

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0201.10
(137 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
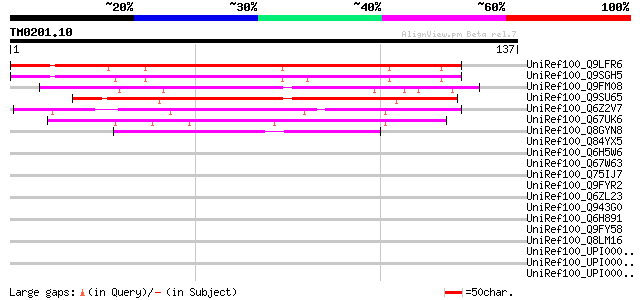
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9LFR6 Hypothetical protein F2G14_10 [Arabidopsis thal... 126 9e-29
UniRef100_Q9SGH5 T13O15.7 protein [Arabidopsis thaliana] 120 9e-27
UniRef100_Q9FM08 Arabidopsis thaliana genomic DNA, chromosome 5,... 99 2e-20
UniRef100_Q9SU65 Hypothetical protein T17F15.110 [Arabidopsis th... 96 2e-19
UniRef100_Q6Z2V7 NHL repeat-containing protein-like [Oryza sativa] 94 7e-19
UniRef100_Q67UK6 NHL repeat-containing protein-like [Oryza sativa] 86 2e-16
UniRef100_Q8GYN8 Hypothetical protein At5g25240/F21J6_107 [Arabi... 51 6e-06
UniRef100_Q84YX5 Hypothetical protein OSJNBb0075O18.125 [Oryza s... 42 0.003
UniRef100_Q6H5W6 Hypothetical protein OSJNBa0014E22.16 [Oryza sa... 39 0.033
UniRef100_Q67W63 Hypothetical protein OJ1226_A12.9 [Oryza sativa] 39 0.033
UniRef100_Q75IJ7 Hypothetical protein B1130G10.5 [Oryza sativa] 38 0.056
UniRef100_Q9FYR2 Arabidopsis thaliana genomic DNA, chromosome 5,... 36 0.16
UniRef100_Q6ZL23 Hypothetical protein OJ1699_E05.13 [Oryza sativa] 35 0.28
UniRef100_Q943G0 P0046E05.20 protein [Oryza sativa] 35 0.36
UniRef100_Q6H891 Hypothetical protein OJ1572_F02.1 [Oryza sativa] 35 0.36
UniRef100_Q9FY58 Hypothetical protein T5K6_60 [Arabidopsis thali... 34 0.62
UniRef100_Q8LM16 Hypothetical protein OSJNAb0008A05.22 [Oryza sa... 33 1.1
UniRef100_UPI000046B0F8 UPI000046B0F8 UniRef100 entry 33 1.8
UniRef100_UPI00002C2001 UPI00002C2001 UniRef100 entry 33 1.8
UniRef100_UPI0000468D60 UPI0000468D60 UniRef100 entry 32 2.3
>UniRef100_Q9LFR6 Hypothetical protein F2G14_10 [Arabidopsis thaliana]
Length = 733
Score = 126 bits (317), Expect = 9e-29
Identities = 68/140 (48%), Positives = 87/140 (61%), Gaps = 19/140 (13%)
Query: 1 MQEVLFAKQACICYLIPCSSSSETS----LSWWERVRSP---ENKEWWWAHGWNKVREWS 53
M E +FAK+ C C+++PC SS+ S WW+R+R+ E E WW GW K+REWS
Sbjct: 579 MHEAMFAKRGC-CFILPCLGSSQPSGPNGSVWWQRIRTVDKLEPDERWWVSGWMKMREWS 637
Query: 54 EIIVGPKWKTFIRRFNRNN----NRAGAASYDKKGSFHYDSLSYALNFDDGE-----EDV 104
EI+ GPKWKTFIRRF RN+ G + + SF YDS SY+LNFDDG+ ED
Sbjct: 638 EIVAGPKWKTFIRRFGRNHCCNGGIDGGCNRPEHVSFRYDSWSYSLNFDDGKQTGHFEDE 697
Query: 105 YSYGGFSTRFA--SVPASTK 122
+ Y +S RFA S+P STK
Sbjct: 698 FPYRDYSMRFAAPSLPVSTK 717
>UniRef100_Q9SGH5 T13O15.7 protein [Arabidopsis thaliana]
Length = 180
Score = 120 bits (300), Expect = 9e-27
Identities = 66/153 (43%), Positives = 88/153 (57%), Gaps = 32/153 (20%)
Query: 1 MQEVLFAKQACICYLIPCSSSSETSLS----WWERVRSP---ENKEWWWAHGWNKVREWS 53
M E LFAK+ C C+L+PC +SS+ S WW+R+ + E E WW GW ++REWS
Sbjct: 17 MHEALFAKRGC-CFLMPCLASSQPSTRGGSVWWQRITTVDKLEPDERWWIRGWRRMREWS 75
Query: 54 EIIVGPKWKTFIRRFNRNN------NRAGAAS-----------YDKKGSFHYDSLSYALN 96
E++ GP+WKT+IRRF R+N R G +S +G F YD LSY+LN
Sbjct: 76 ELVAGPRWKTYIRRFGRSNCCGGGGGRVGNSSGGCGGGAMPNRSSDQGKFRYDQLSYSLN 135
Query: 97 FDDGE-----EDVYSYGGFSTRFA--SVPASTK 122
FDDG +D + Y +S RFA S+P STK
Sbjct: 136 FDDGNQTGHFDDEFPYRDYSMRFAAPSLPVSTK 168
>UniRef100_Q9FM08 Arabidopsis thaliana genomic DNA, chromosome 5, P1 clone:MQB2
[Arabidopsis thaliana]
Length = 167
Score = 99.0 bits (245), Expect = 2e-20
Identities = 60/137 (43%), Positives = 76/137 (54%), Gaps = 20/137 (14%)
Query: 9 QACICYLIPCSSSSETSLSW--WERVRSPENKEW---------WWAHGWNKVREWSEIIV 57
Q C C+ S S T++ + W R+R+ ++ WW K+REWSEI+
Sbjct: 18 QRCCCFPSFRRSRSSTAVGYSSWGRIRTVDDSNHSGDHGDEPRWWIRASLKIREWSEIVA 77
Query: 58 GPKWKTFIRRFNRNNNRAGAASYDKKGSFHYDSLSYALNF-DDGEEDVY-SYGG---FST 112
GP+WKTFIRRFNR+ R +D F YD LSY+LNF DD EED Y GG FST
Sbjct: 78 GPRWKTFIRRFNRDPRR--GRDWDASEKFQYDPLSYSLNFDDDDEEDEYVGLGGLRSFST 135
Query: 113 RFASVP--ASTKPTFTP 127
RFASVP + P +P
Sbjct: 136 RFASVPVYSGKAPAISP 152
>UniRef100_Q9SU65 Hypothetical protein T17F15.110 [Arabidopsis thaliana]
Length = 135
Score = 95.9 bits (237), Expect = 2e-19
Identities = 48/111 (43%), Positives = 68/111 (61%), Gaps = 10/111 (9%)
Query: 18 CSSSSETSLSWWERVRSPENKE-WWWAHGWNKVREWSEIIVGPKWKTFIRRFNRNNNRAG 76
C S++ S SWW+R+ ++E WW + K+REWSEI+ GP+WKTFIRRFNR+ R
Sbjct: 15 CCSTTVKS-SWWQRIHRNNHQEPRWWVRAFLKIREWSEIVAGPRWKTFIRRFNRDPRR-- 71
Query: 77 AASYDKKGSFHYDSLSYALNFDDGEED------VYSYGGFSTRFASVPAST 121
+D F YD +SY L+F+D ++D V FS R+ASVP ++
Sbjct: 72 GQDWDDSDKFRYDPVSYTLSFEDEDKDDDDEAGVGGVRSFSMRYASVPVAS 122
>UniRef100_Q6Z2V7 NHL repeat-containing protein-like [Oryza sativa]
Length = 171
Score = 94.0 bits (232), Expect = 7e-19
Identities = 57/136 (41%), Positives = 73/136 (52%), Gaps = 23/136 (16%)
Query: 2 QEVLFAKQA--CICYLIPCSSSSETSLSWWERVRSPENKEWWW--AHGWNKVREWSEIIV 57
+ FA++ C C+ P S+SS +RV E + WW KVREWSE++
Sbjct: 18 EAAFFARRGRRCCCFPWPSSASSH------QRVGGAEEESWWQRAVDAVLKVREWSELVA 71
Query: 58 GPKWKTFIRRFNRNNNRAGAA------SYDKKGSFHYDSLSYALNFDDG-----EEDVYS 106
GP+WKTFIRRF R G +Y +K +YD+LSYALNFD+G E D
Sbjct: 72 GPRWKTFIRRFGRGGGGGGGGGGPRPHNYGRK--LNYDALSYALNFDEGHGASPEGDYTG 129
Query: 107 YGGFSTRFASVPASTK 122
Y FS RFA+ PAS K
Sbjct: 130 YRDFSARFAAPPASAK 145
>UniRef100_Q67UK6 NHL repeat-containing protein-like [Oryza sativa]
Length = 211
Score = 85.9 bits (211), Expect = 2e-16
Identities = 52/132 (39%), Positives = 66/132 (49%), Gaps = 24/132 (18%)
Query: 11 CICYLIPCSSSSETSLS---------WWERVRSPEN-KEWWWAHGWN---KVREWSEIIV 57
C C+ P +SSS + WW RV + WW G + KVREWSE++
Sbjct: 36 CCCFWGPWASSSYSRAGGPAAAAEEEWWHRVGGGGGERRRWWRRGVDALMKVREWSELVA 95
Query: 58 GPKWKTFIRRFNR----NNNRAGAASYDKKGSFHYDSLSYALNFDDG-------EEDVYS 106
GP+WKTFIRRF R +++ G +YD LSYALNFD+G E D
Sbjct: 96 GPRWKTFIRRFRRSPRHHHHGGGGGGGGGGRKLNYDPLSYALNFDEGHGGACSPEGDYAG 155
Query: 107 YGGFSTRFASVP 118
Y FSTRF + P
Sbjct: 156 YRDFSTRFVAPP 167
>UniRef100_Q8GYN8 Hypothetical protein At5g25240/F21J6_107 [Arabidopsis thaliana]
Length = 131
Score = 50.8 bits (120), Expect = 6e-06
Identities = 27/72 (37%), Positives = 36/72 (49%), Gaps = 5/72 (6%)
Query: 29 WERVRSPENKEWWWAHGWNKVREWSEIIVGPKWKTFIRRFNRNNNRAGAASYDKKGSFHY 88
W E + W + ++E SE I GPKWK FIR F+ +G + F Y
Sbjct: 46 WSGCLQEERRGNWGSEKLKGLKEISEKIAGPKWKNFIRSFS-----SGRKKMRRDVDFTY 100
Query: 89 DSLSYALNFDDG 100
D +Y+LNFDDG
Sbjct: 101 DLKNYSLNFDDG 112
>UniRef100_Q84YX5 Hypothetical protein OSJNBb0075O18.125 [Oryza sativa]
Length = 132
Score = 42.0 bits (97), Expect = 0.003
Identities = 24/49 (48%), Positives = 27/49 (54%), Gaps = 4/49 (8%)
Query: 84 GSFHYDSLSYALNFD---DGEEDVYSYGGFSTRF-ASVPASTKPTFTPV 128
G F YD +SYALNF+ DGE + Y FS R AS P PT PV
Sbjct: 80 GEFRYDPVSYALNFEEDGDGEAQPFKYMAFSARLPASPPPPPPPTALPV 128
>UniRef100_Q6H5W6 Hypothetical protein OSJNBa0014E22.16 [Oryza sativa]
Length = 81
Score = 38.5 bits (88), Expect = 0.033
Identities = 20/37 (54%), Positives = 22/37 (59%), Gaps = 1/37 (2%)
Query: 65 IRRFNRNNNR-AGAASYDKKGSFHYDSLSYALNFDDG 100
IRR R G + SFHYD+LSYALNFDDG
Sbjct: 36 IRRMQSERGRWRGGRRDHARFSFHYDALSYALNFDDG 72
>UniRef100_Q67W63 Hypothetical protein OJ1226_A12.9 [Oryza sativa]
Length = 102
Score = 38.5 bits (88), Expect = 0.033
Identities = 20/46 (43%), Positives = 24/46 (51%), Gaps = 9/46 (19%)
Query: 84 GSFHYDSLSYALNFDDGEED---------VYSYGGFSTRFASVPAS 120
G F YD LSYALNFDDG+ D + Y FS+R P +
Sbjct: 45 GEFGYDPLSYALNFDDGDGDDDAADDAAAAFRYKNFSSRLPPSPVA 90
>UniRef100_Q75IJ7 Hypothetical protein B1130G10.5 [Oryza sativa]
Length = 119
Score = 37.7 bits (86), Expect = 0.056
Identities = 20/53 (37%), Positives = 26/53 (48%), Gaps = 7/53 (13%)
Query: 61 WKTFIRRFNRNN-------NRAGAASYDKKGSFHYDSLSYALNFDDGEEDVYS 106
W +RR R + +R G A +FHYD+ SYA NFDDG Y+
Sbjct: 54 WSRLLRRLVRESRSFCSLGSRHGGAMAAATTTFHYDAASYAKNFDDGRRAHYA 106
>UniRef100_Q9FYR2 Arabidopsis thaliana genomic DNA, chromosome 5, BAC clone:T13C12
[Arabidopsis thaliana]
Length = 159
Score = 36.2 bits (82), Expect = 0.16
Identities = 19/40 (47%), Positives = 22/40 (54%), Gaps = 4/40 (10%)
Query: 86 FHYDSLSYALNFDDGEE----DVYSYGGFSTRFASVPAST 121
FHYD SYALNFD G+E D + FS R P S+
Sbjct: 105 FHYDPSSYALNFDKGDEDDNIDRFPLRNFSARLPHSPPSS 144
>UniRef100_Q6ZL23 Hypothetical protein OJ1699_E05.13 [Oryza sativa]
Length = 119
Score = 35.4 bits (80), Expect = 0.28
Identities = 17/42 (40%), Positives = 23/42 (54%), Gaps = 2/42 (4%)
Query: 60 KWKTFIRRF--NRNNNRAGAASYDKKGSFHYDSLSYALNFDD 99
+WK + R+ G + +K +F YDS SYALNFDD
Sbjct: 72 RWKAVAQEIMARRSGGGGGGSGRRRKTAFSYDSKSYALNFDD 113
>UniRef100_Q943G0 P0046E05.20 protein [Oryza sativa]
Length = 150
Score = 35.0 bits (79), Expect = 0.36
Identities = 23/69 (33%), Positives = 34/69 (48%), Gaps = 3/69 (4%)
Query: 43 AHGWNKVREWSEIIVGPKWKTFIRRFNRNNNRAGAASYDKKGSFHYDSLSYALNFDDGEE 102
A GW R W ++ +T R +++ +GAAS + +F YD+ SYA NFDDG
Sbjct: 67 AAGWCGHRAWRRLLRRLAQET--RCICSSSSPSGAAS-SRPITFGYDAASYAKNFDDGRR 123
Query: 103 DVYSYGGFS 111
Y +
Sbjct: 124 PAAHYAALA 132
>UniRef100_Q6H891 Hypothetical protein OJ1572_F02.1 [Oryza sativa]
Length = 103
Score = 35.0 bits (79), Expect = 0.36
Identities = 17/29 (58%), Positives = 19/29 (64%), Gaps = 4/29 (13%)
Query: 84 GSFHYDSLSYALNFDDG----EEDVYSYG 108
G F YD LSYALNFD+G E+D Y G
Sbjct: 52 GEFRYDPLSYALNFDEGAADDEDDDYEAG 80
>UniRef100_Q9FY58 Hypothetical protein T5K6_60 [Arabidopsis thaliana]
Length = 152
Score = 34.3 bits (77), Expect = 0.62
Identities = 25/68 (36%), Positives = 32/68 (46%), Gaps = 9/68 (13%)
Query: 62 KTFIRRFNRNNNRA-------GAASYDKKGSFHYDSLSYALNFDDG--EEDVYSYGGFST 112
K+ I+ RNNN A S G F YD LSYALNF+D +D S+ F+
Sbjct: 73 KSRIKVTCRNNNCAYNNCVHHHHHSQSYPGDFSYDPLSYALNFEDNVRADDDGSFPNFTA 132
Query: 113 RFASVPAS 120
R P +
Sbjct: 133 RLPQSPVT 140
>UniRef100_Q8LM16 Hypothetical protein OSJNAb0008A05.22 [Oryza sativa]
Length = 184
Score = 33.5 bits (75), Expect = 1.1
Identities = 20/66 (30%), Positives = 28/66 (42%), Gaps = 17/66 (25%)
Query: 82 KKGSFHYDSLSYALNFDDGEE--------------DVYSYGGFSTRFASVPASTKPTFTP 127
+ G F YD+ SYA NFD+G + D Y F++R +P S P +P
Sbjct: 102 RSGDFQYDARSYARNFDEGTDGEASGDEQAGLAAGDTLKYRSFASR---LPPSPTPALSP 158
Query: 128 VMVRRC 133
C
Sbjct: 159 SAAPVC 164
>UniRef100_UPI000046B0F8 UPI000046B0F8 UniRef100 entry
Length = 288
Score = 32.7 bits (73), Expect = 1.8
Identities = 24/70 (34%), Positives = 34/70 (48%), Gaps = 4/70 (5%)
Query: 54 EIIVGPKWKTFIRRFNRNNNR-AGAASYDKKGSFHYDSLSYALNFDDGEEDVYSYGGFST 112
+I V +KT++ F+ N N+ G + K + SLS AL DGE D+ G T
Sbjct: 195 QICVANTFKTYVELFHHNTNKFEGVKALCK---YFNISLSDALVIGDGENDIEMLQGIET 251
Query: 113 RFASVPASTK 122
A AS+K
Sbjct: 252 SIAVHNASSK 261
>UniRef100_UPI00002C2001 UPI00002C2001 UniRef100 entry
Length = 275
Score = 32.7 bits (73), Expect = 1.8
Identities = 17/68 (25%), Positives = 35/68 (51%), Gaps = 2/68 (2%)
Query: 52 WSEIIVGPKWKTFIRRFNRNNNRAGAASYDKKGSFHYDSLSYALNFDDGEEDVYSYGGFS 111
+ ++ V P T+ + +N A + +KG++ L+Y ++FD + + GF
Sbjct: 68 YEDVFVSPSISTYYESL-KTDNSASSILKKQKGTYFDADLNYVIDFDKRNQKFQTTDGFR 126
Query: 112 TRFA-SVP 118
+RF+ S+P
Sbjct: 127 SRFSQSIP 134
>UniRef100_UPI0000468D60 UPI0000468D60 UniRef100 entry
Length = 285
Score = 32.3 bits (72), Expect = 2.3
Identities = 24/70 (34%), Positives = 35/70 (49%), Gaps = 6/70 (8%)
Query: 55 IIVGPKWKTFIRRFNRNNNR-AGAASYDKKGSFHYD-SLSYALNFDDGEEDVYSYGGFST 112
I +G K+KT++ F+ N N+ G + K H+D SL+ AL D D+ G T
Sbjct: 193 ISIGKKFKTYVELFHHNANKFEGVKALCK----HFDISLNDALAIGDAVNDIEMLQGVGT 248
Query: 113 RFASVPASTK 122
A AS+K
Sbjct: 249 SIAVQNASSK 258
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.319 0.133 0.452
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 255,329,107
Number of Sequences: 2790947
Number of extensions: 10631514
Number of successful extensions: 22736
Number of sequences better than 10.0: 63
Number of HSP's better than 10.0 without gapping: 15
Number of HSP's successfully gapped in prelim test: 48
Number of HSP's that attempted gapping in prelim test: 22698
Number of HSP's gapped (non-prelim): 64
length of query: 137
length of database: 848,049,833
effective HSP length: 113
effective length of query: 24
effective length of database: 532,672,822
effective search space: 12784147728
effective search space used: 12784147728
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 67 (30.4 bits)
Lotus: description of TM0201.10


