

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0213.7
(451 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
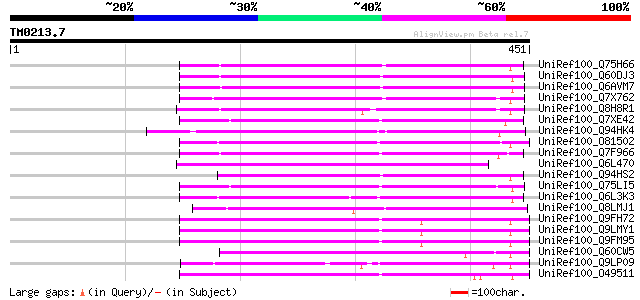
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q75H66 Hypothetical protein OSJNBb0007E22.22 [Oryza sa... 209 1e-52
UniRef100_Q60DJ3 MuDR family transposase protein [Oryza sativa] 199 2e-49
UniRef100_Q6AVM7 Putative polyprotein [Oryza sativa] 197 7e-49
UniRef100_Q7X762 OSJNBb0118P14.3 protein [Oryza sativa] 189 1e-46
UniRef100_Q8H8R1 Putative mutator-like transposase [Oryza sativa] 184 4e-45
UniRef100_Q7XE42 Putative mutator-like transposase [Oryza sativa] 182 2e-44
UniRef100_Q94HK4 Mutator-like transposase [Oryza sativa] 182 2e-44
UniRef100_O81502 F9D12.2 protein [Arabidopsis thaliana] 181 4e-44
UniRef100_Q7F966 OSJNBa0091C07.2 protein [Oryza sativa] 177 5e-43
UniRef100_Q6L470 Putative mutator transposable element [Solanum ... 169 1e-40
UniRef100_Q94HS2 Putative transposon protein [Oryza sativa] 168 3e-40
UniRef100_Q75LI5 Putative transposon protein [Oryza sativa] 163 8e-39
UniRef100_Q6L3K3 Putative transposon MuDR mudrA-like protein [So... 162 2e-38
UniRef100_Q8LMJ1 Putative transposon protein [Oryza sativa] 157 6e-37
UniRef100_Q9FH72 Arabidopsis thaliana genomic DNA, chromosome 5,... 155 2e-36
UniRef100_Q9LMY1 F21F23.10 protein [Arabidopsis thaliana] 155 3e-36
UniRef100_Q9FM95 Similarity to mutator-like transposase [Arabido... 155 3e-36
UniRef100_Q60CW5 Putative mutator transposable element-related p... 152 1e-35
UniRef100_Q9LP09 F9C16.9 [Arabidopsis thaliana] 145 3e-33
UniRef100_O49511 MuDR transposable element - like protein [Arabi... 140 7e-32
>UniRef100_Q75H66 Hypothetical protein OSJNBb0007E22.22 [Oryza sativa]
Length = 981
Score = 209 bits (533), Expect = 1e-52
Identities = 108/307 (35%), Positives = 165/307 (53%), Gaps = 11/307 (3%)
Query: 148 QGLLTVFDDLMGGVEHIFCLRHLYANFKKKFGGGVVIRNLMMAAAKATYHQLWTEKMMEL 207
+GL+ L EH FC+RHLY NF++ F G + ++N + A A+++ Q W K E+
Sbjct: 547 KGLIPAVQQLFPDSEHRFCVRHLYQNFQQSFKGEI-LKNQLWACARSSSVQEWNTKFEEM 605
Query: 208 QALHPEAYTWLMQIPTRAWCKHAFTFYPKCDVLMNNLSESFNATILLVRDKPVLTMVDWI 267
+AL+ +AY WL Q+ W + F+ +PKCD+L+NN E FN IL R+ P+LTM++ I
Sbjct: 606 KALNEDAYNWLEQMAPNTWVRAFFSDFPKCDILLNNSCEVFNKYILEAREMPILTMLEKI 665
Query: 268 RSYIMGRFATMNEKLEKYKGEVMPKPLKRLEHEIEMVRYWVATRCGEDKYEVKHMVSQDQ 327
+ +M RF ++ +K++G + PK K+L E G+ ++V+ + +
Sbjct: 666 KGQLMTRFFNKQKEAQKWQGPICPKIRKKLLKIAEQANICYVLPAGKGVFQVEERGT--K 723
Query: 328 FVVNLAEHTCSCNFWELVGIPCRHAVAAISHSSQRPEDFVHPYYRREAYMATYGHVITPI 387
++V++ C C W+L GIPC HA+A I EDF+ Y A+ A Y I P
Sbjct: 724 YIVDVVTKHCDCRRWDLTGIPCCHAIACIREDHLSEEDFLPHCYSINAFKAVYAENIIPC 783
Query: 388 NGQKLWPTTNDPYILPPLYKRAPGRPKKLRRRDPHEDTTDNQTRLR--------SRCGQF 439
N + W N P ILPP+Y++ GRPKK RR+ P E N +L S C +
Sbjct: 784 NDKANWEKMNGPQILPPVYEKKVGRPKKSRRKQPQEVQGRNGPKLTKHGVTIHCSYCHEA 843
Query: 440 GHNSRSC 446
HN + C
Sbjct: 844 NHNKKGC 850
>UniRef100_Q60DJ3 MuDR family transposase protein [Oryza sativa]
Length = 1030
Score = 199 bits (505), Expect = 2e-49
Identities = 103/309 (33%), Positives = 165/309 (53%), Gaps = 12/309 (3%)
Query: 148 QGLLTVFDDLMGGVEHIFCLRHLYANFKKKFGGGVVIRNLMMAAAKATYHQLWTEKMMEL 207
+GL+ + EH FC+RHLY+NF+ +F G V ++N + A A+++ Q W + M +
Sbjct: 582 KGLIPAVQQVFPESEHRFCVRHLYSNFQLQFKGEV-LKNQLWACARSSSVQEWNKNMDVM 640
Query: 208 QALHPEAYTWLMQIPTRAWCKHAFTFYPKCDVLMNNLSESFNATILLVRDKPVLTMVDWI 267
+ L+ AY WL ++P W + F+ +PKCD+L+NN E FN IL R+ P+LTM++ I
Sbjct: 641 RNLNKSAYEWLEKLPPNTWVRAFFSEFPKCDILLNNNCEVFNKYILEARELPILTMLEKI 700
Query: 268 RSYIMGR-FATMNEKLEKYKGEVMPKPLKRLEHEIEMVRYWVATRCGEDKYEVKHMVSQD 326
+ +M R F E ++++G + PK K++ + A G+ ++V +
Sbjct: 701 KGQLMTRHFNKQKELADQFQGLICPKIRKKVLKNADAANTCYALPAGQGIFQVHER--EY 758
Query: 327 QFVVNLAEHTCSCNFWELVGIPCRHAVAAISHSSQRPEDFVHPYYRREAYMATYGHVITP 386
Q++V++ C C W+L GIPC HA++ + H E + Y +A+ YG I P
Sbjct: 759 QYIVDINAMYCDCRRWDLTGIPCNHAISCLRHERINAESILPNCYTTDAFSKAYGFNIWP 818
Query: 387 INGQKLWPTTNDPYILPPLYKRAPGRPKKLRRRDPHEDTTDNQTRLRSR--------CGQ 438
N + W N P I PP+Y++ GRPKK RR+ P+E N +L CG+
Sbjct: 819 CNDKSKWENVNGPEIKPPVYEKKAGRPKKSRRKAPYEVIGKNGPKLTKHGVMMHYKYCGE 878
Query: 439 FGHNSRSCR 447
HNS C+
Sbjct: 879 ENHNSGGCK 887
>UniRef100_Q6AVM7 Putative polyprotein [Oryza sativa]
Length = 1006
Score = 197 bits (500), Expect = 7e-49
Identities = 103/309 (33%), Positives = 164/309 (52%), Gaps = 12/309 (3%)
Query: 148 QGLLTVFDDLMGGVEHIFCLRHLYANFKKKFGGGVVIRNLMMAAAKATYHQLWTEKMMEL 207
+GL+ + EH FC+RHLY+NF+ +F G V +N + A A+++ Q W + M +
Sbjct: 558 KGLIPAVQQVFPESEHRFCVRHLYSNFQLQFKGEVP-KNQLWACARSSSVQEWNKNMDVM 616
Query: 208 QALHPEAYTWLMQIPTRAWCKHAFTFYPKCDVLMNNLSESFNATILLVRDKPVLTMVDWI 267
+ L+ AY WL ++P W + F+ +PKCD+L+NN E FN IL R+ P+LTM++ I
Sbjct: 617 RNLNKSAYEWLEKLPPNTWVRAFFSEFPKCDILLNNNCEVFNKYILEARELPILTMLEKI 676
Query: 268 RSYIMGR-FATMNEKLEKYKGEVMPKPLKRLEHEIEMVRYWVATRCGEDKYEVKHMVSQD 326
+ +M R F E ++++G + PK K++ + A G+ ++V +
Sbjct: 677 KGQLMTRHFNKQKELADQFQGLICPKIRKKVLKNADAANTCYALPAGQGIFQVHER--EY 734
Query: 327 QFVVNLAEHTCSCNFWELVGIPCRHAVAAISHSSQRPEDFVHPYYRREAYMATYGHVITP 386
Q++V++ C C W+L GIPC HA++ + H E + Y +A+ YG I P
Sbjct: 735 QYIVDINAMHCDCRRWDLTGIPCNHAISCLRHERINAESILPNCYTTDAFSKAYGFNIWP 794
Query: 387 INGQKLWPTTNDPYILPPLYKRAPGRPKKLRRRDPHEDTTDNQTRLRSR--------CGQ 438
N + W N P I PP+Y++ GRPKK RR+ P+E N +L CG+
Sbjct: 795 CNDKSKWENINGPEIKPPVYEKKAGRPKKSRRKAPYEVIGKNGPKLTKHGVMMHCKYCGE 854
Query: 439 FGHNSRSCR 447
HNS C+
Sbjct: 855 ENHNSGGCK 863
>UniRef100_Q7X762 OSJNBb0118P14.3 protein [Oryza sativa]
Length = 939
Score = 189 bits (480), Expect = 1e-46
Identities = 105/309 (33%), Positives = 160/309 (50%), Gaps = 14/309 (4%)
Query: 148 QGLLTVFDDLMGGVEHIFCLRHLYANFKKKFGGGVVIRNLMMAAAKATYHQLWTEKMMEL 207
+GL+ E FC+RHLY NF+ G ++N + A A+++ W M ++
Sbjct: 517 KGLVPAVRREFSDAEQRFCVRHLYQNFQV-LHKGETLKNQLWAIARSSTVPEWNANMEKM 575
Query: 208 QALHPEAYTWLMQIPTRAWCKHAFTFYPKCDVLMNNLSESFNATILLVRDKPVLTMVDWI 267
+AL EAY +L +IP WC+ F+ +PKCD+L+NN SE FN IL R+ P+L+M++ I
Sbjct: 576 KALSSEAYKYLEEIPPNQWCRAFFSDFPKCDILLNNNSEVFNKYILDAREMPILSMLERI 635
Query: 268 RSYIMGRFATMNEKLEK-YKGEVMPKPLKRLEHEIEMVRYWVATRCGEDKYEVKHMVSQD 326
R+ IM R T ++LE+ + + PK +++E EM G ++V + S
Sbjct: 636 RNQIMNRLYTKQKELERNWPCGICPKIKRKVEKNTEMANTCYVFPAGMGAFQVSDIGS-- 693
Query: 327 QFVVNLAEHTCSCNFWELVGIPCRHAVAAISHSSQRPEDFVHPYYRREAYMATYGHVITP 386
Q++V L C C W+L GIPC HA++ + H +PED V Y Y Y + I P
Sbjct: 694 QYIVELNVKRCDCRRWQLTGIPCNHAISCLRHERIKPEDVVSFCYSTRCYEQAYSYNIMP 753
Query: 387 INGQKLWPTTNDPYILPPLYKRAPGRPKKLRRRDPHEDTTDNQTRLR--------SRCGQ 438
+ W + PP+Y++ GRPKK RR+ P E + T++ S C
Sbjct: 754 LRDSIHWEKMQGIEVKPPVYEKKVGRPKKTRRKQPQE--LEGGTKISKHGVEMHCSYCKN 811
Query: 439 FGHNSRSCR 447
GHN SC+
Sbjct: 812 GGHNKTSCK 820
>UniRef100_Q8H8R1 Putative mutator-like transposase [Oryza sativa]
Length = 746
Score = 184 bits (467), Expect = 4e-45
Identities = 103/313 (32%), Positives = 166/313 (52%), Gaps = 19/313 (6%)
Query: 146 ILQGLLTVFDDLMGGVEHIFCLRHLYANFKKKFGGGVVIRNLMMAAAKATYHQLWTEKMM 205
+++GL+ D EH FC+RHLY NF + G ++N + A A++T W
Sbjct: 391 VVEGLIPAIKDEFPDSEHRFCVRHLYQNFAVLYKGEA-LKNQLWAIARSTTVPEWNVNTE 449
Query: 206 ELQALHPEAYTWLMQIPTRAWCKHAFTFYPKCDVLMNNLSESFNATILLVRDKPVLTMVD 265
+++A++ +AY +L +IP WC+ F + KCD+L+NN E IL R+ +L+M++
Sbjct: 450 KMKAVNKDAYGYLEEIPPNQWCRAFFRDFSKCDILLNNNLECHVRYILEARELTILSMLE 509
Query: 266 WIRSYIMGRFATMNEKLEKYKGEVMPKPLKRLEHEIEMVR--YWVATRCGEDKYEVKHMV 323
IRS +M R T E+ +K+ ++ PK +++E IEM Y + +R G V +
Sbjct: 510 KIRSKLMNRIYTKQEECKKWVFDICPKIKQKVEKNIEMSNTCYALPSRMG-----VFQVT 564
Query: 324 SQD-QFVVNLAEHTCSCNFWELVGIPCRHAVAAISHSSQRPEDFVHPYYRREAYMATYGH 382
+D QFVV++ C C W+L+GIPC HA++ + H +PED V Y +++ Y
Sbjct: 565 DRDKQFVVDIKNKQCDCRRWQLIGIPCNHAISCLRHERIKPEDEVSFCYTIQSFKQAYMF 624
Query: 383 VITPINGQKLWPTTNDPYILPPLYKRAPGRPKKLRRRDPHEDTTDNQTRLR--------S 434
I P+ + W N + PP+Y++ GRPK RR+ P E D T++ S
Sbjct: 625 NIMPVRDKTHWEKMNGVPVNPPVYEKKVGRPKTTRRKQPQE--LDAGTKISKHGVQIHCS 682
Query: 435 RCGQFGHNSRSCR 447
C GHN + C+
Sbjct: 683 YCKNVGHNKKGCK 695
>UniRef100_Q7XE42 Putative mutator-like transposase [Oryza sativa]
Length = 1005
Score = 182 bits (461), Expect = 2e-44
Identities = 102/306 (33%), Positives = 158/306 (51%), Gaps = 10/306 (3%)
Query: 148 QGLLTVFDDLMGGVEHIFCLRHLYANFKKKFGGGVVIRNLMMAAAKATYHQLWTEKMMEL 207
+GLL++ L EH C RH+YAN++KK + A KA QL+ +L
Sbjct: 552 KGLLSIVSTLFPFAEHRMCARHIYANWRKKHRLQEYQKRFWKIA-KAPNEQLFNHYKRKL 610
Query: 208 QALHPEAYTWLMQIPTRAWCKHAFTFYPKCDVLMNNLSESFNATILLVRDKPVLTMVDWI 267
A P + L + W + F C+ + NN+SESFN+ I+ R KP++TM++ I
Sbjct: 611 AAKTPRGWQDLEKTNPIHWSRAWFRLGSNCESVDNNMSESFNSWIIESRFKPIITMLEDI 670
Query: 268 RSYIMGRFATMNEKLEKYKGEVMPKPLKRLEHEIEMVRYWVATRCGEDKYEVKHMVSQDQ 327
R + R E++ V P +++ ++ G+ +EV+ + +
Sbjct: 671 RIQVTRRIQENRSNSERWTMTVCPNIIRKFNKIRHRTQFCHVLWNGDAGFEVRD--KKWR 728
Query: 328 FVVNLAEHTCSCNFWELVGIPCRHAVAAISHSSQRPEDFVHPYYRREAYMATYGHVITPI 387
F V+L TCSC +W++ GIPC+HA AA+ +Q P + +H + E Y TY HV+ P+
Sbjct: 729 FTVDLTSKTCSCRYWQVSGIPCQHACAALFKMAQEPNNCIHECFSLERYKKTYQHVLQPV 788
Query: 388 NGQKLWPTTNDPYILPPLYKRAPGRPKKLRRRDPHEDTTDNQ------TRLR-SRCGQFG 440
+ WP + +P LPP K+ PGRPKK RR+DP + T++R RCG +G
Sbjct: 789 EHESAWPVSPNPKPLPPRVKKMPGRPKKNRRKDPSKPVKSGTKSSKVGTKIRCRRCGNYG 848
Query: 441 HNSRSC 446
HNSR+C
Sbjct: 849 HNSRTC 854
>UniRef100_Q94HK4 Mutator-like transposase [Oryza sativa]
Length = 725
Score = 182 bits (461), Expect = 2e-44
Identities = 111/331 (33%), Positives = 170/331 (50%), Gaps = 8/331 (2%)
Query: 120 TEGKMSISLTVHFTNFLVEPIYEVS*ILQGLLTVFDDLMGGVEHIFCLRHLYANFKKKFG 179
T G ++ + N + PI ++ + VF D EH +C HL N K
Sbjct: 386 TNGAQVLAASARDENNNMFPIAFAVGLMNAIPIVFPDS----EHRYCKMHLLQNMGNKGW 441
Query: 180 GGVVIRNLMMAAAKATYHQLWTEKMMELQALHPEAYTWLMQIPTRAWCKHAFTFYPKCDV 239
G + + AA AT + + M +L+ L+ +A+ WL+ I + +HAF+ K D+
Sbjct: 442 RGEKYKGFVDAAIYATTVWDYDKAMEDLKKLNLKAWEWLIAIGKEHFSRHAFSPKAKSDL 501
Query: 240 LMNNLSESFNATILLVRDKPVLTMVDWIRSYIMGRFATMNEKLEKYKGEVMPKPLKRLEH 299
++NNLSE FN IL RDKP++TMV+ IR +M R + + + + E+ P +LE
Sbjct: 502 VVNNLSEVFNKYILDARDKPIVTMVEHIRRKVMVRLSLKRQGGDAAQWEITPIVAGKLEM 561
Query: 300 EIEMVRYWVATRCGEDKYEVKHMVSQDQFVVNLAEHTCSCNFWELVGIPCRHAVAAISHS 359
E RY + +EV H + + F V+++ TC+C+ W+L GIPC+H V A+ +
Sbjct: 562 EKNHARYCWCYQSNLTTWEV-HCLDRS-FAVDISARTCACHKWQLTGIPCKHDVCALYKA 619
Query: 360 SQRPEDFVHPYYRREAYMATYGHVITPINGQKLWPTTNDPYILPPLYKRAPGRPKKLRRR 419
PED+V Y+R++AYM TY VI P+ + W T+ PYI PP + + GRPKK RRR
Sbjct: 620 GHTPEDYVADYFRKDAYMRTYTAVIYPVPDEHRWTKTDSPYIDPPKFDKHVGRPKKSRRR 679
Query: 420 DPHED--TTDNQTRLRSRCGQFGHNSRSCRN 448
P E + S C + +C N
Sbjct: 680 GPDEGPRVQGPARKTTSTCSNCRRDGHTCSN 710
>UniRef100_O81502 F9D12.2 protein [Arabidopsis thaliana]
Length = 940
Score = 181 bits (459), Expect = 4e-44
Identities = 107/313 (34%), Positives = 162/313 (51%), Gaps = 13/313 (4%)
Query: 148 QGLLTVFDDLMGGVEHIFCLRHLYANFKKKFGGGVVIRNLMMAAAKATYHQLWTEKMMEL 207
+GL+ DL+ EH C RH+YAN+KK +G + A A + ++ M L
Sbjct: 507 KGLINAVADLLPQAEHRHCARHVYANWKKVYGD-YCHESYFWAIAYSATEGDYSYNMDAL 565
Query: 208 QALHPEAYTWLMQIPTRAWCKHAFTFYPKCDVLMNNLSESFNATILLVRDKPVLTMVDWI 267
++ P+A L++ WC+ F+ + C+ + NNLSESFN TI R PV+ M++ +
Sbjct: 566 RSYDPDACDDLLKTDPTTWCRAFFSTHSSCEDVSNNLSESFNRTIREARKLPVVNMLEEV 625
Query: 268 RSYIMGRFATMNEKLEKYKGEVMPKPLKRLEHEIEMVRYWVATRCGEDKYEVKHMVSQDQ 327
R M R + + +K K + PK ++ LE + +Y + GE+K+E+ +
Sbjct: 626 RRISMKRISKLCDKTAKCETRFPPKIMQILEGNRKSSKYCQVLKSGENKFEI--LEGSGY 683
Query: 328 FVVNLAEHTCSCNFWELVGIPCRHAVAAISHSSQRPEDFVHPYYRREAYMATYGHVITPI 387
+ V+L TC C WEL GIPC HA+ I+ ++ PED+V + + ATY I P+
Sbjct: 684 YSVDLLTRTCGCRQWELTGIPCPHAIYVITEHNRDPEDYVDRLLETQVWKATYKDNIEPM 743
Query: 388 NGQKLWPTTNDPYILPPLYKRAPGRPKKLRR-RDPHEDTTDNQTRLR--------SRCGQ 438
NG++LW I P + GRPKK R ++P E +T N T+L S C Q
Sbjct: 744 NGERLWKRRGKGRIEVPDKRGKRGRPKKFARIKEPGESST-NPTKLTREGKTVTCSNCKQ 802
Query: 439 FGHNSRSCRNPVV 451
GHN SC+NP V
Sbjct: 803 IGHNKGSCKNPTV 815
>UniRef100_Q7F966 OSJNBa0091C07.2 protein [Oryza sativa]
Length = 879
Score = 177 bits (449), Expect = 5e-43
Identities = 101/306 (33%), Positives = 154/306 (50%), Gaps = 11/306 (3%)
Query: 148 QGLLTVFDDLMGGVEHIFCLRHLYANFKKKFGGGVVIRNLMMAAAKATYHQLWTEKMMEL 207
+GL++ ++ + +EH C RH+YAN++KK+ + AKA+ + +L
Sbjct: 513 KGLVSAVEEFLPQIEHRMCTRHIYANWRKKYRDQA-FQKPFWKCAKASCRPFFNFCRAKL 571
Query: 208 QALHPEAYTWLMQIPTRAWCKHAFTFYPKCDVLMNNLSESFNATILLVRDKPVLTMVDWI 267
L P W + F CD + NN+ ESFN I+ +R P+++M + I
Sbjct: 572 AQLTPAGAKXXXSTEPMHWSRAWFRIGSNCDSVDNNMCESFNNWIIDIRAHPIISMFEGI 631
Query: 268 RSYIMGRFATMNEKLEKYKGEVMPKPLKRLEHEIEMVRYWVATRCGEDKYEVKHMVSQDQ 327
R+ + R K + + G + P LK+L I++ A G+D +EV +
Sbjct: 632 RTKVYVRIQQNRSKAKGWLGRICPNILKKLNKYIDLSGNCEAIWNGKDGFEVTD--KDKR 689
Query: 328 FVVNLAEHTCSCNFWELVGIPCRHAVAAISHSSQRPEDFVHPYYRREAYMATYGHVITPI 387
+ V+L + TCSC +W+L GIPC HA+ A+ SS++PED++ Y E Y Y H + P+
Sbjct: 690 YTVDLEKRTCSCRYWQLAGIPCAHAITALFVSSKQPEDYIADCYSVEVYNKIYDHCMMPM 749
Query: 388 NGQKLWPTTNDPYILPPLYKRAPGRPKKLRRRDPHE-------DTTDNQTRLRSRCGQFG 440
G WP T P PP Y + PGRP+K RRRDP E T + R S+C Q
Sbjct: 750 EGMMQWPITGHPKPGPPGYVKMPGRPRKERRRDPLEAKKGKKLSKTGTKGRC-SQCKQTT 808
Query: 441 HNSRSC 446
HN R+C
Sbjct: 809 HNIRTC 814
>UniRef100_Q6L470 Putative mutator transposable element [Solanum demissum]
Length = 707
Score = 169 bits (429), Expect = 1e-40
Identities = 94/273 (34%), Positives = 146/273 (53%), Gaps = 2/273 (0%)
Query: 146 ILQGLLT-VFDDLMGGVEHIFCLRHLYANFKKKFGGGVVIRNLMMAAAKATYHQLWTEKM 204
I Q LT + +DL +++FC+RHL+ NFK+ G+ ++N + AA AT + M
Sbjct: 428 IWQWFLTYLMNDLEIEEQYLFCVRHLHNNFKRAGYSGMALKNALWKAASATTIDRFDACM 487
Query: 205 MELQALHPEAYTWLMQIPTRAWCKHAFTFYPKCDVLMNNLSESFNATILLVRDKPVLTMV 264
+L L +AY WL W + F+ PKCD+L+NN E FN IL RDKP++ ++
Sbjct: 488 TDLFELDKDAYAWLSAKVPSEWSRSHFSPLPKCDILLNNQCEVFNKFILDARDKPIVKLL 547
Query: 265 DWIRSYIMGRFATMNEKLEKYK-GEVMPKPLKRLEHEIEMVRYWVATRCGEDKYEVKHMV 323
+ IR +M R ++ EK EK+ ++ P K+L ++ ++ R YEV V
Sbjct: 548 ETIRHLLMTRINSIREKAEKWNLNDICPTIKKKLAKTMKKAANYIPQRSNMWNYEVIGPV 607
Query: 324 SQDQFVVNLAEHTCSCNFWELVGIPCRHAVAAISHSSQRPEDFVHPYYRREAYMATYGHV 383
D + V+L TCSC WEL G+PC+HA+++I + ++V Y+ + Y Y
Sbjct: 608 EGDNWAVDLYNRTCSCRQWELSGVPCKHAISSIWLKNDEVLNYVDDCYKVDTYRKIYEAS 667
Query: 384 ITPINGQKLWPTTNDPYILPPLYKRAPGRPKKL 416
I P+NG LWP + +P PP Y + KK+
Sbjct: 668 ILPMNGLDLWPKSLNPPPFPPSYLNNKKKGKKM 700
>UniRef100_Q94HS2 Putative transposon protein [Oryza sativa]
Length = 841
Score = 168 bits (426), Expect = 3e-40
Identities = 86/275 (31%), Positives = 143/275 (51%), Gaps = 11/275 (4%)
Query: 181 GVVIRNLMMAAAKATYHQLWTEKMMELQALHPEAYTWLMQIPTRAWCKHAFTFYPKCDVL 240
G ++N + A A+++ W M E+++L+ +AY WL ++P + W K F+ +PKCD+L
Sbjct: 433 GENLKNQLWACARSSSEVEWNANMEEMKSLNQDAYEWLQKMPPKTWVKAYFSEFPKCDIL 492
Query: 241 MNNLSESFNATILLVRDKPVLTMVDWIRSYIMGR-FATMNEKLEKYKGEVMPKPLKRLEH 299
+NN E FN IL R+ P+L+M + I+S ++ R ++ E E++ G + PK K++
Sbjct: 493 LNNNCEVFNKYILEARELPILSMFEKIKSQLISRHYSKQKEVAEQWHGPICPKIRKKVLK 552
Query: 300 EIEMVRYWVATRCGEDKYEVKHMVSQDQFVVNLAEHTCSCNFWELVGIPCRHAVAAISHS 359
+M G+ ++V+ +++V+L+ C C W+L GIPC HA++ +
Sbjct: 553 NADMANTCYVLPAGKGIFQVEDR--NFKYIVDLSAKHCDCRRWDLTGIPCNHAISCLRSE 610
Query: 360 SQRPEDFVHPYYRREAYMATYGHVITPINGQKLWPTTNDPYILPPLYKRAPGRPKKLRRR 419
E + P Y EA+ Y I P N W N P + PP+Y++ GRP K RR+
Sbjct: 611 RISAESILPPCYSLEAFSRAYAFNIWPYNDMTKWVQVNGPEVKPPIYEKKVGRPPKSRRK 670
Query: 420 DPHEDTTDNQTRLR--------SRCGQFGHNSRSC 446
PHE N ++ S C + HN + C
Sbjct: 671 APHEVQGKNGPKMSKHGVEMHCSFCKEPRHNKKGC 705
>UniRef100_Q75LI5 Putative transposon protein [Oryza sativa]
Length = 839
Score = 163 bits (413), Expect = 8e-39
Identities = 94/308 (30%), Positives = 155/308 (49%), Gaps = 12/308 (3%)
Query: 148 QGLLTVFDDLMGGVEHIFCLRHLYANFKKKFGGGVVIRNLMMAAAKATYHQLWTEKMMEL 207
+GLL+V + L EH C RH+YAN++K+ + A +++ L+ +L
Sbjct: 437 KGLLSVIEHLFPKAEHRMCARHIYANWRKRHRLQEYQKRFWKIA-RSSNAVLFNHYKSKL 495
Query: 208 QALHPEAYTWLMQIPTRAWCKHAFTFYPKCDVLMNNLSESFNATILLVRDKPVLTMVDWI 267
P + L + WC+ F CD + NN+ ESFN I+ R KP++TM++ I
Sbjct: 496 ANKTPMGWEDLEKTNPIHWCRAWFKLGSNCDSVENNICESFNNWIIEARFKPIITMLEDI 555
Query: 268 RSYIMGRFATMNEKLEKYKGEVMPKPLKRLEHEIEMVRYWVATRCGEDKYEVKHMVSQDQ 327
R + R E++ + P LK++ ++ G +EV+ + +
Sbjct: 556 RMKVTRRIQENKTNSERWTMGICPNILKKINKIRHATQFCHVLWNGSSGFEVRE--KKWR 613
Query: 328 FVVNLAEHTCSCNFWELVGIPCRHAVAAISHSSQRPEDFVHPYYRREAYMATYGHVITPI 387
F V+L+ +TCSC +W++ GIPC+HA AA ++ P + V+ + + Y TY V+ P+
Sbjct: 614 FTVDLSANTCSCRYWQISGIPCQHACAAYFKMAEEPNNHVNMCFSIDQYRNTYQDVLQPV 673
Query: 388 NGQKLWPTTNDPYILPPLYKRAPGRPKKLRRRDPHEDTTDNQTRLRSR--------CGQF 439
+ +WP + +P LPP K+ PG PK+ RR+DP E + T+ R C +
Sbjct: 674 EHESVWPLSTNPRPLPPRVKKMPGSPKRARRKDPTE-AAGSSTKSSKRGGSVKCGFCHEK 732
Query: 440 GHNSRSCR 447
GHNSR C+
Sbjct: 733 GHNSRGCK 740
>UniRef100_Q6L3K3 Putative transposon MuDR mudrA-like protein [Solanum demissum]
Length = 873
Score = 162 bits (410), Expect = 2e-38
Identities = 100/309 (32%), Positives = 153/309 (49%), Gaps = 14/309 (4%)
Query: 148 QGLLTVFDDLMGGVEHIFCLRHLYANFKKKFGGGVVIRNLMMAAAKATYHQLWTEKMMEL 207
+GLL + H +C RH+ AN+ K + G V +R L+ +A +TY + + +++ +
Sbjct: 409 KGLLDAVSQVFPKAHHRWCARHIEANWSKAWKG-VQMRKLLWWSAWSTYEEEFHDQLKVM 467
Query: 208 QALHPEAYTWLMQIPTRAWCKHAFTFYPKCDVLMNNLSESFNATILLVRDKPVLTMVDWI 267
A+ +A L+ P + WC+ F K NN +ESFN IL R KP++ M++ I
Sbjct: 468 GAVSKQAAKDLVWYPAQNWCRAYFDTVCKNHSCENNFTESFNKWILEARAKPIIKMLENI 527
Query: 268 RSYIMGRFATMNEKLEKYKGEVMPKPLKRLEHEIEMVRYWVATRC-GEDKYEVKHMVSQD 326
R IM R + E+ + +KG+ P ++ L ++ ++ + G+ YEV + +D
Sbjct: 528 RIKIMNRLQKLEEEGKNWKGDFSPYAME-LYNDFNIIAQCCQVQSNGDQGYEV--VEGED 584
Query: 327 QFVVNLAEHTCSCNFWELVGIPCRHAVAAISHSSQRPEDFVHPYYRREAYMATYGHVITP 386
+ VVNL C+C W+L GI C HA+ A H Q P D + +Y REAYM Y H I P
Sbjct: 585 RHVVNLNRKKCTCRTWDLTGILCPHAIKAYLHDKQEPLDQLSWWYSREAYMLVYMHKIQP 644
Query: 387 INGQKLWPTTNDPYILPPLYKRAPGRPKKLRRRDPHEDTTDNQTRLRSR---------CG 437
+ G+K W + PP + GRPK R+R+ E SR C
Sbjct: 645 VRGEKFWKVDPSHAMEPPEIHKLVGRPKLKRKREKDEARKREGVWSASRKGLKMTCGHCS 704
Query: 438 QFGHNSRSC 446
GHN R C
Sbjct: 705 ATGHNQRRC 713
>UniRef100_Q8LMJ1 Putative transposon protein [Oryza sativa]
Length = 835
Score = 157 bits (397), Expect = 6e-37
Identities = 95/296 (32%), Positives = 142/296 (47%), Gaps = 7/296 (2%)
Query: 160 GVEHIFCLRHLYANFKKKFGGGVVIRNLMMAAAKATYHQLWTEKMMELQALHPEAYTWLM 219
GVEH C+RHL NF+K+F G V RNL +A++A ++ E++ P A W+
Sbjct: 505 GVEHRECMRHLVKNFQKRFSGEVFERNLW-SASRAYRQDIYEGHYNEMKEACPRATEWID 563
Query: 220 QIPTRAWCKHAFTFYPKCDVLMNNLSESFNATILLVRDKPVLTMVDWIRSYIMGRFATMN 279
W + F+ KCD + NN++E+FN+ I + PV+ ++D IR IM R +
Sbjct: 564 DFHKHIWTRSKFSPVSKCDYVTNNIAETFNSWIRHEKSLPVVDLMDKIRHMIMERISIRK 623
Query: 280 EKLEKYKGEVMPKPLKRL---EHEIEMVRYWVATRCGEDKYEVKHMVSQDQFVVNLAEHT 336
EK G+++P +K L ++ Y GE + + + + V+L
Sbjct: 624 RLAEKLTGQILPSVMKTLYARSRDLGYKLYSAHKHLGEIGGTGRDLKTW-RHTVDLNTRE 682
Query: 337 CSCNFWELVGIPCRHAV-AAISHSSQRPEDFVHPYYRREAYMATYGHVITPINGQKLWPT 395
CSC W++ GIPC H + IS E FV YY A+ Y + P+ + W
Sbjct: 683 CSCRQWQITGIPCTHVIFLIISRRGLELEQFVDEYYYVAAFKRAYAGHVVPMTDKAQWAK 742
Query: 396 TNDPYIL-PPLYKRAPGRPKKLRRRDPHEDTTDNQTRLRSRCGQFGHNSRSCRNPV 450
N L PPL KR+ GRP+ R + E + RCGQFGH ++C PV
Sbjct: 743 GNIGLKLHPPLLKRSAGRPRSRRIKGVEEGGVGKRKYRCKRCGQFGHTKKTCNEPV 798
>UniRef100_Q9FH72 Arabidopsis thaliana genomic DNA, chromosome 5, P1 clone:MRD20
[Arabidopsis thaliana]
Length = 733
Score = 155 bits (393), Expect = 2e-36
Identities = 87/313 (27%), Positives = 169/313 (53%), Gaps = 11/313 (3%)
Query: 148 QGLLTVFDDLMGGVEHIFCLRHLYANFKKKFGGGVVIRNLMMAAAKATYHQLWTEKMMEL 207
+GL+ +++ EH C RH++AN +K++ + + A+A ++ +++ ++
Sbjct: 365 KGLIYAIKNVLPYAEHRMCARHIFANLQKRYKQMGPLHKVFWKCARAYNETVFWKQLEKM 424
Query: 208 QALHPEAYTWLMQIPTRAWCKHAFTFYPKCDVLMNNLSESFNATILLVRDKPVLTMVDWI 267
+ + EAY + + W + F+ K + NN+SES+NA + R+KPV+ +++ I
Sbjct: 425 KTIKFEAYDEVKRSVGSNWSRAFFSDITKSAAVENNISESYNAVLKDAREKPVVALLEDI 484
Query: 268 RSYIMGRFATMNEKLEKYKGEVMPKPLKRLEHEIEMVRYWVATRCGEDKYEVKHMVSQDQ 327
R +IM ++++ G + PK + +E + +++ G YEV H +++
Sbjct: 485 RRHIMASNLVKIKEMQNVTGLITPKAIAIMEKRKKSLKWCYPFSNGRGIYEVDH--GKNK 542
Query: 328 FVVNLAEHT-CSCNFWELVGIPCRHAVAAI---SHSSQRPEDFVHPYYRREAYMATYGHV 383
+VV++ + T C+C +++ GIPC H ++A+ ++ PE + +Y E + Y +
Sbjct: 543 YVVHVRDKTSCTCREYDVSGIPCCHIMSAMWAEYKETKLPETAILDWYSVEKWKLCYNSL 602
Query: 384 ITPINGQKLWPTTNDPYILPPLYKRAPGRPKKLRR-RDPHEDTTDNQTRLR----SRCGQ 438
+ P+NG +LW T +D ++PP + PGRPKK R +DP E+ + ++ S CGQ
Sbjct: 603 LFPVNGMELWETHSDVVVMPPPDRIMPGRPKKNDRIKDPSEEASSENSQKALVTCSNCGQ 662
Query: 439 FGHNSRSCRNPVV 451
GHN R+C+ +V
Sbjct: 663 IGHNKRTCQIELV 675
>UniRef100_Q9LMY1 F21F23.10 protein [Arabidopsis thaliana]
Length = 753
Score = 155 bits (391), Expect = 3e-36
Identities = 87/313 (27%), Positives = 169/313 (53%), Gaps = 11/313 (3%)
Query: 148 QGLLTVFDDLMGGVEHIFCLRHLYANFKKKFGGGVVIRNLMMAAAKATYHQLWTEKMMEL 207
+GL+ +++ EH C RH++AN +K++ + + A+A ++ +++ ++
Sbjct: 385 KGLIYAIKNVLPYAEHRMCARHIFANLQKRYKQMGPLHKVFWKCARAYNETVFWKQLEKM 444
Query: 208 QALHPEAYTWLMQIPTRAWCKHAFTFYPKCDVLMNNLSESFNATILLVRDKPVLTMVDWI 267
+ + EAY + + W + F+ K + NN+SES+NA + R+KPV+ +++ I
Sbjct: 445 KTIKFEAYDEVKRSVGSNWSRAFFSDITKSAAVENNISESYNAVLKDAREKPVVALLEDI 504
Query: 268 RSYIMGRFATMNEKLEKYKGEVMPKPLKRLEHEIEMVRYWVATRCGEDKYEVKHMVSQDQ 327
R +IM ++++ G + PK + +E + +++ G YEV H +++
Sbjct: 505 RRHIMASNLVKIKEMQNVTGLITPKAIAIMEKRKKSLKWCYPFSNGRGIYEVDH--GKNK 562
Query: 328 FVVNLAEHT-CSCNFWELVGIPCRHAVAAI---SHSSQRPEDFVHPYYRREAYMATYGHV 383
+VV++ + T C+C +++ GIPC H ++A+ ++ PE + +Y E + Y +
Sbjct: 563 YVVHVRDKTSCTCREYDVSGIPCCHIMSAMWAEYKETKLPETAILDWYSVEKWKLCYNSL 622
Query: 384 ITPINGQKLWPTTNDPYILPPLYKRAPGRPKKLRR-RDPHEDTTDNQTRLR----SRCGQ 438
+ P+NG +LW T +D ++PP + PGRPKK R +DP E+ + ++ S CGQ
Sbjct: 623 LFPVNGMELWETHSDVVVMPPPDRIMPGRPKKNDRIKDPSEEASSENSQKALVTCSNCGQ 682
Query: 439 FGHNSRSCRNPVV 451
GHN R+C+ +V
Sbjct: 683 TGHNKRTCQIELV 695
>UniRef100_Q9FM95 Similarity to mutator-like transposase [Arabidopsis thaliana]
Length = 733
Score = 155 bits (391), Expect = 3e-36
Identities = 87/313 (27%), Positives = 169/313 (53%), Gaps = 11/313 (3%)
Query: 148 QGLLTVFDDLMGGVEHIFCLRHLYANFKKKFGGGVVIRNLMMAAAKATYHQLWTEKMMEL 207
+GL+ +++ EH C RH++AN +K++ + + A+A ++ +++ ++
Sbjct: 365 KGLIYAIKNVLPYAEHRMCARHIFANLQKRYKQMGPLHKVFWKCARAYNETVFWKQLEKM 424
Query: 208 QALHPEAYTWLMQIPTRAWCKHAFTFYPKCDVLMNNLSESFNATILLVRDKPVLTMVDWI 267
+ + EAY + + W + F+ K + NN+SES+NA + R+KPV+ +++ I
Sbjct: 425 KTIKFEAYDEVKRSVGSNWSRAFFSDITKSAAVENNISESYNAVLKDAREKPVVALLEDI 484
Query: 268 RSYIMGRFATMNEKLEKYKGEVMPKPLKRLEHEIEMVRYWVATRCGEDKYEVKHMVSQDQ 327
R +IM ++++ G + PK + +E + +++ G YEV H +++
Sbjct: 485 RRHIMASNLVKLKEMQNVTGLITPKAIAIMEKRKKSLKWCYPFSNGRGIYEVDH--GKNK 542
Query: 328 FVVNLAEHT-CSCNFWELVGIPCRHAVAAI---SHSSQRPEDFVHPYYRREAYMATYGHV 383
+VV++ + T C+C +++ GIPC H ++A+ ++ PE + +Y E + Y +
Sbjct: 543 YVVHVRDKTSCTCREYDVSGIPCCHIMSAMWAEYKETKLPETAILDWYSVEKWKLCYNSL 602
Query: 384 ITPINGQKLWPTTNDPYILPPLYKRAPGRPKKLRR-RDPHEDTTDNQTRLR----SRCGQ 438
+ P+NG +LW T +D ++PP + PGRPKK R +DP E+ + ++ S CGQ
Sbjct: 603 LFPVNGMELWETHSDVVVMPPPDRIMPGRPKKNDRIKDPSEEASSENSQKALVTCSNCGQ 662
Query: 439 FGHNSRSCRNPVV 451
GHN R+C+ +V
Sbjct: 663 TGHNKRTCQIELV 675
>UniRef100_Q60CW5 Putative mutator transposable element-related protein [Solanum
tuberosum]
Length = 493
Score = 152 bits (385), Expect = 1e-35
Identities = 88/280 (31%), Positives = 140/280 (49%), Gaps = 12/280 (4%)
Query: 183 VIRNLMMAAAKATYHQLWTEKMMELQALHPEAYTWLMQIPTRAWCKHAFTFYPKCDVLMN 242
V + AAAKAT + + M+ ++ L P A WL W + + KCD+L+N
Sbjct: 161 VDKTAFWAAAKATTVKEFDACMVRIRELDPNAVDWLNDKEPSQWSRSHLSSDAKCDILLN 220
Query: 243 NLSESFNATILLVRDKPVLTMVDWIRSYIMGRFATMNEKLEKYKG-EVMPKPLKRLEHEI 301
N+ E FN+ I RDKP++T+++ +R +M R EK K+ +V PK L
Sbjct: 221 NICEVFNSMIFDARDKPIVTLLEKLRYLLMARMLANREKAHKWSSNDVCPKIKDILHKNQ 280
Query: 302 EMVRYWVATRCGEDKYEVKHMVSQDQFVVNLAEHTCSCNFWELVGIPCRHAVAAISHSSQ 361
++ + + KYE+ D + V+L CSC W ++GIPC+HA+AAI
Sbjct: 281 TAAGEYIPRKSNQRKYEIIGATIHDSWAVDLENRICSCTKWSIMGIPCKHAIAAIRAKKD 340
Query: 362 RPEDFVHPYYRREAYMATYGHVITPINGQKLWP--TTNDPYILPPLYKRAPGRPKKLRRR 419
D+V Y+ E Y Y H I ING ++WP T P L + + GR +K RR+
Sbjct: 341 NILDYVDDCYKVETYRRIYEHAILSINGPQMWPKSTKVPPLPLTIVGNKKTGRKQKARRK 400
Query: 420 DPHEDTTDNQTRLR--------SRCGQFGHNSRSCRNPVV 451
+ ++ ++T+++ S C + GHN ++C+ VV
Sbjct: 401 EA-DEVGASRTKIKRKQQSLDCSTCNKPGHNKKTCKYNVV 439
>UniRef100_Q9LP09 F9C16.9 [Arabidopsis thaliana]
Length = 946
Score = 145 bits (365), Expect = 3e-33
Identities = 100/323 (30%), Positives = 152/323 (46%), Gaps = 29/323 (8%)
Query: 149 GLLTVFDDLMGGVEHIFCLRHLYANFKKKFGGGVVIRNLMMAAAKATYHQLWTEKMMELQ 208
GL+ ++ EH C RH+ N+K+ + ++ L ++ + M L+
Sbjct: 508 GLVKAIHSVIPQAEHRQCARHIMENWKRN-SHDMELQRLFWKIVRSYTEGEFGAHMRALK 566
Query: 209 ALHPEAYTWLMQIPTRAWCKHAFTFYPKCDVLMNNLSESFNATILLVRDKPVLTMVDWIR 268
+ + A+ L++ W + F C+ +NNLSESFN TI R KP+L M++ IR
Sbjct: 567 SYNASAFELLLKTLPVTWSRAFFKIGSCCNDNLNNLSESFNRTIREARRKPLLDMLEDIR 626
Query: 269 SYIMGRFATMNEKLEKYKGEVMPKPLKRLEHEIEMV-------RYWVATRCGEDKYEVKH 321
M R NEK + + KR EIE + + W+A +K+E+K
Sbjct: 627 RQCMVR----NEKRYVIADRLRTRFTKRAHAEIEKMIAGSQVCQRWMARH---NKHEIK- 678
Query: 322 MVSQDQFVVNLAEHTCSCNFWELVGIPCRHAVAAISHSSQRPEDFVHPYYRREAYMATYG 381
+ D F V++ +TC C W++ GIPC HA + I Q+ ED+V +Y + TY
Sbjct: 679 VGPVDSFTVDMNNNTCGCMKWQMTGIPCIHAASVIIGKRQKVEDYVSDWYTTSMWKQTYN 738
Query: 382 HVITPINGQKLWPTTNDPYILPPLYKRA-PGRPKKLRRR-------DPHEDTTDNQTRLR 433
I P+ G+ LWPT N +LPP ++R PGRPK RR +T +RL
Sbjct: 739 DGIGPVQGKLLWPTVNKVGVLPPPWRRGNPGRPKNHARRKGVFESSTASSSSTTELSRLH 798
Query: 434 -----SRCGQFGHNSRSCRNPVV 451
S C GHN + C+N V
Sbjct: 799 RVMTCSNCQGEGHNKQGCKNETV 821
>UniRef100_O49511 MuDR transposable element - like protein [Arabidopsis thaliana]
Length = 633
Score = 140 bits (353), Expect = 7e-32
Identities = 88/315 (27%), Positives = 148/315 (46%), Gaps = 13/315 (4%)
Query: 148 QGLLTVFDDLMGGVEHIFCLRHLYANFKKKFGGGVVIRNLMMAAAKATYHQLWTEKMMEL 207
+GL+ + +EH C++H+Y N KK +G +I+ L+ A + + + + ++
Sbjct: 303 KGLIKAVQLELPKIEHRMCVQHIYGNLKKTYGSKTMIKPLLWNLAWSYNETEYKQHLEKI 362
Query: 208 QALHPEAYTWLMQIPTRAWCKHAFTFYPKCDVLMNNLSESFNATILLVRDKPVLTMVDWI 267
+ + Y +M+ R+W + C+ + NN ESFN ++ R+K + M++ I
Sbjct: 363 RCYDTKVYESVMKTNPRSWVRAFQKIGSFCEDVDNNSVESFNGSLNKAREKQFVAMLETI 422
Query: 268 RSYIMGRFATMNEKLEKYKGEVMPKPLKRLEHEIEMVRYWVATRCGEDKYEVKHMVSQDQ 327
R M R A + + + G P +K L E ++ YEV+H D
Sbjct: 423 RRMAMVRIAKRSVESHTHTGVCTPYVMKFLAGEHKVASTAKVAPGTNGMYEVRH--GGDT 480
Query: 328 FVVNLAEHTCSCNFWELVGIPCRHAVAAISHSSQRPEDFVHPYYRREAYMATYGHVITPI 387
V+LA +TC+C W++ GIPC HA I H +PEDFV ++R + Y + P
Sbjct: 481 HRVDLAAYTCTCIKWQICGIPCEHAYGVILHKKLQPEDFVCQWFRTAMWKKNYTDGLFPQ 540
Query: 388 NGQKLWPTTNDPYIL---PPLYK--RAPGRPKKLRRRDPHEDTTDNQTRLRSR------C 436
G K WP +N P + PP + + + +K R++ +E T Q + + R C
Sbjct: 541 RGPKFWPESNGPRVFAAEPPEGEEDKKMTKAEKKRKKGVNESPTKKQPKAKKRIMHCGVC 600
Query: 437 GQFGHNSRSCRNPVV 451
G HNSR +N V
Sbjct: 601 GAADHNSRHHKNDKV 615
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.341 0.149 0.525
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 733,453,354
Number of Sequences: 2790947
Number of extensions: 29644859
Number of successful extensions: 99503
Number of sequences better than 10.0: 259
Number of HSP's better than 10.0 without gapping: 183
Number of HSP's successfully gapped in prelim test: 76
Number of HSP's that attempted gapping in prelim test: 98770
Number of HSP's gapped (non-prelim): 338
length of query: 451
length of database: 848,049,833
effective HSP length: 131
effective length of query: 320
effective length of database: 482,435,776
effective search space: 154379448320
effective search space used: 154379448320
T: 11
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.9 bits)
S2: 76 (33.9 bits)
Lotus: description of TM0213.7


