

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0190.3
(329 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
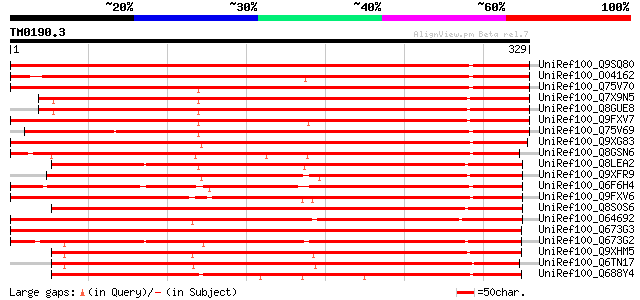
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9SQ80 Gibberellin 2-beta-dioxygenase 1 [Pisum sativum] 528 e-149
UniRef100_O04162 Dioxygenase [Marah macrocarpus] 468 e-130
UniRef100_Q75V70 Gibberellin 2-oxidase 1 [Nicotiana tabacum] 451 e-125
UniRef100_Q7X9N5 Gibberellin 2-oxidase [Cucurbita maxima] 450 e-125
UniRef100_Q8GUE8 Gibberellin 2-oxidase [Cucurbita maxima] 449 e-125
UniRef100_Q9FXV7 Gibberellin 2-oxidase No1 [Lactuca sativa] 438 e-121
UniRef100_Q75V69 Gibberellin 2-oxidase 2 [Nicotiana tabacum] 431 e-119
UniRef100_Q9XG83 Gibberellin 2-beta-dioxygenase [Phaseolus cocci... 417 e-115
UniRef100_Q8GSN6 Gibberellin 2-oxidase 1 [Spinacia oleracea] 384 e-105
UniRef100_Q8LEA2 Gibberellin 2-beta-dioxygenase 1 [Arabidopsis t... 373 e-102
UniRef100_Q9XFR9 Gibberellin 2-beta-dioxygenase 2 [Arabidopsis t... 370 e-101
UniRef100_Q6F6H4 Gibberellin 2-oxidase1 [Daucus carota] 370 e-101
UniRef100_Q9FXV6 Gibberellin 2-oxidase No2 [Lactuca sativa] 362 8e-99
UniRef100_Q8S0S6 Putative GA 2-oxidase [Oryza sativa] 346 4e-94
UniRef100_O64692 Gibberellin 2-beta-dioxygenase 3 [Arabidopsis t... 342 1e-92
UniRef100_Q673G3 GA 2-oxidase 4 [Hordeum vulgare var. distichum] 335 8e-91
UniRef100_Q673G2 GA 2-oxidase 5 [Hordeum vulgare var. distichum] 305 9e-82
UniRef100_Q9XHM5 Gibberellin 2-beta-dioxygenase 2 [Pisum sativum] 305 1e-81
UniRef100_Q6TN17 Gibberellin 2-oxidase [Populus alba x Populus t... 302 9e-81
UniRef100_Q688Y4 Putative GA2-oxidase [Oryza sativa] 296 4e-79
>UniRef100_Q9SQ80 Gibberellin 2-beta-dioxygenase 1 [Pisum sativum]
Length = 327
Score = 528 bits (1360), Expect = e-149
Identities = 256/329 (77%), Positives = 294/329 (88%), Gaps = 2/329 (0%)
Query: 1 MVLLSKQATEQYSYVKNCMATPCSFTIPVVDLSKPDAKSIIVKACEEFGFFKVINHGVSM 60
MVLLSK +EQY+YV+N M S +IP+VDLSKPDAK++IVKACE+FGFFKVINHG+ +
Sbjct: 1 MVLLSKPTSEQYTYVRNNMPITFSSSIPLVDLSKPDAKTLIVKACEDFGFFKVINHGIPL 60
Query: 61 EAISQLESEALNFFSMPVTEKEKTGLANPFGYGNKRIGHNGDVGWVEYLLLKTNQEVNVP 120
+AISQLESEA FFS+P TEKEK G ANPFGYGNKRIG NGD+GW+EYLLL TNQ+ N
Sbjct: 61 DAISQLESEAFKFFSLPQTEKEKAGPANPFGYGNKRIGLNGDIGWIEYLLLTTNQDHNFS 120
Query: 121 VHSQNPEKFRCVLDDYLCAVRNMGCEILDLMAEGLKIQPKNVLSKLLMDKQSDSVFRVNH 180
++ ++ +KFR +L DY CA+RNM CEILDLMAEGLKIQPKNV SKL+MDKQSD +FRVNH
Sbjct: 121 LYGEDIDKFRGLLKDYKCAMRNMACEILDLMAEGLKIQPKNVFSKLVMDKQSDCLFRVNH 180
Query: 181 YPACPEMGLDGENLIGFGEHTDPQIISLLRSNNTSGLQISLRDGSWISVPPDYSSFFINV 240
YPACPE+ ++GENLIGFGEHTDPQIIS+LRSNNTSG QISLRDGSWISVPPD+SSFFINV
Sbjct: 181 YPACPELAINGENLIGFGEHTDPQIISILRSNNTSGFQISLRDGSWISVPPDHSSFFINV 240
Query: 241 GDSLQVMTNGRFRSVKHRVLANGFQSRLSMIYFGGPPLSEKIAPLPSLLKGGKEESLYKE 300
GDSLQVMTNGRF+SV+HRVLANG RLSMIYF GPPLSEKIAPLPSL+KG +ESLYKE
Sbjct: 241 GDSLQVMTNGRFKSVRHRVLANGIDPRLSMIYFCGPPLSEKIAPLPSLMKG--KESLYKE 298
Query: 301 FTWFEYKNSSYGSRLADNRLGHFERIAAS 329
FTWFEYK+S+YGSRLADNRLG++ERIAA+
Sbjct: 299 FTWFEYKSSTYGSRLADNRLGNYERIAAT 327
>UniRef100_O04162 Dioxygenase [Marah macrocarpus]
Length = 322
Score = 468 bits (1204), Expect = e-130
Identities = 234/331 (70%), Positives = 272/331 (81%), Gaps = 11/331 (3%)
Query: 1 MVLLSKQATEQYSYVKNCMATPCSFTIPVVDLSKPDAKSIIVKACEEFGFFKVINHGVSM 60
MV+LSK +Q S + S +P++DLS PDAK +IVKACEE GFFKVINHGV M
Sbjct: 1 MVVLSKPEIQQLS-------SSSSSGVPLIDLSAPDAKHLIVKACEELGFFKVINHGVPM 53
Query: 61 EAISQLESEALNFFSMPVTEKEKTGLANPFGYGNKRIGHNGDVGWVEYLLLKTNQEVNVP 120
E IS LESE+ FFS+P++EK++ G +PFGYGNK+IG NGDVGWVEYLLL++N +
Sbjct: 54 EFISTLESESTKFFSLPLSEKQRAGPPSPFGYGNKQIGRNGDVGWVEYLLLQSNSHAFLS 113
Query: 121 VHSQNPEKFRCVLDDYLCAVRNMGCEILDLMAEGLKIQPKNVLSKLLMDKQSDSVFRVNH 180
+ Q+P+K R L+DY+ AVRNM CEI++LMAEGLKIQ +N LSKLLM ++SDSVFRVNH
Sbjct: 114 IFGQDPQKLRSALNDYIWAVRNMACEIVELMAEGLKIQQRNALSKLLMGEESDSVFRVNH 173
Query: 181 YPACPE--MGLDGENLIGFGEHTDPQIISLLRSNNTSGLQISLRDGSWISVPPDYSSFFI 238
YP CPE L+G N+IGFGEHTDPQIIS+LRSNNTSGLQISL D +WISVPPD +SFFI
Sbjct: 174 YPPCPEELQALEGTNMIGFGEHTDPQIISVLRSNNTSGLQISLPDANWISVPPDQTSFFI 233
Query: 239 NVGDSLQVMTNGRFRSVKHRVLANGFQSRLSMIYFGGPPLSEKIAPLPSLLKGGKEESLY 298
NVGDSLQVMTNGRF+SVKHRVL N +SR+SMIYFGGPPLSEKIAPLPSL+KG EESLY
Sbjct: 234 NVGDSLQVMTNGRFKSVKHRVLTNSLKSRISMIYFGGPPLSEKIAPLPSLMKG--EESLY 291
Query: 299 KEFTWFEYKNSSYGSRLADNRLGHFERIAAS 329
KEFTWFEYK S+Y SRLADNRL HFERIAAS
Sbjct: 292 KEFTWFEYKRSAYNSRLADNRLVHFERIAAS 322
>UniRef100_Q75V70 Gibberellin 2-oxidase 1 [Nicotiana tabacum]
Length = 332
Score = 451 bits (1160), Expect = e-125
Identities = 223/334 (66%), Positives = 271/334 (80%), Gaps = 7/334 (2%)
Query: 1 MVLLSKQATEQYSYVKNCMATPCSFTIPVVDLSKPDAKSIIVKACEEFGFFKVINHGVSM 60
MV+L+K + + VKNC + +P++DLSKP++K++IVKACEEFGFFKVINH V
Sbjct: 1 MVVLTKPGIDHFPIVKNCKLSSFFNGVPLIDLSKPNSKNLIVKACEEFGFFKVINHSVPT 60
Query: 61 EAISQLESEALNFFSMPVTEKEKTGLANPFGYGNKRIGHNGDVGWVEYLLLKTNQEVN-- 118
E I++LESEA+ FFS P++EK+K G A+PFGYGNK+IG NGDVGWVEY+LL TN E N
Sbjct: 61 EFITKLESEAIKFFSSPLSEKQKAGPADPFGYGNKKIGPNGDVGWVEYILLSTNSEFNYH 120
Query: 119 --VPVHSQNPEKFRCVLDDYLCAVRNMGCEILDLMAEGLKIQPKNVLSKLLMDKQSDSVF 176
+ NPE R ++DY+ AV+ M CEIL+++AEGL I P+NV SKLLMD++SDSVF
Sbjct: 121 KFASILGVNPETIRAAVNDYVSAVKKMACEILEMLAEGLNIHPRNVFSKLLMDEKSDSVF 180
Query: 177 RVNHYPACPEMG-LDGENLIGFGEHTDPQIISLLRSNNTSGLQISLRDGSWISVPPDYSS 235
R+NHYP CPE+ NLIGFGEHTDPQIIS+LRSNNTSGLQI L++G WISVPPD +S
Sbjct: 181 RLNHYPPCPEIQQFSDNNLIGFGEHTDPQIISVLRSNNTSGLQILLKNGHWISVPPDPNS 240
Query: 236 FFINVGDSLQVMTNGRFRSVKHRVLANGFQSRLSMIYFGGPPLSEKIAPLPSLLKGGKEE 295
FFINVGDSLQVMTNGRF+SV+HRVLAN +SRLSMIYFGGPPLSEKIAPL SL++G EE
Sbjct: 241 FFINVGDSLQVMTNGRFKSVRHRVLANSVKSRLSMIYFGGPPLSEKIAPLASLMEG--EE 298
Query: 296 SLYKEFTWFEYKNSSYGSRLADNRLGHFERIAAS 329
SLY+EFTWFEYK S+Y +RLADNRL FE++AAS
Sbjct: 299 SLYEEFTWFEYKKSAYKTRLADNRLVLFEKVAAS 332
>UniRef100_Q7X9N5 Gibberellin 2-oxidase [Cucurbita maxima]
Length = 321
Score = 450 bits (1158), Expect = e-125
Identities = 227/322 (70%), Positives = 265/322 (81%), Gaps = 12/322 (3%)
Query: 19 MATPCSFT------IPVVDLSKPDAKSIIVKACEEFGFFKVINHGVSMEAISQLESEALN 72
MA SF+ IP++DLS PDAK +IVKACEE GFFKV+ HGV ME IS LESE+
Sbjct: 1 MAAASSFSAAFYSGIPLIDLSAPDAKQLIVKACEELGFFKVVKHGVPMELISSLESESTK 60
Query: 73 FFSMPVTEKEKTGLANPFGYGNKRIGHNGDVGWVEYLLLKTNQEVN----VPVHSQNPEK 128
FFS+P++EK++ G +PFGYGNK+IG NGDVGWVEYLLL T+ E N + + Q+P+K
Sbjct: 61 FFSLPLSEKQRAGPPSPFGYGNKQIGRNGDVGWVEYLLLNTHLESNSDGFLSIFGQDPQK 120
Query: 129 FRCVLDDYLCAVRNMGCEILDLMAEGLKIQPKNVLSKLLMDKQSDSVFRVNHYPACPEM- 187
R ++DY+ AVRNM EIL+LMAEGLKIQ +NV SKL+MD+QSDSVFRVNHYP CP++
Sbjct: 121 LRSAVNDYISAVRNMAGEILELMAEGLKIQQRNVFSKLVMDEQSDSVFRVNHYPPCPDLQ 180
Query: 188 GLDGENLIGFGEHTDPQIISLLRSNNTSGLQISLRDGSWISVPPDYSSFFINVGDSLQVM 247
L G N+IGFGEHTDPQIIS+LRSNNTSG QISL DG+WISVPPD+SSFFINVGDSLQVM
Sbjct: 181 ALKGTNMIGFGEHTDPQIISVLRSNNTSGFQISLADGNWISVPPDHSSFFINVGDSLQVM 240
Query: 248 TNGRFRSVKHRVLANGFQSRLSMIYFGGPPLSEKIAPLPSLLKGGKEESLYKEFTWFEYK 307
TNGRF+SVKHRVL N +SR+SMIYFGGPPLSEKIAPL SL++ G+E SLYKEFTWFEYK
Sbjct: 241 TNGRFKSVKHRVLTNSSKSRVSMIYFGGPPLSEKIAPLASLMQ-GEERSLYKEFTWFEYK 299
Query: 308 NSSYGSRLADNRLGHFERIAAS 329
S+Y SRLADNRL FERIAAS
Sbjct: 300 RSAYNSRLADNRLVPFERIAAS 321
>UniRef100_Q8GUE8 Gibberellin 2-oxidase [Cucurbita maxima]
Length = 321
Score = 449 bits (1156), Expect = e-125
Identities = 227/322 (70%), Positives = 265/322 (81%), Gaps = 12/322 (3%)
Query: 19 MATPCSFT------IPVVDLSKPDAKSIIVKACEEFGFFKVINHGVSMEAISQLESEALN 72
MA SF+ IP++DLS PDAK +IVKACEE GFFKV+ HGV ME IS LESE+
Sbjct: 1 MAAASSFSAAFYSGIPLIDLSAPDAKQLIVKACEELGFFKVVKHGVPMELISSLESESTK 60
Query: 73 FFSMPVTEKEKTGLANPFGYGNKRIGHNGDVGWVEYLLLKTNQEVN----VPVHSQNPEK 128
FFS+P++EK++ G +PFGYGNK+IG NGDVGWVEYLLL T+ E N + + Q+P+K
Sbjct: 61 FFSLPLSEKQRAGPPSPFGYGNKQIGRNGDVGWVEYLLLNTHLESNSDGFLSMFGQDPQK 120
Query: 129 FRCVLDDYLCAVRNMGCEILDLMAEGLKIQPKNVLSKLLMDKQSDSVFRVNHYPACPEM- 187
R ++DY+ AVRNM EIL+LMAEGLKIQ +NV SKL+MD+QSDSVFRVNHYP CP++
Sbjct: 121 LRSAVNDYISAVRNMAGEILELMAEGLKIQQRNVFSKLVMDEQSDSVFRVNHYPPCPDLQ 180
Query: 188 GLDGENLIGFGEHTDPQIISLLRSNNTSGLQISLRDGSWISVPPDYSSFFINVGDSLQVM 247
L G N+IGFGEHTDPQIIS+LRSNNTSG QISL DG+WISVPPD+SSFFINVGDSLQVM
Sbjct: 181 ALKGTNMIGFGEHTDPQIISVLRSNNTSGFQISLADGNWISVPPDHSSFFINVGDSLQVM 240
Query: 248 TNGRFRSVKHRVLANGFQSRLSMIYFGGPPLSEKIAPLPSLLKGGKEESLYKEFTWFEYK 307
TNGRF+SVKHRVL N +SR+SMIYFGGPPLSEKIAPL SL++ G+E SLYKEFTWFEYK
Sbjct: 241 TNGRFKSVKHRVLTNSSKSRVSMIYFGGPPLSEKIAPLASLMQ-GEERSLYKEFTWFEYK 299
Query: 308 NSSYGSRLADNRLGHFERIAAS 329
S+Y SRLADNRL FERIAAS
Sbjct: 300 RSAYNSRLADNRLVPFERIAAS 321
>UniRef100_Q9FXV7 Gibberellin 2-oxidase No1 [Lactuca sativa]
Length = 337
Score = 438 bits (1126), Expect = e-121
Identities = 223/338 (65%), Positives = 272/338 (79%), Gaps = 10/338 (2%)
Query: 1 MVLLSKQATEQYSYVKNCMAT-PCSF-TIPVVDLSKPDAKSIIVKACEEFGFFKVINHGV 58
MV LSK + EQ+ + T P F TIP++DLSKP++K +VKAC++FGFFKV+NHGV
Sbjct: 1 MVALSKPSIEQFFMKPSKPNTNPLIFPTIPLIDLSKPESKQHLVKACQDFGFFKVVNHGV 60
Query: 59 SMEAISQLESEALNFFSMPVTEKEKTGLANPFGYGNKRIGHNGDVGWVEYLLLKTNQEVN 118
+ I +LESEAL FFS P++ KEK G +PFGYGNKRIG NGDVGWVEYLLL E +
Sbjct: 61 PTKFIKKLESEALKFFSSPLSTKEKAGPPDPFGYGNKRIGRNGDVGWVEYLLLNAKPESD 120
Query: 119 ----VPVHSQNPEKFRCVLDDYLCAVRNMGCEILDLMAEGLKIQPKNVLSKLLMDKQSDS 174
+ V +NPE F+ V++DY+ AV+ M CEIL+L+A+ +K+QP+NV SKLLMD+QSDS
Sbjct: 121 YQRYLSVFEENPEIFQGVVNDYVTAVKKMACEILELLADEMKLQPRNVFSKLLMDEQSDS 180
Query: 175 VFRVNHYPACPEMG---LDGENLIGFGEHTDPQIISLLRSNNTSGLQISLRDGSWISVPP 231
VFRVNHY CPE +G L+GFGEHTDPQIIS+LRSNNTSGL+ISLRDGSW+SVP
Sbjct: 181 VFRVNHYLPCPEFQENERNGRKLVGFGEHTDPQIISVLRSNNTSGLEISLRDGSWMSVPA 240
Query: 232 DYSSFFINVGDSLQVMTNGRFRSVKHRVLANGFQSRLSMIYFGGPPLSEKIAPLPSLLKG 291
D SFFINVGDSLQVMTNGRF+SVKHRV+AN +SR+SMIYFGGPPLSEKIAPLPSL++
Sbjct: 241 DSDSFFINVGDSLQVMTNGRFKSVKHRVVANSTKSRVSMIYFGGPPLSEKIAPLPSLIQ- 299
Query: 292 GKEESLYKEFTWFEYKNSSYGSRLADNRLGHFERIAAS 329
G+E+SLYKEFTWFEYK S++ +RLADNRLG FE+I A+
Sbjct: 300 GEEDSLYKEFTWFEYKKSAFNTRLADNRLGLFEKITAT 337
>UniRef100_Q75V69 Gibberellin 2-oxidase 2 [Nicotiana tabacum]
Length = 323
Score = 431 bits (1108), Expect = e-119
Identities = 216/325 (66%), Positives = 262/325 (80%), Gaps = 8/325 (2%)
Query: 10 EQYSYVKNCMATPCSFTIPVVDLSKPDAKSIIVKACEEFGFFKVINHGVSMEAISQLESE 69
+Q+ NC T +P++DLSKPD+K++IVKACEEFGFFKVINH V E I++L SE
Sbjct: 2 DQHFSKDNCKPTSFFNNVPLIDLSKPDSKNLIVKACEEFGFFKVINHSVPTEFITKL-SE 60
Query: 70 ALNFFSMPVTEKEKTGLANPFGYGNKRIGHNGDVGWVEYLLLKTNQEVN----VPVHSQN 125
A+ FFS P++EKEK G +PFGYGNK+IG NGD GWVE++L+ TN E N + N
Sbjct: 61 AIKFFSSPLSEKEKAGPPDPFGYGNKQIGCNGDSGWVEHILVSTNSEFNYYKFASILGVN 120
Query: 126 PEKFRCVLDDYLCAVRNMGCEILDLMAEGLKIQPKNVLSKLLMDKQSDSVFRVNHYPACP 185
PE R ++DY+ AV+ M CEIL+++AEGLKI P+NV SKLLMD++SDSVFR+NHYP C
Sbjct: 121 PENIRAAVNDYVSAVKKMACEILEMLAEGLKIHPRNVFSKLLMDEKSDSVFRLNHYPPCT 180
Query: 186 EMG-LDGENLIGFGEHTDPQIISLLRSNNTSGLQISLRDGSWISVPPDYSSFFINVGDSL 244
E+ + NLIGFGEHTDPQIIS+LRSNNTSGLQI L++G WISVPPD +SFF+NVGDSL
Sbjct: 181 EIQQFNDNNLIGFGEHTDPQIISVLRSNNTSGLQILLKNGHWISVPPDPNSFFVNVGDSL 240
Query: 245 QVMTNGRFRSVKHRVLANGFQSRLSMIYFGGPPLSEKIAPLPSLLKGGKEESLYKEFTWF 304
QVMTNG+F+SVKHRVLAN +SRLSMIYFGGPPLSEKIAPL SL++G E+SLYKEFTWF
Sbjct: 241 QVMTNGKFKSVKHRVLANSMKSRLSMIYFGGPPLSEKIAPLASLMEG--EDSLYKEFTWF 298
Query: 305 EYKNSSYGSRLADNRLGHFERIAAS 329
EYK S+Y +RLADNRL FE+IAAS
Sbjct: 299 EYKKSAYKTRLADNRLILFEKIAAS 323
>UniRef100_Q9XG83 Gibberellin 2-beta-dioxygenase [Phaseolus coccineus]
Length = 332
Score = 417 bits (1071), Expect = e-115
Identities = 207/332 (62%), Positives = 260/332 (77%), Gaps = 5/332 (1%)
Query: 1 MVLLSKQATEQYSYVKNCMATPCSFTIPVVDLSKPDAKSIIVKACEEFGFFKVINHGVSM 60
MV+LS+ A Q+ +K +TP IPVVDL+ PDAK++IV AC +FGFFK++NHGV +
Sbjct: 1 MVVLSQPALNQFFLLKPFKSTPLFTGIPVVDLTHPDAKNLIVNACRDFGFFKLVNHGVPL 60
Query: 61 EAISQLESEALNFFSMPVTEKEKTGLANPFGYGNKRIGHNGDVGWVEYLLLKTNQEVNVP 120
E ++ LE+EAL FF +EK++ G +PFGYG+KRIG NGDVGWVEYLLL TN +V P
Sbjct: 61 ELMANLENEALRFFKKSQSEKDRAGPPDPFGYGSKRIGPNGDVGWVEYLLLNTNPDVISP 120
Query: 121 ----VHSQNPEKFRCVLDDYLCAVRNMGCEILDLMAEGLKIQPKNVLSKLLMDKQSDSVF 176
+ +NP FR V+++Y+ AV+NM +L+LMAEGL I+ +N LS+LL D++SDS F
Sbjct: 121 KSLCIFRENPHHFRAVVENYITAVKNMCYAVLELMAEGLGIRQRNTLSRLLKDEKSDSCF 180
Query: 177 RVNHYPACPEMGLDGENLIGFGEHTDPQIISLLRSNNTSGLQISLRDGSWISVPPDYSSF 236
R+NHYP CPE+ NL+GFGEHTDPQIIS+LRSN+TSGLQI L DG+W+SVPPD +SF
Sbjct: 181 RLNHYPPCPEVQALNRNLVGFGEHTDPQIISVLRSNSTSGLQICLTDGTWVSVPPDQTSF 240
Query: 237 FINVGDSLQVMTNGRFRSVKHRVLANGFQSRLSMIYFGGPPLSEKIAPLPSLLKGGKEES 296
FINVGD+LQVMTNGRF+SVKHRVLA+ +SRLSMIYFGGP LSE IAPLPS++ G EE
Sbjct: 241 FINVGDALQVMTNGRFKSVKHRVLADTTKSRLSMIYFGGPALSENIAPLPSVMLKG-EEC 299
Query: 297 LYKEFTWFEYKNSSYGSRLADNRLGHFERIAA 328
LYKEFTW EYK ++Y SRLADNRL F++ AA
Sbjct: 300 LYKEFTWCEYKKAAYTSRLADNRLAPFQKSAA 331
>UniRef100_Q8GSN6 Gibberellin 2-oxidase 1 [Spinacia oleracea]
Length = 337
Score = 384 bits (987), Expect = e-105
Identities = 195/334 (58%), Positives = 246/334 (73%), Gaps = 14/334 (4%)
Query: 1 MVLLSKQATEQYSYVKNCMATPCSF---TIPVVDLSKPDAKSIIVKACEEFGFFKVINHG 57
MV+LS A +Q+ +K C IP +DL+ PD KS+IVKAC+EFGFFKV+NHG
Sbjct: 1 MVVLSHVALDQF--MKTCKPIDTHLFPTVIPTIDLNSPDVKSLIVKACKEFGFFKVVNHG 58
Query: 58 VSMEAISQLESEALNFFSMPVTEKEKTGLANPFGYGNKRIGHNGDVGWVEYLLLKTNQE- 116
V +E QLE EA FFS+P KE G NPFGYGNK+IG NGDVGW+EYLLL + E
Sbjct: 59 VPIEMTRQLEDEASKFFSLPQGLKESAGNPNPFGYGNKKIGRNGDVGWIEYLLLSASDEF 118
Query: 117 ---VNVPVHSQNPEKFRCVLDDYLCAVRNMGCEILDLMAEGLKIQPKN--VLSKLLMDKQ 171
+ ++ NP+ FRC L++Y+ ++ MG +L+ MAEGL ++ ++ LS L+ D +
Sbjct: 119 ISQTCLSIYPDNPDVFRCALNNYISKMKKMGVRVLETMAEGLNLKGEDRYALSNLIKDSK 178
Query: 172 SDSVFRVNHYPACPEM--GLDGENLIGFGEHTDPQIISLLRSNNTSGLQISLRDGSWISV 229
SDS FR+NHYP PE G++G NL+GFGEHTDPQIIS+LRSN GLQISL DG+W+SV
Sbjct: 179 SDSYFRLNHYPPRPEAFEGINGRNLVGFGEHTDPQIISVLRSNKIGGLQISLNDGTWVSV 238
Query: 230 PPDYSSFFINVGDSLQVMTNGRFRSVKHRVLANGFQSRLSMIYFGGPPLSEKIAPLPSLL 289
PPD +SFFI VGDSLQVMTNGRF+SVKHRVLA+ +SRLSMI+F GPPL +KIAPLP ++
Sbjct: 239 PPDDNSFFILVGDSLQVMTNGRFKSVKHRVLADNMKSRLSMIFFAGPPLCQKIAPLPCIM 298
Query: 290 KGGKEESLYKEFTWFEYKNSSYGSRLADNRLGHF 323
+ G EESLY+EFTW EYK S+Y SRL++NRL F
Sbjct: 299 QKG-EESLYEEFTWCEYKKSAYKSRLSENRLLRF 331
>UniRef100_Q8LEA2 Gibberellin 2-beta-dioxygenase 1 [Arabidopsis thaliana]
Length = 329
Score = 373 bits (957), Expect = e-102
Identities = 184/306 (60%), Positives = 236/306 (76%), Gaps = 9/306 (2%)
Query: 27 IPVVDLSKPDAKSIIVKACEEFGFFKVINHGVSMEAISQLESEALNFFSMPVTEKEKTGL 86
IPV+D+S P++K +VKACE+FGFFKVINHGVS E +S LE E ++FFS+P +EK +
Sbjct: 18 IPVIDMSDPESKHALVKACEDFGFFKVINHGVSAELVSVLEHETVDFFSLPKSEKTQVA- 76
Query: 87 ANPFGYGNKRIGHNGDVGWVEYLLLKTNQEVN----VPVHSQNPEKFRCVLDDYLCAVRN 142
PFGYGN +IG NGDVGWVEYLL+ N + P ++P FR L++Y +VR
Sbjct: 77 GYPFGYGNSKIGRNGDVGWVEYLLMNANHDSGSGPLFPSLLKSPGTFRNALEEYTTSVRK 136
Query: 143 MGCEILDLMAEGLKIQPKNVLSKLLMDKQSDSVFRVNHYPACP---EMGLDGENLIGFGE 199
M ++L+ + +GL I+P+N LSKL+ D+ +DS+ R+NHYP CP + G+N+IGFGE
Sbjct: 137 MTFDVLEKITDGLGIKPRNTLSKLVSDQNTDSILRLNHYPPCPLSNKKTNGGKNVIGFGE 196
Query: 200 HTDPQIISLLRSNNTSGLQISLRDGSWISVPPDYSSFFINVGDSLQVMTNGRFRSVKHRV 259
HTDPQIIS+LRSNNTSGLQI+L DGSWISVPPD++SFF NVGDSLQVMTNGRF+SV+HRV
Sbjct: 197 HTDPQIISVLRSNNTSGLQINLNDGSWISVPPDHTSFFFNVGDSLQVMTNGRFKSVRHRV 256
Query: 260 LANGFQSRLSMIYFGGPPLSEKIAPLPSLLKGGKEESLYKEFTWFEYKNSSYGSRLADNR 319
LAN +SR+SMIYF GP L+++IAPL L+ ++E LY+EFTW EYKNS+Y SRL+DNR
Sbjct: 257 LANCKKSRVSMIYFAGPSLTQRIAPLTCLI-DNEDERLYEEFTWSEYKNSTYNSRLSDNR 315
Query: 320 LGHFER 325
L FER
Sbjct: 316 LQQFER 321
>UniRef100_Q9XFR9 Gibberellin 2-beta-dioxygenase 2 [Arabidopsis thaliana]
Length = 341
Score = 370 bits (950), Expect = e-101
Identities = 181/308 (58%), Positives = 244/308 (78%), Gaps = 10/308 (3%)
Query: 24 SFTIPVVDLSKPDAKSIIVKACEEFGFFKVINHGVSMEAISQLESEALNFFSMPVTEKEK 83
S +IPVV+L+ P+AK+ IVKACEEFGFFKV+NHGV E +++LE EA+ FF +P + K +
Sbjct: 28 SHSIPVVNLADPEAKTRIVKACEEFGFFKVVNHGVRPELMTRLEQEAIGFFGLPQSLKNR 87
Query: 84 TGLANPFGYGNKRIGHNGDVGWVEYLLLKTNQEVNVP----VHSQNPEKFRCVLDDYLCA 139
G P+GYGNKRIG NGDVGW+EYLLL N +++ P V Q P+ FR +++Y+
Sbjct: 88 AGPPEPYGYGNKRIGPNGDVGWIEYLLLNANPQLSSPKTSAVFRQTPQIFRESVEEYMKE 147
Query: 140 VRNMGCEILDLMAEGLKIQPKNVLSKLLMDKQSDSVFRVNHYPACPEMGLDGENLI--GF 197
++ + ++L+++AE L I+P++ LSK+L D++SDS R+NHYPA E + E ++ GF
Sbjct: 148 IKEVSYKVLEMVAEELGIEPRDTLSKMLRDEKSDSCLRLNHYPAAEE---EAEKMVKVGF 204
Query: 198 GEHTDPQIISLLRSNNTSGLQISLRDGSWISVPPDYSSFFINVGDSLQVMTNGRFRSVKH 257
GEHTDPQIIS+LRSNNT+GLQI ++DGSW++VPPD+SSFFINVGD+LQVMTNGRF+SVKH
Sbjct: 205 GEHTDPQIISVLRSNNTAGLQICVKDGSWVAVPPDHSSFFINVGDALQVMTNGRFKSVKH 264
Query: 258 RVLANGFQSRLSMIYFGGPPLSEKIAPLPSLLKGGKEESLYKEFTWFEYKNSSYGSRLAD 317
RVLA+ +SR+SMIYFGGPPLS+KIAPLP L+ +++ LYKEFTW +YK+S+Y S+L D
Sbjct: 265 RVLADTRRSRISMIYFGGPPLSQKIAPLPCLVP-EQDDWLYKEFTWSQYKSSAYKSKLGD 323
Query: 318 NRLGHFER 325
RLG FE+
Sbjct: 324 YRLGLFEK 331
>UniRef100_Q6F6H4 Gibberellin 2-oxidase1 [Daucus carota]
Length = 317
Score = 370 bits (949), Expect = e-101
Identities = 196/331 (59%), Positives = 239/331 (71%), Gaps = 23/331 (6%)
Query: 1 MVLLSKQATE-QYSYVKNCMATPCSFTIPVVDLSKPDAKSIIVKACEEFGFFKVINHGVS 59
MV+LS+ + +K+C S IPVVDLS PDAK+ IV AC+EFGFFKVINHGV
Sbjct: 1 MVVLSQPVEHVNFPAIKSCNGN--STHIPVVDLSNPDAKTQIVNACQEFGFFKVINHGVP 58
Query: 60 MEAISQLESEALNFFSMPVTEKEKTGLANPFGYGNKRIGHNGDVGWVEYLLLKTNQEVNV 119
ME +++LE++AL+FF P K K A PFGYGNK IG GD GWVEYLL T+ ++
Sbjct: 59 MEIVTKLEAQALSFFKQPQNHKNK---AAPFGYGNKNIGRKGDTGWVEYLLFGTDTQLI- 114
Query: 120 PVHSQN-----PEKFRCVLDDYLCAVRNMGCEILDLMAEGLKIQPKNVLSKLLMDKQSDS 174
SQN P F +++ YL AV+N+ CEIL LMAE L +QPKNVLS+LL DK+SDS
Sbjct: 115 ---SQNSLTILPNDFWAMVNKYLSAVKNLACEILKLMAEELNLQPKNVLSRLLSDKKSDS 171
Query: 175 VFRVNHYPACPEMGLDGENLIGFGEHTDPQIISLLRSNNTSGLQISLRDGSWISVPPDYS 234
FR+NHYPA N +GFGEHTDPQIIS++RSNN SGL+I+L+DG+W VP D S
Sbjct: 172 FFRINHYPA------SDRNELGFGEHTDPQIISVIRSNNASGLEIALKDGTWTQVPADPS 225
Query: 235 SFFINVGDSLQVMTNGRFRSVKHRVLANGFQSRLSMIYFGGPPLSEKIAPLPSLLKGGKE 294
SFFI V D LQVMTNGRFRSVKHRV+ + RLSMIYFGGPPLSEKIAP SL++G E
Sbjct: 226 SFFITVDDCLQVMTNGRFRSVKHRVITESLKERLSMIYFGGPPLSEKIAPSTSLMQG--E 283
Query: 295 ESLYKEFTWFEYKNSSYGSRLADNRLGHFER 325
ESLYKEFTW EYK +++ ++LA NR+ FE+
Sbjct: 284 ESLYKEFTWGEYKKAAFNTKLAFNRISFFEK 314
>UniRef100_Q9FXV6 Gibberellin 2-oxidase No2 [Lactuca sativa]
Length = 338
Score = 362 bits (929), Expect = 8e-99
Identities = 189/339 (55%), Positives = 246/339 (71%), Gaps = 21/339 (6%)
Query: 1 MVLLSKQATEQYSYVKNCMATPCSFT-IPVVDLSKP-DAKSIIVKACEEFGFFKVINHGV 58
MV++S+ + + + +FT IPV+DLS P DAK++IV AC+++GFFKVINHGV
Sbjct: 1 MVMISQVEKKPVVDIFPTKSNKTNFTGIPVIDLSSPEDAKNLIVNACQDYGFFKVINHGV 60
Query: 59 SMEAISQLESEALNFFSMPVTEKEKTGLANPFGYGNKRIGHNGDVGWVEYLLLKTNQEVN 118
S +S+LE+EA+ FF+M EK++ NPFGYG +IG NGDVGW+EYLLL + N
Sbjct: 61 SPLLVSELENEAIEFFNMRQYEKDQYCPPNPFGYGRNKIGTNGDVGWIEYLLLTST---N 117
Query: 119 VPVHSQNPEKFRCVLDDYLCAVRNMGCEILDLMAEGLKIQPKNVLSKLLMDKQSDSVFRV 178
P +S+ F + ++Y+ VR +GC IL+LMAEGLKI+PKNVLS++L D+ +D+VFR+
Sbjct: 118 FPTNSKI---FSSLTNEYVKEVRKLGCTILELMAEGLKIEPKNVLSRMLSDENADTVFRL 174
Query: 179 NHYPAC--PEMGLD----------GENLIGFGEHTDPQIISLLRSNNTSGLQISLRDGSW 226
NHYP C P D G IGFGEHTDPQ+IS+ RSN TSG QI L DG+W
Sbjct: 175 NHYPPCLDPNSNHDSDLNNKSMSHGRTSIGFGEHTDPQLISIARSNATSGFQIYLEDGTW 234
Query: 227 ISVPPDYSSFFINVGDSLQVMTNGRFRSVKHRVLANGFQSRLSMIYFGGPPLSEKIAPLP 286
+ VPPD +S+FINV D L+VMTNGRFRSV+HRV+A+ F+SR+SMIYFGGPPL EKI+PL
Sbjct: 235 VGVPPDETSYFINVDDLLEVMTNGRFRSVRHRVVADSFKSRVSMIYFGGPPLMEKISPLG 294
Query: 287 SLLKGGKEESLYKEFTWFEYKNSSYGSRLADNRLGHFER 325
SL++ G EESLY EFTWFEYK+ +Y +RL DNRL F +
Sbjct: 295 SLMEPG-EESLYNEFTWFEYKSCTYKTRLTDNRLSLFHK 332
>UniRef100_Q8S0S6 Putative GA 2-oxidase [Oryza sativa]
Length = 327
Score = 346 bits (888), Expect = 4e-94
Identities = 169/300 (56%), Positives = 220/300 (73%), Gaps = 2/300 (0%)
Query: 27 IPVVDLSKPDAKSIIVKACEEFGFFKVINHGVSMEAISQLESEALNFFSMPVTEKEKTGL 86
+PVVDL P A +V ACE +GFFKV+NHGV+ + + + ESEA+ FFS +K+++G
Sbjct: 28 VPVVDLGSPGAARAVVDACERYGFFKVVNHGVATDTMDKAESEAVRFFSQTQPDKDRSGP 87
Query: 87 ANPFGYGNKRIGHNGDVGWVEYLLLKTNQEVNVPVHS-QNPEKFRCVLDDYLCAVRNMGC 145
A PFGYG+KRIG NGD+GW+EYLLL + + + FR L++Y+ VR +
Sbjct: 88 AYPFGYGSKRIGFNGDMGWLEYLLLALDDASLADACTVPSCAVFRAALNEYISGVRKVAV 147
Query: 146 EILDLMAEGLKIQPKNVLSKLLMDKQSDSVFRVNHYPACPEMGLDGENLIGFGEHTDPQI 205
+++ M+EGL I + LS L+ + SD VFRVNHYP C + G ++ GFGEHTDPQ+
Sbjct: 148 RVMEAMSEGLGIAQADALSALVTAEGSDQVFRVNHYPPCRALQGLGCSVTGFGEHTDPQL 207
Query: 206 ISLLRSNNTSGLQISLRDGSWISVPPDYSSFFINVGDSLQVMTNGRFRSVKHRVLANGFQ 265
+S+LRSN TSGLQI+LRDG W+SVP D SFF+NVGDSLQV+TNGRF+SVKHRV+AN +
Sbjct: 208 VSVLRSNGTSGLQIALRDGQWVSVPSDRDSFFVNVGDSLQVLTNGRFKSVKHRVVANSLK 267
Query: 266 SRLSMIYFGGPPLSEKIAPLPSLLKGGKEESLYKEFTWFEYKNSSYGSRLADNRLGHFER 325
SR+S IYFGGPPL+++IAPLP LL G E+SLYKEFTW EYK ++Y SRL DNRL FE+
Sbjct: 268 SRVSFIYFGGPPLAQRIAPLPQLL-GEGEQSLYKEFTWDEYKKAAYKSRLGDNRLAQFEK 326
>UniRef100_O64692 Gibberellin 2-beta-dioxygenase 3 [Arabidopsis thaliana]
Length = 335
Score = 342 bits (876), Expect = 1e-92
Identities = 175/329 (53%), Positives = 233/329 (70%), Gaps = 7/329 (2%)
Query: 1 MVLLSKQATEQYSYVKNCMATPCSFTIPVVDLSKPDAKSIIVKACEEFGFFKVINHGVSM 60
MV++ + A+ + N P IPV+DL+ DAK+ IVKACEEFGFFKVINHGV
Sbjct: 1 MVIVLQPASFDSNLYVNPKCKPRPVLIPVIDLTDSDAKTQIVKACEEFGFFKVINHGVRP 60
Query: 61 EAISQLESEALNFFSMPVTEKEKTGLANPFGYGNKRIGHNGDVGWVEYLLLKTN----QE 116
+ ++QLE EA+NFF++ + K+K G +PFGYG KRIG NGD+GW+EY+LL N
Sbjct: 61 DLLTQLEQEAINFFALHHSLKDKAGPPDPFGYGTKRIGPNGDLGWLEYILLNANLCLESH 120
Query: 117 VNVPVHSQNPEKFRCVLDDYLCAVRNMGCEILDLMAEGLKIQPKNVLSKLLMDKQSDSVF 176
+ P FR +++Y+ ++ M + L+++ E LKI+PK LS+L+ K+SDS
Sbjct: 121 KTTAIFRHTPAIFREAVEEYIKEMKRMSSKFLEMVEEELKIEPKEKLSRLVKVKESDSCL 180
Query: 177 RVNHYPACPEMGLDGENLIGFGEHTDPQIISLLRSNNTSGLQISLRDGSWISVPPDYSSF 236
R+NHYP E + E IGFGEHTDPQ+ISLLRSN+T GLQI ++DG+W+ V PD+SSF
Sbjct: 181 RMNHYPEKEETPVKEE--IGFGEHTDPQLISLLRSNDTEGLQICVKDGTWVDVTPDHSSF 238
Query: 237 FINVGDSLQVMTNGRFRSVKHRVLANGFQSRLSMIYFGGPPLSEKIAPLPSLLKGGKEES 296
F+ VGD+LQVMTNGRF+SVKHRV+ N +SR+SMIYF GPPLSEKIAPL S L +++
Sbjct: 239 FVLVGDTLQVMTNGRFKSVKHRVVTNTKRSRISMIYFAGPPLSEKIAPL-SCLVPKQDDC 297
Query: 297 LYKEFTWFEYKNSSYGSRLADNRLGHFER 325
LY EFTW +YK S+Y ++L D RLG FE+
Sbjct: 298 LYNEFTWSQYKLSAYKTKLGDYRLGLFEK 326
>UniRef100_Q673G3 GA 2-oxidase 4 [Hordeum vulgare var. distichum]
Length = 342
Score = 335 bits (860), Expect = 8e-91
Identities = 166/328 (50%), Positives = 230/328 (69%), Gaps = 4/328 (1%)
Query: 1 MVLLSKQAT-EQYSYVKNCMATPCSFTIPVVDLSKPDAKSIIVKACEEFGFFKVINHGVS 59
MV+L+K A EQ + ++ +P VDLS P A + +V+ACE FGFF V+NHGV
Sbjct: 1 MVVLAKPAALEQIALMRTPEPWESFSGVPAVDLSSPGAAADVVRACERFGFFSVVNHGVP 60
Query: 60 MEAISQLESEALNFFSMPVTEKEKTGLANPFGYGNKRIGHNGDVGWVEYLLLKTNQEVNV 119
+ +LE+EA+ FF+ EK+ +G A+PFGYG+KRIG NGD+GWVEYLLL +++
Sbjct: 61 AGVVDRLEAEAVRFFASTQAEKDASGPADPFGYGSKRIGRNGDMGWVEYLLLAIDRDTLS 120
Query: 120 PVHSQNPEKFRCVLDDYLCAVRNMGCEILDLMAEGLKIQPKNVLSKLLMDKQSDSVFRVN 179
R ++ Y+ A+R + +L+++AEGL + P+ L+ ++ + SD VFRVN
Sbjct: 121 KASPAPSSALREAINAYVSAMRGLARTVLEMVAEGLGVSPRGALADMVTGEASDQVFRVN 180
Query: 180 HYPACPEM-GLDGE-NLIGFGEHTDPQIISLLRSNNTSGLQISLRDGSWISVPPDYSSFF 237
HYP CP + GL ++ GFGEHTDPQ++S+L SN T+GLQ++L DG W+SVPP+ +FF
Sbjct: 181 HYPPCPLLQGLPPNCSVTGFGEHTDPQLVSILHSNGTAGLQVALHDGRWVSVPPNRDAFF 240
Query: 238 INVGDSLQVMTNGRFRSVKHRVLA-NGFQSRLSMIYFGGPPLSEKIAPLPSLLKGGKEES 296
+NVGDSLQV+TNGR RSV+HRV+A NG +SR+SMIYF GPPL+++IAPL LL G +
Sbjct: 241 VNVGDSLQVLTNGRLRSVRHRVVAGNGLKSRVSMIYFAGPPLAQRIAPLQQLLAGTQSLP 300
Query: 297 LYKEFTWFEYKNSSYGSRLADNRLGHFE 324
LY++FTW EYK ++Y SRL DNRL FE
Sbjct: 301 LYRDFTWGEYKKAAYRSRLGDNRLAPFE 328
>UniRef100_Q673G2 GA 2-oxidase 5 [Hordeum vulgare var. distichum]
Length = 341
Score = 305 bits (782), Expect = 9e-82
Identities = 170/347 (48%), Positives = 220/347 (62%), Gaps = 30/347 (8%)
Query: 1 MVLLSKQATEQYSYVKNCMATPCSFTIPVVDLS------KPDAKSIIVKACEEFGFFKVI 54
MV+L+K EQ + A P + +DLS + A +V ACEE GFFKV
Sbjct: 1 MVVLAKGELEQIALPA---AQPPLAHVRAIDLSAAPGPGRDAAARALVSACEEQGFFKVT 57
Query: 55 NHGVSMEAISQLESEALNFFSMPVTEKEKTGLANPFGYGNKRIGHNGDVGWVEYLLLKTN 114
HGV +E + +E+ A FF++P EKE P GYG+KRIG +GD+GW+EYLLL
Sbjct: 58 GHGVPLELVRAVEAAAAEFFALPQAEKEAAA-GRPLGYGSKRIGVSGDLGWIEYLLLGVT 116
Query: 115 QEVNVPV-----------------HSQNPEKFRCVLDDYLCAVRNMGCEILDLMAEGLKI 157
+P S++P R +L++Y AVR M C +L+LMAEGL I
Sbjct: 117 PAGALPAASFASWTLPCAGAAVASSSEHPCPLRDLLEEYAAAVRRMACGVLELMAEGLGI 176
Query: 158 QPKNVLSKLLMDKQSDSVFRVNHYPACPEMGLDGENLIGFGEHTDPQIISLLRSNNTSGL 217
P + LS+L+ D +SD++ RVNHYP PEM G L GFGEHTDPQIIS+LRSN TSGL
Sbjct: 177 GPADALSRLVSDGESDNMLRVNHYPPRPEM--QGCLLTGFGEHTDPQIISVLRSNGTSGL 234
Query: 218 QISLRDGSWISVPPDYSSFFINVGDSLQVMTNGRFRSVKHRVLANGFQSRLSMIYFGGPP 277
+I RDG W SVPPD +FF+NV D+LQV+TNGRF SVKHRV+ + + R+SMI+FGGPP
Sbjct: 235 EICARDGEWTSVPPDPDAFFVNVADALQVLTNGRFSSVKHRVVVSSERPRVSMIFFGGPP 294
Query: 278 LSEKIAPLPSLLKGGKEESLYKEFTWFEYKNSSYGSRLADNRLGHFE 324
+ E++APL LL G S Y+EFTW EYK+S++ RLA +RL FE
Sbjct: 295 MGERLAPLRQLL-GDGGRSKYREFTWKEYKSSTHKGRLATDRLCSFE 340
>UniRef100_Q9XHM5 Gibberellin 2-beta-dioxygenase 2 [Pisum sativum]
Length = 345
Score = 305 bits (781), Expect = 1e-81
Identities = 154/314 (49%), Positives = 209/314 (66%), Gaps = 17/314 (5%)
Query: 27 IPVVDLS--KPDAKSIIVKACEEFGFFKVINHGVSMEAISQLESEALNFFSMPVTEKEKT 84
IP +DLS + ++VKACEE+GFFKV+NH V E IS+L+ E + FFS +EK +
Sbjct: 20 IPTIDLSLERSQLSELVVKACEEYGFFKVVNHSVPKEVISRLDEEGIEFFSKNSSEKRQA 79
Query: 85 GLANPFGYGNKRIGHNGDVGWVEYLLLKTN----QEVNVPVHSQNPEKFRCVLDDYLCAV 140
G + PFGYG K IG NGD G +EYLLL +N E + + +P KF C+++DY+ AV
Sbjct: 80 GTSTPFGYGCKNIGPNGDKGELEYLLLHSNPISISERSKTIAKDHPIKFSCIVNDYIKAV 139
Query: 141 RNMGCEILDLMAEGLKIQPKNVLSKLLMDKQSDSVFRVNHYPACPEMGLDG--------- 191
+++ CEIL+L AEGL + K+ LSK++ D+ SDS+ R+NHYP ++G D
Sbjct: 140 KDLTCEILELAAEGLWVPDKSSLSKIIKDEHSDSLLRINHYPPVKKLGNDNWDPSKIQNS 199
Query: 192 -ENLIGFGEHTDPQIISLLRSNNTSGLQISLRDGSWISVPPDYSSFFINVGDSLQVMTNG 250
N IGFGEH+DPQI+++LRSNN GLQIS G WI VPPD S F++ VGD+LQV+TNG
Sbjct: 200 NNNNIGFGEHSDPQILTILRSNNVGGLQISTHHGLWIPVPPDPSEFYVMVGDALQVLTNG 259
Query: 251 RFRSVKHRVLANGFQSRLSMIYFGGPPLSEKIAPLPSLLKGGKEESLYKEFTWFEYKNSS 310
RF SV+HRVL N + R+SM+YF PPL+ I+PL ++ LY+ FTW +YK ++
Sbjct: 260 RFVSVRHRVLTNTTKPRMSMMYFAAPPLNWLISPLSKMVT-AHSPCLYRPFTWAQYKQAA 318
Query: 311 YGSRLADNRLGHFE 324
Y RL D RL F+
Sbjct: 319 YALRLGDTRLDQFK 332
>UniRef100_Q6TN17 Gibberellin 2-oxidase [Populus alba x Populus tremuloides]
Length = 335
Score = 302 bits (773), Expect = 9e-81
Identities = 154/309 (49%), Positives = 209/309 (66%), Gaps = 13/309 (4%)
Query: 27 IPVVDLS--KPDAKSIIVKACEEFGFFKVINHGVSMEAISQLESEALNFFSMPVTEKEKT 84
+PV+DLS + ++IVKACEE+GFFKV NHGV + I+++E+E+ NFF+ EK+K
Sbjct: 19 LPVIDLSGERSMVSNLIVKACEEYGFFKVRNHGVPHDIIARMENESSNFFAKTFDEKQKA 78
Query: 85 GLANPFGYGNKRIGHNGDVGWVEYLLLKTNQ---EVNVPVHSQNPEKFRCVLDDYLCAVR 141
GLAN FGYG K IG NGD G VEYLL TN S +P +F + Y+ AVR
Sbjct: 79 GLANSFGYGCKNIGFNGDTGEVEYLLFNTNPLSIAERSKTISNDPTEFSSAMSGYIEAVR 138
Query: 142 NMGCEILDLMAEGLKIQPKNVLSKLLMDKQSDSVFRVNHYPACPEMGLDGE-------NL 194
+ CE+LDLMAEGL + ++V S+L+ D SDS+ R+NHYP P + D + N
Sbjct: 139 ELACELLDLMAEGLWVPDRSVFSRLIRDDDSDSIIRLNHYPPMPILCKDKDSSSPCNHNR 198
Query: 195 IGFGEHTDPQIISLLRSNNTSGLQISLRDGSWISVPPDYSSFFINVGDSLQVMTNGRFRS 254
+GFGEH+DPQI+++LRSN+ GLQISL DG W+ V PD S+F +NVGD LQ MTNGRF S
Sbjct: 199 VGFGEHSDPQILTILRSNDVGGLQISLNDGVWVPVTPDPSAFCVNVGDLLQAMTNGRFVS 258
Query: 255 VKHRVLANGFQSRLSMIYFGGPPLSEKIAPLPSLLKGGKEESLYKEFTWFEYKNSSYGSR 314
V+H+ L N ++SR+SM YF PPL+ +IA P ++ K +LY+ F+W E+K +++ R
Sbjct: 259 VRHKALTNSYKSRMSMAYFAAPPLNARIAVPPEMVTPIK-PALYRPFSWAEFKKAAFALR 317
Query: 315 LADNRLGHF 323
L D+RLG F
Sbjct: 318 LGDSRLGLF 326
>UniRef100_Q688Y4 Putative GA2-oxidase [Oryza sativa]
Length = 353
Score = 296 bits (759), Expect = 4e-79
Identities = 163/313 (52%), Positives = 209/313 (66%), Gaps = 18/313 (5%)
Query: 27 IPVVDLSKPDAKSIIVKACEEFGFFKVINHGVSMEAISQLESEALNFFSMPVTEK-EKTG 85
IP +DLS P A + + AC GFFK NHGV LES A+ FF++P EK + +G
Sbjct: 30 IPCIDLSAPGAAAAVADACRTLGFFKATNHGVPAGLADALESSAMAFFALPHQEKLDMSG 89
Query: 86 LANPFGYGNKRIGHNGDVGWVEYLLLKTNQEVNVPVHSQNPEKFRCVLDDYLCAVRNMGC 145
A P GYG+K IG NGDVGW+EYLLL + + P R ++ Y AVR +GC
Sbjct: 90 PARPLGYGSKSIGSNGDVGWLEYLLLSAGAASSGG--AALPAALRAAVEAYTGAVRGVGC 147
Query: 146 EILDLMAEGLKI----QPKNVLSKLLMDKQ-SDSVFRVNHYPAC---PEMGLDGENLIGF 197
+++LMAEGL + + + VL ++++ + SD + RVNHYP C P D + GF
Sbjct: 148 RVMELMAEGLGLGASEEGRCVLRRMVVGCEGSDEMLRVNHYPPCLLPPGRDRDECGVTGF 207
Query: 198 GEHTDPQIISLLRSNNTSGLQISLRD-----GSWISVPPDYSSFFINVGDSLQVMTNGRF 252
GEHTDPQIIS+LRSN T+GLQI LR W+ VPPD SFF+NVGDSLQV+TNGRF
Sbjct: 208 GEHTDPQIISVLRSNCTAGLQILLRGDYSSPARWVPVPPDPDSFFVNVGDSLQVLTNGRF 267
Query: 253 RSVKHRVLA-NGFQSRLSMIYFGGPPLSEKIAPLPSLLKGGKEESLYKEFTWFEYKNSSY 311
RSVKHRVLA G +SRLS+IYFGGP S++IAPL +++ G E+SLY+EFTW EYK ++Y
Sbjct: 268 RSVKHRVLAPEGEESRLSVIYFGGPAASQRIAPLEQVMREG-EQSLYREFTWGEYKKAAY 326
Query: 312 GSRLADNRLGHFE 324
+RL DNRLG +E
Sbjct: 327 KTRLGDNRLGPYE 339
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.319 0.137 0.405
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 571,327,170
Number of Sequences: 2790947
Number of extensions: 24127364
Number of successful extensions: 54557
Number of sequences better than 10.0: 1256
Number of HSP's better than 10.0 without gapping: 1110
Number of HSP's successfully gapped in prelim test: 146
Number of HSP's that attempted gapping in prelim test: 51330
Number of HSP's gapped (non-prelim): 1482
length of query: 329
length of database: 848,049,833
effective HSP length: 127
effective length of query: 202
effective length of database: 493,599,564
effective search space: 99707111928
effective search space used: 99707111928
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 75 (33.5 bits)
Lotus: description of TM0190.3


