + T+LFQ +++GLEV TKD KWI KPS + FIV+ D A
Sbjct: 283 EKKLGLPCHTDKNLFTVLFQHEIEGLEVKTKD-EKWIRVKPSPNT--FIVIAGDSLCALM 339
Query: 176 NGRLHSPFHRVMMTGNE-ARYSAALFFVPKGGCIIKAPEELVDEEHPLLYKPFDRED 231
NGR+ +P+HRV +T + RY+AA+F PK +I+AP+ELVDE+HP L++PFD D
Sbjct: 340 NGRIRAPYHRVRVTEKKRTRYTAAIFTCPKPDYVIEAPKELVDEKHPRLFRPFDYRD 396
Score = 44.7 bits (104), Expect = 0.002
Identities = 35/138 (25%), Positives = 54/138 (38%), Gaps = 42/138 (30%)
Query: 39 VEYMQLNVSKKPYFGYAGQFPNIPLFKSMGIDDANVYEKVESLTKILWPNGNPSFRYAS- 97
VE Q NV KPY GY + L +S+GI +AN+ E V T+ LWP+G+ + +
Sbjct: 59 VETKQRNVCPKPYVGYLN---HNNLSESLGISNANILENVNEFTQQLWPDGDGNENISKT 115
Query: 98 ---------------------------------QPITYFLKTMKYKGPQTSDTKVGLDTH 124
+ Y ++ MKY P D +D +
Sbjct: 116 IQLFAEKLVEIDVMVRRMVMESFGIEKYIDDHLKSTEYRMRLMKYIAPPEGDANTTVDDY 175
Query: 125 TDTAILTILFQSQVDGLE 142
D +L + +DG+E
Sbjct: 176 AD-----LLAKLNIDGVE 188
>UniRef100_Q945B0 2-oxoglutarate-dependent dioxygenase [Arabidopsis thaliana]
Length = 404
Score = 113 bits (283), Expect = 3e-24
Identities = 58/117 (49%), Positives = 80/117 (67%), Gaps = 4/117 (3%)
Query: 116 DTKVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMMNDCFHAWS 175
+ K+GL HTD + T+LFQ +++GLEV TKD KWI KPS + FIV+ D A
Sbjct: 273 EKKLGLPCHTDKNLFTVLFQHEIEGLEVKTKD-EKWIRVKPSPNT--FIVIAGDSLCALM 329
Query: 176 NGRLHSPFHRVMMTGNE-ARYSAALFFVPKGGCIIKAPEELVDEEHPLLYKPFDRED 231
NGR+ +P+HRV +T + RY+AA+F PK +I+AP+ELVDE+HP L++PFD D
Sbjct: 330 NGRIRAPYHRVRVTEKKRTRYTAAIFTCPKPDYVIEAPKELVDEKHPRLFRPFDYRD 386
Score = 43.9 bits (102), Expect = 0.003
Identities = 22/52 (42%), Positives = 31/52 (59%), Gaps = 3/52 (5%)
Query: 39 VEYMQLNVSKKPYFGYAGQFPNIPLFKSMGIDDANVYEKVESLTKILWPNGN 90
VE Q NV KPY GY + L +S+GI +AN+ E + T+ LWP+G+
Sbjct: 58 VETKQRNVCPKPYVGYLN---HNNLSESLGISNANILENINEFTQQLWPHGD 106
>UniRef100_Q945A9 2-oxoglutarate-dependent dioxygenase [Arabidopsis thaliana]
Length = 405
Score = 113 bits (283), Expect = 3e-24
Identities = 58/117 (49%), Positives = 80/117 (67%), Gaps = 4/117 (3%)
Query: 116 DTKVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMMNDCFHAWS 175
+ K+GL HTD + T+LFQ +++GLEV TKD KWI KPS + FIV+ D A
Sbjct: 273 EKKLGLPCHTDKNLFTVLFQHEIEGLEVKTKD-EKWIRVKPSPNT--FIVIAGDSLCALM 329
Query: 176 NGRLHSPFHRVMMTGNE-ARYSAALFFVPKGGCIIKAPEELVDEEHPLLYKPFDRED 231
NGR+ +P+HRV +T + RY+AA+F PK +I+AP+ELVDE+HP L++PFD D
Sbjct: 330 NGRIRAPYHRVRVTEKKRTRYTAAIFTCPKPDYVIEAPKELVDEKHPRLFRPFDYRD 386
Score = 43.9 bits (102), Expect = 0.003
Identities = 22/52 (42%), Positives = 31/52 (59%), Gaps = 3/52 (5%)
Query: 39 VEYMQLNVSKKPYFGYAGQFPNIPLFKSMGIDDANVYEKVESLTKILWPNGN 90
VE Q NV KPY GY + L +S+GI +AN+ E + T+ LWP+G+
Sbjct: 58 VETKQRNVCPKPYVGYLN---HNNLSESLGISNANILENINEFTQQLWPHGD 106
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.318 0.135 0.411
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 420,599,619
Number of Sequences: 2790947
Number of extensions: 17430645
Number of successful extensions: 35063
Number of sequences better than 10.0: 1078
Number of HSP's better than 10.0 without gapping: 563
Number of HSP's successfully gapped in prelim test: 515
Number of HSP's that attempted gapping in prelim test: 33545
Number of HSP's gapped (non-prelim): 1178
length of query: 231
length of database: 848,049,833
effective HSP length: 123
effective length of query: 108
effective length of database: 504,763,352
effective search space: 54514442016
effective search space used: 54514442016
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 73 (32.7 bits)
Lotus: description of TM0226.11


