

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0221.5
(190 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
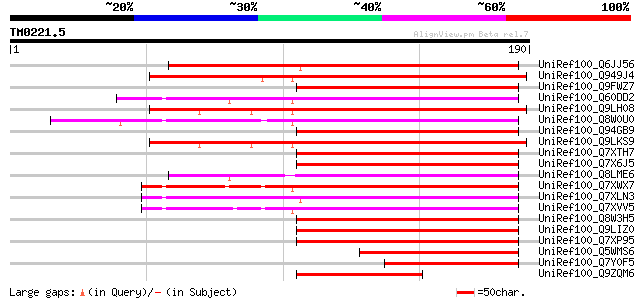
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q6JJ56 Putative copia-like polyprotein [Ipomoea trifida] 100 3e-20
UniRef100_Q949J4 Putative copia-like polyprotein [Lycopersicon e... 100 3e-20
UniRef100_Q9FWZ7 F11O6.4 protein [Arabidopsis thaliana] 91 2e-17
UniRef100_Q60DD2 Integrase core domain containing protein [Oryza... 91 2e-17
UniRef100_Q9LH08 Copia-type pol polyprotein-like [Arabidopsis th... 90 3e-17
UniRef100_Q8W0U0 Putative copia polyprotein [Sorghum bicolor] 90 3e-17
UniRef100_Q94GB9 Putative copia-type pol polyprotein [Oryza sativa] 89 8e-17
UniRef100_Q9LKS9 Hypothetical protein T15F17.l [Arabidopsis thal... 88 1e-16
UniRef100_Q7XTH7 P0041A24.2 protein [Oryza sativa] 87 2e-16
UniRef100_Q7X6J5 OSJNBb0067G11.4 protein [Oryza sativa] 86 5e-16
UniRef100_Q8LME6 Putative retroelement [Oryza sativa] 84 2e-15
UniRef100_Q7XWX7 OSJNBa0036B17.5 protein [Oryza sativa] 84 2e-15
UniRef100_Q7XLN3 OSJNBa0049H08.8 protein [Oryza sativa] 83 4e-15
UniRef100_Q7XVV5 OSJNBa0065J03.17 protein [Oryza sativa] 82 9e-15
UniRef100_Q8W3H5 Putative polyprotein [Oryza sativa] 80 3e-14
UniRef100_Q9LIZ0 Similar to putative copia polyprotein [Oryza sa... 79 5e-14
UniRef100_Q7XP95 OSJNBa0021F22.5 protein [Oryza sativa] 79 6e-14
UniRef100_Q5WMS6 Hypothetical protein OJ1333_C12.5 [Oryza sativa] 66 4e-10
UniRef100_Q7Y0F5 Putative polyprotein [Oryza sativa] 55 1e-06
UniRef100_Q9ZQM6 Putative retroelement pol polyprotein [Arabidop... 46 6e-04
>UniRef100_Q6JJ56 Putative copia-like polyprotein [Ipomoea trifida]
Length = 1190
Score = 100 bits (248), Expect = 3e-20
Identities = 57/131 (43%), Positives = 83/131 (62%), Gaps = 3/131 (2%)
Query: 59 CIFMMRYEIVYVIINVYDNDLLRL*KSSQKL*IC*RKNLR*KS*E*N---FCLAL*IEYL 115
CIF+ + + II+VY +D+ + + + C + + FCL L IE+L
Sbjct: 829 CIFIKKSSNGFCIISVYVDDMNIIGTNKDIIEACSYLKAEFEMKDLGKTKFCLGLQIEHL 888
Query: 116 KDEVFMYQRTYITKVLKGFYMDKSCPLCTLLILRSLNVNKDPFRP*EKNEELLDPEVLYL 175
+ +F++Q Y+ K+L+ FYMDKS PL T +++RSL +KDPFRP E NE++L PEV YL
Sbjct: 889 PNGIFIHQANYVNKLLEKFYMDKSHPLSTPMVVRSLEADKDPFRPKEDNEDILGPEVPYL 948
Query: 176 SAIKALLYLAS 186
SAI ALLYLA+
Sbjct: 949 SAIGALLYLAN 959
>UniRef100_Q949J4 Putative copia-like polyprotein [Lycopersicon esculentum]
Length = 779
Score = 100 bits (248), Expect = 3e-20
Identities = 61/141 (43%), Positives = 85/141 (60%), Gaps = 3/141 (2%)
Query: 52 QNDPTNSCIFMMRYEIVYVIINVYDNDLLRL*KSSQKL*I--C*RKNLR*KS*-E*NFCL 108
+ND CIF+ R +VII VY +DL + + L C ++ K + FCL
Sbjct: 547 KNDSICPCIFIKRLGSEFVIIAVYVDDLNIIGTRKELLEAVECLKREFEMKDLGKTKFCL 606
Query: 109 AL*IEYLKDEVFMYQRTYITKVLKGFYMDKSCPLCTLLILRSLNVNKDPFRP*EKNEELL 168
L IE L + + ++Q TY K+LK FYMD S PL T +++RSL++N DPFRP E +EELL
Sbjct: 607 GLQIENLSNGILVHQSTYTEKILKRFYMDNSHPLSTPMVVRSLDINTDPFRPQENDEELL 666
Query: 169 DPEVLYLSAIKALLYLASHCK 189
E YLSAI AL+YL ++ +
Sbjct: 667 GDETPYLSAIGALMYLVNNTR 687
>UniRef100_Q9FWZ7 F11O6.4 protein [Arabidopsis thaliana]
Length = 484
Score = 90.5 bits (223), Expect = 2e-17
Identities = 44/81 (54%), Positives = 58/81 (71%)
Query: 106 FCLAL*IEYLKDEVFMYQRTYITKVLKGFYMDKSCPLCTLLILRSLNVNKDPFRP*EKNE 165
FCL L IE+ +D +F++Q Y K+LK F MDK+ PL T ++ RSLNV DPFRP E NE
Sbjct: 288 FCLGLQIEHFQDGIFVHQSNYTKKILKRFNMDKANPLSTPMVNRSLNVENDPFRPCEDNE 347
Query: 166 ELLDPEVLYLSAIKALLYLAS 186
+ L P+V Y+SAI L+YLA+
Sbjct: 348 DFLGPKVPYMSAIGGLMYLAN 368
>UniRef100_Q60DD2 Integrase core domain containing protein [Oryza sativa]
Length = 1419
Score = 90.5 bits (223), Expect = 2e-17
Identities = 62/150 (41%), Positives = 89/150 (59%), Gaps = 4/150 (2%)
Query: 40 RQSR*LFS*GRSQNDPTNSCIFMMRYEIVYVIINVYDNDL--LRL*KSSQKL*IC*RKNL 97
R S L G + ND C+F+ + E + II+VY +DL + + ++ +
Sbjct: 1047 RLSNFLLRRGYANNDDC-LCVFIKKSENGFCIISVYVDDLNIIGTTRDIEEASTYLKAEF 1105
Query: 98 R*KS*-E*NFCLAL*IEYLKDEVFMYQRTYITKVLKGFYMDKSCPLCTLLILRSLNVNKD 156
K + FCL L +E+L VF++Q TY KVL+ F MDKS PL T +++RSL V+KD
Sbjct: 1106 EMKDLGKTKFCLGLQLEHLPKGVFVHQSTYTKKVLEMFNMDKSHPLKTPMVVRSLEVDKD 1165
Query: 157 PFRP*EKNEELLDPEVLYLSAIKALLYLAS 186
PFRP E++E+ L PEV YLSAI L+YL +
Sbjct: 1166 PFRPKEEDEKPLGPEVPYLSAIGPLMYLTN 1195
>UniRef100_Q9LH08 Copia-type pol polyprotein-like [Arabidopsis thaliana]
Length = 1123
Score = 90.1 bits (222), Expect = 3e-17
Identities = 57/142 (40%), Positives = 87/142 (61%), Gaps = 4/142 (2%)
Query: 52 QNDPTNSCIFMMRYEIV-YVIINVYDNDLLRL*KSSQ--KL*IC*RKNLR*KS*-E*NFC 107
+NDP + CIF+ ++ +VII VY +DL L S + + +K K + FC
Sbjct: 787 KNDPISPCIFIKKFASKGFVIIAVYVDDLNILGTSGEIAQTVEYLKKEFEMKDLGKTKFC 846
Query: 108 LAL*IEYLKDEVFMYQRTYITKVLKGFYMDKSCPLCTLLILRSLNVNKDPFRP*EKNEEL 167
L L +EY+ + ++Q+ Y VLK F MDK+ PL + + +RSL ++ DPF P + +EE+
Sbjct: 847 LGLQLEYIDKGILVHQKAYTETVLKRFNMDKAHPLTSPMQVRSLGLDSDPFGPKKDDEEI 906
Query: 168 LDPEVLYLSAIKALLYLASHCK 189
L PE+ YLSAI AL+YL+SH +
Sbjct: 907 LGPEMPYLSAIGALMYLSSHTR 928
>UniRef100_Q8W0U0 Putative copia polyprotein [Sorghum bicolor]
Length = 1082
Score = 89.7 bits (221), Expect = 3e-17
Identities = 63/179 (35%), Positives = 101/179 (56%), Gaps = 11/179 (6%)
Query: 16 NSLKIVRCLKHTIHVLNNEQITRL---RQSR*LFS*GRSQNDPTNSCIFMMRYEIVYVII 72
N+ + + C+K + +Q R+ R S L G S +D C+F+ + + + II
Sbjct: 730 NTNRNMYCVKLQKSLYGLKQSGRMWYNRLSEYLLQKGYSNSDGC-PCVFIKKSQTGFCII 788
Query: 73 NVYDNDLLRL*KSSQKL*IC*RKNLR*KS*-----E*NFCLAL*IEYLKDEVFMYQRTYI 127
+VY +DL + C K+L+ + + FCL L +E+L +F++Q Y+
Sbjct: 789 SVYVDDLNIIGHKQDIDEAC--KHLKTEFELKDLGKTKFCLGLQLEHLSTGIFVHQSAYV 846
Query: 128 TKVLKGFYMDKSCPLCTLLILRSLNVNKDPFRP*EKNEELLDPEVLYLSAIKALLYLAS 186
KVL+ F MDK+ P T +++RSL +N DPFRP ++ EE+L P V YLSA+ AL+YLA+
Sbjct: 847 QKVLERFNMDKAYPSKTPMVVRSLGLNTDPFRPQDEGEEILGPHVPYLSAVGALMYLAN 905
>UniRef100_Q94GB9 Putative copia-type pol polyprotein [Oryza sativa]
Length = 1225
Score = 88.6 bits (218), Expect = 8e-17
Identities = 45/81 (55%), Positives = 60/81 (73%)
Query: 106 FCLAL*IEYLKDEVFMYQRTYITKVLKGFYMDKSCPLCTLLILRSLNVNKDPFRP*EKNE 165
FCL L +++L + VFM+Q TY +VL F MDK PL T +++RSL +KDPF+P E +E
Sbjct: 921 FCLGLQLKHLPEGVFMHQSTYTKRVLGKFNMDKCHPLKTPMVVRSLEADKDPFKPKEDDE 980
Query: 166 ELLDPEVLYLSAIKALLYLAS 186
E+L PEV YLSAI AL+YLA+
Sbjct: 981 EVLGPEVPYLSAIGALMYLAN 1001
>UniRef100_Q9LKS9 Hypothetical protein T15F17.l [Arabidopsis thaliana]
Length = 1141
Score = 88.2 bits (217), Expect = 1e-16
Identities = 56/142 (39%), Positives = 86/142 (60%), Gaps = 4/142 (2%)
Query: 52 QNDPTNSCIFMMRYEIV-YVIINVYDNDLLRL*KSSQ--KL*IC*RKNLR*KS*-E*NFC 107
+NDP + CIF+ ++ +VII VY +DL L S + + +K K + FC
Sbjct: 787 KNDPISPCIFIKKFASKGFVIIAVYVDDLNILGTSGEIAQTVEYLKKEFEMKDLGKTKFC 846
Query: 108 LAL*IEYLKDEVFMYQRTYITKVLKGFYMDKSCPLCTLLILRSLNVNKDPFRP*EKNEEL 167
L L +EY+ + ++QR Y +LK F MDK+ PL + + +RSL ++ DPF P + +EE+
Sbjct: 847 LGLQLEYVDKGILVHQRAYTETILKRFNMDKAHPLTSPMQVRSLGLDSDPFGPKKDDEEI 906
Query: 168 LDPEVLYLSAIKALLYLASHCK 189
L PE+ YLSAI A +YL+SH +
Sbjct: 907 LGPEMPYLSAIGAWMYLSSHTR 928
>UniRef100_Q7XTH7 P0041A24.2 protein [Oryza sativa]
Length = 1376
Score = 87.4 bits (215), Expect = 2e-16
Identities = 44/81 (54%), Positives = 60/81 (73%)
Query: 106 FCLAL*IEYLKDEVFMYQRTYITKVLKGFYMDKSCPLCTLLILRSLNVNKDPFRP*EKNE 165
FCL L +E+L + VFM+Q TY +VL F MDK PL T +++RSL +KDPF+P E +E
Sbjct: 1072 FCLGLQLEHLPEGVFMHQSTYTKRVLGKFNMDKCHPLKTPMVVRSLEADKDPFKPKEDDE 1131
Query: 166 ELLDPEVLYLSAIKALLYLAS 186
++L PEV YL+AI AL+YLA+
Sbjct: 1132 KVLGPEVPYLNAIGALMYLAN 1152
>UniRef100_Q7X6J5 OSJNBb0067G11.4 protein [Oryza sativa]
Length = 986
Score = 85.9 bits (211), Expect = 5e-16
Identities = 44/81 (54%), Positives = 60/81 (73%)
Query: 106 FCLAL*IEYLKDEVFMYQRTYITKVLKGFYMDKSCPLCTLLILRSLNVNKDPFRP*EKNE 165
FCL L +E+L + VFM+Q TY +VL F M+K PL T ++++SL +KDPF+P E +E
Sbjct: 836 FCLDLQLEHLPEGVFMHQSTYTKRVLGKFNMNKCHPLKTPMVVQSLEADKDPFKPKEDDE 895
Query: 166 ELLDPEVLYLSAIKALLYLAS 186
E+L PEV YLSAI AL+YLA+
Sbjct: 896 EVLGPEVPYLSAIGALMYLAN 916
>UniRef100_Q8LME6 Putative retroelement [Oryza sativa]
Length = 852
Score = 84.0 bits (206), Expect = 2e-15
Identities = 56/134 (41%), Positives = 77/134 (56%), Gaps = 9/134 (6%)
Query: 59 CIFMMRYEIVYVIINVYDNDL------LRL*KSSQKL*IC*RKNLR*KS*E*NFCLAL*I 112
C+FM R E + II+VY +DL + ++S L K+ +CL L +
Sbjct: 498 CLFMKRSEHGFCIISVYVDDLNIIGTTQVIKEASSYLKTEFEMKELGKT---TYCLGLQL 554
Query: 113 EYLKDEVFMYQRTYITKVLKGFYMDKSCPLCTLLILRSLNVNKDPFRP*EKNEELLDPEV 172
E+ + V ++Q YI K+L+ F M S P T +++RSL V DPFRP E NEELL PE
Sbjct: 555 EHTPEGVLLHQYAYIQKILEKFNMKDSYPTRTPMVVRSLAVESDPFRPQENNEELLGPEC 614
Query: 173 LYLSAIKALLYLAS 186
YLSAI AL+YLA+
Sbjct: 615 PYLSAIGALMYLAN 628
>UniRef100_Q7XWX7 OSJNBa0036B17.5 protein [Oryza sativa]
Length = 443
Score = 83.6 bits (205), Expect = 2e-15
Identities = 55/143 (38%), Positives = 87/143 (60%), Gaps = 8/143 (5%)
Query: 49 GRSQNDPTNSCIFMMRYEIVYVIINVYDNDLLRL*KSSQKL*IC*RKNLR*KS*-----E 103
G ++ND C+F+ + + II+VY +DL + ++Q + R +L+ + +
Sbjct: 77 GYTKNDDC-PCVFIKKSPTGFCIISVYVDDL-NIIGNTQDINEA-RHHLKTEFEMKDLGQ 133
Query: 104 *NFCLAL*IEYLKDEVFMYQRTYITKVLKGFYMDKSCPLCTLLILRSLNVNKDPFRP*EK 163
FCL L +E+L + ++Q Y K+L+ F MDKS P T +++RSL+V KDPFRP E
Sbjct: 134 TKFCLGLQLEHLPSGILVHQSAYAQKMLEKFNMDKSYPSKTPMVVRSLDVEKDPFRPKED 193
Query: 164 NEELLDPEVLYLSAIKALLYLAS 186
E++L P+ YLSAI AL+YLA+
Sbjct: 194 GEDVLGPDFPYLSAIGALMYLAN 216
>UniRef100_Q7XLN3 OSJNBa0049H08.8 protein [Oryza sativa]
Length = 810
Score = 82.8 bits (203), Expect = 4e-15
Identities = 51/141 (36%), Positives = 80/141 (56%), Gaps = 4/141 (2%)
Query: 49 GRSQNDPTNSCIFMMRYEIVYVIINVYDNDLLRL*KSSQKL*IC*RKNLR*KS*E*N--- 105
G ++ND C+F+ + + I+VY +DL + + C + +
Sbjct: 601 GYTKNDDF-PCVFIKKSPTGFCFISVYVDDLNIIRNTQDINEACHHLKTEFEMKDLGQTK 659
Query: 106 FCLAL*IEYLKDEVFMYQRTYITKVLKGFYMDKSCPLCTLLILRSLNVNKDPFRP*EKNE 165
FCL L +++L + ++Q Y K+L+ F MDKS P T +++RSL+V KDPFRP E E
Sbjct: 660 FCLGLQLDHLPSRILVHQSAYTQKILEKFNMDKSYPSKTPMVVRSLDVEKDPFRPKEYGE 719
Query: 166 ELLDPEVLYLSAIKALLYLAS 186
++L P+ YLSAI AL+YLA+
Sbjct: 720 DVLGPDFPYLSAIGALMYLAN 740
>UniRef100_Q7XVV5 OSJNBa0065J03.17 protein [Oryza sativa]
Length = 864
Score = 81.6 bits (200), Expect = 9e-15
Identities = 54/143 (37%), Positives = 86/143 (59%), Gaps = 8/143 (5%)
Query: 49 GRSQNDPTNSCIFMMRYEIVYVIINVYDNDLLRL*KSSQKL*IC*RKNLR*KS*-----E 103
G ++ND C+F+ + + II+VY +DL + ++Q + R +L+ + +
Sbjct: 498 GYTKNDDC-PCVFIKKSPTGFCIISVYVDDLSII-GNTQDINEA-RHHLKTEFEMKDLGQ 554
Query: 104 *NFCLAL*IEYLKDEVFMYQRTYITKVLKGFYMDKSCPLCTLLILRSLNVNKDPFRP*EK 163
FCL L +E+L + ++Q Y K+L+ F MDKS P +++RSL+V KDPFRP E
Sbjct: 555 TKFCLGLQLEHLPSGILVHQSAYTQKILEKFNMDKSYPSKIPMVVRSLDVEKDPFRPKED 614
Query: 164 NEELLDPEVLYLSAIKALLYLAS 186
E++L P+ YLSAI AL+YLA+
Sbjct: 615 GEDVLGPDFPYLSAIGALMYLAN 637
>UniRef100_Q8W3H5 Putative polyprotein [Oryza sativa]
Length = 419
Score = 80.1 bits (196), Expect = 3e-14
Identities = 41/81 (50%), Positives = 58/81 (70%)
Query: 106 FCLAL*IEYLKDEVFMYQRTYITKVLKGFYMDKSCPLCTLLILRSLNVNKDPFRP*EKNE 165
FCL L +E+L + + ++Q TY KVL+ F M K+ PL T +++RSL+ +KD FRP + E
Sbjct: 111 FCLVLQVEHLPNGILVHQSTYTKKVLERFNMSKAHPLKTPMVVRSLDKDKDLFRPRSETE 170
Query: 166 ELLDPEVLYLSAIKALLYLAS 186
E L PE+ YLSAI AL+YLA+
Sbjct: 171 EELGPEIPYLSAIGALMYLAN 191
>UniRef100_Q9LIZ0 Similar to putative copia polyprotein [Oryza sativa]
Length = 1321
Score = 79.3 bits (194), Expect = 5e-14
Identities = 40/81 (49%), Positives = 55/81 (67%)
Query: 106 FCLAL*IEYLKDEVFMYQRTYITKVLKGFYMDKSCPLCTLLILRSLNVNKDPFRP*EKNE 165
+CL L +E+ + V ++Q YI K+L+ F + S P T +++RSL V DPFRP E N+
Sbjct: 1078 YCLGLQLEHTHEGVLLHQSAYIQKILEKFNIKDSYPTRTPMVVRSLAVESDPFRPQENNQ 1137
Query: 166 ELLDPEVLYLSAIKALLYLAS 186
ELL PE YLSAI AL+YLA+
Sbjct: 1138 ELLGPECPYLSAIGALMYLAN 1158
>UniRef100_Q7XP95 OSJNBa0021F22.5 protein [Oryza sativa]
Length = 911
Score = 79.0 bits (193), Expect = 6e-14
Identities = 40/81 (49%), Positives = 57/81 (69%)
Query: 106 FCLAL*IEYLKDEVFMYQRTYITKVLKGFYMDKSCPLCTLLILRSLNVNKDPFRP*EKNE 165
FCL+L +E+L + ++Q Y K+L+ F MDKS P T +++RSL+V KD FRP E E
Sbjct: 779 FCLSLQLEHLPFGILVHQNAYTQKILEKFNMDKSYPSKTPMVVRSLDVEKDLFRPKEDGE 838
Query: 166 ELLDPEVLYLSAIKALLYLAS 186
++L P+ YLSAI AL+YLA+
Sbjct: 839 DVLGPDFPYLSAIGALMYLAN 859
>UniRef100_Q5WMS6 Hypothetical protein OJ1333_C12.5 [Oryza sativa]
Length = 710
Score = 66.2 bits (160), Expect = 4e-10
Identities = 33/58 (56%), Positives = 44/58 (74%)
Query: 129 KVLKGFYMDKSCPLCTLLILRSLNVNKDPFRP*EKNEELLDPEVLYLSAIKALLYLAS 186
K+L+ F MDKS P T +++RSL+V KDPFRP E E++L P+ YLSAI AL+YLA+
Sbjct: 464 KILEKFNMDKSYPSKTPMVVRSLDVEKDPFRPKEDGEDVLGPDFPYLSAIGALMYLAN 521
>UniRef100_Q7Y0F5 Putative polyprotein [Oryza sativa]
Length = 639
Score = 54.7 bits (130), Expect = 1e-06
Identities = 27/49 (55%), Positives = 36/49 (73%)
Query: 138 KSCPLCTLLILRSLNVNKDPFRP*EKNEELLDPEVLYLSAIKALLYLAS 186
KS P T +++RSL+V KDPFRP E E++L P+ YLSAI AL+Y A+
Sbjct: 445 KSYPSKTPMVVRSLDVEKDPFRPKEDGEDVLGPDFPYLSAIGALMYFAN 493
>UniRef100_Q9ZQM6 Putative retroelement pol polyprotein [Arabidopsis thaliana]
Length = 557
Score = 45.8 bits (107), Expect = 6e-04
Identities = 22/46 (47%), Positives = 30/46 (64%)
Query: 106 FCLAL*IEYLKDEVFMYQRTYITKVLKGFYMDKSCPLCTLLILRSL 151
FCL L IE+LK+ + YQ TY VLK FYMD + PL + ++ + L
Sbjct: 401 FCLGLQIEHLKNVILEYQETYTKNVLKRFYMDGAYPLSSPMVGKRL 446
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.359 0.163 0.534
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 246,799,120
Number of Sequences: 2790947
Number of extensions: 7833076
Number of successful extensions: 29567
Number of sequences better than 10.0: 27
Number of HSP's better than 10.0 without gapping: 22
Number of HSP's successfully gapped in prelim test: 5
Number of HSP's that attempted gapping in prelim test: 29530
Number of HSP's gapped (non-prelim): 36
length of query: 190
length of database: 848,049,833
effective HSP length: 120
effective length of query: 70
effective length of database: 513,136,193
effective search space: 35919533510
effective search space used: 35919533510
T: 11
A: 40
X1: 14 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.8 bits)
S2: 71 (32.0 bits)
Lotus: description of TM0221.5


