

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0192.11
(571 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
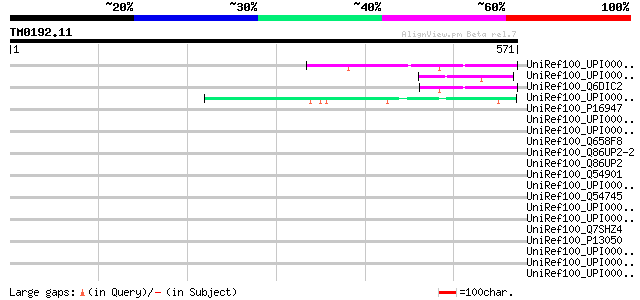
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_UPI00001CFC7D UPI00001CFC7D UniRef100 entry 51 8e-05
UniRef100_UPI000023F6CB UPI000023F6CB UniRef100 entry 49 3e-04
UniRef100_Q6DIC2 Early endosome antigen 1 [Mus musculus] 49 3e-04
UniRef100_UPI0000361937 UPI0000361937 UniRef100 entry 49 5e-04
UniRef100_P16947 M protein, serotype 49 precursor [Streptococcus... 47 0.002
UniRef100_UPI0000369CDB UPI0000369CDB UniRef100 entry 46 0.003
UniRef100_UPI0000369CDA UPI0000369CDA UniRef100 entry 46 0.003
UniRef100_Q658F8 Hypothetical protein P0015E04.9 [Oryza sativa] 46 0.003
UniRef100_Q86UP2-2 Splice isoform 2 of Q86UP2 [Homo sapiens] 46 0.003
UniRef100_Q86UP2 Kinectin [Homo sapiens] 46 0.003
UniRef100_Q54901 Precursor to Protein Sir22 precursor [Streptoco... 45 0.004
UniRef100_UPI000049A29E UPI000049A29E UniRef100 entry 45 0.006
UniRef100_Q54745 Emm50 [Streptococcus pyogenes] 45 0.007
UniRef100_UPI0000456A48 UPI0000456A48 UniRef100 entry 44 0.010
UniRef100_UPI00001CF6F5 UPI00001CF6F5 UniRef100 entry 44 0.010
UniRef100_Q7SHZ4 Hypothetical protein [Neurospora crassa] 44 0.010
UniRef100_P13050 IgA receptor precursor [Streptococcus pyogenes] 44 0.010
UniRef100_UPI0000437334 UPI0000437334 UniRef100 entry 44 0.013
UniRef100_UPI000035EFFA UPI000035EFFA UniRef100 entry 44 0.013
UniRef100_UPI000021BCCB UPI000021BCCB UniRef100 entry 44 0.013
>UniRef100_UPI00001CFC7D UPI00001CFC7D UniRef100 entry
Length = 1804
Score = 51.2 bits (121), Expect = 8e-05
Identities = 60/250 (24%), Positives = 111/250 (44%), Gaps = 16/250 (6%)
Query: 335 ENPKKKRHQRDNEVCENSNPVRQGTGEKESRTALREKEGHFDLAEK-------EARPNLG 387
+N K++ + + ++ N N ++Q T +K+ + + E ++K + +
Sbjct: 1337 QNTLKQKEKEEQQLQGNINQLKQATEQKKKQMEALQGELKNVTSQKAQLESKLQQQAAQA 1396
Query: 388 EKEKEVPSDPSTMLRDATDHVEFLLSRINHDYL--EKEVFGKGIDDAGQECVASLFHTGC 445
+E V + L+ D + L ++ D E E+ D E +L
Sbjct: 1397 AQELAVEKGKLSALQSTYDKCQADLQQLQSDLYGKESELLATRQDLKCVEEKLALAQEDL 1456
Query: 446 IFAHAFQKFGASTAEGEKLQADYTALRTEHAKCDDRL---DKVLTEAARDKITIDKKLVS 502
I + G ++LQA AL + AK ++ L K L +A R+K +K+LV+
Sbjct: 1457 ISNR--NQIGNQNKSIQELQAAKAALEQDLAKKEEVLKEQSKALQDAQREKSVKEKELVT 1514
Query: 503 IANELVEARFELVCRSER-ITDLEEEIKETKQEAQLALAAKDDALALKDQELTSLRAELE 561
++L E E+ CR E+ I L+EE+K KQE+ + DA L Q+ L+ +++
Sbjct: 1515 EKSKLAEME-EIKCRQEKEIAKLKEELKSHKQESIKEVTNLKDAKQLLIQQKLELQGKVD 1573
Query: 562 EKKRDLLKKK 571
K L ++K
Sbjct: 1574 SLKAALEQEK 1583
>UniRef100_UPI000023F6CB UPI000023F6CB UniRef100 entry
Length = 1493
Score = 49.3 bits (116), Expect = 3e-04
Identities = 33/110 (30%), Positives = 56/110 (50%), Gaps = 4/110 (3%)
Query: 461 GEKLQADYTALRTEHAKCDDRLDKVLTEAARDKITIDKKLVSIANELVEARFELVCRSER 520
GEKL+A T + + +++ R K+ +K+ V + + ++ EL S
Sbjct: 1017 GEKLRAQRTKVDAIKEEISSNNEEISNAEVR-KVKAEKQKVKLEKDHAKSSKELAAASRD 1075
Query: 521 ITDLEEEIK---ETKQEAQLALAAKDDALALKDQELTSLRAELEEKKRDL 567
+ LE +I E +E Q +A ++ LA K QEL +L+AEL+EK +L
Sbjct: 1076 LEKLENDINNQGERAEELQAQVAEAEEGLATKKQELKALKAELDEKTDEL 1125
>UniRef100_Q6DIC2 Early endosome antigen 1 [Mus musculus]
Length = 1411
Score = 49.3 bits (116), Expect = 3e-04
Identities = 38/114 (33%), Positives = 61/114 (53%), Gaps = 5/114 (4%)
Query: 462 EKLQADYTALRTEHAKCDDRL---DKVLTEAARDKITIDKKLVSIANELVEARFELVCRS 518
++LQA +L + AK + L K L +A R+K +K+LV+ ++L E E+ CR
Sbjct: 1078 QELQAAKASLEQDSAKKEALLKEQSKALEDAQREKSVKEKELVAEKSKLAEME-EIKCRQ 1136
Query: 519 ER-ITDLEEEIKETKQEAQLALAAKDDALALKDQELTSLRAELEEKKRDLLKKK 571
E+ IT L EE+K KQE+ + DA L Q+ L+ ++ K L ++K
Sbjct: 1137 EKEITKLNEELKSHKQESIKEITNLKDAKQLLIQQKLELQGRVDSLKAALEQEK 1190
Score = 37.0 bits (84), Expect = 1.6
Identities = 34/119 (28%), Positives = 59/119 (49%), Gaps = 24/119 (20%)
Query: 460 EGEKLQADYTALRTEHAKCDDRLDKVLTE-----AARDKI-----TIDKKLVSIANELVE 509
E EKL D +T+H + DR+ +TE A +D + T +KL +++ L
Sbjct: 815 EFEKLSQDS---KTQHKELGDRMQAAVTELTAVKAQKDALLAELSTTKEKLSKVSDSLKN 871
Query: 510 ARFELVCRSER----ITDLEEEIKETKQEAQLALAAKDDALALKDQELTSLRAELEEKK 564
++ E +++ + DLE+ KE K + Q+ + ALK+QE L+ LE++K
Sbjct: 872 SKSEFEKENQKGKAAVLDLEKACKELKHQLQVQAES-----ALKEQE--DLKKSLEKEK 923
>UniRef100_UPI0000361937 UPI0000361937 UniRef100 entry
Length = 1370
Score = 48.5 bits (114), Expect = 5e-04
Identities = 84/391 (21%), Positives = 148/391 (37%), Gaps = 55/391 (14%)
Query: 220 DAKSLKAKGWVSLKARMGRKCLNPFKSNAKTSFSQKYFMVAVHQAYPEVFTLRDG--TAS 277
DAK L K R+GR ++ K +A +Q F + P V L T++
Sbjct: 300 DAKDLICAFLSDRKVRLGRNGVDDIKRHAFFQNNQWTFDIIRETVAPVVPELSSDIDTSN 359
Query: 278 FPFYWTDKPNGIVDPPVDSLTVEDKTIIGF--FAQLPILDCSKLAAARRLAKFPPNFNTE 335
F DK N PP + +GF F + +L + A N +
Sbjct: 360 FDDIENDKGNAETFPPPRAFVGNQLPFVGFTYFKEDQLLKANNKYLAEDSKDGEENSENK 419
Query: 336 NP------KKKRHQRDNEV-------------CENSNPV---------RQGTGEKESRTA 367
+KK HQ + ++ C+N+ ++ +G ++ +
Sbjct: 420 ESDTNTELQKKLHQLEEQLDCEMQAKDELEQKCKNATNRLEKLVKELDKEMSGRQKVEAS 479
Query: 368 LREKEGHFDLAEKEARPNLGEKEKEVPSDPS--TMLRDATDHVEFLLSRINHDYLEKEV- 424
LR+ E L + ++ NL + E E S L + D +E L + + + E
Sbjct: 480 LRQLERERALLQHQSVENLRKVELETDRKRSLENELNNLRDQLEDLRKKNQNSQISNEKN 539
Query: 425 --FGKGIDDAGQECVASLFHTGCIFAHAFQKFGASTAEGEKLQADYTALRTEHAKCDDRL 482
K +DD A A + S AE +K A E + +L
Sbjct: 540 IQLQKQLDDTNAGLQAE--------QEAAARLKKSQAEAQKQAQSLEASLGETKERCSQL 591
Query: 483 DKVLTEAARDKITIDKKLVSIANELVEARFELVCRSERITDLEEEIKETKQEAQLALAAK 542
+K D++ ++K+L ++ EL E R + R+E I DL+ + K+E + ++
Sbjct: 592 EK-------DRMELEKQLTALQAELDEERRDGNLRTETIADLQGRVGRLKEEVKEVKSSH 644
Query: 543 DDALALK---DQELTSLRAELEEKKRDLLKK 570
+ L+ K ++L L E ++ DL+ K
Sbjct: 645 SEVLSEKKKLQEQLGILEKEKSNQEIDLMFK 675
>UniRef100_P16947 M protein, serotype 49 precursor [Streptococcus pyogenes]
Length = 389
Score = 47.0 bits (110), Expect = 0.002
Identities = 58/239 (24%), Positives = 106/239 (44%), Gaps = 31/239 (12%)
Query: 340 KRHQRDNEVCENSNPVRQGTGEKESRTALREKEGHFD-----LAEKEARPNLGEKEKEVP 394
K+HQ + E RQ E+ R RE E + + E + E ++
Sbjct: 115 KKHQEYKQEQEE----RQKNQEQLERKYQREVEKRYQEQLQKQQQLETEKQISEASRKSL 170
Query: 395 SDPSTMLRDATDHVEFLLSRIN--HDYLEKEVFGKGIDDAGQECVASLFHTGCIFAHAFQ 452
S R+A VE L+ + H L++E K I DA ++ ++ +
Sbjct: 171 SRDLEASREAKKKVEADLAALTAEHQKLKEE---KQISDASRQGLS-------------R 214
Query: 453 KFGASTAEGEKLQADYTALRTEHAKCDDRLDKVLTEAARDKITIDKKLVSIANELVEARF 512
AS +K++AD AL EH K + +K +++A+R ++ D + A + VEA
Sbjct: 215 DLEASREAKKKVEADLAALTAEHQKLKE--EKQISDASRQGLSRDLEASREAKKKVEA-- 270
Query: 513 ELVCRSERITDLEEEIKETKQEAQLALAAKDDALALKDQELTSLRAELEEKKRDLLKKK 571
+L + ++ LE+ KE ++ +L+ K + A + E +L+ +L ++ +L K K
Sbjct: 271 DLAEANSKLQALEKLNKELEEGKKLSEKEKAELQARLEAEAKALKEQLAKQAEELAKLK 329
>UniRef100_UPI0000369CDB UPI0000369CDB UniRef100 entry
Length = 1298
Score = 46.2 bits (108), Expect = 0.003
Identities = 56/221 (25%), Positives = 101/221 (45%), Gaps = 32/221 (14%)
Query: 371 KEGHFDLAEKEARPNLGEKEKE--VPSDPSTMLRDATDHVEFLLS-RINHDYLEKEVFGK 427
K+ L E++ + + K +E + +D + L+ ++ L+S + N D +E+
Sbjct: 678 KDDKIRLLEEQLQHEISNKMEEFKILNDQNKALKSEVQKLQTLVSEQPNKDVVEQMEKCI 737
Query: 428 GIDDAGQECVASLFHTGCI-FAHAFQKFGASTAEGEKLQADYTALRTEH------AKCDD 480
D + V L TG I A ++ A E L + L+ + A +
Sbjct: 738 QEKDEKLKTVEELLETGLIQVATKEEELNAIRTENSSLTKEVQDLKAKQNDQVSFASLVE 797
Query: 481 RLDKVLTEAARDKITIDKKLVSIANELVEARFELVCRSER-ITDLEEEIKETKQEA---- 535
L KV+ E D K+ S+ EL+EA V E+ + DL++EIK K+E
Sbjct: 798 ELKKVIHEK-------DGKIKSV-EELLEAELLKVANKEKTVQDLKQEIKALKEEIGNVQ 849
Query: 536 -----QLALAAK----DDALALKDQELTSLRAELEEKKRDL 567
QL++ +K + L K++++ +++A LEEK++DL
Sbjct: 850 LEKAQQLSITSKVQELQNLLKGKEEQMNTMKAVLEEKEKDL 890
>UniRef100_UPI0000369CDA UPI0000369CDA UniRef100 entry
Length = 1355
Score = 46.2 bits (108), Expect = 0.003
Identities = 56/221 (25%), Positives = 101/221 (45%), Gaps = 32/221 (14%)
Query: 371 KEGHFDLAEKEARPNLGEKEKE--VPSDPSTMLRDATDHVEFLLS-RINHDYLEKEVFGK 427
K+ L E++ + + K +E + +D + L+ ++ L+S + N D +E+
Sbjct: 678 KDDKIRLLEEQLQHEISNKMEEFKILNDQNKALKSEVQKLQTLVSEQPNKDVVEQMEKCI 737
Query: 428 GIDDAGQECVASLFHTGCI-FAHAFQKFGASTAEGEKLQADYTALRTEH------AKCDD 480
D + V L TG I A ++ A E L + L+ + A +
Sbjct: 738 QEKDEKLKTVEELLETGLIQVATKEEELNAIRTENSSLTKEVQDLKAKQNDQVSFASLVE 797
Query: 481 RLDKVLTEAARDKITIDKKLVSIANELVEARFELVCRSER-ITDLEEEIKETKQEA---- 535
L KV+ E D K+ S+ EL+EA V E+ + DL++EIK K+E
Sbjct: 798 ELKKVIHEK-------DGKIKSV-EELLEAELLKVANKEKTVQDLKQEIKALKEEIGNVQ 849
Query: 536 -----QLALAAK----DDALALKDQELTSLRAELEEKKRDL 567
QL++ +K + L K++++ +++A LEEK++DL
Sbjct: 850 LEKAQQLSITSKVQELQNLLKGKEEQMNTMKAVLEEKEKDL 890
>UniRef100_Q658F8 Hypothetical protein P0015E04.9 [Oryza sativa]
Length = 569
Score = 46.2 bits (108), Expect = 0.003
Identities = 47/180 (26%), Positives = 67/180 (37%), Gaps = 27/180 (15%)
Query: 164 IKFPFSSFFCDVLADINVAPCQLHPNAWAFMRCFEILCVAIQVTPSPV-------HFFYL 216
++ P FF VLA +AP QL PNAW + F LC V P P HFF
Sbjct: 98 MRLPLQPFFAAVLAHFGLAPSQLTPNAWRLLAGFAALCRRAGVAPPPPPPLTVFRHFF-- 155
Query: 217 YDVDAKSLKAKGWVSLKARMGRKCLNPFKSNAKTSFSQKYFMVAVHQAYPEVFTLRDGTA 276
A W S +AR S + S + V ++ + E F L A
Sbjct: 156 ---TAIGCPPGAWYSFRAR---------GSASTGSLFTRLSNVTMNPWWKEQFILVSSPA 203
Query: 277 SF--PFYWTDKPNGIVDPPVDSLTVEDKTIIGFFAQ----LPILDCSKLAAARRLAKFPP 330
+ P W PPV + +D A+ + + + +AAA+ + PP
Sbjct: 204 PWPCPVRWGKPRKHANFPPVLTTAEKDMATKLLLARGSSLIDLTTYNSIAAAKNIIAPPP 263
>UniRef100_Q86UP2-2 Splice isoform 2 of Q86UP2 [Homo sapiens]
Length = 1300
Score = 46.2 bits (108), Expect = 0.003
Identities = 56/221 (25%), Positives = 101/221 (45%), Gaps = 32/221 (14%)
Query: 371 KEGHFDLAEKEARPNLGEKEKE--VPSDPSTMLRDATDHVEFLLS-RINHDYLEKEVFGK 427
K+ L E++ + + K +E + +D + L+ ++ L+S + N D +E+
Sbjct: 678 KDDKIRLLEEQLQHEISNKMEEFKILNDQNKALKSEVQKLQTLVSEQPNKDVVEQMEKCI 737
Query: 428 GIDDAGQECVASLFHTGCI-FAHAFQKFGASTAEGEKLQADYTALRTEH------AKCDD 480
D + V L TG I A ++ A E L + L+ + A +
Sbjct: 738 QEKDEKLKTVEELLETGLIQVATKEEELNAIRTENSSLTKEVQDLKAKQNDQVSFASLVE 797
Query: 481 RLDKVLTEAARDKITIDKKLVSIANELVEARFELVCRSER-ITDLEEEIKETKQEA---- 535
L KV+ E D K+ S+ EL+EA V E+ + DL++EIK K+E
Sbjct: 798 ELKKVIHEK-------DGKIKSV-EELLEAELLKVANKEKTVQDLKQEIKALKEEIGNVQ 849
Query: 536 -----QLALAAK----DDALALKDQELTSLRAELEEKKRDL 567
QL++ +K + L K++++ +++A LEEK++DL
Sbjct: 850 LEKAQQLSITSKVQELQNLLKGKEEQMNTMKAVLEEKEKDL 890
>UniRef100_Q86UP2 Kinectin [Homo sapiens]
Length = 1357
Score = 46.2 bits (108), Expect = 0.003
Identities = 56/221 (25%), Positives = 101/221 (45%), Gaps = 32/221 (14%)
Query: 371 KEGHFDLAEKEARPNLGEKEKE--VPSDPSTMLRDATDHVEFLLS-RINHDYLEKEVFGK 427
K+ L E++ + + K +E + +D + L+ ++ L+S + N D +E+
Sbjct: 678 KDDKIRLLEEQLQHEISNKMEEFKILNDQNKALKSEVQKLQTLVSEQPNKDVVEQMEKCI 737
Query: 428 GIDDAGQECVASLFHTGCI-FAHAFQKFGASTAEGEKLQADYTALRTEH------AKCDD 480
D + V L TG I A ++ A E L + L+ + A +
Sbjct: 738 QEKDEKLKTVEELLETGLIQVATKEEELNAIRTENSSLTKEVQDLKAKQNDQVSFASLVE 797
Query: 481 RLDKVLTEAARDKITIDKKLVSIANELVEARFELVCRSER-ITDLEEEIKETKQEA---- 535
L KV+ E D K+ S+ EL+EA V E+ + DL++EIK K+E
Sbjct: 798 ELKKVIHEK-------DGKIKSV-EELLEAELLKVANKEKTVQDLKQEIKALKEEIGNVQ 849
Query: 536 -----QLALAAK----DDALALKDQELTSLRAELEEKKRDL 567
QL++ +K + L K++++ +++A LEEK++DL
Sbjct: 850 LEKAQQLSITSKVQELQNLLKGKEEQMNTMKAVLEEKEKDL 890
>UniRef100_Q54901 Precursor to Protein Sir22 precursor [Streptococcus pyogenes]
Length = 365
Score = 45.4 bits (106), Expect = 0.004
Identities = 47/214 (21%), Positives = 99/214 (45%), Gaps = 24/214 (11%)
Query: 376 DLAEKEARPNLGEKEKEVPSDPSTMLRDATDHVEFLLSRINHDYLEKEVFGKGIDDAGQE 435
DL EKE + +K K++ + + + D L +++ ++ E++ K +++ ++
Sbjct: 98 DLREKERKYQ--DKIKKLEEKEKNLEKKSEDVERHYLKKLDQEHKEQQERQKNLEELERQ 155
Query: 436 CVASL-------------FHTGCIFAHAFQK-----FGASTAEGEKLQADYTALRTEHAK 477
+ T + A +K AS A +K++AD AL EH K
Sbjct: 156 SQREIDKRYQEQLQKQQQLETEKQISEASRKSLSRDLEASRAAKKKVEADLAALNAEHQK 215
Query: 478 CDDRLDKVLTEAARDKITIDKKLVSIANELVEARFELVCRSERITDLEEEIKETKQEAQL 537
+ +K +++A+R ++ D + A + VEA +L + ++ LE+ KE ++ +L
Sbjct: 216 LKE--EKQISDASRQGLSRDLEASREAKKKVEA--DLAEANSKLQALEKLNKELEEGKKL 271
Query: 538 ALAAKDDALALKDQELTSLRAELEEKKRDLLKKK 571
+ K + A + E +L+ +L ++ +L K K
Sbjct: 272 SEKEKAELQARLEAEAKALKEQLAKQAEELAKLK 305
>UniRef100_UPI000049A29E UPI000049A29E UniRef100 entry
Length = 1813
Score = 45.1 bits (105), Expect = 0.006
Identities = 50/231 (21%), Positives = 102/231 (43%), Gaps = 26/231 (11%)
Query: 341 RHQRDNEVCENSNPVRQGTGEKESRTALREKEGHFDLAEKEARPNLGEKEKEVPSDPSTM 400
+++RDN + N ++ +KE+ T +E L E N ++EK+ D +
Sbjct: 653 KNERDN-ISNEFNKTKEEIKQKENETIQLNEEKSVLLNEL----NQIKEEKQKIEDEKAV 707
Query: 401 LRDATDHVEFLLSRINHDYLEKEVFGKGIDDAGQECVASLFHTGCIFAHAFQKFGASTAE 460
++ ++ ++++N D E I QE L T E
Sbjct: 708 IQQEKENE---ITKLNEDKTVIENELNQIKTEKQEIENELNQT--------------KDE 750
Query: 461 GEKLQADYTALRTEHAKCDDRLDKVLTEAARDKITIDKKLVSIANELVEARFELVCRSER 520
+K++ + + L TE + +D + K+ E + K ++ ++ NEL + + E E+
Sbjct: 751 KQKIEDEKSKLITELSNGNDGISKLNEELTQTK----QEKENVLNELNQIKNEFASFKEQ 806
Query: 521 ITDLEEEIKETKQEAQLALAAKDDALALKDQELTSLRAELEEKKRDLLKKK 571
T E E+K+ + Q L K++ ++ ++E ++ EL K++L +KK
Sbjct: 807 NTQKENELKDENNKVQQELEQKNNEVSKLEEEKGNISNELSNTKQELEQKK 857
Score = 41.2 bits (95), Expect = 0.083
Identities = 33/183 (18%), Positives = 82/183 (44%), Gaps = 29/183 (15%)
Query: 388 EKEKEVPSDPSTMLRDATDHVEFLLSRINHDYLEKEVFGKGIDDAGQECVASLFHTGCIF 447
++EKE ++ T L+ D E L+++ H+ E
Sbjct: 267 KQEKESINNELTQLKTDNDQKENELNQVRHEKDE-------------------------- 300
Query: 448 AHAFQKFGASTAEGEKLQADYTALRTEHAKCDDRLDKVLTEAARDKITIDKKLVSIANEL 507
+KF S E EK+ + + L+ E + ++ L + + + +K + +L + ++ +
Sbjct: 301 --VIEKFNTSKEENEKIMNELSQLKQEKEEKENELKEQVKKMEEEKSKLITELSNGSDGI 358
Query: 508 VEARFELVCRSERITDLEEEIKETKQEAQLALAAKDDALALKDQELTSLRAELEEKKRDL 567
+ EL + ++ E+ K+E + + + + + +++E+ + ++EE+K++L
Sbjct: 359 SKLNEELTQTKQEKEEINNELNSIKEEKK-RIEEEKNQIINENKEIKEEKEKIEEEKKEL 417
Query: 568 LKK 570
LK+
Sbjct: 418 LKE 420
Score = 40.8 bits (94), Expect = 0.11
Identities = 50/253 (19%), Positives = 109/253 (42%), Gaps = 27/253 (10%)
Query: 343 QRDNEVCENSNPVRQGTGEK----ESRTALREKEGHFDLAEKEARPNLG-------EKEK 391
+RD + E ++ Q G K E+ + E + F+ +E E +L EKEK
Sbjct: 1052 ERDRVISELNDIKLQNEGMKKQVEEAHNRMTEMQKSFEGSENEMINSLNNQITQLNEKEK 1111
Query: 392 EVPSDPSTMLRDATDHVEFLLSRINHDYLEKEVFGKGIDDAGQECVASLFHT-GCIFAHA 450
++ ++ L+ L + D +E + I++ ++CV + +
Sbjct: 1112 QM-NEQVMALQTQLSQSNINLEEVKKDLIESQNKYTQINEE-KDCVEQERNKINEEYKTV 1169
Query: 451 FQKFGASTAEGEKLQADYTALRTEHAKCDDRLDKVLTEAARDKITIDKKLVSIANELVEA 510
++ + E LQ Y E K D L+ ++ +K +++++ + E +
Sbjct: 1170 NEELEKNKKELNDLQTKYDNEILELNKNKDELNSLINNLKEEKTNLEEQVKKMEEEKSKL 1229
Query: 511 RFELVCRSERITDLEEEIKETKQEAQ-------------LALAAKDDALALKDQELTSLR 557
EL S+ ++ L EE+ +TKQE + + + + + +++E+ +
Sbjct: 1230 ITELSNGSDGVSKLNEELTQTKQEKEEINNELNSIKEEKKRIEEEKNQIINENKEIKEEK 1289
Query: 558 AELEEKKRDLLKK 570
++EE+K++LLK+
Sbjct: 1290 EKIEEEKKELLKE 1302
Score = 39.3 bits (90), Expect = 0.31
Identities = 52/251 (20%), Positives = 104/251 (40%), Gaps = 28/251 (11%)
Query: 338 KKKRHQRDNEVCENSNPVRQGTGEKESRTA-LREKEGHFDLAEKEARPNLGEKEKEV--- 393
K++ Q++NE+ + +N V+Q +K + + L E++G+ + L +K++E+
Sbjct: 804 KEQNTQKENELKDENNKVQQELEQKNNEVSKLEEEKGNISNELSNTKQELEQKKQEIITI 863
Query: 394 ---PSDPSTMLRDATDHVEFLLSRINHDYLEKEVFGKGIDDAGQECVASLFHTGCIFAHA 450
+ L++ +E S++ E GI +E +
Sbjct: 864 TQEKEEKENELKEQVKKIEEEKSKL---ITELSNGSDGISKLNEELTQTKQEK-----EE 915
Query: 451 FQKFGASTAEGEKLQADYTALRTEHAKCDDRLDKVLTEAARDKITIDKKL---VSIANEL 507
QK A E EKL+ T L+ E + L++ + +K + ++L I EL
Sbjct: 916 IQK--ALEEEKEKLERIETELK-EIKEAKQELEEEKNKTIEEKTNLQQELNENKKIVEEL 972
Query: 508 VEARFELVCRSERITDLEEEIKETKQEAQLALAAKDD-------ALALKDQELTSLRAEL 560
+ + E + + ++EE K ++E + + ++ K QE+ SL +
Sbjct: 973 TQTKQEKEEINNELNSIKEEKKRIEEEKNQIINENKEIKEENIKSIEEKTQEINSLTTSI 1032
Query: 561 EEKKRDLLKKK 571
EE K L + K
Sbjct: 1033 EELKGRLEESK 1043
Score = 35.8 bits (81), Expect = 3.5
Identities = 43/221 (19%), Positives = 95/221 (42%), Gaps = 15/221 (6%)
Query: 346 NEVCENSNPVRQGTGEKESRTALREKEGHFDLAEKEARPNLGEKEKEVPSDPSTMLRDAT 405
N++ E+ N ++ E + + E + +E E N EKE L+
Sbjct: 1441 NQLNEDLNQIKNDKEELTEKNVQLQNEINKLKSENEELSNNLSFEKEG-------LKQVN 1493
Query: 406 DHVEFLLSRINHDYLEKEVFGKGIDDAGQECVASLFHTGCIFAHAFQKFGASTAEGEKLQ 465
+ V + + D L K++ K I++ ++ L G + ++ E E+L
Sbjct: 1494 EEVNAI--KEERDELVKQI--KKIEEEKRKVEEELNFNG---SEVNEQIAQINNEKEQLN 1546
Query: 466 ADYTALRTEHAKCDDRLDKVLTEAARDKITIDKKLVSIANELVEARFELVCRSERITDLE 525
+ L+ + +++++ E ++I ++L + E+ E ++ E I +E
Sbjct: 1547 QECNELKQNLKELQSKIEEIEQEKESNEIKKKEELQELQEEITEKDNDIKNLKEEIERIE 1606
Query: 526 EEIKETKQEAQLALAAKDDALALKDQELTSLRAELEEKKRD 566
+E++E K+E ++ + L +LT + LEE+K++
Sbjct: 1607 KELQE-KEEDMEQMSNNTEELEELKNKLTETQRLLEEEKKE 1646
Score = 34.7 bits (78), Expect = 7.7
Identities = 53/263 (20%), Positives = 106/263 (40%), Gaps = 53/263 (20%)
Query: 334 TENPKKK------RHQRDNEVCENSNPVRQGTGEKESRTALREKEGHFDLAEKEARPNLG 387
T+N +K+ RH++D EV E N KE + + ++E L
Sbjct: 282 TDNDQKENELNQVRHEKD-EVIEKFNT------SKEENEKIMNELSQLKQEKEEKENELK 334
Query: 388 EKEKEVPSDPS---TMLRDATDHVEFLLSRINHDYLEKEVFGKGIDDAGQECVASLFHTG 444
E+ K++ + S T L + +D + L + EKE ++ +E
Sbjct: 335 EQVKKMEEEKSKLITELSNGSDGISKLNEELTQTKQEKEEINNELNSIKEE--------- 385
Query: 445 CIFAHAFQKFGASTAEGEKLQA--DYTALRTEHAKCDDRLDKVLTEAARDKI---TIDKK 499
E EK Q + ++ E K ++ ++L E ++K + +
Sbjct: 386 -----------KKRIEEEKNQIINENKEIKEEKEKIEEEKKELLKEIEKEKEGNNQLQNE 434
Query: 500 LVSIAN---ELVEARFELVCRS--------ERITDLEEEIKETKQEAQLALAAKDDALAL 548
+ +I E+ E E++C + E +L++E+ + K+E Q K++ + +
Sbjct: 435 INTIQTRMKEIEEKNQEIICDNNKEIAKFKEEQENLQKELNQIKEEKQKTENEKNELVDV 494
Query: 549 KDQELTSLRAELEEKKRDLLKKK 571
K Q+ L +L+E+K + +K
Sbjct: 495 KTQKENELN-KLKEEKEQIFNEK 516
>UniRef100_Q54745 Emm50 [Streptococcus pyogenes]
Length = 413
Score = 44.7 bits (104), Expect = 0.007
Identities = 54/236 (22%), Positives = 111/236 (46%), Gaps = 25/236 (10%)
Query: 338 KKKRHQRDNEVCENSNPVRQGTGEKESRTALREKEGHFDLAEKEARPNLGEKEKEVPSDP 397
K+++ +++ ++ E S E SR A +E E ++E + + + ++ S
Sbjct: 141 KQQQLEKEKQISEASRKSLSRDLEA-SRAAKKELEAEHQKLKEEKQ--ISDASRQGLSRD 197
Query: 398 STMLRDATDHVEFLLSRIN--HDYLEKEVFGKGIDDAGQECVASLFHTGCIFAHAFQKFG 455
R+A VE L+ + H L++E K I +A ++ ++ +
Sbjct: 198 LEASREAKKKVEADLAALTAEHQKLKEE---KQISEASRQGLS-------------RDLE 241
Query: 456 ASTAEGEKLQADYTALRTEHAKCDDRLDKVLTEAARDKITIDKKLVSIANELVEARFELV 515
AS +K++AD AL EH K + +K +++A+R ++ D + A + VEA +L
Sbjct: 242 ASREAKKKVEADLAALTAEHQKLKE--EKQISDASRQGLSRDLEASREAKKKVEA--DLA 297
Query: 516 CRSERITDLEEEIKETKQEAQLALAAKDDALALKDQELTSLRAELEEKKRDLLKKK 571
+ ++ LE+ KE ++ +L+ K + A + E +L+ +L ++ +L K K
Sbjct: 298 EANSKLQALEKLNKELEEGKKLSEKEKAELQARLEAEAKALKEQLAKQAEELAKLK 353
>UniRef100_UPI0000456A48 UPI0000456A48 UniRef100 entry
Length = 1143
Score = 44.3 bits (103), Expect = 0.010
Identities = 53/266 (19%), Positives = 110/266 (40%), Gaps = 27/266 (10%)
Query: 315 DCSKLAAARRLAKFPPNFNTENPKKKRHQRDNEVCENSNPVRQGTGEKESRTA----LRE 370
+C K+ ++ K+ + + R+ D V E ++ +Q + E+++R L
Sbjct: 782 ECGKVVETKQQLKWNLRRLVKELRTVRNNLDLVVQERNDAQKQLSEEQDARILQDQILTS 841
Query: 371 KEGHFDLAEKEARPNLGEKEKEVPSDPSTMLRDATDHVEFLLSRINH-----DYLEKEVF 425
K+ ++A+K+ K + ++ + + + L+ NH D EKE
Sbjct: 842 KQKELEMAQKKMNSEFHSAAKNLQAESEKQILSLQEKNKELMDEYNHLKERMDQCEKEKA 901
Query: 426 GKGID-DAGQECVASLFHTGCIFAHAFQKFGASTAEGEKLQADYT--ALRTEHAKCDDRL 482
G+ ID QE V S C+ A E E LQ T +++ +C+ L
Sbjct: 902 GRKIDLTEAQETVPSR----CLHLDA---------ENEVLQLQQTLFSMKAIQKQCET-L 947
Query: 483 DKVLTEAARDKITIDKKLVSIANELVEA-RFELVCRSERITDLEEEIKETKQEAQLALAA 541
K + ++ + + + E E R +L+ +++E++ E Q+A +
Sbjct: 948 QKNKKQLKQEVVNLKSYMERNMLERGEGERHKLLIEERARKEIQEKLNEAVLTLQVAAVS 1007
Query: 542 KDDALALKDQELTSLRAELEEKKRDL 567
+ L++ TS++ ++E +DL
Sbjct: 1008 HEQLAQLREDNTTSIKTQMELTIKDL 1033
>UniRef100_UPI00001CF6F5 UPI00001CF6F5 UniRef100 entry
Length = 1253
Score = 44.3 bits (103), Expect = 0.010
Identities = 44/189 (23%), Positives = 87/189 (45%), Gaps = 21/189 (11%)
Query: 388 EKEKEVPSDPSTMLRDATDHVEFLLSRINHDYLEK-----EVFGKGIDDAGQECVASLFH 442
E+ K+ + S+ + + D + L+ R N Y+E+ E + A + A + H
Sbjct: 572 ERLKQELLEKSSRIEEQNDKISDLIER-NQRYVEQSNLMMEKRNNSLQTATENTQARILH 630
Query: 443 TGCIFAHAFQKFGASTAEGEKLQADYTALRTEHAKCDDRLDKVLTEAARDKITIDKKLVS 502
A ++ A+TA+ LQ TA H K + L LTE ++ + ++
Sbjct: 631 AEQEKAKVTEELAAATAQVSHLQLKMTA----HQKKETELQLQLTENLKETDLLRGQVTR 686
Query: 503 IANELVEARFELVCRSERITDLEEEIKETKQEAQLALAAKDDALALKDQELTSLRAELEE 562
+ +L+E R SE+ + K+++++ +L + + ++ ELT LRAE E
Sbjct: 687 LQADLLELREA----SEQTQTKFKSEKQSRRQLELKVTSLEE-------ELTDLRAEKES 735
Query: 563 KKRDLLKKK 571
+++L ++K
Sbjct: 736 LEKNLSERK 744
>UniRef100_Q7SHZ4 Hypothetical protein [Neurospora crassa]
Length = 4007
Score = 44.3 bits (103), Expect = 0.010
Identities = 58/259 (22%), Positives = 101/259 (38%), Gaps = 39/259 (15%)
Query: 339 KKRHQRDNEVCENSNPVRQGTGEKE--------------------SRTALREKEGHFDLA 378
K RH ++ E+ + R G E E S+ A++ K+ +LA
Sbjct: 1151 KNRHLQETEILRKQHQSRVGELESEIATIKEKYKKDLDELSRNNTSQDAIKLKQHENELA 1210
Query: 379 EKEARPNLGEKEKEVPSDPSTMLRDATDHVEFLLSRINHDYLEKEVFGKGIDDAGQECVA 438
+A+ +++K++ T + TD Y EKE + Q A
Sbjct: 1211 NFKAKYE--QEKKQLAVQHKTEMESLTDR-----------YHEKEKLATQYQERVQALSA 1257
Query: 439 SLFHTGCIFAHAFQKFGASTAEGEKLQADYTALRTE-HAKCDDRLDKVLTEAARDKITID 497
L A ++ AS A+ +KL+AD+ E AK + KV + + +
Sbjct: 1258 ELADKKTALAEYKEQLSASKAQLDKLKADHGVKVDELQAKLKSEVAKVTADYEGNLSELR 1317
Query: 498 KKLVSIANELV-----EARFELVCRSERITDLEEEIKETKQEAQLALAAKDDALALKDQE 552
K N L E + +E+I +LE I + K E + A D AL + E
Sbjct: 1318 TKHQGEVNVLKVHHQDEIKKLTAGHNEKIRNLEHRINDLKAELKQDRAEFDKKKALLEGE 1377
Query: 553 LTSLRAELEEKKRDLLKKK 571
+ +L+ ++++K L K+
Sbjct: 1378 VATLQGKVDDKSSKLSSKE 1396
Score = 37.0 bits (84), Expect = 1.6
Identities = 32/121 (26%), Positives = 51/121 (41%), Gaps = 13/121 (10%)
Query: 463 KLQADYTALRTEHAKCDDRL---DKVLTEAARDKITIDKKLVSIANELVEARFELVCRSE 519
KL D L+T+ K L D LT+ A + + D L + EL L ++E
Sbjct: 2508 KLNKDIFDLKTDVTKLKQELSTKDANLTQKAGEIGSRDAGLAKLREELRAKEAALAKKTE 2567
Query: 520 RITDLEEEIKETKQEA----------QLALAAKDDALALKDQELTSLRAELEEKKRDLLK 569
+ LE+ +K+ EA LA DA++ ++++ L EL K L +
Sbjct: 2568 EASSLEKNVKKLTDEATGLKKDVTSRDTQLAQDKDAISKLEKDIAKLNQELSTKDASLTQ 2627
Query: 570 K 570
K
Sbjct: 2628 K 2628
Score = 37.0 bits (84), Expect = 1.6
Identities = 29/131 (22%), Positives = 57/131 (43%), Gaps = 11/131 (8%)
Query: 452 QKFGASTAEGEKLQADYTALRTEHAKCDDRLDKVLTEAARDKITIDKKLVSIANELVEAR 511
+K GA+T+ E+ L + +LD+ E K + + + + +
Sbjct: 1497 RKEGAATSSTEQNTVQLNKLNDDVKDKQKKLDEQQAELNNLKTKHQAETTDLNQTIKDTK 1556
Query: 512 FELVCRSERITDLEEEIKETKQEAQLALAAKDDALALKDQELTSLRA-----------EL 560
+L + + DL+++ K+ + +A K LA K+ EL +L+A E+
Sbjct: 1557 AKLKQKETELIDLKKKHKDRLDTLEKTIAEKQTTLAQKETELENLKAQNRTNMMNTNREI 1616
Query: 561 EEKKRDLLKKK 571
+K +LLKK+
Sbjct: 1617 GDKTAELLKKE 1627
Score = 36.2 bits (82), Expect = 2.7
Identities = 55/242 (22%), Positives = 103/242 (41%), Gaps = 43/242 (17%)
Query: 345 DNEVCENSNPVRQGTGEKESRTA----LREKEGHFDLAEKEARPNLGEKEKEVPSDPSTM 400
+ E+ + Q TGE S+ A LRE ++ KE L +K +E+ ++
Sbjct: 2615 NQELSTKDASLTQKTGEVGSKNAELAKLRE-----EIRVKETA--LAKKTEELKGLNQSV 2667
Query: 401 LRDATDHVEFLLSRINHDYLEKEVFGKGIDDAGQECVASLFHTGCIFAHAFQKFGASTAE 460
DA D + +I + LEKEV G D + L +F K +
Sbjct: 2668 --DAKD-TQLAQDKIKIERLEKEVKGLTAD------IVKLREDVAFKDKSFAKKAEAV-- 2716
Query: 461 GEKLQADYTALRTEHAKCDDR---LDKVLTEAARDKITIDKKLVSIANELVEARFE---- 513
+ L+AD T L +E AK D + ++ +++ K + + N+ ++ +
Sbjct: 2717 -DHLKADITELNSEVAKLKKEGTNKDAAILGKEKELVSLRKAVRDLTNQAKQSAQDSKKS 2775
Query: 514 ----------LVCRSERITDLEEEI---KETKQEAQLALAAKDDALALKDQELTSLRAEL 560
L + ++I +L++EI K+T +E +D L+ K++EL LR ++
Sbjct: 2776 AEDLANRDALLKEKEKKIFELQQEIQKVKDTAEELNQTTKTRDSTLSQKNEELRKLREQI 2835
Query: 561 EE 562
++
Sbjct: 2836 KQ 2837
Score = 35.8 bits (81), Expect = 3.5
Identities = 52/245 (21%), Positives = 102/245 (41%), Gaps = 31/245 (12%)
Query: 331 NFNTENPKKKRHQRDNEVCE--NSNPVRQGTGEK---ESRTALREKEGHFDLAEKEARPN 385
N ++ K K Q++ E+ + + R T EK E +T L +KE + + + R N
Sbjct: 1549 NQTIKDTKAKLKQKETELIDLKKKHKDRLDTLEKTIAEKQTTLAQKETELENLKAQNRTN 1608
Query: 386 LGEKEKEVPSDPSTMLRDATDHVEFLLSRINHDYLEKEVFGKGIDDAGQECVASLFHTGC 445
+ +E+ + +L+ + + R +D +K G D
Sbjct: 1609 MMNTNREIGDKTAELLKKEGELRDL---RQKYDDAQKLADGSKEKDLA------------ 1653
Query: 446 IFAHAFQKFGASTAEGEKLQADYTALRTEHAKCDDRLDKVLTEAARDKITIDKKLVSIAN 505
A Q T+E EK + D AL + R+ + + + + + KK I++
Sbjct: 1654 -IAQYKQIIATKTSELEKAKKDVAALTKDVNDQKARIKDLESSVSSKRADLKKKETEISD 1712
Query: 506 ELVEARFELVCRSERITDLEEEIKETKQEAQLALAAKDDALALKDQELTSLRAELEEKKR 565
++ ++E E I L ++ + Q+A L A +++ ALK + L +++EK
Sbjct: 1713 --LKRQYE-----ENIKRLNNDL--SSQKATLT-AKENEIAALKSGNASRLSRDIQEKAS 1762
Query: 566 DLLKK 570
+L +K
Sbjct: 1763 ELAQK 1767
>UniRef100_P13050 IgA receptor precursor [Streptococcus pyogenes]
Length = 386
Score = 44.3 bits (103), Expect = 0.010
Identities = 33/116 (28%), Positives = 62/116 (53%), Gaps = 4/116 (3%)
Query: 456 ASTAEGEKLQADYTALRTEHAKCDDRLDKVLTEAARDKITIDKKLVSIANELVEARFELV 515
AS +K++AD AL EH K + DK +++A+R ++ D + A + VEA +L
Sbjct: 215 ASREAKKKVEADLAALTAEHQKLKE--DKQISDASRQGLSRDLEASREAKKKVEA--DLA 270
Query: 516 CRSERITDLEEEIKETKQEAQLALAAKDDALALKDQELTSLRAELEEKKRDLLKKK 571
+ ++ LE+ KE ++ +L+ K + A + E +L+ +L ++ +L K K
Sbjct: 271 EANSKLQALEKLNKELEEGKKLSEKEKAELQARLEAEAKALKEQLAKQAEELAKLK 326
>UniRef100_UPI0000437334 UPI0000437334 UniRef100 entry
Length = 3065
Score = 43.9 bits (102), Expect = 0.013
Identities = 57/240 (23%), Positives = 101/240 (41%), Gaps = 13/240 (5%)
Query: 333 NTENPKKKRHQRDNEVCENSNPVRQGTGEKESRTALREKEGHFDLA-EKEARPNLGEKEK 391
++E K+K R++E+ + + + GE + + E L ++EA N EK
Sbjct: 454 DSEKEKEKIRLREDELEQLQAEIHKQQGEIKMEKSNNENRDAESLKLDREAFEN----EK 509
Query: 392 EVPSDPSTMLRDATDHVEFLLSRINHDYLEKEVFGKGIDDAGQECVASLFHTGCIFAHAF 451
EV T L D +E + HD E I E + T
Sbjct: 510 EVLKQMKTELEREADEIEKIKLETQHDRQRVEEMAAQIQKDRDEINNLIEETNRKDMVLN 569
Query: 452 QKFGASTAEGEKLQADYTALRTEHAKCDDRLDKVLTEAARDKITIDKKLVSIANELVEAR 511
+K + + +Q+D L E K D L+K +E + K ++K+ + E+ + R
Sbjct: 570 EKNRDIEEKIKSIQSDKDMLEKE--KHD--LEKTRSELYKVKEDLEKQKENTLAEIQKER 625
Query: 512 FELVCRSERITDLEEEIKETKQEAQLALAAKDDALALKDQELTSLRAELEEKKRDLLKKK 571
+L +E IT EIK +++ + K D L E+ +L+ ELE++K ++K +
Sbjct: 626 EDLEKMNENITREMHEIKHQEEQ----MNQKQDELDQLKTEIQNLQQELEKEKEIIMKDR 681
Score = 38.1 bits (87), Expect = 0.70
Identities = 52/261 (19%), Positives = 107/261 (40%), Gaps = 45/261 (17%)
Query: 337 PKKKRHQRDNEVCENSNP-VRQGTGEKESRTALREKEGHFDLAEKEARPNLGEKEKEVPS 395
PK+K + D + E + + + + E +EKE + E + + KE E
Sbjct: 2494 PKEKEMKEDRKSLEETKANILEMKTKAEPEEIKKEKEKEEEEQEMRVKVEMERKEIEQIK 2553
Query: 396 DPSTMLRDATDHVEFLLSRINHDYLEKEVFGKGIDDAGQECVASLFHTGCIFAHAFQKFG 455
+R DH + ++N D E + ++ E +
Sbjct: 2554 SQLERVRSEMDHEQ---KKLNDDKKMIEQEKEDLEKMKSEIMKQ---------------- 2594
Query: 456 ASTAEGEKLQADYTALRTEHAKCD-----DRLDKVLTEAARDKITI-------DKKLVSI 503
E E+ + D +T+ K D + ++K++ E + K I D + I
Sbjct: 2595 RQQMEEERSELDKKIKQTDLEKHDIENSKEIVEKLMVEVEKQKHEIQFQKEELDIERQKI 2654
Query: 504 ANE---LVEARFELVCRSERITDLEEEIK---ETKQEAQLALAAKDDAL-------ALKD 550
A+E L++ + E +ERI +++EEIK ET +E ++ L + + + +
Sbjct: 2655 ADEQDLLIQNKIEQQNENERIKEMDEEIKKERETLKEMEVNLQKEKEEIESVIEETQRRK 2714
Query: 551 QELTSLRAELEEKKRDLLKKK 571
++L + ++ E+K+DL+ ++
Sbjct: 2715 EDLEKMSTDINEQKQDLMNQR 2735
Score = 37.4 bits (85), Expect = 1.2
Identities = 53/280 (18%), Positives = 112/280 (39%), Gaps = 61/280 (21%)
Query: 339 KKRHQRDNEVCENSNPVRQGTGEKESRTALREKEGHFDLAEKEARPNLGEKEKEVPSDPS 398
K Q +NE N N + K+ R L+E E H + ++E + ++ E+ + + D
Sbjct: 1898 KSELQNENERIRNLNEII-----KKERETLKEMEAHLEKKKEEIK-SVSEETRRLKEDLE 1951
Query: 399 TMLRDATDHVEFLLSR----------INHDYLEKEVFGKGIDDAGQECVASLFHTGCIFA 448
M D + L+++ INH + + + + ID+ + + L
Sbjct: 1952 KMSTDVNKQNKDLMNQRDLLEQEREEINHKWKQLQ---QRIDEFDAQIKSQLERVRSEID 2008
Query: 449 HAFQKFGASTAEGEKLQADYTALRTEHAKCDDRLDK------------------------ 484
H +K E+ + D +++E K ++++
Sbjct: 2009 HEQKKLNDDKKMIEQEKEDLEKMKSEIMKQRQQMEEERSELDNKIKQTDLERHDIENSKE 2068
Query: 485 ------VLTEAARDKITIDKKLVSIANE--------LVEARFELVCRSERITDLEEEIKE 530
V E R I + K+ + I + +V+ + +L +ERI +++EEIK+
Sbjct: 2069 IVQKLMVEVEEQRKDIRLQKEELDIERQKIADEQGLVVQNKAKLQNENERIKEMDEEIKK 2128
Query: 531 TKQEAQLALAAKDDALALKDQELTSLRAELEEKKRDLLKK 570
K+ L + L + +E+ S+ E + ++++ L+K
Sbjct: 2129 EKE----TLKEMEAHLRKEKEEMRSVIEETQRRQKEDLEK 2164
Score = 36.2 bits (82), Expect = 2.7
Identities = 46/237 (19%), Positives = 100/237 (41%), Gaps = 21/237 (8%)
Query: 336 NPKKKRHQRDNEVCENSNPVRQGTGEKESRTALREKEGHFDLAEKEARPNLGEKEKEVPS 395
N + Q E+ ++Q E E+ +EK + E N+ E+EK
Sbjct: 2733 NQRDLLKQEREEMNHKLTQLQQRIDEFETTNLQKEKN------KTEELKNILEREKSELY 2786
Query: 396 DPSTMLRDATDHVEFLLSRINHDYLEKEVFGKGIDDAGQECVASLFHTGCIFAHAFQKFG 455
++ +E + + + E ++D + ++ L + H K
Sbjct: 2787 QEKAQVQQQKHEMETIRKTMKSEMENLECLRNELEDEKKSMLSRL----SMEKHEIDKLK 2842
Query: 456 AST-AEGEKLQADYTALRT---EHAKCDDRLDKVLTEAARDKITIDKKLVSIANELVEAR 511
+E E ++ + L+ E+ +LD + + DK++I ++ + L E +
Sbjct: 2843 TEIQSEKESIEEEKKTLKQLKDENESKRTQLDNTIEKMNMDKLSIQRE----KDSLEEIK 2898
Query: 512 FELVCRSERITD-LEEEIKETKQEAQLALAAKDDALALKDQELTSLRAELEEKKRDL 567
+L ++E++ + + + ++ T Q +Q A DD K++EL S+RA++E + + L
Sbjct: 2899 ADLQRKAEKLDEEMRKAVQITSQISQDAKILDDDRE--KEEELRSMRADIERQAKIL 2953
Score = 35.8 bits (81), Expect = 3.5
Identities = 43/184 (23%), Positives = 77/184 (41%), Gaps = 8/184 (4%)
Query: 388 EKEKEVPSDPSTMLRDATDHVEFLLSRINHDYLEKEVFGKGIDDAGQECVASLFHTGCIF 447
E E+EV T L D +E + HD E I + + T
Sbjct: 971 ENEREVLKQMKTELEREADEIEKIKLETQHDRQRVEEMTAQIQKDRDKINNLIEETNRKD 1030
Query: 448 AHAFQKFGASTAEGEKLQADYTALRTEHAKCDDRLDKVLTEAARDKITIDKKLVSIANEL 507
+K + + +Q+D L E K D L+K +E + K ++K+ + ++
Sbjct: 1031 MVLNEKNRDIEKKIKSIQSDKDMLEKE--KHD--LEKTRSELYKVKEDLEKQKENTLAQI 1086
Query: 508 VEARFELVCRSERITDLEEEIKETKQEAQLALAAKDDALALKDQELTSLRAELEEKKRDL 567
+ R +L +E IT EIK +++ + K D L E+ +L+ ELE++K +
Sbjct: 1087 QKEREDLEKMNENITREMHEIKHQEEQ----MNQKQDELDQLKTEIQNLQQELEKEKEII 1142
Query: 568 LKKK 571
+K +
Sbjct: 1143 MKAR 1146
Score = 35.4 bits (80), Expect = 4.5
Identities = 35/129 (27%), Positives = 60/129 (46%), Gaps = 18/129 (13%)
Query: 460 EGEKLQADYTALRTEHAKCD-----DRLDKVLTEAARDKITI-------DKKLVSIANE- 506
E E+ + + +T KCD + + K++ E K I D + IA+E
Sbjct: 1832 EEERSELENKIKQTYIEKCDIENSKEIVQKLMVEVEEQKHEIQFQKEELDIERQKIADEQ 1891
Query: 507 --LVEARFELVCRSERITDLEEEIK---ETKQEAQLALAAKDDALALKDQELTSLRAELE 561
L++ + EL +ERI +L E IK ET +E + L K + + +E L+ +LE
Sbjct: 1892 DLLIQNKSELQNENERIRNLNEIIKKERETLKEMEAHLEKKKEEIKSVSEETRRLKEDLE 1951
Query: 562 EKKRDLLKK 570
+ D+ K+
Sbjct: 1952 KMSTDVNKQ 1960
>UniRef100_UPI000035EFFA UPI000035EFFA UniRef100 entry
Length = 1932
Score = 43.9 bits (102), Expect = 0.013
Identities = 35/111 (31%), Positives = 58/111 (51%), Gaps = 10/111 (9%)
Query: 458 TAEGEKLQADYTALRTEHAKCDDRLDKVLTEAARDKITIDKKLVSIANELVEARFELVCR 517
+AE EK A ++ + K + L K L + K +++K+VS+ E + + ++
Sbjct: 842 SAEAEKEMAQ---MKEDFEKTKEDLAKALAK----KKELEEKMVSLLQEKNDLQLQIQSE 894
Query: 518 SERITDLEEEIKET-KQEAQLALAAKDDALALKDQELTSLRAELEEKKRDL 567
SE ++D EE + K + QL AK+ A L+D+E + AEL KKR L
Sbjct: 895 SETLSDAEERCEGLIKAKIQLEAKAKETAERLEDEE--EINAELTAKKRKL 943
>UniRef100_UPI000021BCCB UPI000021BCCB UniRef100 entry
Length = 1496
Score = 43.9 bits (102), Expect = 0.013
Identities = 30/113 (26%), Positives = 56/113 (49%), Gaps = 9/113 (7%)
Query: 464 LQADYTALRTEHAKCDDRLDKVLTE------AARDKITIDKKLVSIANELVEARFELVCR 517
+Q LR + AK D +++ T A K+ +K+ V +A + +A +L
Sbjct: 1034 MQVGGEKLRAQRAKIDSLKEEIATHNEELSTAEVKKVKAEKQKVKLAKDHAKATKDLQAA 1093
Query: 518 SERITDLEEEIK---ETKQEAQLALAAKDDALALKDQELTSLRAELEEKKRDL 567
+ L++EI+ + + A+ +AL +K +EL+ L+AEL+EK +L
Sbjct: 1094 VRDLESLDDEIQNQGDKSESLSSAVEEAQEALKVKKKELSELKAELDEKTAEL 1146
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.320 0.134 0.403
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 976,425,483
Number of Sequences: 2790947
Number of extensions: 42977407
Number of successful extensions: 146093
Number of sequences better than 10.0: 1093
Number of HSP's better than 10.0 without gapping: 82
Number of HSP's successfully gapped in prelim test: 1031
Number of HSP's that attempted gapping in prelim test: 143324
Number of HSP's gapped (non-prelim): 3676
length of query: 571
length of database: 848,049,833
effective HSP length: 133
effective length of query: 438
effective length of database: 476,853,882
effective search space: 208862000316
effective search space used: 208862000316
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 78 (34.7 bits)
Lotus: description of TM0192.11


