

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0193.4
(1140 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
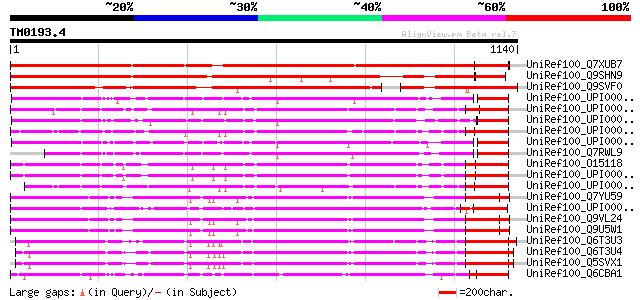
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q7XUB7 OSJNBb0078D11.11 protein [Oryza sativa] 1357 0.0
UniRef100_Q9SHN9 F7F22.1 [Arabidopsis thaliana] 1326 0.0
UniRef100_Q9SVF0 Hypothetical protein F22I13.120 [Arabidopsis th... 979 0.0
UniRef100_UPI000021AE27 UPI000021AE27 UniRef100 entry 602 e-170
UniRef100_UPI00003ABE91 UPI00003ABE91 UniRef100 entry 579 e-163
UniRef100_UPI00002344B9 UPI00002344B9 UniRef100 entry 573 e-162
UniRef100_UPI00003635C8 UPI00003635C8 UniRef100 entry 565 e-159
UniRef100_UPI000023DFE9 UPI000023DFE9 UniRef100 entry 560 e-158
UniRef100_Q7RWL9 Hypothetical protein [Neurospora crassa] 558 e-157
UniRef100_O15118 Niemann-Pick C1 protein precursor [Homo sapiens] 555 e-156
UniRef100_UPI000036A3B7 UPI000036A3B7 UniRef100 entry 551 e-155
UniRef100_UPI0000318C66 UPI0000318C66 UniRef100 entry 550 e-155
UniRef100_Q7YU59 RE56428p [Drosophila melanogaster] 522 e-146
UniRef100_UPI000042D349 UPI000042D349 UniRef100 entry 521 e-146
UniRef100_Q9VL24 CG5722-PA [Drosophila melanogaster] 521 e-146
UniRef100_Q9U5W1 NPC1 protein [Drosophila melanogaster] 521 e-146
UniRef100_Q6T3U3 Niemann-Pick C1-like 1 [Rattus norvegicus] 514 e-144
UniRef100_Q6T3U4 Niemann-Pick C1-like 1 [Mus musculus] 509 e-142
UniRef100_Q5SVX1 Ortholog of human NPC1 (Niemann-Pick disease, t... 509 e-142
UniRef100_Q6CBA1 Similar to tr|Q12200 Saccharomyces cerevisiae Y... 501 e-140
>UniRef100_Q7XUB7 OSJNBb0078D11.11 protein [Oryza sativa]
Length = 1361
Score = 1357 bits (3512), Expect = 0.0
Identities = 684/1125 (60%), Positives = 851/1125 (74%), Gaps = 42/1125 (3%)
Query: 1 MYDICGTRSDGKVLNCPFGSPAVKPDDLLSSKIQSMCPTITGNVCCTKAQFDTLQTQVQQ 60
MY IC RSDGKVLNC + AVKPD L S++IQS+CPTITG+VCCT QFDTL QVQQ
Sbjct: 55 MYGICAQRSDGKVLNCVNATKAVKPDTLFSARIQSLCPTITGDVCCTVDQFDTLHQQVQQ 114
Query: 61 AIPFLVGCPACLRNFLNLFCELTCSPNQSLFINVTSVDKAGGNSTVGGIDYFVSDAFGEG 120
AIPFLVGCPACLRNFLNLFCE++CSPNQSLFINVTSV + TV GIDY+V+ +GE
Sbjct: 115 AIPFLVGCPACLRNFLNLFCEMSCSPNQSLFINVTSVKQVNNTMTVNGIDYYVTSTYGEE 174
Query: 121 LYESCKDVKFGSMNSRAIQFIGAGAQNFKEWFAFIGRKAAPNSPGSPYAIMFRPNATKSS 180
LY SCKDVKFG++N+RA+ F+G GA+N+KEW AFIGR+A N GSPY I F + + S+
Sbjct: 175 LYNSCKDVKFGTLNTRAMDFLGGGAKNYKEWMAFIGRQADLNQIGSPYLITFPSDISGST 234
Query: 181 GMKPMNVSAYSCSDTSLGCSCGDCPSSSVCSNSASTTINKANSCSIKVGSLTVKCVDFIL 240
+KP+N + YSC D SLGCSCGDCPSSSVC+ S +N SCS+K+GSL KC+DF L
Sbjct: 235 AVKPLNATIYSCGDPSLGCSCGDCPSSSVCTGSLLPQLNTETSCSVKMGSLKAKCLDFSL 294
Query: 241 AVLYIILICVFLGWALYHRIRERKMTYRTEPVSNVISGGVLYARNQEKDENLPMQMIEDV 300
V+Y++L+C+FL A HR R + T+P+ N + +++ N K + Q+ E
Sbjct: 295 VVVYLVLLCIFLFGAFLHRTRRSGIFSHTKPLKN--AEDKIHSSNNGKVPDSSAQVSEAA 352
Query: 301 PQNRNGVRLSVVQGYMSNFYRKYGSLVARHPINVLALTLAIVLLLCLGLIRFKVETRPEK 360
SV+Q YMS F+RK+G+ VA+HP+ VL ++L + LLC+GLIRFKVE RPEK
Sbjct: 353 SAPVQSAHPSVIQTYMSTFFRKHGTFVAKHPLLVLFVSLLVPTLLCIGLIRFKVEIRPEK 412
Query: 361 LWVGPGSKAAQEKQFFDSHLAPFYRIEQLILATVPDHMNSTSPRIVSADNIRFLFEVQKK 420
LWV GS+AA EKQ+FDSHLAPFYRIEQL+LAT S +P IV+ +N++ LF++QKK
Sbjct: 413 LWVSSGSRAADEKQYFDSHLAPFYRIEQLVLAT-SAFGGSEAPTIVNDNNMKLLFQIQKK 471
Query: 421 VDAIRANYSGLMVSLQDICMKPLDKDCATQSVLQYFKMDPRNFDDSGAVEHLNYCFQQYS 480
+D +RANYSG VSL DIC+KPL +CATQSVLQ+ Y+
Sbjct: 472 IDDLRANYSGSTVSLADICLKPLGTECATQSVLQH-----------------------YT 508
Query: 481 SADQCMSAFKAPLDPSTVLGGFSGKDYSGASAFIVTYPVNNAIDEEGNETAKAVAWEKAF 540
+ + C+S F++P+DPST+LGGF G +++ ASAF++TYPVNN ++ G E KAVAWE+A+
Sbjct: 509 TEETCLSTFQSPIDPSTILGGFPGNNFTEASAFVITYPVNNKVETTGQENGKAVAWERAY 568
Query: 541 IQLVKDELLPMAQSRNLTLAFSSESSIEEELKRESTADAITILVSYLVMFAYISLTLGDT 600
+ LVK+E+LPM + NLT++FSSESSI++EL RESTADAITI++SY+VMFAYIS TLGD
Sbjct: 569 VNLVKEEILPMVLAHNLTMSFSSESSIQDELNRESTADAITIVISYIVMFAYISFTLGDR 628
Query: 601 P-HPSSFYISSKVLLGLSGVILVMLSVLGSVAIFSALGVKSTLIIMEVIPFLVLAVGVDN 659
P H S ++SSKVLLGLSGV+LVMLSVLGS+ FSA+GVKSTLIIMEVIPFLVLAVGVDN
Sbjct: 629 PSHLLSLFVSSKVLLGLSGVVLVMLSVLGSMGFFSAIGVKSTLIIMEVIPFLVLAVGVDN 688
Query: 660 MCILVHAVKRQPLELPLEGRISNALVEVGPSITLASLSEVLAFAVGSFISMPACRVFSMF 719
MCILVHAVKRQP L LE RIS ALVEVGPSITLASL+EVLAFAV + MPA RVFSMF
Sbjct: 689 MCILVHAVKRQPDGLDLEERISTALVEVGPSITLASLAEVLAFAVSAINPMPATRVFSMF 748
Query: 720 AALAVLLDFVLQVTAFVALIVLDSQRAEDKRVDCFPCIKVHSFHADPDKGIRQRKPGLLA 779
AALAVLLDF+LQV+AFVALIVLD +RA+D R+DC PC +V S D G Q P LLA
Sbjct: 749 AALAVLLDFLLQVSAFVALIVLDFRRAQDGRIDCMPCARVKSSVVASDGGNHQGLP-LLA 807
Query: 780 RYMKEVHAPILSIWGVKIVVIAIFVAFALASIALSTRIEPGLEQEIVLPRDSYLQGYFNN 839
RYMK VHAPIL VK VVIA+FV F+ ASIALSTR++PGLEQ+IVLPRDSYLQ YF++
Sbjct: 808 RYMKNVHAPILGYRAVKFVVIAVFVGFSFASIALSTRLQPGLEQKIVLPRDSYLQDYFDD 867
Query: 840 VSEYLRIGPPLYFVVKNYNYSSESTHTNQLCSISQCNSDSLLNEISKASLVPETSYIAKP 899
++ Y+++GPPLYFV+KN+NYSS S HTN++CSI+QC+S+SLLNEI+K SL PETSYIAKP
Sbjct: 868 LATYMKVGPPLYFVIKNFNYSSASEHTNKICSINQCDSNSLLNEIAKQSLSPETSYIAKP 927
Query: 900 AASWLDDFLVWISPEAFGCCRKFTNGSYCPPDDQPPCCAPEDDS---CVSGACKDCTTCF 956
AASWLDDFL+W+SPEAFGCCRKF NGSYCPPDDQPPCC + DS SGAC +CTTCF
Sbjct: 928 AASWLDDFLIWMSPEAFGCCRKFVNGSYCPPDDQPPCCQHDQDSSSCSASGACNNCTTCF 987
Query: 957 RHSDLRNDRTSTMQFRDKLPWFLSALPSADCAKGGHGAYTSSVDLKGYDSGIIQASSFRT 1016
SDL N R ST QF++KLPWFL ALPS+DC+KGG GAY++S+DL GY++GIIQAS+FRT
Sbjct: 988 LRSDLHNGRPSTTQFKEKLPWFLDALPSSDCSKGGKGAYSTSLDLNGYENGIIQASAFRT 1047
Query: 1017 YHTPLNKQVDYVNSMRAAREFSSKVSDSLKVASGDKDQRVKEALGTMGASVFSGITLTKL 1076
YHTPLNKQ DYVNSM+AAR+FSSK+S L++ Q ++ + + G+ T +
Sbjct: 1048 YHTPLNKQSDYVNSMKAARDFSSKMSKELQM------QMFPYSVFYIFFEQYLGVWKTAI 1101
Query: 1077 VGVIVLYFSRTEVFVIYYFQMYLSLVLLGFLHGLVFLPVVLSIFG 1121
+ + V + V + ++ S+++L +V +VL + G
Sbjct: 1102 MNICVCLGTVFVVCFVVTSSLWASIIIL-----IVLAMIVLDLMG 1141
Score = 110 bits (274), Expect = 3e-22
Identities = 50/80 (62%), Positives = 68/80 (84%)
Query: 1045 LKVASGDKDQRVKEALGTMGASVFSGITLTKLVGVIVLYFSRTEVFVIYYFQMYLSLVLL 1104
+++ G+++ R ++AL TMGASVFSGITLTKLVGVIVL F+++EVFV+YYFQMYL+LV++
Sbjct: 1194 MQIGIGNRESRARQALSTMGASVFSGITLTKLVGVIVLRFAKSEVFVVYYFQMYLALVII 1253
Query: 1105 GFLHGLVFLPVVLSIFGPPS 1124
GFLHGL+FLP ++ PS
Sbjct: 1254 GFLHGLIFLPCYIATSSHPS 1273
>UniRef100_Q9SHN9 F7F22.1 [Arabidopsis thaliana]
Length = 1275
Score = 1326 bits (3431), Expect = 0.0
Identities = 697/1136 (61%), Positives = 813/1136 (71%), Gaps = 171/1136 (15%)
Query: 1 MYDICGTRSDGKVLNCPFGSPAVKPDDLLSSKIQSMCPTITGNVCCTKAQFDTLQTQVQQ 60
MYDICG RSDGKVLNCPF P+VKPDDLLSSKIQS+CPTITGNVCCT+ QFDTL++QVQQ
Sbjct: 21 MYDICGARSDGKVLNCPFNIPSVKPDDLLSSKIQSLCPTITGNVCCTETQFDTLRSQVQQ 80
Query: 61 AIPFLVGCPACLRNFLNLFCELTCSPNQSLFINVTSVDKAGGNSTVGGIDYFVSDAFGEG 120
AIPF+VGCPACLRNFLNLFCELTCSP+QSLFINVTS K NSTV GI Y+++D FG G
Sbjct: 81 AIPFIVGCPACLRNFLNLFCELTCSPDQSLFINVTSTTKVKNNSTVDGIQYYITDDFGAG 140
Query: 121 LYESCKDVKFGSMNSRAIQFIGAGAQNFKEWFAFIGRKAAPNSPGSPYAIMFRPNATKSS 180
+YESCK+VKFGS NSRA+ F+GAGA+NFKEWF FIG+KA N PGSPY I F P SS
Sbjct: 141 MYESCKNVKFGSSNSRALDFLGAGAKNFKEWFTFIGQKAGVNLPGSPYGIAFLPTPPVSS 200
Query: 181 GMKPMNVSAYSCSDTSLGCSCGDCPSSSVCSNSASTTINKANSCSIKVGSLTVKCVDFIL 240
GM+PMNVS YSC D SLGCSCGDCPS++ CS+ A K +SCSIK+GSL VKCVDFIL
Sbjct: 201 GMRPMNVSIYSCGDESLGCSCGDCPSAATCSSKAEVPTQKKHSCSIKIGSLEVKCVDFIL 260
Query: 241 AVLYIILICVFLGWALYHRIRERKMTYRTEPVSNVISGGVLYARNQEKDENLPMQMIEDV 300
A+LYI+L+ +FLG L H +R +K T + +S + G + NQ+K + + QM+++
Sbjct: 261 AILYIVLVSLFLGGGLLHPVRGKKKTSQMGTLSE--ASGERNSVNQQKPDTIQSQMLQNT 318
Query: 301 PQNRNGVRLSVVQGYMSNFYRKYGSLVARHPINVLALTLAIVLLLCLGLIRFKVETRPEK 360
PQ RN +LS VQG+++NFY KYG VARHP VL L++++VLLLC+GLIRFKVETRP+K
Sbjct: 319 PQ-RNWGQLSTVQGHLANFYGKYGIWVARHPTLVLCLSVSVVLLLCVGLIRFKVETRPDK 377
Query: 361 LWVGPGSKAAQEKQFFDSHLAPFYRIEQLILATVPDHMNSTSPRIVSADNIRFLFEVQKK 420
LWVG GS+AA+EKQFFD+HLAPFYRIEQLI+ATV + +P I++ DNI+ LF++QKK
Sbjct: 378 LWVGSGSRAAEEKQFFDTHLAPFYRIEQLIIATVQTSSHEKAPEILTDDNIKLLFDIQKK 437
Query: 421 VDAIRANYSG--LMVSLQDICMKPLDKDCATQSVLQYFKMDPRNFDDSGAVEHLNYCFQQ 478
V + +N S V + C K + YFKM P N+DD G V+H+ YCF+
Sbjct: 438 VSQLFSNPSNHPYNVFMYRTCKKLFN---------MYFKMKPENYDDYGGVDHVKYCFEH 488
Query: 479 YSSADQCMSAFKAPLDPSTVLGGFSGKDYSGASAFIVTYPVNNAIDEEGNETAKAVAWEK 538
++S + C+SAFK PLDP+T LGGFSG +S ASAF+VTYPV+N +D +GN+T KAVAWEK
Sbjct: 489 FTSTESCLSAFKGPLDPTTALGGFSGNSFSEASAFLVTYPVDNFVDNKGNKTEKAVAWEK 548
Query: 539 AFIQLVKDELLPMAQSRNLTLAFSSESSIEEELKRESTADAITILV-------------- 584
AFIQL KDELLPM Q++NLTL+FSSESSIEEELKRESTAD ITI V
Sbjct: 549 AFIQLAKDELLPMVQAKNLTLSFSSESSIEEELKRESTADVITIAVICFSFILYWVSNMS 608
Query: 585 -------------SYLVMFAYISLTLGDTPHPSSFYISSKVLLGLSGVILVMLSVLGSVA 631
SYLVMFAYISLTLGD+P SFYI+SKVLLGLSGV+LVMLSVLGSV
Sbjct: 609 FMSSISHVSLLQISYLVMFAYISLTLGDSPRLKSFYITSKVLLGLSGVLLVMLSVLGSVG 668
Query: 632 IFSALGVKSTLIIMEVIPFLVLAV------------------------------------ 655
FSA+G+KSTLIIMEVIPFLVLAV
Sbjct: 669 FFSAVGMKSTLIIMEVIPFLVLAVIVSISNIACNFNMLADVVATFFILFLIFFYFYLEYF 728
Query: 656 ----GVDNMCILVHAVKRQPLELPLEGRISNALVEVGPSITLASLSEVLAFAVGSFISMP 711
GVDNMCILVHAVKRQ ELPLE RISNAL+EVGPSITLASL+E+LAFAVG+FI MP
Sbjct: 729 YRQVGVDNMCILVHAVKRQEQELPLERRISNALMEVGPSITLASLAEILAFAVGAFIKMP 788
Query: 712 ACRVFSMFA--------------------ALAVLLDFVLQVTAFVALIVLDSQRAEDKRV 751
A RVFSMFA ALAVLLDF+LQ+TAFVALIV D +R EDKRV
Sbjct: 789 AVRVFSMFAVYSLNAFIIYFLTSICIMLAALAVLLDFLLQITAFVALIVFDFRRTEDKRV 848
Query: 752 DCFPCIKVHSFHADPDKGIRQRKPGLLARYMKEVHAPILSIWGVKIVVIAIFVAFALASI 811
DCFPCIK +KG+ QRK GLL RYMKEVHAP+LS W VKIVVIA F A+A I
Sbjct: 849 DCFPCIKTSKSSISAEKGVGQRKAGLLTRYMKEVHAPVLSHWIVKIVVIAFFFGLAMAGI 908
Query: 812 ALSTRIEPGLEQEIVLPRDSYLQGYFNNVSEYLRIGPPLYFVVKNYNYSSESTHTNQLCS 871
ALSTRIEPGLEQ+IVLP+DSYLQ
Sbjct: 909 ALSTRIEPGLEQQIVLPQDSYLQ------------------------------------- 931
Query: 872 ISQCNSDSLLNEISKASLVPETSYIAKPAASWLDDFLVWISPEAFGCCRKFTNGSYCPPD 931
I++ASL PE SYIAKPAASWLDDFLVW+SPEAFGCCRKFTNG++CPPD
Sbjct: 932 ------------IARASLTPELSYIAKPAASWLDDFLVWLSPEAFGCCRKFTNGTFCPPD 979
Query: 932 DQPPCCAPEDDSCVSGACKDCTTCFRHSDLRNDRTSTMQFRDKLPWFLSALPSADCAKGG 991
DQ CFRH+DL +DR ST QF++KLPWFL+ALPSADCAKGG
Sbjct: 980 DQ---------------------CFRHADLSSDRPSTTQFKEKLPWFLNALPSADCAKGG 1018
Query: 992 HGAYTSSVDLKGYDSGIIQASSFRTYHTPLNKQVDYVNSMRAAREFSSKVSDSLKV 1047
HGAY+SSVDL+GY +GIIQASSFRTYHTPLNKQVD+VNSMRAA+EFS+KVS SLK+
Sbjct: 1019 HGAYSSSVDLQGYANGIIQASSFRTYHTPLNKQVDFVNSMRAAQEFSAKVSRSLKM 1074
Score = 115 bits (289), Expect = 6e-24
Identities = 56/70 (80%), Positives = 67/70 (95%)
Query: 1046 KVASGDKDQRVKEALGTMGASVFSGITLTKLVGVIVLYFSRTEVFVIYYFQMYLSLVLLG 1105
++++GD++ R+KEALG MGASVFSGITLTKLVGVIVL FSR+EVFV+YYF+MYL+LVLLG
Sbjct: 1163 QISTGDRNHRMKEALGGMGASVFSGITLTKLVGVIVLGFSRSEVFVVYYFKMYLALVLLG 1222
Query: 1106 FLHGLVFLPV 1115
FLHGLVFLPV
Sbjct: 1223 FLHGLVFLPV 1232
>UniRef100_Q9SVF0 Hypothetical protein F22I13.120 [Arabidopsis thaliana]
Length = 1055
Score = 979 bits (2531), Expect = 0.0
Identities = 512/860 (59%), Positives = 627/860 (72%), Gaps = 85/860 (9%)
Query: 1 MYDICGTRSDGKVLNCPFGSPAVKPDDLLSSKIQSMCPTITGNVCCTKAQFDTLQTQVQQ 60
MYDICG RSDGKVLNCP+ SP+++PD+L S+KIQS+CPTI+GNVCCT+ QFDTL++QVQQ
Sbjct: 1 MYDICGHRSDGKVLNCPYASPSIQPDELFSAKIQSLCPTISGNVCCTETQFDTLRSQVQQ 60
Query: 61 AIPFLVGCPACLRNFLNLFCELTCSPNQSLFINVTSVDKAGGNSTVGGIDYFVSDAFGEG 120
A+PFLVGCPACLRNFLNLFCEL+CSPNQSLFINVTSV + GN TV GIDY ++D FGEG
Sbjct: 61 AVPFLVGCPACLRNFLNLFCELSCSPNQSLFINVTSVAEVSGNLTVDGIDYHITDTFGEG 120
Query: 121 LYESCKDVKFGSMNSRAIQFIGAGAQNFKEWFAFIGRKAAPNSPGSPYAIMFRPNATKSS 180
LYESCK+VKFG+MN+RAI F+G GA+NF+EWF FIG+KA PGSPYAI F+ + +SS
Sbjct: 121 LYESCKEVKFGTMNTRAINFVGGGAKNFREWFTFIGQKAPSGFPGSPYAINFKSSIPESS 180
Query: 181 GMKPMNVSAYSCSDTSLGCSCGDCPSSSVCSNSASTTINKANSCSIKVGSLTVKCVDFIL 240
M PMNVS YSC+ CS+ + +SCSI++G L V+C++ +
Sbjct: 181 AMVPMNVSVYSCA----------------CSSPEPLPPHDEDSCSIRIGPLKVRCIELSM 224
Query: 241 AVLYIILICVFLGWALYHRIRERKMTYRTEPVSNVISGGVLYARNQEKDENLPMQMIEDV 300
A++Y++L+ F GWA +R R T+P+ + S +L+ ++ + + I V
Sbjct: 225 ALVYVLLVSCFFGWAGLNRRRNT-----TQPLDS--SKPLLHPVEEDGINSEMKENILGV 277
Query: 301 PQNRNGVRLSVVQGYMSNFYRKYGSLVARHPINVLALTLAIVLLLCLGLIRFKVETRPEK 360
R+ +LS VQ YM+ FYR YGS +AR+P VL +++AIVL LC GL FKVETRPEK
Sbjct: 278 KVQRHA-QLSPVQRYMAKFYRSYGSWIARNPSLVLFMSVAIVLALCSGLYNFKVETRPEK 336
Query: 361 LWVGPGSKAAQEKQFFDSHLAPFYRIEQLILATVPDHMNSTSPRIVSADNIRFLFEVQKK 420
LWVGP SKAA+EK+FFD+HL+PFYRIEQLILATVPD + +P IV+ +NI LF++Q+K
Sbjct: 337 LWVGPESKAAEEKKFFDTHLSPFYRIEQLILATVPDPKSGRAPSIVTDENILLLFDIQQK 396
Query: 421 VDAIRANYSGLMVSLQDICMKPLDKDCATQSVLQYFKMDPRNFDDSGAVEHLNYCFQQYS 480
YFKMD FDD G VEH YCFQ Y+
Sbjct: 397 ----------------------------------YFKMDSGTFDDYGGVEHAEYCFQHYT 422
Query: 481 SADQCMSAFKAPLDPSTVLGGFSGKDYSG------------------------ASAFIVT 516
S++ C+SAF+AP+DPS VLGGFSG +YS A+AF+VT
Sbjct: 423 SSETCLSAFQAPVDPSAVLGGFSGNNYSEVMVSELGCSVPFDCYSDVKRTLFQATAFVVT 482
Query: 517 YPVNNAIDEEGNETAKAVAWEKAFIQLVKDELLPMAQSRNLTLAFSSESSIEEELKREST 576
YPVNN I + NE A+AVAWEK+FIQL K+ELLPM +S+NL+L+FSSESSIEEELKREST
Sbjct: 483 YPVNNVIGDSSNENARAVAWEKSFIQLAKEELLPMVRSKNLSLSFSSESSIEEELKREST 542
Query: 577 ADAITILVSYLVMFAYISLTLGDTPHPSSFYISSKVLLGLSGVILVMLSVLGSVAIFSAL 636
AD ITI SYLVMF YIS+TLGD P +FYISSKVLLGLSGV+LV+LSVLGSV +FSAL
Sbjct: 543 ADVITIAASYLVMFVYISVTLGDAPQFYTFYISSKVLLGLSGVVLVLLSVLGSVGVFSAL 602
Query: 637 GVKSTLIIMEVIPFLVLAVGVDNMCILVHAVKRQPLELPLEGRISNALVEVGPSITLASL 696
GVKSTLIIMEVIPFLVLAVGVDNMCILVHAVKRQP E+ LE RIS+ALVEVGPSITLASL
Sbjct: 603 GVKSTLIIMEVIPFLVLAVGVDNMCILVHAVKRQPREVSLEQRISSALVEVGPSITLASL 662
Query: 697 SEVLAFAVGSFISMPACRVFSMFAALAVLLDFVLQVTAFVALIVLDSQRAEDKRVDCFPC 756
SEVLAFAVG+F+ MPACR+FSMFAALA++LDF LQ+TAFVALIV D +R+ D R+DCFPC
Sbjct: 663 SEVLAFAVGAFVPMPACRIFSMFAALAIMLDFFLQITAFVALIVFDCKRSADNRIDCFPC 722
Query: 757 IKVHSFHADPDKGIRQRKPGLLARYMKEVHAPILSIWGVKIVVIAIFVAFALASIALSTR 816
IKV S + +G R+PG L RYMKEVHAP+L +WGVK+VV+A+F AFALASI +S
Sbjct: 723 IKVPSSSRESVEG--GREPGFLERYMKEVHAPVLGLWGVKMVVVAVFFAFALASI-ISRA 779
Query: 817 IEPGLEQEIVLPRDSYLQGY 836
+ I P S+L +
Sbjct: 780 SQASDTSYIAKPAASWLDDF 799
Score = 334 bits (856), Expect = 1e-89
Identities = 175/303 (57%), Positives = 208/303 (67%), Gaps = 62/303 (20%)
Query: 879 SLLNEISKASLVPETSYIAKPAASWLDDFLVWISPEAFGCCRKFTNGSYCPPDDQPPCCA 938
+L + IS+AS +TSYIAKPAASWLDDFLVW+SPEAFGCCRKFTNGSYCPPDDQ
Sbjct: 771 ALASIISRASQASDTSYIAKPAASWLDDFLVWLSPEAFGCCRKFTNGSYCPPDDQ----- 825
Query: 939 PEDDSCVSGACKDCTTCFRHSDLRNDRTSTMQFRDKLPWFLSALPSADCAKGGHGAYTSS 998
CFRHSDL DR ST QFR+KLPWFL+ALPSADCAKGGHGAYT+S
Sbjct: 826 ----------------CFRHSDLVQDRPSTAQFREKLPWFLNALPSADCAKGGHGAYTNS 869
Query: 999 VDLKGYDSGIIQASSFRTYHTPLNKQVD-------------YVN---------------- 1029
VDLKGY+SG+IQAS FRTYHTPLN Q+D Y+N
Sbjct: 870 VDLKGYESGVIQASEFRTYHTPLNTQIDIFPYSVFYIFFEQYLNIWTVALTNLAIAIVGI 929
Query: 1030 ------------SMRAAREFSSKVSDSLKVASGDKDQRVKEALGTMGASVFSGITLTKLV 1077
S+ A EF +S + ++SGD++ R +EAL TMGASVFSGITLTKLV
Sbjct: 930 QLNAVSVVNLIMSIGIAVEFCVHISHAFLMSSGDREHRAREALETMGASVFSGITLTKLV 989
Query: 1078 GVIVLYFSRTEVFVIYYFQMYLSLVLLGFLHGLVFLPVVLSIFGPPSRCTIIEQEENRSS 1137
GVIVL F+R+E+FV+YYFQMYL+LV++GFLHGLVFLPV+LS+ GPP IEQ++ +
Sbjct: 990 GVIVLCFARSEIFVVYYFQMYLALVIIGFLHGLVFLPVILSLAGPPQLNLDIEQQQTDEA 1049
Query: 1138 TSS 1140
+SS
Sbjct: 1050 SSS 1052
>UniRef100_UPI000021AE27 UPI000021AE27 UniRef100 entry
Length = 1275
Score = 602 bits (1551), Expect = e-170
Identities = 382/1076 (35%), Positives = 585/1076 (53%), Gaps = 104/1076 (9%)
Query: 5 CGTRS-DGKVLNCPFGSPAVKPDDLLSSKIQSMCPTI--TGNVCCTKAQFDTLQTQVQQA 61
CG +S GK L C PA PD+ + +C TG VCC K+Q ++L++++
Sbjct: 40 CGKKSWFGKELPCVDNGPAENPDEDFRKLLVDLCGPKWETGPVCCDKSQVESLKSELSTP 99
Query: 62 IPFLVGCPACLRNFLNLFCELTCSPNQSLFINVTSVDKAGGNSTVGGIDYFVSDAFGEGL 121
+ CPAC NF NLFC TCSP+QSLF+NVT + G + +D +S +G G
Sbjct: 100 RQIVSSCPACKDNFYNLFCTFTCSPDQSLFVNVTKAQEKNGKLQITELDQLISSEYGTGF 159
Query: 122 YESCKDVKFGSMNSRAIQFIGAGAQNFKEWFAFIGRKAAPNSPGSPYAIMFRPNATKSSG 181
Y+SCKDVKFG NS+A+ FIG GA+N+ + F+G + A GSP+ I F P
Sbjct: 160 YDSCKDVKFGPSNSKAMDFIGGGAKNYTQLLKFLGDEKA---IGSPFQINF-PTEYSEPA 215
Query: 182 MKPMNVSAYSCS--DTSLGCSCGDCPSSSVCSNSASTTINKANSCSIKVGSLTVKCVDFI 239
M P + C+ D + C+C DCP VC + + ++ SC VGSL C+ F
Sbjct: 216 MSPREMKPKRCNDDDPNFRCACVDCP--QVCPELPA--VKESGSC--HVGSL--PCLSFA 267
Query: 240 L-----AVLYIILICVFLGWAL--YHRIRERKMTYRTEPVSNVISGGVLYARNQEKDEN- 291
+L+ + + + G+ L ++ R ++ +PV R+ ++DE
Sbjct: 268 AIFTYSIILFSLAVALTGGFVLKKHNERRRERLRLLQDPV-----------RSDDEDEGD 316
Query: 292 -LPMQMIEDVPQNRNGVRLSVVQGYMSNFYRKYGSLVARHPINVLALTLAIVLLLCLGLI 350
+ + D PQ+ V + + + K G AR P + +L IV +L LGL
Sbjct: 317 LVHNNAMLDRPQSN-----YPVNSWCDSAFSKLGHTAARFPGITIISSLIIVAVLSLGLF 371
Query: 351 RFKVETRPEKLWVGPGSKAAQEKQFFDSHLAPFYRIEQLILATVPDHMNSTSPRIVSADN 410
RF +E P +LWV P S AAQEK FFD++ PFYR E++ L V D S+ ++S DN
Sbjct: 372 RFDIEKDPARLWVSPTSAAAQEKAFFDANFGPFYRAEKIFL--VNDTNPSSPGPVLSYDN 429
Query: 411 IRFLFEVQKKVDAIRANYSGLMVSLQDICMKPLDKDCATQSVLQYFKMDPRNFDDSGAVE 470
+ + +V+ V ++ G M LQD+C+KP C QSV YF D N G
Sbjct: 430 LIWWIDVENSVKQLKGPRFGAM--LQDVCLKPTGSACVVQSVAAYFGNDADNVSKGGWKG 487
Query: 471 HLNYCFQQYSSADQCMSAFKAPLDPSTVLGGF-SGKDYSGASAFIVTYPVNNAIDEEGNE 529
+L C + S +C F P+DP +LGG+ +G D + A A VT+ +NN E +E
Sbjct: 488 NLRDCAR---SPVECRPDFGQPIDPGMILGGYGAGDDIADAQAMTVTWVLNN-FPEGTSE 543
Query: 530 TAKAVAWEKAFIQLVKDELLPM---AQSRNLTLAFSSESSIEEELKRESTADAITILVSY 586
A+A+ +E+A +K+ LL + A R L L+FS+E S+E+EL + + DA I++SY
Sbjct: 544 EARAMDFEEA----LKNRLLKLQEEAADRGLRLSFSTEISLEQELNKSTNTDAKIIVISY 599
Query: 587 LVMFAYISLTLGDTP--------HPSSFYISSKVLLGLSGVILVMLSVLGSVAIFSALGV 638
+VMF Y S+ LG T + S F++ SK LG+ G+ +V+LS++ S+ +FS G+
Sbjct: 600 IVMFLYASIALGSTTLNFREFFRNKSLFFVQSKFGLGIVGIAIVLLSIMASIGLFSWFGL 659
Query: 639 KSTLIIMEVIPFLVLAVGVDNMCILVHAVKRQPL---ELPLEGRISNALVEVGPSITLAS 695
K TLII++VIPF+VLAVGVDN+ ++VH +R + +L +E RI+ AL +GPSI ++
Sbjct: 660 KVTLIIVDVIPFIVLAVGVDNIFLIVHEFERVNISHPDLDVELRIAKALGRMGPSILFSA 719
Query: 696 LSEVLAFAVGSFISMPACRVFSMFAALAVLLDFVLQVTAFVALIVLDSQRAEDKRVDCFP 755
++E +FA+G+F+ MPA R F+++AA AV ++ +LQVT FV+ + L+ QR ED R+D FP
Sbjct: 720 VTETASFALGAFVGMPAVRNFAIYAAGAVFINALLQVTMFVSFLTLNQQRVEDCRMDLFP 779
Query: 756 CIKVHS--FHADPDKGIRQR-----KPGLLARYMKEVHAPILSIWGVKIVVIAIFVAFAL 808
C+++ S H + + R + +L +++++ +AP L VK VV+ +F+
Sbjct: 780 CVQLKSARIHLNGTGNLGPRYHEAPQESMLQQFIRKYYAPALLGKKVKAVVVLVFLGVFT 839
Query: 809 ASIALSTRIEPGLEQEIVLPRDSYLQGYFNNVSEYLRIGPPLYFVVKNYNYSSESTHTNQ 868
A ++L +E GL+Q + +P DSYL YFN++ Y GPP+YFV K N+ ++ H +
Sbjct: 840 AGVSLIPEVELGLDQRVAIPDDSYLIPYFNDLYAYFESGPPVYFVTKESNF-TQREHQQE 898
Query: 869 LCS-ISQCNSDSLLNEISKASLVPETSYIAKPAASWLDDFLVWISPE-AFGCCRKFTNGS 926
+C+ + CN S+ N + + PE SYIA P ASW+DDF +W+ P+ CC NG
Sbjct: 899 VCARFTTCNELSMTNILEQERKRPEISYIASPTASWIDDFFLWLDPDLGESCC--VENGK 956
Query: 927 YCPPDDQPPCCAPEDDSCVSGACKDCTTCFRHSDLRNDRTSTMQFRDKLPWFLSALPSAD 986
C D PP +SG K +F L F+ + + +
Sbjct: 957 ACFADRNPPW-----SITMSGMPKG-----------------QEFVHYLDKFIQSPTTEE 994
Query: 987 CAKGGHGAYTSSVDLKGYDSGIIQASSFRTYHTPLNKQVDYVNSMRAAREFSSKVS 1042
C GG AY +V + ++ I AS FRT HTPL Q D++ + +AR ++ +S
Sbjct: 995 CPLGGQAAYGDAVVI-DHEKTTIGASHFRTMHTPLRSQSDFIKAYASARRIANDIS 1049
Score = 80.5 bits (197), Expect = 3e-13
Identities = 37/70 (52%), Positives = 53/70 (74%)
Query: 1052 KDQRVKEALGTMGASVFSGITLTKLVGVIVLYFSRTEVFVIYYFQMYLSLVLLGFLHGLV 1111
+D R AL +G SVFSGIT+TKL+GV VL F+R+++F IYYF+++++LV+ H L+
Sbjct: 1170 RDARTWTALANVGGSVFSGITVTKLLGVTVLAFTRSKIFEIYYFRVWVALVIFAATHALI 1229
Query: 1112 FLPVVLSIFG 1121
FLPV LS+ G
Sbjct: 1230 FLPVALSLLG 1239
>UniRef100_UPI00003ABE91 UPI00003ABE91 UniRef100 entry
Length = 1283
Score = 579 bits (1493), Expect = e-163
Identities = 381/1106 (34%), Positives = 582/1106 (52%), Gaps = 106/1106 (9%)
Query: 2 YDICGTRSDGKVLNCPFGSPAVKPDDLLSSKIQSMCPTIT-GNV--CCTKAQFDTLQTQV 58
Y CG S K NC + P + +Q +CP + GNV CC Q TL+ +
Sbjct: 33 YGECGVASGDKRYNCAYDGPPIALPKGGYDLMQELCPGLFFGNVSTCCDILQLQTLKNNL 92
Query: 59 QQAIPFLVGCPACLRNFLNLFCELTCSPNQSLFINVTS-------VDKAGGNSTVGGIDY 111
Q + FL CP+C N +NLFCELTCSPNQS F+NVTS V K S++ + Y
Sbjct: 93 QLPLQFLSRCPSCFYNLINLFCELTCSPNQSEFLNVTSTIPYYDPVSKEN-KSSITELQY 151
Query: 112 FVSDAFGEGLYESCKDVKFGSMNSRAIQFI-GAGAQ--NFKEWFAFIGRKAAPNSPGSPY 168
F+ + F +Y +CKDV+ S N +A+ + G + N W ++ K ++ +P+
Sbjct: 152 FIGERFANEMYNACKDVEAPSSNVKALGLLCGKDVKDCNATNWIEYMFNK---DNGQTPF 208
Query: 169 AIMFRPNATKSSGMKPMNVSAYSCS---DTSLG-CSCGDCPSSSVCSNSASTTINKANSC 224
+I+ + GM PMN + C+ D S G CSC DC S VC A
Sbjct: 209 SIIPIFSDAPVHGMNPMNNATKGCNESVDDSTGPCSCQDC--SIVCGPKPQPPPLPAPWL 266
Query: 225 SIKVGSLTVKCVDFILAVLYIILICVFLGWALYHRIRERKMTYRTEPVSNVISGGVLYAR 284
+ ++ + + L I VF W R R P+ + V ++
Sbjct: 267 LFGLDAVYIIMWISYMGFLLIFFALVFGVWCY----RRRHFVSEYTPIDS----NVAFSA 318
Query: 285 NQEKDENLPMQMIEDVPQN-RNGVRLSVVQGYMSNFYRKYGSLVARHPINVLALTLAIVL 343
N +D N + E + + NG+R++ + +G+ R+P V+ ++ +
Sbjct: 319 NSHRD-NGKITCGERLGERFENGLRMT---------FTSWGAFCVRNPRPVILFSVVFIA 368
Query: 344 LLCLGLIRFKVETRPEKLWVGPGSKAAQEKQFFDSHLAPFYRIEQLILATVPDHMNSTSP 403
+ C G + K T P LW P S+A +EK++FD H PF+R EQLI+ H + SP
Sbjct: 369 MCCSGFVYIKATTNPVDLWSAPSSQARKEKEYFDKHFGPFFRTEQLIIQAPNSHPDIYSP 428
Query: 404 RIVSAD-------NIRFLFEVQKKVDAI---RANYSGLMVSLQDICMKPL---DKDCATQ 450
AD N L +V DAI A++ V+L+DIC+ PL + +C
Sbjct: 429 YPSGADVPFGPPLNKDILHQVLDLQDAIVNIAASFDNETVTLKDICLAPLAPYNNNCTIL 488
Query: 451 SVLQYFKMDPRNFDDSGAVE---------HLNYCFQQYSSA-------DQCMSAFKAPLD 494
SVL YF+ D S + H YC + +S D C+ AF P+
Sbjct: 489 SVLNYFQNSHSVLDHSVGDDFFVYADYHTHFLYCVRAPASLNDTSLLHDPCLGAFGGPVF 548
Query: 495 PSTVLGGFSGKDYSGASAFIVTYPVNNAIDEEGNETAKAVAWEKAFIQLVKDELLPMAQS 554
P VLGG+ G +Y+ A+A ++T PVNN ++ + KA+AWEK FI +K+ P
Sbjct: 549 PWLVLGGYDGDNYNNATALVITLPVNNYYNDS-KKIMKALAWEKEFINFLKNYDNP---- 603
Query: 555 RNLTLAFSSESSIEEELKRESTADAITILVSYLVMFAYISLTLGDTPHPSSFYISSKVLL 614
NLT++FS+E SIE+E+ RES +D T+++SY+VMF YIS+ LG + SK+ L
Sbjct: 604 -NLTISFSAERSIEDEINRESRSDVSTVVISYVVMFVYISIALGHIQSWGRLLVDSKISL 662
Query: 615 GLSGVILVMLSVLGSVAIFSALGVKSTLIIMEVIPFLVLAVGVDNMCILVHAVKR-QPLE 673
G++G+++V+ SV S+ IFS GV TLI+ EVIPFLVLA+GVDN+ I+V ++R + LE
Sbjct: 663 GIAGILIVLSSVACSIGIFSYFGVPVTLIVAEVIPFLVLAIGVDNLFIIVQTLQRDERLE 722
Query: 674 -LPLEGRISNALVEVGPSITLASLSEVLAFAVGSFISMPACRVFSMFAALAVLLDFVLQV 732
L+ +I L +V PS+ L+S SE +AF +G+ +MPA R FS+FA +AVL+DF+LQV
Sbjct: 723 GETLDKQIGRVLGDVAPSMFLSSFSETVAFFLGTLSTMPAVRTFSLFAGMAVLIDFILQV 782
Query: 733 TAFVALIVLDSQRAEDKRVDCFPCIKVHSFHADPDKGIRQRKPGLLARYMKEVHAPILSI 792
T FV+L+ LD +R E R+D CIK ++ +G+ QR +L + K +++P L
Sbjct: 783 TCFVSLLGLDIRRQERNRLDILCCIK----GSEEMRGV-QRSESILFLFFKNLYSPYLLK 837
Query: 793 WGVKIVVIAIFVAFALASIALSTRIEPGLEQEIVLPRDSYLQGYFNNVSEYLRIGPPLYF 852
++ +VIA+FV S A+ +E GL+Q + +P DSY+ YF + +Y+ GPP+YF
Sbjct: 838 DWMRPIVIALFVGLLSFSTAVIHNVEIGLDQSLSMPDDSYVMNYFKQLGKYMHAGPPVYF 897
Query: 853 VV-KNYNYSSESTHTNQLCSISQCNSDSLLNEISKASLVPETSYIAKPAASWLDDFLVWI 911
V+ + +NY+S N +C + CN+DSL+ ++ A+ + + I +SW+DD+ W+
Sbjct: 898 VLEEGHNYTSLEGQ-NMVCGGTGCNNDSLVQQVFNAAEISSYTRIGYAPSSWIDDYFDWV 956
Query: 912 SPEAFGCCRKF-TNGSYCPPDDQPPCCAPEDDSCVSGACKDCTTCFRHSDLRNDRTSTMQ 970
P++ CCR + T G +C P CT C S R
Sbjct: 957 KPQS-SCCRVYNTTGQFCNASVTDP---------------SCTRCRPLSQEGKQRPQGED 1000
Query: 971 FRDKLPWFLSALPSADCAKGGHGAYTSSVDLKGYDSGIIQASSFRTYHTPLNKQVDYVNS 1030
F LP FLS P+ C KGGH AY+S+VDL + + A+ F TYHT L K DY+++
Sbjct: 1001 FMTFLPMFLSDNPNPKCGKGGHAAYSSAVDLIKNKTD-VGATYFMTYHTVLKKSSDYIDA 1059
Query: 1031 MRAAREFSSKVSDSLKVASGDKDQRV 1056
M+ AR+ + +++++ + +K+ RV
Sbjct: 1060 MKKARDIADNITETMGIK--EKNYRV 1083
Score = 102 bits (255), Expect = 5e-20
Identities = 53/99 (53%), Positives = 74/99 (74%), Gaps = 1/99 (1%)
Query: 1025 VDYVNSMRAAREFSSKVSDSLKVAS-GDKDQRVKEALGTMGASVFSGITLTKLVGVIVLY 1083
V+ V S A EF S V+ + V++ G + +R +EAL MG+SVFSGITLTK G++VL
Sbjct: 1160 VNLVMSCGIAVEFCSHVTRAFTVSTKGSRVERAEEALSHMGSSVFSGITLTKFGGIVVLA 1219
Query: 1084 FSRTEVFVIYYFQMYLSLVLLGFLHGLVFLPVVLSIFGP 1122
FS++++F I+YF+MYL++VLLG HGL+FLPV+LS GP
Sbjct: 1220 FSKSQIFKIFYFRMYLAMVLLGATHGLIFLPVLLSYIGP 1258
>UniRef100_UPI00002344B9 UPI00002344B9 UniRef100 entry
Length = 1271
Score = 573 bits (1478), Expect = e-162
Identities = 372/1080 (34%), Positives = 563/1080 (51%), Gaps = 103/1080 (9%)
Query: 5 CGTRSD-GKVLNCPFGSPAVKPDDLLSSKIQSMCPTI--TGNVCCTKAQFDTLQTQVQQA 61
CG +S G L CP A +P+ + K+ ++C G VCC + Q D L ++ A
Sbjct: 39 CGKQSFFGGELPCPDNDAAREPEAAVREKLVNLCGAKWQEGPVCCEEEQIDALSKNLKLA 98
Query: 62 IPFLVGCPACLRNFLNLFCELTCSPNQSLFINVTSVDKAG-GNSTVGGIDYFVSDAFGEG 120
+ CPAC NF N+FC TCSP+QSLFINVT +K G V +D S+ + G
Sbjct: 99 EGIISSCPACKDNFFNIFCTFTCSPDQSLFINVTKTEKGNSGKELVTELDNIWSEEYQSG 158
Query: 121 LYESCKDVKFGSMNSRAIQFIGAGAQNFKEWFAFIGRKAAPNSPGSPYAIMFRPNATKSS 180
Y+SCK+VK G+ +AI FIG GA++++++ F+G K GSP+ I + + S
Sbjct: 159 FYDSCKNVKNGASGGKAIDFIGGGAKDYQQFLKFLGDK---KFLGSPFQINYHTEPPEDS 215
Query: 181 -GMKPMNVSAYSCSDT--SLGCSCGDCPSSSVCSNSASTTINKANSCSIKVGSLTVKCVD 237
GM+ + + +C+D + CSC DCP VC + I C + + + C+
Sbjct: 216 QGMQALPIHPKACNDADPAYRCSCVDCPD--VCPELPA--IKTEEHCHVGL----LPCLS 267
Query: 238 FILAVLYIILICVFLGWALYHRIRERKMTYRTEPVSNVISGGVLYARNQEKDENLPMQMI 297
F + ++Y + + G++ Y RER+ + E V +L N DE
Sbjct: 268 FSVILIYSVFLLGVAGFSSYFTYRERRYR-KPERVR------LLQDPNPSDDE------- 313
Query: 298 EDVPQNRNGVRLSVVQGY------MSNFYRKYGSLVARHPINVLALTLAIVLLLCLGLIR 351
++ G L GY + + + GS+ AR P + ++ V+LL LG +R
Sbjct: 314 DEGDIVHAGGHLEYPHGYYKLNSMLDTVFSRIGSVCARFPALTIISSVVAVVLLSLGWLR 373
Query: 352 FKVETRPEKLWVGPGSKAAQEKQFFDSHLAPFYRIEQLILATVPDHMNSTSPRIVSADNI 411
F VET P +LWV P S AA+EK FFD + PFYR EQ L + + R++ D +
Sbjct: 374 FAVETDPVRLWVSPTSAAAKEKAFFDENFGPFYRAEQAFLVNDDE---TGDGRVLDYDTL 430
Query: 412 RFLFEVQKKVDAIRANYSGLMVSLQDICMKPLDKDCATQSVLQYFKMDPRNFDDSGAVEH 471
+ F V+ ++ + + GL SL DIC KP C QSV YF N D +
Sbjct: 431 TWWFGVESRIRRVISLDRGL--SLDDICYKPTGDACVIQSVTGYFGGSLSNLDPDTWQDR 488
Query: 472 LNYCFQQYSSADQCMSAFKAPLDPSTVLGGFSGK-DYSGASAFIVTYPVNNAIDEEGNET 530
L +C A C+ F PL P +LGG+ + A A IVT+ VNN E
Sbjct: 489 LTHCASSPGDAS-CLPDFSQPLRPEMILGGYEDSGNVLDAKALIVTWVVNNHAPGS-EEE 546
Query: 531 AKAVAWEKAF---IQLVKDELLPMAQSRNLTLAFSSESSIEEELKRESTADAITILVSYL 587
A+A+ WE F Q+V++E A++R L ++F++E+S+E+EL + S DA +++SY+
Sbjct: 547 AEAIDWEDTFRGIFQVVQEE----AKNRGLRVSFTTEASVEQELNKSSNTDAKIVVISYI 602
Query: 588 VMFAYISLTLGDTP--------HPSSFYISSKVLLGLSGVILVMLSVLGSVAIFSALGVK 639
+MF Y SL LG +P++ + SK LG+ G+++V++SV SV +FSA GVK
Sbjct: 603 IMFIYASLALGSVTMTWRSLINNPANALVQSKFTLGVVGIVIVLMSVSASVGLFSAAGVK 662
Query: 640 STLIIMEVIPFLVLAVGVDNMCILVHAVKRQPLELP---LEGRISNALVEVGPSITLASL 696
TLII EVIPFLVLAVGVDN+ ++V+ +R + P ++ RIS A+ +GPSI L+++
Sbjct: 663 VTLIIAEVIPFLVLAVGVDNIFLIVYEFERLNVSHPDEEIDERISRAIGRIGPSIFLSAI 722
Query: 697 SEVLAFAVGSFISMPACRVFSMFAALAVLLDFVLQVTAFVALIVLDSQRAEDKRVDCFPC 756
+E +AFA+G F+ MPA R F+++AA AV ++ VLQ+T FV+++ L+ +R E R DC PC
Sbjct: 723 TETVAFALGVFVGMPAVRNFAIYAAGAVFINAVLQITMFVSVLALNQKRVESLRADCIPC 782
Query: 757 IKVHSFHADPDKGIR---QRKPGLLARYMKEVHAPILSIWGVKIVVIAIFVAFALASIAL 813
+ V H+ + + Q + G+L +++++V+AP+L VK+VV+ F+ A +AL
Sbjct: 783 LTVRKAHSGMPEDLAFDDQDREGILQKFIRKVYAPLLLNRRVKVVVVITFLGILAAGLAL 842
Query: 814 STRIEPGLEQEIVLPRDSYLQGYFNNVSEYLRIGPPLYFVVKNYNYSSESTHTNQLCS-I 872
+ + GL+Q I LP DSYL YF+++SEY GPP+YFV +N N + H QLC
Sbjct: 843 TPEVAMGLDQRIALPSDSYLIDYFDDLSEYFNSGPPVYFVTRNVNITKRE-HQRQLCGRF 901
Query: 873 SQCNSDSLLNEISKASLVPETSYIAKPAASWLDDFLVWISPEAFGCCRKFTNGSYCPPDD 932
+ C SL + + S SYIA ASW+DDF W++P
Sbjct: 902 TTCEEYSLPFVLEQESKRSNVSYIAGATASWIDDFFYWLNP------------------- 942
Query: 933 QPPCCAPEDDSCVSGACKDCTTCFRHSDLRNDRTSTMQFRDKLPWFLSALPSADCAKGGH 992
Q CC + C G +F L ++ + A C GG
Sbjct: 943 QQDCCYEDGKLCFEGRTPGWNISL------TGMPEGAEFIHYLEKWIKSPTDASCPLGGK 996
Query: 993 GAYTSSV--DLKGYDSGIIQASSFRTYHTPLNKQVDYVNSMRAAREFSSKVSDSLKVASG 1050
Y++++ D K + AS FRT HTPL Q D++ S +AR +++D L G
Sbjct: 997 APYSNALVFDPKRITTN---ASHFRTSHTPLRTQDDFIKSYISAR----RIADGLSAEHG 1049
Score = 81.6 bits (200), Expect = 1e-13
Identities = 39/70 (55%), Positives = 53/70 (75%)
Query: 1052 KDQRVKEALGTMGASVFSGITLTKLVGVIVLYFSRTEVFVIYYFQMYLSLVLLGFLHGLV 1111
KD R AL +G SVFSGIT+TKL+GV VL F+R+++F IYYF+++L+L++ H L+
Sbjct: 1166 KDARSWTALVNVGGSVFSGITVTKLLGVCVLAFTRSKIFEIYYFRVWLALIIFAATHALI 1225
Query: 1112 FLPVVLSIFG 1121
FLPV LS FG
Sbjct: 1226 FLPVALSYFG 1235
>UniRef100_UPI00003635C8 UPI00003635C8 UniRef100 entry
Length = 1245
Score = 565 bits (1456), Expect = e-159
Identities = 378/1094 (34%), Positives = 577/1094 (52%), Gaps = 112/1094 (10%)
Query: 2 YDICGT--RSDGKVLNCPFGSPAVKPDDLLSSKIQ---SMCPTI---TGNVCCTKAQFDT 53
Y CG + GK NC + P P LL + +CP ++CC Q T
Sbjct: 10 YGECGESEKVPGKKYNCNYTGP---PKPLLPEGYELLTELCPGYDYENRSLCCDVNQLHT 66
Query: 54 LQTQVQQAIPFLVGCPACLRNFLNLFCELTCSPNQSLFINVTSVDKAGGNSTVGGIDYFV 113
L+ +Q + FL CPAC N +NLFCELTCSP+QS F+N T K G + V + Y++
Sbjct: 67 LKGSLQLPLQFLSRCPACFFNLMNLFCELTCSPHQSQFMNGT---KFSGPNVVE-VQYYI 122
Query: 114 SDAFGEGLYESCKDVKFGSMNSRAIQFI-GAGAQ--NFKEWFAFIGRKAAPNSPGSPYAI 170
F +Y +C+DV+ S N +A+ + G A+ N W ++ N+ +P+ I
Sbjct: 123 GQTFANAMYNACRDVQAPSSNVKALSLLCGKDAKDCNATNWIQYM---FDTNNGQAPFPI 179
Query: 171 MFRPNATKSSGMKPMNVSAYSCS----DTSLGCSCGDCPSSSVCSNSASTTINKANSCSI 226
+ SG PMN Y+C+ D S CSC DC + T +
Sbjct: 180 TAIFSDVPVSGYTPMNNKTYACTEGLEDGSGPCSCQDCTDAC-----GPTPVPPPPPLPW 234
Query: 227 KVGSLTVKCVDFILAVLYIILICVFLGWALYHRIRERKMTYRTEPVSNVISGGVLYARNQ 286
K+ L + + I+ + Y+ + +F G +L RK T ++E + S L + N+
Sbjct: 235 KI--LGIDAMTIIMWLSYMAFLFIFAG-SLLVAWCHRKQTIKSEYEPILDSNNPL-SLNR 290
Query: 287 EKDENLPMQMIEDVPQNRNGVRLSVVQGYMSNFYRKYGSLVARHPINVLALTLAIVLLLC 346
+ E + E + + + ++ + +GS HP VL ++ +V
Sbjct: 291 DNQEQVDASCCETLGER--------FENFLRTCFSVWGSFCVLHPFIVLLGSIVLVAASS 342
Query: 347 LGLIRFKVETRPEKLWVGPGSKAAQEKQFFDSHLAPFYRIEQLILAT--------VPDHM 398
GL+ ++ T P LW P S+A QEK +FDSH PF+R QLI+ + P
Sbjct: 343 GGLVYMQITTDPVDLWSSPKSQARQEKDYFDSHFGPFFRTAQLIITSPLNDTFIYTPYFG 402
Query: 399 NSTSP--RIVSADNIRFLFEVQKKVDAIRANYSGLMVSLQDICMKPL---DKDCATQSVL 453
S P I+S D + + ++Q ++ + A Y G V+L+DIC+ PL + +C SVL
Sbjct: 403 GSDVPFKAILSKDILDQVLDLQLDIENLVATYEGQNVTLKDICLAPLSPYNDNCTILSVL 462
Query: 454 QYFKMDPRNFDDSGAVE---------HLNYCFQQYSSA-------DQCMSAFKAPLDPST 497
YF+ + S E H YC +S D C+ F P+ P
Sbjct: 463 NYFQNSHSVLNHSRGDEFYVYADYHSHFLYCVSAPASLNDTTLLHDPCLGTFGGPVFPWL 522
Query: 498 VLGGFSGKDYSGASAFIVTYPVNNAIDEEGNETAKAVAWEKAFIQLVKDELLPMAQSRNL 557
LGG+ +Y+ A+A +VT+P+NN D K +AWEK FI+ +KD P NL
Sbjct: 523 ALGGYDDTNYNNATALVVTFPLNNNYDPA--RLGKTLAWEKEFIRFMKDYKNP-----NL 575
Query: 558 TLAFSSESSIEEELKRESTADAITILVSYLVMFAYISLTLGDTPHPSSFYISSKVLLGLS 617
T++FS+E SIE+E+ RES +D TI+VSY++MF YISL LG + SK+ LG+S
Sbjct: 576 TISFSAERSIEDEINRESNSDISTIVVSYVIMFVYISLALGHIQSFRRLLVDSKISLGIS 635
Query: 618 GVILVMLSVLGSVAIFSALGVKSTLIIMEVIPFLVLAVGVDNMCILVHAV---KRQPLEL 674
G+++V+ SV S+ IFS +G+ TLI++EVIPFLVLAVGVDN+ ILV + +R P E
Sbjct: 636 GILIVLSSVSSSLGIFSYIGIPLTLIVIEVIPFLVLAVGVDNIFILVQTLQRDERMPQE- 694
Query: 675 PLEGRISNALVEVGPSITLASLSEVLAFAVGSFISMPACRVFSMFAALAVLLDFVLQVTA 734
+ +I L +V PS+ L+S SE +AF +G+ +MPA R FS+FA LAV +DF+LQ++
Sbjct: 695 EIHHQIGRILGDVAPSMFLSSFSETVAFFLGALSNMPAVRTFSLFAGLAVFIDFLLQISC 754
Query: 735 FVALIVLDSQRAEDKRVDCFPCIKVHSFHADPDKGIRQRKPGLLARYMKEVHAP-ILSIW 793
FV+L+ LD+ R E R+D C+K+ +G +K L + K+++AP IL+ W
Sbjct: 755 FVSLLGLDASRQEGNRMDIVCCVKI--------QGEEVKKDSFLFLFFKKIYAPFILNDW 806
Query: 794 GVKIVVIAIFVAFALASIALSTRIEPGLEQEIVLPRDSYLQGYFNNVSEYLRIGPPLYFV 853
V+ V+A+FV SIA+ ++E GL+Q++ +P DSY+ YF N+SEYL G P+YFV
Sbjct: 807 -VRPFVVAVFVGMLSFSIAVMDKVEIGLDQKLSMPDDSYVLDYFKNLSEYLHTGAPVYFV 865
Query: 854 VKN-YNYSSESTHTNQLCSISQCNSDSLLNEISKASLVPETSYIAKPAASWLDDFLVWIS 912
V++ NYSS N +C CN++SL+ ++ ASL+ + +A +SWLDD+ W+
Sbjct: 866 VEDGLNYSSLEGQ-NAVCGGVGCNNNSLVQQVYTASLISNYTTVAYTPSSWLDDYFDWVK 924
Query: 913 PEAFGCCRKFT-NGSYCPPDDQPPCCAPEDDSCVSGACKDCTTCFRHSDLRNDRTSTMQF 971
P++ CCR + G++C + S V+ +C C R M+F
Sbjct: 925 PQS-TCCRYYNGTGAFC------------NASVVNSSCVHCRPMTPSGKERPVGDDFMRF 971
Query: 972 RDKLPWFLSALPSADCAKGGHGAYTSSVDLKGYDSGIIQASSFRTYHTPLNKQVDYVNSM 1031
LP FLS P+ C KGGH AY ++VDL ++G + A+ F TYHT L D++ ++
Sbjct: 972 ---LPMFLSDNPNVKCGKGGHAAYGTAVDLYPQNTG-VGATYFMTYHTILKDSPDFIKAL 1027
Query: 1032 RAAREFSSKVSDSL 1045
+ AR ++ ++ SL
Sbjct: 1028 KMARNLANNITQSL 1041
Score = 92.4 bits (228), Expect = 7e-17
Identities = 46/99 (46%), Positives = 70/99 (70%), Gaps = 1/99 (1%)
Query: 1025 VDYVNSMRAAREFSSKVSDSLKVA-SGDKDQRVKEALGTMGASVFSGITLTKLVGVIVLY 1083
V+ V S + EF S + + ++ + +R +EAL MG+SVFSGITLTK G+++L
Sbjct: 1122 VNLVMSCGISVEFCSHIVRAFSISLMTSRVKRAEEALAHMGSSVFSGITLTKFGGILILA 1181
Query: 1084 FSRTEVFVIYYFQMYLSLVLLGFLHGLVFLPVVLSIFGP 1122
S++++F ++YF+MYL++VLLG HGL+FLPV+LS GP
Sbjct: 1182 LSKSQIFQVFYFRMYLAIVLLGAAHGLIFLPVLLSYIGP 1220
>UniRef100_UPI000023DFE9 UPI000023DFE9 UniRef100 entry
Length = 1295
Score = 560 bits (1444), Expect = e-158
Identities = 374/1083 (34%), Positives = 572/1083 (52%), Gaps = 119/1083 (10%)
Query: 5 CGTRSD-GKVLNCPFGSPAVKPDDLLSSKIQSMCPTI--TGNVCCTKAQFDTLQTQVQQA 61
CG +S GK L C A P++ L +++ +C TG VCCT Q +L++++
Sbjct: 62 CGKQSFFGKELPCVDNGLAEDPEEELRNELVGLCGEKWRTGPVCCTLDQVKSLKSELGTP 121
Query: 62 IPFLVGCPACLRNFLNLFCELTCSPNQSLFINVTSVDKAGGNSTVGGIDYFVSDAFGEGL 121
+ CPAC NF NLFC TCSP+QS FINVT G + V +D+ VS+ +G G
Sbjct: 122 NTLIGSCPACKDNFFNLFCTFTCSPDQSTFINVTDSAPKNGKNLVTELDHLVSEKYGSGF 181
Query: 122 YESCKDVKFGSMNSRAIQFIGAGAQNFKEWFAFIGRKAAPNSPGSPYAIMFRPNATKSSG 181
Y+SCK+VKFG NSRA+ IG GA+N+ E F+G K GSP+ I F P K
Sbjct: 182 YDSCKEVKFGGANSRAMDLIGGGAKNYTEMLTFLGNKKP--FAGSPFQINF-PTQQKVPK 238
Query: 182 MKPMNVSAYSCSDT--SLGCSCGDCPSSSVCSNSASTTINKANSCSIKVGSLTVKCVDFI 239
++P+++ C+D + C C DCP VC+ +K S KVG L C+ F
Sbjct: 239 LQPVDMKPKKCNDEDPNYRCVCVDCPE--VCAKLPEVKDSK----SCKVGLLP--CLSFA 290
Query: 240 LAVLYIILICVFLGWALYHRIRERKMTYRTEPVSNVISGGVLYARNQEKDENLPM--QMI 297
+Y +L+ + H ++ +R E + + + ++DE P+ + +
Sbjct: 291 SIFVYGVLLSTLILAVTGHIAYQKYSQHRVERTRLLHES----SHSDDEDEGGPVDTEAM 346
Query: 298 EDVPQNRNGVRLSVVQGYMSNFYRKYGSLVARHPINVLALTLAIVLLLCLGLIRFKVETR 357
+ P R V +G+ + G + AR P + L+L V +L +GL RF +E
Sbjct: 347 RERPTKRYWVNDRCDRGFY-----QLGHIAARFPGWCIGLSLLFVGILSIGLFRFDLEKE 401
Query: 358 PEKLWVGPGSKAAQEKQFFDSHLAPFYRIEQLILATVPDHMNSTSPR----IVSADNIRF 413
P +LWV P S AAQEK +FD + PFYR E++ LA N T+P ++S D +++
Sbjct: 402 PARLWVSPSSAAAQEKAYFDENFGPFYRAEKIFLA------NDTNPSGPGPVLSYDTLKW 455
Query: 414 LFEVQKKVDAIRANYSGLMVSLQDICMKPLDKDCATQSVLQYFK----MDPRNFDDSGAV 469
EV++ V I + G QD+C KP + C QSV Y+ +DP+ + D
Sbjct: 456 WIEVEESVKKIESPVYGKY--FQDLCFKPSNDACVVQSVSAYWHSKGGLDPQTWKDD--- 510
Query: 470 EHLNYCFQQYSSADQCMSAFKAPLDPSTVLGGFSGKDYSGASAFIVTYPVNNAIDEEGNE 529
+ C + S C F P++P+ + GG+ G D A A VT+ VNNA +EG +
Sbjct: 511 --IRACAK---SPVDCRPDFGQPIEPNMIFGGY-GDDIVDAHAITVTWVVNNA--KEGTD 562
Query: 530 T-AKAVAWEKAFIQLVKDELLPM---AQSRNLTLAFSSESSIEEELKRESTADAITILVS 585
A+AV WE A ++D LL + A+ R L L+F++E S+E+EL + + DA I++S
Sbjct: 563 AIARAVDWETA----LRDRLLEVQEEAKERGLRLSFNTEISLEQELNKSTNTDAKIIVIS 618
Query: 586 YLVMFAYISLTLGDTP------HPSSFYISSKVLLGLSGVILVMLSVLGSVAIFSALGVK 639
Y+VMF Y + LG TP +P+ + SKV LGL G+I+V++S+ S+ FS +G+K
Sbjct: 619 YIVMFVYACMALG-TPLKHIFRNPAVLLVESKVTLGLVGIIIVLMSIAASIGFFSWVGLK 677
Query: 640 STLIIMEVIPFLVLAVGVDNMCILVHAVKRQPLELP---LEGRISNALVEVGPSITLASL 696
+TLII+EVIPF+VLAVGVDN+ ++VH ++R P +E R++ AL +GPSI ++L
Sbjct: 678 ATLIIVEVIPFIVLAVGVDNIFLIVHELERVNTSFPDQMVEERVARALGRMGPSILFSAL 737
Query: 697 SEVLAFAVGSFISMPACRVFSMFAALAVLLDFVLQVTAFVALIVLDSQRAEDKRVDCFPC 756
+E +AFA+G+ + MPA R F+ +AA AV ++ VLQ+T FV+ + L+ R ED R + +P
Sbjct: 738 TETVAFALGTAVGMPAVRNFAAYAAGAVFVNAVLQMTMFVSFLSLNQMRVEDHRCELWPW 797
Query: 757 IKV----------HSFHADPDKGIRQRKPGLLARYMKEVHAPILSIWGVKIVVIAIFVAF 806
++ + F +G + LL ++K +AP L VK+ V+ IF+
Sbjct: 798 WQITKARIHLNGSNGFAQGGGRGSDMAEESLLQVFIKNTYAPRLLGKKVKLAVVTIFLGM 857
Query: 807 ALASIALSTRIEPGLEQEIVLPRDSYLQGYFNNVSEYLRIGPPLYFVVKNYNYSSESTHT 866
+AL +I+ GL+Q + +P SYL YFN++ YL GPP+YFV + + +S+
Sbjct: 858 FAGGLALLPKIQLGLDQRVAIPDGSYLIPYFNDLYGYLETGPPVYFVTREVD-ASKRKEQ 916
Query: 867 NQLCS-ISQCNSDSLLNEISKASLVPETSYIAKPAASWLDDFLVWISPEAFGCCRKFTNG 925
+CS + C SL N + PE SYIA PAASW+DD+ +W++P CC + +G
Sbjct: 917 QAICSRFTTCQDLSLPNTLELERQRPEVSYIASPAASWIDDYFLWLNPIFEDCCVE--HG 974
Query: 926 SYCPPDDQPPCCA-----PEDDSCVSGACKDCTTCFRHSDLRNDRTSTMQFRDKLPWFLS 980
C D P PED+ +F L FLS
Sbjct: 975 QTCFADRVPAWNTTLYGMPEDE---------------------------EFIHYLKKFLS 1007
Query: 981 ALPSADCAKGGHGAYTSSVDLKGYDSGIIQASSFRTYHTPLNKQVDYVNSMRAAREFSSK 1040
+ +C G AY +V L ++ I +++ FRT H+PL Q D++ + AAR +S
Sbjct: 1008 SPTGEECPLAGQAAYGQAVVLDSKENHI-KSTHFRTMHSPLRSQEDFIAAYSAARRIASD 1066
Query: 1041 VSD 1043
+ +
Sbjct: 1067 IGE 1069
Score = 83.6 bits (205), Expect = 3e-14
Identities = 42/70 (60%), Positives = 54/70 (77%)
Query: 1052 KDQRVKEALGTMGASVFSGITLTKLVGVIVLYFSRTEVFVIYYFQMYLSLVLLGFLHGLV 1111
+D R AL +G SVFSGIT+TKL+GV VL F+R+++F IYYF+++LSLV+ LH LV
Sbjct: 1189 RDARAWTALVNVGGSVFSGITVTKLLGVSVLAFTRSKIFEIYYFRVWLSLVIFAALHALV 1248
Query: 1112 FLPVVLSIFG 1121
FLPV LSI G
Sbjct: 1249 FLPVALSIAG 1258
>UniRef100_Q7RWL9 Hypothetical protein [Neurospora crassa]
Length = 1162
Score = 558 bits (1437), Expect = e-157
Identities = 349/993 (35%), Positives = 536/993 (53%), Gaps = 87/993 (8%)
Query: 78 LFCELTCSPNQSLFINVTSVDKAGGNSTVGGIDYFVSDAFGEGLYESCKDVKFGSMNSRA 137
+FC TCSPNQSLF+NVT + G V +D +S+ +G G Y SCKDVKFG NSRA
Sbjct: 1 MFCTFTCSPNQSLFVNVTKTIEKKGKELVTELDQLISEEYGTGFYNSCKDVKFGPTNSRA 60
Query: 138 IQFIGAGAQNFKEWFAFIGRKAAPNSPGSPYAIMFRPNATKSSGMKPMNVSAYSCSDT-- 195
+ IG GA+N+ + F+G++ GSP+ I F P MKP+ + C+D
Sbjct: 61 MDLIGGGAKNYTQLLKFLGQE---RFGGSPFQINF-PVEYAEPDMKPLPMKPKKCNDEDP 116
Query: 196 SLGCSCGDCPSSSVCSNSASTTINKANSCSIKVGSLTVKCVDFILAVLYIILICVFLGWA 255
+ C+C DCP +C + +A SC VG+L C+ F + Y +++ + +
Sbjct: 117 NFRCACVDCPE--ICPTLPD--VEQAGSCH--VGALP--CLSFASILTYSVILFISIAAV 168
Query: 256 LYHRIRERKMTYRTEPVSNVISGGVLYARNQEKDENLPMQMIEDVPQNRNGVRLSVVQGY 315
+ H +R R+E + + A + ++DE + ++V + ++ +
Sbjct: 169 VGHVAWKRHAKRRSERLRLLTDA----APSDDEDEG---DLTQNVAMIDRPQKTYIINTW 221
Query: 316 MSNFYRKYGSLVARHPINVLALTLAIVLLLCLGLIRFKVETRPEKLWVGPGSKAAQEKQF 375
+ + K G + A P + ++ I +L LG F++E P +LWV P S AA+EK F
Sbjct: 222 CDSAFSKLGYVAATFPAITIVTSILIASILSLGWFHFELEKNPARLWVSPTSPAAEEKAF 281
Query: 376 FDSHLAPFYRIEQLILATVPDHMNSTSPRIVSADNIRFLFEVQKKVDAIRANYSGLMVSL 435
FDSH FYR E++ L V D S ++S D + + +V+K V A++ + G S
Sbjct: 282 FDSHFGAFYRAEKVFL--VNDTQPSGPGPVLSRDTLLWWMDVEKSVAALKGSNYGS--SF 337
Query: 436 QDICMKPLDKDCATQSVLQYFKMDPRNFDDSGAVEHLNYCFQQYSSADQCMSAFKAPLDP 495
QD+C+KP C QSV YF+ DP + D L C +S +C A+ PLDP
Sbjct: 338 QDLCLKPTGDACVVQSVAAYFQDDPDSVDPETWQSTLRTCA---ASPVECRPAYGQPLDP 394
Query: 496 STVLGGF-SGKDYSGASAFIVTYPVNNAIDEEGNETAKAVAWEKAFIQLVKDELLPM--- 551
S +LGG+ G + + ASA VT+ + N E E +A+ WE A +K+ LL +
Sbjct: 395 SMILGGYPEGGNVAEASAMTVTWVLINP-SENSPEVDRAMDWEVA----LKNRLLEVQDE 449
Query: 552 AQSRNLTLAFSSESSIEEELKRESTADAITILVSYLVMFAYISLTLGDTP--------HP 603
A+ R L L+FS+E S+EEEL + + DA I++SY++MF Y SL LG T +P
Sbjct: 450 AKERGLRLSFSTEISLEEELNKSTNTDAKIIVISYIIMFLYASLALGSTTLTFKDLIRNP 509
Query: 604 SSFYISSKVLLGLSGVILVMLSVLGSVAIFSALGVKSTLIIMEVIPFLVLAVGVDNMCIL 663
+ + SK LG+ G+++V++S+ S+ +FS G+++TLII++VIPF+VLAVGVDN+ ++
Sbjct: 510 AVSLVESKFTLGIVGIVIVLMSITASIGLFSWAGLRATLIIVDVIPFIVLAVGVDNIFLI 569
Query: 664 VHAVKRQPLELP---LEGRISNALVEVGPSITLASLSEVLAFAVGSFISMPACRVFSMFA 720
VH +R + P +E RIS AL +GPSI ++L+E +FA+G+F+ MPA R F+++A
Sbjct: 570 VHEFERVNVSYPDDMVEARISRALGRMGPSILFSALTETASFALGAFVGMPAVRNFAIYA 629
Query: 721 ALAVLLDFVLQVTAFVALIVLDSQRAEDKRVDCFPCIKVHSFH---ADPDKG-----IRQ 772
A AV ++ +LQVT FV+++ L+ R ED R DCFPCI++ S A G +
Sbjct: 630 AGAVFINAILQVTMFVSVLTLNQIRVEDSRADCFPCIQIKSARVHLASNGAGPAPVYLEA 689
Query: 773 RKPGLLARYMKEVHAPILSIWGVKIVVIAIFVAFALASIALSTRIEPGLEQEIVLPRDSY 832
+ L +++++V+AP L K V++ IF+ A +AL ++ GL+Q + +P DSY
Sbjct: 690 PEESYLQQFIRKVYAPRLLGKKTKAVIVMIFLGVFAAGVALIPEVKLGLDQRVAIPDDSY 749
Query: 833 LQGYFNNVSEYLRIGPPLYFVVKNYNYSSESTHTNQLCSISQCNSDSLLNEISKASLVPE 892
L YFN++ EYL GPP+YFV + +N + + + C SL N + + E
Sbjct: 750 LIPYFNDLYEYLNTGPPVYFVTREFNATDRAQQQKVCARYTTCEQMSLSNILEQERKRTE 809
Query: 893 TSYIAKPAASWLDDFLVWISPEAFGCCRKFTNGSYCPPDDQPPCCA---PEDDSCVSGAC 949
SYI+ P ASW+DDF W++PE CC + + PC A P + +SG
Sbjct: 810 VSYISTPTASWIDDFFQWLNPENERCCM----------ERRRPCFANRTPAWNITLSG-- 857
Query: 950 KDCTTCFRHSDLRNDRTSTMQFRDKLPWFLSALPSADCAKGGHGAYTSSVDLKGYDSGII 1009
+F L FLSA + DC GG +Y S+V L D I
Sbjct: 858 ---------------MPEGDEFVYYLKKFLSAPTNEDCPLGGQASYGSAV-LVDSDRDTI 901
Query: 1010 QASSFRTYHTPLNKQVDYVNSMRAAREFSSKVS 1042
AS FRT H PL Q D++++ AAR ++++S
Sbjct: 902 PASHFRTSHIPLRSQEDFIDAYAAARRIANEIS 934
Score = 80.5 bits (197), Expect = 3e-13
Identities = 38/70 (54%), Positives = 53/70 (75%)
Query: 1052 KDQRVKEALGTMGASVFSGITLTKLVGVIVLYFSRTEVFVIYYFQMYLSLVLLGFLHGLV 1111
+D R AL +G SVFSGIT+TKL+GV VL F+R+++F IYYF+++++LV+ H LV
Sbjct: 1055 RDARAWTALSNVGGSVFSGITVTKLLGVFVLGFTRSKIFEIYYFRIWVALVIFAATHALV 1114
Query: 1112 FLPVVLSIFG 1121
FLPV LS+ G
Sbjct: 1115 FLPVALSLVG 1124
>UniRef100_O15118 Niemann-Pick C1 protein precursor [Homo sapiens]
Length = 1278
Score = 555 bits (1430), Expect = e-156
Identities = 370/1100 (33%), Positives = 566/1100 (50%), Gaps = 110/1100 (10%)
Query: 2 YDICGTRSDGKVLNCPF-GSPAVKPDDLLSSKIQSMCPTIT-GNV--CCTKAQFDTLQTQ 57
Y CG K NC + G P P D +Q +CP GNV CC Q TL+
Sbjct: 28 YGECGIAYGDKRYNCEYSGPPKPLPKDGYDL-VQELCPGFFFGNVSLCCDVRQLQTLKDN 86
Query: 58 VQQAIPFLVGCPACLRNFLNLFCELTCSPNQSLFINVTS----VDKAGGNS--TVGGIDY 111
+Q + FL CP+C N LNLFCELTCSP QS F+NVT+ VD + V + Y
Sbjct: 87 LQLPLQFLSRCPSCFYNLLNLFCELTCSPRQSQFLNVTATEDYVDPVTNQTKTNVKELQY 146
Query: 112 FVSDAFGEGLYESCKDVKFGSMNSRAIQFI---GAGAQNFKEWFAFIGRKAAPNSPGSPY 168
+V +F +Y +C+DV+ S N +A+ + A A N W ++ K ++ +P+
Sbjct: 147 YVGQSFANAMYNACRDVEAPSSNDKALGLLCGKDADACNATNWIEYMFNK---DNGQAPF 203
Query: 169 AIMFRPNATKSSGMKPMNVSAYSCSDT----SLGCSCGDCPSSSVCSNSASTTINKANSC 224
I + GM+PMN + C ++ + CSC DC S VC A
Sbjct: 204 TITPVFSDFPVHGMEPMNNATKGCDESVDEVTAPCSCQDC--SIVCGPKPQPPPPPAPWT 261
Query: 225 SIKVGSLTVKCVDFILAVLYIILICVFLG-----WALYHRIRERKMTYRTEPVSNVISGG 279
+ + ++ V I+ + Y+ + VF G W R+R P+ + I+
Sbjct: 262 ILGLDAMYV-----IMWITYMAFLLVFFGAFFAVWCY----RKRYFVSEYTPIDSNIAFS 312
Query: 280 VLYARNQEKDENLPMQMIEDVPQNRNGVRLSVVQGYMSNFYRKYGSLVARHPINVLALTL 339
V + E P+ + +G + + ++GS R+P V+ +L
Sbjct: 313 VNASDKGEASCCDPVS--------------AAFEGCLRRLFTRWGSFCVRNPGCVIFFSL 358
Query: 340 AIVLLLCLGLIRFKVETRPEKLWVGPGSKAAQEKQFFDSHLAPFYRIEQLILATVPDHMN 399
+ GL+ +V T P LW P S+A EK++FD H PF+R EQLI+ +
Sbjct: 359 VFITACSSGLVFVRVTTNPVDLWSAPSSQARLEKEYFDQHFGPFFRTEQLIIRAPLTDKH 418
Query: 400 STSPRIVSAD-------NIRFLFEV---QKKVDAIRANYSGLMVSLQDICMKPL---DKD 446
P AD +I+ L +V Q ++ I A+Y V+LQDIC+ PL + +
Sbjct: 419 IYQPYPSGADVPFGPPLDIQILHQVLDLQIAIENITASYDNETVTLQDICLAPLSPYNTN 478
Query: 447 CATQSVLQYFK-----MDPRNFDD----SGAVEHLNYCFQQYSSA-------DQCMSAFK 490
C SVL YF+ +D + DD + H YC + +S D C+ F
Sbjct: 479 CTILSVLNYFQNSHSVLDHKKGDDFFVYADYHTHFLYCVRAPASLNDTSLLHDPCLGTFG 538
Query: 491 APLDPSTVLGGFSGKDYSGASAFIVTYPVNNAIDEEGNETAKAVAWEKAFIQLVKDELLP 550
P+ P VLGG+ ++Y+ A+A ++T+PVNN ++ + +A AWEK FI VK+ P
Sbjct: 539 GPVFPWLVLGGYDDQNYNNATALVITFPVNNYYNDT-EKLQRAQAWEKEFINFVKNYKNP 597
Query: 551 MAQSRNLTLAFSSESSIEEELKRESTADAITILVSYLVMFAYISLTLGDTPHPSSFYISS 610
NLT++F++E SIE+EL RES +D T+++SY +MF YISL LG + S
Sbjct: 598 -----NLTISFTAERSIEDELNRESDSDVFTVVISYAIMFLYISLALGHIKSCRRLLVDS 652
Query: 611 KVLLGLSGVILVMLSVLGSVAIFSALGVKSTLIIMEVIPFLVLAVGVDNMCILVHAVKRQ 670
KV LG++G+++V+ SV S+ +FS +G+ TLI++EVIPFLVLAVGVDN+ ILV A +R
Sbjct: 653 KVSLGIAGILIVLSSVACSLGVFSYIGLPLTLIVIEVIPFLVLAVGVDNIFILVQAYQRD 712
Query: 671 P--LELPLEGRISNALVEVGPSITLASLSEVLAFAVGSFISMPACRVFSMFAALAVLLDF 728
L+ ++ L EV PS+ L+S SE +AF +G+ MPA FS+FA LAV +DF
Sbjct: 713 ERLQGETLDQQLGRVLGEVAPSMFLSSFSETVAFFLGALSVMPAVHTFSLFAGLAVFIDF 772
Query: 729 VLQVTAFVALIVLDSQRAEDKRVDCFPCIKVHSFHADPDKGIRQRKPGLLARYMKEVHAP 788
+LQ+T FV+L+ LD +R E R+D F C++ D Q L R+ K ++P
Sbjct: 773 LLQITCFVSLLGLDIKRQEKNRLDIFCCVR-----GAEDGTSVQASESCLFRFFKNSYSP 827
Query: 789 ILSIWGVKIVVIAIFVAFALASIALSTRIEPGLEQEIVLPRDSYLQGYFNNVSEYLRIGP 848
+L ++ +VIAIFV SIA+ +++ GL+Q + +P DSY+ YF ++S+YL GP
Sbjct: 828 LLLKDWMRPIVIAIFVGVLSFSIAVLNKVDIGLDQSLSMPDDSYMVDYFKSISQYLHAGP 887
Query: 849 PLYFVVKNYNYSSESTHTNQLCSISQCNSDSLLNEISKASLVPETSYIAKPAASWLDDFL 908
P+YFV++ + + S N +C CN+DSL+ +I A+ + + I +SW+DD+
Sbjct: 888 PVYFVLEEGHDYTSSKGQNMVCGGMGCNNDSLVQQIFNAAQLDNYTRIGFAPSSWIDDYF 947
Query: 909 VWISPEAFGCCR-KFTNGSYCPPDDQPPCCAPEDDSCVSGACKDCTTCFRHSDLRNDRTS 967
W+ P++ CCR +C + S V AC C R
Sbjct: 948 DWVKPQS-SCCRVDNITDQFC------------NASVVDPACVRCRPLTPEGKQRPQGGD 994
Query: 968 TMQFRDKLPWFLSALPSADCAKGGHGAYTSSVDLKGYDSGIIQASSFRTYHTPLNKQVDY 1027
M+F LP FLS P+ C KGGH AY+S+V++ + A+ F TYHT L D+
Sbjct: 995 FMRF---LPMFLSDNPNPKCGKGGHAAYSSAVNILLGHGTRVGATYFMTYHTVLQTSADF 1051
Query: 1028 VNSMRAAREFSSKVSDSLKV 1047
+++++ AR +S V++++ +
Sbjct: 1052 IDALKKARLIASNVTETMGI 1071
Score = 99.8 bits (247), Expect = 4e-19
Identities = 50/99 (50%), Positives = 73/99 (73%), Gaps = 1/99 (1%)
Query: 1025 VDYVNSMRAAREFSSKVSDSLKVA-SGDKDQRVKEALGTMGASVFSGITLTKLVGVIVLY 1083
V+ V S + EF S ++ + V+ G + +R +EAL MG+SVFSGITLTK G++VL
Sbjct: 1155 VNLVMSCGISVEFCSHITRAFTVSMKGSRVERAEEALAHMGSSVFSGITLTKFGGIVVLA 1214
Query: 1084 FSRTEVFVIYYFQMYLSLVLLGFLHGLVFLPVVLSIFGP 1122
F+++++F I+YF+MYL++VLLG HGL+FLPV+LS GP
Sbjct: 1215 FAKSQIFQIFYFRMYLAMVLLGATHGLIFLPVLLSYIGP 1253
>UniRef100_UPI000036A3B7 UPI000036A3B7 UniRef100 entry
Length = 1277
Score = 551 bits (1419), Expect = e-155
Identities = 368/1100 (33%), Positives = 569/1100 (51%), Gaps = 111/1100 (10%)
Query: 2 YDICGTRSDGKVLNCPF-GSPAVKPDDLLSSKIQSMCPTIT-GNV--CCTKAQFDTLQTQ 57
Y CG K NC + G P P D +Q +CP GNV CC Q TL+
Sbjct: 28 YGECGIAYGDKRYNCEYSGPPKPLPKDGYDL-VQELCPGFFFGNVSLCCDVQQLQTLKDN 86
Query: 58 VQQAIPFLVGCPACLRNFLNLFCELTCSPNQSLFINVTS----VDKAGGNS--TVGGIDY 111
+Q + FL CP+C N LNLFCELTCSP QS F+NVT+ VD + V + Y
Sbjct: 87 LQLPLQFLSRCPSCFYNLLNLFCELTCSPRQSQFLNVTATEDYVDPVTNQTKTNVKELQY 146
Query: 112 FVSDAFGEGLYESCKDVKFGSMNSRAIQFI---GAGAQNFKEWFAFIGRKAAPNSPGSPY 168
+V +F +Y +C+DV+ S N +A+ + A A N W ++ K ++ +P+
Sbjct: 147 YVGQSFANAMYNACRDVEAPSSNDKALGLLCGKDADACNATNWIEYMFNK---DNGQAPF 203
Query: 169 AIMFRPNATKSSGMKPMNVSAYSCSDT----SLGCSCGDCPSSSVCSNSASTTINKANSC 224
I + GM+PMN + C ++ + CSC DC S VC A
Sbjct: 204 TITPVFSDFPVHGMEPMNNATKGCDESVDEVTAPCSCQDC--SIVCGPKPQPPPPPAPWT 261
Query: 225 SIKVGSLTVKCVDFILAVLYIILICVFLG-----WALYHRIRERKMTYRTEPVSNVISGG 279
+ + ++ V I+ + Y+ + VF G W R+R P+ + I+
Sbjct: 262 ILGLDAMYV-----IMWITYMAFLLVFFGAFFAVWCY----RKRYFVSEYTPIDSNIAFS 312
Query: 280 VLYARNQEKDENLPMQMIEDVPQNRNGVRLSVVQGYMSNFYRKYGSLVARHPINVLALTL 339
V + E P+ + +G + + ++GS R+P V+ +L
Sbjct: 313 VNASDKGEASCCDPVS--------------AAFEGCLRRLFTRWGSFCVRNPGCVIFFSL 358
Query: 340 AIVLLLCLGLIRFKVETRPEKLWVGPGSKAAQEKQFFDSHLAPFYRIEQLILATVPDHMN 399
+ GL+ V T P LW P S+A EK++FD H PF+R EQLI+ +
Sbjct: 359 VFITACSSGLVFVWVTTNPVDLWSAPSSQARLEKEYFDQHFGPFFRTEQLIIRAPLTDKH 418
Query: 400 STSPRIVSAD-------NIRFLFEV---QKKVDAIRANYSGLMVSLQDICMKPL---DKD 446
+ P AD +I+ L +V Q ++ I A+ + V+LQDIC+ PL + +
Sbjct: 419 TYQPYPSGADVPFGPPLDIQILHQVLDLQIAIENITASSNNETVTLQDICLAPLSPYNTN 478
Query: 447 CATQSVLQYFK-----MDPRNFDD----SGAVEHLNYCFQQYSSA-------DQCMSAFK 490
C SVL YF+ +D + DD + H YC + +S D C+ F
Sbjct: 479 CTILSVLNYFQNSHSVLDHKKGDDFFVYADYHTHFLYCVRAPASLNDTSLLHDPCLGTFG 538
Query: 491 APLDPSTVLGGFSGKDYSGASAFIVTYPVNNAIDEEGNETAKAVAWEKAFIQLVKDELLP 550
P+ P VLGG+ ++Y+ A+A ++T+PVNN ++ + +A AWEK FI VK+ P
Sbjct: 539 GPVFPWLVLGGYDDQNYNNATALVITFPVNNYYNDT-EKLQRAQAWEKEFINFVKNYKNP 597
Query: 551 MAQSRNLTLAFSSESSIEEELKRESTADAITILVSYLVMFAYISLTLGDTPHPSSFYISS 610
NL+++F++E SIE+EL+RES +D T+++SY +MF YISL LG + S
Sbjct: 598 -----NLSISFTAERSIEDELRRESDSDVFTVVISYAIMFLYISLALGHIKSCRRLLVDS 652
Query: 611 KVLLGLSGVILVMLSVLGSVAIFSALGVKSTLIIMEVIPFLVLAVGVDNMCILVHAVKRQ 670
KV LG++G+++V+ SV S+ +FS +G+ TLI++EVIPFLVLAVGVDN+ ILV A +R
Sbjct: 653 KVSLGIAGILIVLSSVACSLGVFSYIGLPLTLIVIEVIPFLVLAVGVDNIFILVQAYQRD 712
Query: 671 P--LELPLEGRISNALVEVGPSITLASLSEVLAFAVGSFISMPACRVFSMFAALAVLLDF 728
L+ ++ L EV PS+ L+S SE +AF +G+ MPA FS+FA LAV +DF
Sbjct: 713 ERLQGETLDQQLGRVLGEVAPSMFLSSFSETVAFFLGALSVMPAVHTFSLFAGLAVFIDF 772
Query: 729 VLQVTAFVALIVLDSQRAEDKRVDCFPCIKVHSFHADPDKGIRQRKPGLLARYMKEVHAP 788
+LQ+T FV+L+ LD +R E R+D F C++ D Q L R+ K ++P
Sbjct: 773 LLQITCFVSLLGLDIKRQEKNRLDIFCCVR-----GAEDGTSVQASESCLFRFFKNSYSP 827
Query: 789 ILSIWGVKIVVIAIFVAFALASIALSTRIEPGLEQEIVLPRDSYLQGYFNNVSEYLRIGP 848
+L ++ +VIA+FV SIA+ +++ GL+Q + +P DSY+ YF ++S+YL GP
Sbjct: 828 LLLKDWMRPIVIAVFVGVLSFSIAVLNKVDIGLDQSLSMPDDSYVVDYFKSISQYLHAGP 887
Query: 849 PLYFVVKNYNYSSESTHTNQLCSISQCNSDSLLNEISKASLVPETSYIAKPAASWLDDFL 908
P+YFV++ + + S N +C CN+DSL+ +I A+ + + I +SW+DD+
Sbjct: 888 PVYFVLEEGHDYTSSKGQNMVCGGMGCNNDSLVQQIFNAAQLDNYTRIGFAPSSWIDDYF 947
Query: 909 VWISPEAFGCCR-KFTNGSYCPPDDQPPCCAPEDDSCVSGACKDCTTCFRHSDLRNDRTS 967
W+ P++ CCR +C + S V AC C R
Sbjct: 948 DWVKPQS-SCCRVDNITDQFC------------NASVVDPACVRCRPLTPEGKQRPQGGD 994
Query: 968 TMQFRDKLPWFLSALPSADCAKGGHGAYTSSVDLKGYDSGIIQASSFRTYHTPLNKQVDY 1027
M+F LP FLS P+ C KGGH AY+S+V++ G + + A+ F TYHT L D+
Sbjct: 995 FMRF---LPMFLSDNPNPKCGKGGHAAYSSAVNILG-NGTRVGATYFMTYHTVLQTSADF 1050
Query: 1028 VNSMRAAREFSSKVSDSLKV 1047
+++++ AR +S V++++ +
Sbjct: 1051 IDALKKARLIASNVTETMGI 1070
Score = 100 bits (248), Expect = 3e-19
Identities = 50/99 (50%), Positives = 74/99 (74%), Gaps = 1/99 (1%)
Query: 1025 VDYVNSMRAAREFSSKVSDSLKVAS-GDKDQRVKEALGTMGASVFSGITLTKLVGVIVLY 1083
V+ V S + EF S ++ + V++ G + +R +EAL MG+SVFSGITLTK G++VL
Sbjct: 1154 VNLVMSCGISVEFCSHITRAFTVSTKGSRVERAEEALAHMGSSVFSGITLTKFGGIVVLA 1213
Query: 1084 FSRTEVFVIYYFQMYLSLVLLGFLHGLVFLPVVLSIFGP 1122
F+++++F I+YF+MYL++VLLG HGL+FLPV+LS GP
Sbjct: 1214 FAKSQIFQIFYFRMYLAMVLLGATHGLIFLPVLLSYIGP 1252
>UniRef100_UPI0000318C66 UPI0000318C66 UniRef100 entry
Length = 1209
Score = 550 bits (1417), Expect = e-155
Identities = 370/1084 (34%), Positives = 569/1084 (52%), Gaps = 126/1084 (11%)
Query: 33 IQSMCPTIT-GN--VCCTKAQFDTLQTQVQQAIPFLVGCPACLRNFLNLFCELTCSPNQS 89
+Q +CP GN +CC Q TL+ ++ + FL CPAC N +NLFCELTCSP+QS
Sbjct: 3 LQELCPGYDYGNRSLCCDVNQLHTLKESLEVPLQFLSRCPACFFNLVNLFCELTCSPHQS 62
Query: 90 LFINVTSVDKAGGNSTVGGIDYFVSDAFGEGLYESCKDVKFGSMNSRAIQFI-GAGAQ-- 146
F+N T K G V + Y++ F +Y +C+DV+ S N +A+ + G A+
Sbjct: 63 QFMNAT---KLSGPDVVE-VQYYIGLTFANAMYNACRDVQAPSSNVKALSLLCGKDAKDC 118
Query: 147 NFKEWFAFIGRKAAPNSPGSPYAIMFRPNATKSSGMKPMNVSAYSCSDT----SLGCSCG 202
N W ++ ++ +P+ I + SG PMN +C+D S CSC
Sbjct: 119 NATNWIQYMFNT---DNEQAPFPITPIFSDVPVSGYTPMNNDTAACTDGLEDGSGPCSCQ 175
Query: 203 DCPSSSVCSNSASTTINKANSCSIKVGSLTVKCVDFILAVLYIILICVFLGWALYHRIRE 262
DC ++ C A + + ++TV +A L I + + + W
Sbjct: 176 DC--TNACGPRPVPPPTPAPWKILGMDAMTVIMWFSYMAFLLIFVGSLLIAWC------H 227
Query: 263 RKMTYRTEPVSNVISGGVLYARNQEKDENLPMQMIEDVPQNRN-GVRLSVVQGYMSNFYR 321
RK T +E ++ + N++ +++P +++D G R + Y+ + +
Sbjct: 228 RKETIMSE-YGPILDSKNRPSLNRDNPDHVPFFILDDASCCETLGERF---ESYLRSCFS 283
Query: 322 KYGSLVARHPINVLALTLAIVLLLCLGLIRFKVETRPEKLWVGPGSKAAQEKQFFDSHLA 381
+GS HP VL +L +V+ GLI ++ T P LW P S+A QE+++FDSH
Sbjct: 284 CWGSFCVLHPCVVLLGSLILVVASSGGLIYMRITTDPVDLWSSPSSQARQEREYFDSHFG 343
Query: 382 PFYRIEQLILATVPDHMNSTSP----------RIVSADNIRFLFEVQKKVDAIRANYSGL 431
PF+R QLI+ + + SP ++S D + + ++Q ++++ A Y G
Sbjct: 344 PFFRTAQLIITSPLNDTFLYSPVMGGPDIPFKAVLSKDILHQVLDLQLDIESLVATYEGQ 403
Query: 432 MVSLQDICMKPL---DKDCATQSVLQYFKMDPRNFD----DSGAV-----EHLNYCFQQY 479
V+L+DIC+ PL + +C SVL YF+ D D V H YC
Sbjct: 404 SVTLKDICLAPLSPYNDNCTILSVLNYFQNSHSTLDHVLKDEFLVYADFHSHFLYCVSAP 463
Query: 480 SSA-------DQCMSAFKAPLDPSTVLGGFSGKDYSGASAFIVTYPVNNAIDEEGNETAK 532
+S D C+ F P+ P LGG+ +Y+ A+A +VT+P+NN D + K
Sbjct: 464 ASLNDTTPLHDPCLGTFGGPVFPWLALGGYDDTNYNNATALVVTFPINNNYDP--TKLGK 521
Query: 533 AVAWEKAFIQLVKDELLPMAQSRNLTLAFSSESSIEEELKRESTADAITILVSYLVMFAY 592
+AWEK FI+ +K+ P NLT+AFS+E SIE+E+ RES +D TI+VSY++MF Y
Sbjct: 522 TLAWEKEFIRFMKNYSNP-----NLTIAFSAERSIEDEINRESNSDISTIVVSYVIMFVY 576
Query: 593 ISLTLGDTPHPSSF-------------YISSKVLLGLSGVILVMLSVLGSVAIFSALGVK 639
ISL LG H SF + SKV LG+SG+++V+ SV S+ IFS G+
Sbjct: 577 ISLALG---HIQSFTRLLPHVLLLLLLLVDSKVSLGISGILIVLSSVSSSLGIFSYFGIP 633
Query: 640 STLIIMEVIPFLVLAVGVDNMCILVHAVKRQPLELP---LEGRISNALVEVGPSITLASL 696
TLI++EVIPFLVLAVGVDN+ I+V ++R +P L +I L +V PS+ L+S
Sbjct: 634 LTLIVIEVIPFLVLAVGVDNIFIIVQTLQRDE-RMPHEELHQQIGRILGDVAPSMFLSSF 692
Query: 697 SEVLAF-------------AVGSFISMPACRVFSMFAALAVLLDFVLQVTAFVALIVLDS 743
SE +AF A+G+ +MPA R FS+FA LAV +DF+LQ++ FV+L+ LD+
Sbjct: 693 SETVAFFLGKFNSSLKLFEAIGALSNMPAARTFSLFAGLAVFIDFLLQISCFVSLLGLDA 752
Query: 744 QRAEDKRVDCFPCIKVHSFHADPDKGIRQRKPGLLARYMKEVHAPILSIWGVKIVVIAIF 803
R ED R+D C+K+ +K L + K+++AP L V+ V+A+F
Sbjct: 753 SRQEDNRLDIVCCVKLQDRE-------EVKKDSFLFLFFKKIYAPFLLKDWVRPFVVAVF 805
Query: 804 VAFALASIALSTRIEPGLEQEIVLPRDSYLQGYFNNVSEYLRIGPPLYFVVKN-YNYSSE 862
V SIA+ ++E GL+Q++ +P DSY+ YF N+SEYL G P+YFVV+ NYSS
Sbjct: 806 VGMLSFSIAVVDKVEIGLDQKLSMPDDSYVLDYFKNLSEYLHTGAPVYFVVEEGLNYSSL 865
Query: 863 STHTNQLCSISQCNSDSLLNEISKASLVPETSYIAKPAASWLDDFLVWISPEAFGCCRKF 922
N +C C++DSL+ ++ ASL+ S IA +SWLDD+ W+ P + CCR +
Sbjct: 866 EGQ-NAVCGGVGCSNDSLVQQVYYASLISNYSTIANTPSSWLDDYFDWVKPRS-SCCRYY 923
Query: 923 T-NGSYCPPDDQPPCCAPEDDSCVSGACKDCTTCFRHSDLRNDRTSTMQFRDKLPWFLSA 981
G++C + S V+ +C C R M+F LP FLS
Sbjct: 924 NGTGAFC------------NASVVNSSCVHCRPMTPSGMQRPVGDDFMRF---LPMFLSD 968
Query: 982 LPSADCAKGGHGAYTSSVDLKGYDSGIIQASSFRTYHTPLNKQVDYVNSMRAAREFSSKV 1041
P+ C KGGH AY+++VDL ++G + A+ F TYHT + + D++ ++ AR +S +
Sbjct: 969 NPNVKCGKGGHAAYSTAVDLYPGNTG-VGATYFMTYHTIMKESPDFIKALERARSLASNI 1027
Query: 1042 SDSL 1045
+ ++
Sbjct: 1028 TQAV 1031
Score = 90.5 bits (223), Expect = 3e-16
Identities = 47/98 (47%), Positives = 68/98 (68%), Gaps = 1/98 (1%)
Query: 1025 VDYVNSMRAAREFSSKVSDSLKVASGDKDQ-RVKEALGTMGASVFSGITLTKLVGVIVLY 1083
V+ V S + EF S + + ++ K R +EAL MG+SVFSGITLTK G+++L
Sbjct: 1112 VNLVMSCGISVEFCSHIVRAFSISMKKKKVGRAEEALAHMGSSVFSGITLTKFGGILILA 1171
Query: 1084 FSRTEVFVIYYFQMYLSLVLLGFLHGLVFLPVVLSIFG 1121
S++++F ++YF+MYL++VLLG HGLVFLPV+LS G
Sbjct: 1172 LSKSQIFQVFYFRMYLAIVLLGAAHGLVFLPVLLSYIG 1209
>UniRef100_Q7YU59 RE56428p [Drosophila melanogaster]
Length = 1287
Score = 522 bits (1344), Expect = e-146
Identities = 360/1155 (31%), Positives = 571/1155 (49%), Gaps = 142/1155 (12%)
Query: 2 YDICGTRSDGKVLNCPFGSPAVKPD----DLLSSKIQSMCPTITGNVCCTKAQFDTLQTQ 57
Y +C T NCP+ A + +LL + + CC K Q + L
Sbjct: 36 YGVCNTNDFSHSQNCPYNGTAKEMATDGLELLKKRCGFLLENSENKFCCDKNQVELLNKN 95
Query: 58 VQQAIPFLVGCPACLRNFLNLFCELTCSPNQSLFINVTSVDK-AGGNSTVGGIDYFVSDA 116
V+ A L CP+C+ N + C+ TCSP Q+ F++V + K G+ + +D +S
Sbjct: 96 VELAGNILDRCPSCMENLVRHICQFTCSPKQAEFMHVVATQKNKKGDEYISSVDLHISTE 155
Query: 117 FGEGLYESCKDVKFGSMNSRAIQFI----GAGAQNFKEWFAFIGRKAAPNSPGSPYAIMF 172
+ Y+SC V A + A N +WF F+G P P I
Sbjct: 156 YINKTYKSCSQVSVPQTGQLAFDLMCGAYSASRCNPTKWFNFMGDATNPYVPFQITYIQH 215
Query: 173 RPNATKSSGMKPMNVSAYSC----SDTSLGCSCGDCPSSSVCSNSASTTINKANSCSIKV 228
P + S+ P+NV+ C S CSC DC S C K+
Sbjct: 216 EPKSN-SNNFTPLNVTTVPCNQAVSSKLPACSCSDCDLS--CPQGPPEPPRPE---PFKI 269
Query: 229 GSLTVKCVDFILAVLYIILICVFLGWALYHRIRERKMTYRTEPVSNVISGGVLYARNQEK 288
L V I+A ++++ + VFL G L+ +
Sbjct: 270 VGLDAYFV--IMAAVFLVGVLVFL------------------------MGSFLFTQGSSM 303
Query: 289 DENLPMQ---MIEDVPQNRNGVRLSVV----QGYMSNFYRKYGSLVARHPINVLALTLAI 341
D+N + + +++P + N + + ++ F+ K+G+ A +P L ++
Sbjct: 304 DDNFQVDGNDVSDEMPYSENDSYFEKLGAHTETFLETFFTKWGTYFASNPGLTLIAGASL 363
Query: 342 VLLLCLGLIRFKVETRPEKLWVGPGSKAAQEKQFFDSHLAPFYRIEQLILATV--PDHMN 399
V++L G+ ++ T P KLW P SK+ E++FFD+ +PFYR+EQ+I+ V P ++
Sbjct: 364 VVILGYGINFIEITTDPVKLWASPNSKSRLEREFFDTKFSPFYRLEQIIIKAVNLPQIVH 423
Query: 400 STSPR------IVSADNIRFLFEVQKKVDAIRANYSGLMVSLQDICMKPLDKD------- 446
+TS + + + + ++Q+ + I AN + L+DIC PL D
Sbjct: 424 NTSNGPYTFGPVFDREFLTKVLDLQEGIKEINANGT----QLKDICYAPLSDDGSEIDVS 479
Query: 447 -CATQSVLQYF-----KMDPRNFDDSGAVEHLNYCFQQYSSADQCMSAFKAPLDPSTVLG 500
C QS+ YF ++D + D+ V +L+ + S+ C++ + P+DP+ LG
Sbjct: 480 QCVVQSIWGYFGDDRERLDDHDEDNGFNVTYLDALYDCISNPYLCLAPYGGPVDPAIALG 539
Query: 501 GF--SGKDYSG------ASAFIVTYPVNNAIDEEGNETAKAVAWEKAFIQLVKDELLPMA 552
GF G +G A+A I+T+ V N ++ E A + WEK F++ + +
Sbjct: 540 GFLPPGDQLTGSTKFELANAIILTFLVKNHHNKTDLENA--LTWEKKFVEFMTN-YTKNN 596
Query: 553 QSRNLTLAFSSESSIEEELKRESTADAITILVSYLVMFAYISLTLGDTPHPSSFYISSKV 612
S+ + +AF+SE SIE+EL RES +D +TILVSYL+MF YI+++LG +I SK+
Sbjct: 597 MSQYMDIAFTSERSIEDELNRESQSDVLTILVSYLIMFMYIAISLGHVKEFKRVFIDSKI 656
Query: 613 LLGLSGVILVMLSVLGSVAIFSALGVKSTLIIMEVIPFLVLAVGVDNMCILVHA---VKR 669
LG+ GVI+V+ SV+ SV +F +G+ +TLII+EVIPFLVLAVGVDN+ ILV +R
Sbjct: 657 TLGIGGVIIVLASVVSSVGVFGYIGLPATLIIVEVIPFLVLAVGVDNIFILVQTHQRDQR 716
Query: 670 QPLELPLEGRISNALVEVGPSITLASLSEVLAFAVGSFISMPACRVFSMFAALAVLLDFV 729
+P E LE ++ L +VGPSI L SLSE F +G MPA R F+++A +A+++DF+
Sbjct: 717 KPNE-TLEQQVGRILGKVGPSILLTSLSESFCFFLGGLSDMPAVRAFALYAGVALIIDFL 775
Query: 730 LQVTAFVALIVLDSQRAEDKRVDCFPCIKVHSFHADPDKGIRQRKPGLLARYMKEVHAPI 789
LQ+T FV+L LD++R E+ R+D IK PD GLL ++ V+ P
Sbjct: 776 LQITCFVSLFTLDTKRREENRMDICCFIK----GKKPDS--ITSNEGLLYKFFSSVYVPF 829
Query: 790 LSIWGVKIVVIAIFVAFALASIALSTRIEPGLEQEIVLPRDSYLQGYFNNVSEYLRIGPP 849
L V+ V+ IF A+ SIA++ RI+ GL+QE+ +P+DS++ YF ++ E L IGPP
Sbjct: 830 LMKKIVRASVMVIFFAWLCFSIAIAPRIDIGLDQELAMPQDSFVLHYFQSLDENLNIGPP 889
Query: 850 LYFVVKNYNYSSESTHTNQLCSISQCNSDSLLNEISKASLVPETSYIAKPAASWLDDFLV 909
+YFV+K + S+ N +C+ CN DS+L +I AS +YIA+PA+SW+DD+
Sbjct: 890 VYFVLKGDLAYTNSSDQNLVCAGQYCNDDSVLTQIYLASRHSNQTYIARPASSWIDDYFD 949
Query: 910 WISPEAFGCCRKFTNGSYCPPDDQPPCCAPEDDSCVSGACKDCTTCFRHSDLRND--RTS 967
W + + C + +G +CP D T+C R + +N R
Sbjct: 950 WAAAASSCCKYRKDSGDFCPHQD--------------------TSCLRCNITKNSLLRPE 989
Query: 968 TMQFRDKLPWFLSALPSADCAKGGHGAYTSSVDLKGYDSGI-IQASSFRTYHTPLNKQVD 1026
+F LP+FL P CAK GH AY +V + I+AS F YHT L D
Sbjct: 990 EKEFVKYLPFFLKDNPDDTCAKAGHAAYGGAVRYSNSHERLNIEASYFMAYHTILKSSAD 1049
Query: 1027 YVNSMRAAREFSSKVSDSLKVASGDKDQRVKEALGTMGASVFSGITLTKLVGVIVLYFSR 1086
Y ++ +AR+ S+ ++ L+ G + +G+ + + V V +S
Sbjct: 1050 YFLALESARKISANITQMLQ-----------------GRLMSNGVPMASALTVEVFPYS- 1091
Query: 1087 TEVFVIYYFQMYLSL 1101
VF ++Y + YL++
Sbjct: 1092 --VFYVFY-EQYLTM 1103
Score = 89.0 bits (219), Expect = 8e-16
Identities = 43/100 (43%), Positives = 69/100 (69%), Gaps = 1/100 (1%)
Query: 1025 VDYVNSMRAAREFSSKVSDSLKVA-SGDKDQRVKEALGTMGASVFSGITLTKLVGVIVLY 1083
V+ V ++ + EF S + S + S + R ++L MG+S+FSGITLTK G++VL
Sbjct: 1164 VNLVMAVGISVEFCSHLVHSFATSKSVSQIDRAADSLSKMGSSIFSGITLTKFAGILVLA 1223
Query: 1084 FSRTEVFVIYYFQMYLSLVLLGFLHGLVFLPVVLSIFGPP 1123
F+++++F ++YF+MYL +V++G HGL+FLPV+LS G P
Sbjct: 1224 FAKSQIFQVFYFRMYLGIVVIGAAHGLIFLPVLLSYIGAP 1263
>UniRef100_UPI000042D349 UPI000042D349 UniRef100 entry
Length = 1256
Score = 521 bits (1343), Expect = e-146
Identities = 334/1059 (31%), Positives = 549/1059 (51%), Gaps = 74/1059 (6%)
Query: 2 YDICGTRSD-GKVLNCPFGSPAVKPDDLLSSKIQSMCPTITGNVCCTKAQFDTLQTQVQQ 60
Y CG +S GK L C PAVK K++S+C +CC+ Q D L++ +++
Sbjct: 42 YGNCGKKSVFGKPLPCAEFVPAVKASQESREKLKSICGKDFDYICCSPEQIDILESNLKR 101
Query: 61 AIPFLVGCPACLRNFLNLFCELTCSPNQSLFINVTSVDKAG--GNSTVGGIDYFVSDAFG 118
P + CPAC +NF + FC+ +CSPN+S F+ + + A G V I+ +V
Sbjct: 102 VDPLISSCPACRKNFYDFFCQFSCSPNESQFVEIIKTETARDTGKEIVTEINQYVEPGMA 161
Query: 119 EGLYESCKDVKFGSMNSRAIQFIGAGAQNFKEWFAFIGRKAAPNSPGSPYAIMFRPNATK 178
++SCK+VKF + N A+ IG GA+N+ ++ F+G + P GSPY I F +
Sbjct: 162 NQFFDSCKNVKFSATNGYAMDLIGGGAKNYSQFLKFLGDEK-PLLGGSPYQINFVYKLPE 220
Query: 179 S-SGMKPMNVSAYSCSDTSLGCSCGDCPSSSVCSNSASTTINKANSCSIKVGSLTVKCVD 237
+ SG+ N C+D C+C DC S A K C++ V + C
Sbjct: 221 TDSGLVLRNEPLRDCNDKEYKCACTDCEESCPKLPHAKDLTKK---CTVGV----LPCFS 273
Query: 238 FILAVLYIILICVFLGWALY----HRIRERKMTYRTEPVSNVISGGVLYARNQEKDENLP 293
F + +++ +I + G+ +Y + R R + +E + + + YA +K
Sbjct: 274 FSIIIIWSCMIVLLGGYHVYLAKLKKERRRSIAEDSEDDESTMINPLFYAGLGKKRAK-- 331
Query: 294 MQMIEDVPQNRNGVRLSVVQGYMSNFYRKYGSLVARHPINVLALTLAIVLLLCLGLIRFK 353
Q ++ S +Q + +N G ++ P + +LA+V+LL LGL + +
Sbjct: 332 -QFSSEIG--------SKIQDWFANI----GYFCSKFPGISIGTSLAVVVLLSLGLFKLQ 378
Query: 354 VETRPEKLWVGPGSKAAQEKQFFDSHLAPFYRIEQLILATVPDHMNSTSPRIVSADNIRF 413
+ET P KLWV P A + +Q+F+S+ ++RIEQ+I+++ D +++ D +++
Sbjct: 379 LETDPVKLWVSPNDPAYKNQQYFESNFGEWFRIEQVIVSSKDDGP------VLNWDIVKW 432
Query: 414 LFEVQKKVDAIRANYSGLMVSLQDICMKPLDKDCATQSVLQYFKMDPRNFDDSGAVEHLN 473
F+ + +++ + N V L DIC KPLD+ CA QS QYF+ D ++ L
Sbjct: 433 WFDKESQLETLNEN-----VRLSDICFKPLDETCALQSFTQYFQGDISGLTETNWKSKLQ 487
Query: 474 YCFQQYSSADQCMSAFKAPLDPSTVLGGFSGKDYSGASAFIVTYPVNNAIDEEGNETAKA 533
C S C+ F+ PL P+ + F D S A AF VT VN+ E N T+
Sbjct: 488 SCVD---SPVNCLPTFQQPLKPNIL---FDSNDISQAKAFTVTVLVNSDTQNE-NYTSNT 540
Query: 534 VAWEKAFIQLVKDELLPMAQSRNLTLAFSSESSIEEELKRESTADAITILVSYLVMFAYI 593
+++E +F + D + NL +A+S+E S++EEL + S D TI +SYLVMF Y
Sbjct: 541 ISYEHSFQKWAADL---QTEYPNLNIAYSTEISLKEELNQSSNTDIKTIAISYLVMFIYA 597
Query: 594 SLTLGDTPHPSSFY--ISSKVLLGLSGVILVMLSVLGSVAIFSALGVKSTLIIMEVIPFL 651
SL LG ++ Y + ++ LG S +I+++LSV SV FS +G++STLII EVIPFL
Sbjct: 598 SLALGGKLPSANLYSLVKTRFTLGFSSIIIILLSVTASVGFFSIIGLRSTLIIAEVIPFL 657
Query: 652 VLAVGVDNMCILVH---AVKRQPLELPLEGRISNALVEVGPSITLASLSEVLAFAVGSFI 708
VLA+G+DN+ ++VH + L LE RIS AL +GPS ++++ +V F + + +
Sbjct: 658 VLAIGIDNIFLIVHELHVISEGNPNLALEVRISQALKHIGPSCFISAVLQVCMFLLATSV 717
Query: 709 SMPACRVFSMFAALAVLLDFVLQVTAFVALIVLDSQRAEDKRVDCFPCIKVHSFHADPDK 768
MPA + F+ + A AVL++F LQ+T F+ L+ LD +R ED RVD P + + +
Sbjct: 718 GMPAVKNFAYYGAGAVLINFSLQMTCFIGLLALDQRRLEDNRVDYVPWVTISPIQLQDND 777
Query: 769 GIRQ--RKPGLLARYMKEVHAPILSIWGVKIVVIAIFVAFALASIALSTRIEPGLEQEIV 826
I + +R++ + +AP L K VI +FV + S++L +I+ GL+Q I
Sbjct: 778 EIDEPVHLEYNFSRWIGDHYAPFLLKKTTKPKVITLFVLWVGISLSLFPKIQLGLDQRIA 837
Query: 827 LPRDSYLQGYFNNVSEYLRIGPPLYFVVKNYNYSSESTHTNQLCSISQCNSDSLLNEISK 886
+P SYL YFN+V +YL +GPP++FVVK+ +YS S S C+ SL N + +
Sbjct: 838 IPSKSYLVNYFNSVYDYLNVGPPVFFVVKDLDYSERSNQQKICGGFSACDEFSLANILEQ 897
Query: 887 ASLVPETSYIAKPAASWLDDFLVWISPEAFGCCRKFTNGSYCPPDDQPPCCAPEDDSCVS 946
+ S +++PA++WLDDF W++P+ CCR + + + P C+P +
Sbjct: 898 EFKRSDISMLSEPASNWLDDFFSWLNPDLDQCCRFKKSTVF---EKTPEFCSP------N 948
Query: 947 GACKDCTTCF-RHSDLRNDRTSTMQFRDKLPWFLSAL--PSADCAKGGHGAYTSSVDLKG 1003
+ C +C+ H+ + RD + +F + PS C GG A+ ++
Sbjct: 949 APQRQCQSCYLNHNPPYDSSMKAFPERDFMFYFNDWIQEPSDPCPLGGKAAHGQAI---S 1005
Query: 1004 YDSGIIQASSFRTYHTPLNKQVDYVNSMRAAREFSSKVS 1042
+ I +S FRT PL Q +++N+ ++ +++
Sbjct: 1006 RTTEKIDSSYFRTSFAPLRGQDEFINAYKSGNNIVKEIT 1044
Score = 78.2 bits (191), Expect = 1e-12
Identities = 43/111 (38%), Positives = 69/111 (61%), Gaps = 8/111 (7%)
Query: 1015 RTYHTPLNKQVD-------YVNSMRAAREFSSKVSDSLKVASGDKDQRVKEALGTMGASV 1067
R Y P K D Y N + A E +++ S + + ++ + AL ++G S+
Sbjct: 1146 RAYCVPKVKMFDNPAEEELYNNLVNAEPE-NTRRSSITSLNAEFRNTKAHNALCSVGGSL 1204
Query: 1068 FSGITLTKLVGVIVLYFSRTEVFVIYYFQMYLSLVLLGFLHGLVFLPVVLS 1118
SG+TLTKL+G+ VL F+R+++F +YYF+M+LSLV++ F+H V LPV+LS
Sbjct: 1205 ISGVTLTKLIGISVLAFTRSQIFEVYYFRMWLSLVVISFVHAFVLLPVLLS 1255
>UniRef100_Q9VL24 CG5722-PA [Drosophila melanogaster]
Length = 1287
Score = 521 bits (1342), Expect = e-146
Identities = 359/1155 (31%), Positives = 572/1155 (49%), Gaps = 142/1155 (12%)
Query: 2 YDICGTRSDGKVLNCPFGSPAVKPD----DLLSSKIQSMCPTITGNVCCTKAQFDTLQTQ 57
Y +C T NCP+ A + +LL + + CC K Q + L
Sbjct: 36 YGVCNTNDFSHSQNCPYNGTAKEMATDGLELLKKRCGFLLENSENKFCCDKNQVELLNKN 95
Query: 58 VQQAIPFLVGCPACLRNFLNLFCELTCSPNQSLFINVTSVDK-AGGNSTVGGIDYFVSDA 116
V+ A L CP+C+ N + C+ TCSP Q+ F++V + K G+ + +D +S
Sbjct: 96 VELAGNILDRCPSCMENLVRHICQFTCSPKQAEFMHVVATQKNKKGDEYISSVDLHISTE 155
Query: 117 FGEGLYESCKDVKFGSMNSRAIQFI----GAGAQNFKEWFAFIGRKAAPNSPGSPYAIMF 172
+ Y+SC V A + A N +WF F+G P P I
Sbjct: 156 YINKTYKSCSQVSVPQTGQLAFDLMCGAYSASRCNPTKWFNFMGDATNPYVPFQITYIQH 215
Query: 173 RPNATKSSGMKPMNVSAYSC----SDTSLGCSCGDCPSSSVCSNSASTTINKANSCSIKV 228
P + S+ P+NV+ C S CSC DC S C K+
Sbjct: 216 EPKSN-SNNFTPLNVTTVPCNQAVSSKLPACSCSDCDLS--CPQGPPEPPRPE---PFKI 269
Query: 229 GSLTVKCVDFILAVLYIILICVFLGWALYHRIRERKMTYRTEPVSNVISGGVLYARNQEK 288
L V I+A ++++ + VFL G L+ +
Sbjct: 270 VGLDAYFV--IMAAVFLVGVLVFL------------------------MGSFLFTQGSSM 303
Query: 289 DENLPMQ---MIEDVPQNRNGVRLSVV----QGYMSNFYRKYGSLVARHPINVLALTLAI 341
D+N + + +++P + N + + ++ F+ K+G+ A +P L ++
Sbjct: 304 DDNFQVDGNDVSDEMPYSENDSYFEKLGAHTETFLETFFTKWGTYFASNPGLTLIAGASL 363
Query: 342 VLLLCLGLIRFKVETRPEKLWVGPGSKAAQEKQFFDSHLAPFYRIEQLILATV--PDHMN 399
V++L G+ ++ T P KLW P SK+ E++FFD+ +PFYR+EQ+I+ V P ++
Sbjct: 364 VVILGYGINFIEITTDPVKLWASPNSKSRLEREFFDTKFSPFYRLEQIIIKAVNLPQIVH 423
Query: 400 STSPR------IVSADNIRFLFEVQKKVDAIRANYSGLMVSLQDICMKPLDKD------- 446
+TS + + + + ++Q+ + I AN + L+DIC PL D
Sbjct: 424 NTSNGPYTFGPVFDREFLTKVLDLQEGIKEINANGT----QLKDICYAPLSDDGSEIDVS 479
Query: 447 -CATQSVLQYF-----KMDPRNFDDSGAVEHLNYCFQQYSSADQCMSAFKAPLDPSTVLG 500
C QS+ YF ++D + D+ V +L+ + S+ C++ + P+DP+ LG
Sbjct: 480 QCVVQSIWGYFGDDRERLDDHDEDNGFNVTYLDALYDCISNPYLCLAPYGGPVDPAIALG 539
Query: 501 GF--SGKDYSG------ASAFIVTYPVNNAIDEEGNETAKAVAWEKAFIQLVKDELLPMA 552
GF G +G A+A I+T+ V N ++ E A + WEK F++ + +
Sbjct: 540 GFLPPGDQLTGSTKFELANAIILTFLVKNHHNKTDLENA--LTWEKKFVEFMTN-YTKNN 596
Query: 553 QSRNLTLAFSSESSIEEELKRESTADAITILVSYLVMFAYISLTLGDTPHPSSFYISSKV 612
S+ + +AF+SE SIE+EL RES +D +TILVSYL+MF YI+++LG +I SK+
Sbjct: 597 MSQYMDIAFTSERSIEDELNRESQSDVLTILVSYLIMFMYIAISLGHVKEFKRVFIDSKI 656
Query: 613 LLGLSGVILVMLSVLGSVAIFSALGVKSTLIIMEVIPFLVLAVGVDNMCILVHA---VKR 669
LG+ GVI+V+ SV+ SV +F +G+ +TLII+EVIPFLVLAVGVDN+ ILV +R
Sbjct: 657 TLGIGGVIIVLASVVSSVGVFGYIGLPATLIIVEVIPFLVLAVGVDNIFILVQTHQRDQR 716
Query: 670 QPLELPLEGRISNALVEVGPSITLASLSEVLAFAVGSFISMPACRVFSMFAALAVLLDFV 729
+P E LE ++ L +VGPS+ L SLSE F +G MPA R F+++A +A+++DF+
Sbjct: 717 KPNE-TLEQQVGRILGKVGPSMLLTSLSESFCFFLGGLSDMPAVRAFALYAGVALIIDFL 775
Query: 730 LQVTAFVALIVLDSQRAEDKRVDCFPCIKVHSFHADPDKGIRQRKPGLLARYMKEVHAPI 789
LQ+T FV+L LD++R E+ R+D IK PD GLL ++ V+ P
Sbjct: 776 LQITCFVSLFTLDTKRREENRMDICCFIK----GKKPDS--ITSNEGLLYKFFSSVYVPF 829
Query: 790 LSIWGVKIVVIAIFVAFALASIALSTRIEPGLEQEIVLPRDSYLQGYFNNVSEYLRIGPP 849
L V+ V+ IF A+ SIA++ RI+ GL+QE+ +P+DS++ YF +++E L IGPP
Sbjct: 830 LMKKIVRASVMVIFFAWLCFSIAIAPRIDIGLDQELAMPQDSFVLHYFQSLNENLNIGPP 889
Query: 850 LYFVVKNYNYSSESTHTNQLCSISQCNSDSLLNEISKASLVPETSYIAKPAASWLDDFLV 909
+YFV+K + S+ N +C+ CN DS+L +I AS +YIA+PA+SW+DD+
Sbjct: 890 VYFVLKGDLAYTNSSDQNLVCAGQYCNDDSVLTQIYLASRHSNQTYIARPASSWIDDYFD 949
Query: 910 WISPEAFGCCRKFTNGSYCPPDDQPPCCAPEDDSCVSGACKDCTTCFRHSDLRND--RTS 967
W + + C + +G +CP D T+C R + +N R
Sbjct: 950 WAAAASSCCKYRKDSGDFCPHQD--------------------TSCLRCNITKNSLLRPE 989
Query: 968 TMQFRDKLPWFLSALPSADCAKGGHGAYTSSVDLKGYDSGI-IQASSFRTYHTPLNKQVD 1026
+F LP+FL P CAK GH AY +V + I+AS F YHT L D
Sbjct: 990 EKEFVKYLPFFLKDNPDDTCAKAGHAAYGGAVRYSNSHERLNIEASYFMAYHTILKSSAD 1049
Query: 1027 YVNSMRAAREFSSKVSDSLKVASGDKDQRVKEALGTMGASVFSGITLTKLVGVIVLYFSR 1086
Y ++ +AR+ S+ ++ L+ G + +G+ + + V V +S
Sbjct: 1050 YFLALESARKISANITQMLQ-----------------GRLMSNGVPMASALTVEVFPYS- 1091
Query: 1087 TEVFVIYYFQMYLSL 1101
VF ++Y + YL++
Sbjct: 1092 --VFYVFY-EQYLTM 1103
Score = 89.0 bits (219), Expect = 8e-16
Identities = 43/100 (43%), Positives = 69/100 (69%), Gaps = 1/100 (1%)
Query: 1025 VDYVNSMRAAREFSSKVSDSLKVA-SGDKDQRVKEALGTMGASVFSGITLTKLVGVIVLY 1083
V+ V ++ + EF S + S + S + R ++L MG+S+FSGITLTK G++VL
Sbjct: 1164 VNLVMAVGISVEFCSHLVHSFATSKSVSQIDRAADSLSKMGSSIFSGITLTKFAGILVLA 1223
Query: 1084 FSRTEVFVIYYFQMYLSLVLLGFLHGLVFLPVVLSIFGPP 1123
F+++++F ++YF+MYL +V++G HGL+FLPV+LS G P
Sbjct: 1224 FAKSQIFQVFYFRMYLGIVVIGAAHGLIFLPVLLSYIGAP 1263
>UniRef100_Q9U5W1 NPC1 protein [Drosophila melanogaster]
Length = 1287
Score = 521 bits (1341), Expect = e-146
Identities = 359/1155 (31%), Positives = 572/1155 (49%), Gaps = 142/1155 (12%)
Query: 2 YDICGTRSDGKVLNCPFGSPAVKPD----DLLSSKIQSMCPTITGNVCCTKAQFDTLQTQ 57
Y +C T NCP+ A + +LL + + CC K Q + L
Sbjct: 36 YGVCNTNDFSHSQNCPYNGTAKEMATDGLELLKKRCGFLLENSENKFCCDKNQVELLNKN 95
Query: 58 VQQAIPFLVGCPACLRNFLNLFCELTCSPNQSLFINVTSVDK-AGGNSTVGGIDYFVSDA 116
V+ A L CP+C+ N + C+ TCSP Q+ F++V + K G+ + +D +S
Sbjct: 96 VELAGNILDRCPSCMENLVRHICQFTCSPKQAEFMHVVATQKNKKGDEYISSVDLHISTE 155
Query: 117 FGEGLYESCKDVKFGSMNSRAIQFI----GAGAQNFKEWFAFIGRKAAPNSPGSPYAIMF 172
+ Y+SC V A + A N +WF F+G P P I
Sbjct: 156 YINKTYKSCSQVSVPQTGQLAFDLMCGAYSASRCNPTKWFNFMGDATNPYVPFQITYIQH 215
Query: 173 RPNATKSSGMKPMNVSAYSC----SDTSLGCSCGDCPSSSVCSNSASTTINKANSCSIKV 228
P + S+ P+NV+ C S CSC DC S C K+
Sbjct: 216 EPKSN-SNNFTPLNVTTVPCNQAVSSKLPACSCSDCDLS--CPQGPPEPPRPE---PFKI 269
Query: 229 GSLTVKCVDFILAVLYIILICVFLGWALYHRIRERKMTYRTEPVSNVISGGVLYARNQEK 288
L V I+A ++++ + VFL G L+ +
Sbjct: 270 VGLDAYFV--IMAAVFLVGVLVFL------------------------MGSFLFTQGSSM 303
Query: 289 DENLPMQ---MIEDVPQNRNGVRLSVV----QGYMSNFYRKYGSLVARHPINVLALTLAI 341
D+N + + +++P + N + + ++ F+ K+G+ A +P L ++
Sbjct: 304 DDNFQVDGNDVSDEMPYSENDSYFEKLGAHTETFLETFFTKWGTYFASNPGLTLIAGASL 363
Query: 342 VLLLCLGLIRFKVETRPEKLWVGPGSKAAQEKQFFDSHLAPFYRIEQLILATV--PDHMN 399
V++L G+ ++ T P KLW P SK+ E++FFD+ +PFYR+EQ+I+ V P ++
Sbjct: 364 VVILGYGINFIEITTDPVKLWASPNSKSRLEREFFDTKFSPFYRLEQIIIKAVNLPQIVH 423
Query: 400 STSPR------IVSADNIRFLFEVQKKVDAIRANYSGLMVSLQDICMKPLDKD------- 446
+TS + + + + ++Q+ + I AN + L+DIC PL D
Sbjct: 424 NTSNGPYTFGPVFDREFLTKVLDLQEGIKEINANGT----QLKDICYAPLSDDGSEIDVS 479
Query: 447 -CATQSVLQYF-----KMDPRNFDDSGAVEHLNYCFQQYSSADQCMSAFKAPLDPSTVLG 500
C QS+ YF ++D + D+ V +L+ + S+ C++ + P+DP+ LG
Sbjct: 480 QCVVQSIWGYFGDDRERLDDHDEDNGFNVTYLDALYDCISNPYLCLAPYGGPVDPAIALG 539
Query: 501 GF--SGKDYSG------ASAFIVTYPVNNAIDEEGNETAKAVAWEKAFIQLVKDELLPMA 552
GF G +G A+A I+T+ V N ++ E A + WEK F++ + +
Sbjct: 540 GFLPPGDQLTGSTKFELANAIILTFLVKNHHNKTDLENA--LTWEKKFVEFMTN-YTKNN 596
Query: 553 QSRNLTLAFSSESSIEEELKRESTADAITILVSYLVMFAYISLTLGDTPHPSSFYISSKV 612
S+ + +AF+SE SIE+EL RES +D +TILVSYL+MF YI+++LG +I SK+
Sbjct: 597 MSQYMDIAFTSERSIEDELTRESQSDVLTILVSYLIMFMYIAISLGHVKEFKRVFIDSKI 656
Query: 613 LLGLSGVILVMLSVLGSVAIFSALGVKSTLIIMEVIPFLVLAVGVDNMCILVHA---VKR 669
LG+ GVI+V+ SV+ SV +F +G+ +TLII+EVIPFLVLAVGVDN+ ILV +R
Sbjct: 657 TLGIGGVIIVLASVVSSVGVFGYIGLPATLIIVEVIPFLVLAVGVDNIFILVQTHQRDQR 716
Query: 670 QPLELPLEGRISNALVEVGPSITLASLSEVLAFAVGSFISMPACRVFSMFAALAVLLDFV 729
+P E LE ++ L +VGPS+ L SLSE F +G MPA R F+++A +A+++DF+
Sbjct: 717 KPNE-TLEQQVGRILGKVGPSMLLTSLSESFCFFLGGLSDMPAVRAFALYAGVALIIDFL 775
Query: 730 LQVTAFVALIVLDSQRAEDKRVDCFPCIKVHSFHADPDKGIRQRKPGLLARYMKEVHAPI 789
LQ+T FV+L LD++R E+ R+D IK PD GLL ++ V+ P
Sbjct: 776 LQITCFVSLFTLDTKRREENRMDICCFIK----GKKPDS--ITSNEGLLYKFFSSVYVPF 829
Query: 790 LSIWGVKIVVIAIFVAFALASIALSTRIEPGLEQEIVLPRDSYLQGYFNNVSEYLRIGPP 849
L V+ V+ IF A+ SIA++ RI+ GL+QE+ +P+DS++ YF +++E L IGPP
Sbjct: 830 LMKKIVRASVMVIFFAWLCFSIAIAPRIDIGLDQELAMPQDSFVLHYFQSLNENLNIGPP 889
Query: 850 LYFVVKNYNYSSESTHTNQLCSISQCNSDSLLNEISKASLVPETSYIAKPAASWLDDFLV 909
+YFV+K + S+ N +C+ CN DS+L +I AS +YIA+PA+SW+DD+
Sbjct: 890 VYFVLKGDLAYTNSSDQNLVCAGQYCNDDSVLTQIYLASRHSNQTYIARPASSWIDDYFD 949
Query: 910 WISPEAFGCCRKFTNGSYCPPDDQPPCCAPEDDSCVSGACKDCTTCFRHSDLRND--RTS 967
W + + C + +G +CP D T+C R + +N R
Sbjct: 950 WAAAASSCCKYRKDSGDFCPHQD--------------------TSCLRCNITKNSLLRPE 989
Query: 968 TMQFRDKLPWFLSALPSADCAKGGHGAYTSSVDLKGYDSGI-IQASSFRTYHTPLNKQVD 1026
+F LP+FL P CAK GH AY +V + I+AS F YHT L D
Sbjct: 990 EKEFVKYLPFFLKDNPDDTCAKAGHAAYGGAVRYSNSHERLNIEASYFMAYHTILKSSAD 1049
Query: 1027 YVNSMRAAREFSSKVSDSLKVASGDKDQRVKEALGTMGASVFSGITLTKLVGVIVLYFSR 1086
Y ++ +AR+ S+ ++ L+ G + +G+ + + V V +S
Sbjct: 1050 YFLALESARKISANITQMLQ-----------------GRLMSNGVPMASALTVEVFPYS- 1091
Query: 1087 TEVFVIYYFQMYLSL 1101
VF ++Y + YL++
Sbjct: 1092 --VFYVFY-EQYLTM 1103
Score = 89.0 bits (219), Expect = 8e-16
Identities = 43/100 (43%), Positives = 69/100 (69%), Gaps = 1/100 (1%)
Query: 1025 VDYVNSMRAAREFSSKVSDSLKVA-SGDKDQRVKEALGTMGASVFSGITLTKLVGVIVLY 1083
V+ V ++ + EF S + S + S + R ++L MG+S+FSGITLTK G++VL
Sbjct: 1164 VNLVMAVGISVEFCSHLVHSFATSKSVSQIDRAADSLSKMGSSIFSGITLTKFAGILVLA 1223
Query: 1084 FSRTEVFVIYYFQMYLSLVLLGFLHGLVFLPVVLSIFGPP 1123
F+++++F ++YF+MYL +V++G HGL+FLPV+LS G P
Sbjct: 1224 FAKSQIFQVFYFRMYLGIVVIGAAHGLIFLPVLLSYIGAP 1263
>UniRef100_Q6T3U3 Niemann-Pick C1-like 1 [Rattus norvegicus]
Length = 1331
Score = 514 bits (1324), Expect = e-144
Identities = 382/1172 (32%), Positives = 575/1172 (48%), Gaps = 134/1172 (11%)
Query: 14 LNCPFGSPAVKPDDLLSSKIQSMCPTI-----TGNVCCTKAQFDTLQTQVQQAIPFLVGC 68
++C +PA + +Q +CP + T CC+ Q +L++ + L C
Sbjct: 54 VSCLSNTPARHVTGEHLALLQRICPRLYNGPNTTFACCSTKQLLSLESSMSITKALLTRC 113
Query: 69 PACLRNFLNLFCELTCSPNQSLFINVTSVDKAGGNSTVGGIDY--FVSDAFGEGLYESCK 126
PAC NF++L C TCSP+QSLFINVT V + G + Y F +F E YESC
Sbjct: 114 PACSDNFVSLHCHNTCSPDQSLFINVTRVVERGAGEPPAVVAYEAFYQRSFAEKAYESCS 173
Query: 127 DVKFGSMNSRAIQFI----GAGAQNFKEWFAFIGRKAAPNSPGSPYAIMFRPNATKSSGM 182
V+ + S A+ + G+ N + W F G +P + P G+
Sbjct: 174 QVRIPAAASLAVGSMCGVYGSALCNAQRWLNFQGDTGNGLAPLDITFHLLEPGQALPDGI 233
Query: 183 KPMNVSAYSCS----DTSLGCSCGDCPSSSVCSNSASTTINKANSCSIKVGSLTVKCVDF 238
+P+N C+ D S CSC DC +S +A S +G +
Sbjct: 234 QPLNGKIAPCNESQGDDSAVCSCQDCAASCPVIPPP-----EALRPSFYMGRMPGWLALI 288
Query: 239 ILAVLYIILICVFLGWALYHRIRERKMTYRTEPVSNVISGGVLYARNQEKDENLPMQMIE 298
I+ +L+ L R VSN RN+ K E +
Sbjct: 289 IIFTAVFVLLSAVL--------------VRLRVVSN---------RNKNKAEGP-----Q 320
Query: 299 DVPQNRNGVRLSVVQGYMSNFYRKYGSLVARHPINVLALTLAIVLLLCLGLIRFKVETRP 358
+ P+ + +LS + F++ +G+ VA P+ VLAL+ +V+ L GL ++ T P
Sbjct: 321 EAPKLPHKHKLSP-HTILGRFFQNWGTRVASWPLTVLALSFIVVIALAAGLTFIELTTDP 379
Query: 359 EKLWVGPGSKAAQEKQFFDSHLAPFYRIEQLILATVPDHMNSTSPR-------------I 405
+LW P S+A +EK F D H PF+R Q+ + N +S + I
Sbjct: 380 VELWSAPKSQARKEKSFHDEHFGPFFRTNQIFVTA----RNRSSYKYDSLLLGSKNFSGI 435
Query: 406 VSADNIRFLFEVQKKVDAIR--ANYSGLMVSLQDICMKPLD------KDCATQSVLQYFK 457
+S D + L E+Q+++ ++ + + +SLQDIC PL+ DC S+LQYF+
Sbjct: 436 LSLDFLLELLELQERLRHLQVWSPEAERNISLQDICYAPLNPYNTSLSDCCVNSLLQYFQ 495
Query: 458 ------MDPRNFDDSGAV------EHLNYC------FQQYSS-ADQCMSAFKAPLDPSTV 498
M N +G +H YC F+ +S A CM+ + AP+ P
Sbjct: 496 NNRTLLMLTANQTLNGQTSLVDWKDHFLYCANAPLTFKDGTSLALSCMADYGAPVFPFLA 555
Query: 499 LGGFSGKDYSGASAFIVTYPVNNAIDEEGNETAKAVAWEKAFIQLVKDELLPMAQSRNLT 558
+GG+ G DYS A A I+T+ +NN + A+A WE+AF++ + E S
Sbjct: 556 VGGYQGTDYSEAEALIITFSLNN-YPADDPRMAQAKLWEEAFLKEM--ESFQRNTSDKFQ 612
Query: 559 LAFSSESSIEEELKRESTADAITILVSYLVMFAYISLTLGDTPHPSSFYISSKVLLGLSG 618
+AFS+E S+E+E+ R + D VSY+++F YISL LG S + SK LGL G
Sbjct: 613 VAFSAERSLEDEINRTTIQDLPVFAVSYIIVFLYISLALGSYSRCSRVAVESKATLGLGG 672
Query: 619 VILVMLSVLGSVAIFSALGVKSTLIIMEVIPFLVLAVGVDNMCILVHAVKRQPLELPLEG 678
VI+V+ +VL ++ +S LGV S+L+I++V+PFLVLAVG DN+ I V +R P +P E
Sbjct: 673 VIVVLGAVLAAMGFYSYLGVPSSLVIIQVVPFLVLAVGADNIFIFVLEYQRLP-RMPGEQ 731
Query: 679 R---ISNALVEVGPSITLASLSEVLAFAVGSFISMPACRVFSMFAALAVLLDFVLQVTAF 735
R I L V PS+ L SLSE + F +G+ MPA R F++ + LA++LDF+LQ+TAF
Sbjct: 732 REAHIGRTLGSVAPSMLLCSLSEAICFFLGALTPMPAVRTFALTSGLAIILDFLLQMTAF 791
Query: 736 VALIVLDSQRAEDKRVDCFPCIKVHSFHADPDKGIRQRKPGLLARYMKEVHAPILSIWGV 795
VAL+ LDS+R E R D C P K K GLL R+ ++++AP L +
Sbjct: 792 VALLSLDSKRQEASRPDVLCCFSTRKL--PPPK----EKEGLLLRFFRKIYAPFLLHRFI 845
Query: 796 KIVVIAIFVAFALASIALSTRIEPGLEQEIVLPRDSYLQGYFNNVSEYLRIGPPLYFV-V 854
+ VV+ +F+ A++ L I GL+QE+ LP+DSYL YF ++ YL +GPP+YFV
Sbjct: 846 RPVVMLLFLTLFGANLYLMCNINVGLDQELALPKDSYLIDYFLFLNRYLEVGPPVYFVTT 905
Query: 855 KNYNYSSESTHTNQLCSISQCNSDSLLNEISKASLVPETSYIAKPAASWLDDFLVWISPE 914
+N+SSE+ N CS + C S SL +I AS P+ SY+A A+SW+DDF+ W++P
Sbjct: 906 SGFNFSSEA-GMNATCSSAGCKSFSLTQKIQYASEFPDQSYVAIAASSWVDDFIDWLTPS 964
Query: 915 AFGCCRKFTNG----SYCPPDDQPPCCAPEDDSCVSGACKDCTTCFRHSDLRNDRTSTMQ 970
+ CCR + G +CP D C K+C + L R + Q
Sbjct: 965 S-SCCRLYIRGPHKDEFCPSTDTSFNC-----------LKNC----MNRTLGPVRPTAEQ 1008
Query: 971 FRDKLPWFLSALPSADCAKGGHGAYTSSVDLKGYDSGIIQASSFRTYHTPLNKQVDYVNS 1030
F LPWFL+ P+ C KGG AY +SV+L G + AS F YH PL D+ +
Sbjct: 1009 FHKYLPWFLNDPPNIRCPKGGLAAYRTSVNLS--SDGQVIASQFMAYHKPLRNSQDFTEA 1066
Query: 1031 MRAAREFSSKVSDSLKVASGDKDQRVKEALGTMGASVFSGITLTKL-VGVIVLYFSRTEV 1089
+RA+R ++ ++ L+ G E ++VF LT L G+ L
Sbjct: 1067 LRASRLLAANITADLRKVPGTDPN--FEVFPYTISNVFYQQYLTVLPEGIFTLALCFVPT 1124
Query: 1090 FVIYYFQMYLSLVLLGFLHGLVFLPVVLSIFG 1121
FV+ Y + L + G L+ L + +++ G
Sbjct: 1125 FVVCYLLLGLDM-CSGILNLLSIIMILVDTIG 1155
Score = 81.6 bits (200), Expect = 1e-13
Identities = 40/116 (34%), Positives = 75/116 (64%), Gaps = 1/116 (0%)
Query: 1025 VDYVNSMRAAREFSSKVSDSLKVASGD-KDQRVKEALGTMGASVFSGITLTKLVGVIVLY 1083
++ V ++ + EF S ++ S V++ + +R K+A MG++VF+G+ +T G+++L
Sbjct: 1170 INLVTAVGMSVEFVSHITRSFAVSTKPTRLERAKDATVFMGSAVFAGVAMTNFPGILILG 1229
Query: 1084 FSRTEVFVIYYFQMYLSLVLLGFLHGLVFLPVVLSIFGPPSRCTIIEQEENRSSTS 1139
F++ ++ I++F++ L + LLG LHGLVFLPVVLS GP ++++E+ S +
Sbjct: 1230 FAQAQLIQIFFFRLNLLITLLGLLHGLVFLPVVLSYLGPDVNQALVQEEKLASEAA 1285
>UniRef100_Q6T3U4 Niemann-Pick C1-like 1 [Mus musculus]
Length = 1333
Score = 509 bits (1312), Expect = e-142
Identities = 374/1174 (31%), Positives = 579/1174 (48%), Gaps = 138/1174 (11%)
Query: 14 LNCPFGSPA--VKPDDLLSSKIQSMCPTITGN-----VCCTKAQFDTLQTQVQQAIPFLV 66
++C +PA V D L + +Q +CP + CC+ Q +L + + L
Sbjct: 54 ISCLSNTPARHVTGDHL--ALLQRVCPRLYNGPNDTYACCSTKQLVSLDSSLSITKALLT 111
Query: 67 GCPACLRNFLNLFCELTCSPNQSLFINVTSVDKA--GGNSTVGGIDYFVSDAFGEGLYES 124
CPAC NF+++ C TCSP+QSLFINVT V + G V + F +F E YES
Sbjct: 112 RCPACSENFVSIHCHNTCSPDQSLFINVTRVVQRDPGQLPAVVAYEAFYQRSFAEKAYES 171
Query: 125 CKDVKFGSMNSRAIQFI----GAGAQNFKEWFAFIGRKAAPNSPGSPYAIMFRPNATKSS 180
C V+ + S A+ + G+ N + W F G +P + P +
Sbjct: 172 CSRVRIPAAASLAVGSMCGVYGSALCNAQRWLNFQGDTGNGLAPLDITFHLLEPGQALAD 231
Query: 181 GMKPMNVSAYSCSDT----SLGCSCGDCPSSSVCSNSASTTINKANSCSIKVGSLTVKCV 236
GMKP++ C+++ S CSC DC +S A S +G +
Sbjct: 232 GMKPLDGKITPCNESQGEDSAACSCQDCAASCPVIPPPP-----ALRPSFYMGRMPGWLA 286
Query: 237 DFILAVLYIILICVFLGWALYHRIRERKMTYRTEPVSNVISGGVLYARNQEKDENLPMQM 296
I+ +L+ V L +Y R+ A N+ K++ Q
Sbjct: 287 LIIIFTAVFVLLSVVL---VYLRV----------------------ASNRNKNKTAGSQE 321
Query: 297 IEDVPQNRNGVRLSVVQGYMSNFYRKYGSLVARHPINVLALTLAIVLLLCLGLIRFKVET 356
++P+ R +V + F+ +G+ VA P+ VLAL+ +V+ L +GL ++ T
Sbjct: 322 APNLPRKRRFSPHTV----LGRFFESWGTRVASWPLTVLALSFIVVIALSVGLTFIELTT 377
Query: 357 RPEKLWVGPGSKAAQEKQFFDSHLAPFYRIEQLILATVPDHMNSTSPR------------ 404
P +LW P S+A +EK F D H PF+R Q+ + N +S +
Sbjct: 378 DPVELWSAPKSQARKEKAFHDEHFGPFFRTNQIFVTA----KNRSSYKYDSLLLGPKNFS 433
Query: 405 -IVSADNIRFLFEVQKKVDAIR--ANYSGLMVSLQDICMKPLDK------DCATQSVLQY 455
I+S D ++ L E+Q+++ ++ ++ + +SLQDIC PL+ DC S+LQY
Sbjct: 434 GILSLDLLQELLELQERLRHLQVWSHEAQRNISLQDICYAPLNPHNTSLTDCCVNSLLQY 493
Query: 456 FKMD------PRNFDDSGAV------EHLNYCFQQ-------YSSADQCMSAFKAPLDPS 496
F+ + N +G +H YC + A C++ + AP+ P
Sbjct: 494 FQNNHTLLLLTANQTLNGQTSLVDWKDHFLYCANAPLTYKDGTALALSCIADYGAPVFPF 553
Query: 497 TVLGGFSGKDYSGASAFIVTYPVNNAIDEEGNETAKAVAWEKAFIQLVKDELLPMAQSRN 556
+GG+ G DYS A A I+T+ +NN + A A WE+AF++ ++ + +
Sbjct: 554 LAVGGYQGTDYSEAEALIITFSINN-YPADDPRMAHAKLWEEAFLKEMQS--FQRSTADK 610
Query: 557 LTLAFSSESSIEEELKRESTADAITILVSYLVMFAYISLTLGDTPHPSSFYISSKVLLGL 616
+AFS+E S+E+E+ R + D +SYL++F YISL LG S + SK LGL
Sbjct: 611 FQIAFSAERSLEDEINRTTIQDLPVFAISYLIVFLYISLALGSYSRWSRVAVDSKATLGL 670
Query: 617 SGVILVMLSVLGSVAIFSALGVKSTLIIMEVIPFLVLAVGVDNMCILVHAVKRQPLELPL 676
GV +V+ +V+ ++ +S LGV S+L+I++V+PFLVLAVG DN+ I V +R P +P
Sbjct: 671 GGVAVVLGAVVAAMGFYSYLGVPSSLVIIQVVPFLVLAVGADNIFIFVLEYQRLP-RMPG 729
Query: 677 EGR---ISNALVEVGPSITLASLSEVLAFAVGSFISMPACRVFSMFAALAVLLDFVLQVT 733
E R I L V PS+ L SLSE + F +G+ SMPA R F++ + LA++ DF+LQ+T
Sbjct: 730 EQREAHIGRTLGSVAPSMLLCSLSEAICFFLGALTSMPAVRTFALTSGLAIIFDFLLQMT 789
Query: 734 AFVALIVLDSQRAEDKRVDCFPCIKVHSFHADPDKGIRQRKPGLLARYMKEVHAPILSIW 793
AFVAL+ LDS+R E R D C S + P K +K GLL + ++++ P L
Sbjct: 790 AFVALLSLDSKRQEASRPDVVCCFS--SRNLPPPK----QKEGLLLCFFRKIYTPFLLHR 843
Query: 794 GVKIVVIAIFVAFALASIALSTRIEPGLEQEIVLPRDSYLQGYFNNVSEYLRIGPPLYF- 852
++ VV+ +F+ A++ L I GL+Q++ LP+DSYL YF ++ YL +GPP+YF
Sbjct: 844 FIRPVVLLLFLVLFGANLYLMCNISVGLDQDLALPKDSYLIDYFLFLNRYLEVGPPVYFD 903
Query: 853 VVKNYNYSSESTHTNQLCSISQCNSDSLLNEISKASLVPETSYIAKPAASWLDDFLVWIS 912
YN+S+E+ N +CS + C S SL +I AS P SY+A A+SW+DDF+ W++
Sbjct: 904 TTSGYNFSTEAG-MNAICSSAGCESFSLTQKIQYASEFPNQSYVAIAASSWVDDFIDWLT 962
Query: 913 PEAFGCCRKFTNG----SYCPPDDQPPCCAPEDDSCVSGACKDCTTCFRHSDLRNDRTST 968
P + CCR +T G +CP D C K+C + L R +T
Sbjct: 963 PSS-SCCRIYTRGPHKDEFCPSTDTSFNC-----------LKNC----MNRTLGPVRPTT 1006
Query: 969 MQFRDKLPWFLSALPSADCAKGGHGAYTSSVDLKGYDSGIIQASSFRTYHTPLNKQVDYV 1028
QF LPWFL+ P+ C KGG AY +SV+L G I AS F YH PL D+
Sbjct: 1007 EQFHKYLPWFLNDTPNIRCPKGGLAAYRTSVNLS--SDGQIIASQFMAYHKPLRNSQDFT 1064
Query: 1029 NSMRAAREFSSKVSDSLKVASGDKDQRVKEALGTMGASVFSGITLTKL-VGVIVLYFSRT 1087
++RA+R ++ ++ L+ G E ++VF LT L G+ L
Sbjct: 1065 EALRASRLLAANITAELRKVPGTDPN--FEVFPYTISNVFYQQYLTVLPEGIFTLALCFV 1122
Query: 1088 EVFVIYYFQMYLSLVLLGFLHGLVFLPVVLSIFG 1121
FV+ Y + L + G L+ L + +++ G
Sbjct: 1123 PTFVVCYLLLGLD-IRSGILNLLSIIMILVDTIG 1155
Score = 79.0 bits (193), Expect = 8e-13
Identities = 39/110 (35%), Positives = 72/110 (65%), Gaps = 1/110 (0%)
Query: 1025 VDYVNSMRAAREFSSKVSDSLKVASGD-KDQRVKEALGTMGASVFSGITLTKLVGVIVLY 1083
++ V ++ + EF S ++ S V++ + +R K+A MG++VF+G+ +T G+++L
Sbjct: 1170 INLVTAVGMSVEFVSHITRSFAVSTKPTRLERAKDATIFMGSAVFAGVAMTNFPGILILG 1229
Query: 1084 FSRTEVFVIYYFQMYLSLVLLGFLHGLVFLPVVLSIFGPPSRCTIIEQEE 1133
F++ ++ I++F++ L + LLG LHGLVFLPVVLS GP ++ +E+
Sbjct: 1230 FAQAQLIQIFFFRLNLLITLLGLLHGLVFLPVVLSYLGPDVNQALVLEEK 1279
>UniRef100_Q5SVX1 Ortholog of human NPC1 (Niemann-Pick disease, type C1, gene)-like 1
NPC1L1 [Mus musculus]
Length = 1333
Score = 509 bits (1310), Expect = e-142
Identities = 374/1174 (31%), Positives = 578/1174 (48%), Gaps = 138/1174 (11%)
Query: 14 LNCPFGSPA--VKPDDLLSSKIQSMCPTITGN-----VCCTKAQFDTLQTQVQQAIPFLV 66
++C +PA V D L + +Q +CP + CC+ Q +L + + L
Sbjct: 54 ISCLSNTPARHVTGDHL--ALLQRVCPRLYNGPNDTYACCSTKQLVSLDSSLSITKALLT 111
Query: 67 GCPACLRNFLNLFCELTCSPNQSLFINVTSVDKA--GGNSTVGGIDYFVSDAFGEGLYES 124
CPAC NF+++ C TCSP+QSLFINVT V + G V + F +F E YES
Sbjct: 112 RCPACSENFVSIHCHNTCSPDQSLFINVTRVVQRDPGQLPAVVAYEAFYQRSFAEKAYES 171
Query: 125 CKDVKFGSMNSRAIQFI----GAGAQNFKEWFAFIGRKAAPNSPGSPYAIMFRPNATKSS 180
C V+ + S A+ + G+ N + W F G +P + P +
Sbjct: 172 CSRVRIPAAASLAVGSMCGVYGSALCNAQRWLNFQGDTGNGLAPLDITFHLLEPGQALAD 231
Query: 181 GMKPMNVSAYSCSDT----SLGCSCGDCPSSSVCSNSASTTINKANSCSIKVGSLTVKCV 236
GMKP++ C+++ S CSC DC +S A S +G +
Sbjct: 232 GMKPLDGKITPCNESQGEDSAACSCQDCAASCPVIPPPP-----ALRPSFYMGRMPGWLA 286
Query: 237 DFILAVLYIILICVFLGWALYHRIRERKMTYRTEPVSNVISGGVLYARNQEKDENLPMQM 296
I+ +L+ V L +Y R+ A N+ K++ Q
Sbjct: 287 LIIIFTAVFVLLSVVL---VYLRV----------------------ASNRNKNKTAGSQE 321
Query: 297 IEDVPQNRNGVRLSVVQGYMSNFYRKYGSLVARHPINVLALTLAIVLLLCLGLIRFKVET 356
++P+ R +V + F+ +G+ VA P+ VLAL+ +V+ L +GL ++ T
Sbjct: 322 APNLPRKRRFSPHTV----LGRFFESWGTRVASWPLTVLALSFIVVIALSVGLTFIELTT 377
Query: 357 RPEKLWVGPGSKAAQEKQFFDSHLAPFYRIEQLILATVPDHMNSTSPR------------ 404
P +LW P S+A +EK F D H PF+R Q+ + N +S +
Sbjct: 378 DPVELWSAPKSQARKEKAFHDEHFGPFFRTNQIFVTA----KNRSSYKYDSLLLGPKNFS 433
Query: 405 -IVSADNIRFLFEVQKKVDAIR--ANYSGLMVSLQDICMKPLDK------DCATQSVLQY 455
I+S D ++ L E+Q+++ ++ ++ + +SLQDIC PL DC S+LQY
Sbjct: 434 GILSLDLLQELLELQERLRHLQVWSHEAQRNISLQDICYAPLKPHNTSLTDCCVNSLLQY 493
Query: 456 FKMD------PRNFDDSGAV------EHLNYCFQQ-------YSSADQCMSAFKAPLDPS 496
F+ + N +G +H YC + A C++ + AP+ P
Sbjct: 494 FQNNHTLLLLTANQTLNGQTSLVDWKDHFLYCANAPLTYKDGTALALSCIADYGAPVFPF 553
Query: 497 TVLGGFSGKDYSGASAFIVTYPVNNAIDEEGNETAKAVAWEKAFIQLVKDELLPMAQSRN 556
+GG+ G DYS A A I+T+ +NN + A A WE+AF++ ++ + +
Sbjct: 554 LAVGGYQGTDYSEAEALIITFSINN-YPADDPRMAHAKLWEEAFLKEMQS--FQRSTADK 610
Query: 557 LTLAFSSESSIEEELKRESTADAITILVSYLVMFAYISLTLGDTPHPSSFYISSKVLLGL 616
+AFS+E S+E+E+ R + D +SYL++F YISL LG S + SK LGL
Sbjct: 611 FQIAFSAERSLEDEINRTTIQDLPVFAISYLIVFLYISLALGSYSRWSRVAVDSKATLGL 670
Query: 617 SGVILVMLSVLGSVAIFSALGVKSTLIIMEVIPFLVLAVGVDNMCILVHAVKRQPLELPL 676
GV +V+ +V+ ++ +S LGV S+L+I++V+PFLVLAVG DN+ I V +R P +P
Sbjct: 671 GGVAVVLGAVVAAMGFYSYLGVPSSLVIIQVVPFLVLAVGADNIFIFVLEYQRLP-RMPG 729
Query: 677 EGR---ISNALVEVGPSITLASLSEVLAFAVGSFISMPACRVFSMFAALAVLLDFVLQVT 733
E R I L V PS+ L SLSE + F +G+ SMPA R F++ + LA++ DF+LQ+T
Sbjct: 730 EQREAHIGRTLGSVAPSMLLCSLSEAICFFLGALTSMPAVRTFALTSGLAIIFDFLLQMT 789
Query: 734 AFVALIVLDSQRAEDKRVDCFPCIKVHSFHADPDKGIRQRKPGLLARYMKEVHAPILSIW 793
AFVAL+ LDS+R E R D C S + P K +K GLL + ++++ P L
Sbjct: 790 AFVALLSLDSKRQEASRPDVVCCFS--SRNLPPPK----QKEGLLLCFFRKIYTPFLLHR 843
Query: 794 GVKIVVIAIFVAFALASIALSTRIEPGLEQEIVLPRDSYLQGYFNNVSEYLRIGPPLYF- 852
++ VV+ +F+ A++ L I GL+Q++ LP+DSYL YF ++ YL +GPP+YF
Sbjct: 844 FIRPVVLLLFLVLFGANLYLMCNISVGLDQDLALPKDSYLIDYFLFLNRYLEVGPPVYFD 903
Query: 853 VVKNYNYSSESTHTNQLCSISQCNSDSLLNEISKASLVPETSYIAKPAASWLDDFLVWIS 912
YN+S+E+ N +CS + C S SL +I AS P SY+A A+SW+DDF+ W++
Sbjct: 904 TTSGYNFSTEAG-MNAICSSAGCESFSLTQKIQYASEFPNQSYVAIAASSWVDDFIDWLT 962
Query: 913 PEAFGCCRKFTNG----SYCPPDDQPPCCAPEDDSCVSGACKDCTTCFRHSDLRNDRTST 968
P + CCR +T G +CP D C K+C + L R +T
Sbjct: 963 PSS-SCCRIYTRGPHKDEFCPSTDTSFNC-----------LKNC----MNRTLGPVRPTT 1006
Query: 969 MQFRDKLPWFLSALPSADCAKGGHGAYTSSVDLKGYDSGIIQASSFRTYHTPLNKQVDYV 1028
QF LPWFL+ P+ C KGG AY +SV+L G I AS F YH PL D+
Sbjct: 1007 EQFHKYLPWFLNDTPNIRCPKGGLAAYRTSVNLS--SDGQIIASQFMAYHKPLRNSQDFT 1064
Query: 1029 NSMRAAREFSSKVSDSLKVASGDKDQRVKEALGTMGASVFSGITLTKL-VGVIVLYFSRT 1087
++RA+R ++ ++ L+ G E ++VF LT L G+ L
Sbjct: 1065 EALRASRLLAANITAELRKVPGTDPN--FEVFPYTISNVFYQQYLTVLPEGIFTLALCFV 1122
Query: 1088 EVFVIYYFQMYLSLVLLGFLHGLVFLPVVLSIFG 1121
FV+ Y + L + G L+ L + +++ G
Sbjct: 1123 PTFVVCYLLLGLD-IRSGILNLLSIIMILVDTIG 1155
Score = 79.0 bits (193), Expect = 8e-13
Identities = 39/110 (35%), Positives = 72/110 (65%), Gaps = 1/110 (0%)
Query: 1025 VDYVNSMRAAREFSSKVSDSLKVASGD-KDQRVKEALGTMGASVFSGITLTKLVGVIVLY 1083
++ V ++ + EF S ++ S V++ + +R K+A MG++VF+G+ +T G+++L
Sbjct: 1170 INLVTAVGMSVEFVSHITRSFAVSTKPTRLERAKDATIFMGSAVFAGVAMTNFPGILILG 1229
Query: 1084 FSRTEVFVIYYFQMYLSLVLLGFLHGLVFLPVVLSIFGPPSRCTIIEQEE 1133
F++ ++ I++F++ L + LLG LHGLVFLPVVLS GP ++ +E+
Sbjct: 1230 FAQAQLIQIFFFRLNLLITLLGLLHGLVFLPVVLSYLGPDVNQALVLEEK 1279
>UniRef100_Q6CBA1 Similar to tr|Q12200 Saccharomyces cerevisiae YPL006W NCR1 [Yarrowia
lipolytica]
Length = 1239
Score = 501 bits (1290), Expect = e-140
Identities = 333/1079 (30%), Positives = 549/1079 (50%), Gaps = 113/1079 (10%)
Query: 1 MYDICGTRS-DGKVLNCPFGSPAVKPDDLLSSK-----IQSMCPTITGN---VCCTKAQF 51
+Y CG +S G L CP +P K SK + +C + + +CC AQ
Sbjct: 29 IYGNCGKKSLFGSELPCP--TPQDKAGPFEPSKGDLDLLGEICGEVWTHEKLLCCDTAQI 86
Query: 52 DTLQTQVQQAIPFLVGCPACLRNFLNLFCELTCSPNQSLFINVTSVDKA-GGNSTVGGID 110
L+ +++A + CPAC NF FC+ TC+P+Q ++NVT + K+ G V +
Sbjct: 87 KDLKNSLKKADGLISSCPACKSNFYEFFCKFTCAPDQRDYVNVTQLGKSTDGRDIVTELS 146
Query: 111 YFVSDAFGEGLYESCKDVKFGSMNSRAIQFIGAGAQNFKEWFAFIGRKAAPNSPGSPYAI 170
Y+V + +G Y+SCKD+KF + N A+ +G G +K + F+G + P GSP+ I
Sbjct: 147 YYVKPEWAQGFYDSCKDIKFSATNGYAMDLLGGGRHGYKNFLKFLGDEK-PMLGGSPFQI 205
Query: 171 MFR-PNATKSS-----GMKPMNVSAYSCSDTSLGCSCGDCPSSSVCSNSASTTINKANSC 224
F P K G+KP+ C+ + C+C DC + C + I K +C
Sbjct: 206 NFPWPEDEKDKNYPPKGIKPVEPVLRDCATGAYKCACSDCEGA--CPELPN--IKKHGAC 261
Query: 225 SIKVGSLTVKCVDFILAVLY-IILICVFLGWALYHRIRERKMTYRTEPVSNVISGGVLYA 283
KVG L C+ F + ++Y ++L+ G H + SG L
Sbjct: 262 --KVGHLP--CLSFAVIIIYSVVLLGAIAGSWFVHS-------------HTLFSGATL-- 302
Query: 284 RNQEKDENLPMQMIEDVPQNRNGVRLSVVQGYMSNFYRKYGSLVARHPINVLALTLAIVL 343
+D+ + + ++ P + ++ + + A +P + T+ + +
Sbjct: 303 ----EDQYMTSRWYQN-PHASEKYTTYPINRFLQYWVSRIALFCASYPAVTIGFTVTLSI 357
Query: 344 LLCLGLIRFKVETRPEKLWVGPGSKAAQEKQFFDSHLAPFYRIEQLILATVPDHMNSTSP 403
L+ +GL RF+VE P +LWV S +K+ FD++ PFYR Q+ L +T+
Sbjct: 358 LMSVGLTRFQVEENPVRLWVSEDSTPYLQKENFDNNFGPFYRTAQVYLV-------NTTG 410
Query: 404 RIVSADNIRFLFEVQKKVDAIRANYSGLMVSLQDICMKPLDKDCATQSVLQYFKMDPRNF 463
+V+ +N+ + V++++ +++ + S D C+KP + C QS QY N
Sbjct: 411 SVVNQENLVWWQGVEQQIISLQVEGT----SFNDFCLKPTNDACVIQSYTQY----GINL 462
Query: 464 DDSGAVEHLNYCFQQYSSADQCMSAFKAPLDPSTVLGGFS--GKDYSGASAFIVTYPVNN 521
+ L C SSA QC+ F PL+ + + GG++ +D A A ++T
Sbjct: 463 NSPDWATQLQTCT---SSAVQCLPPFGQPLNMNLLFGGYNETSRDPLSAQALVITLVGEG 519
Query: 522 AIDEEGNETAKAVAWEKAFIQLVKDELLPMAQSRNLTLAFSSESSIEEELKRESTADAIT 581
++++ E A+A WEK I ++ D + A R L L+FS+E+S+ EEL + + D
Sbjct: 520 YLEDDPQE-ARAQKWEKGLIDVLLD-VQHEAWRRGLQLSFSTEASLTEELNKSTNTDVKI 577
Query: 582 ILVSYLVMFAYISLTLGDTPHPSSFYISSKVLLGLSGVILVMLSVLGSVAIFSALGVKST 641
+++SYLVMF Y S+ LG + F LGL G+I+V+LSV S I +A+G+K+T
Sbjct: 578 VVISYLVMFLYASMALGGGSGKAKFG------LGLCGIIIVLLSVAASAGICAAIGIKAT 631
Query: 642 LIIMEVIPFLVLAVGVDNMCILVHAVKRQPLELP---LEGRISNALVEVGPSITLASLSE 698
LII EVIPFLVLAVGVDN+ +L H + + P ++ R+S A+ +GPSI +++++E
Sbjct: 632 LIIAEVIPFLVLAVGVDNIFLLCHEMDAANIAYPNDSVDTRVSKAVGRIGPSIVISAITE 691
Query: 699 VLAFAVGSFISMPACRVFSMFAALAVLLDFVLQVTAFVALIVLDSQRAEDKRVDCFPCIK 758
LAF + + + MPA R F+++AA AV ++ +LQ+T FV+++ LD +R P
Sbjct: 692 TLAFGLAATVKMPAVRNFAIYAAGAVFINAILQLTIFVSVMALDQRRQSA------PIQL 745
Query: 759 VHSFHADPDKGIRQRKPGLLARYMKEVHAPILSIWGVKIVVIAIFVAFALASIALSTRIE 818
S+ D D I + + L +R ++ +AP L K +V+A+F +A S+ L ++
Sbjct: 746 ADSY--DLDFNILEHRENLFSRLIRRYYAPFLLKKKTKKIVLAVFGTWAAVSLILWPMLQ 803
Query: 819 PGLEQEIVLPRDSYLQGYFNNVSEYLRIGPPLYFVVKNYNYSSESTHTNQLCSISQCNSD 878
GL+Q + +P DSYL YF+++ +YL +GPP+YFVV N +S + + + + C
Sbjct: 804 LGLDQRLAVPSDSYLVQYFDDIYDYLNVGPPVYFVVSGLNATSRNGQQSLCGTFTTCEDY 863
Query: 879 SLLNEISKASLVPETSYIAKPAASWLDDFLVWISPEAFGCCR--KFTNGSYCPPDDQPPC 936
SL+N + + PE SYI++PA+SW+DD+L W++P+ CCR K CPP P
Sbjct: 864 SLVNIVEQERKRPELSYISEPASSWIDDYLKWLNPDLDECCRVKKTDKDVACPPRASP-- 921
Query: 937 CAPEDDSCVSGACKDCTTCFRHSDLRNDRTST-----MQFRDKLPWFLSALPSADCAKGG 991
+ C CF+ D + T + +F +++ A PS C GG
Sbjct: 922 -------------RACNVCFKDRDPAWNITMSGLPQGPEFMHYFDFWIDA-PSDPCPLGG 967
Query: 992 HGAYTSSVDLKGYDSGIIQASSFRTYHTPLNKQVDYVNSMRAAREFSSKVSDSLKVASG 1050
Y+ +V YDS + S FR++HTPL Q D++N++ +A+ S + +L G
Sbjct: 968 KAPYSDAV---VYDSDDVLTSHFRSFHTPLRSQKDFINALASAKRISKDIEKTLGGTEG 1023
Score = 75.1 bits (183), Expect = 1e-11
Identities = 38/89 (42%), Positives = 61/89 (67%), Gaps = 1/89 (1%)
Query: 1034 AREFSSKVSDSLKVA-SGDKDQRVKEALGTMGASVFSGITLTKLVGVIVLYFSRTEVFVI 1092
AR F++ ++S + S K+ R EAL +G SVF GI +TK +GV VL F+R+++F +
Sbjct: 1118 ARAFTTVSANSSHIRMSPTKNTRTFEALVGVGGSVFGGIAMTKFLGVFVLAFTRSKIFEV 1177
Query: 1093 YYFQMYLSLVLLGFLHGLVFLPVVLSIFG 1121
YYF+++L+LV++ H L+ LPV+L+ G
Sbjct: 1178 YYFRVWLALVIVATTHSLILLPVLLTYIG 1206
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.322 0.136 0.406
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,825,932,231
Number of Sequences: 2790947
Number of extensions: 75217472
Number of successful extensions: 199053
Number of sequences better than 10.0: 416
Number of HSP's better than 10.0 without gapping: 284
Number of HSP's successfully gapped in prelim test: 132
Number of HSP's that attempted gapping in prelim test: 197313
Number of HSP's gapped (non-prelim): 941
length of query: 1140
length of database: 848,049,833
effective HSP length: 138
effective length of query: 1002
effective length of database: 462,899,147
effective search space: 463824945294
effective search space used: 463824945294
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 81 (35.8 bits)
Lotus: description of TM0193.4


