

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0219.2
(365 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
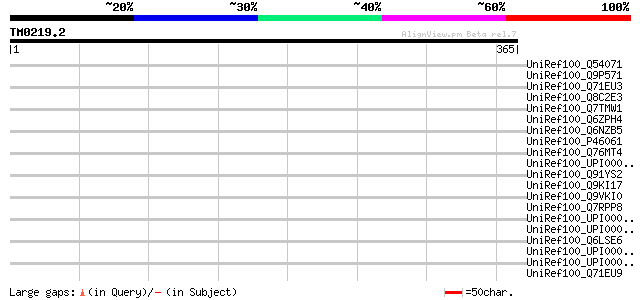
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q54071 M protein precursor; MSzW60 [Streptococcus equi] 45 0.002
UniRef100_Q9P571 Related to actin-interacting protein AIP3 [Neur... 45 0.004
UniRef100_Q71EU3 Szp protein [Streptococcus equi subsp. zooepide... 44 0.005
UniRef100_Q8C2E3 Mus musculus 2 days neonate thymus thymic cells... 44 0.009
UniRef100_Q7TMW1 Rangap1 protein [Mus musculus] 44 0.009
UniRef100_Q6ZPH4 MKIAA1835 protein [Mus musculus] 44 0.009
UniRef100_Q6NZB5 Rangap1 protein [Mus musculus] 44 0.009
UniRef100_P46061 Ran GTPase-activating protein 1 [Mus musculus] 44 0.009
UniRef100_Q76MT4 ABTAP [Rattus rattus] 43 0.012
UniRef100_UPI000023EDFF UPI000023EDFF UniRef100 entry 43 0.016
UniRef100_Q91YS2 Rangap1 protein [Mus musculus] 42 0.020
UniRef100_Q9KI17 M-like protein Szp3 precursor [Streptococcus eq... 42 0.027
UniRef100_Q9VKI0 CG6392-PA [Drosophila melanogaster] 42 0.027
UniRef100_Q7RPP8 Hypothetical protein [Plasmodium yoelii yoelii] 42 0.027
UniRef100_UPI00003640AE UPI00003640AE UniRef100 entry 42 0.035
UniRef100_UPI00003640AD UPI00003640AD UniRef100 entry 42 0.035
UniRef100_Q6LSE6 Hypothetical methyl-accepting chemotaxis protei... 42 0.035
UniRef100_UPI000023DA4F UPI000023DA4F UniRef100 entry 41 0.046
UniRef100_UPI0000219DB0 UPI0000219DB0 UniRef100 entry 41 0.046
UniRef100_Q71EU9 Szp protein [Streptococcus equi subsp. zooepide... 41 0.046
>UniRef100_Q54071 M protein precursor; MSzW60 [Streptococcus equi]
Length = 376
Score = 45.4 bits (106), Expect = 0.002
Identities = 44/187 (23%), Positives = 81/187 (42%), Gaps = 19/187 (10%)
Query: 163 SAASIVAGMAQCVKELIATKNRYEKKAADYKTAYEQAKADAETANKNLKAAEERCAKLTD 222
+A + +A + Q + +L EK+ A + EQ K A ++N A E+R +L+
Sbjct: 94 TAEATIASLEQKMNDL--ESKIQEKQKAFEQVVLEQKKEQANPVDRNGDAEEDRLDELSG 151
Query: 223 DL----AASDLLLQKTKSLKETINDKHTAVQAKCQKLEKKYERLNASILG------RASL 272
++ AA ++ LQKTK +T + ++ + Q K + N + G +A
Sbjct: 152 EIRREKAALEVELQKTKEALDTAKRAYAGIEERKQVAAVKLDAANKAFAGVEEKHDQAMA 211
Query: 273 LFAQGFLAAKEQISVVEPGFDLSRIGWLKEIKDGQVVGDDDISLDLL------PQFDDES 326
FA+ F A KE + S + K+I V + +++ P+ + E+
Sbjct: 212 KFAEAFAAYKEAVKAELKAAGASDF-YTKKIDSANTVAGVNTLREMILDSIAKPEVEPEA 270
Query: 327 EPEEDGE 333
+PE E
Sbjct: 271 KPEPKPE 277
>UniRef100_Q9P571 Related to actin-interacting protein AIP3 [Neurospora crassa]
Length = 1127
Score = 44.7 bits (104), Expect = 0.004
Identities = 28/91 (30%), Positives = 45/91 (48%), Gaps = 8/91 (8%)
Query: 282 KEQISVVEPGFDLSRIGWLKEIKDGQVVGDDDISLD-------LLPQFDDESEPEEDGED 334
KE S VE G L + G ++E++ + DD I + L+ + DD+ E EED E+
Sbjct: 984 KELASFVEEG-KLKKSGGVEEVERARKAKDDQIRREVWERMNGLIAENDDDGEDEEDDEE 1042
Query: 335 GNEQHKNEDHEKEDPQAGTSQGNNANNENLA 365
E+ + E+ E+E+ + G Q E A
Sbjct: 1043 EEEEEEEEEGEEEEEEEGEEQEEGEEGEEKA 1073
>UniRef100_Q71EU3 Szp protein [Streptococcus equi subsp. zooepidemicus]
Length = 375
Score = 44.3 bits (103), Expect = 0.005
Identities = 46/187 (24%), Positives = 82/187 (43%), Gaps = 19/187 (10%)
Query: 163 SAASIVAGMAQCVKELIATKNRYEKKAADYKTAYEQAKADAETANKNLKAAEERCAKLTD 222
+A + +A + Q + +L EK+ A + EQ K A ++N A E+R +L+
Sbjct: 93 TAEATIAELEQKMNDL--ESKIQEKQKAFEQVVLEQKKEQANPVDRNGDAEEDRLDELSG 150
Query: 223 DL----AASDLLLQKTKSLKETINDKHTAVQAKCQKLEKKYERLNASILG------RASL 272
++ AA ++ LQKTK +T + ++ + Q K + N + G +A
Sbjct: 151 EIRREKAALEVELQKTKEALDTAKRAYAGIEERKQVAAAKLDAANKAFAGVEEKHAQAMA 210
Query: 273 LFAQGFLAAKEQISVVEPGFDLSRIGWLKEIKDGQVVGD----DDISLDLL--PQFDDES 326
FA+ F A KE + S + K+I V ++ LD + P+ + E+
Sbjct: 211 KFAEAFAAYKEAVKAELKAAGASDF-YTKKIDSANTVDGVKTLREMILDSIAKPEVEPEA 269
Query: 327 EPEEDGE 333
+PE E
Sbjct: 270 KPEPKPE 276
>UniRef100_Q8C2E3 Mus musculus 2 days neonate thymus thymic cells cDNA, RIKEN full-
length enriched library, clone:E430025K03 product:RAN
GTPase activating protein 1, full insert sequence [Mus
musculus]
Length = 589
Score = 43.5 bits (101), Expect = 0.009
Identities = 29/107 (27%), Positives = 53/107 (49%), Gaps = 14/107 (13%)
Query: 249 QAKCQKLEKKYERLNASILGRASLLFAQGFLAAKEQISVVEPGFDLSRIGWLKEIKDGQV 308
+A K E + LN + LG EQ+ V F+++++ L + D +
Sbjct: 315 EAVADKAELEKLDLNGNALGEEGC----------EQLQEVMDSFNMAKV--LASLSDDE- 361
Query: 309 VGDDDISLDLLPQFDDESEPEEDGEDGNEQHKNEDHEKEDPQAGTSQ 355
G+D+ + + D+E E EED ED +E+ + ++ E+E PQ G+ +
Sbjct: 362 -GEDEDEEEEGEEDDEEEEDEEDEEDDDEEEEEQEEEEEPPQRGSGE 407
>UniRef100_Q7TMW1 Rangap1 protein [Mus musculus]
Length = 589
Score = 43.5 bits (101), Expect = 0.009
Identities = 29/107 (27%), Positives = 53/107 (49%), Gaps = 14/107 (13%)
Query: 249 QAKCQKLEKKYERLNASILGRASLLFAQGFLAAKEQISVVEPGFDLSRIGWLKEIKDGQV 308
+A K E + LN + LG EQ+ V F+++++ L + D +
Sbjct: 315 EAVADKAELEKLDLNGNALGEEGC----------EQLQEVMDSFNMAKV--LASLSDDE- 361
Query: 309 VGDDDISLDLLPQFDDESEPEEDGEDGNEQHKNEDHEKEDPQAGTSQ 355
G+D+ + + D+E E EED ED +E+ + ++ E+E PQ G+ +
Sbjct: 362 -GEDEDEEEEGEEDDEEEEDEEDEEDDDEEEEEQEEEEEPPQRGSGE 407
>UniRef100_Q6ZPH4 MKIAA1835 protein [Mus musculus]
Length = 646
Score = 43.5 bits (101), Expect = 0.009
Identities = 29/107 (27%), Positives = 53/107 (49%), Gaps = 14/107 (13%)
Query: 249 QAKCQKLEKKYERLNASILGRASLLFAQGFLAAKEQISVVEPGFDLSRIGWLKEIKDGQV 308
+A K E + LN + LG EQ+ V F+++++ L + D +
Sbjct: 372 EAVADKAELEKLDLNGNALGEEGC----------EQLQEVMDSFNMAKV--LASLSDDE- 418
Query: 309 VGDDDISLDLLPQFDDESEPEEDGEDGNEQHKNEDHEKEDPQAGTSQ 355
G+D+ + + D+E E EED ED +E+ + ++ E+E PQ G+ +
Sbjct: 419 -GEDEDEEEEGEEDDEEEEDEEDEEDDDEEEEEQEEEEEPPQRGSGE 464
>UniRef100_Q6NZB5 Rangap1 protein [Mus musculus]
Length = 589
Score = 43.5 bits (101), Expect = 0.009
Identities = 29/107 (27%), Positives = 53/107 (49%), Gaps = 14/107 (13%)
Query: 249 QAKCQKLEKKYERLNASILGRASLLFAQGFLAAKEQISVVEPGFDLSRIGWLKEIKDGQV 308
+A K E + LN + LG EQ+ V F+++++ L + D +
Sbjct: 315 EAVADKAELEKLDLNGNALGEEGC----------EQLQEVMDSFNMAKV--LASLSDDE- 361
Query: 309 VGDDDISLDLLPQFDDESEPEEDGEDGNEQHKNEDHEKEDPQAGTSQ 355
G+D+ + + D+E E EED ED +E+ + ++ E+E PQ G+ +
Sbjct: 362 -GEDEDEEEEGEEDDEEEEDEEDEEDDDEEEEEQEEEEEPPQRGSGE 407
>UniRef100_P46061 Ran GTPase-activating protein 1 [Mus musculus]
Length = 589
Score = 43.5 bits (101), Expect = 0.009
Identities = 29/107 (27%), Positives = 53/107 (49%), Gaps = 14/107 (13%)
Query: 249 QAKCQKLEKKYERLNASILGRASLLFAQGFLAAKEQISVVEPGFDLSRIGWLKEIKDGQV 308
+A K E + LN + LG EQ+ V F+++++ L + D +
Sbjct: 315 EAVADKAELEKLDLNGNALGEEGC----------EQLQEVMDSFNMAKV--LASLSDDE- 361
Query: 309 VGDDDISLDLLPQFDDESEPEEDGEDGNEQHKNEDHEKEDPQAGTSQ 355
G+D+ + + D+E E EED ED +E+ + ++ E+E PQ G+ +
Sbjct: 362 -GEDEDEEEEGEEDDEEEEDEEDEEDDDEEEEEQEEEEEPPQRGSGE 407
>UniRef100_Q76MT4 ABTAP [Rattus rattus]
Length = 842
Score = 43.1 bits (100), Expect = 0.012
Identities = 40/169 (23%), Positives = 72/169 (41%), Gaps = 27/169 (15%)
Query: 193 KTAYEQAKADAETANKNLKAAEERCAKLTDDLAASDLLLQKTKSLKETINDKHTAVQAKC 252
K +++ K D+E + K+ + + + K D +DL ++ + K D T+ K
Sbjct: 142 KNSHKSPKIDSEVSPKDSEESLQSRKKKRD---TTDLSVEASPKGKLRTKDPSTSAMVK- 197
Query: 253 QKLEKKYERLNASILGRASLLFAQGFLAAKEQISVV-------EPGFDLSRIGWLKEIKD 305
++++ G + Q + AK+ + E S +G +E +
Sbjct: 198 ----------SSTVSGSKAKREKQAVIMAKDNAGRMLHEEAPEEDSDSASELGRDEE-SE 246
Query: 306 GQVVGDDDISLDLLPQFDDESEPEEDGEDGNEQHKNEDHEKEDPQAGTS 354
G++ DD S D DDE+E EE+ EDG E+ + E+ E E S
Sbjct: 247 GEITSDDRASAD-----DDENEDEEEEEDGEEEEEEEEEEDESDDESDS 290
>UniRef100_UPI000023EDFF UPI000023EDFF UniRef100 entry
Length = 1116
Score = 42.7 bits (99), Expect = 0.016
Identities = 72/316 (22%), Positives = 119/316 (36%), Gaps = 59/316 (18%)
Query: 58 AGVKPLHQSTLDPKGRPTEKKKGHDNVP--------PH-----QPDSGALINRPSTPFIQ 104
AG K L T GRP + G+ +P PH + D +P P ++
Sbjct: 167 AGSKVLRLRTFGKDGRP---QLGNSRLPTSFANNDHPHPKNGDEDDDSDSQKKPPVPTVE 223
Query: 105 AGPSSAIGGEALPPLLNLSDPRFNGLDFMNRTFDNRIHKDVSGQGPPNIASVAIHHA-LS 163
A P G A + SD T + R V+ P I + LS
Sbjct: 224 AAPVLD-GNNA-----STSDVTATVPSQRRNTVNRRHSPSVASMDQPQIKLPLVDGPELS 277
Query: 164 AASIVAGMAQCVKELIATKNRYEKKAADYKTAYEQAKADAETANKNLKAAEERCAKLTDD 223
+ KE+ T +Y K+ A+ + E+ K + ET + LK EE+ A+L +
Sbjct: 278 LDELNRRFESTRKEVDETLAQYAKEEAELQQQEEELKQEKETKRQALKEKEEQTAQLKAN 337
Query: 224 L--------AASDLLLQKTKSLKETINDKHTAVQAKCQKLEKKYERLNASILGRASLLFA 275
+ AA +K + LK+ N K + V+ KLE ER+
Sbjct: 338 VRTTMEQMRAAEKDRAKKEQQLKDKEN-KKSKVRDTATKLENDIERMKKE---------R 387
Query: 276 QGFLAAKEQISVVEPGFDLSRIGWLKEIKDGQVVGDDDISL-----DLLPQFDDESEPEE 330
+GF A K++++ K +DGQ + D + L +L + ++ + +
Sbjct: 388 EGFEAQKKELAA-------------KRDRDGQDLDDSNAELQENCAELEAELKEKGKQLQ 434
Query: 331 DGEDGNEQHKNEDHEK 346
D + E+ D E+
Sbjct: 435 DLKATREELPGADDEQ 450
>UniRef100_Q91YS2 Rangap1 protein [Mus musculus]
Length = 589
Score = 42.4 bits (98), Expect = 0.020
Identities = 28/107 (26%), Positives = 53/107 (49%), Gaps = 14/107 (13%)
Query: 249 QAKCQKLEKKYERLNASILGRASLLFAQGFLAAKEQISVVEPGFDLSRIGWLKEIKDGQV 308
+A K E + LN + LG EQ+ V F+++++ L + D +
Sbjct: 315 EAVADKAELEKLDLNGNALGEEGC----------EQLQEVMDSFNMAKV--LASLSDDE- 361
Query: 309 VGDDDISLDLLPQFDDESEPEEDGEDGNEQHKNEDHEKEDPQAGTSQ 355
G+D+ + + D+E E EED ED +++ + ++ E+E PQ G+ +
Sbjct: 362 -GEDEDEEEEGEEDDEEEEDEEDEEDDDDEEEEQEEEEEPPQRGSGE 407
>UniRef100_Q9KI17 M-like protein Szp3 precursor [Streptococcus equi subsp.
zooepidemicus]
Length = 224
Score = 42.0 bits (97), Expect = 0.027
Identities = 34/131 (25%), Positives = 60/131 (44%), Gaps = 12/131 (9%)
Query: 163 SAASIVAGMAQCVKELIATKNRYEKKAADYKTAYEQAKADAETANKNLKAAEERCAKLTD 222
+A + +A + Q + +L EK+ A + EQ K A ++N A E+R +L+
Sbjct: 95 TAEATIASLEQKMNDL--ESKIQEKQKAFEQVVLEQKKEQANPVDRNGDAEEDRLDELSG 152
Query: 223 DL----AASDLLLQKTKSLKETINDKHTAVQAKCQKLEKKYERLNASILG------RASL 272
++ AA ++ LQKTK +T + ++ + Q K + N + G +A
Sbjct: 153 EIRREKAALEVELQKTKEALDTAKRAYAGIEERKQVAATKLDAANKAFAGVEEKHAQAMA 212
Query: 273 LFAQGFLAAKE 283
F + F A KE
Sbjct: 213 AFEEAFAAYKE 223
Score = 33.5 bits (75), Expect = 9.5
Identities = 35/121 (28%), Positives = 58/121 (47%), Gaps = 13/121 (10%)
Query: 153 IASVAIHHALSAASIVAGMAQCVKELIATKNRYEKKAADYKTAYEQAKADAETANKNLKA 212
+ASV+ +S+A + A A + + +AD + +A+A+ TA +L A
Sbjct: 17 LASVSAALLVSSAVVAADTADSATPAVTAT--VKDSSADKEAVALKAEAELATAKADLAA 74
Query: 213 AEE--RCAKLTDDLAASDLLLQKT--KSLKETINDKHTAVQAKCQK------LEKKYERL 262
AE AK + A +DL + SL++ +ND + +Q K QK LE+K E+
Sbjct: 75 AEAAITAAKAEYETAQADLATAEATIASLEQKMNDLESKIQEK-QKAFEQVVLEQKKEQA 133
Query: 263 N 263
N
Sbjct: 134 N 134
>UniRef100_Q9VKI0 CG6392-PA [Drosophila melanogaster]
Length = 2013
Score = 42.0 bits (97), Expect = 0.027
Identities = 26/101 (25%), Positives = 50/101 (48%), Gaps = 3/101 (2%)
Query: 177 ELIATKNRYEKKAADYKTAYE---QAKADAETANKNLKAAEERCAKLTDDLAASDLLLQK 233
E+++ +++ E + + A Q D E +K L ++ +L + A D Q
Sbjct: 704 EIVSVRDKLESVESAFNLASSEIIQKATDCERLSKELSTSQNAFGQLQERYDALDQQWQA 763
Query: 234 TKSLKETINDKHTAVQAKCQKLEKKYERLNASILGRASLLF 274
++ T+++KH VQ K QKL+++YE+L + +S F
Sbjct: 764 QQAGITTLHEKHEHVQEKYQKLQEEYEQLESRARSASSAEF 804
>UniRef100_Q7RPP8 Hypothetical protein [Plasmodium yoelii yoelii]
Length = 483
Score = 42.0 bits (97), Expect = 0.027
Identities = 36/127 (28%), Positives = 59/127 (46%), Gaps = 3/127 (2%)
Query: 163 SAASIVAGMAQCVKELIATKNRYEKKAADYKTAYEQAKADAETANKN-LKAAEERCAKLT 221
S + A + + ELI +N EK + + T + K +T NKN +K E KL
Sbjct: 78 SKDELYADLIKFETELIKEQNENEKYSLEILTLTNKNKILTDTINKNNIKIKENEIEKLK 137
Query: 222 DDLAASDLLLQKTKSLKETINDKHTAVQAKCQKLEKKYERLNASILGRASLLFAQGFLAA 281
+ L K K L T +++ K + +EK ++LN L + +LLF L+
Sbjct: 138 LKELKDEYL--KEKLLLTTKVEEYDKELKKYKIIEKYLDKLNKEDLNKFNLLFGLNILSL 195
Query: 282 KEQISVV 288
+EQ +V+
Sbjct: 196 EEQQNVI 202
>UniRef100_UPI00003640AE UPI00003640AE UniRef100 entry
Length = 1932
Score = 41.6 bits (96), Expect = 0.035
Identities = 29/143 (20%), Positives = 62/143 (43%), Gaps = 9/143 (6%)
Query: 152 NIASVAIHHALSAASI-------VAGMAQCVKELIATKNRYEKKAADYKTAYEQAKADAE 204
++ +HH + A++ VA +++ + L K + EK+ ++ K + + E
Sbjct: 1183 DLEEAMLHHEATTAALRKKHADSVAELSEQIDSLQRVKQKLEKERSEAKMEIDDLASTVE 1242
Query: 205 TANKNLKAAEERCAKLTDDLAASDLLLQKTKSLKETINDKHTAVQAKCQKLEKKYERLNA 264
+KN +AE+ C D + + + + + N + Q + +L +K E A
Sbjct: 1243 QLSKNKASAEKTCRLYEDQMNEAKAKVDELQRQLNDSNSQRARAQTESGELSRKLEEREA 1302
Query: 265 SI--LGRASLLFAQGFLAAKEQI 285
+ L R+ F+Q K+Q+
Sbjct: 1303 MVAQLQRSKNSFSQSVEELKKQL 1325
>UniRef100_UPI00003640AD UPI00003640AD UniRef100 entry
Length = 1930
Score = 41.6 bits (96), Expect = 0.035
Identities = 29/143 (20%), Positives = 62/143 (43%), Gaps = 9/143 (6%)
Query: 152 NIASVAIHHALSAASI-------VAGMAQCVKELIATKNRYEKKAADYKTAYEQAKADAE 204
++ +HH + A++ VA +++ + L K + EK+ ++ K + + E
Sbjct: 1173 DLEEAMLHHEATTAALRKKHADSVAELSEQIDSLQRVKQKLEKERSEAKMEIDDLASTVE 1232
Query: 205 TANKNLKAAEERCAKLTDDLAASDLLLQKTKSLKETINDKHTAVQAKCQKLEKKYERLNA 264
+KN +AE+ C D + + + + + N + Q + +L +K E A
Sbjct: 1233 QLSKNKASAEKTCRLYEDQMNEAKAKVDELQRQLNDSNSQRARAQTESGELSRKLEEREA 1292
Query: 265 SI--LGRASLLFAQGFLAAKEQI 285
+ L R+ F+Q K+Q+
Sbjct: 1293 MVAQLQRSKNSFSQSVEELKKQL 1315
>UniRef100_Q6LSE6 Hypothetical methyl-accepting chemotaxis protein [Photobacterium
profundum)]
Length = 646
Score = 41.6 bits (96), Expect = 0.035
Identities = 27/90 (30%), Positives = 45/90 (50%), Gaps = 4/90 (4%)
Query: 157 AIHHALSAA----SIVAGMAQCVKELIATKNRYEKKAADYKTAYEQAKADAETANKNLKA 212
A+ H+L A + + +A V EL+AT N+ +D E+ KADA+ + K +
Sbjct: 388 AVSHSLENAQQQQAESSSVATAVNELLATSNQIAANISDAALTAEKVKADAQESLKINQQ 447
Query: 213 AEERCAKLTDDLAASDLLLQKTKSLKETIN 242
AE L D++ S +L++K +IN
Sbjct: 448 AESSIKGLVSDISQSQVLIEKLAQESVSIN 477
>UniRef100_UPI000023DA4F UPI000023DA4F UniRef100 entry
Length = 1075
Score = 41.2 bits (95), Expect = 0.046
Identities = 41/178 (23%), Positives = 75/178 (42%), Gaps = 19/178 (10%)
Query: 186 EKKAADYKTAYEQAKADAETANKNLKAAEERCAKLTDDLAASDLLLQKTKSLKETINDKH 245
+KK A+ + +Q+K D E AEE+C L L+ D ++ ++ + +
Sbjct: 891 KKKDAEVEALQQQSKDDLEQQKSMTLKAEEKCQHLEKQLSGGDAGMKDMQARLARMESEL 950
Query: 246 TAVQAKCQKLEKKYERLNASILGRASLLFAQGFLAAKEQISVVEPGFDLSR--------- 296
+A EK +L + A A+ ++Q+S + ++++
Sbjct: 951 SAANKARDAAEKNLSKLKEAAKEEA----AKAKSEVEKQLSELREAREMTQSELDDLLMV 1006
Query: 297 IGWLKE----IKDGQVVGDDDISLDLLPQFDDESEPEEDGE--DGNEQHKNEDHEKED 348
G L+E KD + + +S DD+ E EE+G DG+E N D EK++
Sbjct: 1007 FGDLEEKMTKYKDRLIELGETVSDGEDDDEDDDDEEEENGSDADGSEGSDNNDDEKDE 1064
>UniRef100_UPI0000219DB0 UPI0000219DB0 UniRef100 entry
Length = 709
Score = 41.2 bits (95), Expect = 0.046
Identities = 25/82 (30%), Positives = 43/82 (51%), Gaps = 4/82 (4%)
Query: 188 KAADYKTAYEQAKADAETANKNLKAAEERCAKLTDDLAA----SDLLLQKTKSLKETIND 243
K A + E A ETA + + +KL +DLAA SD L Q+ ++ K++++D
Sbjct: 256 KIAKLEEDLEAANKSTETAQAEAATLKTKISKLEEDLAAAKSQSDKLTQEAEAQKKSLDD 315
Query: 244 KHTAVQAKCQKLEKKYERLNAS 265
+ +QAK ++ E +L A+
Sbjct: 316 ANAQIQAKTKETEDLLAKLKAA 337
Score = 39.3 bits (90), Expect = 0.17
Identities = 64/266 (24%), Positives = 101/266 (37%), Gaps = 45/266 (16%)
Query: 4 SKVDANPI----KMKEYLAQSAAAAKKRA-AETEQKKKNEGTSGSDNVRDPKRQKTSSAA 58
SK DAN K AQ A A+K+A AET+ + E T+ +D + + +K+++
Sbjct: 473 SKADANSALETQKSAVADAQKAVEAEKKAHAETKSALEAEKTAAADAKKAVESEKSTAQE 532
Query: 59 GVKPLHQSTLDPKGRPTEKKKGHDNVPPHQPDSGALINRPSTPFIQAGPSSAIGGEALPP 118
K L K E KK + ++ + +A +S AL
Sbjct: 533 AQKALEAE----KAAHAETKKSSEAGTSAVTEAQKALESEKAAHAEAKKASEADKAALAD 588
Query: 119 LLNLSDPRFNGLDFMNRTFDNRIHKDVSGQGPPNIASVAIHHALSAASIVAGMAQCVKEL 178
D A+ AL A A A K+
Sbjct: 589 AKKALDAE-------------------------KAAAADAQKALEAEKAAAADA---KKA 620
Query: 179 IATKNRYEKKAADYKTAY-EQAKADAETANKNLKAAEERCAKLTDDLAASDLLLQKTKSL 237
+ + KKA + +TA ++AK AE A LK AEE D A + + K
Sbjct: 621 LEAEKASAKKALETETAATKEAKKAAEDAQSKLKKAEE------DTKALNKRVADAEKES 674
Query: 238 KETINDKHTAVQAKCQKLEKKYERLN 263
KE ++ K TA +++ ++LE K + L+
Sbjct: 675 KE-LSTKATAAESRTKELEAKVKELS 699
>UniRef100_Q71EU9 Szp protein [Streptococcus equi subsp. zooepidemicus]
Length = 379
Score = 41.2 bits (95), Expect = 0.046
Identities = 46/187 (24%), Positives = 78/187 (41%), Gaps = 19/187 (10%)
Query: 163 SAASIVAGMAQCVKELIATKNRYEKKAADYKTAYEQAKADAETANKNLKAAEERCAKLTD 222
+A + +AG+ Q + EL K EK+ + EQ A N+ A ++R +L+
Sbjct: 93 TAEATIAGLEQKMAEL--EKKIQEKEKEFEQVVLEQKMEQANPVNRGGDAEKDRLDELSG 150
Query: 223 DL----AASDLLLQKTKSLKETINDKHTAVQAKCQKLEKKYERLNASILG------RASL 272
++ AA LQKTK E + ++ + Q K + N + G +A
Sbjct: 151 EIRRAKAALKAELQKTKESLEVAKKAYAGIEERKQVAAVKLDAANKAFAGVEEKHDQAMA 210
Query: 273 LFAQGFLAAKEQISVVEPGFDLSRIGWLKEIKDGQVVGD----DDISLDLL--PQFDDES 326
FA+ F A KE + S + K+I V ++ LD + P+ + E+
Sbjct: 211 KFAEAFAAYKEAVKAELKAAGASDF-YTKKIDSANTVAGVKTLREMILDSIAKPEVEPEA 269
Query: 327 EPEEDGE 333
+PE E
Sbjct: 270 KPEPKPE 276
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.308 0.127 0.350
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 629,878,895
Number of Sequences: 2790947
Number of extensions: 28031215
Number of successful extensions: 174123
Number of sequences better than 10.0: 1102
Number of HSP's better than 10.0 without gapping: 417
Number of HSP's successfully gapped in prelim test: 719
Number of HSP's that attempted gapping in prelim test: 166455
Number of HSP's gapped (non-prelim): 5183
length of query: 365
length of database: 848,049,833
effective HSP length: 129
effective length of query: 236
effective length of database: 488,017,670
effective search space: 115172170120
effective search space used: 115172170120
T: 11
A: 40
X1: 16 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.6 bits)
S2: 75 (33.5 bits)
Lotus: description of TM0219.2


