

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0203.8
(124 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
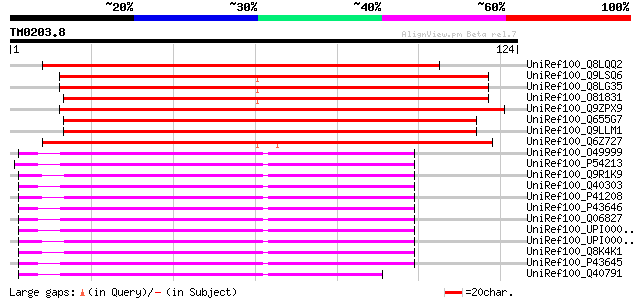
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q8LQQ2 Putative EF-hand Ca2+-binding protein CCD1 [Ory... 136 9e-32
UniRef100_Q9LSQ6 Arabidopsis thaliana genomic DNA, chromosome 5,... 136 1e-31
UniRef100_Q8LG35 EF-hand Ca2+-binding protein CCD1 [Arabidopsis ... 136 1e-31
UniRef100_O81831 Hypothetical protein AT4g27280 [Arabidopsis tha... 133 1e-30
UniRef100_Q9ZPX9 Putative caltractin [Arabidopsis thaliana] 130 7e-30
UniRef100_Q655G7 Putative EF-hand Ca2+-binding protein CCD1 [Ory... 125 2e-28
UniRef100_Q9LLM1 EF-hand Ca2+-binding protein CCD1 [Triticum aes... 125 2e-28
UniRef100_Q6Z727 Putative EF-hand Ca2+-binding protein CCD1 [Ory... 118 3e-26
UniRef100_O49999 Centrin [Marsilea vestita] 60 1e-08
UniRef100_P54213 Caltractin [Dunaliella salina] 58 6e-08
UniRef100_Q9R1K9 Centrin 2 [Mus musculus] 57 7e-08
UniRef100_Q40303 Centrin [Micromonas pusilla] 57 1e-07
UniRef100_P41208 Centrin 2 [Homo sapiens] 57 1e-07
UniRef100_P43646 Caltractin [Tetraselmis striata] 57 1e-07
UniRef100_Q06827 Caltractin [Scherffelia dubia] 57 1e-07
UniRef100_UPI00003AD860 UPI00003AD860 UniRef100 entry 57 1e-07
UniRef100_UPI00001CEE3A UPI00001CEE3A UniRef100 entry 56 2e-07
UniRef100_Q8K4K1 Centrin 4 [Mus musculus] 56 2e-07
UniRef100_P43645 Caltractin [Spermatozopsis similis] 56 2e-07
UniRef100_Q40791 Centrin [Pterosperma cristatum] 56 2e-07
>UniRef100_Q8LQQ2 Putative EF-hand Ca2+-binding protein CCD1 [Oryza sativa]
Length = 111
Score = 136 bits (343), Expect = 9e-32
Identities = 66/97 (68%), Positives = 79/97 (81%)
Query: 9 GKGLEFEDLLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAAVLGLQDMKE 68
G G++FED LP MA KLG EGLI+ELC GF++LMD G IT SLK NAA+LGL ++++
Sbjct: 7 GTGVQFEDFLPSMARKLGVEGLIEELCKGFELLMDPGAGKITFRSLKRNAAMLGLGELRD 66
Query: 69 DELVSMIREGDLDGDGALTQMEFCVLMFRLSPELMEE 105
DEL M+REGDLDGDGAL QMEFCVLM RLSPELM++
Sbjct: 67 DELSEMMREGDLDGDGALDQMEFCVLMVRLSPELMQD 103
>UniRef100_Q9LSQ6 Arabidopsis thaliana genomic DNA, chromosome 5, BAC clone:F24B18
[Arabidopsis thaliana]
Length = 127
Score = 136 bits (342), Expect = 1e-31
Identities = 67/106 (63%), Positives = 83/106 (78%), Gaps = 1/106 (0%)
Query: 13 EFEDLLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAA-VLGLQDMKEDEL 71
+F+D P MA KLGGEGLI+E+C GF++LMDKD+GVIT ESL+ NA+ VLGL D+ +D++
Sbjct: 18 QFQDFFPTMAGKLGGEGLIEEICKGFELLMDKDKGVITFESLRRNASTVLGLGDLTDDDV 77
Query: 72 VSMIREGDLDGDGALTQMEFCVLMFRLSPELMEESWIWLEEALQHE 117
MI EGD D DGAL QMEFCVLMFRLSPELME S + E ++ E
Sbjct: 78 RYMINEGDFDRDGALNQMEFCVLMFRLSPELMEASRCVVTEVIEEE 123
>UniRef100_Q8LG35 EF-hand Ca2+-binding protein CCD1 [Arabidopsis thaliana]
Length = 127
Score = 136 bits (342), Expect = 1e-31
Identities = 67/106 (63%), Positives = 83/106 (78%), Gaps = 1/106 (0%)
Query: 13 EFEDLLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAA-VLGLQDMKEDEL 71
+F+D P MA KLGGEGLI+E+C GF++LMDKD+GVIT ESL+ NA+ VLGL D+ +D++
Sbjct: 18 QFQDFFPTMAGKLGGEGLIEEICKGFELLMDKDKGVITFESLRRNASTVLGLGDLTDDDV 77
Query: 72 VSMIREGDLDGDGALTQMEFCVLMFRLSPELMEESWIWLEEALQHE 117
MI EGD D DGAL QMEFCVLMFRLSPELME S + E ++ E
Sbjct: 78 RYMINEGDFDRDGALNQMEFCVLMFRLSPELMEASRCVVTEVIEEE 123
>UniRef100_O81831 Hypothetical protein AT4g27280 [Arabidopsis thaliana]
Length = 130
Score = 133 bits (334), Expect = 1e-30
Identities = 66/105 (62%), Positives = 82/105 (77%), Gaps = 1/105 (0%)
Query: 14 FEDLLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAA-VLGLQDMKEDELV 72
F D LP MA LGGEGLI ELCNGF++LMD+++GVIT ESL+ NAA VLGL D+ ++++
Sbjct: 19 FHDFLPTMAGNLGGEGLIGELCNGFELLMDREKGVITFESLRRNAAAVLGLGDLTDEDVR 78
Query: 73 SMIREGDLDGDGALTQMEFCVLMFRLSPELMEESWIWLEEALQHE 117
MI+EGD D DGAL QMEFCVLMFRLSP+LME S + E ++ E
Sbjct: 79 CMIKEGDFDCDGALNQMEFCVLMFRLSPDLMEASRCLVTEVIEEE 123
>UniRef100_Q9ZPX9 Putative caltractin [Arabidopsis thaliana]
Length = 135
Score = 130 bits (327), Expect = 7e-30
Identities = 60/109 (55%), Positives = 82/109 (75%)
Query: 13 EFEDLLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAAVLGLQDMKEDELV 72
++ED+LPVMA K+ E + ELC GF +L D +R +IT ESL+ N+ +LG++ M +++
Sbjct: 21 KYEDMLPVMAEKMDVEEFVSELCKGFSLLADPERHLITAESLRRNSGILGIEGMSKEDAQ 80
Query: 73 SMIREGDLDGDGALTQMEFCVLMFRLSPELMEESWIWLEEALQHELDNN 121
M+REGDLDGDGAL Q EFCVLM RLSPE+ME++ WLE+AL EL N+
Sbjct: 81 GMVREGDLDGDGALNQTEFCVLMVRLSPEMMEDAETWLEKALTQELCNH 129
>UniRef100_Q655G7 Putative EF-hand Ca2+-binding protein CCD1 [Oryza sativa]
Length = 125
Score = 125 bits (314), Expect = 2e-28
Identities = 60/101 (59%), Positives = 79/101 (77%)
Query: 14 FEDLLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAAVLGLQDMKEDELVS 73
FED LPVMA +LG EGL++EL +GF +LMD G+IT +SL+ NA +LGL M +D+L
Sbjct: 16 FEDYLPVMAERLGEEGLMQELASGFRLLMDPASGLITFDSLRRNAPLLGLGGMSDDDLRG 75
Query: 74 MIREGDLDGDGALTQMEFCVLMFRLSPELMEESWIWLEEAL 114
M+ EGD DGDGAL++MEFCVLM RLSP+LM+E WL++A+
Sbjct: 76 MLAEGDFDGDGALSEMEFCVLMVRLSPDLMDEPRRWLDDAV 116
>UniRef100_Q9LLM1 EF-hand Ca2+-binding protein CCD1 [Triticum aestivum]
Length = 129
Score = 125 bits (314), Expect = 2e-28
Identities = 61/101 (60%), Positives = 78/101 (76%)
Query: 14 FEDLLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAAVLGLQDMKEDELVS 73
FED LPVMA +LG EGL++EL GF +LMD G+IT +SL+ NA +LGL M +D+L
Sbjct: 19 FEDYLPVMAERLGEEGLMEELAAGFRLLMDPASGLITFDSLRRNAPLLGLGGMSDDDLRG 78
Query: 74 MIREGDLDGDGALTQMEFCVLMFRLSPELMEESWIWLEEAL 114
M+ EGD DGDGAL+QMEFCVLM RLSP+LM+E WL++A+
Sbjct: 79 MLAEGDFDGDGALSQMEFCVLMVRLSPDLMDEPRRWLDDAV 119
>UniRef100_Q6Z727 Putative EF-hand Ca2+-binding protein CCD1 [Oryza sativa]
Length = 147
Score = 118 bits (296), Expect = 3e-26
Identities = 65/115 (56%), Positives = 82/115 (70%), Gaps = 5/115 (4%)
Query: 9 GKGLEFEDLLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAA-VLGLQ--- 64
G G E+EDL+PVMA +LG EGL+ EL GF +L D RG IT ESL+ +AA VLGL
Sbjct: 23 GGGEEYEDLMPVMAGRLGAEGLLSELRAGFRLLADPARGAITAESLRRSAASVLGLGGGG 82
Query: 65 -DMKEDELVSMIREGDLDGDGALTQMEFCVLMFRLSPELMEESWIWLEEALQHEL 118
+M +E +M+REGD DGDGAL++ EFCVLM RLSP +M ++ WLEEA+ EL
Sbjct: 83 GEMTVEEAAAMVREGDQDGDGALSEAEFCVLMVRLSPGIMGDAEGWLEEAIADEL 137
>UniRef100_O49999 Centrin [Marsilea vestita]
Length = 170
Score = 59.7 bits (143), Expect = 1e-08
Identities = 35/97 (36%), Positives = 55/97 (56%), Gaps = 6/97 (6%)
Query: 3 GTGTVNGKGLEFEDLLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAAVLG 62
G+GT+ +FED L +M K+G +E+ F + D + G I+ ++LK A LG
Sbjct: 78 GSGTI-----DFEDFLQMMTTKMGERDSKEEIMKAFRLFDDDETGKISFKNLKRVAKELG 132
Query: 63 LQDMKEDELVSMIREGDLDGDGALTQMEFCVLMFRLS 99
++M ++EL MI E D DGDG + + EF +M + S
Sbjct: 133 -ENMTDEELQEMIDEADRDGDGEINEEEFYRIMKKTS 168
Score = 33.5 bits (75), Expect = 1.1
Identities = 20/64 (31%), Positives = 30/64 (46%), Gaps = 1/64 (1%)
Query: 32 KELCNGFDMLMDKDRGVITLESLKTNAAVLGLQDMKEDELVSMIREGDLDGDGALTQMEF 91
+E+ FD+ G I + LK LG + KE E+ MI + D DG G + +F
Sbjct: 29 QEIREAFDLFDTDGSGTIDAKELKVAMRALGFEPKKE-EIKKMIADIDKDGSGTIDFEDF 87
Query: 92 CVLM 95
+M
Sbjct: 88 LQMM 91
>UniRef100_P54213 Caltractin [Dunaliella salina]
Length = 169
Score = 57.8 bits (138), Expect = 6e-08
Identities = 34/98 (34%), Positives = 57/98 (57%), Gaps = 6/98 (6%)
Query: 2 AGTGTVNGKGLEFEDLLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAAVL 61
AG+GT+ +FE+ L +M +K+G +E+ F + D + G ITL++LK A L
Sbjct: 76 AGSGTI-----DFEEFLQMMTSKMGERDSREEIIKAFKLFDDDNTGFITLKNLKRVAKEL 130
Query: 62 GLQDMKEDELVSMIREGDLDGDGALTQMEFCVLMFRLS 99
G +++ ++EL M E D +GDG + + EF +M + S
Sbjct: 131 G-ENLTDEELQEMTDEADRNGDGQIDEDEFYRIMKKTS 167
Score = 31.6 bits (70), Expect = 4.3
Identities = 20/64 (31%), Positives = 29/64 (45%), Gaps = 1/64 (1%)
Query: 32 KELCNGFDMLMDKDRGVITLESLKTNAAVLGLQDMKEDELVSMIREGDLDGDGALTQMEF 91
+E+ FD+ G I + LK LG + KE E+ MI + D G G + EF
Sbjct: 28 QEIREAFDLFDTDGSGTIDAKELKVAMRALGFEPKKE-EIKKMIADIDKAGSGTIDFEEF 86
Query: 92 CVLM 95
+M
Sbjct: 87 LQMM 90
>UniRef100_Q9R1K9 Centrin 2 [Mus musculus]
Length = 172
Score = 57.4 bits (137), Expect = 7e-08
Identities = 34/97 (35%), Positives = 53/97 (54%), Gaps = 6/97 (6%)
Query: 3 GTGTVNGKGLEFEDLLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAAVLG 62
GTG +N F D L VM K+ + +E+ F + D + G I+ ++LK A LG
Sbjct: 80 GTGKMN-----FSDFLTVMTQKMSEKDTKEEILKAFKLFDDDETGKISFKNLKRVAKELG 134
Query: 63 LQDMKEDELVSMIREGDLDGDGALTQMEFCVLMFRLS 99
+++ ++EL MI E D DGDG + + EF +M + S
Sbjct: 135 -ENLTDEELQEMIDEADRDGDGEVNEQEFLRIMKKTS 170
Score = 34.3 bits (77), Expect = 0.66
Identities = 20/64 (31%), Positives = 31/64 (48%), Gaps = 1/64 (1%)
Query: 32 KELCNGFDMLMDKDRGVITLESLKTNAAVLGLQDMKEDELVSMIREGDLDGDGALTQMEF 91
+E+ FD+ G I ++ LK LG + KE E+ MI E D +G G + +F
Sbjct: 31 QEIREAFDLFDADGTGTIDIKELKVAMRALGFEPKKE-EIKKMISEIDKEGTGKMNFSDF 89
Query: 92 CVLM 95
+M
Sbjct: 90 LTVM 93
>UniRef100_Q40303 Centrin [Micromonas pusilla]
Length = 148
Score = 57.0 bits (136), Expect = 1e-07
Identities = 34/97 (35%), Positives = 55/97 (56%), Gaps = 6/97 (6%)
Query: 3 GTGTVNGKGLEFEDLLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAAVLG 62
G+GT+ +FE+ L +M K+G +E+ F + D + G I+ ++LK A LG
Sbjct: 56 GSGTI-----DFEEFLTMMTAKMGERDSREEIMKAFRLFDDDETGKISFKNLKRVAKELG 110
Query: 63 LQDMKEDELVSMIREGDLDGDGALTQMEFCVLMFRLS 99
++M ++EL MI E D DGDG + + EF +M + S
Sbjct: 111 -ENMTDEELQEMIDEADRDGDGEVNEEEFFRIMKKTS 146
Score = 35.0 bits (79), Expect = 0.39
Identities = 21/64 (32%), Positives = 30/64 (46%), Gaps = 1/64 (1%)
Query: 32 KELCNGFDMLMDKDRGVITLESLKTNAAVLGLQDMKEDELVSMIREGDLDGDGALTQMEF 91
+E+ FD+ G I + LK LG + KE E+ MI + D DG G + EF
Sbjct: 7 QEIRXAFDLFDTDGSGTIDAKELKVAMRALGFEPKKE-EIKKMIADIDKDGSGTIDFEEF 65
Query: 92 CVLM 95
+M
Sbjct: 66 LTMM 69
>UniRef100_P41208 Centrin 2 [Homo sapiens]
Length = 172
Score = 57.0 bits (136), Expect = 1e-07
Identities = 34/97 (35%), Positives = 54/97 (55%), Gaps = 6/97 (6%)
Query: 3 GTGTVNGKGLEFEDLLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAAVLG 62
GTG +N F D L VM K+ + +E+ F + D + G I+ ++LK A LG
Sbjct: 80 GTGKMN-----FGDFLTVMTQKMSEKDTKEEILKAFKLFDDDETGKISFKNLKRVAKELG 134
Query: 63 LQDMKEDELVSMIREGDLDGDGALTQMEFCVLMFRLS 99
+++ ++EL MI E D DGDG +++ EF +M + S
Sbjct: 135 -ENLTDEELQEMIDEADRDGDGEVSEQEFLRIMKKTS 170
Score = 33.1 bits (74), Expect = 1.5
Identities = 20/64 (31%), Positives = 31/64 (48%), Gaps = 1/64 (1%)
Query: 32 KELCNGFDMLMDKDRGVITLESLKTNAAVLGLQDMKEDELVSMIREGDLDGDGALTQMEF 91
+E+ FD+ G I ++ LK LG + KE E+ MI E D +G G + +F
Sbjct: 31 QEIREAFDLFDADGTGTIDVKELKVAMRALGFEPKKE-EIKKMISEIDKEGTGKMNFGDF 89
Query: 92 CVLM 95
+M
Sbjct: 90 LTVM 93
>UniRef100_P43646 Caltractin [Tetraselmis striata]
Length = 148
Score = 57.0 bits (136), Expect = 1e-07
Identities = 34/97 (35%), Positives = 55/97 (56%), Gaps = 6/97 (6%)
Query: 3 GTGTVNGKGLEFEDLLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAAVLG 62
G+GT+ +FE+ L +M K+G +E+ F + D + G I+ ++LK A LG
Sbjct: 56 GSGTI-----DFEEFLQMMTAKMGERDSREEIMKAFRLFDDDETGKISFKNLKRVAKELG 110
Query: 63 LQDMKEDELVSMIREGDLDGDGALTQMEFCVLMFRLS 99
++M ++EL MI E D DGDG + + EF +M + S
Sbjct: 111 -ENMTDEELQEMIDEADRDGDGEVNEEEFFRIMKKTS 146
Score = 33.5 bits (75), Expect = 1.1
Identities = 20/64 (31%), Positives = 30/64 (46%), Gaps = 1/64 (1%)
Query: 32 KELCNGFDMLMDKDRGVITLESLKTNAAVLGLQDMKEDELVSMIREGDLDGDGALTQMEF 91
+++ FD+ G I + LK LG + KE E+ MI + D DG G + EF
Sbjct: 7 QDIREAFDLFDTDGSGTIDAKELKVAMRALGFEPKKE-EIKKMIADIDKDGSGTIDFEEF 65
Query: 92 CVLM 95
+M
Sbjct: 66 LQMM 69
>UniRef100_Q06827 Caltractin [Scherffelia dubia]
Length = 168
Score = 57.0 bits (136), Expect = 1e-07
Identities = 34/97 (35%), Positives = 55/97 (56%), Gaps = 6/97 (6%)
Query: 3 GTGTVNGKGLEFEDLLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAAVLG 62
G+GT+ +FE+ L +M K+G +E+ F + D + G I+ ++LK A LG
Sbjct: 76 GSGTI-----DFEEFLQMMTAKMGERDSREEIMKAFRLFDDDETGKISFKNLKRVAKELG 130
Query: 63 LQDMKEDELVSMIREGDLDGDGALTQMEFCVLMFRLS 99
++M ++EL MI E D DGDG + + EF +M + S
Sbjct: 131 -ENMTDEELQEMIDEADRDGDGEVNEEEFFRIMKKTS 166
Score = 34.7 bits (78), Expect = 0.51
Identities = 21/64 (32%), Positives = 30/64 (46%), Gaps = 1/64 (1%)
Query: 32 KELCNGFDMLMDKDRGVITLESLKTNAAVLGLQDMKEDELVSMIREGDLDGDGALTQMEF 91
+E+ FD+ G I + LK LG + KE E+ MI + D DG G + EF
Sbjct: 27 QEIREAFDLFDTDGSGTIDAKELKVAMRALGFEPKKE-EIKKMIADIDKDGSGTIDFEEF 85
Query: 92 CVLM 95
+M
Sbjct: 86 LQMM 89
>UniRef100_UPI00003AD860 UPI00003AD860 UniRef100 entry
Length = 170
Score = 56.6 bits (135), Expect = 1e-07
Identities = 33/97 (34%), Positives = 55/97 (56%), Gaps = 6/97 (6%)
Query: 3 GTGTVNGKGLEFEDLLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAAVLG 62
G+GT+ +FED L +M K+ + +E+ F + D G I+ ++LK A LG
Sbjct: 78 GSGTI-----DFEDFLAMMTQKMSEKDSKEEILKAFRLFDDDGTGKISFKNLKRVAKELG 132
Query: 63 LQDMKEDELVSMIREGDLDGDGALTQMEFCVLMFRLS 99
+++ ++EL MI E D DGDG +++ EF +M + S
Sbjct: 133 -ENLTDEELQEMIDEADRDGDGEVSEQEFLRIMKKTS 168
Score = 33.1 bits (74), Expect = 1.5
Identities = 19/64 (29%), Positives = 31/64 (47%), Gaps = 1/64 (1%)
Query: 32 KELCNGFDMLMDKDRGVITLESLKTNAAVLGLQDMKEDELVSMIREGDLDGDGALTQMEF 91
+E+ FD+ G I ++ LK LG + KE E+ MI + D +G G + +F
Sbjct: 29 QEIREAFDLFDTDGSGSIDIKELKVAMRALGFEPKKE-EIKKMIADIDKEGSGTIDFEDF 87
Query: 92 CVLM 95
+M
Sbjct: 88 LAMM 91
>UniRef100_UPI00001CEE3A UPI00001CEE3A UniRef100 entry
Length = 337
Score = 56.2 bits (134), Expect = 2e-07
Identities = 34/97 (35%), Positives = 53/97 (54%), Gaps = 6/97 (6%)
Query: 3 GTGTVNGKGLEFEDLLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAAVLG 62
GTGT+ FED +M+ K+ + +E+ F + D G I+L ++K A LG
Sbjct: 245 GTGTIC-----FEDFFAIMSIKMSEKDEKEEILKAFKLFDDDATGSISLNNIKRVAKELG 299
Query: 63 LQDMKEDELVSMIREGDLDGDGALTQMEFCVLMFRLS 99
+++ EDEL M+ E D DGDG + + EF +M + S
Sbjct: 300 -ENLTEDELQEMLDEADRDGDGEINEEEFLKMMRKTS 335
>UniRef100_Q8K4K1 Centrin 4 [Mus musculus]
Length = 168
Score = 56.2 bits (134), Expect = 2e-07
Identities = 34/97 (35%), Positives = 53/97 (54%), Gaps = 6/97 (6%)
Query: 3 GTGTVNGKGLEFEDLLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAAVLG 62
GTGT+ FED +M+ K+ + +E+ F + D G I+L ++K A LG
Sbjct: 76 GTGTIC-----FEDFFAIMSVKMSEKDEKEEILKAFKLFDDDATGSISLNNIKRVAKELG 130
Query: 63 LQDMKEDELVSMIREGDLDGDGALTQMEFCVLMFRLS 99
+++ EDEL M+ E D DGDG + + EF +M + S
Sbjct: 131 -ENLTEDELQEMLDEADRDGDGEINEEEFLKMMKKTS 166
Score = 32.3 bits (72), Expect = 2.5
Identities = 20/64 (31%), Positives = 31/64 (48%), Gaps = 1/64 (1%)
Query: 32 KELCNGFDMLMDKDRGVITLESLKTNAAVLGLQDMKEDELVSMIREGDLDGDGALTQMEF 91
+E+ FD+ G I L+ LK LG + KE E+ +I E D +G G + +F
Sbjct: 27 QEIKEAFDLFDIDGSGTIDLKELKIAMRALGFEPKKE-EVKQLIAEIDKEGTGTICFEDF 85
Query: 92 CVLM 95
+M
Sbjct: 86 FAIM 89
>UniRef100_P43645 Caltractin [Spermatozopsis similis]
Length = 148
Score = 56.2 bits (134), Expect = 2e-07
Identities = 33/97 (34%), Positives = 54/97 (55%), Gaps = 6/97 (6%)
Query: 3 GTGTVNGKGLEFEDLLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAAVLG 62
G+GT+ +FE+ L +M K+G +E+ F + D G IT ++LK A LG
Sbjct: 56 GSGTI-----DFEEFLQMMTAKMGERDSREEIMKAFRLFDDDQTGKITFKNLKRVAKELG 110
Query: 63 LQDMKEDELVSMIREGDLDGDGALTQMEFCVLMFRLS 99
+++ ++E+ MI E D DGDG + + EF +M + S
Sbjct: 111 -ENLTDEEIQEMIDEADRDGDGEINEEEFFRIMKKTS 146
Score = 34.3 bits (77), Expect = 0.66
Identities = 21/64 (32%), Positives = 29/64 (44%), Gaps = 1/64 (1%)
Query: 32 KELCNGFDMLMDKDRGVITLESLKTNAAVLGLQDMKEDELVSMIREGDLDGDGALTQMEF 91
+E FD+ G I + LK LG + KE E+ MI + D DG G + EF
Sbjct: 7 QEXREAFDLFDTDGSGTIDAKELKVXMXALGFEPKKE-EIQKMISDIDKDGSGTIDFEEF 65
Query: 92 CVLM 95
+M
Sbjct: 66 LQMM 69
>UniRef100_Q40791 Centrin [Pterosperma cristatum]
Length = 133
Score = 55.8 bits (133), Expect = 2e-07
Identities = 32/89 (35%), Positives = 51/89 (56%), Gaps = 6/89 (6%)
Query: 3 GTGTVNGKGLEFEDLLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAAVLG 62
G+GT+ +FE+ L +M K+G +E+ F + D + G I+ ++LK A LG
Sbjct: 48 GSGTI-----DFEEFLQMMTAKMGERDSREEIMKAFRLFDDDETGKISFKNLKRVAKELG 102
Query: 63 LQDMKEDELVSMIREGDLDGDGALTQMEF 91
++M ++EL MI E D DGDG + + EF
Sbjct: 103 -ENMSDEELQEMIDEADRDGDGEVNEEEF 130
Score = 32.7 bits (73), Expect = 1.9
Identities = 20/58 (34%), Positives = 27/58 (46%), Gaps = 1/58 (1%)
Query: 38 FDMLMDKDRGVITLESLKTNAAVLGLQDMKEDELVSMIREGDLDGDGALTQMEFCVLM 95
FD+ G I + LK LG + KE E+ MI + D DG G + EF +M
Sbjct: 5 FDLFDTDGSGTIDAKELKVPMRALGFEPKKE-EIKKMIADIDKDGSGTIDFEEFLQMM 61
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.317 0.138 0.398
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 218,520,040
Number of Sequences: 2790947
Number of extensions: 8403196
Number of successful extensions: 21849
Number of sequences better than 10.0: 744
Number of HSP's better than 10.0 without gapping: 315
Number of HSP's successfully gapped in prelim test: 429
Number of HSP's that attempted gapping in prelim test: 20889
Number of HSP's gapped (non-prelim): 1321
length of query: 124
length of database: 848,049,833
effective HSP length: 100
effective length of query: 24
effective length of database: 568,955,133
effective search space: 13654923192
effective search space used: 13654923192
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 67 (30.4 bits)
Lotus: description of TM0203.8


