

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0225.5
(295 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
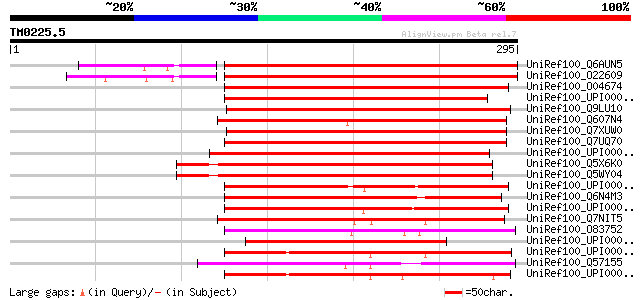
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q6AUN5 Putative DegP protease [Oryza sativa] 316 4e-85
UniRef100_O22609 Protease Do-like 1, chloroplast precursor [Arab... 307 2e-82
UniRef100_O04674 HtrA-like protein [Haematococcus pluvialis] 217 3e-55
UniRef100_UPI00002FFA81 UPI00002FFA81 UniRef100 entry 201 1e-50
UniRef100_Q9LU10 Protease Do-like 8, chloroplast precursor [Arab... 162 8e-39
UniRef100_Q607N4 Serine protease, putative [Methylococcus capsul... 159 6e-38
UniRef100_Q7XUW0 OSJNBa0072F16.5 protein [Oryza sativa] 159 6e-38
UniRef100_Q7UQ70 Protease Do-like [Rhodopirellula baltica] 155 2e-36
UniRef100_UPI000033647B UPI000033647B UniRef100 entry 153 6e-36
UniRef100_Q5X6K0 Hypothetical protein [Legionella pneumophila st... 142 8e-33
UniRef100_Q5WY04 Hypothetical protein [Legionella pneumophila st... 142 8e-33
UniRef100_UPI00002952CE UPI00002952CE UniRef100 entry 139 1e-31
UniRef100_Q6N4M3 Putative DegP protease [Rhodopseudomonas palust... 139 1e-31
UniRef100_UPI00002748C3 UPI00002748C3 UniRef100 entry 134 3e-30
UniRef100_Q7NIT5 Gll2097 protein [Gloeobacter violaceus] 131 2e-29
UniRef100_O83752 Periplasmic serine protease DO [Treponema palli... 129 1e-28
UniRef100_UPI000034BB2A UPI000034BB2A UniRef100 entry 124 4e-27
UniRef100_UPI00002ADF0E UPI00002ADF0E UniRef100 entry 122 1e-26
UniRef100_Q57155 Serine protease MucD [Pseudomonas aeruginosa] 120 3e-26
UniRef100_UPI00002D1A2D UPI00002D1A2D UniRef100 entry 120 4e-26
>UniRef100_Q6AUN5 Putative DegP protease [Oryza sativa]
Length = 437
Score = 316 bits (810), Expect = 4e-85
Identities = 157/170 (92%), Positives = 166/170 (97%)
Query: 126 LVSSDAAINPGNSGGPLLDSSGNLIGINTAIYSPSGASSGVGFSIPLDTVNGIVDQLVKF 185
++ +DAAINPGNSGGPLLDSSGNLIG+NTAIYSPSGASSGVGFSIP+DTV GIVDQL+KF
Sbjct: 268 VIQTDAAINPGNSGGPLLDSSGNLIGVNTAIYSPSGASSGVGFSIPVDTVGGIVDQLIKF 327
Query: 186 GKVTRPILGIKFAPDQSVEQLGVSGVLVLDAPANGPAGKAGLQSTKRDSYGRLILGDIIT 245
GKVTRPILGIKFAPDQSVEQLG+SGVLVLDAP NGPAGKAGLQSTKRDSYGRLILGDIIT
Sbjct: 328 GKVTRPILGIKFAPDQSVEQLGLSGVLVLDAPPNGPAGKAGLQSTKRDSYGRLILGDIIT 387
Query: 246 SVNGTKVTSGSDLYRILDQCKVGDKVTVEVLRGDHKEKIPVILEPKPDES 295
SVNGTKVT+GSDLYRILDQCKVG+KVTVEVLRGD KEKIPVILEPKPDES
Sbjct: 388 SVNGTKVTNGSDLYRILDQCKVGEKVTVEVLRGDQKEKIPVILEPKPDES 437
Score = 65.1 bits (157), Expect = 2e-09
Identities = 46/103 (44%), Positives = 55/103 (52%), Gaps = 25/103 (24%)
Query: 41 SILILCTSIALSFTNADSAYAFVVTPPRKLQSDELAT----------VVYITNLAVKMRL 90
S+++ S AL +A SA AFVV PRKLQ+DELAT VVYITNLAV+
Sbjct: 83 SLIVALASAALILGDAGSASAFVVATPRKLQADELATVRLFQENTPSVVYITNLAVRQDA 142
Query: 91 -------------RWMCWRFLKEGNIVTNYHVIPGASDLSLDL 120
W K G+IVTN+HVI GASDL + L
Sbjct: 143 FTLDVLEVPQGSGSGFVWD--KSGHIVTNFHVIRGASDLRVTL 183
>UniRef100_O22609 Protease Do-like 1, chloroplast precursor [Arabidopsis thaliana]
Length = 439
Score = 307 bits (787), Expect = 2e-82
Identities = 153/170 (90%), Positives = 163/170 (95%)
Query: 126 LVSSDAAINPGNSGGPLLDSSGNLIGINTAIYSPSGASSGVGFSIPLDTVNGIVDQLVKF 185
++ +DAAINPGNSGGPLLDSSG LIGINTAIYSPSGASSGVGFSIP+DTV GIVDQLV+F
Sbjct: 270 VIQTDAAINPGNSGGPLLDSSGTLIGINTAIYSPSGASSGVGFSIPVDTVGGIVDQLVRF 329
Query: 186 GKVTRPILGIKFAPDQSVEQLGVSGVLVLDAPANGPAGKAGLQSTKRDSYGRLILGDIIT 245
GKVTRPILGIKFAPDQSVEQLGVSGVLVLDAP +GPAGKAGLQSTKRD YGRL+LGDIIT
Sbjct: 330 GKVTRPILGIKFAPDQSVEQLGVSGVLVLDAPPSGPAGKAGLQSTKRDGYGRLVLGDIIT 389
Query: 246 SVNGTKVTSGSDLYRILDQCKVGDKVTVEVLRGDHKEKIPVILEPKPDES 295
SVNGTKV++GSDLYRILDQCKVGD+VTVEVLRGDHKEKI V LEPKPDES
Sbjct: 390 SVNGTKVSNGSDLYRILDQCKVGDEVTVEVLRGDHKEKISVTLEPKPDES 439
Score = 73.2 bits (178), Expect = 8e-12
Identities = 51/116 (43%), Positives = 64/116 (54%), Gaps = 31/116 (26%)
Query: 34 TPKTCFNSILILCTSIALSFT------NADSAYAFVVTPPRKLQSDELATV--------- 78
TP + +LCTS+ALSF+ +SA AFVV+ P+KLQ+DELATV
Sbjct: 72 TPFSAVKPFFLLCTSVALSFSLFAASPAVESASAFVVSTPKKLQTDELATVRLFQENTPS 131
Query: 79 -VYITNLAVKMRLRWM-------------CWRFLKEGNIVTNYHVIPGASDLSLDL 120
VYITNLAV+ + W K+G+IVTNYHVI GASDL + L
Sbjct: 132 VVYITNLAVRQDAFTLDVLEVPQGSGSGFVWD--KQGHIVTNYHVIRGASDLRVTL 185
>UniRef100_O04674 HtrA-like protein [Haematococcus pluvialis]
Length = 398
Score = 217 bits (552), Expect = 3e-55
Identities = 105/165 (63%), Positives = 131/165 (78%)
Query: 126 LVSSDAAINPGNSGGPLLDSSGNLIGINTAIYSPSGASSGVGFSIPLDTVNGIVDQLVKF 185
++ +DAAINPGNSGGPLLDSSG +IGINTAIYSPSG +SGVGF+IP DTV V Q+++F
Sbjct: 231 VIQTDAAINPGNSGGPLLDSSGCVIGINTAIYSPSGTNSGVGFAIPADTVRSSVTQILEF 290
Query: 186 GKVTRPILGIKFAPDQSVEQLGVSGVLVLDAPANGPAGKAGLQSTKRDSYGRLILGDIIT 245
GKV RP+LGI FAPDQ+VE LGV G++VL+A GPA KAG+ T RD YGRL+LGDII
Sbjct: 291 GKVVRPMLGIAFAPDQAVEALGVKGIMVLNAREGGPAWKAGIVGTSRDEYGRLVLGDIIR 350
Query: 246 SVNGTKVTSGSDLYRILDQCKVGDKVTVEVLRGDHKEKIPVILEP 290
+VNGT + S +DLYR+LD+ +VG+ + +EVLRG E + V L P
Sbjct: 351 TVNGTVIRSSTDLYRVLDKAQVGETLDIEVLRGSSTEHVNVTLAP 395
>UniRef100_UPI00002FFA81 UPI00002FFA81 UniRef100 entry
Length = 255
Score = 201 bits (512), Expect = 1e-50
Identities = 97/153 (63%), Positives = 124/153 (80%)
Query: 126 LVSSDAAINPGNSGGPLLDSSGNLIGINTAIYSPSGASSGVGFSIPLDTVNGIVDQLVKF 185
++ +DAAINPGNSGGPLL+S G LIG+NTAIYS SG SSGVGF++P D V+GIVDQL++F
Sbjct: 65 IIQTDAAINPGNSGGPLLNSGGQLIGVNTAIYSASGTSSGVGFALPSDMVSGIVDQLIRF 124
Query: 186 GKVTRPILGIKFAPDQSVEQLGVSGVLVLDAPANGPAGKAGLQSTKRDSYGRLILGDIIT 245
G+VTRPILG+ FAPD +++QLG+ GVLVLDA A GPA +AG++ST RD GRLILGDII
Sbjct: 125 GRVTRPILGVSFAPDGALDQLGLGGVLVLDARAGGPADRAGVRSTTRDESGRLILGDIIV 184
Query: 246 SVNGTKVTSGSDLYRILDQCKVGDKVTVEVLRG 278
+ G + SDL+R LD+ VG+ V V+++RG
Sbjct: 185 ELAGYPINDSSDLFRALDKLSVGEIVDVKLVRG 217
>UniRef100_Q9LU10 Protease Do-like 8, chloroplast precursor [Arabidopsis thaliana]
Length = 448
Score = 162 bits (411), Expect = 8e-39
Identities = 86/166 (51%), Positives = 114/166 (67%), Gaps = 1/166 (0%)
Query: 127 VSSDAAINPGNSGGPLLDSSGNLIGINTAIYSPSGASSGVGFSIPLDTVNGIVDQLVKFG 186
+ +DAAINPGNSGGPLLDS GNLIGINTAI++ +G S+GVGF+IP TV IV QL++F
Sbjct: 281 IQTDAAINPGNSGGPLLDSKGNLIGINTAIFTQTGTSAGVGFAIPSSTVLKIVPQLIQFS 340
Query: 187 KVTRPILGIKFAPDQSVEQLGV-SGVLVLDAPANGPAGKAGLQSTKRDSYGRLILGDIIT 245
KV R + I+ APD QL V +G LVL P A KAGL T R G ++LGDII
Sbjct: 341 KVLRAGINIELAPDPVANQLNVRNGALVLQVPGKSLAEKAGLHPTSRGFAGNIVLGDIIV 400
Query: 246 SVNGTKVTSGSDLYRILDQCKVGDKVTVEVLRGDHKEKIPVILEPK 291
+V+ V + ++L +ILD+ VGDKVT+++ RG+ ++ + LE K
Sbjct: 401 AVDDKPVKNKAELMKILDEYSVGDKVTLKIKRGNEDLELKISLEEK 446
>UniRef100_Q607N4 Serine protease, putative [Methylococcus capsulatus]
Length = 374
Score = 159 bits (403), Expect = 6e-38
Identities = 90/171 (52%), Positives = 115/171 (66%), Gaps = 3/171 (1%)
Query: 122 TLLQLVSSDAAINPGNSGGPLLDSSGNLIGINTAIYSPSGASSGVGFSIPLDTVNGIVDQ 181
T+ L+ +DAAINPGNSGGPLLDS+G L+GINTAIYSPSGA SGVGF++P+DTVN +V Q
Sbjct: 201 TIEHLIQTDAAINPGNSGGPLLDSAGRLVGINTAIYSPSGAFSGVGFAVPVDTVNRVVPQ 260
Query: 182 LVKFGKVTRPILGI---KFAPDQSVEQLGVSGVLVLDAPANGPAGKAGLQSTKRDSYGRL 238
L+ G+ RP LGI + ++V++LGV+GVLVL A AGL+ GRL
Sbjct: 261 LIGRGQYIRPALGIAVDEGLNQRAVQRLGVTGVLVLKVNPGSAAEAAGLKGATLLPDGRL 320
Query: 239 ILGDIITSVNGTKVTSGSDLYRILDQCKVGDKVTVEVLRGDHKEKIPVILE 289
I GDII +V G V S S L +LD ++G KV + V RGD + I V L+
Sbjct: 321 IPGDIIVAVEGRPVDSVSKLSALLDDYQIGQKVRLSVRRGDTEMDIAVQLQ 371
>UniRef100_Q7XUW0 OSJNBa0072F16.5 protein [Oryza sativa]
Length = 420
Score = 159 bits (403), Expect = 6e-38
Identities = 83/164 (50%), Positives = 111/164 (67%), Gaps = 1/164 (0%)
Query: 127 VSSDAAINPGNSGGPLLDSSGNLIGINTAIYSPSGASSGVGFSIPLDTVNGIVDQLVKFG 186
+ +DAAINPGNSGGPLLDS G++IGINTAI++ +G S+GVGF+IP TV I QL++FG
Sbjct: 253 IQTDAAINPGNSGGPLLDSKGHMIGINTAIFTQTGTSAGVGFAIPSSTVLKIAPQLIQFG 312
Query: 187 KVTRPILGIKFAPDQSVEQLGV-SGVLVLDAPANGPAGKAGLQSTKRDSYGRLILGDIIT 245
KV R L ++FAPD QL V +G L+L P A KAGL T R G ++LGD+I
Sbjct: 313 KVRRAGLNVEFAPDPIAYQLNVRTGSLILQVPGGSAAAKAGLVPTSRGFAGNIVLGDVIV 372
Query: 246 SVNGTKVTSGSDLYRILDQCKVGDKVTVEVLRGDHKEKIPVILE 289
+V+G + SDL R+LD VGDKV++ + RG ++ + LE
Sbjct: 373 AVDGKPIKGKSDLSRVLDDYGVGDKVSLTIQRGAETLEVTLPLE 416
>UniRef100_Q7UQ70 Protease Do-like [Rhodopirellula baltica]
Length = 399
Score = 155 bits (391), Expect = 2e-36
Identities = 80/165 (48%), Positives = 114/165 (68%), Gaps = 1/165 (0%)
Query: 126 LVSSDAAINPGNSGGPLLDSSGNLIGINTAIYSPSGASSGVGFSIPLDTVNGIVDQLVKF 185
++ +DAAINPGNSGGPLLD SG LIG+NTAIYSPSGA +G+GF+IP+DTV +V +L++
Sbjct: 232 VIQTDAAINPGNSGGPLLDRSGQLIGVNTAIYSPSGAYAGIGFAIPVDTVRWVVPELIEH 291
Query: 186 GKVTRPILGIKFAPDQSVEQLGV-SGVLVLDAPANGPAGKAGLQSTKRDSYGRLILGDII 244
G++ RP + I A D ++ + GVL+LD P G A +AGL+ T+R +G ++LGDII
Sbjct: 292 GRIIRPGIAITVASDSMSKRFKLPPGVLILDMPERGNAERAGLRPTRRTRFGDIVLGDII 351
Query: 245 TSVNGTKVTSGSDLYRILDQCKVGDKVTVEVLRGDHKEKIPVILE 289
+V+ V S +DL I + + GD V + V+R + +PV LE
Sbjct: 352 VAVDEMPVASTADLTLIFENYESGDVVDLTVIRQGTELVLPVELE 396
>UniRef100_UPI000033647B UPI000033647B UniRef100 entry
Length = 190
Score = 153 bits (386), Expect = 6e-36
Identities = 72/163 (44%), Positives = 109/163 (66%)
Query: 117 SLDLTTLLQLVSSDAAINPGNSGGPLLDSSGNLIGINTAIYSPSGASSGVGFSIPLDTVN 176
SL+ T+ ++ +DAAINPGNSGGPLL+SSG +IG+NTAI SPSGAS+G+GF+IP++T+
Sbjct: 12 SLNQRTIRDVIQTDAAINPGNSGGPLLNSSGQIIGVNTAIRSPSGASAGIGFAIPINTLK 71
Query: 177 GIVDQLVKFGKVTRPILGIKFAPDQSVEQLGVSGVLVLDAPANGPAGKAGLQSTKRDSYG 236
IV QL++ GKV RP++G++ D ++LG GV + PA +AG+ + D G
Sbjct: 72 NIVPQLIEHGKVVRPVMGVELLSDYWTKRLGARGVAIFSVVEGLPAERAGMVGVRYDKQG 131
Query: 237 RLILGDIITSVNGTKVTSGSDLYRILDQCKVGDKVTVEVLRGD 279
++ LGD++ +NG V+ L L++ + GDK+ V R +
Sbjct: 132 KIHLGDVLIDINGALVSDEDTLLEQLEKFEPGDKIQVTTTRNE 174
>UniRef100_Q5X6K0 Hypothetical protein [Legionella pneumophila str. Paris]
Length = 363
Score = 142 bits (359), Expect = 8e-33
Identities = 73/185 (39%), Positives = 112/185 (60%), Gaps = 6/185 (3%)
Query: 98 LKEGNIVTNYHVIPGASDLSLDLTTLLQLVSSDAAINPGNSGGPLLDSSGNLIGINTAIY 157
L +G I +PG + T+ ++ +D INPGNSGGPLL+S+G LIG+NT IY
Sbjct: 171 LSKGVISALGRKVPGIGGV-----TIYDMIQTDTPINPGNSGGPLLNSAGQLIGMNTMIY 225
Query: 158 SPSGASSGVGFSIPLDTVNGIVDQLVKFGKVTRPILGIKFAPDQSVEQLGV-SGVLVLDA 216
S SG+S+G+GF++P + + I Q++ G+V +GI+ E+LGV G+L+ D
Sbjct: 226 SRSGSSAGIGFAVPAEDIQKIASQIINHGRVVLSGIGIQRVEPHLAERLGVKKGILIADV 285
Query: 217 PANGPAGKAGLQSTKRDSYGRLILGDIITSVNGTKVTSGSDLYRILDQCKVGDKVTVEVL 276
PA K L+ T R+ +GR++LGD+I VN V + LY +L + KVG+++TV ++
Sbjct: 286 VPGTPADKLKLRGTHRNQWGRIVLGDVIVGVNAHPVPNYDALYNLLTEIKVGEQITVSII 345
Query: 277 RGDHK 281
R K
Sbjct: 346 RNGKK 350
>UniRef100_Q5WY04 Hypothetical protein [Legionella pneumophila str. Lens]
Length = 363
Score = 142 bits (359), Expect = 8e-33
Identities = 73/185 (39%), Positives = 112/185 (60%), Gaps = 6/185 (3%)
Query: 98 LKEGNIVTNYHVIPGASDLSLDLTTLLQLVSSDAAINPGNSGGPLLDSSGNLIGINTAIY 157
L +G I +PG + T+ ++ +D INPGNSGGPLL+S+G LIG+NT IY
Sbjct: 171 LSKGVISALGRKVPGIGGV-----TIYDMIQTDTPINPGNSGGPLLNSAGQLIGMNTMIY 225
Query: 158 SPSGASSGVGFSIPLDTVNGIVDQLVKFGKVTRPILGIKFAPDQSVEQLGV-SGVLVLDA 216
S SG+S+G+GF++P + + I Q++ G+V +GI+ E+LGV G+L+ D
Sbjct: 226 SRSGSSAGIGFAVPAEDIQKIASQIINHGRVVLSGIGIQRVEPHLAERLGVKKGILIADV 285
Query: 217 PANGPAGKAGLQSTKRDSYGRLILGDIITSVNGTKVTSGSDLYRILDQCKVGDKVTVEVL 276
PA K L+ T R+ +GR++LGD+I VN V + LY +L + KVG+++TV ++
Sbjct: 286 VPGTPADKLKLRGTHRNQWGRIVLGDVIVGVNAHPVPNYDALYNLLTEIKVGEQITVSII 345
Query: 277 RGDHK 281
R K
Sbjct: 346 RNGKK 350
>UniRef100_UPI00002952CE UPI00002952CE UniRef100 entry
Length = 248
Score = 139 bits (349), Expect = 1e-31
Identities = 82/170 (48%), Positives = 104/170 (60%), Gaps = 8/170 (4%)
Query: 126 LVSSDAAINPGNSGGPLLDSSGNLIGINTAIYSPSGASSGVGFSIPLDTVNGIVDQLVKF 185
LV +DAAINPGNSGGPLLDS+G LIGINTAIYSPSGAS+G+GF++P+DTV +V QL+K
Sbjct: 80 LVQTDAAINPGNSGGPLLDSAGRLIGINTAIYSPSGASAGIGFAVPVDTVMRVVPQLIKT 139
Query: 186 GKVTRPILGIKFAPDQSVEQ-----LGVSGVLVLDAPANGPAGKAGLQSTKRDSYGRLIL 240
G+ RP LGI+ D+ + Q G GV VL A KAGL + G ++
Sbjct: 140 GRYIRPALGIEV--DEQLNQRLLALTGSKGVFVLRVTPGSAAHKAGLAGVEVTPQG-IVP 196
Query: 241 GDIITSVNGTKVTSGSDLYRILDQCKVGDKVTVEVLRGDHKEKIPVILEP 290
GD I V+G + L LD KVGD V + V R ++ V L+P
Sbjct: 197 GDRIIGVDGKATDDVAKLLARLDDRKVGDVVVLSVERAGKSREVRVELQP 246
>UniRef100_Q6N4M3 Putative DegP protease [Rhodopseudomonas palustris]
Length = 399
Score = 139 bits (349), Expect = 1e-31
Identities = 75/161 (46%), Positives = 101/161 (62%), Gaps = 4/161 (2%)
Query: 126 LVSSDAAINPGNSGGPLLDSSGNLIGINTAIYSPSGASSGVGFSIPLDTVNGIVDQLVKF 185
++ +DAAINPGNSGGPLLDS+G LIGINTAI S SGAS+G+GF+IP+D VN +V L+
Sbjct: 232 VIQTDAAINPGNSGGPLLDSAGRLIGINTAIISGSGASAGIGFAIPVDAVNRVVTALITN 291
Query: 186 GKVTRPILGIKFAPDQSVEQLGVSGVLVLDAPANGPAGKAGLQSTKRDSYGRLILGDIIT 245
G V P +GI A + QLG+ GV++L + PA +AGL+ D Y R D+IT
Sbjct: 292 GSVPVPGIGIVAARETETAQLGIDGVVILRTLPDSPAAQAGLEGATDDGYVR----DVIT 347
Query: 246 SVNGTKVTSGSDLYRILDQCKVGDKVTVEVLRGDHKEKIPV 286
NG+ + S SDL L++ +G V + V R + V
Sbjct: 348 GANGSDIHSMSDLAAALEEAGIGRDVKLTVERDGRARTVTV 388
>UniRef100_UPI00002748C3 UPI00002748C3 UniRef100 entry
Length = 383
Score = 134 bits (337), Expect = 3e-30
Identities = 78/168 (46%), Positives = 103/168 (60%), Gaps = 4/168 (2%)
Query: 126 LVSSDAAINPGNSGGPLLDSSGNLIGINTAIYSPSGASSGVGFSIPLDTVNGIVDQLVKF 185
L+ +DAAINPGNSGGPLLDS+G LIGINTAIYSPSGAS+G+GF++P+DTV +V QL+K
Sbjct: 215 LIQTDAAINPGNSGGPLLDSAGRLIGINTAIYSPSGASAGIGFAVPVDTVMRVVPQLIKT 274
Query: 186 GKVTRPILGIKFAPDQSVE---QLGVSGVLVLDAPANGPAGKAGLQSTKRDSYGRLILGD 242
GK RP LGI+ + G GV VL A +AGL + + G ++ GD
Sbjct: 275 GKYIRPALGIEVDEQLNARLQALTGSKGVFVLRVTPGSAAHRAGLVGVEVTA-GGIVPGD 333
Query: 243 IITSVNGTKVTSGSDLYRILDQCKVGDKVTVEVLRGDHKEKIPVILEP 290
+ S++G V + L LD VGD V + V R ++ V L+P
Sbjct: 334 RVISIDGIAVDDVTTLQARLDDKNVGDVVVLLVERAGKTREMLVELQP 381
>UniRef100_Q7NIT5 Gll2097 protein [Gloeobacter violaceus]
Length = 400
Score = 131 bits (329), Expect = 2e-29
Identities = 74/176 (42%), Positives = 111/176 (63%), Gaps = 9/176 (5%)
Query: 122 TLLQLVSSDAAINPGNSGGPLLDSSGNLIGINTAIYSPSGASSGVGFSIPLDTVNGIVDQ 181
TL L+ +DAAINPGNSGGPLLDS G LIG+NTAI+S SG+S+G+GF++P+DTV ++ +
Sbjct: 215 TLRNLIQTDAAINPGNSGGPLLDSQGRLIGVNTAIFSTSGSSAGIGFAVPVDTVRQVLPE 274
Query: 182 LVKFGKVTRPILGIKFAP--DQSVEQLGVS---GVLVLDAPANGPAGKAGLQSTKRDSY- 235
L+ G V R LG++ P VE L +S G LV G A +AGL++ + ++
Sbjct: 275 LISRGTVRRASLGVQVLPLSPMVVETLKLSVKEGALVAAVVPGGAAARAGLRAGRLETID 334
Query: 236 GRLIL---GDIITSVNGTKVTSGSDLYRILDQCKVGDKVTVEVLRGDHKEKIPVIL 288
G L L D+I +++ + DL + + K GDKVT+ ++R + + ++PV L
Sbjct: 335 GNLQLPVGADVIVAIDRVAIKDAQDLINQIQKHKPGDKVTLTIVRNNAEVQVPVTL 390
>UniRef100_O83752 Periplasmic serine protease DO [Treponema pallidum]
Length = 398
Score = 129 bits (323), Expect = 1e-28
Identities = 77/182 (42%), Positives = 109/182 (59%), Gaps = 13/182 (7%)
Query: 126 LVSSDAAINPGNSGGPLLDSSGNLIGINTAIYSPSGASSGVGFSIPLDTVNGIVDQLVKF 185
++ +DAAINPGNSGGPLLD+ G +IGINT IYS SG+SSGVGF++P+DT IV +L+++
Sbjct: 217 MIQTDAAINPGNSGGPLLDTQGRMIGINTVIYSTSGSSSGVGFAVPVDTAKRIVSELIRY 276
Query: 186 GKVTRPILGIKF----APDQSVEQLGV-SGVLVLDAPANGPAGKAGLQ---STKRDSYGR 237
G+V R + + A QL V G+LV PA +AGL+ + R GR
Sbjct: 277 GRVRRGKIDAELVQVNASIAHYAQLTVGKGLLVSQVKRGSPAAQAGLRGGTTAVRYGLGR 336
Query: 238 -----LILGDIITSVNGTKVTSGSDLYRILDQCKVGDKVTVEVLRGDHKEKIPVILEPKP 292
+ GD+IT+++ V + SD Y +L+ K D+V V VLRG + + V L +
Sbjct: 337 RAAVIYLGGDVITAIDNQPVANLSDYYSVLEDKKPDDEVRVTVLRGRRQHVVAVRLTERS 396
Query: 293 DE 294
DE
Sbjct: 397 DE 398
>UniRef100_UPI000034BB2A UPI000034BB2A UniRef100 entry
Length = 182
Score = 124 bits (310), Expect = 4e-27
Identities = 63/118 (53%), Positives = 83/118 (69%), Gaps = 1/118 (0%)
Query: 138 SGGPLLDSSGNLIGINTAIYSPSGASSGVGFSIPLDTVNGIVDQLVKFGKVTRPILGIKF 197
+GG LLDSSG LIG+NTAIYSPSGAS+G+GF+IP+DT+ ++K+G+V RP +GI +
Sbjct: 5 TGGALLDSSGRLIGMNTAIYSPSGASAGIGFAIPVDTLKQQAATIIKYGRVLRPAIGISY 64
Query: 198 APDQSVEQLGVS-GVLVLDAPANGPAGKAGLQSTKRDSYGRLILGDIITSVNGTKVTS 254
A LGV GVLV+ P A KAGL+ + R GR+ LGD+IT+VNG +V S
Sbjct: 65 AQGSQARALGVQRGVLVITVPPGSNAEKAGLRGSFRTEDGRISLGDVITAVNGVQVRS 122
>UniRef100_UPI00002ADF0E UPI00002ADF0E UniRef100 entry
Length = 241
Score = 122 bits (305), Expect = 1e-26
Identities = 73/174 (41%), Positives = 107/174 (60%), Gaps = 8/174 (4%)
Query: 126 LVSSDAAINPGNSGGPLLDSSGNLIGINTAIYSPSGASSGVGFSIPLDTVNGIVDQLVKF 185
++ +DAAINPGNSGGPLLDS G +IGINTAI SG S G+GF++P DT I+ L+++
Sbjct: 67 IIQTDAAINPGNSGGPLLDSEGKVIGINTAIIGTSG-SVGIGFAVPSDTALRILPDLLEY 125
Query: 186 GKVTRPILGIKFAPDQSVEQLGV---SGVLVLDAPANGPAGKAGL-QSTKRDSYGRLIL- 240
G V RP LGI+ P ++++G+ GVL+ A AGL +TK G +
Sbjct: 126 GYVRRPWLGIEPIPTVYLQRIGIDVPDGVLIARVVKGASADLAGLIGATKTVKVGNYKIP 185
Query: 241 --GDIITSVNGTKVTSGSDLYRILDQCKVGDKVTVEVLRGDHKEKIPVILEPKP 292
GDIIT +N TKVT+ DL + ++ + GD + ++++R + + I V L +P
Sbjct: 186 WGGDIITHINQTKVTTMEDLAQQIETRRSGDIINIKLIRNERELSIKVKLSLRP 239
>UniRef100_Q57155 Serine protease MucD [Pseudomonas aeruginosa]
Length = 474
Score = 120 bits (302), Expect = 3e-26
Identities = 69/190 (36%), Positives = 104/190 (54%), Gaps = 16/190 (8%)
Query: 110 IPGASDLSLDLTTLLQLVSSDAAINPGNSGGPLLDSSGNLIGINTAIYSPSGASSGVGFS 169
I A SL + + + +D AINPGNSGGPLL+ G ++GIN+ I++ SG G+ F+
Sbjct: 189 IVSAKGRSLPNESYVPFIQTDVAINPGNSGGPLLNLQGEVVGINSQIFTRSGGFMGLSFA 248
Query: 170 IPLDTVNGIVDQLVKFGKVTRPILG--IKFAPDQSVEQLGV---SGVLVLDAPANGPAGK 224
IP+D + DQL K GKV+R LG I+ E G+ SG LV +GPA K
Sbjct: 249 IPIDVALNVADQLKKAGKVSRGWLGVVIQEVNKDLAESFGLDKPSGALVAQLVEDGPAAK 308
Query: 225 AGLQSTKRDSYGRLILGDIITSVNGTKVTSGSDLYRILDQCKVGDKVTVEVLRGDHKEKI 284
GLQ +GD+I S+NG + +DL ++ K GDK+ ++V+R ++ +
Sbjct: 309 GGLQ-----------VGDVILSLNGQSINESADLPHLVGNMKPGDKINLDVIRNGQRKSL 357
Query: 285 PVILEPKPDE 294
+ + PD+
Sbjct: 358 SMAVGSLPDD 367
>UniRef100_UPI00002D1A2D UPI00002D1A2D UniRef100 entry
Length = 338
Score = 120 bits (301), Expect = 4e-26
Identities = 72/175 (41%), Positives = 107/175 (61%), Gaps = 10/175 (5%)
Query: 126 LVSSDAAINPGNSGGPLLDSSGNLIGINTAIYSPSGASSGVGFSIPLDTVNGIVDQLVKF 185
++ +DAAINPGNSGGPLLDS G +IGINTAI SG S G+GF++P DT I+ L+++
Sbjct: 164 IIQTDAAINPGNSGGPLLDSEGKVIGINTAIIGTSG-SVGIGFAVPSDTALRILPDLLEY 222
Query: 186 GKVTRPILGIKFAPDQSVEQLGV---SGVLVLDAPANGPAGKAGL----QSTKRDSYGRL 238
G V RP LGI+ P + ++G+ GVL+ A +AGL ++ K +Y
Sbjct: 223 GYVRRPWLGIEPIPTVYLRRIGIDVPEGVLISRVVKGASADQAGLLGANKTVKVGNYKIP 282
Query: 239 ILGDIITSVNGTKVTSGSDLYRILDQCKVGDKVTVEVLRGDH--KEKIPVILEPK 291
GDIIT +N TKVT+ DL + ++ + GD + ++++R + K+ + L PK
Sbjct: 283 WGGDIITHINQTKVTTMEDLAKQIETRRSGDVINIKLIRNERVLTIKVELFLRPK 337
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.317 0.136 0.396
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 503,192,670
Number of Sequences: 2790947
Number of extensions: 21233750
Number of successful extensions: 66615
Number of sequences better than 10.0: 1510
Number of HSP's better than 10.0 without gapping: 1206
Number of HSP's successfully gapped in prelim test: 304
Number of HSP's that attempted gapping in prelim test: 63057
Number of HSP's gapped (non-prelim): 2484
length of query: 295
length of database: 848,049,833
effective HSP length: 126
effective length of query: 169
effective length of database: 496,390,511
effective search space: 83889996359
effective search space used: 83889996359
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 74 (33.1 bits)
Lotus: description of TM0225.5


