

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0225.8
(423 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
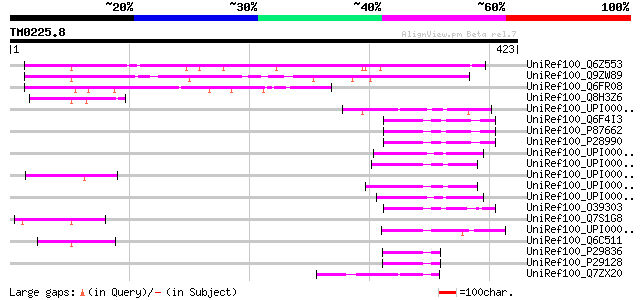
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q6Z553 Hypothetical protein OSJNBa0007M04.36 [Oryza sa... 212 1e-53
UniRef100_Q9ZW89 F5A8.9 protein [Arabidopsis thaliana] 205 2e-51
UniRef100_Q6FR08 Candida glabrata strain CBS138 chromosome I com... 68 6e-10
UniRef100_Q8H3Z6 Hypothetical protein P0597G07.117 [Oryza sativa] 57 8e-07
UniRef100_UPI0000439BA7 UPI0000439BA7 UniRef100 entry 55 5e-06
UniRef100_Q6F4I3 Transactivator protein [Equid herpesvirus 1] 54 6e-06
UniRef100_P87662 ICPO protein [Equid herpesvirus 1] 54 6e-06
UniRef100_P28990 Trans-acting transcriptional protein ICP0 [Equi... 54 6e-06
UniRef100_UPI0000438BD9 UPI0000438BD9 UniRef100 entry 53 1e-05
UniRef100_UPI000034DCFD UPI000034DCFD UniRef100 entry 53 1e-05
UniRef100_UPI00004988AA UPI00004988AA UniRef100 entry 53 2e-05
UniRef100_UPI0000363644 UPI0000363644 UniRef100 entry 52 2e-05
UniRef100_UPI00003656F9 UPI00003656F9 UniRef100 entry 52 3e-05
UniRef100_O39303 63 [Equid herpesvirus 4] 52 3e-05
UniRef100_Q7S1G8 Hypothetical protein [Neurospora crassa] 52 3e-05
UniRef100_UPI00003AFF25 UPI00003AFF25 UniRef100 entry 52 4e-05
UniRef100_Q6C511 Similar to sp|P32372 Schizosaccharomyces pombe ... 52 4e-05
UniRef100_P29836 Trans-acting transcriptional protein ICP0 [Bovi... 51 5e-05
UniRef100_P29128 Trans-acting transcriptional protein ICP0 [Bovi... 51 5e-05
UniRef100_Q7ZX20 Rnf8-prov protein [Xenopus laevis] 51 7e-05
>UniRef100_Q6Z553 Hypothetical protein OSJNBa0007M04.36 [Oryza sativa]
Length = 531
Score = 212 bits (540), Expect = 1e-53
Identities = 151/461 (32%), Positives = 215/461 (45%), Gaps = 80/461 (17%)
Query: 13 IVATVTGYHGLERFNLIKLINYAGGSYIGSMSKSITHL---KFEGKKYDIARRLSIPVVN 69
+VATV+GYHG ER L++LI G SY+G+MS+SITHL + EGKKYDIARRL VV+
Sbjct: 10 VVATVSGYHGDERHRLVRLIAETGASYVGAMSRSITHLVCWRLEGKKYDIARRLRTRVVS 69
Query: 70 HRWIEDCIREGKRLPIDSYTLQSGQEVGPLLLEVPASFGPNSWIKNKVVSDGLCDIESER 129
HRW EDC++EG+RLP Y L+SG+E GP + E+P P S K + C E
Sbjct: 70 HRWFEDCLKEGRRLPEKPYMLESGEEAGP-VPELPTF--PRSRSKRNASMEDRCLKELPD 126
Query: 130 QNTVISFGAPSYVLEDS---CLMKKYEETSL-----------------HSSRRLKKVKRN 169
S+ V+ DS C +++ ++SL H R K++K
Sbjct: 127 DFCNTSYATDVLVVADSGSDCNHQRWSDSSLLKENFVGDRDNSKIGATHVKERRKRLKHA 186
Query: 170 ISCDNEVS---------TAAGPSRKEKKFVKKVVDEVVSDPVILDLSPDDRLCRMDRDRL 220
+ NE + A R E + ++ L DD R+ L
Sbjct: 187 QNSTNEDALDAEDNISRLMARQGRYESSYTSSRSASNQKGDLLKLLHNDDASMMRKRNSL 246
Query: 221 H------------TEAAATSSLSGGVDVNDLQENSSGPDARLCNQNRAMNGSSDGIEQIT 268
E+ SL+ D + + D R + R + D I
Sbjct: 247 MKKETRTKLAGYLIESCENGSLTDSFDEPQMSDTPPTEDRRKIRKTRLRQSTLDSIYDYG 306
Query: 269 DSNQIEDPQTVEQTSIDLCSSDGKF---------------------ID-------GDQVD 300
++++ + ++ +Q + +L S F ID GD
Sbjct: 307 EASEHDPEKSEDQENFELGESSRSFQSSDSSRQEPAFCTEKTNQGSIDIAADDDKGDDEK 366
Query: 301 NVAGSPIS----NEWSCVICFEDFSSTLGVLPCEHTFCLTCIQKWAHQRISMGKISTCPL 356
P S E SCVIC+ DFSST G+LPC H FC +CIQ+WA S GK+STCPL
Sbjct: 367 ATLEEPTSCQGQAELSCVICWTDFSSTRGILPCGHRFCYSCIQEWADSLSSRGKVSTCPL 426
Query: 357 CKASFAWYKIVEHAATADKELNSQTVPCDNLASVTFISMDQ 397
CK SFAW ++ A T+D+++ SQT+PC + ++ TFI D+
Sbjct: 427 CKTSFAWISKIDEAGTSDQKIYSQTIPC-STSTDTFIFDDR 466
>UniRef100_Q9ZW89 F5A8.9 protein [Arabidopsis thaliana]
Length = 453
Score = 205 bits (521), Expect = 2e-51
Identities = 139/390 (35%), Positives = 211/390 (53%), Gaps = 42/390 (10%)
Query: 13 IVATVTGYHGLERFNLIKLINYAGGSYIGSMSKSITHL---KFEGKKYDIARRLSIPVVN 69
+VATV+GYHG +RF LIKLI+++G SY+G+MS+SITHL KFEGKKYD+A++ VVN
Sbjct: 4 VVATVSGYHGSDRFKLIKLISHSGASYVGAMSRSITHLVCWKFEGKKYDLAKKFGTVVVN 63
Query: 70 HRWIEDCIREGKRLPIDSYTLQSGQEVGPLLLEVPASFGPNSWIKNKVVSDGLCDIESER 129
HRW+E+C++EG+R+ Y SG+EVGPL++E+PA + + KV + SE
Sbjct: 64 HRWVEECVKEGRRVSETPYMFDSGEEVGPLMIELPA-VSEEAKVTKKV------NKASET 116
Query: 130 QNTVISFGAPSYVLEDSCL---MKKYEETSLHSSRRLKKVKRNISCDNEVSTAAGPSRKE 186
+ S G + S L M+K E + HS R K +I + E S A SRK
Sbjct: 117 FDKYFSNGGENRSGSTSELATWMEKNVEANRHSVRLRTKRPSSILENKENSGVAESSRKG 176
Query: 187 KKFVKKVVDEVVSDPVILDLSPDDRLCRMDRDRLHTEAAATSSLSGGVDVNDLQENSSGP 246
KK V K S ++DL D+ + D H + + + N+ Q++
Sbjct: 177 KKRVVKQR----SYRNLIDLESDE-----ESDNNHHDNSDENQ-------NETQDHREPA 220
Query: 247 DARLCN------QNRAMNGSSDGIEQITDSNQIEDPQTVEQTSI----DLCSSDGKFIDG 296
D + + A+ D D ++IE+ + +++ S + K D
Sbjct: 221 DENVRGFVFEQGETSALRHPGDLATPNWDVDEIEESENWSHSAVFKRPRSFSPEIKPQDD 280
Query: 297 DQV---DNVAGSPISNEWSCVICFEDFSSTLGVLPCEHTFCLTCIQKWAHQRISMGKIST 353
D+ + A + SC+IC+ +FSS+ G+LPC H FC +CIQKWA + +S K +T
Sbjct: 281 DESTREETEATEKAPAQVSCIICWTEFSSSRGILPCGHRFCYSCIQKWADRLVSERKKTT 340
Query: 354 CPLCKASFAWYKIVEHAATADKELNSQTVP 383
CPLCK++F +E A ++D+++ SQTVP
Sbjct: 341 CPLCKSNFITITKIEDADSSDQKIYSQTVP 370
>UniRef100_Q6FR08 Candida glabrata strain CBS138 chromosome I complete sequence
[Candida glabrata]
Length = 1099
Score = 67.8 bits (164), Expect = 6e-10
Identities = 71/291 (24%), Positives = 128/291 (43%), Gaps = 40/291 (13%)
Query: 13 IVATVTGYHGLERFNLIKLINYAGGSYIGSMSKSITHLKFE---GKKYDIARRLS----- 64
+ T T Y G +R L +LI GGS +SK THL GKKY++A +
Sbjct: 376 LAVTYTNYFGEQRTYLQQLIELLGGSATMELSKQNTHLICNLPFGKKYEVAMKWKESGSK 435
Query: 65 IPVVNHRWIEDCIREGKRLPID-----------SYTLQSGQEVGPLLLEVPASFGPNSWI 113
I + +HRW+E+C GK++P+D +Y+L GQ++ P L +S
Sbjct: 436 IVICSHRWLEECYISGKKVPVDEAYTRLERDDNTYSLTLGQQLSPKLDSENSSDEETDIE 495
Query: 114 KNKVVSDGLCDIESERQNTVISFGAPSYVLEDSCLMKKYEETSLHSSRRLKK-----VKR 168
+++V + R + + S+ LE S KKY +++ + + + + +
Sbjct: 496 ESQVFALEYLKTSQSRAHDLTGLDDISH-LESSIRKKKYGKSNANENASISRTPSASISN 554
Query: 169 NISCDNEVSTAAGPS-------RKEKKFVKKVVDEVVSDPVILDLSPDD----RLCRMDR 217
+ S D+E A + K K K+++ DP L L D LC +D
Sbjct: 555 DSSRDDEAYETAPTNFLTSTELNKSAKLDKQLLITSTVDPSNLRLKDTDYAGSALC-LDE 613
Query: 218 DRLHTEAAATSSLSGGVDVNDLQENSSGPDARLCNQNRAMNGSSDGIEQIT 268
D+ H + L+ ++ + +N + + + + N+ G+SD E+++
Sbjct: 614 DK-HESNKTDAKLN--IEGHVALKNGAKRTSDVSSTNKVSAGNSDKNEKLS 661
>UniRef100_Q8H3Z6 Hypothetical protein P0597G07.117 [Oryza sativa]
Length = 1335
Score = 57.4 bits (137), Expect = 8e-07
Identities = 34/90 (37%), Positives = 52/90 (57%), Gaps = 11/90 (12%)
Query: 17 VTGYHGLERFNLIKLINYAGGSYIGSMSK-SITHL---KFEGKKYDIARR------LSIP 66
+TGY R +++K+ + G + S THL KFEG+KY +A+R ++
Sbjct: 119 LTGYQKNWRDDIMKMASLMGAEFSKSFDALKDTHLICYKFEGEKYKVAKRENTAKRANVS 178
Query: 67 VVNHRWIEDCIREGKRLPIDSYTLQSGQEV 96
+VNH+W+EDC+ K LP D YT +SG E+
Sbjct: 179 LVNHQWLEDCLMAWKILPADDYT-KSGWEI 207
>UniRef100_UPI0000439BA7 UPI0000439BA7 UniRef100 entry
Length = 455
Score = 54.7 bits (130), Expect = 5e-06
Identities = 40/132 (30%), Positives = 65/132 (48%), Gaps = 14/132 (10%)
Query: 278 TVEQTSIDLCSSDGKF--IDGDQVDNVAGSPISNEWSCVICFEDFSSTLGVLPCEHTFCL 335
T++ T L ++D F I G Q +A +S + SC +C E F + + VL C H+ C
Sbjct: 218 TIKDTEEMLKANDVCFLKISGAQTVEMASLNVSEDLSCPVCCEIFKNPV-VLSCSHSVCK 276
Query: 336 TCIQKWAHQRISMGKISTCPLCKASFAWYKIVEHAATADKELNSQT-----VPCDNLASV 390
C+Q++ + + CP+C+ + K V ++A L+ T V D+L +V
Sbjct: 277 ECLQQYWRTKTTQ----ECPVCRRRSS--KAVNCVSSAPVILDPNTANPRLVLSDDLTTV 330
Query: 391 TFISMDQELPDN 402
F DQ +PDN
Sbjct: 331 RFTGKDQPVPDN 342
Score = 33.9 bits (76), Expect = 9.0
Identities = 17/50 (34%), Positives = 26/50 (52%), Gaps = 7/50 (14%)
Query: 310 EWSCVICFEDFSSTLGVLPCEHTFCLTCIQK-WAHQRISMGKISTCPLCK 358
E SC +C E F + + VL C H+ C C+Q+ W + CP+C+
Sbjct: 2 ELSCPVCCEIFKNPV-VLSCSHSVCKECLQQFWGTK-----NTQECPVCR 45
>UniRef100_Q6F4I3 Transactivator protein [Equid herpesvirus 1]
Length = 532
Score = 54.3 bits (129), Expect = 6e-06
Identities = 33/93 (35%), Positives = 39/93 (41%), Gaps = 16/93 (17%)
Query: 313 CVICFEDFSSTLGVLPCEHTFCLTCIQKWAHQRISMGKISTCPLCKASFAWYKIVEHAAT 372
C IC ED S+ LPC H FC CI +W Q TCPLCK + V H
Sbjct: 8 CPICLEDPSNYSMALPCLHAFCYVCITRWIRQN------PTCPLCKVP---VESVVHTIE 58
Query: 373 ADKELNSQTVPCDNLASVTFISMDQELPDNSFE 405
+D E V D D E ++SFE
Sbjct: 59 SDSEFKETKVSVD-------FDYDSEEDEDSFE 84
>UniRef100_P87662 ICPO protein [Equid herpesvirus 1]
Length = 419
Score = 54.3 bits (129), Expect = 6e-06
Identities = 33/93 (35%), Positives = 39/93 (41%), Gaps = 16/93 (17%)
Query: 313 CVICFEDFSSTLGVLPCEHTFCLTCIQKWAHQRISMGKISTCPLCKASFAWYKIVEHAAT 372
C IC ED S+ LPC H FC CI +W Q TCPLCK + V H
Sbjct: 8 CPICLEDPSNYSMALPCLHAFCYVCITRWIRQN------PTCPLCKVP---VESVVHTIE 58
Query: 373 ADKELNSQTVPCDNLASVTFISMDQELPDNSFE 405
+D E V D D E ++SFE
Sbjct: 59 SDSEFKETKVSVD-------FDYDSEEDEDSFE 84
>UniRef100_P28990 Trans-acting transcriptional protein ICP0 [Equine herpesvirus 1]
Length = 532
Score = 54.3 bits (129), Expect = 6e-06
Identities = 33/93 (35%), Positives = 39/93 (41%), Gaps = 16/93 (17%)
Query: 313 CVICFEDFSSTLGVLPCEHTFCLTCIQKWAHQRISMGKISTCPLCKASFAWYKIVEHAAT 372
C IC ED S+ LPC H FC CI +W Q TCPLCK + V H
Sbjct: 8 CPICLEDPSNYSMALPCLHAFCYVCITRWIRQN------PTCPLCKVP---VESVVHTIE 58
Query: 373 ADKELNSQTVPCDNLASVTFISMDQELPDNSFE 405
+D E V D D E ++SFE
Sbjct: 59 SDSEFKETKVSVD-------FDYDSEEDEDSFE 84
>UniRef100_UPI0000438BD9 UPI0000438BD9 UniRef100 entry
Length = 241
Score = 53.1 bits (126), Expect = 1e-05
Identities = 28/93 (30%), Positives = 47/93 (50%), Gaps = 7/93 (7%)
Query: 304 GSPISNEWSCVICFEDFSSTLGVLPCEHTFCLTCIQKWAHQRISMGKISTCPLCKASFAW 363
G + +++SC IC E + + + C HTFC C+Q + CPLC+ F
Sbjct: 28 GESLESQFSCPICLEVYHKPVSIASCAHTFCGDCLQPCLQVTSPL-----CPLCRMPFD- 81
Query: 364 YKIVEHAATADKELNSQTVPCDNLA-SVTFISM 395
K V+ +++ +K+L+S PC + VT + M
Sbjct: 82 PKKVDKSSSVEKQLSSYKAPCRGCSKKVTLVKM 114
>UniRef100_UPI000034DCFD UPI000034DCFD UniRef100 entry
Length = 473
Score = 53.1 bits (126), Expect = 1e-05
Identities = 29/89 (32%), Positives = 44/89 (48%), Gaps = 10/89 (11%)
Query: 303 AGSPISNEWSCVICFEDFSSTLGVLPCEHTFCLTCIQKWAHQRISMGKISTCPLCKASFA 362
A S + C IC + F++ + C H FC CIQ+W+H + + CPLCK FA
Sbjct: 2 AAEDASPDSKCPICLDRFNNLAYLDRCLHRFCFPCIQEWSHNK------AECPLCKQPFA 55
Query: 363 WYKIVEHAATADKELNSQTV-PCDNLASV 390
+ H+ A+ + T+ P N +SV
Sbjct: 56 ---SILHSVRAEDDFKEYTLQPAPNTSSV 81
>UniRef100_UPI00004988AA UPI00004988AA UniRef100 entry
Length = 102
Score = 52.8 bits (125), Expect = 2e-05
Identities = 31/81 (38%), Positives = 45/81 (55%), Gaps = 4/81 (4%)
Query: 14 VATVTGYHGLERFNLIKLINYAGGSYIGSM-SKSITHLKFEGKKYDIAR---RLSIPVVN 69
+ V+GY ER L ++ GG Y+ M SKS+T L +G D A R +PV++
Sbjct: 18 IICVSGYSSDERLLLRGMVELCGGIYMEDMESKSVTFLLSKGLTSDKASHALRWGVPVLS 77
Query: 70 HRWIEDCIREGKRLPIDSYTL 90
H+W+ DCI E + L I+ Y L
Sbjct: 78 HQWLFDCIAERRLLSINQYVL 98
>UniRef100_UPI0000363644 UPI0000363644 UniRef100 entry
Length = 1042
Score = 52.4 bits (124), Expect = 2e-05
Identities = 29/94 (30%), Positives = 46/94 (48%), Gaps = 10/94 (10%)
Query: 298 QVDNVAGSPISNEWSCVICFEDFSSTLGVLPCEHTFCLTCIQKWAHQRISMGKISTCPLC 357
Q + + S + C IC + F++ + C H FC CIQ+W+H + + CPLC
Sbjct: 73 QAEVMMAEDASPDSKCPICLDRFNNLAYLDRCLHRFCFPCIQEWSHNK------AECPLC 126
Query: 358 KASFAWYKIVEHAATADKELNSQTV-PCDNLASV 390
K FA + H+ A+ + T+ P N +SV
Sbjct: 127 KQPFA---SILHSVRAEDDFKEYTLQPAPNNSSV 157
>UniRef100_UPI00003656F9 UPI00003656F9 UniRef100 entry
Length = 229
Score = 52.0 bits (123), Expect = 3e-05
Identities = 29/90 (32%), Positives = 45/90 (49%), Gaps = 7/90 (7%)
Query: 307 ISNEWSCVICFEDFSSTLGVLPCEHTFCLTCIQKWAHQRISMGKISTCPLCKASFAWYKI 366
I +++SC IC E + + + C HTFC C+Q + CPLC+ F K
Sbjct: 19 IESQFSCPICLEVYHKPVSIASCAHTFCGECLQPCLQVTSPL-----CPLCRVPFD-PKK 72
Query: 367 VEHAATADKELNSQTVPCDNLA-SVTFISM 395
VE +++ +K+L S PC + VT + M
Sbjct: 73 VERSSSVEKQLASFKAPCRGCSKKVTLVKM 102
>UniRef100_O39303 63 [Equid herpesvirus 4]
Length = 536
Score = 52.0 bits (123), Expect = 3e-05
Identities = 34/93 (36%), Positives = 40/93 (42%), Gaps = 16/93 (17%)
Query: 313 CVICFEDFSSTLGVLPCEHTFCLTCIQKWAHQRISMGKISTCPLCKASFAWYKIVEHAAT 372
C IC ED S+ LPC H FC CI +W Q TCPLCK V H
Sbjct: 9 CPICLEDPSNYSMALPCLHAFCYVCITRWIRQN------PTCPLCKVP---VNSVVHTIE 59
Query: 373 ADKELNSQTVPCDNLASVTFISMDQELPDNSFE 405
+D E V SV F D + ++SFE
Sbjct: 60 SDSEFKETKV------SVEF-DYDSDEDEDSFE 85
>UniRef100_Q7S1G8 Hypothetical protein [Neurospora crassa]
Length = 1048
Score = 52.0 bits (123), Expect = 3e-05
Identities = 31/86 (36%), Positives = 46/86 (53%), Gaps = 10/86 (11%)
Query: 5 EENLV------RLKIVATVTGY-HGLERFNLIKLINYAGGSYIGSMSKSITHL---KFEG 54
E NLV R K++ TG+ ER ++I ++ GG Y G +++ +THL K EG
Sbjct: 128 EPNLVDGTIYPRRKLLVCSTGFMEPEERQHIIDMVEKGGGIYTGDLTRRVTHLVVYKPEG 187
Query: 55 KKYDIARRLSIPVVNHRWIEDCIREG 80
+KY A+ I V+ WI DC+ G
Sbjct: 188 RKYQAAQNWGIKNVSVEWINDCVERG 213
Score = 35.8 bits (81), Expect = 2.4
Identities = 19/79 (24%), Positives = 36/79 (45%), Gaps = 6/79 (7%)
Query: 16 TVTGYHGLERFNLIKLINYAGGSYIGSMSKSITHLK------FEGKKYDIARRLSIPVVN 69
+ G+ G++ ++ K I G Y + ++ L +K D+A R +P+V
Sbjct: 462 STAGFTGVDLNHVDKAIRQLGARYEERFTADVSVLVAPSVSVIRKQKLDMALRWKVPIVT 521
Query: 70 HRWIEDCIREGKRLPIDSY 88
W+ CI G + PI+ +
Sbjct: 522 AEWLWTCINSGFKCPIEDF 540
>UniRef100_UPI00003AFF25 UPI00003AFF25 UniRef100 entry
Length = 333
Score = 51.6 bits (122), Expect = 4e-05
Identities = 30/105 (28%), Positives = 46/105 (43%), Gaps = 12/105 (11%)
Query: 311 WSCVICFEDFSSTLGVLPCEHTFCLTCIQKWAHQRISMGKISTCPLCKASFAWYKIVEHA 370
W+C IC + +PC H FCL CIQ+WA + +CPLC+A+ ++
Sbjct: 237 WTCAICRDVRQDVAYAIPCHHMFCLGCIQRWARLK------DSCPLCRAAMQTIRVPVRG 290
Query: 371 ATADKE--LNSQTVPCDNLASVTFISMDQELPDNSFERREEPRVP 413
E ++ VP ++F + PDN+ E P P
Sbjct: 291 DNQYVECIVSPPAVP----VPISFRTPTDTGPDNAEEFLPPPSPP 331
>UniRef100_Q6C511 Similar to sp|P32372 Schizosaccharomyces pombe S-M checkpoint
control protein rad4 [Yarrowia lipolytica]
Length = 570
Score = 51.6 bits (122), Expect = 4e-05
Identities = 28/69 (40%), Positives = 39/69 (55%), Gaps = 4/69 (5%)
Query: 24 ERFNLIKLINYAGGSYIGSMSKSITHL----KFEGKKYDIARRLSIPVVNHRWIEDCIRE 79
ER +I I GG + G++ +T++ K +GKKYD A R SIPVV+ WIE CI+
Sbjct: 109 ERQPMIDTIEENGGIFDGNVKTLVTNILVTPKAQGKKYDFAMRKSIPVVHPIWIEHCIKR 168
Query: 80 GKRLPIDSY 88
G L +
Sbjct: 169 GAMLDTSDF 177
>UniRef100_P29836 Trans-acting transcriptional protein ICP0 [Bovine herpesvirus 1.2]
Length = 676
Score = 51.2 bits (121), Expect = 5e-05
Identities = 22/48 (45%), Positives = 26/48 (53%), Gaps = 6/48 (12%)
Query: 312 SCVICFEDFSSTLGVLPCEHTFCLTCIQKWAHQRISMGKISTCPLCKA 359
SC IC + + LPC H FCL CI++W R TCPLCKA
Sbjct: 12 SCCICLDAITGAARALPCLHAFCLACIRRWLEGR------PTCPLCKA 53
>UniRef100_P29128 Trans-acting transcriptional protein ICP0 [Bovine herpesvirus 1.1]
Length = 676
Score = 51.2 bits (121), Expect = 5e-05
Identities = 22/48 (45%), Positives = 26/48 (53%), Gaps = 6/48 (12%)
Query: 312 SCVICFEDFSSTLGVLPCEHTFCLTCIQKWAHQRISMGKISTCPLCKA 359
SC IC + + LPC H FCL CI++W R TCPLCKA
Sbjct: 12 SCCICLDAITGAARALPCLHAFCLACIRRWLEGR------PTCPLCKA 53
>UniRef100_Q7ZX20 Rnf8-prov protein [Xenopus laevis]
Length = 540
Score = 50.8 bits (120), Expect = 7e-05
Identities = 29/102 (28%), Positives = 47/102 (45%), Gaps = 14/102 (13%)
Query: 257 MNGSSDGIEQITDSNQIEDPQTVEQTSIDLCSSDGKFIDGDQVDNVAGSPISNEWSCVIC 316
+N S + EQI ++ E +T E+ + F ++V N + NE C+IC
Sbjct: 333 LNRSKNDFEQIIEAKNKELQETKEE-------KEKVFAQKEEVLNHMNDVLDNELQCIIC 385
Query: 317 FEDFSSTLGVLPCEHTFCLTCIQKWAHQRISMGKISTCPLCK 358
E F + L C H+FC CI+ W ++ CP+C+
Sbjct: 386 SEHFIEAV-TLNCAHSFCSYCIKSWKKRK------EECPICR 420
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.316 0.133 0.393
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 723,556,917
Number of Sequences: 2790947
Number of extensions: 30398596
Number of successful extensions: 78707
Number of sequences better than 10.0: 2292
Number of HSP's better than 10.0 without gapping: 228
Number of HSP's successfully gapped in prelim test: 2064
Number of HSP's that attempted gapping in prelim test: 77724
Number of HSP's gapped (non-prelim): 2661
length of query: 423
length of database: 848,049,833
effective HSP length: 130
effective length of query: 293
effective length of database: 485,226,723
effective search space: 142171429839
effective search space used: 142171429839
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 76 (33.9 bits)
Lotus: description of TM0225.8


