

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0195a.11
(377 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
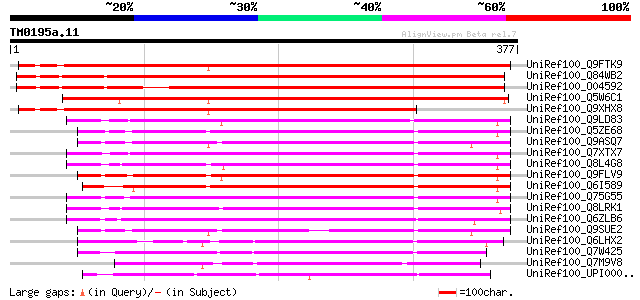
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9FTK9 P0034C11.13 protein [Oryza sativa] 428 e-118
UniRef100_Q84WB2 Hypothetical protein At1g62280 [Arabidopsis tha... 423 e-117
UniRef100_O04592 F19K23.20 protein [Arabidopsis thaliana] 395 e-108
UniRef100_Q5W6C1 Hypothetical protein OSJNBa0037H03.8 [Oryza sat... 389 e-107
UniRef100_Q9XHX8 OSJNBa0049B20.15 protein [Oryza sativa] 333 4e-90
UniRef100_Q9LD83 T12C24.3 [Arabidopsis thaliana] 256 9e-67
UniRef100_Q5ZE68 Hypothetical protein P0501G01.17 [Oryza sativa] 254 4e-66
UniRef100_Q9ASQ7 AT4g27970/T13J8_80 [Arabidopsis thaliana] 251 3e-65
UniRef100_Q7XTX7 OSJNBa0019K04.20 protein [Oryza sativa] 251 3e-65
UniRef100_Q8L4G8 C4-dicarboxylate transporter-like protein [Oryz... 250 5e-65
UniRef100_Q9FLV9 Similarity to unknown protein [Arabidopsis thal... 249 7e-65
UniRef100_Q6I589 Hypothetical protein OSJNBa0009C07.7 [Oryza sat... 247 3e-64
UniRef100_Q75G55 Hypothetical protein B1003C08.2 [Oryza sativa] 246 6e-64
UniRef100_Q8LRK1 P0443E07.21 protein [Oryza sativa] 245 1e-63
UniRef100_Q6ZLB6 Hypothetical protein OJ1014_E09.31 [Oryza sativa] 236 1e-60
UniRef100_Q9SUE2 Hypothetical protein T13J8.80 [Arabidopsis thal... 218 2e-55
UniRef100_Q6LHX2 Hypothetical protein WS0635 [Photobacterium pro... 114 6e-24
UniRef100_Q7W425 Putative inner membrane transport protein [Bord... 102 1e-20
UniRef100_Q7M9V8 SIMILARITY TO mviN PROTEIN [Wolinella succinoge... 98 3e-19
UniRef100_UPI00002B2C06 UPI00002B2C06 UniRef100 entry 91 5e-17
>UniRef100_Q9FTK9 P0034C11.13 protein [Oryza sativa]
Length = 374
Score = 428 bits (1100), Expect = e-118
Identities = 217/368 (58%), Positives = 266/368 (71%), Gaps = 10/368 (2%)
Query: 7 KPEIALVIDNTVSTIISREPPSFIVARRLLASLNSVLTKLHAGYFRISLSLGGQALLWKT 66
KP A ID V+ S +P + V R +LT+ HAGYFRISL+L GQALLW+T
Sbjct: 9 KPFAASSIDGGVA---STKPAAAPVVARF-----GMLTRFHAGYFRISLALSGQALLWRT 60
Query: 67 LIGPTHDKSTLRHVVHMVPSSAFLVLWSLALFTLALLSLLYLLRCLFHFNMVKAEFLHHV 126
L + D L VV +PS+AF++LWSLAL TL L LY RCL F V+AEF HHV
Sbjct: 61 LSDASTDPRALGPVVRSLPSAAFVLLWSLALLTLVALCALYAARCLLRFPAVRAEFRHHV 120
Query: 127 GVNYLFAPWISWFLLLQSAP--FVAPKTATYLVLWWVFAVPVVVLDVKIYGQWFTKGKRF 184
+NYLFAPWISW LLLQ+AP + P Y LWW F++P++ LDVK+YGQWFT+G++F
Sbjct: 121 AMNYLFAPWISWLLLLQAAPPLLLRPDARPYRALWWAFSLPILALDVKVYGQWFTRGRKF 180
Query: 185 LSTAANPTSQLSVIGNLVGAQAAAHMGWKESAVCLFSLGMVHYLVLFVTLYQRLSGGNRL 244
LS ANP S ++VIGNLV A+AAA MGW E AV +F++G HYLVLFVTLYQR G + L
Sbjct: 181 LSMVANPASHITVIGNLVTARAAARMGWHEGAVAMFAVGAAHYLVLFVTLYQRFLGSDSL 240
Query: 245 PVLLRPVFFLFFAAPGVASLAWGSIVGGFDTLSKMLFFLSLFLFMSLVCRPTLFRRSMKR 304
P +LRPVFFLFFAAP +ASLAW +I FDT KMLFFLSLFLF SLV RPTLF+R+M+R
Sbjct: 241 PAMLRPVFFLFFAAPSMASLAWDAISASFDTCCKMLFFLSLFLFASLVSRPTLFKRAMRR 300
Query: 305 FNVAWWAYSFPVTVLAMASTDYAEEVKGTISHVLMLVLLALSVLVSLALTVFTFINSKML 364
F+VAWWAYSFP+TVLA+A+ +YA+EV+ + VLML L LSV V+LAL VFT + + L
Sbjct: 301 FSVAWWAYSFPLTVLALAAAEYAQEVREVAASVLMLALAILSVAVTLALMVFTVLRTNDL 360
Query: 365 LPDDDPIA 372
LP DDP +
Sbjct: 361 LPHDDPFS 368
>UniRef100_Q84WB2 Hypothetical protein At1g62280 [Arabidopsis thaliana]
Length = 385
Score = 423 bits (1087), Expect = e-117
Identities = 212/363 (58%), Positives = 265/363 (72%), Gaps = 5/363 (1%)
Query: 6 PKPEIALVIDNTVSTIISREPPSFIVARRLLASLNSVLTKLHAGYFRISLSLGGQALLWK 65
P+ EI + IDN++ + S+E + + + + L S L LHAGYFRISLSL QALLWK
Sbjct: 4 PRQEIHIEIDNSIPS--SKEFKTGLADAKPVV-LMSALRSLHAGYFRISLSLCSQALLWK 60
Query: 66 TLIGPTHDKSTLRHVVHMVPSSAFLVLWSLALFTLALLSLLYLLRCLFHFNMVKAEFLHH 125
+I P + ++ H+ +PS AF +LW LAL T L LY L+C+F F+ VK EFLH+
Sbjct: 61 IMIAP--ESPSMSHMHSKLPSMAFHLLWYLALVTQVSLCFLYALKCIFFFDKVKEEFLHY 118
Query: 126 VGVNYLFAPWISWFLLLQSAPFVAPKTATYLVLWWVFAVPVVVLDVKIYGQWFTKGKRFL 185
+GVNYL+AP ISW L+LQSAP + P + Y L+W+FAVPV+ LD+K+YGQWFT KRFL
Sbjct: 119 IGVNYLYAPSISWLLMLQSAPMMEPNSVLYQTLFWIFAVPVLTLDIKLYGQWFTTEKRFL 178
Query: 186 STAANPTSQLSVIGNLVGAQAAAHMGWKESAVCLFSLGMVHYLVLFVTLYQRLSGGNRLP 245
S ANP SQ+SVI NLV A+ AA MGW E A+C+FS+GMVHYLV+FVTLYQRL GGN P
Sbjct: 179 SMLANPASQVSVIANLVAARGAAEMGWNECALCMFSIGMVHYLVIFVTLYQRLPGGNNFP 238
Query: 246 VLLRPVFFLFFAAPGVASLAWGSIVGGFDTLSKMLFFLSLFLFMSLVCRPTLFRRSMKRF 305
LRP+FFLF AAP +ASLAW SI G FD ++KMLFFLSLF+FMSLVCRP LF++SMKRF
Sbjct: 239 AKLRPIFFLFVAAPAMASLAWNSICGTFDAVAKMLFFLSLFIFMSLVCRPNLFKKSMKRF 298
Query: 306 NVAWWAYSFPVTVLAMASTDYAEEVKGTISHVLMLVLLALSVLVSLALTVFTFINSKMLL 365
NVAWWAYSFP+T LA+ S YA+EVK + LML+ ++SVL+ L + V T NS LL
Sbjct: 299 NVAWWAYSFPLTFLALDSVQYAQEVKDPVGSGLMLIFSSISVLIFLGMMVLTAANSNRLL 358
Query: 366 PDD 368
D
Sbjct: 359 RHD 361
>UniRef100_O04592 F19K23.20 protein [Arabidopsis thaliana]
Length = 366
Score = 395 bits (1014), Expect = e-108
Identities = 204/363 (56%), Positives = 254/363 (69%), Gaps = 24/363 (6%)
Query: 6 PKPEIALVIDNTVSTIISREPPSFIVARRLLASLNSVLTKLHAGYFRISLSLGGQALLWK 65
P+ EI + IDN++ + S+E + + + + L S L LHAGYFRISLSL QALLWK
Sbjct: 4 PRQEIHIEIDNSIPS--SKEFKTGLADAKPVV-LMSALRSLHAGYFRISLSLCSQALLWK 60
Query: 66 TLIGPTHDKSTLRHVVHMVPSSAFLVLWSLALFTLALLSLLYLLRCLFHFNMVKAEFLHH 125
+I P + ++ H+ +PS AF +LW LAL T + EFLH+
Sbjct: 61 IMIAP--ESPSMSHMHSKLPSMAFHLLWYLALVT-------------------QEEFLHY 99
Query: 126 VGVNYLFAPWISWFLLLQSAPFVAPKTATYLVLWWVFAVPVVVLDVKIYGQWFTKGKRFL 185
+GVNYL+AP ISW L+LQSAP + P + Y L+W+FAVPV+ LD+K+YGQWFT KRFL
Sbjct: 100 IGVNYLYAPSISWLLMLQSAPMMEPNSVLYQTLFWIFAVPVLTLDIKLYGQWFTTEKRFL 159
Query: 186 STAANPTSQLSVIGNLVGAQAAAHMGWKESAVCLFSLGMVHYLVLFVTLYQRLSGGNRLP 245
S ANP SQ+SVI NLV A+ AA MGW E A+C+FSLGMVHYLV+FVTLYQRL GGN P
Sbjct: 160 SMLANPASQVSVIANLVAARGAAEMGWNECALCMFSLGMVHYLVIFVTLYQRLPGGNNFP 219
Query: 246 VLLRPVFFLFFAAPGVASLAWGSIVGGFDTLSKMLFFLSLFLFMSLVCRPTLFRRSMKRF 305
LRP+FFLF AAP +ASLAW SI G FD ++KMLFFLSLF+FMSLVCRP LF++SMKRF
Sbjct: 220 AKLRPIFFLFVAAPAMASLAWNSICGTFDAVAKMLFFLSLFIFMSLVCRPNLFKKSMKRF 279
Query: 306 NVAWWAYSFPVTVLAMASTDYAEEVKGTISHVLMLVLLALSVLVSLALTVFTFINSKMLL 365
NVAWWAYSFP+T LA+ S YA+EVK + LML+ ++SVL+ L + V T NS LL
Sbjct: 280 NVAWWAYSFPLTFLALDSVQYAQEVKDPVGSGLMLIFSSISVLIFLGMMVLTAANSNRLL 339
Query: 366 PDD 368
D
Sbjct: 340 RHD 342
>UniRef100_Q5W6C1 Hypothetical protein OSJNBa0037H03.8 [Oryza sativa]
Length = 399
Score = 389 bits (998), Expect = e-107
Identities = 201/344 (58%), Positives = 248/344 (71%), Gaps = 12/344 (3%)
Query: 40 NSVLTKLHAGYFRISLSLGGQALLWKTLIGPTHDKSTLRHV--------VHMVPSSAFLV 91
+S+LT AGYFRISLSL GQALLW+TL G D H+ +P +A ++
Sbjct: 33 SSILTHFDAGYFRISLSLCGQALLWRTLCGGGGDGDGDEHMQPRALGALARHLPPAASVL 92
Query: 92 LWSLALFTLALLSLLYLLRCLFHFNMVKAEFLHHVGVNYLFAPWISWFLLLQSAP--FVA 149
LWSLAL +L L+ LY RCL F V+AEF H + VNYLFAPW SW LLLQSAP ++
Sbjct: 93 LWSLALLSLVALTALYAARCLLRFAAVRAEFRHRIAVNYLFAPWASWLLLLQSAPSSLLS 152
Query: 150 PKTATYLVLWWVFAVPVVVLDVKIYGQWFTKGKRFLSTAANPTSQLSVIGNLVGAQAAAH 209
P A VLW FA PV+ LDV +YGQWFT+G+ LS AANPT ++V+ NLV A+AAA
Sbjct: 153 PGAAPRRVLWCAFAAPVLALDVTVYGQWFTEGRTALSMAANPTGHITVVANLVTARAAAE 212
Query: 210 MGWKESAVCLFSLGMVHYLVLFVTLYQRLSGGNRLPVLLRPVFFLFFAAPGVASLAWGSI 269
+GW+E AV +F++ + HY VLFVTLYQRL G N LP +LRPVFFLFFAAP +ASLAWG+I
Sbjct: 213 LGWREGAVAVFAVAVAHYAVLFVTLYQRLLGANALPAMLRPVFFLFFAAPSMASLAWGAI 272
Query: 270 VGGFDTLSKMLFFLSLFLFMSLVCRPTLFRRSMKRFNVAWWAYSFPVTVLAMASTDYAEE 329
FDT KMLFFLSLFLF SLV RPTLFRR+M+RF+VAWWA+ FP+T LA+AS +YA E
Sbjct: 273 SSSFDTACKMLFFLSLFLFASLVSRPTLFRRAMRRFSVAWWAFPFPLTALAVASVEYARE 332
Query: 330 VKGTISHVLMLVLLALSVLVSLALTVFTFINSKMLLP--DDDPI 371
V+ + VL+LVL ALSV+V++A+ V T I + LLP DDDP+
Sbjct: 333 VEDHAAVVLVLVLSALSVVVTVAVVVCTAIRTSDLLPHGDDDPL 376
>UniRef100_Q9XHX8 OSJNBa0049B20.15 protein [Oryza sativa]
Length = 338
Score = 333 bits (855), Expect = 4e-90
Identities = 171/298 (57%), Positives = 208/298 (69%), Gaps = 10/298 (3%)
Query: 7 KPEIALVIDNTVSTIISREPPSFIVARRLLASLNSVLTKLHAGYFRISLSLGGQALLWKT 66
KP A ID V+ S +P + V R +LT+ HAGYFRISL+L GQALLW+T
Sbjct: 9 KPFAASSIDGGVA---STKPAAAPVVARF-----GMLTRFHAGYFRISLALSGQALLWRT 60
Query: 67 LIGPTHDKSTLRHVVHMVPSSAFLVLWSLALFTLALLSLLYLLRCLFHFNMVKAEFLHHV 126
L + D L VV +PS+AF++LWSLAL TL L LY RCL F V+AEF HHV
Sbjct: 61 LSDASTDPRALGPVVRSLPSAAFVLLWSLALLTLVALCALYAARCLLRFPAVRAEFRHHV 120
Query: 127 GVNYLFAPWISWFLLLQSAP--FVAPKTATYLVLWWVFAVPVVVLDVKIYGQWFTKGKRF 184
+NYLFAPWISW LLLQ+AP + P Y LWW F++P++ LDVK+YGQWFT+G++F
Sbjct: 121 AMNYLFAPWISWLLLLQAAPPLLLRPDARPYRALWWAFSLPILALDVKVYGQWFTRGRKF 180
Query: 185 LSTAANPTSQLSVIGNLVGAQAAAHMGWKESAVCLFSLGMVHYLVLFVTLYQRLSGGNRL 244
LS ANP S ++VIGNLV A+AAA MGW E AV +F++G HYLVLFVTLYQR G + L
Sbjct: 181 LSMVANPASHITVIGNLVTARAAARMGWHEGAVAMFAVGAAHYLVLFVTLYQRFLGSDSL 240
Query: 245 PVLLRPVFFLFFAAPGVASLAWGSIVGGFDTLSKMLFFLSLFLFMSLVCRPTLFRRSM 302
P +LRPVFFLFFAAP +ASLAW +I FDT KMLFFLSLFLF SLV + L +S+
Sbjct: 241 PAMLRPVFFLFFAAPSMASLAWDAISASFDTCCKMLFFLSLFLFASLVIKLALHSKSL 298
>UniRef100_Q9LD83 T12C24.3 [Arabidopsis thaliana]
Length = 556
Score = 256 bits (653), Expect = 9e-67
Identities = 144/335 (42%), Positives = 197/335 (57%), Gaps = 13/335 (3%)
Query: 43 LTKLHAGYFRISLSLGGQALLWKTLIGPTHDKSTLRHVVHMVPSSAFLVLWSLALFTLAL 102
L + G F I L L QA+LW L KS + +H+ P LV+W +L L
Sbjct: 185 LLRFPIGCFGICLGLSSQAVLWLALA-----KSPATNFLHITPLIN-LVVWLFSLVVLVS 238
Query: 103 LSLLYLLRCLFHFNMVKAEFLHHVGVNYLFAPWISWFLLLQSAPFVAPKTATYL--VLWW 160
+S Y+L+C+F+F VK E+ H V VN+ FAPW+ L S P + YL +W
Sbjct: 239 VSFTYILKCIFYFEAVKREYFHPVRVNFFFAPWVVCMFLAISVPPMFSPNRKYLHPAIWC 298
Query: 161 VFAVPVVVLDVKIYGQWFTKGKRFLSTAANPTSQLSVIGNLVGAQAAAHMGWKESAVCLF 220
VF P L++KIYGQW + GKR L ANP+S LSV+GN VGA A+ +GW E A L+
Sbjct: 299 VFMGPYFFLELKIYGQWLSGGKRRLCKVANPSSHLSVVGNFVGAILASKVGWDEVAKFLW 358
Query: 221 SLGMVHYLVLFVTLYQRLSGGNRLPVLLRPVFFLFFAAPGVASLAWGSIVGGFDTLSKML 280
++G HYLV+FVTLYQRL LP L PV+ +F AAP AS+AW +I G FD S+
Sbjct: 359 AVGFAHYLVVFVTLYQRLPTSEALPKELHPVYSMFIAAPSAASIAWNTIYGQFDGCSRTC 418
Query: 281 FFLSLFLFMSLVCRPTLFRRSMKRFNVAWWAYSFPVTVLAMASTDYAEEVKGTISHVLML 340
FF++LFL++SLV R F + +F+VAWW+Y+FP+T ++A+ YAE V G S L L
Sbjct: 419 FFIALFLYISLVARINFF--TGFKFSVAWWSYTFPMTTASVATIKYAEAVPGYPSRALAL 476
Query: 341 VLLALSVLVSLALTVFTFINS---KMLLPDDDPIA 372
L +S + L V T +++ + L P+D IA
Sbjct: 477 TLSFISTAMVCVLFVSTLLHAFVWQTLFPNDLAIA 511
>UniRef100_Q5ZE68 Hypothetical protein P0501G01.17 [Oryza sativa]
Length = 686
Score = 254 bits (648), Expect = 4e-66
Identities = 136/325 (41%), Positives = 197/325 (59%), Gaps = 13/325 (4%)
Query: 51 FRISLSLGGQALLWKTLIGPTHDKSTLRHVVHMVPSSAFLVLWSLALFTLALLSLLYLLR 110
F + L + QA+LWKTL ++ HV +V VLW ++L + +S +YLL+
Sbjct: 312 FGMCLGVSSQAMLWKTLASAP--PTSFLHVSPVVNH----VLWWISLALMGFVSFIYLLK 365
Query: 111 CLFHFNMVKAEFLHHVGVNYLFAPWISWFLLLQSAPFVAPKTATYLVLWWVFAVPVVVLD 170
+F+F V+ EF H + N+ FAPWI+ L+Q P P T + +W+ P+ L+
Sbjct: 366 VVFYFEAVRREFYHPIRANFFFAPWIACLFLVQGVP--RPVTEVHHGVWYALMAPIFCLE 423
Query: 171 VKIYGQWFTKGKRFLSTAANPTSQLSVIGNLVGAQAAAHMGWKESAVCLFSLGMVHYLVL 230
+KIYGQW + G+R LS ANP++ LS++GN VGA A MG +E + F++G+ HY+VL
Sbjct: 424 LKIYGQWMSGGQRRLSKVANPSNHLSIVGNFVGALLGAKMGLREGPIFYFAVGLAHYMVL 483
Query: 231 FVTLYQRLSGGNRLPVLLRPVFFLFFAAPGVASLAWGSIVGGFDTLSKMLFFLSLFLFMS 290
FVTLYQRL LP L PVFFLF AAP VAS+AW I+G FD +++ +F++LFL+MS
Sbjct: 484 FVTLYQRLPTNVTLPKELHPVFFLFVAAPSVASMAWAKILGEFDYGARIAYFIALFLYMS 543
Query: 291 LVCRPTLFRRSMKRFNVAWWAYSFPVTVLAMASTDYAEEVKGTISHVLMLVLLALSVLVS 350
L R FR RF++AWWAY+FP+T A+A+ YA EV ++ L + L ++ +
Sbjct: 544 LAVRINFFRGF--RFSLAWWAYTFPMTGAAIATITYATEVTNVLTRALSIGLSGIATVTV 601
Query: 351 LALTVFTFINS---KMLLPDDDPIA 372
L V T ++ K L P+D IA
Sbjct: 602 AGLLVTTMFHAFVLKDLFPNDVSIA 626
>UniRef100_Q9ASQ7 AT4g27970/T13J8_80 [Arabidopsis thaliana]
Length = 519
Score = 251 bits (640), Expect = 3e-65
Identities = 138/327 (42%), Positives = 196/327 (59%), Gaps = 16/327 (4%)
Query: 51 FRISLSLGGQALLWKTLIGPTHDKSTLRHVVHMVPSSAFLVLWSLALFTLALLSLLYLLR 110
+ + L + QA++WKTL T + HV ++ VLW ++L L +S+ YL +
Sbjct: 148 YGMCLGVSSQAIMWKTLA--TTEAEKFLHVTQVINH----VLWWISLLLLLAVSITYLFK 201
Query: 111 CLFHFNMVKAEFLHHVGVNYLFAPWISWFLLLQSAP--FVAPKTATYLVLWWVFAVPVVV 168
+ F V+ EF H + VN+ FAP IS L P ++ +T LW+ P++
Sbjct: 202 TILFFEAVRREFRHPIRVNFFFAPLISILFLALGIPHSIISHLPST---LWYFLMAPILF 258
Query: 169 LDVKIYGQWFTKGKRFLSTAANPTSQLSVIGNLVGAQAAAHMGWKESAVCLFSLGMVHYL 228
L++KIYGQW + G+R LS ANPT+ LS++GN GA A MG KE + F++G+ +YL
Sbjct: 259 LEMKIYGQWMSGGQRRLSKVANPTNHLSIVGNFAGALLGASMGLKEGPIFFFAIGLAYYL 318
Query: 229 VLFVTLYQRLSGGNRLPVLLRPVFFLFFAAPGVASLAWGSIVGGFDTLSKMLFFLSLFLF 288
VLFVTLYQRL LP L PVFFLF AAP VAS+AW I FD S++ +F+SLFL+
Sbjct: 319 VLFVTLYQRLPTNETLPKELHPVFFLFVAAPAVASMAWTKISASFDLGSRLAYFISLFLY 378
Query: 289 MSLVCRPTLFRRSMKRFNVAWWAYSFPVTVLAMASTDYAEEVKGTISHVLMLVL---LAL 345
SLVCR LFR +F++AWWAY+FP+T +A A+ Y++EV G + +L +V+ L
Sbjct: 379 FSLVCRINLFRGF--KFSLAWWAYTFPMTAVASATIKYSDEVTGVATKILSVVMSGAATL 436
Query: 346 SVLVSLALTVFTFINSKMLLPDDDPIA 372
+V+ L LTV + L P+D IA
Sbjct: 437 TVIAVLGLTVMHAFVQRDLFPNDVVIA 463
>UniRef100_Q7XTX7 OSJNBa0019K04.20 protein [Oryza sativa]
Length = 578
Score = 251 bits (640), Expect = 3e-65
Identities = 136/333 (40%), Positives = 192/333 (56%), Gaps = 11/333 (3%)
Query: 43 LTKLHAGYFRISLSLGGQALLWKTLIGPTHDKSTLRHVVHMVPSSAFLVLWSLALFTLAL 102
L + G F + L LG QA+LW L S +H+ P + LW LAL L
Sbjct: 198 LLRFPVGCFGVCLGLGSQAILWGALAA-----SPAMRFLHVTPMIN-VALWLLALAVLVA 251
Query: 103 LSLLYLLRCLFHFNMVKAEFLHHVGVNYLFAPWISWFLLLQSAPFVAPKTATYLVLWWVF 162
+S+ Y L+C+F+F ++ E+ H V VN+ FAP I+ L P + +W F
Sbjct: 252 VSVTYALKCVFYFEAIRREYFHPVRVNFFFAPSIAAMFLTIGLPRAVAPERLHPAVWCAF 311
Query: 163 AVPVVVLDVKIYGQWFTKGKRFLSTAANPTSQLSVIGNLVGAQAAAHMGWKESAVCLFSL 222
P+ L++KIYGQW + GKR L ANP+S LSV+GN VGA AA +GW E+ L+++
Sbjct: 312 VAPLFGLELKIYGQWLSGGKRRLCKVANPSSHLSVVGNFVGAILAARVGWAEAGKFLWAI 371
Query: 223 GMVHYLVLFVTLYQRLSGGNRLPVLLRPVFFLFFAAPGVASLAWGSIVGGFDTLSKMLFF 282
G+ HY+V+FVTLYQRL LP L PV+ +F A P ASLAW +I G FD +++ FF
Sbjct: 372 GVAHYIVVFVTLYQRLPTNEALPKELHPVYSMFIATPSAASLAWAAIYGSFDAVARTFFF 431
Query: 283 LSLFLFMSLVCRPTLFRRSMKRFNVAWWAYSFPVTVLAMASTDYAEEVKGTISHVLMLVL 342
++LFL+MSLV R FR RF++AWW+Y+FP+T ++A+ YAE S L L L
Sbjct: 432 MALFLYMSLVVRINFFRGF--RFSIAWWSYTFPMTTASLATVKYAEAEPCFTSRALALSL 489
Query: 343 LALSVLVSLALTVFTFINS---KMLLPDDDPIA 372
+S + L V T +++ + L P+D IA
Sbjct: 490 SLMSTTMVSLLLVSTLLHAFVWRSLFPNDLAIA 522
>UniRef100_Q8L4G8 C4-dicarboxylate transporter-like protein [Oryza sativa]
Length = 625
Score = 250 bits (638), Expect = 5e-65
Identities = 140/335 (41%), Positives = 195/335 (57%), Gaps = 16/335 (4%)
Query: 43 LTKLHAGYFRISLSLGGQALLWKTLIGPTHDKSTLRHVVHMVPSSAFLVLWSLALFTLAL 102
L + F I L + QA+LWKT+ ST +H V + LVLW ++L + +
Sbjct: 262 LLRFPVSAFGICLGVSSQAILWKTVA-----TSTPTRFLH-VTTKVNLVLWCVSLALMCV 315
Query: 103 LSLLYLLRCLFHFNMVKAEFLHHVGVNYLFAPWISWFLLLQSAPFVAPKTATYLV--LWW 160
++ +Y + +F F V+ E+ H + VN+ FAPWI+ L P P AT L LW+
Sbjct: 316 IAAIYACKVVFFFEAVRREYYHPIRVNFFFAPWIACLFLAIGVP---PSVATELPRWLWY 372
Query: 161 VFAVPVVVLDVKIYGQWFTKGKRFLSTAANPTSQLSVIGNLVGAQAAAHMGWKESAVCLF 220
P++ +++KIYGQW + G+R LS ANP++ LSV+GN VGA A MG KE V F
Sbjct: 373 ALMTPILCMELKIYGQWMSGGQRRLSKVANPSNHLSVVGNFVGALLGASMGLKEGPVFFF 432
Query: 221 SLGMVHYLVLFVTLYQRLSGGNRLPVLLRPVFFLFFAAPGVASLAWGSIVGGFDTLSKML 280
S+G+ HY VLFVTLYQRL LP L PVFFLF AAP VA +AW I G F S++
Sbjct: 433 SVGLAHYTVLFVTLYQRLPTNETLPKELHPVFFLFVAAPSVACMAWAKITGEFGLGSRVA 492
Query: 281 FFLSLFLFMSLVCRPTLFRRSMKRFNVAWWAYSFPVTVLAMASTDYAEEVKGTISHVLML 340
+F+++FL+ SL R FR RF++AWWAY+FP+T A+AS Y+ EV + L +
Sbjct: 493 YFIAMFLYASLAVRINFFRGF--RFSLAWWAYTFPMTGAAIASIRYSTEVDNAFTKALCV 550
Query: 341 VLLALSVLVSLALTVFTFINS---KMLLPDDDPIA 372
L L++L LAL T ++ + L P+D IA
Sbjct: 551 ALSVLAMLTVLALLATTIVHGFVLRNLFPNDISIA 585
>UniRef100_Q9FLV9 Similarity to unknown protein [Arabidopsis thaliana]
Length = 635
Score = 249 bits (637), Expect = 7e-65
Identities = 136/327 (41%), Positives = 199/327 (60%), Gaps = 16/327 (4%)
Query: 51 FRISLSLGGQALLWKTLIGPTHDKSTLRHVVHMVPSSAFLVLWSLALFTLALLSLLYLLR 110
F + L + QA++WKTL T + + HV + LW +++ + ++ +YLL+
Sbjct: 263 FGMCLGVSSQAIMWKTLA--TAEPTKFLHVPLWINQG----LWFISVALILTIATIYLLK 316
Query: 111 CLFHFNMVKAEFLHHVGVNYLFAPWISWFLLLQSAPFVAPKTATYL--VLWWVFAVPVVV 168
+ F V+ E+ H + +N+ FAP+IS L P P T L LW++ P +
Sbjct: 317 IILFFEAVRREYYHPIRINFFFAPFISLLFLALGVP---PSIITDLPHFLWYLLMFPFIC 373
Query: 169 LDVKIYGQWFTKGKRFLSTAANPTSQLSVIGNLVGAQAAAHMGWKESAVCLFSLGMVHYL 228
L++KIYGQW + G+R LS ANPT+ LSV+GN VGA A MG +E + +++GM HYL
Sbjct: 374 LELKIYGQWMSGGQRRLSRVANPTNHLSVVGNFVGALLGASMGLREGPIFFYAVGMAHYL 433
Query: 229 VLFVTLYQRLSGGNRLPVLLRPVFFLFFAAPGVASLAWGSIVGGFDTLSKMLFFLSLFLF 288
VLFVTLYQRL LP L PVFFLF AAP VAS+AW + G FD SK+ +F+++FL+
Sbjct: 434 VLFVTLYQRLPTNETLPKDLHPVFFLFVAAPSVASMAWAKVTGSFDYGSKVCYFIAIFLY 493
Query: 289 MSLVCRPTLFRRSMKRFNVAWWAYSFPVTVLAMASTDYAEEVKGTISHVLMLVLLALSVL 348
SL R FR +F+++WWAY+FP+T A+A+ YA VK T++ ++ +VL A++ L
Sbjct: 494 FSLAVRINFFRGI--KFSLSWWAYTFPMTGAAIATIRYATVVKSTMTQIMCVVLCAIATL 551
Query: 349 VSLALTVFTFINS---KMLLPDDDPIA 372
V AL V T I++ + L P+D IA
Sbjct: 552 VVFALLVTTIIHAFVLRDLFPNDLAIA 578
>UniRef100_Q6I589 Hypothetical protein OSJNBa0009C07.7 [Oryza sativa]
Length = 377
Score = 247 bits (631), Expect = 3e-64
Identities = 133/329 (40%), Positives = 200/329 (60%), Gaps = 29/329 (8%)
Query: 55 LSLGGQALLWKTLIGPTHDKSTLRHVVHMVPSSAFL--------VLWSLALFTLALLSLL 106
L + QA+LWKTL PS+AFL VLW +++ +AL+S +
Sbjct: 3 LGVSSQAMLWKTLASE--------------PSTAFLHISLDVNHVLWWVSVALMALVSAI 48
Query: 107 YLLRCLFHFNMVKAEFLHHVGVNYLFAPWISWFLLLQSAPFVAPKTATYLVLWWVFAVPV 166
YLL+ +F+F V+ EF H + VN+ FAPWI+ L++ P + V+W++ P+
Sbjct: 49 YLLKVVFYFEAVRREFHHPIRVNFFFAPWIACLFLVKGLP--RQVWTIHHVVWFLLMAPI 106
Query: 167 VVLDVKIYGQWFTKGKRFLSTAANPTSQLSVIGNLVGAQAAAHMGWKESAVCLFSLGMVH 226
++LD+KIYGQW + G+R LS ANP++ L+++GN VGA A MG +E + ++G+VH
Sbjct: 107 LLLDLKIYGQWMSGGERRLSKVANPSNHLAIVGNFVGALLGARMGLREGPIFFLAVGLVH 166
Query: 227 YLVLFVTLYQRLSGGNRLPVLLRPVFFLFFAAPGVASLAWGSIVGGFDTLSKMLFFLSLF 286
Y+VLFVTLYQRL +LP L PVFFLF AAP VAS+AW + G FD +++ +F++LF
Sbjct: 167 YIVLFVTLYQRLPTNVQLPKELHPVFFLFIAAPSVASMAWARLTGEFDFGARIAYFVALF 226
Query: 287 LFMSLVCRPTLFRRSMKRFNVAWWAYSFPVTVLAMASTDYAEEVKGTISHVLMLVLLALS 346
L+MSL R +FR RF++AWWAY+FP+T A+A+ YA EV + + + L ++
Sbjct: 227 LYMSLAVRVNMFRGF--RFSLAWWAYTFPMTSAAIATVLYASEVTNVATRAMAVGLSGIA 284
Query: 347 VLVSLALTVFTFINS---KMLLPDDDPIA 372
+ + V T ++ + L P+D IA
Sbjct: 285 TVTVTGVLVTTMYHAFVRRDLFPNDVSIA 313
>UniRef100_Q75G55 Hypothetical protein B1003C08.2 [Oryza sativa]
Length = 622
Score = 246 bits (629), Expect = 6e-64
Identities = 139/333 (41%), Positives = 191/333 (56%), Gaps = 11/333 (3%)
Query: 43 LTKLHAGYFRISLSLGGQALLWKTLIGPTHDKSTLRHVVHMVPSSAFLVLWSLALFTLAL 102
L + F + L + QA+LWKT+ T + HV V LVLW +++ +
Sbjct: 256 LLRFPVSAFGMCLGVSSQAILWKTIA--TSGPTAFLHVTTKVN----LVLWCVSVALMCA 309
Query: 103 LSLLYLLRCLFHFNMVKAEFLHHVGVNYLFAPWISWFLLLQSAPFVAPKTATYLVLWWVF 162
+S Y + +F F V+ E+ H + VN+ FAPWI+ L P T LW+
Sbjct: 310 ISATYGAKVVFFFEAVRREYYHPIRVNFFFAPWIACLFLTIGVPDSVAPTLLPHWLWYAL 369
Query: 163 AVPVVVLDVKIYGQWFTKGKRFLSTAANPTSQLSVIGNLVGAQAAAHMGWKESAVCLFSL 222
PV+ L++KIYGQW + G+R LS ANP++ LSV+GN VGA A MG +E V F++
Sbjct: 370 MAPVLCLELKIYGQWMSGGQRRLSKVANPSNHLSVVGNFVGALLGASMGLREGPVFFFAV 429
Query: 223 GMVHYLVLFVTLYQRLSGGNRLPVLLRPVFFLFFAAPGVASLAWGSIVGGFDTLSKMLFF 282
GM HY VLFVTLYQRL LP L PVFFLF AAP VAS+AW I G F S++ +F
Sbjct: 430 GMAHYSVLFVTLYQRLPTNETLPKELHPVFFLFVAAPSVASMAWARITGEFGLGSRVAYF 489
Query: 283 LSLFLFMSLVCRPTLFRRSMKRFNVAWWAYSFPVTVLAMASTDYAEEVKGTISHVLMLVL 342
+++FL+ SL R FR RF++AWWAY+FP+T A+AS Y+ EV ++ L + L
Sbjct: 490 IAMFLYASLAVRINFFRGF--RFSLAWWAYTFPMTGAAIASIRYSTEVDNALTRALCVAL 547
Query: 343 LALSVLVSLALTVFTFINS---KMLLPDDDPIA 372
A++ LV AL T I++ L P+D IA
Sbjct: 548 SAVATLVVTALFATTMIHAFVLHKLFPNDIAIA 580
>UniRef100_Q8LRK1 P0443E07.21 protein [Oryza sativa]
Length = 714
Score = 245 bits (626), Expect = 1e-63
Identities = 137/334 (41%), Positives = 197/334 (58%), Gaps = 14/334 (4%)
Query: 43 LTKLHAGYFRISLSLGGQALLWKTLIGPTHDKSTLRHVVHMVPSSAFLVLWSLALFTLAL 102
L + F + + + QA+LWK + ST +H + LVLW +++ +
Sbjct: 320 LLRFPVSAFGMCMGMSSQAILWKNIA-----ISTSTRFLH-ITVKINLVLWCVSVALMCA 373
Query: 103 LSLLYLLRCLFHFNMVKAEFLHHVGVNYLFAPWISW-FLLLQSAPFVAPKTATYLVLWWV 161
+S LY + F+F V+ E+ H V VN+ FAPWI+ FL + P VA +L W++
Sbjct: 374 VSALYACKVAFYFEAVRREYYHPVRVNFFFAPWIACLFLAIGVPPMVAASLPHWL--WYL 431
Query: 162 FAVPVVVLDVKIYGQWFTKGKRFLSTAANPTSQLSVIGNLVGAQAAAHMGWKESAVCLFS 221
P+V L++KIYGQW + G+R LS ANP++ LS++GN VGA A MG +E + F+
Sbjct: 432 LMAPIVCLELKIYGQWISGGQRRLSRVANPSNHLSIVGNFVGALLGATMGLREGPIFFFA 491
Query: 222 LGMVHYLVLFVTLYQRLSGGNRLPVLLRPVFFLFFAAPGVASLAWGSIVGGFDTLSKMLF 281
+G+ HY+VLFVTLYQRL LP L PVFFLF AAP VA LAW I G F S++ +
Sbjct: 492 VGLAHYIVLFVTLYQRLPTSETLPRDLHPVFFLFVAAPSVACLAWARITGEFGYGSRVAY 551
Query: 282 FLSLFLFMSLVCRPTLFRRSMKRFNVAWWAYSFPVTVLAMASTDYAEEVKGTISHVLMLV 341
F+++FL+ SL R LFR RF++AWWAY+FP+T A+AS Y+ EVK + L +
Sbjct: 552 FIAMFLYASLAVRINLFRGF--RFSLAWWAYTFPMTSAAIASIRYSSEVKNAFTQSLCIA 609
Query: 342 LLALSVLVSLALTVFTFINSKM---LLPDDDPIA 372
L L+ L AL + T +++ + L P+D IA
Sbjct: 610 LSVLATLTVTALFLTTLLHAAVHRDLFPNDISIA 643
>UniRef100_Q6ZLB6 Hypothetical protein OJ1014_E09.31 [Oryza sativa]
Length = 624
Score = 236 bits (601), Expect = 1e-60
Identities = 140/336 (41%), Positives = 196/336 (57%), Gaps = 12/336 (3%)
Query: 43 LTKLHAGYFRISLSLGGQALLWKTLIGPTHDKSTLRHVVHMVPSSAFLVLWSLALFTLAL 102
L + F I L +G QA+LWK + T R++ V + LVLW L++
Sbjct: 247 LLRFPVSAFGICLGMGSQAILWKRI---AESPPTTRYL--HVAADVNLVLWWLSVALTCA 301
Query: 103 LSLLYLLRCLFHFNMVKAEFLHHVGVNYLFAPWISWFLLLQSAP-FVAPKTATYLV-LWW 160
+S +Y + +F F V+ E+LH V VN+ FAP I+ L P VAP TA LW+
Sbjct: 302 VSAVYACKVVFFFEAVRREYLHPVRVNFFFAPLIACLFLAIGVPRSVAPSTAALPAWLWY 361
Query: 161 VFAVPVVVLDVKIYGQWFTKGKRFLSTAANPTSQLSVIGNLVGAQAAAHMGWKESAVCLF 220
P++ L++KIYGQW + G+R LS ANP++ LSV+GN VGA A MG +E AV F
Sbjct: 362 ALMAPMLCLELKIYGQWMSSGQRRLSMVANPSNHLSVVGNFVGALLGASMGIREGAVFFF 421
Query: 221 SLGMVHYLVLFVTLYQRLSGGNRLPVLLRPVFFLFFAAPGVASLAWGSIVGGFDTLSKML 280
++G+ HY+VLFVTLYQRL LP L PVFFLF A P VAS+AW +I G F +++
Sbjct: 422 AVGVAHYVVLFVTLYQRLPTNEALPRELHPVFFLFVATPSVASVAWAAIAGEFALGARLA 481
Query: 281 FFLSLFLFMSLVCRPTLFRRSMKRFNVAWWAYSFPVTVLAMASTDYAEEVKGT-ISHVLM 339
+F+++FL+ SL R + RF++AWWAY+FP+T A A+ YA EV+ T ++ L
Sbjct: 482 YFVAMFLYASLAARAVSLFGGV-RFSLAWWAYTFPMTSAAAATIRYAAEVEDTRLARALC 540
Query: 340 LVLLA---LSVLVSLALTVFTFINSKMLLPDDDPIA 372
+ L A L+V A TV + + L P+D IA
Sbjct: 541 VALAAAATLTVGCLFATTVVHAVVLRSLFPNDVAIA 576
>UniRef100_Q9SUE2 Hypothetical protein T13J8.80 [Arabidopsis thaliana]
Length = 505
Score = 218 bits (556), Expect = 2e-55
Identities = 127/327 (38%), Positives = 183/327 (55%), Gaps = 30/327 (9%)
Query: 51 FRISLSLGGQALLWKTLIGPTHDKSTLRHVVHMVPSSAFLVLWSLALFTLALLSLLYLLR 110
+ + L + QA++WKTL T + HV ++ VLW ++L L +S+ YL +
Sbjct: 148 YGMCLGVSSQAIMWKTLA--TTEAEKFLHVTQVINH----VLWWISLLLLLAVSITYLFK 201
Query: 111 CLFHFNMVKAEFLHHVGVNYLFAPWISWFLLLQSAP--FVAPKTATYLVLWWVFAVPVVV 168
+ F V+ EF H + VN+ FAP IS L P ++ +T LW+ P++
Sbjct: 202 TILFFEAVRREFRHPIRVNFFFAPLISILFLALGIPHSIISHLPST---LWYFLMAPILF 258
Query: 169 LDVKIYGQWFTKGKRFLSTAANPTSQLSVIGNLVGAQAAAHMGWKESAVCLFSLGMVHYL 228
L++KIYGQW + G+R LS ANPT+ LS++GN GA A MG KE + F++G
Sbjct: 259 LEMKIYGQWMSGGQRRLSKVANPTNHLSIVGNFAGALLGASMGLKEGPIFFFAIG----- 313
Query: 229 VLFVTLYQRLSGGNRLPVLLRPVFFLFFAAPGVASLAWGSIVGGFDTLSKMLFFLSLFLF 288
L LP L PVFFLF AAP VAS+AW I FD S++ +F+SLFL+
Sbjct: 314 ---------LPTNETLPKELHPVFFLFVAAPAVASMAWTKISASFDLGSRLAYFISLFLY 364
Query: 289 MSLVCRPTLFRRSMKRFNVAWWAYSFPVTVLAMASTDYAEEVKGTISHVLMLVL---LAL 345
SLVCR LFR +F++AWWAY+FP+T +A A+ Y++EV G + +L +V+ L
Sbjct: 365 FSLVCRINLFRGF--KFSLAWWAYTFPMTAVASATIKYSDEVTGVATKILSVVMSGAATL 422
Query: 346 SVLVSLALTVFTFINSKMLLPDDDPIA 372
+V+ L LTV + L P+D IA
Sbjct: 423 TVIAVLGLTVMHAFVQRDLFPNDVVIA 449
>UniRef100_Q6LHX2 Hypothetical protein WS0635 [Photobacterium profundum)]
Length = 311
Score = 114 bits (284), Expect = 6e-24
Identities = 89/325 (27%), Positives = 155/325 (47%), Gaps = 30/325 (9%)
Query: 51 FRISLSLGGQALLWKTLIGPTHDKSTLRHVVHMVPSSAFLVLWSLALFTLALLSLLYLLR 110
F I + L G + WK + TL + +V S+ F++L Y +
Sbjct: 9 FSIVMGLIGLGIAWKECFQTSSVAQTLSLAIGIVASALFIILLGA-----------YGWK 57
Query: 111 CLFHFNMVKAEFLHHVGVNYLFAPWISWFLLL-----QSAPFVAPKTATYLVLWWVFAVP 165
H + V+AE H V +N F P IS +LL QS P +A + LW +
Sbjct: 58 IFRHASDVRAELHHPVRLN--FFPTISVNMLLLSVFWQSIPILA------ITLWTAGMLL 109
Query: 166 VVVLDVKIYGQWFTKGKRFLSTAANPTSQLSVIGNLVGAQAAAHMGWKESAVCLFSLGMV 225
+VL + + W F NP+ + ++GN+V ++G+ E + FS+G++
Sbjct: 110 QLVLTIYVMSSWLNHN-HFTIEHVNPSWFIPMVGNIVVPINGMYLGYVEISWFFFSVGLI 168
Query: 226 HYLVLFVTLYQRLSGGNRLPVLLRPVFFLFFAAPGVASLAWGSIVGGFDTLSKMLFFLSL 285
+LVL + RL +LP+ L P +F+ A P + +++ ++GG D +++L++ +L
Sbjct: 169 FWLVLLTIVMYRLLFHEQLPLHLTPTYFILLAPPSIGFVSYTGLIGGLDNAARLLYYCAL 228
Query: 286 FLFMSLVCRPTLFRRSMKRFNVAWWAYSFPVTVLAMASTDYAE-EVKGTISHVLMLVLLA 344
FL M L F R F ++ WA+SFP+ L++A+ AE + + + L++
Sbjct: 229 FLMMLLGSNAIRFWR--VPFFISSWAFSFPLAALSIATIKMAELSQQSCLIWLARLLVTL 286
Query: 345 LSVLVSLAL--TVFTFINSKMLLPD 367
LSV+VS + TV K+ +P+
Sbjct: 287 LSVIVSWLVFRTVQAARQGKICIPE 311
>UniRef100_Q7W425 Putative inner membrane transport protein [Bordetella
parapertussis]
Length = 353
Score = 102 bits (255), Expect = 1e-20
Identities = 85/305 (27%), Positives = 136/305 (43%), Gaps = 17/305 (5%)
Query: 51 FRISLSLGGQALLWKTLIGPTHDKSTLRHVVHMVPSSAFLVLWSLALFTLALLSLLYLLR 110
F L L G + W+ H V +P L + +L L+LLR
Sbjct: 46 FASVLGLSGLGMAWRKA-----------HQVWGLPLQVALAIQVAVAAVYGVLLALFLLR 94
Query: 111 CLFHFNMVKAEFLHHVGVNYLFAPWISWFLLLQSAPFVAPKTATYLVLWWVFAVPVVVLD 170
L H + + E+ H V + + A + L+ + AP+ A LWW+ A +
Sbjct: 95 LLRHPDEIAREWNHPVQLAFFSAISLGVVLIGTAWADDAPRAAA--ALWWLGATVHLGFT 152
Query: 171 VKIYGQWFTKGKRFLSTAANPTSQLSVIGNLVGAQAAAHMGWKESAVCLFSLGMVHYLVL 230
+ I W + NP+ + V+GN+V A E A FSLG+V ++VL
Sbjct: 153 LVIVRSWIFH-THYQLDHVNPSWFVPVVGNIVIPIAGVRFAPAECAWFFFSLGIVFWIVL 211
Query: 231 FVTLYQRLSGGNRLPVLLRPVFFLFFAAPGVASLAWGSIVGGFDTLSKMLFFLSLFLFMS 290
+ RL G L L P F+ A P VA LA+ ++ G D +++L+++SLF +
Sbjct: 212 KAIILYRLMFGAPLARALTPTLFIMIAPPAVAFLAYMALTGSLDGFARVLYYISLFWTLL 271
Query: 291 LVCRPTLFRRSMKRFNVAWWAYSFPVTVLAMASTDYAEEVKGTIS-HVLMLVLLALSVLV 349
L F R F++A W+YSFP+ + +A+ +AE ++ H L L+A+ V
Sbjct: 272 LSTAAPRFAR--LPFSLAAWSYSFPLAAITLATFSFAEHSGHALAFHTLAGFLVAVLSTV 329
Query: 350 SLALT 354
+ LT
Sbjct: 330 LVLLT 334
>UniRef100_Q7M9V8 SIMILARITY TO mviN PROTEIN [Wolinella succinogenes]
Length = 322
Score = 98.2 bits (243), Expect = 3e-19
Identities = 76/277 (27%), Positives = 128/277 (45%), Gaps = 16/277 (5%)
Query: 79 HVVHMVPSSAFLVLWSLALFTLALLSLLYLLRCLFHFNMVKAEFLHHVGVNYLFAPWISW 138
H + +P F L +L ++ LY ++ L H K EF H + +N+ A I+
Sbjct: 37 HEILGIPLLVFEGLRALTSLLFLVVFSLYAMKFLTHRAGAKKEFSHPIRINFFAAFSIAL 96
Query: 139 FLLL---QSAPFVAPKTATYLVLWWVFAVPVVVLDVKIYGQWFTKGKRFLSTAANPTSQL 195
LL + P + L+W+ A L W F T +NP +
Sbjct: 97 LLLAILWRGVPSLGGS------LFWLGAGVHTFLTFYTVSFWINNN--FEITHSNPAWFI 148
Query: 196 SVIGNLVGAQAAAHMGWKESAVCLFSLGMVHYLVLFVTLYQRLSGGNRLPVLLRPVFFLF 255
++GNL+ A M F++G+ ++VLF ++ R+ N+LP P F+F
Sbjct: 149 PIVGNLIVPVAGDGMASATFLGYYFAIGIFFWVVLFTVVFYRIIFHNQLPQKFMPTLFIF 208
Query: 256 FAAPGVASLAWGSIVGGFDTLSKMLFFLSLFLFMSLVCRPTLFRRSMK-RFNVAWWAYSF 314
A P V LA+ + GGFD ++ L L+LF L+ +++ MK +F ++WWA++F
Sbjct: 209 IAPPAVGFLAYLKVGGGFDLFAQFLLNLALFFAFLLL---FMYKNFMKLQFFISWWAFTF 265
Query: 315 PVTVLAMA-STDYAEEVKGTISHVLMLVLLALSVLVS 350
P+ L +A Y + + H+ L LL ++LV+
Sbjct: 266 PMAALTIALLVAYEKSHEAVYLHLGELFLLFATLLVA 302
>UniRef100_UPI00002B2C06 UPI00002B2C06 UniRef100 entry
Length = 314
Score = 90.9 bits (224), Expect = 5e-17
Identities = 85/307 (27%), Positives = 130/307 (41%), Gaps = 20/307 (6%)
Query: 55 LSLGGQALLWKTLIGPTHDKSTLRHVVHMVPSSAFLVLWSLALFTLALLSLLYLLRCLFH 114
+ L G A+ W+ L V PS V+ ++AL +L+ YL + H
Sbjct: 10 MGLTGLAVAWR-----------LAQVHFGAPSWVAQVVGAIALAVFVVLAAAYLAKLAIH 58
Query: 115 FNMVKAEFLHHVGVNYLFAPWISWFLLLQSAPFVAPKTATYLVLWWVFAVPVVVLDVKIY 174
+ V+AEF H + N AP IS LLL V VLW + + +L +
Sbjct: 59 PDRVRAEFAHPIAGNMFGAPMIS--LLLIPLLLVDQHLLLARVLWCIGVAGMTLLAWWMV 116
Query: 175 GQWFTKGKRFLSTAANPTSQLSVIGNLVGAQAAAHMGWKESAVCLFS----LGMVHYLVL 230
+W + ++ A P + V+G L AA +GW L + +G+ + L
Sbjct: 117 MRWISVRQQ--PEHATPAWIVPVVGMLDIPLAAPALGWAHGMHGLMAFGTAVGLFFAVPL 174
Query: 231 FVTLYQRLSGGNRLPVLLRPVFFLFFAAPGVASLAWGSIVGGFDTLSKMLFFLSLFLFMS 290
F + RL P +RP + A V A+ + G D + L+ + LF+
Sbjct: 175 FTMIVSRLMFAEPFPDAMRPSLLILCAPFAVGFSAYVATTGVIDDFATGLYMVMLFVLAV 234
Query: 291 LVCRPTLFRRSMKRFNVAWWAYSFPVTVLAMASTDYAEEVKGTISHVLMLVLLALSVLVS 350
LV R R F V+WWA SFP+ A+A+ YA + + + VLL L+ LV
Sbjct: 235 LVTRLWHLGRCCP-FRVSWWAVSFPLAASAVAALKYAGFAQHPSADAIAAVLLGLASLVI 293
Query: 351 LALTVFT 357
AL + T
Sbjct: 294 AALFLRT 300
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.330 0.141 0.438
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 587,447,234
Number of Sequences: 2790947
Number of extensions: 22740029
Number of successful extensions: 72596
Number of sequences better than 10.0: 114
Number of HSP's better than 10.0 without gapping: 41
Number of HSP's successfully gapped in prelim test: 74
Number of HSP's that attempted gapping in prelim test: 72431
Number of HSP's gapped (non-prelim): 143
length of query: 377
length of database: 848,049,833
effective HSP length: 129
effective length of query: 248
effective length of database: 488,017,670
effective search space: 121028382160
effective search space used: 121028382160
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.9 bits)
S2: 75 (33.5 bits)
Lotus: description of TM0195a.11


