

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0192.7
(263 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
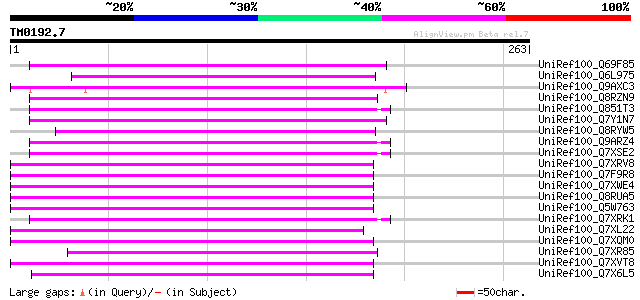
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q69F85 Gag-pol polyprotein [Phaseolus vulgaris] 114 3e-24
UniRef100_Q6L975 GAG-POL [Vitis vinifera] 108 1e-22
UniRef100_Q9AXC3 Hypothetical protein [Antirrhinum hispanicum] 106 7e-22
UniRef100_Q8RZN9 Putative polyprotein [Oryza sativa] 93 6e-18
UniRef100_Q851T3 Putative gag-pol [Oryza sativa] 90 7e-17
UniRef100_Q7Y1N7 Putative polyprotein [Oryza sativa] 89 9e-17
UniRef100_Q8RYW5 OSJNBa0066C06.12 protein [Oryza sativa] 89 1e-16
UniRef100_Q9ARZ4 Putative polyprotein [Oryza sativa] 87 4e-16
UniRef100_Q7XSE2 OSJNBb0033G08.12 protein [Oryza sativa] 87 4e-16
UniRef100_Q7XRV8 OSJNBb0049I21.2 protein [Oryza sativa] 87 4e-16
UniRef100_Q7F9R8 OSJNBb0054B09.2 protein [Oryza sativa] 87 4e-16
UniRef100_Q7XWE4 OSJNBa0035O13.3 protein [Oryza sativa] 87 6e-16
UniRef100_Q8RUA5 Putative gag-pol [Oryza sativa] 86 7e-16
UniRef100_Q5W763 Putative polyprotein [Oryza sativa] 86 7e-16
UniRef100_Q7XRK1 OSJNBa0042D13.9 protein [Oryza sativa] 86 1e-15
UniRef100_Q7XL22 OSJNBa0079M09.7 protein [Oryza sativa] 86 1e-15
UniRef100_Q7XQM0 OSJNBa0089E12.4 protein [Oryza sativa] 86 1e-15
UniRef100_Q7XR85 OSJNBa0011L07.16 protein [Oryza sativa] 86 1e-15
UniRef100_Q7XVT8 OSJNBa0035B13.13 protein [Oryza sativa] 86 1e-15
UniRef100_Q7X6L5 OSJNBb0093G06.3 protein [Oryza sativa] 86 1e-15
>UniRef100_Q69F85 Gag-pol polyprotein [Phaseolus vulgaris]
Length = 1859
Score = 114 bits (285), Expect = 3e-24
Identities = 51/181 (28%), Positives = 99/181 (54%)
Query: 11 PFSEDVESVAILDNMKALVLDSYSGDSDPKDHLLYFNTKMVIIAASDAVKCRMFPSTFKS 70
PF++D+ + + D + L ++ Y G +DP +HL F T+M + V C++FP++ +
Sbjct: 211 PFTDDIIATPLPDKWRGLTINLYDGSTDPDEHLNIFRTQMTLYTTDRTVWCKVFPTSLRE 270
Query: 71 TAMAWFTTLPRGSISNFRDFSSKFLVQFSANKNQPVTINDLYSIRQQEGESLKEYVARYS 130
+ WF+ LP SI++F KF Q++ ++ + L +++Q+ GESL+ ++ R+S
Sbjct: 271 GPLGWFSDLPPNSIASFDALELKFTTQYATSRPHRTSSMSLLNVKQERGESLRTFMNRFS 330
Query: 131 AASVKVEDEEPRACALAFKNGLLPGGLNSKLTRKPARSMEEMRARASTYILDEEDDAFKR 190
+ + + P + +LPG L ++P +M+E+R RA+ ++ EE + R
Sbjct: 331 KVCMNIRNLNPEIAMHHLVSAILPGRFTESLIKRPPCNMDELRTRATKFMQIEEHIDYHR 390
Query: 191 K 191
K
Sbjct: 391 K 391
>UniRef100_Q6L975 GAG-POL [Vitis vinifera]
Length = 549
Score = 108 bits (270), Expect = 1e-22
Identities = 53/155 (34%), Positives = 90/155 (57%), Gaps = 1/155 (0%)
Query: 32 SYSGDSDPKDHLLYFNTKMVIIAASDAVKCRMFPSTFKSTAMAWFTTLPRGSISNFRDFS 91
+Y G SDP DH++++ M + +DA+ C++FP++ + A++WF LP S+ NFRD S
Sbjct: 26 TYDGTSDPFDHIMHYRQLMTLDIGNDALLCKVFPASLQGQALSWFHRLPPNSVGNFRDLS 85
Query: 92 SKFLVQFSANKNQPVTINDLYSIRQQEGESLKEYVARYSAASVKVEDEEPRACALAFKNG 151
F+ Q+ + I+ L +I+ Q+ ESL+E+V R+ A ++VE A FK
Sbjct: 86 EAFVGQYLCSARHKQNISTLQNIKMQDNESLREFVKRFGQAVLQVEACSMDAVLQIFKRS 145
Query: 152 LLPG-GLNSKLTRKPARSMEEMRARASTYILDEED 185
+ PG L +KP +M+++ RA+ Y + E+D
Sbjct: 146 ICPGTPFFESLAKKPPTTMDDLFRRANKYSMLEDD 180
>UniRef100_Q9AXC3 Hypothetical protein [Antirrhinum hispanicum]
Length = 1455
Score = 106 bits (264), Expect = 7e-22
Identities = 69/214 (32%), Positives = 106/214 (49%), Gaps = 13/214 (6%)
Query: 1 READSVAEF----RPFSEDVESVAILDNMKALVLDSYSGDS---DPKDHLLYFNTKMVII 53
R+ D AE PF+ + + I +M+ Y+GD DP+DH+ F M I+
Sbjct: 142 RDRDGTAEEDEMGSPFAPHILAPEIPVSMRLSNPVVYTGDRNARDPRDHINQFLAVMDIL 201
Query: 54 AASDAVKCRMFPSTFKSTAMAWFTTLPRGSISNFRDFSSKFLVQFSANKNQPVTINDLYS 113
+ D C +F ST A AWF+ LPRGSI +F + F+ QF+ N+ P TI L +
Sbjct: 202 STPDHALCWIFRSTLTGRAQAWFSQLPRGSIRSFDELRELFVRQFACNRRYPKTIGHLMT 261
Query: 114 IRQQEGESLKEYVARYSAASVKVEDEEPRACALAFKNGLLPGGLNSKLTRKPARSMEEMR 173
+ + EGESL+EY+ R++ + V P + G + L +PARS+EE+
Sbjct: 262 MVEAEGESLREYMRRFANTVIDVRHVGPHILSEFVAQGTRSVEFANSLVGRPARSLEELM 321
Query: 174 ARASTYILDEEDDAFK------RKRAKLEKGDTS 201
RA Y+L E+ K R++ +GD +
Sbjct: 322 HRAERYMLQEDTRLAKLGIQGHEDRSRNRRGDNT 355
>UniRef100_Q8RZN9 Putative polyprotein [Oryza sativa]
Length = 2001
Score = 93.2 bits (230), Expect = 6e-18
Identities = 44/176 (25%), Positives = 87/176 (49%)
Query: 11 PFSEDVESVAILDNMKALVLDSYSGDSDPKDHLLYFNTKMVIIAASDAVKCRMFPSTFKS 70
P S++++ N K + Y GD+DP ++L + T + D K ++ P+ +
Sbjct: 342 PLSQNLQMAPWPINFKLSNITKYKGDTDPNEYLRVYETAVEAAGGDDTTKAKILPTMLEG 401
Query: 71 TAMAWFTTLPRGSISNFRDFSSKFLVQFSANKNQPVTINDLYSIRQQEGESLKEYVARYS 130
A++W+TT+P +I ++ F F +P +DLY+++Q GE+L+ ++ ++S
Sbjct: 402 VALSWYTTIPPMTIYSWEHMRDTFRAGFIGAYEEPKEADDLYAMKQLPGETLRSFIVKFS 461
Query: 131 AASVKVEDEEPRACALAFKNGLLPGGLNSKLTRKPARSMEEMRARASTYILDEEDD 186
++ + A K LLPG L L R ++ +++ R ++ EED+
Sbjct: 462 RVRCQIRHVDDEMLIAAAKRALLPGPLRFDLARNRPKTTKDLFERMESFARGEEDE 517
>UniRef100_Q851T3 Putative gag-pol [Oryza sativa]
Length = 971
Score = 89.7 bits (221), Expect = 7e-17
Identities = 53/183 (28%), Positives = 88/183 (47%), Gaps = 2/183 (1%)
Query: 11 PFSEDVESVAILDNMKALVLDSYSGDSDPKDHLLYFNTKMVIIAASDAVKCRMFPSTFKS 70
P S +++ I K L + YSG++DPK+ L + + + + K ++
Sbjct: 357 PLSGGIKNRPIPPQFKFLPVPRYSGETDPKEFLSIYESAIEAAHGDENTKAKVIHLALDG 416
Query: 71 TAMAWFTTLPRGSISNFRDFSSKFLVQFSANKNQPVTINDLYSIRQQEGESLKEYVARYS 130
A +W+ LP SI ++ F++ F +P T L IRQ+ GES++EY+ R+S
Sbjct: 417 IARSWYFNLPANSIYSWEQLRDVFVLNFRGTYEEPKTQQHLLGIRQRPGESIREYMRRFS 476
Query: 131 AASVKVEDEEPRACALAFKNGLLPGGLNSKLTRKPARSMEEMRARASTYILDEEDDAFKR 190
A +VED + A GLL G L K+ K +++E + + EED KR
Sbjct: 477 QARCQVEDITEASVINAASAGLLEGELTRKIANKEPQTLEHLVRIIDGFAKSEEDS--KR 534
Query: 191 KRA 193
++A
Sbjct: 535 RQA 537
>UniRef100_Q7Y1N7 Putative polyprotein [Oryza sativa]
Length = 518
Score = 89.4 bits (220), Expect = 9e-17
Identities = 52/181 (28%), Positives = 85/181 (46%)
Query: 11 PFSEDVESVAILDNMKALVLDSYSGDSDPKDHLLYFNTKMVIIAASDAVKCRMFPSTFKS 70
P S +++ I K + YSG++D K+ L + + + + K ++
Sbjct: 282 PLSWGIKNHPIPPQFKYPPVPQYSGETDSKEFLSIYESAIEAAHGDENTKAKLIHLVLDG 341
Query: 71 TAMAWFTTLPRGSISNFRDFSSKFLVQFSANKNQPVTINDLYSIRQQEGESLKEYVARYS 130
A +W+ LP SI ++ F++ F +P T DL I Q++GES +EY+ R+
Sbjct: 342 IARSWYFNLPVNSIYSWEQLRDVFVLNFRGTYEEPKTQQDLVGIHQRQGESTREYIRRFL 401
Query: 131 AASVKVEDEEPRACALAFKNGLLPGGLNSKLTRKPARSMEEMRARASTYILDEEDDAFKR 190
A +V+D + A GLL G L KL RK R++E + Y DEED +R
Sbjct: 402 QARCQVQDITEASVIHAASAGLLEGELTRKLVRKEPRTLEHLFRIIDGYARDEEDTKRRR 461
Query: 191 K 191
+
Sbjct: 462 E 462
>UniRef100_Q8RYW5 OSJNBa0066C06.12 protein [Oryza sativa]
Length = 359
Score = 88.6 bits (218), Expect = 1e-16
Identities = 43/162 (26%), Positives = 80/162 (48%)
Query: 24 NMKALVLDSYSGDSDPKDHLLYFNTKMVIIAASDAVKCRMFPSTFKSTAMAWFTTLPRGS 83
N K + Y GD DP ++L + T + D VK ++ P+ + A++W+TT+P +
Sbjct: 7 NFKLSNITKYRGDIDPNEYLRVYETIVEETGGDDTVKAKILPTMLEGVALSWYTTIPPMT 66
Query: 84 ISNFRDFSSKFLVQFSANKNQPVTINDLYSIRQQEGESLKEYVARYSAASVKVEDEEPRA 143
I ++ F F +P +DLY+++Q GE+L+ ++ ++S ++ +
Sbjct: 67 IYSWEHMMDTFRAGFVGAYEEPKETDDLYAMKQLPGETLRSFIVKFSRVRCQIRQVDDEM 126
Query: 144 CALAFKNGLLPGGLNSKLTRKPARSMEEMRARASTYILDEED 185
A K LLPG L L R ++ +++ R ++ EED
Sbjct: 127 LIAAAKRALLPGPLRFDLARNIPKTAKDLFERMESFTKGEED 168
>UniRef100_Q9ARZ4 Putative polyprotein [Oryza sativa]
Length = 1579
Score = 87.0 bits (214), Expect = 4e-16
Identities = 51/183 (27%), Positives = 89/183 (47%), Gaps = 2/183 (1%)
Query: 11 PFSEDVESVAILDNMKALVLDSYSGDSDPKDHLLYFNTKMVIIAASDAVKCRMFPSTFKS 70
P S +++ I K + YSG++DPK+ L + + + + K ++ T
Sbjct: 49 PLSGGIKTRPIPPQFKFPPVPCYSGETDPKEFLSIYESAIEAAHGDENTKAKVIHLTLDG 108
Query: 71 TAMAWFTTLPRGSISNFRDFSSKFLVQFSANKNQPVTINDLYSIRQQEGESLKEYVARYS 130
A +W+ LP SI ++ F++ F +P T L IRQ+ GES++EY+ R+S
Sbjct: 109 IARSWYFNLPANSIYSWEQLRDVFVLNFRGTYEEPKTQQHLLGIRQRPGESIREYMRRFS 168
Query: 131 AASVKVEDEEPRACALAFKNGLLPGGLNSKLTRKPARSMEEMRARASTYILDEEDDAFKR 190
A +V+D + A GLL G L K+ K ++++++ + EED KR
Sbjct: 169 QARCQVQDITEASVINAASAGLLEGELTRKIANKEPQTLKQLLRIIDGFARGEEDS--KR 226
Query: 191 KRA 193
++A
Sbjct: 227 RQA 229
>UniRef100_Q7XSE2 OSJNBb0033G08.12 protein [Oryza sativa]
Length = 1847
Score = 87.0 bits (214), Expect = 4e-16
Identities = 51/183 (27%), Positives = 88/183 (47%), Gaps = 2/183 (1%)
Query: 11 PFSEDVESVAILDNMKALVLDSYSGDSDPKDHLLYFNTKMVIIAASDAVKCRMFPSTFKS 70
P S +++ I K + YSG++DPK+ L + + + + K ++
Sbjct: 417 PLSGGIKTRPIPPQFKFPPVPRYSGETDPKEFLSIYESAIEAAHGDENTKAKVIHLALDG 476
Query: 71 TAMAWFTTLPRGSISNFRDFSSKFLVQFSANKNQPVTINDLYSIRQQEGESLKEYVARYS 130
A +W+ LP SI ++ F++ F +P T L IRQ++GES++EY+ R+S
Sbjct: 477 IARSWYFNLPANSIYSWEQLRDVFVLNFRGTYKEPKTQQHLLGIRQKQGESIREYMRRFS 536
Query: 131 AASVKVEDEEPRACALAFKNGLLPGGLNSKLTRKPARSMEEMRARASTYILDEEDDAFKR 190
A +V+D + A GLL G L K+ K +++E + + EED KR
Sbjct: 537 QARCQVQDITKASVINAASAGLLEGELTRKIANKEPQTLEHLLRIIDGFARGEEDS--KR 594
Query: 191 KRA 193
++A
Sbjct: 595 RQA 597
>UniRef100_Q7XRV8 OSJNBb0049I21.2 protein [Oryza sativa]
Length = 1843
Score = 87.0 bits (214), Expect = 4e-16
Identities = 49/184 (26%), Positives = 83/184 (44%)
Query: 1 READSVAEFRPFSEDVESVAILDNMKALVLDSYSGDSDPKDHLLYFNTKMVIIAASDAVK 60
R D + F++D+ V K ++ Y G ++P+ L + +
Sbjct: 416 RHDDDLDGVAAFTDDLRRVDWPAGFKPTGIEKYDGTTNPESWLTVYGLAIRAAGGDSKAM 475
Query: 61 CRMFPSTFKSTAMAWFTTLPRGSISNFRDFSSKFLVQFSANKNQPVTINDLYSIRQQEGE 120
P +A +W LPRG+I ++ + +F+ F +P T DLY++ Q+ GE
Sbjct: 476 ANYLPVALADSARSWLHGLPRGTIGSWAELRDRFIANFQGTFERPGTQYDLYNVIQKSGE 535
Query: 121 SLKEYVARYSAASVKVEDEEPRACALAFKNGLLPGGLNSKLTRKPARSMEEMRARASTYI 180
SL+EY+ R+S K+ D AF G+ L K RKP R+++ M +A+ Y
Sbjct: 536 SLREYIRRFSEQRNKISDITDDVIIAAFTKGIRHEELVGKFGRKPPRTVKLMFEKANEYA 595
Query: 181 LDEE 184
E+
Sbjct: 596 KAED 599
>UniRef100_Q7F9R8 OSJNBb0054B09.2 protein [Oryza sativa]
Length = 1992
Score = 87.0 bits (214), Expect = 4e-16
Identities = 48/184 (26%), Positives = 84/184 (45%)
Query: 1 READSVAEFRPFSEDVESVAILDNMKALVLDSYSGDSDPKDHLLYFNTKMVIIAASDAVK 60
R D + F++D+ V K ++ Y G ++P+ L + + +
Sbjct: 409 RHDDDLDGVAAFTDDLRRVDWPAGFKPTGIEKYDGTTNPESWLTVYGLAIRAVGGDSKAM 468
Query: 61 CRMFPSTFKSTAMAWFTTLPRGSISNFRDFSSKFLVQFSANKNQPVTINDLYSIRQQEGE 120
P + +A +W LPRG+I ++ + F+ F +P T DLY++ Q+ GE
Sbjct: 469 ANYLPVALEDSARSWLHGLPRGTIGSWAELRDHFIANFQGTFERPGTQYDLYNVIQKSGE 528
Query: 121 SLKEYVARYSAASVKVEDEEPRACALAFKNGLLPGGLNSKLTRKPARSMEEMRARASTYI 180
SL++Y+ R+S K+ D AF G+ L K RKP R+++ M +A+ Y
Sbjct: 529 SLRDYIRRFSEQRNKISDITDDVIIAAFTKGIRHEELVGKFGRKPPRTVKLMFEKANEYA 588
Query: 181 LDEE 184
E+
Sbjct: 589 KAED 592
>UniRef100_Q7XWE4 OSJNBa0035O13.3 protein [Oryza sativa]
Length = 2008
Score = 86.7 bits (213), Expect = 6e-16
Identities = 49/184 (26%), Positives = 83/184 (44%)
Query: 1 READSVAEFRPFSEDVESVAILDNMKALVLDSYSGDSDPKDHLLYFNTKMVIIAASDAVK 60
R D + E F++D+ V K ++ Y G ++P L + + +
Sbjct: 410 RHDDDLDEVAAFTDDLRRVDWPAGFKPTGIEKYDGTTNPGSWLTVYGLAIRAAGGDNKAM 469
Query: 61 CRMFPSTFKSTAMAWFTTLPRGSISNFRDFSSKFLVQFSANKNQPVTINDLYSIRQQEGE 120
P +A +W LPRG+I ++ + F+ F +P T DLY++ Q+ GE
Sbjct: 470 ANYLPVALADSARSWLHGLPRGTIGSWAELRDHFIANFQGTFERPGTQYDLYNVIQKSGE 529
Query: 121 SLKEYVARYSAASVKVEDEEPRACALAFKNGLLPGGLNSKLTRKPARSMEEMRARASTYI 180
SL++Y+ R+S K+ D AF G+ L K RKP R+++ M +A+ Y
Sbjct: 530 SLRDYIRRFSEQRNKISDITDDVIIAAFTKGIRHEELVGKFGRKPPRTVKLMFEKANEYA 589
Query: 181 LDEE 184
E+
Sbjct: 590 KAED 593
>UniRef100_Q8RUA5 Putative gag-pol [Oryza sativa]
Length = 1964
Score = 86.3 bits (212), Expect = 7e-16
Identities = 49/184 (26%), Positives = 83/184 (44%)
Query: 1 READSVAEFRPFSEDVESVAILDNMKALVLDSYSGDSDPKDHLLYFNTKMVIIAASDAVK 60
R D + F++D+ V K ++ Y G ++P+ L + + +
Sbjct: 410 RHDDDLDGVAAFTDDLRRVDWPAGFKPTGIEKYDGTTNPESWLTVYGLAIRAAGGDNKAM 469
Query: 61 CRMFPSTFKSTAMAWFTTLPRGSISNFRDFSSKFLVQFSANKNQPVTINDLYSIRQQEGE 120
P +A +W LPRG+I ++ + F+ F +P T DLY++ Q+ GE
Sbjct: 470 ANYLPVALADSARSWLHGLPRGTIGSWAELRDHFIANFQGTFERPGTQYDLYNVIQKSGE 529
Query: 121 SLKEYVARYSAASVKVEDEEPRACALAFKNGLLPGGLNSKLTRKPARSMEEMRARASTYI 180
SL+EY+ R+S K+ D AF G+ L K RKP R+++ M +A+ Y
Sbjct: 530 SLREYIRRFSEQRNKISDITDDVIIAAFTKGIRHEELVGKFGRKPPRTVKLMFEKANEYA 589
Query: 181 LDEE 184
E+
Sbjct: 590 KAED 593
>UniRef100_Q5W763 Putative polyprotein [Oryza sativa]
Length = 1551
Score = 86.3 bits (212), Expect = 7e-16
Identities = 49/184 (26%), Positives = 82/184 (43%)
Query: 1 READSVAEFRPFSEDVESVAILDNMKALVLDSYSGDSDPKDHLLYFNTKMVIIAASDAVK 60
R D V F+ D+ V K ++ Y G ++P+ L + +
Sbjct: 396 RYDDDVDGVAAFTSDLRRVDWPAGFKPTGIEKYEGTTNPESWLTAYGLAIRAAGGDSKAM 455
Query: 61 CRMFPSTFKSTAMAWFTTLPRGSISNFRDFSSKFLVQFSANKNQPVTINDLYSIRQQEGE 120
P +A +W LPRG+I ++ + F+ F +P T DLY+I Q+ GE
Sbjct: 456 ANYLPVALADSAQSWLHGLPRGTIGSWAELRDHFIANFQGTFERPGTQFDLYNIIQKSGE 515
Query: 121 SLKEYVARYSAASVKVEDEEPRACALAFKNGLLPGGLNSKLTRKPARSMEEMRARASTYI 180
SL++Y+ R+S K+ D AF G+ L K RKP +++++M +A+ Y
Sbjct: 516 SLRDYIRRFSEQRNKISDITDNVIIAAFTKGIRHEDLVGKFGRKPPKTVKQMFEKANEYA 575
Query: 181 LDEE 184
E+
Sbjct: 576 KAED 579
>UniRef100_Q7XRK1 OSJNBa0042D13.9 protein [Oryza sativa]
Length = 303
Score = 85.9 bits (211), Expect = 1e-15
Identities = 51/183 (27%), Positives = 87/183 (46%), Gaps = 2/183 (1%)
Query: 11 PFSEDVESVAILDNMKALVLDSYSGDSDPKDHLLYFNTKMVIIAASDAVKCRMFPSTFKS 70
P S +++ I K + YSG++DPK+ L + + + + K ++
Sbjct: 49 PLSGGIKTRPIPPQFKFPPVSRYSGETDPKEFLSIYESAIEAAHGDENTKAKVIHLALDG 108
Query: 71 TAMAWFTTLPRGSISNFRDFSSKFLVQFSANKNQPVTINDLYSIRQQEGESLKEYVARYS 130
A +W+ LP SI ++ F++ F +P T L IRQ+ GES++EY+ R+S
Sbjct: 109 IARSWYFNLPANSIYSWEQLRDVFVLNFRGTYEEPKTQQHLLGIRQRPGESIREYMRRFS 168
Query: 131 AASVKVEDEEPRACALAFKNGLLPGGLNSKLTRKPARSMEEMRARASTYILDEEDDAFKR 190
A +V+D + A GLL G L K+ K +++E + + EED KR
Sbjct: 169 QARCQVQDITEASVINAALAGLLEGELTKKIANKEPQTLEHLLRIIDGFARGEEDS--KR 226
Query: 191 KRA 193
++A
Sbjct: 227 RQA 229
>UniRef100_Q7XL22 OSJNBa0079M09.7 protein [Oryza sativa]
Length = 877
Score = 85.9 bits (211), Expect = 1e-15
Identities = 47/179 (26%), Positives = 81/179 (44%)
Query: 1 READSVAEFRPFSEDVESVAILDNMKALVLDSYSGDSDPKDHLLYFNTKMVIIAASDAVK 60
R D V F+ D+ V K ++ Y G ++P+ L + +
Sbjct: 322 RYDDDVDRVAAFTNDLRRVDWPAGFKPTGIEKYDGTTNPESWLTVYGLAIRAAGGDSKAM 381
Query: 61 CRMFPSTFKSTAMAWFTTLPRGSISNFRDFSSKFLVQFSANKNQPVTINDLYSIRQQEGE 120
P +A +W LPRG+I ++ + F+ F +P T DLY++ Q+ GE
Sbjct: 382 ANYLPVALADSAWSWLHGLPRGTIGSWAELRDHFITNFQGTFERPGTQFDLYNVIQKSGE 441
Query: 121 SLKEYVARYSAASVKVEDEEPRACALAFKNGLLPGGLNSKLTRKPARSMEEMRARASTY 179
SL++Y+ R+S K+ D +AF G+ L K RKP +++++M +A+ Y
Sbjct: 442 SLRDYIRRFSEQRNKISDITDDVIIVAFTKGIRHEDLVGKFGRKPPKTVKQMFEKANEY 500
>UniRef100_Q7XQM0 OSJNBa0089E12.4 protein [Oryza sativa]
Length = 1965
Score = 85.9 bits (211), Expect = 1e-15
Identities = 49/184 (26%), Positives = 84/184 (45%)
Query: 1 READSVAEFRPFSEDVESVAILDNMKALVLDSYSGDSDPKDHLLYFNTKMVIIAASDAVK 60
R D + F++D+ V + K ++ Y G ++P+ L + + +
Sbjct: 415 RYDDDLDGVAAFTDDLRRVDWPASFKPTGIEKYDGTTNPESWLTVYGLAIRAAGGDNKAM 474
Query: 61 CRMFPSTFKSTAMAWFTTLPRGSISNFRDFSSKFLVQFSANKNQPVTINDLYSIRQQEGE 120
P +A +W LPRG+I ++ + F+ F +P T DLY++ Q+ GE
Sbjct: 475 ANYLPVALADSARSWLHGLPRGTIGSWAELRDHFIANFQGTFERPGTQYDLYNVIQKSGE 534
Query: 121 SLKEYVARYSAASVKVEDEEPRACALAFKNGLLPGGLNSKLTRKPARSMEEMRARASTYI 180
SL+EY+ R+S K+ D AF G+ L K RKP R+++ M +A+ Y
Sbjct: 535 SLREYIRRFSEQRNKISDITDDVIIAAFTKGIRHEELVGKFGRKPPRTVKLMFEKANEYA 594
Query: 181 LDEE 184
E+
Sbjct: 595 KAED 598
>UniRef100_Q7XR85 OSJNBa0011L07.16 protein [Oryza sativa]
Length = 393
Score = 85.5 bits (210), Expect = 1e-15
Identities = 38/157 (24%), Positives = 79/157 (50%)
Query: 30 LDSYSGDSDPKDHLLYFNTKMVIIAASDAVKCRMFPSTFKSTAMAWFTTLPRGSISNFRD 89
+ Y GD+DP ++L + T + D K ++ P+ + A++W+TT+P +I ++
Sbjct: 186 IPQYKGDTDPNEYLRVYETAVEAAGGDDTTKAKILPTMLEGVALSWYTTIPPMTIYSWEH 245
Query: 90 FSSKFLVQFSANKNQPVTINDLYSIRQQEGESLKEYVARYSAASVKVEDEEPRACALAFK 149
F F +P ++LY+++Q GE+L+ ++ ++S ++ + + K
Sbjct: 246 MRDTFRAGFIGVYEEPKEADNLYAMKQMPGETLRSFMVKFSRIRCQIRQVDEEMLIASAK 305
Query: 150 NGLLPGGLNSKLTRKPARSMEEMRARASTYILDEEDD 186
LLPG L L R ++ +++ R ++ EED+
Sbjct: 306 RALLPGPLRFDLARNRPKTAKDLFERMESFARGEEDE 342
>UniRef100_Q7XVT8 OSJNBa0035B13.13 protein [Oryza sativa]
Length = 1736
Score = 85.5 bits (210), Expect = 1e-15
Identities = 48/184 (26%), Positives = 84/184 (45%)
Query: 1 READSVAEFRPFSEDVESVAILDNMKALVLDSYSGDSDPKDHLLYFNTKMVIIAASDAVK 60
R D++ F++D+ V K ++ Y G ++P+ L + + +
Sbjct: 315 RRDDNLDGVAAFTDDLRRVDWPAGFKPTRIEKYDGTTNPESWLTVYGLAIRAAGGDNKAM 374
Query: 61 CRMFPSTFKSTAMAWFTTLPRGSISNFRDFSSKFLVQFSANKNQPVTINDLYSIRQQEGE 120
P +A +W LPRG+I ++ + F+ F +P T DLY++ Q+ GE
Sbjct: 375 ANYLPVALADSARSWLHGLPRGTIGSWAELRDHFIANFQGTFERPGTQYDLYNVIQKSGE 434
Query: 121 SLKEYVARYSAASVKVEDEEPRACALAFKNGLLPGGLNSKLTRKPARSMEEMRARASTYI 180
SL++Y+ R+S K+ D AF G+ L K RKP R+++ M +A+ Y
Sbjct: 435 SLRDYIRRFSEQRNKISDITDDVIIAAFTKGIRHEELVGKFGRKPPRTVKLMFEKANEYA 494
Query: 181 LDEE 184
E+
Sbjct: 495 KAED 498
>UniRef100_Q7X6L5 OSJNBb0093G06.3 protein [Oryza sativa]
Length = 1986
Score = 85.5 bits (210), Expect = 1e-15
Identities = 47/173 (27%), Positives = 80/173 (46%)
Query: 12 FSEDVESVAILDNMKALVLDSYSGDSDPKDHLLYFNTKMVIIAASDAVKCRMFPSTFKST 71
F++D+ V K ++ Y G ++P+ L + + + P +
Sbjct: 421 FTDDLRRVDWPAGFKPTGIEKYDGTTNPESWLTVYGLAIRAAGGDNKAMANYLPVALADS 480
Query: 72 AMAWFTTLPRGSISNFRDFSSKFLVQFSANKNQPVTINDLYSIRQQEGESLKEYVARYSA 131
A +W LPRG+I ++ + F+ F +P T DLY++ Q+ GESL+EY+ R+S
Sbjct: 481 ARSWLHGLPRGTIGSWAELRDHFIANFQGTFERPGTQYDLYNVIQKSGESLREYIRRFSE 540
Query: 132 ASVKVEDEEPRACALAFKNGLLPGGLNSKLTRKPARSMEEMRARASTYILDEE 184
K+ D AF G+ L K RKP R+++ M +A+ Y E+
Sbjct: 541 QRNKISDISDDVIIAAFTKGIRHEELVGKFGRKPPRTVKLMFEKANEYAKAED 593
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.320 0.134 0.373
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 393,884,533
Number of Sequences: 2790947
Number of extensions: 15538924
Number of successful extensions: 39998
Number of sequences better than 10.0: 531
Number of HSP's better than 10.0 without gapping: 456
Number of HSP's successfully gapped in prelim test: 76
Number of HSP's that attempted gapping in prelim test: 39372
Number of HSP's gapped (non-prelim): 611
length of query: 263
length of database: 848,049,833
effective HSP length: 125
effective length of query: 138
effective length of database: 499,181,458
effective search space: 68887041204
effective search space used: 68887041204
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 73 (32.7 bits)
Lotus: description of TM0192.7


