

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0197.5
(317 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
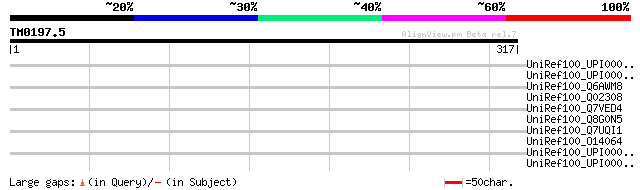
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_UPI000029C4AC UPI000029C4AC UniRef100 entry 35 2.0
UniRef100_UPI0000124881 UPI0000124881 UniRef100 entry 35 2.7
UniRef100_Q6AWM8 RE41617p [Drosophila melanogaster] 35 2.7
UniRef100_Q02308 Hairless protein [Drosophila melanogaster] 35 2.7
UniRef100_Q7VED4 ABC-type uncharacterized transport system perme... 35 3.5
UniRef100_Q8G0N5 Hypothetical protein [Brucella suis] 34 4.5
UniRef100_Q7UQI1 Probable response-regulator [Rhodopirellula bal... 34 5.9
UniRef100_O14064 Bir1 protein [Schizosaccharomyces pombe] 34 5.9
UniRef100_UPI00001CF8CF UPI00001CF8CF UniRef100 entry 33 7.7
UniRef100_UPI000023F610 UPI000023F610 UniRef100 entry 33 7.7
>UniRef100_UPI000029C4AC UPI000029C4AC UniRef100 entry
Length = 477
Score = 35.4 bits (80), Expect = 2.0
Identities = 42/221 (19%), Positives = 88/221 (39%), Gaps = 29/221 (13%)
Query: 65 EMEGLVVFSNGVSTARKP--ETVVGE-SSSERDQYRPESDWNRSSVSRKANIAREVSSPV 121
E EG V + A K + +VG ++++ D+ R + + ++ S++ + E S P
Sbjct: 140 EAEGEGVAAEEARRADKAIEQLIVGHLTAAQGDEVRERDEEDETTRSQERGWSTEESGPE 199
Query: 122 AVEQVAGSKPQAENANHEEKPLDLGRMSHLVCGESKGATEEAMSVTAEARHVVKNADVGL 181
+V + G E + K DLG + + G+ + +S+ + R +N D+GL
Sbjct: 200 SVSEAEGK----EKDQSQGKGTDLGALL-----KETGSKGDELSLQRKIRGYFQNIDLGL 250
Query: 182 MRKEVTVLMVG----------------WVHNWLVGSLRMGTRWVVQINNVWLR-SEQVCG 224
E+ + G W NW+ G ++ E+V
Sbjct: 251 QDNEILPPLKGYKAYNTQLARAGKKPHWQENWMGKQPAKGGNFMDDFEGEGEELEEEVED 310
Query: 225 DTQKAAKEDSWGASDASLQQVMVGEHEVKDFSPDQARVSQV 265
+ Q + + + A Q+V+ + E + ++ R++ +
Sbjct: 311 EEQSLTRMEEEARAQAEKQEVLRQQEEAERAREEEQRLADI 351
>UniRef100_UPI0000124881 UPI0000124881 UniRef100 entry
Length = 1077
Score = 35.0 bits (79), Expect = 2.7
Identities = 23/97 (23%), Positives = 49/97 (49%), Gaps = 3/97 (3%)
Query: 64 LEMEGLVVFSNGVSTAR--KPETVVGESSSERDQYRPESDWNRSSVSRKANIAREVSSPV 121
+E+E + V + + KP+T+ GE +ER + P+ + S S++A+ ++V P
Sbjct: 391 VEIENVAVADTTTNEIKIEKPDTIKGEDDAERLEKEPKKAVSDDSESKEASPGQQV-EPQ 449
Query: 122 AVEQVAGSKPQAENANHEEKPLDLGRMSHLVCGESKG 158
++ + + + EE +L R+++ V G+ G
Sbjct: 450 PKDETVDVEMKMNTSEDEEPMTELPRITNAVNGDLNG 486
>UniRef100_Q6AWM8 RE41617p [Drosophila melanogaster]
Length = 1077
Score = 35.0 bits (79), Expect = 2.7
Identities = 23/97 (23%), Positives = 49/97 (49%), Gaps = 3/97 (3%)
Query: 64 LEMEGLVVFSNGVSTAR--KPETVVGESSSERDQYRPESDWNRSSVSRKANIAREVSSPV 121
+E+E + V + + KP+T+ GE +ER + P+ + S S++A+ ++V P
Sbjct: 391 VEIENVAVADTTTNEIKIEKPDTIKGEDDAERLEKEPKKAVSDDSESKEASPGQQV-EPQ 449
Query: 122 AVEQVAGSKPQAENANHEEKPLDLGRMSHLVCGESKG 158
++ + + + EE +L R+++ V G+ G
Sbjct: 450 PKDETVDVEMKMNTSEDEEPMTELPRITNAVNGDLNG 486
>UniRef100_Q02308 Hairless protein [Drosophila melanogaster]
Length = 1077
Score = 35.0 bits (79), Expect = 2.7
Identities = 23/97 (23%), Positives = 49/97 (49%), Gaps = 3/97 (3%)
Query: 64 LEMEGLVVFSNGVSTAR--KPETVVGESSSERDQYRPESDWNRSSVSRKANIAREVSSPV 121
+E+E + V + + KP+T+ GE +ER + P+ + S S++A+ ++V P
Sbjct: 391 VEIENVAVADTTTNEIKIEKPDTIKGEDDAERLEKEPKKAVSDDSESKEASPGQQV-EPQ 449
Query: 122 AVEQVAGSKPQAENANHEEKPLDLGRMSHLVCGESKG 158
++ + + + EE +L R+++ V G+ G
Sbjct: 450 PKDETVDVEMKMNTSEDEEPMTELPRITNAVNGDLNG 486
>UniRef100_Q7VED4 ABC-type uncharacterized transport system permease and ATPase
component [Prochlorococcus marinus]
Length = 662
Score = 34.7 bits (78), Expect = 3.5
Identities = 20/72 (27%), Positives = 39/72 (53%), Gaps = 2/72 (2%)
Query: 43 RFMLAELTIEAMIDGWLFIVLLEMEGLVVFSNGVSTARKPETVVGESSSERDQYRPESD- 101
RF+ A M++G LF ++ ++E L F+ G+S ++ V + S ++D + D
Sbjct: 385 RFIQASFAF-GMVEGSLFFIVNQIEELAKFTAGISRLEGFQSKVEKVSRQKDSSQENIDS 443
Query: 102 WNRSSVSRKANI 113
WN S + + A++
Sbjct: 444 WNNSIIIKNADL 455
>UniRef100_Q8G0N5 Hypothetical protein [Brucella suis]
Length = 247
Score = 34.3 bits (77), Expect = 4.5
Identities = 23/99 (23%), Positives = 44/99 (44%), Gaps = 8/99 (8%)
Query: 86 VGESSSERDQYRPESDWNRSSVSRKANIAREVSSPVAVEQVAG----SKPQAENANHEEK 141
VGE S D P+ + + SV+ K + ++PV++ ++ + P + +
Sbjct: 126 VGEDESRVDFDLPDQSYTKESVTEKLMELVQANAPVSIHWISDEEFLANPDIVKSKNVRP 185
Query: 142 PLDLGRMSHLVCGESKGATEEAMSVTAEARHVVKNADVG 180
P+ LGR+ + GE+ + T HV + +VG
Sbjct: 186 PVGLGRIRLVAIGENGSVDSQPCGGT----HVSETQEVG 220
>UniRef100_Q7UQI1 Probable response-regulator [Rhodopirellula baltica]
Length = 257
Score = 33.9 bits (76), Expect = 5.9
Identities = 50/172 (29%), Positives = 74/172 (42%), Gaps = 19/172 (11%)
Query: 23 RVKRLGAIKVLLEFEIGEDMRFMLAELTIEAMIDGWLFIVL--LEMEGLV-VFSNGVSTA 79
R++R G + V +E IG+ + + L+ EA L +V L +E +V F NG T
Sbjct: 56 RLERPGCLVVDVEL-IGDGFQRVQKALS-EASCSAPLILVAGELPIETVVHAFENGAWTV 113
Query: 80 --RKPETVVGESSSERDQYRPESDWNRSSVS-----RKAN------IAREVSSPVAVEQV 126
+ + S S RD R DW+R V RK N R+ V +
Sbjct: 114 VLKSADNAQKFSGSLRDHIRQAIDWDRFQVGMEKAHRKRNRILDGLTERQRKVLNCVMEG 173
Query: 127 AGSKPQAENANHEEKPLDLGRMSHLVCGESKGATEEAMSVTAEARHVVKNAD 178
+K A N N ++ ++ R SHL+ + G T E +V E R V K D
Sbjct: 174 MPTKAIAANYNVSKRLIEFER-SHLLSAFNVGGTAELTAVVGEHRIVEKLLD 224
>UniRef100_O14064 Bir1 protein [Schizosaccharomyces pombe]
Length = 997
Score = 33.9 bits (76), Expect = 5.9
Identities = 25/67 (37%), Positives = 32/67 (47%), Gaps = 9/67 (13%)
Query: 76 VSTARKPETVVGESSSE--RDQYRPESDWNRSSVSRKANIAREVSSPVAVEQVAGSKPQA 133
V KP+T + E E R ES R S+ R R+VSSPV+ E ++
Sbjct: 642 VDFIEKPKTEISEVLPEEKRKAICDESQTVRVSIDRGVTKTRDVSSPVSDE-------KS 694
Query: 134 ENANHEE 140
EN NHEE
Sbjct: 695 ENVNHEE 701
>UniRef100_UPI00001CF8CF UPI00001CF8CF UniRef100 entry
Length = 346
Score = 33.5 bits (75), Expect = 7.7
Identities = 37/135 (27%), Positives = 64/135 (47%), Gaps = 12/135 (8%)
Query: 123 VEQVAGSKPQAEN---ANHEEKPLDLGRMSHLVCGESKGATEEAMSVTAEARHVVKNADV 179
+EQ A K ++E+ ++ E +LG V +S G TE+++SVT ++ V +++ V
Sbjct: 107 LEQKAKKKRKSEDEGRSSREPGAEELGDERQCVTEDSVGVTEDSVSVTEDSVGVTEDS-V 165
Query: 180 GLMRKEVTVLMVGWVHNWLVGSLRMGTRWVVQINNVWLRSEQVCGDTQKAAKEDSWGASD 239
G+ VT VG + + T V + +E G T+ + EDS G ++
Sbjct: 166 GVTEDSVTEDSVGVTEDSVTEDSVGVTEDSVSV------TEDSVGVTEDSVTEDSVGITE 219
Query: 240 AS--LQQVMVGEHEV 252
S + + V EH V
Sbjct: 220 DSVGVTEDSVTEHSV 234
>UniRef100_UPI000023F610 UPI000023F610 UniRef100 entry
Length = 545
Score = 33.5 bits (75), Expect = 7.7
Identities = 22/50 (44%), Positives = 26/50 (52%), Gaps = 1/50 (2%)
Query: 229 AAKEDSWGASDASLQQVMVGEHEVKDFSPDQARVSQVVDARNTNIQDGFR 278
AA+ DS G S A + GE D QAR+SQV +A T QDG R
Sbjct: 114 AAQADS-GFSLAPTPALRTGEERPPDIGSLQARLSQVAEAERTTWQDGAR 162
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.316 0.131 0.375
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 479,744,850
Number of Sequences: 2790947
Number of extensions: 18014621
Number of successful extensions: 44582
Number of sequences better than 10.0: 10
Number of HSP's better than 10.0 without gapping: 0
Number of HSP's successfully gapped in prelim test: 10
Number of HSP's that attempted gapping in prelim test: 44580
Number of HSP's gapped (non-prelim): 10
length of query: 317
length of database: 848,049,833
effective HSP length: 127
effective length of query: 190
effective length of database: 493,599,564
effective search space: 93783917160
effective search space used: 93783917160
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 75 (33.5 bits)
Lotus: description of TM0197.5


