

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0214.6
(398 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
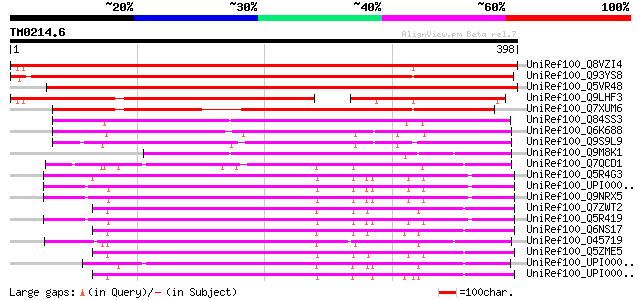
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q8VZI4 AT3g24470/MXP5_4 [Arabidopsis thaliana] 620 e-176
UniRef100_Q93YS8 Hypothetical protein At4g13345 [Arabidopsis tha... 615 e-175
UniRef100_Q5VR48 Hypothetical protein OJ1174_D05.1 [Oryza sativa] 515 e-145
UniRef100_Q9LHF3 Arabidopsis thaliana genomic DNA, chromosome 3,... 334 3e-90
UniRef100_Q7XUM6 OSJNBb0002J11.6 protein [Oryza sativa] 330 6e-89
UniRef100_Q84SS3 Putative membrane protein [Oryza sativa] 207 3e-52
UniRef100_Q6K688 Putative tumor differentially expressed protein... 206 9e-52
UniRef100_Q9S9L9 T24D18.26 protein [Arabidopsis thaliana] 194 5e-48
UniRef100_Q9M8K1 F28L1.11 protein [Arabidopsis thaliana] 176 8e-43
UniRef100_Q7QCD1 ENSANGP00000002975 [Anopheles gambiae str. PEST] 144 6e-33
UniRef100_Q5R4G3 Hypothetical protein DKFZp459I033 [Pongo pygmaeus] 141 4e-32
UniRef100_UPI000036D955 UPI000036D955 UniRef100 entry 140 5e-32
UniRef100_Q9NRX5 Tumor differentially expressed protein 2 [Homo ... 140 5e-32
UniRef100_Q7ZWT2 TDE2 protein [Xenopus laevis] 140 6e-32
UniRef100_Q5R419 Hypothetical protein DKFZp459J102 [Pongo pygmaeus] 140 6e-32
UniRef100_Q6NS17 Hypothetical protein [Xenopus laevis] 139 1e-31
UniRef100_O45719 Hypothetical protein R11H6.2 [Caenorhabditis el... 138 2e-31
UniRef100_Q5ZME5 Hypothetical protein [Gallus gallus] 138 3e-31
UniRef100_UPI00001A07CE UPI00001A07CE UniRef100 entry 137 5e-31
UniRef100_UPI00003614DC UPI00003614DC UniRef100 entry 133 8e-30
>UniRef100_Q8VZI4 AT3g24470/MXP5_4 [Arabidopsis thaliana]
Length = 409
Score = 620 bits (1599), Expect = e-176
Identities = 287/407 (70%), Positives = 344/407 (84%), Gaps = 9/407 (2%)
Query: 1 METG--VSNNG----NEACTISKDTSWWGQFRNASNPGMARYVYALMFLVANLLAWAARD 54
METG VSNN N++ K+ SW+ QFRN NP MARYVY L+FL+ANLLAWAARD
Sbjct: 1 METGTSVSNNNQSIRNDSYEAIKNGSWFNQFRNGCNPWMARYVYGLIFLIANLLAWAARD 60
Query: 55 YGRGALTEMERLKGCNGGKDCLGAEGVLRVSLGCFIFYFIMFLTTARTSKLNEVRDTWHS 114
YGRGAL ++ R K C GG++CLG +GVLRVSLGCF+FYF+MFL+T TSK + RD WHS
Sbjct: 61 YGRGALRKVTRFKNCKGGENCLGTDGVLRVSLGCFLFYFVMFLSTLGTSKTHSSRDRWHS 120
Query: 115 GWWSVKIVLWVGMTVIPFLLPSEFIQIYGEVAHFGAGVFLLIQLISIISFITWLNDCCES 174
GWW VK+++W +T+IPFLLPS I +YGE+AHFGAGVFLLIQLIS+ISFI WLN+C +S
Sbjct: 121 GWWFVKLIMWPALTIIPFLLPSSIIHLYGEIAHFGAGVFLLIQLISVISFIQWLNECYQS 180
Query: 175 EKYAAKCQIHVMLFATTAYVVCLVGIILMFIWYAPKPSCLLNIFFIAWTLVLLQLMTSVS 234
+K A +C+++VML +TT+Y VC+VG+ILM+IWYAP SCLLNIFFI WTL L+QLMTS++
Sbjct: 181 QKDAERCRVYVMLLSTTSYTVCIVGVILMYIWYAPDSSCLLNIFFITWTLFLIQLMTSIA 240
Query: 235 LHPKVNAGILTPGLMGLYVVFLCWNAIRSEPDGGSCIRKSDSSTKTDWLSIISFIIAILA 294
LHPKVNAG LTP LMGLYVVF+CW AIRSEP G SC RK+ +S +TDWL+IISF++A+LA
Sbjct: 241 LHPKVNAGYLTPALMGLYVVFICWCAIRSEPVGESCNRKAAASNRTDWLTIISFVVALLA 300
Query: 295 IVIATFSTGIDSKCFQFRKDD---DIPAEDDVPYGYGFFHFVFATGAMYFAMLLIGWNSH 351
+VIATFSTGIDS+CFQF+KD+ + AEDDVPYGYGFFHFVFATGAMYFAMLLIGWN+H
Sbjct: 301 MVIATFSTGIDSQCFQFKKDENDQEEEAEDDVPYGYGFFHFVFATGAMYFAMLLIGWNTH 360
Query: 352 HSMRKWTIDVGWTSTWVRIVNEWLAVCVYLWMLVAPIIWKSRQTDST 398
H M+KWTIDVGWTSTWVR+VNEWLAVCVY+WMLVAP+I KSR+ T
Sbjct: 361 HPMKKWTIDVGWTSTWVRVVNEWLAVCVYIWMLVAPLILKSRRQTPT 407
>UniRef100_Q93YS8 Hypothetical protein At4g13345 [Arabidopsis thaliana]
Length = 394
Score = 615 bits (1586), Expect = e-175
Identities = 286/397 (72%), Positives = 335/397 (84%), Gaps = 6/397 (1%)
Query: 1 METGVS--NNGNEACTISKDTSWWGQFRNASNPGMARYVYALMFLVANLLAWAARDYGRG 58
METG S N G E K+ SW+ QFRN NP MARYVY L+FL+ANLLAWA RDYGRG
Sbjct: 1 METGTSIDNTGYEGI---KNGSWFIQFRNGCNPWMARYVYGLIFLLANLLAWALRDYGRG 57
Query: 59 ALTEMERLKGCNGGKDCLGAEGVLRVSLGCFIFYFIMFLTTARTSKLNEVRDTWHSGWWS 118
ALTEM + K C G DCLG EGVLRVS GCF+FYFIMFL+T TSK + RD WHSGWW
Sbjct: 58 ALTEMRKFKNCKEGGDCLGTEGVLRVSFGCFLFYFIMFLSTVGTSKTHSSRDKWHSGWWF 117
Query: 119 VKIVLWVGMTVIPFLLPSEFIQIYGEVAHFGAGVFLLIQLISIISFITWLNDCCESEKYA 178
K+ + +G+T+ PFLLPS IQ YGE+AHFGAGVFLLIQLISIISFITWLN+C +++K A
Sbjct: 118 AKLFMLLGLTIFPFLLPSSIIQFYGEIAHFGAGVFLLIQLISIISFITWLNECFQAQKDA 177
Query: 179 AKCQIHVMLFATTAYVVCLVGIILMFIWYAPKPSCLLNIFFIAWTLVLLQLMTSVSLHPK 238
+C +HVML ATTAY VC++G+ILM+IWY P+PSCLLNIFFI WTL L+QLMTS+SLHPK
Sbjct: 178 ERCHVHVMLLATTAYTVCILGVILMYIWYVPEPSCLLNIFFITWTLFLIQLMTSISLHPK 237
Query: 239 VNAGILTPGLMGLYVVFLCWNAIRSEPDGGSCIRKSDSSTKTDWLSIISFIIAILAIVIA 298
+NAG LTP LMGLYVVF+CW AIRSEP G +C RK++ S++TDWL+IISF++A+LA+VIA
Sbjct: 238 INAGFLTPALMGLYVVFICWCAIRSEPVGETCNRKAEGSSRTDWLTIISFVVALLAMVIA 297
Query: 299 TFSTGIDSKCFQFRKDDDIPAEDDVPYGYGFFHFVFATGAMYFAMLLIGWNSHHSMRKWT 358
TFSTG+DS+CFQFRKD++ ED +PYGYGFFHFVFATGAMYFAMLL+GWN HHSM+KWT
Sbjct: 298 TFSTGVDSQCFQFRKDEN-HEEDAIPYGYGFFHFVFATGAMYFAMLLVGWNIHHSMKKWT 356
Query: 359 IDVGWTSTWVRIVNEWLAVCVYLWMLVAPIIWKSRQT 395
IDVGWTSTWVRIVNEWLAV VY+WMLVAP++ KSRQT
Sbjct: 357 IDVGWTSTWVRIVNEWLAVGVYIWMLVAPMVLKSRQT 393
>UniRef100_Q5VR48 Hypothetical protein OJ1174_D05.1 [Oryza sativa]
Length = 421
Score = 515 bits (1327), Expect = e-145
Identities = 229/371 (61%), Positives = 290/371 (77%), Gaps = 2/371 (0%)
Query: 30 NPGMARYVYALMFLVANLLAWAARDYGRGALTEMERLKG-CNGGKDCLGAEGVLRVSLGC 88
NP MARYVYA +FL NLLAW RD+G L E+ RL+G C G CLGAEGVLRVSLGC
Sbjct: 47 NPMMARYVYAFVFLATNLLAWTLRDFGHPVLAELRRLRGSCQGAGYCLGAEGVLRVSLGC 106
Query: 89 FIFYFIMFLTTARTSKLNEVRDTWHSGWWSVKIVLWVGMTVIPFLLPSEFIQIYGEVAHF 148
F+F+F+MFL+T RT K ++ R++WHS WW KIVLW+G TV+PF LPS IQ+YG++AHF
Sbjct: 107 FLFFFVMFLSTVRTRKTHDRRNSWHSEWWPAKIVLWMGFTVVPFFLPSPLIQLYGKIAHF 166
Query: 149 GAGVFLLIQLISIISFITWLNDCCESEKYAAKCQIHVMLFATTAYVVCLVGIILMFIWYA 208
GAG FL+IQL+S+ FITWLNDCC SE +C + V + + AYV ++G++LM++WYA
Sbjct: 167 GAGAFLVIQLVSVTRFITWLNDCCRSETNLKRCHMQVQVVSIAAYVGSILGVVLMYVWYA 226
Query: 209 PKPSCLLNIFFIAWTLVLLQLMTSVSLHPKVNAGILTPGLMGLYVVFLCWNAIRSEPDGG 268
P+PSC LNI FI TLVL+Q+MT VSL KV AG L PGLMG+Y+VFLCW AIRSEP
Sbjct: 227 PRPSCKLNILFITVTLVLVQIMTGVSLSSKVKAGYLAPGLMGVYIVFLCWTAIRSEPHTE 286
Query: 269 SCIRKSDSSTKTDWLSIISFIIAILAIVIATFSTGIDSKCFQFRK-DDDIPAEDDVPYGY 327
C +K++ +T DW++I SF+IA++ IV ATF+TGIDSKC QF+K + + P +DD+PYG+
Sbjct: 287 ICNKKAEVATSADWVNIASFVIAVIVIVTATFATGIDSKCLQFKKAESEQPEDDDIPYGF 346
Query: 328 GFFHFVFATGAMYFAMLLIGWNSHHSMRKWTIDVGWTSTWVRIVNEWLAVCVYLWMLVAP 387
GFFHFVFA GAMYFAML +GWN++ +M KWTIDVGW STWVR+VNEWLA VY+WM++AP
Sbjct: 347 GFFHFVFAMGAMYFAMLFVGWNANQTMEKWTIDVGWASTWVRVVNEWLAAIVYIWMVIAP 406
Query: 388 IIWKSRQTDST 398
I+WK Q S+
Sbjct: 407 IVWKGGQVGSS 417
>UniRef100_Q9LHF3 Arabidopsis thaliana genomic DNA, chromosome 3, P1 clone: MXP5
[Arabidopsis thaliana]
Length = 528
Score = 334 bits (856), Expect = 3e-90
Identities = 156/245 (63%), Positives = 194/245 (78%), Gaps = 12/245 (4%)
Query: 1 METG--VSNNG----NEACTISKDTSWWGQFRNASNPGMARYVYALMFLVANLLAWAARD 54
METG VSNN N++ K+ SW+ QFRN NP MARYVY L+FL+ANLLAWAARD
Sbjct: 1 METGTSVSNNNQSIRNDSYEAIKNGSWFNQFRNGCNPWMARYVYGLIFLIANLLAWAARD 60
Query: 55 YGRGALTEMERLKGCNGGKDCLGAEGVLRVSLGCFIFYFIMFLTTARTSKLNEVRDTWHS 114
YGRGAL ++ R K C GG++CLG +GVLR +FYF+MFL+T TSK + RD WHS
Sbjct: 61 YGRGALRKVTRFKNCKGGENCLGTDGVLR------LFYFVMFLSTLGTSKTHSSRDRWHS 114
Query: 115 GWWSVKIVLWVGMTVIPFLLPSEFIQIYGEVAHFGAGVFLLIQLISIISFITWLNDCCES 174
GWW VK+++W +T+IPFLLPS I +YGE+AHFGAGVFLLIQLIS+ISFI WLN+C +S
Sbjct: 115 GWWFVKLIMWPALTIIPFLLPSSIIHLYGEIAHFGAGVFLLIQLISVISFIQWLNECYQS 174
Query: 175 EKYAAKCQIHVMLFATTAYVVCLVGIILMFIWYAPKPSCLLNIFFIAWTLVLLQLMTSVS 234
+K A +C+++VML +TT+Y VC+VG+ILM+IWYAP SCLLNIFFI WTL L+QLMTS++
Sbjct: 175 QKDAERCRVYVMLLSTTSYTVCIVGVILMYIWYAPDSSCLLNIFFITWTLFLIQLMTSIA 234
Query: 235 LHPKV 239
LHPKV
Sbjct: 235 LHPKV 239
Score = 181 bits (459), Expect = 3e-44
Identities = 89/133 (66%), Positives = 105/133 (78%), Gaps = 11/133 (8%)
Query: 268 GSCIRKSDSSTKTDWLSII--SFIIAILAIVIATFSTGIDSKCFQFRKDD---DIPAEDD 322
G RK D+ + L+ I SF++A+LA+VIATFSTGIDS+CFQF+KD+ + AEDD
Sbjct: 393 GYLTRKKDNVSNMLILNNIVQSFVVALLAMVIATFSTGIDSQCFQFKKDENDQEEEAEDD 452
Query: 323 VPYGYGFFHFVFATGAMYFAMLLIGWNSHHSMRKWTIDVGWTSTWVRIVNEWLAVCVYL- 381
VPYGYGFFHFVFATGAMYFAMLLIGWN+HH M+KWTIDVGWTSTWVR+VNEWLAVCVY
Sbjct: 453 VPYGYGFFHFVFATGAMYFAMLLIGWNTHHPMKKWTIDVGWTSTWVRVVNEWLAVCVYKQ 512
Query: 382 -----WMLVAPII 389
W L+ I+
Sbjct: 513 KTNANWDLIGKIV 525
>UniRef100_Q7XUM6 OSJNBb0002J11.6 protein [Oryza sativa]
Length = 477
Score = 330 bits (845), Expect = 6e-89
Identities = 162/348 (46%), Positives = 214/348 (60%), Gaps = 38/348 (10%)
Query: 34 ARYVYALMFLVANLLAWAARDYGRGALTEMERLKGCNGGKD-CLGAEGVLRVSLGCFIFY 92
ARY Y L+F NLLAW RDYG L + + C G C + GVLR IF+
Sbjct: 27 ARYAYGLIFFATNLLAWFVRDYGAKLLRGLHHVPVCGAGDSKCFQSGGVLR------IFF 80
Query: 93 FIMFLTTARTSKLNEVRDTWHSGWWSVKIVLWVGMTVIPFLLPSEFIQIYGEVAHFGAGV 152
++MF TT T KL+EVR++WHSG W +K +++ +IPF++P+ FIQ+YGE+A GAG
Sbjct: 81 WVMFATTFGTRKLHEVRNSWHSGCWILKFLVYAVSIIIPFIVPNIFIQLYGEIARMGAG- 139
Query: 153 FLLIQLISIISFITWLNDCCESEKYAAKCQIHVMLFATTAYVVCLVGIILMFIWYAPKPS 212
+ + +T +++ GI ++++ Y P S
Sbjct: 140 -----------------------------GLFGLFLSTISFIASFAGIAVLYVLYVPNSS 170
Query: 213 CLLNIFFIAWTLVLLQLMTSVSLHPKVNAGILTPGLMGLYVVFLCWNAIRSEPDGGSCIR 272
C NIF I WT L+ +M +VSLH KVN G+L+ G+MGLY+VFLCW+A+ SEP G C
Sbjct: 171 CAFNIFTITWTATLVAVMMAVSLHSKVNEGLLSSGIMGLYIVFLCWSALHSEPQTGKCHT 230
Query: 273 KSDSSTKTDWLSIISFIIAILAIVIATFSTGIDSKCFQFRKDDDIPAEDDVPYGYGFFHF 332
+ + DW +I+SFIIAI AIV+ATFSTGID++ FQFR D+D EDDVPY Y FH
Sbjct: 231 RLIFANDGDWATIVSFIIAICAIVMATFSTGIDTRSFQFRNDED-QLEDDVPYSYEIFHI 289
Query: 333 VFATGAMYFAMLLIGWNSHHSMRKWTIDVGWTSTWVRIVNEWLAVCVY 380
VFA GAMYFAML I W +H RKW+IDVGW STWV+I+NEW A +Y
Sbjct: 290 VFAMGAMYFAMLFINWELNHPTRKWSIDVGWVSTWVKIINEWFAASIY 337
>UniRef100_Q84SS3 Putative membrane protein [Oryza sativa]
Length = 417
Score = 207 bits (528), Expect = 3e-52
Identities = 120/385 (31%), Positives = 192/385 (49%), Gaps = 26/385 (6%)
Query: 34 ARYVYALMFLVANLLAWAARDYGRGALTEMERLKGCNGGK--DCLGAEGVLRVSLGCFIF 91
AR Y +F + +L++ R + L ++ + + + + VLRVSLG F+F
Sbjct: 31 ARLAYCGLFAASLILSFLMRQFATPLLKQIPWINTFDYTQPDEWFQMNAVLRVSLGNFLF 90
Query: 92 YFIMFLTTARTSKLNEVRDTWHSGWWSVKIVLWVGMTVIPFLLPSEFIQIYGEVAHFGAG 151
+ I L N+ RD WH G W KIV+WV + V+ F +P+ I IY ++ FG+G
Sbjct: 91 FAIFALMMIGVKDQNDRRDAWHHGGWIAKIVVWVVLIVLMFCVPNVVITIYEVLSKFGSG 150
Query: 152 VFLLIQLISIISFITWLNDCCESEKYAAKCQIHVMLFATTAYVVCLVGIILMFIWYAPKP 211
+FLL+Q++ ++ F ND EK K +I +++ Y+ L+F W+ P
Sbjct: 151 LFLLVQVVMLLDFTNNWNDSW-IEKDEQKWEIALLVVTVVCYLSTFAFSGLLFTWFNPSG 209
Query: 212 -SCLLNIFFIAWTLVLLQLMTSVSLHPKVNAGILTPGLMGLYVVFLCWNAIRSEPDGGSC 270
C LN+FFI T++L ++LHP+VN ++ ++ +Y +LC+ ++ SEPD +C
Sbjct: 210 HDCGLNVFFITMTIILAFAFAIIALHPQVNGSVMPASVISVYCAYLCYTSLSSEPDDYAC 269
Query: 271 IRKSDSSTKTDWLSII-SFIIAILAIVIATFSTGIDSKCFQ------------FRKDDDI 317
S + ++I + +L++V + G + DD++
Sbjct: 270 NGLHRHSKQVSMSALILGMLTTVLSVVYSAVRAGSSTTFLSPPSSPRSGIKNPLLGDDNV 329
Query: 318 PAEDD---------VPYGYGFFHFVFATGAMYFAMLLIGWNSHHSMRKWTIDVGWTSTWV 368
A V Y Y FFH +FA +MY AMLL GW S S +DVGWT+ WV
Sbjct: 330 EAGKSNSKEIDARPVSYSYTFFHVIFALASMYSAMLLTGWTSAASDSSELMDVGWTTVWV 389
Query: 369 RIVNEWLAVCVYLWMLVAPIIWKSR 393
RI EW +Y+W LVAP+++ R
Sbjct: 390 RICTEWATAALYIWTLVAPLLFPDR 414
>UniRef100_Q6K688 Putative tumor differentially expressed protein 1 [Oryza sativa]
Length = 414
Score = 206 bits (524), Expect = 9e-52
Identities = 118/389 (30%), Positives = 194/389 (49%), Gaps = 34/389 (8%)
Query: 34 ARYVYALMFLVANLLAWAARDYGRGALTEMERLKGCNGGKD--CLGAEGVLRVSLGCFIF 91
AR Y +F ++ +WA R+ L + + + D + VLRVSLG F+F
Sbjct: 31 ARIAYCGLFALSLFASWALREVAAPLLQSIPWINHFHKTPDREWFETDAVLRVSLGNFVF 90
Query: 92 YFIMFLTTARTSKLNEVRDTWHSGWWSVKIVLWVGMTVIPFLLPSEFIQIYGEVAHFGAG 151
+ I+ + A + RD H G W KI WV + + F +P+ + Y ++ FG+G
Sbjct: 91 FTILAIIMAGIKDQKDPRDKIHHGGWMAKIFCWVVIVFLMFFVPNGVVSFYESISKFGSG 150
Query: 152 VFLLIQLISIISFITWLNDCCESEKYAAKCQ----IHVMLFATTAYVVCLVGIILMFIWY 207
+FLL+Q++ ++ F+ N E + AK + + +++ + Y+V L+F W+
Sbjct: 151 LFLLVQVVLLLDFVHGWN-----ENWVAKDEQFWYMALLVVSVVCYIVTFSFSGLLFHWF 205
Query: 208 APKP-SCLLNIFFIAWTLVLLQLMTSVSLHPKVNAGILTPGLMGLYVVFLCWNAIRSEPD 266
P C +N+FFI +TL+L+ + V+LHPK+N +L ++ LY +LC++ + SEP
Sbjct: 206 TPSGHDCGINLFFIVFTLILVFVFAIVALHPKINGSLLPASVIALYCTYLCYSGLSSEPR 265
Query: 267 GGSC--IRKSDSSTKTDWLSIISFIIAILAIVIATFSTG---------------IDSKCF 309
C + + T LS+ + IL++V + G D
Sbjct: 266 DYECNGLHNHSKAVSTGSLSL-GLLTTILSVVYSAVRAGSSATVLSAPDSPRAGADKPLL 324
Query: 310 QFRKDDDIPAEDDVP----YGYGFFHFVFATGAMYFAMLLIGWNSHHSMRKWTIDVGWTS 365
F K D+ + DVP Y Y FFH +F+ +MY AMLL GW++ +DVGW S
Sbjct: 325 PFSKADEEAEKKDVPRPVTYSYSFFHLIFSLASMYSAMLLTGWSTSVGESGKLVDVGWPS 384
Query: 366 TWVRIVNEWLAVCVYLWMLVAPIIWKSRQ 394
WVRI +W +Y+W LVAP+++ R+
Sbjct: 385 VWVRIATQWATAGLYIWSLVAPLLFPDRE 413
>UniRef100_Q9S9L9 T24D18.26 protein [Arabidopsis thaliana]
Length = 412
Score = 194 bits (492), Expect = 5e-48
Identities = 118/393 (30%), Positives = 191/393 (48%), Gaps = 44/393 (11%)
Query: 34 ARYVYALMFLVANLLAWAARDYGRGALTEMERLKGCNG-----GKDCLGAEGVLRVSLGC 88
AR Y +F ++ +++W R+ A ME+L N ++ + VLRVSLG
Sbjct: 31 ARIAYCGLFALSLIVSWILREV---AAPLMEKLPWINHFHKTPDREWFETDAVLRVSLGN 87
Query: 89 FIFYFIMFLTTARTSKLNEVRDTWHSGWWSVKIVLWVGMTVIPFLLPSEFIQIYGEVAHF 148
F+F+ I+ + + RD H G W +KI+ W + + F LP+E I Y ++ F
Sbjct: 88 FLFFSILSVMMIGVKNQKDPRDGIHHGGWMMKIICWCILVIFMFFLPNEIISFYESMSKF 147
Query: 149 GAGVFLLIQLISIISFITWLNDCC----ESEKYAAKCQIHVMLFATTAYVVCLVGIILMF 204
GAG FLL+Q++ ++ F+ ND E YAA +++ + Y+ V +F
Sbjct: 148 GAGFFLLVQVVLLLDFVHGWNDTWVGYDEQFWYAA-----LLVVSLVCYLATFVFSGFLF 202
Query: 205 IWYAPKP-SCLLNIFFIAWTLVLLQLMTSVSLHPKVNAGILTPGLMGLYVVFLCWNAIRS 263
W+ P C LN FFI TL+ + + V LHP V IL ++ LY ++LC++ + S
Sbjct: 203 HWFTPSGHDCGLNTFFIIMTLIFVFVFAIVVLHPTVGGSILPASVISLYCMYLCYSGLAS 262
Query: 264 EPDGGSC--IRKSDSSTKTDWLSIISFIIAILAIVIATFSTG------------------ 303
EP C + + T ++I + +L++V + G
Sbjct: 263 EPRDYECNGLHNHSKAVSTGTMTI-GLLTTVLSVVYSAVRAGSSTTLLSPPDSPRAEKPL 321
Query: 304 --IDSKCFQFRKDDDIPAEDDVPYGYGFFHFVFATGAMYFAMLLIGWNSHHSMRKWTIDV 361
ID K + + ++ + V Y Y FFH +F+ +MY AMLL GW++ +DV
Sbjct: 322 LPIDGKAEEKEEKEN---KKPVSYSYAFFHIIFSLASMYSAMLLTGWSTSVGESGKLVDV 378
Query: 362 GWTSTWVRIVNEWLAVCVYLWMLVAPIIWKSRQ 394
GW S WVR+V W +++W LVAPI++ R+
Sbjct: 379 GWPSVWVRVVTSWATAGLFIWSLVAPILFPDRE 411
>UniRef100_Q9M8K1 F28L1.11 protein [Arabidopsis thaliana]
Length = 315
Score = 176 bits (447), Expect = 8e-43
Identities = 93/309 (30%), Positives = 155/309 (50%), Gaps = 23/309 (7%)
Query: 106 NEVRDTWHSGWWSVKIVLWVGMTVIPFLLPSEFIQIYGEVAHFGAGVFLLIQLISIISFI 165
N+ RD+WH G W +K+++W + V+ F +P+ + +YG ++ FGAG FLL+Q++ ++
Sbjct: 8 NDRRDSWHHGGWGLKMIVWFLLVVLMFFVPNVIVSLYGTLSKFGAGAFLLVQVVLLLDAT 67
Query: 166 TWLNDCCESEKYAAKCQIHVMLFATTAYVVCLVGIILMFIWYAPK-PSCLLNIFFIAWTL 224
ND EK K I +++ + Y+ ++FIW+ P C LN+FFI +
Sbjct: 68 HNWNDSW-VEKDEKKWYIALLVISIVCYIATYTFSGILFIWFNPSGQDCGLNVFFIVMPM 126
Query: 225 VLLQLMTSVSLHPKVNAGILTPGLMGLYVVFLCWNAIRSEPDGGSCIRKSDSSTKTDWLS 284
+L + ++LHP VN +L ++ +Y ++C+ + SEP C + S
Sbjct: 127 ILAFVFAIIALHPAVNGSLLPASVISVYCAYVCYTGLSSEPHDYVCNGLNKSKAVNASTL 186
Query: 285 IISFIIAILAIVIATFSTGIDSKCF-------------------QFRKDDDIPAEDDVPY 325
I+ + +L+++ + G + +K + A V Y
Sbjct: 187 ILGMLTTVLSVLYSALRAGSSTTFLSPPSSPRSGVKDALLGDPEDGKKSGEAEAR-PVSY 245
Query: 326 GYGFFHFVFATGAMYFAMLLIGWNSHHSMRKWTIDVGWTSTWVRIVNEWLAVCVYLWMLV 385
Y FFH +FA +MY AMLL GW + S IDVGWTS WV+I W+ +Y+W L+
Sbjct: 246 SYSFFHIIFALASMYAAMLLSGW-TDSSESATLIDVGWTSVWVKICTGWVTAGLYIWTLI 304
Query: 386 APIIWKSRQ 394
AP+I R+
Sbjct: 305 APLILPDRE 313
>UniRef100_Q7QCD1 ENSANGP00000002975 [Anopheles gambiae str. PEST]
Length = 435
Score = 144 bits (362), Expect = 6e-33
Identities = 110/423 (26%), Positives = 192/423 (45%), Gaps = 67/423 (15%)
Query: 29 SNPGMARYVYALMFLVANLLAWAARDYGRGALTEMERLKGCN-----------GGK--DC 75
SN R++YALM ++ ++ A G ++++ C GG+ DC
Sbjct: 22 SNSTSTRFMYALMLVLGAIVG--AIMLAPGLQDFLQKVPFCANSTSTASNFVPGGETIDC 79
Query: 76 LGAEGVLRV-----SLGCFIFYFIMFLTTARTSKLNEVRDTWHSGWWSVKIVLWVGMTVI 130
A G L V +L CF + + + R+SK + R +G+W +K ++ + + +
Sbjct: 80 SSAVGYLAVYRICFALVCFFALWALMMIGVRSSK--DPRAALQNGFWGIKFMIVICIAIG 137
Query: 131 PFLLPSE-FIQIYGEVAHFGAGVFLLIQLISIISFI-----TWLNDCCESEK---YAAKC 181
F +P F + V G F+L+QL+ II F W+++ E E +AA C
Sbjct: 138 AFFIPETGFGPAWMWVGLIGGFAFILVQLVYIIDFAHNWAEAWVSNYEEDESRGWFAALC 197
Query: 182 QIHVMLFATTAYVVCLVGIILMFIWYAPKPSCLLNIFFIAWTLVLLQLMTSVSLHPKV-- 239
YV+ L GI+L+++++ C LN FFI + ++L ++ +S+ P+V
Sbjct: 198 -----CATGVQYVLSLTGIVLLYVYFTQADDCSLNKFFITFNMLLCIAVSFLSILPRVQE 252
Query: 240 ---NAGILTPGLMGLYVVFLCWNAIRSEPDGG------SCIRKSDSSTKTDWLSIISFII 290
+G+L ++ LY V+L W+A+ + PD I + + D SI+ II
Sbjct: 253 YQPRSGLLQSAMVTLYTVYLTWSAVANNPDAECNPGFLGIIGEKSNKVHFDKTSIVGLII 312
Query: 291 AILAIVIATFSTGIDSKCFQFRKDDDIPAEDD-------------------VPYGYGFFH 331
+L I+ ++ + + ++ D+ A DD V Y + FH
Sbjct: 313 WLLCILYSSLRSASNVSRLPDLENQDLTASDDGSNAGGRHGNEVRDNEESAVAYNWSLFH 372
Query: 332 FVFATGAMYFAMLLIGWNSHHSMRKWTIDVGWTSTWVRIVNEWLAVCVYLWMLVAPIIWK 391
VF T +Y M L W +S T++ S WV++V+ W+ V +Y W LVAP++
Sbjct: 373 IVFITATLYVMMTLTNWYQPNSSLD-TLNANAASMWVKVVSSWMCVALYGWTLVAPMVLT 431
Query: 392 SRQ 394
R+
Sbjct: 432 DRE 434
>UniRef100_Q5R4G3 Hypothetical protein DKFZp459I033 [Pongo pygmaeus]
Length = 453
Score = 141 bits (355), Expect = 4e-32
Identities = 109/425 (25%), Positives = 188/425 (43%), Gaps = 57/425 (13%)
Query: 27 NASNPGMARYVYALMFLVANLLAWAARDYGRGALTEM-----ERLKGCNGGKDCLGAEGV 81
+ +N + R +YAL LV +A G E KG +G + V
Sbjct: 31 SGNNSAVTRLIYALFLLVGVCVACVMLIPGMEEQLNKIPGFCENKKGVVPCNILVGYKAV 90
Query: 82 LRVSLGCFIFYFIMFLTTARTSKLNEVRDTWHSGWWSVKIVLWVGMTVIPFLLPS-EFIQ 140
R+ G +FY ++ L + ++ R H+G+W K + + + F +P F
Sbjct: 91 YRLCFGLAMFYLLLSLLMIKVKSSSDPRAAVHNGFWFFKFAAAIAIIIGAFFIPEGTFTT 150
Query: 141 IYGEVAHFGAGVFLLIQLISIISFI-TWLNDCCES-EKYAAKCQIHVMLFATTA-YVVCL 197
++ V GA F+LIQL+ +I F +W E E+ ++C +L AT Y++ L
Sbjct: 151 VWFYVGMAGAFCFILIQLVLLIDFAHSWNESWVEKMEEGNSRCWYAALLSATALNYLLSL 210
Query: 198 VGIILMFIWYAPKPSCLLNIFFIAWTLVLLQLMTSVSLHPKVN-----AGILTPGLMGLY 252
V I+L F++Y SC N FI+ ++L + +S+ PK+ +G+L ++ +Y
Sbjct: 211 VAIVLFFVYYTHPASCSENKAFISVNMLLCIGASVMSILPKIQESQPRSGLLQSSVITVY 270
Query: 253 VVFLCWNAIRSEPDGG------SCIRKSDSST------KTDWL---SIISFIIAILAIVI 297
++L W+A+ +EP+ S I + +ST W II I+ +L +
Sbjct: 271 TMYLTWSAMTNEPETNCNPSLLSIIGYNTTSTVPKEGQSVQWWHAQGIIGLILFLLCVFY 330
Query: 298 ATFSTGIDSKCFQF---------------RKDDDIPAEDDV-----------PYGYGFFH 331
++ T +S+ + R D + DDV Y Y FFH
Sbjct: 331 SSIRTSNNSQVNKLTLTSDESTLIEDGGARSDGSLEDGDDVHRAVDNERDGVTYSYSFFH 390
Query: 332 FVFATGAMYFAMLLIGWNSHHSMRKWTIDVGWTSTWVRIVNEWLAVCVYLWMLVAPIIWK 391
F+ ++Y M L W + R+ WT+ WV+I + W+ + +Y+W LVAP++
Sbjct: 391 FMLFLASLYIMMTLTNWYRYEPSREMKSQ--WTAVWVKISSSWIGIVLYVWTLVAPLVLT 448
Query: 392 SRQTD 396
+R D
Sbjct: 449 NRDFD 453
>UniRef100_UPI000036D955 UPI000036D955 UniRef100 entry
Length = 477
Score = 140 bits (354), Expect = 5e-32
Identities = 109/427 (25%), Positives = 192/427 (44%), Gaps = 61/427 (14%)
Query: 27 NASNPGMARYVYALMFLVANLLAWAARDYGRGALTEMERLKG-CNGGKDCL------GAE 79
+ +N + R +YAL LV +A G ++ ++ G C K + G +
Sbjct: 55 SGNNSTVTRLIYALFLLVGVCVACVMLIPGMEE--QLNKIPGFCENEKGVVPCNILVGYK 112
Query: 80 GVLRVSLGCFIFYFIMFLTTARTSKLNEVRDTWHSGWWSVKIVLWVGMTVIPFLLPS-EF 138
V R+ G +FY ++ L + ++ R H+G+W K + + + F +P F
Sbjct: 113 AVYRLCFGLAMFYLLLSLLMIKVKSSSDPRAAVHNGFWFFKFAAAIAIIIGAFFIPEGTF 172
Query: 139 IQIYGEVAHFGAGVFLLIQLISIISFI-TWLNDCCES-EKYAAKCQIHVMLFATTA-YVV 195
++ V GA F+LIQL+ +I F +W E E+ ++C +L AT Y++
Sbjct: 173 TTVWFYVGMAGAFCFILIQLVLLIDFAHSWNESWVEKMEEGNSRCWYAALLSATALNYLL 232
Query: 196 CLVGIILMFIWYAPKPSCLLNIFFIAWTLVLLQLMTSVSLHPKVN-----AGILTPGLMG 250
LV I+L F++Y SC N FI+ ++L + +S+ PK+ +G+L ++
Sbjct: 233 SLVAIVLFFVYYTHPASCSENKAFISVNMLLCVGASVMSILPKIQESQPRSGLLQSSVIT 292
Query: 251 LYVVFLCWNAIRSEPDGG------SCIRKSDSST------KTDWL---SIISFIIAILAI 295
+Y ++L W+A+ +EP+ S I + +ST W II I+ +L +
Sbjct: 293 VYTMYLTWSAMTNEPETNCNPSLLSIIGYNTTSTVPKEGQSVQWWHAQGIIGLILFLLCV 352
Query: 296 VIATFSTGIDSKCFQF---------------RKDDDIPAEDDV-----------PYGYGF 329
++ T +S+ + R D + DDV Y Y F
Sbjct: 353 FYSSIRTSNNSQVNKLTLTSDESTLIEDGGARSDGSLEDGDDVHRAVDNERDGVTYSYSF 412
Query: 330 FHFVFATGAMYFAMLLIGWNSHHSMRKWTIDVGWTSTWVRIVNEWLAVCVYLWMLVAPII 389
FHF+ ++Y M L W + R+ WT+ WV+I + W+ + +Y+W LVAP++
Sbjct: 413 FHFMLFLASLYIMMTLTNWYRYEPSREMKSQ--WTAVWVKISSSWIGIVLYVWTLVAPLV 470
Query: 390 WKSRQTD 396
+R D
Sbjct: 471 LTNRDFD 477
>UniRef100_Q9NRX5 Tumor differentially expressed protein 2 [Homo sapiens]
Length = 453
Score = 140 bits (354), Expect = 5e-32
Identities = 109/427 (25%), Positives = 192/427 (44%), Gaps = 61/427 (14%)
Query: 27 NASNPGMARYVYALMFLVANLLAWAARDYGRGALTEMERLKG-CNGGKDCL------GAE 79
+ +N + R +YAL LV +A G ++ ++ G C K + G +
Sbjct: 31 SGNNSTVTRLIYALFLLVGVCVACVMLIPGMEE--QLNKIPGFCENEKGVVPCNILVGYK 88
Query: 80 GVLRVSLGCFIFYFIMFLTTARTSKLNEVRDTWHSGWWSVKIVLWVGMTVIPFLLPS-EF 138
V R+ G +FY ++ L + ++ R H+G+W K + + + F +P F
Sbjct: 89 AVYRLCFGLAMFYLLLSLLMIKVKSSSDPRAAVHNGFWFFKFAAAIAIIIGAFFIPEGTF 148
Query: 139 IQIYGEVAHFGAGVFLLIQLISIISFI-TWLNDCCES-EKYAAKCQIHVMLFATTA-YVV 195
++ V GA F+LIQL+ +I F +W E E+ ++C +L AT Y++
Sbjct: 149 TTVWFYVGMAGAFCFILIQLVLLIDFAHSWNESWVEKMEEGNSRCWYAALLSATALNYLL 208
Query: 196 CLVGIILMFIWYAPKPSCLLNIFFIAWTLVLLQLMTSVSLHPKVN-----AGILTPGLMG 250
LV I+L F++Y SC N FI+ ++L + +S+ PK+ +G+L ++
Sbjct: 209 SLVAIVLFFVYYTHPASCSENKAFISVNMLLCVGASVMSILPKIQESQPRSGLLQSSVIT 268
Query: 251 LYVVFLCWNAIRSEPDGG------SCIRKSDSST------KTDWL---SIISFIIAILAI 295
+Y ++L W+A+ +EP+ S I + +ST W II I+ +L +
Sbjct: 269 VYTMYLTWSAMTNEPETNCNPSLLSIIGYNTTSTVPKEGQSVQWWHAQGIIGLILFLLCV 328
Query: 296 VIATFSTGIDSKCFQF---------------RKDDDIPAEDDV-----------PYGYGF 329
++ T +S+ + R D + DDV Y Y F
Sbjct: 329 FYSSIRTSNNSQVNKLTLTSDESTLIEDGGARSDGSLEDGDDVHRAVDNERDGVTYSYSF 388
Query: 330 FHFVFATGAMYFAMLLIGWNSHHSMRKWTIDVGWTSTWVRIVNEWLAVCVYLWMLVAPII 389
FHF+ ++Y M L W + R+ WT+ WV+I + W+ + +Y+W LVAP++
Sbjct: 389 FHFMLFLASLYIMMTLTNWYRYEPSREMKSQ--WTAVWVKISSSWIGIVLYVWTLVAPLV 446
Query: 390 WKSRQTD 396
+R D
Sbjct: 447 LTNRDFD 453
>UniRef100_Q7ZWT2 TDE2 protein [Xenopus laevis]
Length = 476
Score = 140 bits (353), Expect = 6e-32
Identities = 101/382 (26%), Positives = 178/382 (46%), Gaps = 52/382 (13%)
Query: 66 LKGCNGGKDC---LGAEGVLRVSLGCFIFYFIMFLTTARTSKLNEVRDTWHSGWWSVKIV 122
+ G +G +C +G + V RV G +F+ + L + + R + H+G+W K
Sbjct: 96 IPGISGHVNCDVLVGYKAVYRVCFGMAMFFLLFSLLMIKVKSSQDPRASLHNGFWFFKFA 155
Query: 123 LWVGMTVIPFLLP-SEFIQIYGEVAHFGAGVFLLIQLISIISFI-TWLNDCCES-EKYAA 179
VG+TV F +P F ++ V GA F+LIQL+ +I F +W E E+ +
Sbjct: 156 AAVGITVGAFFIPEGPFTTVWFYVGMGGAFCFILIQLVLLIDFAHSWNESWVEKMEEGNS 215
Query: 180 KCQIHVMLFAT-TAYVVCLVGIILMFIWYAPKPSCLLNIFFIAWTLVLLQLMTSVSLHPK 238
+C +L AT YV+ LV I+L++++Y C N FI+ ++L ++ +S+ PK
Sbjct: 216 RCWYVALLSATGLNYVLSLVAIVLLYVYYTHPEGCAENKAFISVNMLLCIGVSIMSVLPK 275
Query: 239 V-----NAGILTPGLMGLYVVFLCWNAIRSEPDGG------SCIRKSDSSTK-------- 279
+ +G+L ++ +Y ++L W+A+ +EPD S I + +ST
Sbjct: 276 IQESQPRSGLLQSSVITVYTMYLTWSAMTNEPDRKCNPSLLSIIGYNSTSTPGQVKVVQW 335
Query: 280 TDWLSIISFIIAILAIVIATFSTGIDSKCFQFRKDDDIPA-------------------- 319
D I+ ++ +L ++ ++ T +S+ + D
Sbjct: 336 WDAQGIVGLVLFLLCVLYSSIRTSNNSQVNKLTLTSDESTLIEDGGRSEGSMDDSDNAHR 395
Query: 320 -----EDDVPYGYGFFHFVFATGAMYFAMLLIGWNSHHSMRKWTIDVGWTSTWVRIVNEW 374
D V Y Y FFHF+ ++Y M L W S S + + W S WV+I + W
Sbjct: 396 AVDNERDGVTYSYSFFHFMLFLASLYIMMTLTNWYSPDSSYE-MMTSKWPSVWVKISSSW 454
Query: 375 LAVCVYLWMLVAPIIWKSRQTD 396
+ + +Y+W LVAP++ +R+ D
Sbjct: 455 VCIVLYVWTLVAPLVLTNREFD 476
>UniRef100_Q5R419 Hypothetical protein DKFZp459J102 [Pongo pygmaeus]
Length = 453
Score = 140 bits (353), Expect = 6e-32
Identities = 109/427 (25%), Positives = 192/427 (44%), Gaps = 61/427 (14%)
Query: 27 NASNPGMARYVYALMFLVANLLAWAARDYGRGALTEMERLKG-CNGGKDCL------GAE 79
+ +N + R +YAL LV +A G ++ ++ G C K + G +
Sbjct: 31 SGNNSTVTRLIYALFLLVGVCVACVMLIPGMEE--QLNKIPGFCENEKGVVPCNILVGYK 88
Query: 80 GVLRVSLGCFIFYFIMFLTTARTSKLNEVRDTWHSGWWSVKIVLWVGMTVIPFLLPS-EF 138
V R+ G +FY ++ L + ++ R H+G+W K + + + F +P F
Sbjct: 89 AVYRLCFGLAMFYLLLSLLMIKVKSSSDPRAAVHNGFWFFKFAAAIAIIIGAFFIPEGTF 148
Query: 139 IQIYGEVAHFGAGVFLLIQLISIISFI-TWLNDCCES-EKYAAKCQIHVMLFATTA-YVV 195
++ V GA F+LIQL+ +I F +W E E+ ++C +L AT Y++
Sbjct: 149 TTVWFYVGMAGAFCFILIQLVLLIDFAHSWNESWVEKMEEGNSRCWYAALLSATALNYLL 208
Query: 196 CLVGIILMFIWYAPKPSCLLNIFFIAWTLVLLQLMTSVSLHPKVN-----AGILTPGLMG 250
LV I+L F++Y SC N FI+ ++L + +S+ PK+ +G+L ++
Sbjct: 209 SLVAIVLFFVYYTHPASCSENKAFISVNMLLCIGASVMSILPKIQESQPRSGLLQSSVIT 268
Query: 251 LYVVFLCWNAIRSEPDGG------SCIRKSDSST------KTDWL---SIISFIIAILAI 295
+Y ++L W+A+ +EP+ S I + +ST W II I+ +L +
Sbjct: 269 VYTMYLTWSAMTNEPETNCNPSLLSIIGYNTTSTVPKEGQSVQWWHAQGIIGLILFLLCV 328
Query: 296 VIATFSTGIDSKCFQF---------------RKDDDIPAEDDV-----------PYGYGF 329
++ T +S+ + R D + DDV Y Y F
Sbjct: 329 FYSSIRTSNNSQVNKLTLTSDESTLIEDGGARSDGSLEDGDDVHRAVDNERDGVTYSYSF 388
Query: 330 FHFVFATGAMYFAMLLIGWNSHHSMRKWTIDVGWTSTWVRIVNEWLAVCVYLWMLVAPII 389
FHF+ ++Y M L W + R+ WT+ WV+I + W+ + +Y+W LVAP++
Sbjct: 389 FHFMLFLASLYIMMTLTNWYRYEPSREMKSQ--WTAVWVKISSSWIGIVLYVWTLVAPLV 446
Query: 390 WKSRQTD 396
+R D
Sbjct: 447 LTNRDFD 453
>UniRef100_Q6NS17 Hypothetical protein [Xenopus laevis]
Length = 460
Score = 139 bits (351), Expect = 1e-31
Identities = 103/382 (26%), Positives = 178/382 (45%), Gaps = 52/382 (13%)
Query: 66 LKGCNGGKDC---LGAEGVLRVSLGCFIFYFIMFLTTARTSKLNEVRDTWHSGWWSVKIV 122
+ G +G +C +G + V RV G +F+ + L + + R + H+G+W K
Sbjct: 80 IPGISGHANCDVLVGYKAVYRVCFGMAMFFLLFSLLMIKVKSSQDPRASVHNGFWFFKFA 139
Query: 123 LWVGMTVIPFLLP-SEFIQIYGEVAHFGAGVFLLIQLISIISFI-TWLNDCCES-EKYAA 179
VG+TV F +P F ++ V GA F+LIQL+ +I F +W E E+ +
Sbjct: 140 AAVGITVGAFFIPEGPFTTVWFYVGMGGAFCFILIQLVLLIDFAHSWNESWVEKMEEGNS 199
Query: 180 KCQIHVMLFAT-TAYVVCLVGIILMFIWYAPKPSCLLNIFFIAWTLVLLQLMTSVSLHPK 238
+C +L AT YV+ LV I+L +++Y C N FI+ ++L + +S+ PK
Sbjct: 200 RCWYAALLSATGLNYVLSLVAIVLFYVYYTHPEGCAENKAFISVNMLLCIGASIMSVLPK 259
Query: 239 V-----NAGILTPGLMGLYVVFLCWNAIRSEPDGG------SCIRKSDSSTK-------- 279
+ +G+L ++ +Y ++L W+A+ +EPD S I + +ST
Sbjct: 260 IQESQPRSGLLQSSVITVYTMYLTWSAMTNEPDRKCNPSLLSIIGYNSTSTPGQVKVVQW 319
Query: 280 TDWLSIISFIIAILAIVIATFSTGIDSKC------------------FQFRKDDDIPA-- 319
D I+ ++ +L ++ ++ T +S+ + DD A
Sbjct: 320 WDAQGIVGLVLFLLCVLYSSIRTSNNSQVNKLTLTSDESTLIEDGGRSEVSMDDSDNAHR 379
Query: 320 -----EDDVPYGYGFFHFVFATGAMYFAMLLIGWNSHHSMRKWTIDVGWTSTWVRIVNEW 374
D V Y Y FFHF+ ++Y M L W S S + + W S WV+I + W
Sbjct: 380 AVDNERDGVTYSYSFFHFMLFLASLYIMMTLTNWYSPDSSYE-MMTSKWPSVWVKISSSW 438
Query: 375 LAVCVYLWMLVAPIIWKSRQTD 396
+ + +Y+W LVAP++ +R+ D
Sbjct: 439 VCIVLYVWTLVAPLVLTNREFD 460
>UniRef100_O45719 Hypothetical protein R11H6.2 [Caenorhabditis elegans]
Length = 442
Score = 138 bits (348), Expect = 2e-31
Identities = 110/413 (26%), Positives = 188/413 (44%), Gaps = 50/413 (12%)
Query: 28 ASNPGMARYVYALMFLVANLLAWAARDYG-RGALTEMERLKGCNG-----GKDC---LGA 78
A N R +YALM + A +A G + L E + L C+G G +C +G
Sbjct: 33 AKNSTTTRIMYALMLISATFMAVVMLLPGVQKKLVENKWL--CDGLNEYAGVNCEHAIGY 90
Query: 79 EGVLRVSLGCFIFYFIMFLTTARTSKLNEVRDTWHSGWWSVKIVLWVGMTVIPFLLPSEF 138
+ V RV G F+F+ L S + R + +G+W K +L G+ F + SE
Sbjct: 91 QAVYRVCAGAASFFFLFMLLMFGVSSSKDGRSSIQNGFWFFKYLLMFGIIGGFFFIGSET 150
Query: 139 IQI-YGEVAHFGAGVFLLIQLISIISFITWLNDCCESEKYAAKCQIHVMLFATTAYVVCL 197
+ + GA +F+LIQLI I+ F L + + + C +++ ++VCL
Sbjct: 151 LATPLMYIGMLGAFLFILIQLILIVDFAHGLAESQYEDNDSRACYAGLLITTFGGFLVCL 210
Query: 198 VGIILMFIWYAPKPSCLLNIFFIAWTLVLLQLMTSVSLHPKV-----NAGILTPGLMGLY 252
+ + +FI YA C L FF+ + +++ ++ +S+ P V +G+L P ++ Y
Sbjct: 211 IAAVYVFINYAIGDGCGLPKFFVIFNVLICVAISLLSVSPMVQEVNPRSGLLQPVVISAY 270
Query: 253 VVFLCWNAIRSEPDGGSC----------------IRKSDS-STKTDWLSIISFIIAILAI 295
+++L W+A+ S P+ SC + K DS T S+IS +I ++ +
Sbjct: 271 IIYLTWSALLSNPN-ESCNPTLANVTQSAIPTGGVTKDDSFVTPLPVHSLISLLIWLICL 329
Query: 296 VIATFSTGIDSKCFQFRKDDDIPA--------------EDDVPYGYGFFHFVFATGAMYF 341
V A+ ++ + D++ E+ V Y Y FFHF+F ++Y
Sbjct: 330 VYASIRNSSNTSLGKITGDNEEHVQLNDVEGGKAWDNEEEGVAYSYSFFHFMFCLASLYV 389
Query: 342 AMLLIGWNSHHSMRKWTIDVGWTSTWVRIVNEWLAVCVYLWMLVAPIIWKSRQ 394
M L W H ++ S WV++ + W+ +Y W LVAPII+ R+
Sbjct: 390 MMTLTSW-YHPDSDLAHLNSNMASVWVKMFSSWICGGLYAWTLVAPIIFPDRE 441
>UniRef100_Q5ZME5 Hypothetical protein [Gallus gallus]
Length = 461
Score = 138 bits (347), Expect = 3e-31
Identities = 100/383 (26%), Positives = 178/383 (46%), Gaps = 53/383 (13%)
Query: 66 LKGCNGGKDC---LGAEGVLRVSLGCFIFYFIMFLTTARTSKLNEVRDTWHSGWWSVKIV 122
+ G +G +C +G + V RV G +F+ + L + N+ R H+G+W K
Sbjct: 80 IPGVHGHVNCDVLVGYKAVYRVCFGMAMFFLLFSLLMIKVKSSNDPRAAVHNGFWFFKFA 139
Query: 123 LWVGMTVIPFLLPS-EFIQIYGEVAHFGAGVFLLIQLISIISFI-TWLNDCCES-EKYAA 179
+ ++V F +P F ++ V GA F+LIQL+ +I F +W E E+ +
Sbjct: 140 TALAISVGAFFIPEGPFTTVWFYVGMSGAFCFILIQLVLLIDFAHSWNESWVEKMEEGNS 199
Query: 180 KCQIHVMLFATTA-YVVCLVGIILMFIWYAPKPSCLLNIFFIAWTLVLLQLMTSVSLHPK 238
+C +L AT Y++ LV I+L +++Y C N FI+ ++L + +S+ P+
Sbjct: 200 RCWYAALLSATAVNYLLSLVAIVLFYVYYTHPEGCSENKTFISVNMLLCIGASVMSILPR 259
Query: 239 VN-----AGILTPGLMGLYVVFLCWNAIRSEPDGG------SCIRKSDSSTKT------- 280
+ +G+L ++ +Y ++L W+A+ +EPD S I + ++ T
Sbjct: 260 IQESQPRSGLLQSSVITIYTMYLTWSAMTNEPDRRCNPSLLSIIGYNTTTIPTQGQVVQW 319
Query: 281 -DWLSIISFIIAILAIVIATFSTGIDSKCFQF---------------RKDDDIPAEDDV- 323
D I+ I+ +L ++ ++ T +S+ + R D + DDV
Sbjct: 320 WDAQGIVGLILFLLCVLYSSIRTSNNSQVNKLMLTSDESTLIEDGMPRNDGSLDDGDDVH 379
Query: 324 ----------PYGYGFFHFVFATGAMYFAMLLIGWNSHHSMRKWTIDVGWTSTWVRIVNE 373
Y Y FFHF+ ++Y M L W S S + T+ W S WV+I +
Sbjct: 380 RAIDNERDGVTYSYSFFHFMLFLASLYIMMTLTNWYSPDSTYE-TMTSKWPSVWVKISSS 438
Query: 374 WLAVCVYLWMLVAPIIWKSRQTD 396
W+ + +Y+W LVAP++ +R D
Sbjct: 439 WIGIVLYVWTLVAPLVLTNRDFD 461
>UniRef100_UPI00001A07CE UPI00001A07CE UniRef100 entry
Length = 458
Score = 137 bits (345), Expect = 5e-31
Identities = 102/399 (25%), Positives = 180/399 (44%), Gaps = 66/399 (16%)
Query: 58 GALTEMERLKG-CNGGKDCLGAEGVLRV--------------SLGCFIFYFIMFLTTART 102
G T+++++ G C GG G EG + ++ CF F F + + R+
Sbjct: 60 GMETQLKKIPGFCEGGSSIPGFEGKVNCDVIVGYKSVYRMCFAMACFFFLFSIIMIRVRS 119
Query: 103 SKLNEVRDTWHSGWWSVKIVLWVGMTVIPFLLPSE-FIQIYGEVAHFGAGVFLLIQLISI 161
SK + R +G+W K ++ VG+TV F +P F ++ G+ +F+LIQLI +
Sbjct: 120 SK--DPRAAIQNGFWFFKFLILVGITVGAFFIPDGMFNTVWYYFGVVGSFIFILIQLILL 177
Query: 162 ISFI-TWLNDCCES-EKYAAKCQIHVMLFATTAYVVC-LVGIILMFIWYAPKPSCLLNIF 218
+ F +W E+ E +KC +L T + +C ++L +I+Y C N
Sbjct: 178 VDFAHSWNQKWVENAEDGNSKCWYAALLSFTLVHYICAFAAVVLFYIFYTQPDDCTENKV 237
Query: 219 FIAWTLVLLQLMTSVSLHPKV-----NAGILTPGLMGLYVVFLCWNAIRSEPDG------ 267
FI+ L+ +++ V++ PKV ++G+L L+ LY ++L W+A+ + P+
Sbjct: 238 FISLNLIFCIIVSVVAILPKVQEAQPSSGLLQASLISLYTMYLTWSAMSNNPNRKCNPSL 297
Query: 268 -----GSCIRKSDSSTK---TDWL---SIISFIIAILAIVIATFSTGIDSKCFQFRKDDD 316
G + +ST T W S++ +I +L + A+ + +S+ + + ++
Sbjct: 298 LSIVKGQSAAPTPTSTPGVFTQWWDAQSVVGLVIFLLCTLYASIRSSNNSQVNKLMQTEE 357
Query: 317 IPA----------------------EDDVPYGYGFFHFVFATGAMYFAMLLIGWNSHHSM 354
+ ED V YGY FFHF ++Y M L W +
Sbjct: 358 VQGLASSDANDAISEDGVRRAVDNEEDGVTYGYSFFHFSLFLASLYIMMTLTNWYQPETD 417
Query: 355 RKWTIDVGWTSTWVRIVNEWLAVCVYLWMLVAPIIWKSR 393
+ S WV+I + WL + +YLW LVAP++ R
Sbjct: 418 YA-AMTTTMPSVWVKICSSWLGLGLYLWTLVAPLVLTDR 455
>UniRef100_UPI00003614DC UPI00003614DC UniRef100 entry
Length = 471
Score = 133 bits (335), Expect = 8e-30
Identities = 97/381 (25%), Positives = 174/381 (45%), Gaps = 51/381 (13%)
Query: 66 LKGCNGGKDC---LGAEGVLRVSLGCFIFYFIMFLTTARTSKLNEVRDTWHSGWWSVKIV 122
L G G +C +G V RV G F+ + L + + R H+G+W K
Sbjct: 92 LPGVGGHVNCDVLVGYRAVYRVCFGMASFFLLFSLLMIKVKSSQDPRAAVHNGFWFFKFA 151
Query: 123 LWVGMTVIPFLLPS-EFIQIYGEVAHFGAGVFLLIQLISIISFI-TWLNDCCES-EKYAA 179
V +TV F + F ++ + GA F+LIQL+ +I F +W E E+ +
Sbjct: 152 AAVSLTVGAFFISEGPFTTVWFYIGMAGAFCFILIQLVLLIDFAHSWNESWVEKMEEGNS 211
Query: 180 KCQIHVMLFATTA-YVVCLVGIILMFIWYAPKPSCLLNIFFIAWTLVLLQLMTSVSLHPK 238
+C +L TT Y++ +V ++L +++Y C N FI+ ++L + +S+ P+
Sbjct: 212 RCWYAALLSVTTVNYLLSVVSLVLFYLYYTHSDGCTENKVFISINMLLCVAASLLSILPQ 271
Query: 239 VN-----AGILTPGLMGLYVVFLCWNAIRSEPDG-------GSCIRKSDSSTKTDWL--- 283
+ +G+L L+ L+ ++L W+A+ +EP+ G S S+T+ D +
Sbjct: 272 IQESQPRSGLLQSSLVTLFTMYLTWSAMTNEPERKCNPSLLGIIGLNSTSATEQDHVVQW 331
Query: 284 ----SIISFIIAILAIVIATFSTGIDSKC--FQFRKDD-----DIPA------------- 319
I+ ++ +L ++ ++ +++ R D+ D PA
Sbjct: 332 WDAQGIVGLVLFLLCVLYSSIRNSSNAQVNKLTLRSDESALIEDGPAVDSFEEDSSPNRA 391
Query: 320 ----EDDVPYGYGFFHFVFATGAMYFAMLLIGWNSHHSMRKWTIDVGWTSTWVRIVNEWL 375
+D V Y Y FFHF+ ++Y M L W S S + + W S WV+I + W+
Sbjct: 392 LDNEKDGVTYSYSFFHFMLFLASLYIMMTLTNWYSPDSNNE-KMTSRWPSVWVKICSSWV 450
Query: 376 AVCVYLWMLVAPIIWKSRQTD 396
+ +Y+W LVAP++ +R D
Sbjct: 451 CIALYVWTLVAPLVLVNRDFD 471
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.328 0.140 0.472
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 701,859,507
Number of Sequences: 2790947
Number of extensions: 29169689
Number of successful extensions: 86721
Number of sequences better than 10.0: 164
Number of HSP's better than 10.0 without gapping: 113
Number of HSP's successfully gapped in prelim test: 51
Number of HSP's that attempted gapping in prelim test: 86099
Number of HSP's gapped (non-prelim): 267
length of query: 398
length of database: 848,049,833
effective HSP length: 129
effective length of query: 269
effective length of database: 488,017,670
effective search space: 131276753230
effective search space used: 131276753230
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.8 bits)
S2: 76 (33.9 bits)
Lotus: description of TM0214.6


