

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0197.8
(242 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
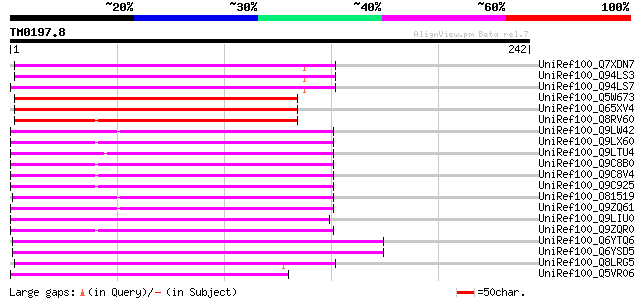
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q7XDN7 Hypothetical protein [Oryza sativa] 107 3e-22
UniRef100_Q94LS3 Putative helicase [Oryza sativa] 107 3e-22
UniRef100_Q94LS7 Putative helicase [Oryza sativa] 106 5e-22
UniRef100_Q5W673 Putative helicase [Oryza sativa] 105 1e-21
UniRef100_Q65XV4 Hypothetical protein P0016H04.14 [Oryza sativa] 104 2e-21
UniRef100_Q8RV60 Hypothetical protein At2g05640 [Arabidopsis tha... 104 2e-21
UniRef100_Q9LW42 Helicase-like protein [Arabidopsis thaliana] 102 7e-21
UniRef100_Q9LX60 Hypothetical protein F4M19_60 [Arabidopsis thal... 102 1e-20
UniRef100_Q9LTU4 Helicase-like protein [Arabidopsis thaliana] 101 2e-20
UniRef100_Q9C8B0 Hypothetical protein F10O5.11 [Arabidopsis thal... 100 3e-20
UniRef100_Q9C8V4 Hypothetical protein T22A15.15 [Arabidopsis tha... 100 3e-20
UniRef100_Q9C925 Hypothetical protein F14G24.23 [Arabidopsis tha... 100 6e-20
UniRef100_O81519 T24M8.10 protein [Arabidopsis thaliana] 98 2e-19
UniRef100_Q9ZQ61 Putative helicase [Arabidopsis thaliana] 97 3e-19
UniRef100_Q9LIU0 Similar to Arabidopsis thaliana chromosome II B... 96 8e-19
UniRef100_Q9ZQR0 Putative helicase [Arabidopsis thaliana] 93 5e-18
UniRef100_Q6YTQ6 Helicase-like protein [Oryza sativa] 92 2e-17
UniRef100_Q6YSD5 Helicase-like protein [Oryza sativa] 91 2e-17
UniRef100_Q8LRG5 Putative helicase [Oryza sativa] 91 2e-17
UniRef100_Q5VR06 Helicase-like protein [Oryza sativa] 91 3e-17
>UniRef100_Q7XDN7 Hypothetical protein [Oryza sativa]
Length = 1477
Score = 107 bits (267), Expect = 3e-22
Identities = 58/152 (38%), Positives = 91/152 (59%), Gaps = 2/152 (1%)
Query: 3 FFHDRVILTPTLKAVEMINTYMLGMIAAPEHEYLSSNSALRSYEHSEI*GEWFTT*FLNN 62
+ +R IL P + V +N Y++ I + YLS +S +S +S + T FLN+
Sbjct: 1261 YLEERAILCPVNENVNELNEYIMDQIQGDKVTYLSRDSVSKSVSYSHEMEMLYPTEFLNS 1320
Query: 63 TKS*GMSNHKLLLKVDIPIMLLRNIDQAAGLCNGTRLIVVELGERYIGATVITGTNANDK 122
G+ NH+L LKV +P+MLLRNI+Q+AGLCNGTR+ + LG + I A +ITGT++ D
Sbjct: 1321 LNHSGIPNHQLKLKVGLPVMLLRNINQSAGLCNGTRMTITRLGNKVIEAQIITGTHSGDM 1380
Query: 123 VHISIMDLVPSDPN--FQLNSEEDSFQLQYAL 152
V I + + P++P F LN ++ + +A+
Sbjct: 1381 VCIPQIIMSPTEPKWPFMLNRKQFPLSVCFAM 1412
>UniRef100_Q94LS3 Putative helicase [Oryza sativa]
Length = 1501
Score = 107 bits (267), Expect = 3e-22
Identities = 58/152 (38%), Positives = 91/152 (59%), Gaps = 2/152 (1%)
Query: 3 FFHDRVILTPTLKAVEMINTYMLGMIAAPEHEYLSSNSALRSYEHSEI*GEWFTT*FLNN 62
+ +R IL P + V +N Y++ I + YLS +S +S +S + T FLN+
Sbjct: 1285 YLEERAILCPVNENVNELNEYIMDQIQGDKVTYLSRDSVSKSVSYSHEMEMLYPTEFLNS 1344
Query: 63 TKS*GMSNHKLLLKVDIPIMLLRNIDQAAGLCNGTRLIVVELGERYIGATVITGTNANDK 122
G+ NH+L LKV +P+MLLRNI+Q+AGLCNGTR+ + LG + I A +ITGT++ D
Sbjct: 1345 LNHSGIPNHQLKLKVGLPVMLLRNINQSAGLCNGTRMTITRLGNKVIEAQIITGTHSGDM 1404
Query: 123 VHISIMDLVPSDPN--FQLNSEEDSFQLQYAL 152
V I + + P++P F LN ++ + +A+
Sbjct: 1405 VCIPQIIMSPTEPKWPFMLNRKQFPLSVCFAM 1436
>UniRef100_Q94LS7 Putative helicase [Oryza sativa]
Length = 1573
Score = 106 bits (265), Expect = 5e-22
Identities = 60/154 (38%), Positives = 91/154 (58%), Gaps = 2/154 (1%)
Query: 1 PQFFHDRVILTPTLKAVEMINTYMLGMIAAPEHEYLSSNSALRSYEHSEI*GEWFTT*FL 60
P++ +R IL PT V +N Y++ I + YLS +S +S +S + T FL
Sbjct: 1355 PKYLEERAILCPTNDDVNELNEYIMDQIQGDKVTYLSHDSVSKSMSYSHEMEMLYPTEFL 1414
Query: 61 NNTKS*GMSNHKLLLKVDIPIMLLRNIDQAAGLCNGTRLIVVELGERYIGATVITGTNAN 120
N+ K G+ NH+L LKV +P+MLLRNI+Q AGLCNGTR+ + G+R I A +ITGT+
Sbjct: 1415 NSLKHPGIPNHQLKLKVGLPVMLLRNINQNAGLCNGTRMRITRFGKRVIEAEIITGTHIG 1474
Query: 121 DKVHISIMDLVPSDPN--FQLNSEEDSFQLQYAL 152
D V I + + P++ F LN ++ + +A+
Sbjct: 1475 DMVCIPQIIMSPNERKWPFVLNRKQFPLSVCFAM 1508
>UniRef100_Q5W673 Putative helicase [Oryza sativa]
Length = 1634
Score = 105 bits (262), Expect = 1e-21
Identities = 53/132 (40%), Positives = 82/132 (61%)
Query: 3 FFHDRVILTPTLKAVEMINTYMLGMIAAPEHEYLSSNSALRSYEHSEI*GEWFTT*FLNN 62
F R IL P + IN +++ MI E YLS ++ ++ + + T FLN+
Sbjct: 1419 FLEQRAILCPRNETAREINEFIMNMIEGEEITYLSCDTVCKATTNDSETDVLYPTEFLNS 1478
Query: 63 TKS*GMSNHKLLLKVDIPIMLLRNIDQAAGLCNGTRLIVVELGERYIGATVITGTNANDK 122
GM NH L LKV +P+MLLRNI+Q++GLCNGTR+ + +LG+R+I A +ITGT+ +K
Sbjct: 1479 LNFPGMPNHVLKLKVGLPVMLLRNINQSSGLCNGTRMTITQLGKRFIEAQIITGTHVGEK 1538
Query: 123 VHISIMDLVPSD 134
V+I + + P++
Sbjct: 1539 VYIPRIIMTPTE 1550
>UniRef100_Q65XV4 Hypothetical protein P0016H04.14 [Oryza sativa]
Length = 1525
Score = 104 bits (260), Expect = 2e-21
Identities = 52/132 (39%), Positives = 82/132 (61%)
Query: 3 FFHDRVILTPTLKAVEMINTYMLGMIAAPEHEYLSSNSALRSYEHSEI*GEWFTT*FLNN 62
F R IL P + IN +++ MI E YLS ++ ++ + + T FLN+
Sbjct: 1310 FLEQRAILCPRNETARKINEFIMNMIEGEEITYLSCDTVCKATTNDSETDVLYPTEFLNS 1369
Query: 63 TKS*GMSNHKLLLKVDIPIMLLRNIDQAAGLCNGTRLIVVELGERYIGATVITGTNANDK 122
GM NH L LK+ +P+MLLRNI+Q++GLCNGTR+ + +LG+R+I A +ITGT+ +K
Sbjct: 1370 LNFPGMPNHVLKLKLGLPVMLLRNINQSSGLCNGTRMTITQLGKRFIEAQIITGTHVGEK 1429
Query: 123 VHISIMDLVPSD 134
V+I + + P++
Sbjct: 1430 VYIPRIIMTPTE 1441
>UniRef100_Q8RV60 Hypothetical protein At2g05640 [Arabidopsis thaliana]
Length = 1308
Score = 104 bits (259), Expect = 2e-21
Identities = 56/132 (42%), Positives = 83/132 (62%), Gaps = 1/132 (0%)
Query: 3 FFHDRVILTPTLKAVEMINTYMLGMIAAPEHEYLSSNSALRSYEHSEI*GEWFTT*FLNN 62
FF +R IL PT V +N +++ ++ EY SS+ + ++ S +T FLN
Sbjct: 1084 FFRERAILCPTNDDVSEVNNHIMDLLPGEVKEYFSSDK-ICDFDTSVERDANMSTEFLNA 1142
Query: 63 TKS*GMSNHKLLLKVDIPIMLLRNIDQAAGLCNGTRLIVVELGERYIGATVITGTNANDK 122
K G+ NH L LK+ +P+ML+RN+DQ GLCNGTRL V +LG+R I A V+TG+NA +K
Sbjct: 1143 IKCSGVPNHVLRLKLGVPVMLIRNLDQKYGLCNGTRLQVTQLGDRVIEAKVLTGSNAGNK 1202
Query: 123 VHISIMDLVPSD 134
V++ + L P+D
Sbjct: 1203 VYLPRLVLTPAD 1214
>UniRef100_Q9LW42 Helicase-like protein [Arabidopsis thaliana]
Length = 1669
Score = 102 bits (255), Expect = 7e-21
Identities = 62/151 (41%), Positives = 79/151 (52%), Gaps = 1/151 (0%)
Query: 1 PQFFHDRVILTPTLKAVEMINTYMLGMIAAPEHEYLSSNSALRSYEHSEI*GEWFTT*FL 60
P+FF DR IL PT V IN +ML + E Y SS+S S ++ +T FL
Sbjct: 1450 PKFFQDRAILCPTNDDVNSINDHMLSKLTGEEKIYRSSDSIDPSDTRADK-NPVYTPDFL 1508
Query: 61 NNTKS*GMSNHKLLLKVDIPIMLLRNIDQAAGLCNGTRLIVVELGERYIGATVITGTNAN 120
N K G+ NH L LKV P+MLLRN+D GL NGTRL +V LG++ + ++TGT
Sbjct: 1509 NKIKISGLPNHLLWLKVGCPVMLLRNLDSHGGLMNGTRLQIVRLGDKLVQGRILTGTRVG 1568
Query: 121 DKVHISIMDLVPSDPNFQLNSEEDSFQLQYA 151
V I M L PSD + F L A
Sbjct: 1569 KLVIIPRMPLTPSDRRLPFKMKRRQFPLSVA 1599
>UniRef100_Q9LX60 Hypothetical protein F4M19_60 [Arabidopsis thaliana]
Length = 1752
Score = 102 bits (253), Expect = 1e-20
Identities = 60/151 (39%), Positives = 82/151 (53%), Gaps = 1/151 (0%)
Query: 1 PQFFHDRVILTPTLKAVEMINTYMLGMIAAPEHEYLSSNSALRSYEHSEI*GEWFTT*FL 60
P+FF R IL PT + V IN YML + + E YLS++S + + + T FL
Sbjct: 1532 PKFFQRRAILAPTNEDVNTINQYMLEHLKSEERIYLSADS-IDPTDSDSLANPVITPDFL 1590
Query: 61 NNTKS*GMSNHKLLLKVDIPIMLLRNIDQAAGLCNGTRLIVVELGERYIGATVITGTNAN 120
N+ + GM +H L LKV P+MLLRN+D GLCNGTRL + +L ++ + A VIT
Sbjct: 1591 NSIQLTGMPHHALRLKVGAPVMLLRNLDPKGGLCNGTRLQITQLAKQVVQAKVITRDRIG 1650
Query: 121 DKVHISIMDLVPSDPNFQLNSEEDSFQLQYA 151
D V I +++L PSD F L A
Sbjct: 1651 DIVLIPLINLTPSDTKLPFKMRRRQFPLSVA 1681
>UniRef100_Q9LTU4 Helicase-like protein [Arabidopsis thaliana]
Length = 1428
Score = 101 bits (251), Expect = 2e-20
Identities = 60/151 (39%), Positives = 80/151 (52%), Gaps = 1/151 (0%)
Query: 1 PQFFHDRVILTPTLKAVEMINTYMLGMIAAPEHEYLSSNSALRSYEHSEI*GEWFTT*FL 60
P+FF R IL P + V IN YML + + E YLS++S S + + T FL
Sbjct: 1209 PKFFQRRAILAPKNEDVNTINQYMLEHLDSEERIYLSADSIDPS-DSDSLKNPVITPDFL 1267
Query: 61 NNTKS*GMSNHKLLLKVDIPIMLLRNIDQAAGLCNGTRLIVVELGERYIGATVITGTNAN 120
N+ K GM +H L LKV P+MLLRN+D GLCNGTRL + +L + A VITG
Sbjct: 1268 NSIKVSGMPHHSLRLKVGAPVMLLRNLDPKGGLCNGTRLQITQLCSHIVEAKVITGDRIG 1327
Query: 121 DKVHISIMDLVPSDPNFQLNSEEDSFQLQYA 151
V+I ++++ PSD F L A
Sbjct: 1328 QIVYIPLINITPSDTKLPFKMRRRQFPLSVA 1358
>UniRef100_Q9C8B0 Hypothetical protein F10O5.11 [Arabidopsis thaliana]
Length = 1678
Score = 100 bits (249), Expect = 3e-20
Identities = 59/151 (39%), Positives = 80/151 (52%), Gaps = 1/151 (0%)
Query: 1 PQFFHDRVILTPTLKAVEMINTYMLGMIAAPEHEYLSSNSALRSYEHSEI*GEWFTT*FL 60
P+FF R IL P + V IN Y+L + A E YLS++S + + + T FL
Sbjct: 1458 PKFFQGRAILAPKNEDVNTINEYLLEQLDAEERIYLSADS-IDPTDSDSLNNPVITPDFL 1516
Query: 61 NNTKS*GMSNHKLLLKVDIPIMLLRNIDQAAGLCNGTRLIVVELGERYIGATVITGTNAN 120
N+ K G+ NH L LKV P+MLLRN+D GLCNGTRL + +L + + A VITG
Sbjct: 1517 NSIKLPGLPNHSLCLKVGAPVMLLRNLDPKGGLCNGTRLQITQLCTQIVEAKVITGDRIG 1576
Query: 121 DKVHISIMDLVPSDPNFQLNSEEDSFQLQYA 151
+ V I ++L P+D F L A
Sbjct: 1577 NIVLIPTVNLTPTDTKLPFKMRRRQFPLSVA 1607
>UniRef100_Q9C8V4 Hypothetical protein T22A15.15 [Arabidopsis thaliana]
Length = 729
Score = 100 bits (249), Expect = 3e-20
Identities = 59/151 (39%), Positives = 80/151 (52%), Gaps = 1/151 (0%)
Query: 1 PQFFHDRVILTPTLKAVEMINTYMLGMIAAPEHEYLSSNSALRSYEHSEI*GEWFTT*FL 60
P+FF R IL P + V IN Y+L + A E YLS++S + + + T FL
Sbjct: 509 PKFFQGRAILAPKNEDVNTINEYLLEQLDAEERIYLSADS-IDPTDSDSLNNPVITPDFL 567
Query: 61 NNTKS*GMSNHKLLLKVDIPIMLLRNIDQAAGLCNGTRLIVVELGERYIGATVITGTNAN 120
N+ K G+ NH L LKV P+MLLRN+D GLCNGTRL + +L + + A VITG
Sbjct: 568 NSIKLPGLPNHSLCLKVGAPVMLLRNLDPKGGLCNGTRLQITQLCTQIVEAKVITGDRIG 627
Query: 121 DKVHISIMDLVPSDPNFQLNSEEDSFQLQYA 151
+ V I ++L P+D F L A
Sbjct: 628 NIVLIPTVNLTPTDTKLPFKMRRRQFPLSVA 658
>UniRef100_Q9C925 Hypothetical protein F14G24.23 [Arabidopsis thaliana]
Length = 996
Score = 99.8 bits (247), Expect = 6e-20
Identities = 60/151 (39%), Positives = 80/151 (52%), Gaps = 1/151 (0%)
Query: 1 PQFFHDRVILTPTLKAVEMINTYMLGMIAAPEHEYLSSNSALRSYEHSEI*GEWFTT*FL 60
P+FF R IL PT + V IN +M+ M+ E YLSS+S + + S + ++ FL
Sbjct: 801 PKFFQGRAILCPTNEDVNSINEHMMSMLDGEERIYLSSDS-IDPADTSSANNDAYSADFL 859
Query: 61 NNTKS*GMSNHKLLLKVDIPIMLLRNIDQAAGLCNGTRLIVVELGERYIGATVITGTNAN 120
N+ + G+ NH L LKV P+MLLRN+D GLCNGTRL V ++ + I A ITG
Sbjct: 860 NSVRVSGLPNHCLRLKVGCPVMLLRNMDPNKGLCNGTRLQVTQMADTVIQARFITGNRVG 919
Query: 121 DKVHISIMDLVPSDPNFQLNSEEDSFQLQYA 151
V I M + PSD F L A
Sbjct: 920 KIVLIPRMLITPSDTRLPFKMRRRQFPLSVA 950
>UniRef100_O81519 T24M8.10 protein [Arabidopsis thaliana]
Length = 258
Score = 97.8 bits (242), Expect = 2e-19
Identities = 60/150 (40%), Positives = 81/150 (54%), Gaps = 1/150 (0%)
Query: 2 QFFHDRVILTPTLKAVEMINTYMLGMIAAPEHEYLSSNSALRSYEHSEI*GEWFTT*FLN 61
+F +R IL+PT + V IN +L + E +YLS +S S SE FT FLN
Sbjct: 61 KFIQERAILSPTNEDVNTINQRLLEKLPGEEIQYLSIDSIDLSDTTSEY-NPVFTPDFLN 119
Query: 62 NTKS*GMSNHKLLLKVDIPIMLLRNIDQAAGLCNGTRLIVVELGERYIGATVITGTNAND 121
+ K G+ NH L L + PIMLLRN+D GLCNGTRL ++++ + A +ITG D
Sbjct: 120 SIKISGLPNHCLRLNIGAPIMLLRNLDPKGGLCNGTRLQMIQMTPPILQAVIITGGRIGD 179
Query: 122 KVHISIMDLVPSDPNFQLNSEEDSFQLQYA 151
KV IS + + PSD N F + A
Sbjct: 180 KVLISKILITPSDTKLPFNMRRKQFPIVVA 209
>UniRef100_Q9ZQ61 Putative helicase [Arabidopsis thaliana]
Length = 1230
Score = 97.4 bits (241), Expect = 3e-19
Identities = 60/151 (39%), Positives = 77/151 (50%), Gaps = 1/151 (0%)
Query: 1 PQFFHDRVILTPTLKAVEMINTYMLGMIAAPEHEYLSSNSALRSYEHSEI*GEWFTT*FL 60
P+FF + IL PT V IN +ML + E Y SSNS S ++ +T FL
Sbjct: 1053 PKFFQHKAILCPTNDDVNSINDHMLSKLTGEERIYRSSNSIDPSDTRADK-NPIYTPDFL 1111
Query: 61 NNTKS*GMSNHKLLLKVDIPIMLLRNIDQAAGLCNGTRLIVVELGERYIGATVITGTNAN 120
N K G++NH L LKV P+MLLRN GL NGTRL +V LG++ + ++TGT
Sbjct: 1112 NKIKISGLANHLLRLKVGCPVMLLRNFYPHGGLMNGTRLQIVRLGDKLVQGRILTGTRVG 1171
Query: 121 DKVHISIMDLVPSDPNFQLNSEEDSFQLQYA 151
V I M L PSD + F L A
Sbjct: 1172 KLVIIPRMSLTPSDRRLPFKMKRRHFPLSVA 1202
>UniRef100_Q9LIU0 Similar to Arabidopsis thaliana chromosome II BAC T13P21 genomic
sequence [Oryza sativa]
Length = 1278
Score = 95.9 bits (237), Expect = 8e-19
Identities = 51/149 (34%), Positives = 80/149 (53%)
Query: 1 PQFFHDRVILTPTLKAVEMINTYMLGMIAAPEHEYLSSNSALRSYEHSEI*GEWFTT*FL 60
P + I+ P V+ IN M+ M+ EYLS ++ +S EH + T FL
Sbjct: 1023 PSYLASCAIVCPNNSTVDDINDRMVDMVPGEVKEYLSCDTISKSSEHIPDFDLLYPTEFL 1082
Query: 61 NNTKS*GMSNHKLLLKVDIPIMLLRNIDQAAGLCNGTRLIVVELGERYIGATVITGTNAN 120
N+ + HKL LK + +MLLRN++Q+ GLCNGTRL+ + LG+R + ++TGTN
Sbjct: 1083 NSISANNFPAHKLALKKGVTVMLLRNLNQSMGLCNGTRLLALSLGQRLLECQILTGTNIG 1142
Query: 121 DKVHISIMDLVPSDPNFQLNSEEDSFQLQ 149
D+V I + L + P + + F ++
Sbjct: 1143 DRVFIPRIALTTTSPKWPFTLQRRQFPVR 1171
>UniRef100_Q9ZQR0 Putative helicase [Arabidopsis thaliana]
Length = 1265
Score = 93.2 bits (230), Expect = 5e-18
Identities = 55/151 (36%), Positives = 78/151 (51%), Gaps = 1/151 (0%)
Query: 1 PQFFHDRVILTPTLKAVEMINTYMLGMIAAPEHEYLSSNSALRSYEHSEI*GEWFTT*FL 60
P+FF R IL + V IN Y+L + A E YLS++S + + + T FL
Sbjct: 1062 PKFFQGRAILASKNEDVNTINEYLLDQLHAEERIYLSADS-IDPTDSDSLSNPVITPDFL 1120
Query: 61 NNTKS*GMSNHKLLLKVDIPIMLLRNIDQAAGLCNGTRLIVVELGERYIGATVITGTNAN 120
N+ K G+ NH L LKV P++LLRN+D GLCNGTRL + +L + + A VITG
Sbjct: 1121 NSIKLPGLPNHSLRLKVGAPVLLLRNLDPKGGLCNGTRLQITQLCTQIVEAKVITGDRIG 1180
Query: 121 DKVHISIMDLVPSDPNFQLNSEEDSFQLQYA 151
+ I ++L P++ F L A
Sbjct: 1181 HIILIPTVNLTPTNTKLPFKMRRRQFPLSVA 1211
>UniRef100_Q6YTQ6 Helicase-like protein [Oryza sativa]
Length = 1430
Score = 91.7 bits (226), Expect = 2e-17
Identities = 53/173 (30%), Positives = 88/173 (50%)
Query: 2 QFFHDRVILTPTLKAVEMINTYMLGMIAAPEHEYLSSNSALRSYEHSEI*GEWFTT*FLN 61
++ R I+ PT V+ IN Y++G++ YLS ++ +S E + FLN
Sbjct: 1172 EYLASRAIVCPTNTTVDEINDYIIGLVPGDSRVYLSCDTISKSSEQIPDFDLLYPPEFLN 1231
Query: 62 NTKS*GMSNHKLLLKVDIPIMLLRNIDQAAGLCNGTRLIVVELGERYIGATVITGTNAND 121
+ + HKL+LK + +MLLRN++Q+ GLCNGTRL+V LG+R + V+TG+N +
Sbjct: 1232 SINATNFPTHKLVLKEGVVVMLLRNLNQSIGLCNGTRLLVTVLGDRILQCIVLTGSNIGE 1291
Query: 122 KVHISIMDLVPSDPNFQLNSEEDSFQLQYALR*P*TKANDKHYLRWGSSFRDP 174
V+I + L + + + F ++ K+ + R G R P
Sbjct: 1292 TVYIPRITLGTTKMKWPFTLQRRQFPVRVCYSMTINKSQGQTLQRVGVYLRKP 1344
>UniRef100_Q6YSD5 Helicase-like protein [Oryza sativa]
Length = 1516
Score = 91.3 bits (225), Expect = 2e-17
Identities = 52/173 (30%), Positives = 88/173 (50%)
Query: 2 QFFHDRVILTPTLKAVEMINTYMLGMIAAPEHEYLSSNSALRSYEHSEI*GEWFTT*FLN 61
++ R I+ PT V+ IN Y++G++ YLS ++ +S E + FLN
Sbjct: 1258 EYLASRAIVCPTNTTVDEINDYIIGLVPGDSRVYLSCDTISKSSEQIPDFDLLYPPEFLN 1317
Query: 62 NTKS*GMSNHKLLLKVDIPIMLLRNIDQAAGLCNGTRLIVVELGERYIGATVITGTNAND 121
+ + HKL+LK + +MLLRN++Q+ GLCNGTRL+V LG+R + ++TG+N +
Sbjct: 1318 SINATNFPTHKLVLKEGVVVMLLRNLNQSIGLCNGTRLLVTVLGDRILQCIILTGSNIGE 1377
Query: 122 KVHISIMDLVPSDPNFQLNSEEDSFQLQYALR*P*TKANDKHYLRWGSSFRDP 174
V+I + L + + + F ++ K+ + R G R P
Sbjct: 1378 TVYIPRITLGTTKMKWPFTLQRRQFPVRVCYSMTINKSQGQTLQRVGVYLRKP 1430
>UniRef100_Q8LRG5 Putative helicase [Oryza sativa]
Length = 1453
Score = 91.3 bits (225), Expect = 2e-17
Identities = 51/152 (33%), Positives = 81/152 (52%), Gaps = 2/152 (1%)
Query: 3 FFHDRVILTPTLKAVEMINTYMLGMIAAPEHEYLSSNSALRSYEHSEI*GEWFTT*FLNN 62
+ R I+ P V+ IN YM+ MI EYLS ++ ++ EH + T FLN+
Sbjct: 1223 YLASRAIVCPNNSTVDEINDYMVAMIPGEMKEYLSCDTISKTSEHIPDFDILYPTEFLNS 1282
Query: 63 TKS*GMSNHKLLLKVDIPIMLLRNIDQAAGLCNGTRLIVVELGERYIGATVITGTNANDK 122
+ H+L LK +MLLRN++Q+ GLCNGTRL+V+ LG R + ++TG+N ++
Sbjct: 1283 INANNFPTHRLALKKGATVMLLRNLNQSLGLCNGTRLLVLSLGHRLLECVILTGSNVGER 1342
Query: 123 VHIS--IMDLVPSDPNFQLNSEEDSFQLQYAL 152
I ++ S F L + ++ YA+
Sbjct: 1343 AFIPRIVLSTTSSKWPFVLQRRQFPVRVCYAM 1374
>UniRef100_Q5VR06 Helicase-like protein [Oryza sativa]
Length = 1427
Score = 90.9 bits (224), Expect = 3e-17
Identities = 46/130 (35%), Positives = 76/130 (58%)
Query: 1 PQFFHDRVILTPTLKAVEMINTYMLGMIAAPEHEYLSSNSALRSYEHSEI*GEWFTT*FL 60
P + +R IL T + V+ +N Y+L ++ E EY S+++ + + + + +L
Sbjct: 1205 PNYLKNRAILATTNEIVDEVNNYILSLVPNQEKEYYSADTLSQCMDTTNDADILYPVEYL 1264
Query: 61 NNTKS*GMSNHKLLLKVDIPIMLLRNIDQAAGLCNGTRLIVVELGERYIGATVITGTNAN 120
N+ + H L LKV +PI+LLRN++Q GLCNGTRLI+ LG+ I +ITGT+
Sbjct: 1265 NSLNANNFPTHVLKLKVGVPIILLRNLNQNLGLCNGTRLIITNLGDNIIEGIIITGTHIG 1324
Query: 121 DKVHISIMDL 130
+K +I ++L
Sbjct: 1325 EKAYIPRINL 1334
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.341 0.150 0.478
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 363,423,057
Number of Sequences: 2790947
Number of extensions: 13486677
Number of successful extensions: 36847
Number of sequences better than 10.0: 89
Number of HSP's better than 10.0 without gapping: 84
Number of HSP's successfully gapped in prelim test: 5
Number of HSP's that attempted gapping in prelim test: 36704
Number of HSP's gapped (non-prelim): 103
length of query: 242
length of database: 848,049,833
effective HSP length: 124
effective length of query: 118
effective length of database: 501,972,405
effective search space: 59232743790
effective search space used: 59232743790
T: 11
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.9 bits)
S2: 73 (32.7 bits)
Lotus: description of TM0197.8


