

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0213.11
(579 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
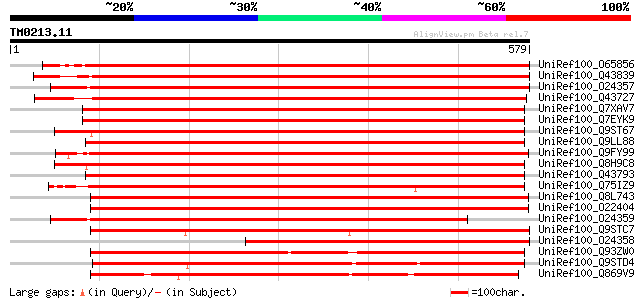
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_O65856 Glucose-6-phosphate dehydrogenase precursor [Ni... 893 0.0
UniRef100_Q43839 Glucose-6-phosphate 1-dehydrogenase, chloroplas... 892 0.0
UniRef100_O24357 Glucose-6-phosphate 1-dehydrogenase, chloroplas... 863 0.0
UniRef100_Q43727 Glucose-6-phosphate 1-dehydrogenase 1, chloropl... 855 0.0
UniRef100_Q7XAV7 Putative plastidic glucose-6-phosphate dehydrog... 833 0.0
UniRef100_Q7EYK9 Putative plastidic glucose 6-phosphate dehydrog... 833 0.0
UniRef100_Q9ST67 Glucose-6-phosphate 1-dehydrogenase precursor [... 833 0.0
UniRef100_Q9LL88 Plastidic glucose 6-phosphate dehydrogenase [Ni... 832 0.0
UniRef100_Q9FY99 Glucose-6-phosphate 1-dehydrogenase 2, chloropl... 832 0.0
UniRef100_Q8H9C8 Glucose 6-phosphate dehydrogenase [Solanum tube... 832 0.0
UniRef100_Q43793 Glucose-6-phosphate 1-dehydrogenase, chloroplas... 830 0.0
UniRef100_Q75IZ9 Putative glucose-6-phosphate dehydrogenase [Ory... 827 0.0
UniRef100_Q8L743 Glucose-6-phosphate 1-dehydrogenase 3, chloropl... 820 0.0
UniRef100_O22404 Plastidic glucose-6-phosphate dehydrogenase [Pe... 820 0.0
UniRef100_O24359 Glucose-6-phosphate dehydrogenase [Spinacia ole... 733 0.0
UniRef100_Q9STC7 Plastidic glucose-6-phosphate dehydrogenase [Du... 696 0.0
UniRef100_O24358 Glucose-6-phosphate dehydrogenase [Spinacia ole... 573 e-162
UniRef100_Q93ZW0 Glucose-6-phosphate 1-dehydrogenase 4, chloropl... 538 e-151
UniRef100_Q9STD4 Glucose-6-phosphate 1-dehydrogenase precursor [... 513 e-144
UniRef100_Q869V9 Similar to Oryza sativa (Rice). glucose-6-phosp... 482 e-134
>UniRef100_O65856 Glucose-6-phosphate dehydrogenase precursor [Nicotiana tabacum]
Length = 588
Score = 893 bits (2308), Expect = 0.0
Identities = 444/543 (81%), Positives = 485/543 (88%), Gaps = 12/543 (2%)
Query: 37 PRNHFQLKSSNGHPLNAVSSHSHDGLAGSSLTKEDSKPQPVDGPFLSSDSECTGSNLSIT 96
PR HF++ SSNG LNAVS L GS+ SK P P ++ E + +SIT
Sbjct: 58 PRKHFEIMSSNGFHLNAVSL-----LDGSA-----SKSMPEQVPL--TELENAETTVSIT 105
Query: 97 VVGASGDLAKKKIFPALFALFYEDCLPENFLVFGFARTKMTSEELRNMISRTLTCRIDKR 156
V+GASGDLAKKKIF ALFALFYEDCLPENF+VFG++RTKM+ EELRNMIS+TLTCRID+R
Sbjct: 106 VIGASGDLAKKKIFTALFALFYEDCLPENFIVFGYSRTKMSDEELRNMISKTLTCRIDQR 165
Query: 157 ANCEDKMDQFLKRCFYHSGLYNSEGSFSDLDCKMKEKEGGKRSNRLFYLSIPPNIFVNVV 216
NCE KMD FL+RCFYHSG Y+SE F++LD K+K KEG + SNRLFYLSIPPNIFV+VV
Sbjct: 166 ENCEAKMDHFLERCFYHSGQYHSEDDFAELDYKLKAKEGSRVSNRLFYLSIPPNIFVDVV 225
Query: 217 RCASLKASSKKGWTRVIVEKPFGRDSESSNELTRCLKQYLTEDQIFRIDHYLGKELVENL 276
RCASLKASS GWTRVIVEKPFGRD ESS+ELTRCLK+YLTE+QIFRIDHYLGKELVENL
Sbjct: 226 RCASLKASSTSGWTRVIVEKPFGRDLESSSELTRCLKKYLTEEQIFRIDHYLGKELVENL 285
Query: 277 SVLRFSNLVFEPLWSRAYIRNVQFIFSEDFGTEGRGGYFDNYGIIRDIMQNHLLQILALF 336
SVLRFSNLVFEPLWSR YIRNVQFIFSED GTEGRGGYFDNYGIIRDIMQNHLLQILALF
Sbjct: 286 SVLRFSNLVFEPLWSRNYIRNVQFIFSEDSGTEGRGGYFDNYGIIRDIMQNHLLQILALF 345
Query: 337 AMEPPVSLDAEDIRNEKVKVLRSMRPLQLENVVTGQYKGHSKGGKSYPGYADDPSVPKGS 396
AME PVS+DAEDIRNEKVKVLRSMRPLQLE+VV GQYKGHSKGGK YP Y DDP+VP GS
Sbjct: 346 AMETPVSMDAEDIRNEKVKVLRSMRPLQLEDVVLGQYKGHSKGGKLYPAYTDDPTVPNGS 405
Query: 397 LTPTFAAAALFIDNARWDGVPFLMKAGKALHTKRAEIRVQFRHVPGNLYKRNFGADMDKA 456
+TPTF+AAALFI+NARWDGVPFLMKAGKALHT+RAEIRVQFRHVPGNLYKRNFG D+DKA
Sbjct: 406 VTPTFSAAALFINNARWDGVPFLMKAGKALHTRRAEIRVQFRHVPGNLYKRNFGTDLDKA 465
Query: 457 TNELVLRVQPDEAIYLKVNNKIPGLGMRLDRSDLNLLYRARYPTEIPDAYERLLLDAIEG 516
TNELVLR+QPDEAIYLK+NNK+PGLGMRLDRSDLNLLY+A+Y EIPDAYERLLLDAIEG
Sbjct: 466 TNELVLRLQPDEAIYLKINNKVPGLGMRLDRSDLNLLYKAKYRGEIPDAYERLLLDAIEG 525
Query: 517 ERRLFIRSDELDAAWALFTPLLKEVEDKKIAPELYPHGSRGPIGAHYLAAKHNVRWGDTG 576
ERRLFIRSDELDAAWALFTPLLKE+E+KKIAPELYP+GSRGP+GAHYLAAKHNVRWGD
Sbjct: 526 ERRLFIRSDELDAAWALFTPLLKELEEKKIAPELYPYGSRGPVGAHYLAAKHNVRWGDLS 585
Query: 577 NDD 579
DD
Sbjct: 586 GDD 588
>UniRef100_Q43839 Glucose-6-phosphate 1-dehydrogenase, chloroplast precursor [Solanum
tuberosum]
Length = 577
Score = 892 bits (2304), Expect = 0.0
Identities = 447/553 (80%), Positives = 481/553 (86%), Gaps = 20/553 (3%)
Query: 27 WFSCIRPGNLPRNHFQLKSSNGHPLNAVSSHSHDGLAGSSLTKEDSKPQPVDGPFLSSDS 86
W S I PR HF++ SSNG PLNAVS Q V P S
Sbjct: 45 WVSGIYSRIQPRKHFEVFSSNGFPLNAVSV------------------QDVQVPLTELGS 86
Query: 87 ECTGSNLSITVVGASGDLAKKKIFPALFALFYEDCLPENFLVFGFARTKMTSEELRNMIS 146
T +SITV+GASGDLAKKKI PALFALFYEDCLPENF+VFG++RTK++ EELRNMIS
Sbjct: 87 GDT--TVSITVIGASGDLAKKKILPALFALFYEDCLPENFVVFGYSRTKLSDEELRNMIS 144
Query: 147 RTLTCRIDKRANCEDKMDQFLKRCFYHSGLYNSEGSFSDLDCKMKEKEGGKRSNRLFYLS 206
TLTCRIDKR NC+ KM+ FL+RCFYHSG YNSE F++LD K+KEKEG + SNRLFYLS
Sbjct: 145 TTLTCRIDKRENCDAKMEHFLERCFYHSGQYNSEDDFAELDYKLKEKEGCRVSNRLFYLS 204
Query: 207 IPPNIFVNVVRCASLKASSKKGWTRVIVEKPFGRDSESSNELTRCLKQYLTEDQIFRIDH 266
IPPNIFV+VVRCASLKASS GWTRVIVEKPFGRD ESS+ELTR LK+YLTE+QIFRIDH
Sbjct: 205 IPPNIFVDVVRCASLKASSTSGWTRVIVEKPFGRDLESSSELTRSLKKYLTEEQIFRIDH 264
Query: 267 YLGKELVENLSVLRFSNLVFEPLWSRAYIRNVQFIFSEDFGTEGRGGYFDNYGIIRDIMQ 326
YLGKELVENLSVLRFSNLVFEPLWSR YIRNVQFIFSEDFGTEGRGGYFD+YGIIRDIMQ
Sbjct: 265 YLGKELVENLSVLRFSNLVFEPLWSRNYIRNVQFIFSEDFGTEGRGGYFDHYGIIRDIMQ 324
Query: 327 NHLLQILALFAMEPPVSLDAEDIRNEKVKVLRSMRPLQLENVVTGQYKGHSKGGKSYPGY 386
NHLLQILALFAME PVSLDAEDIRNEKVKVLRSMRPLQLE+VV GQYKGHS G KSYP Y
Sbjct: 325 NHLLQILALFAMETPVSLDAEDIRNEKVKVLRSMRPLQLEDVVLGQYKGHSNGAKSYPAY 384
Query: 387 ADDPSVPKGSLTPTFAAAALFIDNARWDGVPFLMKAGKALHTKRAEIRVQFRHVPGNLYK 446
DDP+VP GS+TPTF+AAALFIDNARWDGVPFLMKAGKALHTKRAEIRVQFRHVPGNLYK
Sbjct: 385 TDDPTVPNGSITPTFSAAALFIDNARWDGVPFLMKAGKALHTKRAEIRVQFRHVPGNLYK 444
Query: 447 RNFGADMDKATNELVLRVQPDEAIYLKVNNKIPGLGMRLDRSDLNLLYRARYPTEIPDAY 506
RNFG DMDKATNELVLR+QPDEAIYLK+NNK+PGLGMRLDRSDLNLLY+A+Y EIPDAY
Sbjct: 445 RNFGTDMDKATNELVLRLQPDEAIYLKINNKVPGLGMRLDRSDLNLLYKAKYRGEIPDAY 504
Query: 507 ERLLLDAIEGERRLFIRSDELDAAWALFTPLLKEVEDKKIAPELYPHGSRGPIGAHYLAA 566
ERLLLDAIEGERRLFIRSDELDAAWALFTPLLKE+E+KKIAPELYP+GSRGP+GAHYLAA
Sbjct: 505 ERLLLDAIEGERRLFIRSDELDAAWALFTPLLKELEEKKIAPELYPYGSRGPVGAHYLAA 564
Query: 567 KHNVRWGDTGNDD 579
KHNVRWGD DD
Sbjct: 565 KHNVRWGDLSGDD 577
>UniRef100_O24357 Glucose-6-phosphate 1-dehydrogenase, chloroplast precursor
[Spinacia oleracea]
Length = 574
Score = 863 bits (2229), Expect = 0.0
Identities = 428/534 (80%), Positives = 462/534 (86%), Gaps = 2/534 (0%)
Query: 46 SNGHPLNAVSSHSHDGLAGSSLTKEDSKPQPVDGPFLSSDSECTGSNLSITVVGASGDLA 105
SNGHPLN VS + + D P L S E LSI VVGASGDLA
Sbjct: 42 SNGHPLNDVSLQNDVAVNPIVAKSIDPSADLQLLPLLESVKE--EPTLSIIVVGASGDLA 99
Query: 106 KKKIFPALFALFYEDCLPENFLVFGFARTKMTSEELRNMISRTLTCRIDKRANCEDKMDQ 165
KKKIFPALFALFYE+CLPENF VFGF+RT+M EELR MIS+TLTCRID+R NC +KMD
Sbjct: 100 KKKIFPALFALFYENCLPENFTVFGFSRTEMNDEELRTMISKTLTCRIDQRENCGEKMDH 159
Query: 166 FLKRCFYHSGLYNSEGSFSDLDCKMKEKEGGKRSNRLFYLSIPPNIFVNVVRCASLKASS 225
FL+RCFYHSG YNSE FS LDCK+KEKE G+ NRLFYLSIPPNIFV+VVRC S +ASS
Sbjct: 160 FLQRCFYHSGQYNSEDDFSGLDCKLKEKEAGRLQNRLFYLSIPPNIFVDVVRCVSHRASS 219
Query: 226 KKGWTRVIVEKPFGRDSESSNELTRCLKQYLTEDQIFRIDHYLGKELVENLSVLRFSNLV 285
GWTRVIVEKPFGRDS+SS ELTR KQYL+EDQIFRIDHYLGKELVENLSVLRFSNLV
Sbjct: 220 ASGWTRVIVEKPFGRDSDSSRELTRSFKQYLSEDQIFRIDHYLGKELVENLSVLRFSNLV 279
Query: 286 FEPLWSRAYIRNVQFIFSEDFGTEGRGGYFDNYGIIRDIMQNHLLQILALFAMEPPVSLD 345
FEPLWSR YIRNVQ IFSEDFGTEGRGGYFDNYGIIRDIMQNHLLQILALFAME PVSLD
Sbjct: 280 FEPLWSRNYIRNVQLIFSEDFGTEGRGGYFDNYGIIRDIMQNHLLQILALFAMETPVSLD 339
Query: 346 AEDIRNEKVKVLRSMRPLQLENVVTGQYKGHSKGGKSYPGYADDPSVPKGSLTPTFAAAA 405
AEDIRNEKVKVLRSM+PL+L++VV GQYKGHSKG KSY GY DDP+VP S+TPTFAAAA
Sbjct: 340 AEDIRNEKVKVLRSMKPLKLQDVVVGQYKGHSKGNKSYSGYTDDPTVPNNSVTPTFAAAA 399
Query: 406 LFIDNARWDGVPFLMKAGKALHTKRAEIRVQFRHVPGNLYKRNFGADMDKATNELVLRVQ 465
LFIDNARWDGVPFLMKAGKALHTKRAEIRVQFRHVPGNLYK+ FG D+DKATNELVLRVQ
Sbjct: 400 LFIDNARWDGVPFLMKAGKALHTKRAEIRVQFRHVPGNLYKKTFGTDLDKATNELVLRVQ 459
Query: 466 PDEAIYLKVNNKIPGLGMRLDRSDLNLLYRARYPTEIPDAYERLLLDAIEGERRLFIRSD 525
PDEAIYLK+NNK+PGLGMRLDR+DLNL Y RY EIPDAYERLLLDAIEGERRLFIRSD
Sbjct: 460 PDEAIYLKINNKVPGLGMRLDRTDLNLCYSTRYRGEIPDAYERLLLDAIEGERRLFIRSD 519
Query: 526 ELDAAWALFTPLLKEVEDKKIAPELYPHGSRGPIGAHYLAAKHNVRWGDTGNDD 579
+LDAAW+LFTPLLKE+E+KK+APELYP+GSRGP+GAHYLAAKHNVRWGD +D
Sbjct: 520 KLDAAWSLFTPLLKELEEKKVAPELYPYGSRGPVGAHYLAAKHNVRWGDLSGED 573
>UniRef100_Q43727 Glucose-6-phosphate 1-dehydrogenase 1, chloroplast precursor
[Arabidopsis thaliana]
Length = 576
Score = 855 bits (2208), Expect = 0.0
Identities = 427/550 (77%), Positives = 472/550 (85%), Gaps = 20/550 (3%)
Query: 28 FSCIRPGNLPRNHFQLKSSNGHPLNAVS-SHSHDGLAGSSLTKEDSKPQPVDGPFLSSDS 86
FS +R H QL +SNG N S S D L +TK +S
Sbjct: 44 FSQVRLRFFAEKHSQLDTSNGCATNFASLQDSGDQLTEEHVTKGES-------------- 89
Query: 87 ECTGSNLSITVVGASGDLAKKKIFPALFALFYEDCLPENFLVFGFARTKMTSEELRNMIS 146
LSITVVGASGDLAKKKIFPALFALFYE CLP++F VFG+ARTK+T EELR+MIS
Sbjct: 90 -----TLSITVVGASGDLAKKKIFPALFALFYEGCLPQDFSVFGYARTKLTHEELRDMIS 144
Query: 147 RTLTCRIDKRANCEDKMDQFLKRCFYHSGLYNSEGSFSDLDCKMKEKEGGKRSNRLFYLS 206
TLTCRID+R C DKM+QFLKRCFYHSG YNSE F++L+ K+KEKE GK SNRL+YLS
Sbjct: 145 STLTCRIDQREKCGDKMEQFLKRCFYHSGQYNSEEDFAELNKKLKEKEAGKISNRLYYLS 204
Query: 207 IPPNIFVNVVRCASLKASSKKGWTRVIVEKPFGRDSESSNELTRCLKQYLTEDQIFRIDH 266
IPPNIFV+VVRCASL+ASS+ GWTRVIVEKPFGRDSESS ELTRCLKQYLTE+QIFRIDH
Sbjct: 205 IPPNIFVDVVRCASLRASSENGWTRVIVEKPFGRDSESSGELTRCLKQYLTEEQIFRIDH 264
Query: 267 YLGKELVENLSVLRFSNLVFEPLWSRAYIRNVQFIFSEDFGTEGRGGYFDNYGIIRDIMQ 326
YLGKELVENLSVLRFSNLVFEPLWSR YIRNVQ IFSEDFGTEGRGGYFD YGIIRDIMQ
Sbjct: 265 YLGKELVENLSVLRFSNLVFEPLWSRNYIRNVQLIFSEDFGTEGRGGYFDQYGIIRDIMQ 324
Query: 327 NHLLQILALFAMEPPVSLDAEDIRNEKVKVLRSMRPLQLENVVTGQYKGHSKGGKSYPGY 386
NHLLQILALFAME PVSLDAEDIR+EKVKVLRSM+PL+LE+VV GQYKGH+KGGK+YPGY
Sbjct: 325 NHLLQILALFAMETPVSLDAEDIRSEKVKVLRSMKPLRLEDVVVGQYKGHNKGGKTYPGY 384
Query: 387 ADDPSVPKGSLTPTFAAAALFIDNARWDGVPFLMKAGKALHTKRAEIRVQFRHVPGNLYK 446
DDP+VP SLTPTFAAAA+FI+NARWDGVPFLMKAGKALHT+ AEIRVQFRHVPGNLYK
Sbjct: 385 TDDPTVPNHSLTPTFAAAAMFINNARWDGVPFLMKAGKALHTRGAEIRVQFRHVPGNLYK 444
Query: 447 RNFGADMDKATNELVLRVQPDEAIYLKVNNKIPGLGMRLDRSDLNLLYRARYPTEIPDAY 506
++F ++D ATNELV+RVQPDE IYL++NNK+PGLGMRLDRSDLNLLYR+RYP EIPDAY
Sbjct: 445 KSFATNLDNATNELVIRVQPDEGIYLRINNKVPGLGMRLDRSDLNLLYRSRYPREIPDAY 504
Query: 507 ERLLLDAIEGERRLFIRSDELDAAWALFTPLLKEVEDKKIAPELYPHGSRGPIGAHYLAA 566
ERLLLDAIEGERRLFIRSDELDAAW LFTP LKE+E+KKI PELYP+GSRGP+GAHYLA+
Sbjct: 505 ERLLLDAIEGERRLFIRSDELDAAWDLFTPALKELEEKKIIPELYPYGSRGPVGAHYLAS 564
Query: 567 KHNVRWGDTG 576
K+NVRWGD G
Sbjct: 565 KYNVRWGDLG 574
>UniRef100_Q7XAV7 Putative plastidic glucose-6-phosphate dehydrogenase [Oryza sativa]
Length = 588
Score = 833 bits (2153), Expect = 0.0
Identities = 408/493 (82%), Positives = 449/493 (90%)
Query: 82 LSSDSECTGSNLSITVVGASGDLAKKKIFPALFALFYEDCLPENFLVFGFARTKMTSEEL 141
LS SE S +SITVVGASGDLAKKKIFPALFAL+YEDCLP++F +FG+AR+KMT EL
Sbjct: 89 LSPLSENDDSTVSITVVGASGDLAKKKIFPALFALYYEDCLPKHFTIFGYARSKMTDAEL 148
Query: 142 RNMISRTLTCRIDKRANCEDKMDQFLKRCFYHSGLYNSEGSFSDLDCKMKEKEGGKRSNR 201
RNM+S+TLTCRIDKR NC +KM++FLKRCFYHSG Y+SE F DLD K+K+ EG + SNR
Sbjct: 149 RNMVSKTLTCRIDKRENCNEKMEEFLKRCFYHSGQYDSEEHFMDLDKKLKQHEGSRVSNR 208
Query: 202 LFYLSIPPNIFVNVVRCASLKASSKKGWTRVIVEKPFGRDSESSNELTRCLKQYLTEDQI 261
LFYLSIPPNIF++VV+CAS ASS GWTRVIVEKPFGRDS+SS+ LTR LKQYL EDQI
Sbjct: 209 LFYLSIPPNIFLDVVKCASKSASSGNGWTRVIVEKPFGRDSDSSSALTRGLKQYLVEDQI 268
Query: 262 FRIDHYLGKELVENLSVLRFSNLVFEPLWSRAYIRNVQFIFSEDFGTEGRGGYFDNYGII 321
FRIDHYLGKELVENLSVLRFSNLVFEPLWSR YIRNVQ IFSEDFGTEGRGGYFD YGII
Sbjct: 269 FRIDHYLGKELVENLSVLRFSNLVFEPLWSRQYIRNVQLIFSEDFGTEGRGGYFDRYGII 328
Query: 322 RDIMQNHLLQILALFAMEPPVSLDAEDIRNEKVKVLRSMRPLQLENVVTGQYKGHSKGGK 381
RDIMQNHLLQILALFAME PVSL+AEDIRNEKVKVLRSM+PLQLE+VV GQYK H+KGG
Sbjct: 329 RDIMQNHLLQILALFAMETPVSLEAEDIRNEKVKVLRSMKPLQLEDVVIGQYKSHTKGGT 388
Query: 382 SYPGYADDPSVPKGSLTPTFAAAALFIDNARWDGVPFLMKAGKALHTKRAEIRVQFRHVP 441
+YPGY +D +VPK S+TPTFAAAALFI+NARWDGVPFLMKAGKALHTK AEIRVQFRHVP
Sbjct: 389 TYPGYTEDKTVPKDSVTPTFAAAALFINNARWDGVPFLMKAGKALHTKGAEIRVQFRHVP 448
Query: 442 GNLYKRNFGADMDKATNELVLRVQPDEAIYLKVNNKIPGLGMRLDRSDLNLLYRARYPTE 501
GNLYKR+FG D+D ATNELV+RVQPDEAIYLK+NNKIPGLGMRLDRS+LNL Y ARY E
Sbjct: 449 GNLYKRSFGTDLDTATNELVIRVQPDEAIYLKINNKIPGLGMRLDRSNLNLHYAARYSKE 508
Query: 502 IPDAYERLLLDAIEGERRLFIRSDELDAAWALFTPLLKEVEDKKIAPELYPHGSRGPIGA 561
IPDAYERLLLDAIEGERRLFIRSDELDAAW LFTPLLKE+E+K+IAPELYP+GSRGP+GA
Sbjct: 509 IPDAYERLLLDAIEGERRLFIRSDELDAAWELFTPLLKELEEKRIAPELYPYGSRGPVGA 568
Query: 562 HYLAAKHNVRWGD 574
HYLAAK+NVRWGD
Sbjct: 569 HYLAAKYNVRWGD 581
>UniRef100_Q7EYK9 Putative plastidic glucose 6-phosphate dehydrogenase [Oryza sativa]
Length = 588
Score = 833 bits (2153), Expect = 0.0
Identities = 408/493 (82%), Positives = 449/493 (90%)
Query: 82 LSSDSECTGSNLSITVVGASGDLAKKKIFPALFALFYEDCLPENFLVFGFARTKMTSEEL 141
LS SE S +SITVVGASGDLAKKKIFPALFAL+YEDCLP++F +FG+AR+KMT EL
Sbjct: 89 LSPLSENDDSTVSITVVGASGDLAKKKIFPALFALYYEDCLPKHFTIFGYARSKMTDAEL 148
Query: 142 RNMISRTLTCRIDKRANCEDKMDQFLKRCFYHSGLYNSEGSFSDLDCKMKEKEGGKRSNR 201
RNM+S+TLTCRIDKR NC +KM++FLKRCFYHSG Y+SE F DLD K+K+ EG + SNR
Sbjct: 149 RNMVSKTLTCRIDKRENCNEKMEEFLKRCFYHSGQYDSEEHFMDLDKKLKQHEGSRVSNR 208
Query: 202 LFYLSIPPNIFVNVVRCASLKASSKKGWTRVIVEKPFGRDSESSNELTRCLKQYLTEDQI 261
LFYLSIPPNIF++VV+CAS ASS GWTRVIVEKPFGRDS+SS+ LTR LKQYL EDQI
Sbjct: 209 LFYLSIPPNIFLDVVKCASKSASSGNGWTRVIVEKPFGRDSDSSSALTRGLKQYLVEDQI 268
Query: 262 FRIDHYLGKELVENLSVLRFSNLVFEPLWSRAYIRNVQFIFSEDFGTEGRGGYFDNYGII 321
FRIDHYLGKELVENLSVLRFSNLVFEPLWSR YIRNVQ IFSEDFGTEGRGGYFD YGII
Sbjct: 269 FRIDHYLGKELVENLSVLRFSNLVFEPLWSRQYIRNVQLIFSEDFGTEGRGGYFDRYGII 328
Query: 322 RDIMQNHLLQILALFAMEPPVSLDAEDIRNEKVKVLRSMRPLQLENVVTGQYKGHSKGGK 381
RDIMQNHLLQILALFAME PVSL+AEDIRNEKVKVLRSM+PLQLE+VV GQYK H+KGG
Sbjct: 329 RDIMQNHLLQILALFAMETPVSLEAEDIRNEKVKVLRSMKPLQLEDVVIGQYKSHTKGGT 388
Query: 382 SYPGYADDPSVPKGSLTPTFAAAALFIDNARWDGVPFLMKAGKALHTKRAEIRVQFRHVP 441
+YPGY +D +VPK S+TPTFAAAALFI+NARWDGVPFLMKAGKALHTK AEIRVQFRHVP
Sbjct: 389 TYPGYTEDKTVPKDSVTPTFAAAALFINNARWDGVPFLMKAGKALHTKGAEIRVQFRHVP 448
Query: 442 GNLYKRNFGADMDKATNELVLRVQPDEAIYLKVNNKIPGLGMRLDRSDLNLLYRARYPTE 501
GNLYKR+FG D+D ATNELV+RVQPDEAIYLK+NNKIPGLGMRLDRS+LNL Y ARY E
Sbjct: 449 GNLYKRSFGTDLDTATNELVIRVQPDEAIYLKINNKIPGLGMRLDRSNLNLHYAARYSKE 508
Query: 502 IPDAYERLLLDAIEGERRLFIRSDELDAAWALFTPLLKEVEDKKIAPELYPHGSRGPIGA 561
IPDAYERLLLDAIEGERRLFIRSDELDAAW LFTPLLKE+E+K+IAPELYP+GSRGP+GA
Sbjct: 509 IPDAYERLLLDAIEGERRLFIRSDELDAAWELFTPLLKELEEKRIAPELYPYGSRGPVGA 568
Query: 562 HYLAAKHNVRWGD 574
HYLAAK+NVRWGD
Sbjct: 569 HYLAAKYNVRWGD 581
>UniRef100_Q9ST67 Glucose-6-phosphate 1-dehydrogenase precursor [Solanum tuberosum]
Length = 582
Score = 833 bits (2151), Expect = 0.0
Identities = 410/532 (77%), Positives = 463/532 (86%), Gaps = 8/532 (1%)
Query: 51 LNAVSSHSHDGLAGSSLTKEDSKPQPVDGPFLSSDSECT--------GSNLSITVVGASG 102
+NAV +A S T++++ + + LS + + T S +SITVVGASG
Sbjct: 49 INAVRMQDGAVVAQPSKTQDETPLKKLKDGILSKEQKHTFDFDCNKDKSTVSITVVGASG 108
Query: 103 DLAKKKIFPALFALFYEDCLPENFLVFGFARTKMTSEELRNMISRTLTCRIDKRANCEDK 162
DLAKKKIFPALFAL+YE CLPE+F +FG+AR+KM+ +ELRNM+S+TLTCRIDKR NC +K
Sbjct: 109 DLAKKKIFPALFALYYEGCLPEHFTIFGYARSKMSDDELRNMVSKTLTCRIDKRENCGEK 168
Query: 163 MDQFLKRCFYHSGLYNSEGSFSDLDCKMKEKEGGKRSNRLFYLSIPPNIFVNVVRCASLK 222
M+QFL+RCFYHSG Y+S+ F++LD K+KE E G+ SNRLFYLSIPPNIF+N VRCASL
Sbjct: 169 MEQFLERCFYHSGQYDSQEHFAELDKKLKEHEAGRFSNRLFYLSIPPNIFINAVRCASLS 228
Query: 223 ASSKKGWTRVIVEKPFGRDSESSNELTRCLKQYLTEDQIFRIDHYLGKELVENLSVLRFS 282
ASS GWTRVIVEKPFGRDSESS LT LKQYL EDQIFRIDHYLGKELVENLSVLRF
Sbjct: 229 ASSAHGWTRVIVEKPFGRDSESSAALTGALKQYLKEDQIFRIDHYLGKELVENLSVLRFF 288
Query: 283 NLVFEPLWSRAYIRNVQFIFSEDFGTEGRGGYFDNYGIIRDIMQNHLLQILALFAMEPPV 342
NL+FEPLWSR YIRNVQFIFSEDFGTEGRGGYFD+YGIIRDIMQNHLLQILALFAME PV
Sbjct: 289 NLIFEPLWSRQYIRNVQFIFSEDFGTEGRGGYFDHYGIIRDIMQNHLLQILALFAMETPV 348
Query: 343 SLDAEDIRNEKVKVLRSMRPLQLENVVTGQYKGHSKGGKSYPGYADDPSVPKGSLTPTFA 402
SLDAEDIRNEKVKVLRSMRPLQL++V+ GQYK H+KGG +YPGY DD +VPK SLTPTFA
Sbjct: 349 SLDAEDIRNEKVKVLRSMRPLQLDDVIVGQYKSHTKGGVNYPGYTDDKTVPKDSLTPTFA 408
Query: 403 AAALFIDNARWDGVPFLMKAGKALHTKRAEIRVQFRHVPGNLYKRNFGADMDKATNELVL 462
AAALFIDNARWDGVPFLMKAGKALHT+ AEIRVQFRHVPGNLY +NFG+D+D+ATNELV+
Sbjct: 409 AAALFIDNARWDGVPFLMKAGKALHTRSAEIRVQFRHVPGNLYNKNFGSDLDQATNELVI 468
Query: 463 RVQPDEAIYLKVNNKIPGLGMRLDRSDLNLLYRARYPTEIPDAYERLLLDAIEGERRLFI 522
RVQPDEAIYLK+NNK+PGLGMRLDRS+LNLLY ARY EIPDAYERLLLDAIEGERRLFI
Sbjct: 469 RVQPDEAIYLKINNKVPGLGMRLDRSNLNLLYSARYAKEIPDAYERLLLDAIEGERRLFI 528
Query: 523 RSDELDAAWALFTPLLKEVEDKKIAPELYPHGSRGPIGAHYLAAKHNVRWGD 574
RSDELDAAW+LFTP+LKE+E KKI PE YP+GSRGPIGAHYLAA++ VRWGD
Sbjct: 529 RSDELDAAWSLFTPVLKELEHKKIVPESYPYGSRGPIGAHYLAARYKVRWGD 580
>UniRef100_Q9LL88 Plastidic glucose 6-phosphate dehydrogenase [Nicotiana tabacum]
Length = 593
Score = 832 bits (2150), Expect = 0.0
Identities = 404/490 (82%), Positives = 446/490 (90%)
Query: 85 DSECTGSNLSITVVGASGDLAKKKIFPALFALFYEDCLPENFLVFGFARTKMTSEELRNM 144
DS S +SITVVGASGDLAKKKIFPALFAL+YE CLPE+F +FG+AR+KMT ELRNM
Sbjct: 102 DSNKAKSTVSITVVGASGDLAKKKIFPALFALYYEGCLPEHFTIFGYARSKMTDAELRNM 161
Query: 145 ISRTLTCRIDKRANCEDKMDQFLKRCFYHSGLYNSEGSFSDLDCKMKEKEGGKRSNRLFY 204
+S+TLTCRIDKR NC +KM+QFL+RCFYHSG Y+S +F++LD K+KE E G+ SNRLFY
Sbjct: 162 VSKTLTCRIDKRENCGEKMEQFLERCFYHSGQYDSLENFAELDKKLKEHEAGRFSNRLFY 221
Query: 205 LSIPPNIFVNVVRCASLKASSKKGWTRVIVEKPFGRDSESSNELTRCLKQYLTEDQIFRI 264
LSIPPNIF+N VRCASL ASS GWTRVIVEKPFGRDSESS LTR LKQYL EDQIFRI
Sbjct: 222 LSIPPNIFINAVRCASLSASSAHGWTRVIVEKPFGRDSESSAALTRSLKQYLNEDQIFRI 281
Query: 265 DHYLGKELVENLSVLRFSNLVFEPLWSRAYIRNVQFIFSEDFGTEGRGGYFDNYGIIRDI 324
DHYLGKELVENLSVLRFSNL+FEPLWSR YIRNVQFIFSEDFGTEGRGGYFD+YGIIRDI
Sbjct: 282 DHYLGKELVENLSVLRFSNLIFEPLWSRQYIRNVQFIFSEDFGTEGRGGYFDHYGIIRDI 341
Query: 325 MQNHLLQILALFAMEPPVSLDAEDIRNEKVKVLRSMRPLQLENVVTGQYKGHSKGGKSYP 384
MQNHLLQILALFAME PVSLDAEDIRNEKVKVLRSMRPLQL++V+ GQYK H+KG +YP
Sbjct: 342 MQNHLLQILALFAMETPVSLDAEDIRNEKVKVLRSMRPLQLDDVIIGQYKSHTKGDVTYP 401
Query: 385 GYADDPSVPKGSLTPTFAAAALFIDNARWDGVPFLMKAGKALHTKRAEIRVQFRHVPGNL 444
GY DD +VPK SLTPTFAAAALFIDNARWDGVPFLMKAGKALHT+ AEIRVQFRHVPGNL
Sbjct: 402 GYTDDKTVPKDSLTPTFAAAALFIDNARWDGVPFLMKAGKALHTRSAEIRVQFRHVPGNL 461
Query: 445 YKRNFGADMDKATNELVLRVQPDEAIYLKVNNKIPGLGMRLDRSDLNLLYRARYPTEIPD 504
Y +NFG+D+D+ATNELV+RVQP+EAIYLK+NNK+PGLGMRLDRS+LNLLY ARY EIPD
Sbjct: 462 YNKNFGSDLDQATNELVIRVQPNEAIYLKINNKVPGLGMRLDRSNLNLLYSARYSKEIPD 521
Query: 505 AYERLLLDAIEGERRLFIRSDELDAAWALFTPLLKEVEDKKIAPELYPHGSRGPIGAHYL 564
YERLLLDAIEGERRLFIRSDELDAAW+LFTP+LKE+EDKKI PE YP+GSRGPIGAHYL
Sbjct: 522 PYERLLLDAIEGERRLFIRSDELDAAWSLFTPVLKELEDKKIVPEYYPYGSRGPIGAHYL 581
Query: 565 AAKHNVRWGD 574
AA++ VRWGD
Sbjct: 582 AARYKVRWGD 591
>UniRef100_Q9FY99 Glucose-6-phosphate 1-dehydrogenase 2, chloroplast precursor
[Arabidopsis thaliana]
Length = 596
Score = 832 bits (2149), Expect = 0.0
Identities = 413/533 (77%), Positives = 465/533 (86%), Gaps = 13/533 (2%)
Query: 52 NAVSSHSHDGLAG------SSLTKEDSKPQPVDGPFLSSDSECTGSNLSITVVGASGDLA 105
N+ SS S L G S+L++E +K + SD + + S +SITVVGASGDLA
Sbjct: 70 NSNSSESKTSLKGLKDEVLSALSQEAAKVG------VESDGQ-SQSTVSITVVGASGDLA 122
Query: 106 KKKIFPALFALFYEDCLPENFLVFGFARTKMTSEELRNMISRTLTCRIDKRANCEDKMDQ 165
KKKIFPALFAL+YE CLPE+F +FG++R+KMT ELRNM+S+TLTCRIDKRANC +KM++
Sbjct: 123 KKKIFPALFALYYEGCLPEHFTIFGYSRSKMTDVELRNMVSKTLTCRIDKRANCGEKMEE 182
Query: 166 FLKRCFYHSGLYNSEGSFSDLDCKMKEKEGGKRSNRLFYLSIPPNIFVNVVRCASLKASS 225
FLKRCFYHSG Y+S+ F++LD K+KE E G+ SNRLFYLSIPPNIFV+ V+CAS ASS
Sbjct: 183 FLKRCFYHSGQYDSQEHFTELDKKLKEHEAGRISNRLFYLSIPPNIFVDAVKCASTSASS 242
Query: 226 KKGWTRVIVEKPFGRDSESSNELTRCLKQYLTEDQIFRIDHYLGKELVENLSVLRFSNLV 285
GWTRVIVEKPFGRDSE+S LT+ LKQYL EDQIFRIDHYLGKELVENLSVLRFSNL+
Sbjct: 243 VNGWTRVIVEKPFGRDSETSAALTKSLKQYLEEDQIFRIDHYLGKELVENLSVLRFSNLI 302
Query: 286 FEPLWSRAYIRNVQFIFSEDFGTEGRGGYFDNYGIIRDIMQNHLLQILALFAMEPPVSLD 345
FEPLWSR YIRNVQFIFSEDFGTEGRGGYFDNYGIIRDIMQNHLLQILALFAME PVSLD
Sbjct: 303 FEPLWSRQYIRNVQFIFSEDFGTEGRGGYFDNYGIIRDIMQNHLLQILALFAMETPVSLD 362
Query: 346 AEDIRNEKVKVLRSMRPLQLENVVTGQYKGHSKGGKSYPGYADDPSVPKGSLTPTFAAAA 405
AEDIRNEKVKVLRSMRP+++E+VV GQYK H+KGG +YP Y DD +VPKGSLTPTFAAAA
Sbjct: 363 AEDIRNEKVKVLRSMRPIRVEDVVIGQYKSHTKGGVTYPAYTDDKTVPKGSLTPTFAAAA 422
Query: 406 LFIDNARWDGVPFLMKAGKALHTKRAEIRVQFRHVPGNLYKRNFGADMDKATNELVLRVQ 465
LFIDNARWDGVPFLMKAGKALHT+ AEIRVQFRHVPGNLY RN G+D+D+ATNELV+RVQ
Sbjct: 423 LFIDNARWDGVPFLMKAGKALHTRSAEIRVQFRHVPGNLYNRNTGSDLDQATNELVIRVQ 482
Query: 466 PDEAIYLKVNNKIPGLGMRLDRSDLNLLYRARYPTEIPDAYERLLLDAIEGERRLFIRSD 525
PDEAIYLK+NNK+PGLGMRLDRS+LNLLY ARY EIPDAYERLLLDAIEGERRLFIRSD
Sbjct: 483 PDEAIYLKINNKVPGLGMRLDRSNLNLLYSARYSKEIPDAYERLLLDAIEGERRLFIRSD 542
Query: 526 ELDAAWALFTPLLKEVEDKKIAPELYPHGSRGPIGAHYLAAKHNVRWGDTGND 578
ELDAAW+LFTPLLKE+E+KK PE YP+GSRGP+GAHYLAAKH V+WGD D
Sbjct: 543 ELDAAWSLFTPLLKEIEEKKRIPEYYPYGSRGPVGAHYLAAKHKVQWGDVSID 595
>UniRef100_Q8H9C8 Glucose 6-phosphate dehydrogenase [Solanum tuberosum]
Length = 581
Score = 832 bits (2148), Expect = 0.0
Identities = 411/532 (77%), Positives = 464/532 (86%), Gaps = 9/532 (1%)
Query: 51 LNAVSSHSHDGLAGSSLTKEDSKPQPVDGPFLSS--------DSECTGSNLSITVVGASG 102
+NA+ +A S T++++ + + LS DS S +SITVVGASG
Sbjct: 49 INAIRMQDGAVVAQPSKTQDETPLKKLKDGILSKEQKHTFDFDSNKDKSTVSITVVGASG 108
Query: 103 DLAKKKIFPALFALFYEDCLPENFLVFGFARTKMTSEELRNMISRTLTCRIDKRANCEDK 162
DLAKKKIFPALFAL+YE CLPE+F +FG+AR+KMT +ELRNM+S+TLTCRIDKR NC +K
Sbjct: 109 DLAKKKIFPALFALYYEGCLPEHFTIFGYARSKMTDDELRNMVSKTLTCRIDKRENCGEK 168
Query: 163 MDQFLKRCFYHSGLYNSEGSFSDLDCKMKEKEGGKRSNRLFYLSIPPNIFVNVVRCASLK 222
M+QFL+RCFYHSG Y+S+ +F++LD K+KE E G+ SNRLFYLSIPPNIF+N VRCASL
Sbjct: 169 MEQFLERCFYHSGQYDSQENFAELDKKLKEHEAGRFSNRLFYLSIPPNIFINAVRCASL- 227
Query: 223 ASSKKGWTRVIVEKPFGRDSESSNELTRCLKQYLTEDQIFRIDHYLGKELVENLSVLRFS 282
ASS GWTRVIVEKPFGRDSESS LT LKQYL EDQIFRIDHYLGKELVENLSVLRFS
Sbjct: 228 ASSAHGWTRVIVEKPFGRDSESSAALTGALKQYLKEDQIFRIDHYLGKELVENLSVLRFS 287
Query: 283 NLVFEPLWSRAYIRNVQFIFSEDFGTEGRGGYFDNYGIIRDIMQNHLLQILALFAMEPPV 342
NL+FEPLWSR YIRNVQFIFSEDFGTEGRGGYFD+YGIIRDIMQNHLLQILALFAME PV
Sbjct: 288 NLIFEPLWSRQYIRNVQFIFSEDFGTEGRGGYFDHYGIIRDIMQNHLLQILALFAMETPV 347
Query: 343 SLDAEDIRNEKVKVLRSMRPLQLENVVTGQYKGHSKGGKSYPGYADDPSVPKGSLTPTFA 402
SLDAEDIRNEKVKVLRSMRPLQL++V+ GQYK H+KGG +YPGY DD +VPK SLTPTFA
Sbjct: 348 SLDAEDIRNEKVKVLRSMRPLQLDDVIVGQYKSHTKGGVNYPGYTDDKTVPKDSLTPTFA 407
Query: 403 AAALFIDNARWDGVPFLMKAGKALHTKRAEIRVQFRHVPGNLYKRNFGADMDKATNELVL 462
AAALFIDNARWDGVPFLMKAGKALHT+ AEIRVQFRHVPGNLY +NFG+D+D+ATNELV+
Sbjct: 408 AAALFIDNARWDGVPFLMKAGKALHTRSAEIRVQFRHVPGNLYNKNFGSDLDQATNELVI 467
Query: 463 RVQPDEAIYLKVNNKIPGLGMRLDRSDLNLLYRARYPTEIPDAYERLLLDAIEGERRLFI 522
RVQPDEAIYLK+NNK+PGLGMRLDRS+LNLLY ARY EIPDAYERLLLDAIEGERRLFI
Sbjct: 468 RVQPDEAIYLKINNKVPGLGMRLDRSNLNLLYSARYSKEIPDAYERLLLDAIEGERRLFI 527
Query: 523 RSDELDAAWALFTPLLKEVEDKKIAPELYPHGSRGPIGAHYLAAKHNVRWGD 574
RSDELDAAW+LFTP+LK++E KKI PE YP+GSRGPIGAHYLAA++ VRWGD
Sbjct: 528 RSDELDAAWSLFTPVLKDLEHKKIVPESYPYGSRGPIGAHYLAARYKVRWGD 579
>UniRef100_Q43793 Glucose-6-phosphate 1-dehydrogenase, chloroplast precursor
[Nicotiana tabacum]
Length = 593
Score = 830 bits (2143), Expect = 0.0
Identities = 404/490 (82%), Positives = 446/490 (90%)
Query: 85 DSECTGSNLSITVVGASGDLAKKKIFPALFALFYEDCLPENFLVFGFARTKMTSEELRNM 144
DS S +SITVVGASGDLAKKKIFPALFAL+YE CLPE+F +FG+AR+KMT ELRNM
Sbjct: 102 DSNKAKSTVSITVVGASGDLAKKKIFPALFALYYEGCLPEHFTIFGYARSKMTDAELRNM 161
Query: 145 ISRTLTCRIDKRANCEDKMDQFLKRCFYHSGLYNSEGSFSDLDCKMKEKEGGKRSNRLFY 204
+S+TLTCRIDKR NC +KM+QFL+RCFYHSG Y+S +F++LD K+KE E G+ SNRLFY
Sbjct: 162 VSKTLTCRIDKRENCGEKMEQFLERCFYHSGQYDSLENFAELDKKLKEHEAGRFSNRLFY 221
Query: 205 LSIPPNIFVNVVRCASLKASSKKGWTRVIVEKPFGRDSESSNELTRCLKQYLTEDQIFRI 264
LSIPPNIF+N VRCASL ASS GWTRVIVEKPFGRDSESS LTR LKQYL EDQIFRI
Sbjct: 222 LSIPPNIFINAVRCASLSASSAHGWTRVIVEKPFGRDSESSAALTRSLKQYLNEDQIFRI 281
Query: 265 DHYLGKELVENLSVLRFSNLVFEPLWSRAYIRNVQFIFSEDFGTEGRGGYFDNYGIIRDI 324
DHYLGKELVENLSVLRFSNL+FEPLWSR IRNVQFIFSEDFGTEGRGGYFD+YGIIRDI
Sbjct: 282 DHYLGKELVENLSVLRFSNLIFEPLWSRQCIRNVQFIFSEDFGTEGRGGYFDHYGIIRDI 341
Query: 325 MQNHLLQILALFAMEPPVSLDAEDIRNEKVKVLRSMRPLQLENVVTGQYKGHSKGGKSYP 384
MQNHLLQILALFAME PVSLDAEDIRNEKVKVLRSMRPLQL++V+ GQYK H+KG +YP
Sbjct: 342 MQNHLLQILALFAMETPVSLDAEDIRNEKVKVLRSMRPLQLDDVIIGQYKCHTKGDVTYP 401
Query: 385 GYADDPSVPKGSLTPTFAAAALFIDNARWDGVPFLMKAGKALHTKRAEIRVQFRHVPGNL 444
GY DD +VPK SLTPTFAAAALFIDNARWDGVPFLMKAGKALHT+ AEIRVQFRHVPGNL
Sbjct: 402 GYTDDKTVPKDSLTPTFAAAALFIDNARWDGVPFLMKAGKALHTRSAEIRVQFRHVPGNL 461
Query: 445 YKRNFGADMDKATNELVLRVQPDEAIYLKVNNKIPGLGMRLDRSDLNLLYRARYPTEIPD 504
Y +NFG+D+D+ATNELV+RVQP+EAIYLK+NNK+PGLGMRLDRS+LNLLY ARY EIPD
Sbjct: 462 YNKNFGSDLDQATNELVIRVQPNEAIYLKINNKVPGLGMRLDRSNLNLLYSARYSKEIPD 521
Query: 505 AYERLLLDAIEGERRLFIRSDELDAAWALFTPLLKEVEDKKIAPELYPHGSRGPIGAHYL 564
AYERLLLDAIEGERRLFIRSDELDAAW+LFTP+LKE+EDKKI PE YP+GSRGPIGAHYL
Sbjct: 522 AYERLLLDAIEGERRLFIRSDELDAAWSLFTPVLKELEDKKIVPEYYPYGSRGPIGAHYL 581
Query: 565 AAKHNVRWGD 574
AA++ VRWGD
Sbjct: 582 AARYKVRWGD 591
>UniRef100_Q75IZ9 Putative glucose-6-phosphate dehydrogenase [Oryza sativa]
Length = 577
Score = 827 bits (2135), Expect = 0.0
Identities = 420/541 (77%), Positives = 458/541 (84%), Gaps = 25/541 (4%)
Query: 44 KSSNGHPLNAVSSHSHDGLAGSSLTKEDSKPQPVDGPFLSSDSECTGSNLSITVVGASGD 103
KS+NG P +S+ D +A ED P E GS +SITVVGASGD
Sbjct: 50 KSANGRP--QISASFRD-VAIDGAQSEDGAP------------EQGGSTVSITVVGASGD 94
Query: 104 LAKKKIFPALFALFYEDCLPENFLVFGFARTKMTSEELRNMISRTLTCRIDKRANCEDKM 163
LAKKKIFPALFAL+YEDCLPE+F VFG+AR+KM+ EELRNMIS TLTCRID+R NC DKM
Sbjct: 95 LAKKKIFPALFALYYEDCLPEHFTVFGYARSKMSDEELRNMISLTLTCRIDQRENCSDKM 154
Query: 164 DQFLKRCFYHSGLYNSEGSFSDLDCKMKEKEGGKRSNRLFYLSIPPNIFVNVVRCASLKA 223
+QFLKRCFY SG YNSE FS+LD K+KEKE GK NRLFYLSIPPNIFV+VVR AS A
Sbjct: 155 EQFLKRCFYQSGQYNSEEGFSELDRKLKEKEAGKVPNRLFYLSIPPNIFVDVVRSASRTA 214
Query: 224 SSKKGWTRVIVEKPFGRDSESSNELTRCLKQYLTEDQIFRIDHYLGKELVENLSVLRFSN 283
SS+ GWTR IVEKPFGRDSESS ELTR LK+YL E+QIFRIDHYLGKELVENLSVLRFSN
Sbjct: 215 SSQDGWTRFIVEKPFGRDSESSGELTRNLKKYLAEEQIFRIDHYLGKELVENLSVLRFSN 274
Query: 284 LVFEPLWSRAYIRNVQFIFSEDFGTEGRGGYFDNYGIIRDIMQNHLLQILALFAMEPPVS 343
LVFEPLWSR YIRNVQ IFSEDFGTEGRGGYFDNYGIIRDIMQNHLLQILALFAME PVS
Sbjct: 275 LVFEPLWSRNYIRNVQLIFSEDFGTEGRGGYFDNYGIIRDIMQNHLLQILALFAMETPVS 334
Query: 344 LDAEDIRNEKVKVLRSMRPLQLENVVTGQYKGHSKGGKSYPGYADDPSVPKGSLTPTFAA 403
LDAEDIRNEKVKVLRSMR L+LE+VV GQYKGHSKGGK+YP Y DDP+VP GS+TPTFAA
Sbjct: 335 LDAEDIRNEKVKVLRSMRQLRLEDVVVGQYKGHSKGGKTYPAYVDDPTVPSGSITPTFAA 394
Query: 404 AALFIDNARWDGVPFLMKAGKALHTKRAEIRVQFRHVPGNLYKRNFGA----------DM 453
AALFIDNARWDGVPFLMKAGKALHT+RAEIRVQFR VPGNLY R + ++
Sbjct: 395 AALFIDNARWDGVPFLMKAGKALHTRRAEIRVQFRRVPGNLYGRRSRSVGGGGTTATREL 454
Query: 454 DKATNELVLRVQPDEAIYLKVNNKIPGLGMRLDRSDLNLLYRARYPTEIPDAYERLLLDA 513
+KATNELVLRVQPDEAIYLK+N+K+PGLGMRLD SDLNLLY RYP EIPDAYERLLLDA
Sbjct: 455 EKATNELVLRVQPDEAIYLKINSKVPGLGMRLDSSDLNLLYSERYPAEIPDAYERLLLDA 514
Query: 514 IEGERRLFIRSDELDAAWALFTPLLKEVEDKKIAPELYPHGSRGPIGAHYLAAKHNVRWG 573
IEGERRLFIRSDELDAAWA+FTP+L ++E K+APELYP+GSRGP+GAHYLAA HNVRWG
Sbjct: 515 IEGERRLFIRSDELDAAWAIFTPVLADLEANKVAPELYPYGSRGPVGAHYLAANHNVRWG 574
Query: 574 D 574
D
Sbjct: 575 D 575
>UniRef100_Q8L743 Glucose-6-phosphate 1-dehydrogenase 3, chloroplast precursor
[Arabidopsis thaliana]
Length = 599
Score = 820 bits (2118), Expect = 0.0
Identities = 400/488 (81%), Positives = 439/488 (88%)
Query: 91 SNLSITVVGASGDLAKKKIFPALFALFYEDCLPENFLVFGFARTKMTSEELRNMISRTLT 150
S +SITVVGASGDLAKKKIFPALFAL+YE CLPE+F +FG+AR+KMT ELR M+S+TLT
Sbjct: 111 STVSITVVGASGDLAKKKIFPALFALYYEGCLPEHFTIFGYARSKMTDAELRVMVSKTLT 170
Query: 151 CRIDKRANCEDKMDQFLKRCFYHSGLYNSEGSFSDLDCKMKEKEGGKRSNRLFYLSIPPN 210
CRIDKRANC +KM++FLKRCFYHSG Y+S+ F LD K+KE EGG+ SNRLFYLSIPPN
Sbjct: 171 CRIDKRANCGEKMEEFLKRCFYHSGQYDSQEHFVALDEKLKEHEGGRLSNRLFYLSIPPN 230
Query: 211 IFVNVVRCASLKASSKKGWTRVIVEKPFGRDSESSNELTRCLKQYLTEDQIFRIDHYLGK 270
IFV+ V+CAS ASS GWTRVIVEKPFGRDS++S LT+ LKQYL EDQIFRIDHYLGK
Sbjct: 231 IFVDAVKCASSSASSVNGWTRVIVEKPFGRDSKTSAALTKSLKQYLEEDQIFRIDHYLGK 290
Query: 271 ELVENLSVLRFSNLVFEPLWSRAYIRNVQFIFSEDFGTEGRGGYFDNYGIIRDIMQNHLL 330
ELVENLSVLRFSNL+FEPLWSR YIRNVQFIFSEDFGTEGRGGYFDNYGIIRDIMQNHLL
Sbjct: 291 ELVENLSVLRFSNLIFEPLWSRQYIRNVQFIFSEDFGTEGRGGYFDNYGIIRDIMQNHLL 350
Query: 331 QILALFAMEPPVSLDAEDIRNEKVKVLRSMRPLQLENVVTGQYKGHSKGGKSYPGYADDP 390
QILALFAME PVSLDAEDIRNEKVKVLRSMRP++LE+VV GQYK HS GG +YP Y DD
Sbjct: 351 QILALFAMETPVSLDAEDIRNEKVKVLRSMRPIKLEDVVIGQYKSHSIGGVTYPSYTDDK 410
Query: 391 SVPKGSLTPTFAAAALFIDNARWDGVPFLMKAGKALHTKRAEIRVQFRHVPGNLYKRNFG 450
+VPKGSLTPTFAAAALFIDNARWDGVPFLMKAGKAL+T+ AEIRVQFRHVPGNLY RN G
Sbjct: 411 TVPKGSLTPTFAAAALFIDNARWDGVPFLMKAGKALNTRSAEIRVQFRHVPGNLYNRNSG 470
Query: 451 ADMDKATNELVLRVQPDEAIYLKVNNKIPGLGMRLDRSDLNLLYRARYPTEIPDAYERLL 510
D D+ TNELV+RVQPDEAIYLK+NNK+PGLGMRLD+S+LNLLY ARY EIPDAYERLL
Sbjct: 471 TDRDQTTNELVIRVQPDEAIYLKINNKVPGLGMRLDQSNLNLLYSARYSKEIPDAYERLL 530
Query: 511 LDAIEGERRLFIRSDELDAAWALFTPLLKEVEDKKIAPELYPHGSRGPIGAHYLAAKHNV 570
LDAIEGERRLFIRSDELDAAWALFTPLLKE+E+KK PE YP+GSRGP+GAHYLAAKH V
Sbjct: 531 LDAIEGERRLFIRSDELDAAWALFTPLLKEIEEKKTTPEFYPYGSRGPVGAHYLAAKHKV 590
Query: 571 RWGDTGND 578
+WGD D
Sbjct: 591 QWGDLSLD 598
>UniRef100_O22404 Plastidic glucose-6-phosphate dehydrogenase [Petroselinum crispum]
Length = 604
Score = 820 bits (2117), Expect = 0.0
Identities = 393/488 (80%), Positives = 440/488 (89%)
Query: 91 SNLSITVVGASGDLAKKKIFPALFALFYEDCLPENFLVFGFARTKMTSEELRNMISRTLT 150
S+++ITVVGASGDLAKKKIFPALFAL+YEDCLPE+F +FG+AR+KMT ELR+M+S+TLT
Sbjct: 116 SSVTITVVGASGDLAKKKIFPALFALYYEDCLPEHFTIFGYARSKMTDAELRDMVSKTLT 175
Query: 151 CRIDKRANCEDKMDQFLKRCFYHSGLYNSEGSFSDLDCKMKEKEGGKRSNRLFYLSIPPN 210
CRIDKRANC +KM+QFLKRCFYHSG Y+SE +F++LD K+KE E G +NRLFYLSIPPN
Sbjct: 176 CRIDKRANCGEKMEQFLKRCFYHSGQYDSEANFAELDKKLKEHEAGTIANRLFYLSIPPN 235
Query: 211 IFVNVVRCASLKASSKKGWTRVIVEKPFGRDSESSNELTRCLKQYLTEDQIFRIDHYLGK 270
IF+N V+ AS+ ASS GWTRVIVEKPFGRDSESS LT LKQYL EDQIFRIDHYLGK
Sbjct: 236 IFINAVKSASISASSANGWTRVIVEKPFGRDSESSAALTTALKQYLEEDQIFRIDHYLGK 295
Query: 271 ELVENLSVLRFSNLVFEPLWSRAYIRNVQFIFSEDFGTEGRGGYFDNYGIIRDIMQNHLL 330
ELVENLSVLRFSNL+FEPLWSR +IRNVQ IFSEDFGTEGRGGYFDNYGI+RDIMQNHLL
Sbjct: 296 ELVENLSVLRFSNLIFEPLWSRQFIRNVQLIFSEDFGTEGRGGYFDNYGIVRDIMQNHLL 355
Query: 331 QILALFAMEPPVSLDAEDIRNEKVKVLRSMRPLQLENVVTGQYKGHSKGGKSYPGYADDP 390
QILALFAME PVSLDAEDIRNEKVKVLRSMRP+QL++VV GQYK H++GG +YP Y DD
Sbjct: 356 QILALFAMETPVSLDAEDIRNEKVKVLRSMRPIQLDDVVIGQYKSHTRGGVNYPAYTDDK 415
Query: 391 SVPKGSLTPTFAAAALFIDNARWDGVPFLMKAGKALHTKRAEIRVQFRHVPGNLYKRNFG 450
+VP SLTPTFAAAALFIDNARWDGVPFLMKAGKALH +R EIRVQFRHVPGNLYKRNFG
Sbjct: 416 TVPHNSLTPTFAAAALFIDNARWDGVPFLMKAGKALHDRRTEIRVQFRHVPGNLYKRNFG 475
Query: 451 ADMDKATNELVLRVQPDEAIYLKVNNKIPGLGMRLDRSDLNLLYRARYPTEIPDAYERLL 510
++D TNELV+RVQPDEAIYLK+NNK+PGLGMRLDRS+LNLLY RY EIPDAYERLL
Sbjct: 476 TNLDHETNELVIRVQPDEAIYLKINNKVPGLGMRLDRSNLNLLYSTRYSGEIPDAYERLL 535
Query: 511 LDAIEGERRLFIRSDELDAAWALFTPLLKEVEDKKIAPELYPHGSRGPIGAHYLAAKHNV 570
LDA+EGERRLFIRSDELDAAW+LFTP+LK++EDKK PE YP+GSRGP+GAHYLAAK+ V
Sbjct: 536 LDAVEGERRLFIRSDELDAAWSLFTPVLKDLEDKKTVPEYYPYGSRGPVGAHYLAAKYKV 595
Query: 571 RWGDTGND 578
RWGD D
Sbjct: 596 RWGDFAGD 603
>UniRef100_O24359 Glucose-6-phosphate dehydrogenase [Spinacia oleracea]
Length = 465
Score = 733 bits (1891), Expect = 0.0
Identities = 367/465 (78%), Positives = 394/465 (83%), Gaps = 2/465 (0%)
Query: 46 SNGHPLNAVSSHSHDGLAGSSLTKEDSKPQPVDGPFLSSDSECTGSNLSITVVGASGDLA 105
SNGHPLN VS + + D P L S E LSI VVGASGDLA
Sbjct: 3 SNGHPLNDVSLQNDVAVNPIVAKSIDPSADLQLLPLLESVKE--EPTLSIIVVGASGDLA 60
Query: 106 KKKIFPALFALFYEDCLPENFLVFGFARTKMTSEELRNMISRTLTCRIDKRANCEDKMDQ 165
KK+IFP LFALFYE+CLPENF VFGF+RT+M EELR MIS+TLTCRID+R NC +KMD
Sbjct: 61 KKRIFPTLFALFYENCLPENFTVFGFSRTEMNDEELRTMISKTLTCRIDQRENCGEKMDH 120
Query: 166 FLKRCFYHSGLYNSEGSFSDLDCKMKEKEGGKRSNRLFYLSIPPNIFVNVVRCASLKASS 225
FL+RCFYHSG YNSE FS LDCK+KEKE G+ NRLFYLSIPPNIFV+VVRC S +ASS
Sbjct: 121 FLQRCFYHSGQYNSEDDFSGLDCKLKEKEAGRLQNRLFYLSIPPNIFVDVVRCVSHRASS 180
Query: 226 KKGWTRVIVEKPFGRDSESSNELTRCLKQYLTEDQIFRIDHYLGKELVENLSVLRFSNLV 285
GWTRVIVEKPFGRDS+SS ELTR KQYL+EDQIFRIDHYLGKELVENLSVLRFSNLV
Sbjct: 181 ASGWTRVIVEKPFGRDSDSSRELTRSFKQYLSEDQIFRIDHYLGKELVENLSVLRFSNLV 240
Query: 286 FEPLWSRAYIRNVQFIFSEDFGTEGRGGYFDNYGIIRDIMQNHLLQILALFAMEPPVSLD 345
FEPLWSR YIRNVQ IFSEDFGTEGRGGYFDNYGIIRDIMQNHLLQILALFAME PVSLD
Sbjct: 241 FEPLWSRNYIRNVQLIFSEDFGTEGRGGYFDNYGIIRDIMQNHLLQILALFAMETPVSLD 300
Query: 346 AEDIRNEKVKVLRSMRPLQLENVVTGQYKGHSKGGKSYPGYADDPSVPKGSLTPTFAAAA 405
EDIRNEKVKVLRSM+PL+L++VV GQYKGHSKG KSY GY DDP+VP S+TPTFAAAA
Sbjct: 301 TEDIRNEKVKVLRSMKPLKLQDVVVGQYKGHSKGNKSYSGYTDDPTVPNNSVTPTFAAAA 360
Query: 406 LFIDNARWDGVPFLMKAGKALHTKRAEIRVQFRHVPGNLYKRNFGADMDKATNELVLRVQ 465
LFIDNARWDGVPFLMKAGKALHTKRAEIRVQFRHVPGNLYK+ FG D+DKATNELVLRVQ
Sbjct: 361 LFIDNARWDGVPFLMKAGKALHTKRAEIRVQFRHVPGNLYKKTFGTDLDKATNELVLRVQ 420
Query: 466 PDEAIYLKVNNKIPGLGMRLDRSDLNLLYRARYPTEIPDAYERLL 510
PDEAIYLK+NNK+PGLGMRLDR+DLNL Y RY EIPDAYERLL
Sbjct: 421 PDEAIYLKINNKVPGLGMRLDRTDLNLCYSTRYRGEIPDAYERLL 465
>UniRef100_Q9STC7 Plastidic glucose-6-phosphate dehydrogenase [Dunaliella bioculata]
Length = 590
Score = 696 bits (1797), Expect = 0.0
Identities = 341/496 (68%), Positives = 402/496 (80%), Gaps = 7/496 (1%)
Query: 91 SNLSITVVGASGDLAKKKIFPALFALFYEDCLPENFLVFGFARTKMTSEELRNMISRTLT 150
S+LS+ VVGASGDLAKKKIFPALFALFYE LP +F +FG+AR+KMT EE R++I TLT
Sbjct: 94 SSLSVVVVGASGDLAKKKIFPALFALFYEGLLPPDFQLFGYARSKMTDEEFRDLIGNTLT 153
Query: 151 CRIDKRANCEDKMDQFLKRCFYHSGLYNSEGSFSDLDCKMKEKE---GGKRSNRLFYLSI 207
CRID R+ CED FL RCFY G Y++ +++LD K KE+E G +NR+F+LSI
Sbjct: 154 CRIDARSRCEDSQAAFLSRCFYCPGQYDAPEGYANLDKKCKEQEALTGKMVANRMFFLSI 213
Query: 208 PPNIFVNVVRCASLKASSKKGWTRVIVEKPFGRDSESSNELTRCLKQYLTEDQIFRIDHY 267
PPN+FV A+ SS GWTRVIVEKPFGRDS SS EL R L ++LTEDQI+RIDHY
Sbjct: 214 PPNVFVQAAGGAADNCSSPTGWTRVIVEKPFGRDSASSAELGRGLARHLTEDQIYRIDHY 273
Query: 268 LGKELVENLSVLRFSNLVFEPLWSRAYIRNVQFIFSEDFGTEGRGGYFDNYGIIRDIMQN 327
LGKEL+ENL+VLRFSNLVFEPLWSR YIRNVQ IFSEDFGTEGRGGYFD YGIIRD+MQN
Sbjct: 274 LGKELIENLTVLRFSNLVFEPLWSRQYIRNVQVIFSEDFGTEGRGGYFDRYGIIRDVMQN 333
Query: 328 HLLQILALFAMEPPVSLDAEDIRNEKVKVLRSMRPLQLENVVTGQYKGHS----KGGKSY 383
HLLQI+ALFAMEPPVSLD E IRNEKVKVL+SM + LE+V GQY+G S GG
Sbjct: 334 HLLQIVALFAMEPPVSLDGEAIRNEKVKVLQSMSQVALEDVTLGQYRGRSGAGRSGGADL 393
Query: 384 PGYADDPSVPKGSLTPTFAAAALFIDNARWDGVPFLMKAGKALHTKRAEIRVQFRHVPGN 443
PGY DD +VPKGSL PTFAA AL I+NARWDGVPFL+KAGKALHT+ AEIRVQFRHVPGN
Sbjct: 394 PGYLDDATVPKGSLCPTFAAIALHINNARWDGVPFLLKAGKALHTRGAEIRVQFRHVPGN 453
Query: 444 LYKRNFGADMDKATNELVLRVQPDEAIYLKVNNKIPGLGMRLDRSDLNLLYRARYPTEIP 503
++K G ++D TNELV+R+QP E+IYLK+NNK+PGLGMRLD + L+L+Y Y E+P
Sbjct: 454 IFKHKVGPNIDMTTNELVIRIQPRESIYLKINNKVPGLGMRLDTTKLDLVYNDAYSKELP 513
Query: 504 DAYERLLLDAIEGERRLFIRSDELDAAWALFTPLLKEVEDKKIAPELYPHGSRGPIGAHY 563
DAYERLLLD + G++RLFIR DEL+ AW +FTP+L E+E +K+APELYP+GSRGP+GAHY
Sbjct: 514 DAYERLLLDVVNGDKRLFIRDDELEQAWNIFTPVLHEIERRKVAPELYPYGSRGPVGAHY 573
Query: 564 LAAKHNVRWGDTGNDD 579
LAAK+NVRWGD D+
Sbjct: 574 LAAKYNVRWGDLTEDE 589
>UniRef100_O24358 Glucose-6-phosphate dehydrogenase [Spinacia oleracea]
Length = 317
Score = 573 bits (1477), Expect = e-162
Identities = 276/316 (87%), Positives = 295/316 (93%)
Query: 264 IDHYLGKELVENLSVLRFSNLVFEPLWSRAYIRNVQFIFSEDFGTEGRGGYFDNYGIIRD 323
IDHYLGKELVENLSVL FSNLVFEPLWSR YIRNVQ IFSEDFGTEGRGGYFDNYGIIRD
Sbjct: 1 IDHYLGKELVENLSVLHFSNLVFEPLWSRNYIRNVQLIFSEDFGTEGRGGYFDNYGIIRD 60
Query: 324 IMQNHLLQILALFAMEPPVSLDAEDIRNEKVKVLRSMRPLQLENVVTGQYKGHSKGGKSY 383
IMQNHLLQILALFAME PVSLDAEDIRNEKVKVLRSM+PL+L++VV GQYKGHSKG KSY
Sbjct: 61 IMQNHLLQILALFAMETPVSLDAEDIRNEKVKVLRSMKPLKLQDVVVGQYKGHSKGNKSY 120
Query: 384 PGYADDPSVPKGSLTPTFAAAALFIDNARWDGVPFLMKAGKALHTKRAEIRVQFRHVPGN 443
GY DDP+VP S+TP FAAAALFIDNARWDGVPFLMKAGKALHTKRAEIRVQFRHVPGN
Sbjct: 121 SGYTDDPTVPNNSVTPAFAAAALFIDNARWDGVPFLMKAGKALHTKRAEIRVQFRHVPGN 180
Query: 444 LYKRNFGADMDKATNELVLRVQPDEAIYLKVNNKIPGLGMRLDRSDLNLLYRARYPTEIP 503
LYK+ FG D++KATNELVLRVQPDEAIYLK+NNK+PGLGMRLDR+DLNL Y RY EIP
Sbjct: 181 LYKKTFGTDLEKATNELVLRVQPDEAIYLKINNKVPGLGMRLDRTDLNLCYSTRYRGEIP 240
Query: 504 DAYERLLLDAIEGERRLFIRSDELDAAWALFTPLLKEVEDKKIAPELYPHGSRGPIGAHY 563
DAYERLLLDAIEGERRLFIRSDELDAAW+LFTPLLKE+E+KK+APELYP+GSRGP+GAHY
Sbjct: 241 DAYERLLLDAIEGERRLFIRSDELDAAWSLFTPLLKELEEKKVAPELYPYGSRGPVGAHY 300
Query: 564 LAAKHNVRWGDTGNDD 579
LAAKHNVRWGD +D
Sbjct: 301 LAAKHNVRWGDLSGED 316
>UniRef100_Q93ZW0 Glucose-6-phosphate 1-dehydrogenase 4, chloroplast precursor
[Arabidopsis thaliana]
Length = 625
Score = 538 bits (1387), Expect = e-151
Identities = 264/484 (54%), Positives = 346/484 (70%), Gaps = 11/484 (2%)
Query: 91 SNLSITVVGASGDLAKKKIFPALFALFYEDCLPENFLVFGFARTKMTSEELRNMISRTLT 150
++L I VVGA+G+LA+ KIFPALFAL+Y LPE+ +FG +R +T E+LR++I+ TLT
Sbjct: 152 ASLCIAVVGATGELARGKIFPALFALYYSGYLPEDVAIFGVSRKNLTDEDLRSIIASTLT 211
Query: 151 CRIDKRANCEDKMDQFLKRCFYHSGLYNSEGSFSDLDCKMKEKEGGKRSNRLFYLSIPPN 210
CR+D + NC KMD F R +Y +G YN+ S L +MK+ EG +NR+FYLS+P
Sbjct: 212 CRVDHQENCGGKMDAFQSRTYYINGGYNNRDGMSRLAERMKQIEGESEANRIFYLSVPQE 271
Query: 211 IFVNVVRCASLKASSKKGWTRVIVEKPFGRDSESSNELTRCLKQYLTEDQIFRIDHYLGK 270
V+V A + +GWTR+IVEKPFG +S SS++LT+ L E QI+RIDH LG+
Sbjct: 272 ALVDVACTIGDNAQAPRGWTRIIVEKPFGFNSHSSHQLTKSLLSKFEEKQIYRIDHMLGR 331
Query: 271 ELVENLSVLRFSNLVFEPLWSRAYIRNVQFIFSEDFGTEGRGGYFDNYGIIRDIMQNHLL 330
L+ENL+VLRFSNLVFEPLW+R YIRN+Q I SE + + D YGIIRDI+ +H+L
Sbjct: 332 NLIENLTVLRFSNLVFEPLWNRTYIRNIQVIISESIAQTEK--FSDGYGIIRDIVHSHIL 389
Query: 331 QILALFAMEPPVSLDAEDIRNEKVKVLRSMRPLQLENVVTGQYKGHSKGGKSYPGYADDP 390
Q +AL AMEPP+SLD EDIRNEKVKVLRS+R + +V+ GQYK S+ D
Sbjct: 390 QTIALLAMEPPISLDGEDIRNEKVKVLRSIRKIDPRDVILGQYKSSSR---------DKN 440
Query: 391 SVPKGSLTPTFAAAALFIDNARWDGVPFLMKAGKALHTKRAEIRVQFRHVPGNLYKRNFG 450
V + PT+ AAAL+IDNARWDGVPFL++ G L R EI VQFRHVPGNLY+ N G
Sbjct: 441 GVILNGVDPTYCAAALYIDNARWDGVPFLVRVGTGLIKHRVEIHVQFRHVPGNLYRENIG 500
Query: 451 ADMDKATNELVLRVQPDEAIYLKVNNKIPGLGMRLDRSDLNLLYRARYPTEIPDAYERLL 510
++D TNEL+LR +PDEAI +K+NNK+PGLG++LD S+LNLLY+ RY TE+PD+YE L+
Sbjct: 501 INIDLGTNELILRDEPDEAILVKINNKVPGLGLQLDASELNLLYKDRYKTEVPDSYEHLI 560
Query: 511 LDAIEGERRLFIRSDELDAAWALFTPLLKEVEDKKIAPELYPHGSRGPIGAHYLAAKHNV 570
D I+G+ LF+RSDE+ AAW + +P+L+E++ APELY G RGP+ A+YL AKH V
Sbjct: 561 HDVIDGDNHLFMRSDEVAAAWNILSPVLEEIDKHHTAPELYEFGGRGPVAAYYLWAKHGV 620
Query: 571 RWGD 574
W D
Sbjct: 621 PWAD 624
>UniRef100_Q9STD4 Glucose-6-phosphate 1-dehydrogenase precursor [Cyanidium caldarium]
Length = 600
Score = 513 bits (1322), Expect = e-144
Identities = 259/489 (52%), Positives = 340/489 (68%), Gaps = 12/489 (2%)
Query: 93 LSITVVGASGDLAKKKIFPALFALFYEDCLPENFLVFGFARTKMTSEELRNMISRTLTCR 152
L I V+GASGDLAKKK FPALF+L+Y D LP++FL+ G+AR +MT EE RN I +LTCR
Sbjct: 115 LCIVVIGASGDLAKKKTFPALFSLYYHDLLPKDFLIVGYARRQMTQEEFRNSIMESLTCR 174
Query: 153 IDKRANCEDKMDQFLKRCFYHSGLYNSEGSFSDLDCKMKEKEGG---KRSNRLFYLSIPP 209
+ C+ KMD+FL +C Y SG+Y+ F LD + E R +RL+YL++P
Sbjct: 175 VIDGPQCQRKMDEFLPKCHYMSGMYDRTEDFVRLDQFLNNFEQSFPNTRVDRLYYLAVPS 234
Query: 210 NIFVNVVRCASLKASSKKGWTRVIVEKPFGRDSESSNELTRCLKQYLTEDQIFRIDHYLG 269
+F NVV +++GW R+++EKPFG+D S +L L+ ++ED+I+RIDHYLG
Sbjct: 235 QVFENVVHHVHESGRTQRGWNRIVMEKPFGKDITSYLQLRNSLRNCISEDEIYRIDHYLG 294
Query: 270 KELVENLSVLRFSNLVFEPLWSRAYIRNVQFIFSEDFGTEGRGGYFDNYGIIRDIMQNHL 329
KELV+NL VLRF+N +FEPLW+R +I ++Q +F E+FG EGR GYFD YGIIRDIMQNHL
Sbjct: 295 KELVQNLMVLRFANYLFEPLWNRDHIASIQIVFKENFGVEGRAGYFDEYGIIRDIMQNHL 354
Query: 330 LQILALFAMEPPVSLDAEDIRNEKVKVLRSMRPLQLENVVTGQYKGHSKGGKSYPGYADD 389
LQ++AL ME PV+L AEDIR+EKVK LRS+RPL+ + V GQY+ +S Y +
Sbjct: 355 LQVMALLGMEQPVTLHAEDIRDEKVKFLRSIRPLKASDFVLGQYRDRQNPQRS---YLSE 411
Query: 390 PSVPKGSLTPTFAAAALFIDNARWDGVPFLMKAGKALHTKRAEIRVQFRHVPGNLYKRNF 449
P V S TPTFAA +DN RW GVPFLMKAGKAL ++AEIR+QF+ VPG L+ +
Sbjct: 412 PGVMNDSHTPTFAACVFQVDNRRWSGVPFLMKAGKALDERKAEIRIQFQSVPGGLFSQVV 471
Query: 450 GADMDKATNELVLRVQPDEAIYLKVNNKIPGLGMRLDRSDLNLLYRARY--PTEIPDAYE 507
+ + NELV+ VQPDEAIY+++ +K PG RL+ + LNL YR + +IPDAYE
Sbjct: 472 SSHLPH--NELVIIVQPDEAIYMRILSKAPGFTSRLEEARLNLFYRTAWEDSKDIPDAYE 529
Query: 508 RLLLDAIEGERRLFIRSDELDAAWALFTPLLKEVE--DKKIAPELYPHGSRGPIGAHYLA 565
RL+LD I GE+ LFIR DEL+ AW +FTP LKE+E P LY +G RGPI + YLA
Sbjct: 530 RLILDVIHGEKSLFIRDDELEVAWNIFTPSLKEMEMAQDSWKPILYDYGGRGPIESDYLA 589
Query: 566 AKHNVRWGD 574
AK+ V+W +
Sbjct: 590 AKYAVQWSE 598
>UniRef100_Q869V9 Similar to Oryza sativa (Rice). glucose-6-phosphate dehydrogenase
[Dictyostelium discoideum]
Length = 497
Score = 482 bits (1241), Expect = e-134
Identities = 243/482 (50%), Positives = 335/482 (69%), Gaps = 19/482 (3%)
Query: 91 SNLSITVVGASGDLAKKKIFPALFALFYEDCLPENFLVFGFARTKMTSEELRNMISRTLT 150
S L++ ++GASGDLAKKK +PALF L+ D LP N +++G+AR+ + + + IS+ L
Sbjct: 9 SVLTVIILGASGDLAKKKTYPALFGLYLRDLLPSNTIIYGYARSHIEIGDFKARISKGLK 68
Query: 151 CRIDKRANCEDKMDQFLKRCFYHSGLYNSEGSFSDLD----CKMKEKEGGKRSNRLFYLS 206
E+K QFL YHSG Y+ + S+ + + + K+++G + NRLFY++
Sbjct: 69 -------GDEEKKKQFLNLLHYHSGKYDEKASYDEFEKLILAEEKKQQGVDKFNRLFYMA 121
Query: 207 IPPNIFVNVVRCASLKASSKKGWTRVIVEKPFGRDSESSNELTRCLKQYLTEDQIFRIDH 266
IPP+IF+ V SK GW+RVIVEKPFGRD SS EL L + E +FRIDH
Sbjct: 122 IPPSIFIEVSIGIHGSLISKNGWSRVIVEKPFGRDLASSRELVSELGKLFKEKDLFRIDH 181
Query: 267 YLGKELVENLSVLRFSNLVFEPLWSRAYIRNVQFIFSEDFGTEGRGGYFDNYGIIRDIMQ 326
YLGKE+V+NL VLRF+N VFEPLWS+++I ++ F ED GTEGRGGYFD +GIIRD+MQ
Sbjct: 182 YLGKEMVQNLMVLRFANAVFEPLWSKSHISSITITFKEDIGTEGRGGYFDQFGIIRDVMQ 241
Query: 327 NHLLQILALFAMEPPVSLDAEDIRNEKVKVLRSMRPLQLENVVTGQYKGHSKGGKSYPGY 386
NHLLQ+L+L AMEPPVSL+A+DI NEKVK+LR ++P+++ VV GQY +G P Y
Sbjct: 242 NHLLQVLSLVAMEPPVSLNADDITNEKVKLLRCIQPIKMSEVVLGQYTSDPEG--KIPAY 299
Query: 387 ADDPSVPKGSLTPTFAAAALFIDNARWDGVPFLMKAGKALHTKRAEIRVQFRHVPGNLYK 446
DD VPK S TPT+AAA I+N RW G+PF++K GKAL ++ E+R+QF+ L+
Sbjct: 300 LDDEGVPKDSTTPTYAAAVFHINNPRWRGMPFILKCGKALDERKTEVRIQFKRPDNFLF- 358
Query: 447 RNFGADMDKATNELVLRVQPDEAIYLKVNNKIPGLGMRLDRSDLNLLYRARYPT-EIPDA 505
+D D + NELV+R+QP EA+YLK+ +K PGL ++++++L+L YR R+ ++PDA
Sbjct: 359 ----SDDDISRNELVMRIQPGEAVYLKLLSKKPGLENKIEQTELDLSYRHRFENLDLPDA 414
Query: 506 YERLLLDAIEGERRLFIRSDELDAAWALFTPLLKEVEDKKIAPELYPHGSRGPIGAHYLA 565
YERL+LD+I+G+ LF+R DELD AW +FTPLL ++E +KI PE Y GSRGP A L+
Sbjct: 415 YERLILDSIKGDHNLFVRDDELDVAWQIFTPLLDQIEKEKIKPEPYSFGSRGPKSADELS 474
Query: 566 AK 567
K
Sbjct: 475 KK 476
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.320 0.138 0.412
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,005,594,497
Number of Sequences: 2790947
Number of extensions: 44319674
Number of successful extensions: 92665
Number of sequences better than 10.0: 520
Number of HSP's better than 10.0 without gapping: 511
Number of HSP's successfully gapped in prelim test: 9
Number of HSP's that attempted gapping in prelim test: 90654
Number of HSP's gapped (non-prelim): 559
length of query: 579
length of database: 848,049,833
effective HSP length: 133
effective length of query: 446
effective length of database: 476,853,882
effective search space: 212676831372
effective search space used: 212676831372
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 78 (34.7 bits)
Lotus: description of TM0213.11


