

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0178.5
(770 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
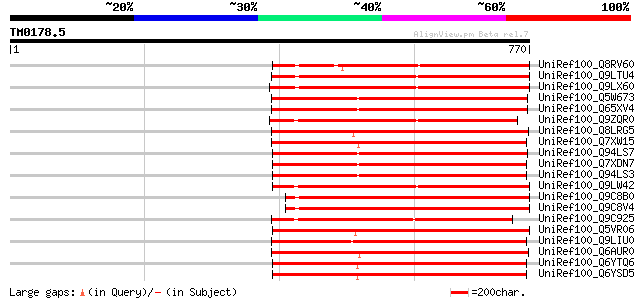
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q8RV60 Hypothetical protein At2g05640 [Arabidopsis tha... 412 e-113
UniRef100_Q9LTU4 Helicase-like protein [Arabidopsis thaliana] 392 e-107
UniRef100_Q9LX60 Hypothetical protein F4M19_60 [Arabidopsis thal... 386 e-105
UniRef100_Q5W673 Putative helicase [Oryza sativa] 385 e-105
UniRef100_Q65XV4 Hypothetical protein P0016H04.14 [Oryza sativa] 382 e-104
UniRef100_Q9ZQR0 Putative helicase [Arabidopsis thaliana] 371 e-101
UniRef100_Q8LRG5 Putative helicase [Oryza sativa] 369 e-100
UniRef100_Q7XW15 OSJNBb0013O03.3 protein [Oryza sativa] 367 e-100
UniRef100_Q94LS7 Putative helicase [Oryza sativa] 367 e-100
UniRef100_Q7XDN7 Hypothetical protein [Oryza sativa] 363 1e-98
UniRef100_Q94LS3 Putative helicase [Oryza sativa] 363 1e-98
UniRef100_Q9LW42 Helicase-like protein [Arabidopsis thaliana] 360 7e-98
UniRef100_Q9C8B0 Hypothetical protein F10O5.11 [Arabidopsis thal... 358 3e-97
UniRef100_Q9C8V4 Hypothetical protein T22A15.15 [Arabidopsis tha... 358 3e-97
UniRef100_Q9C925 Hypothetical protein F14G24.23 [Arabidopsis tha... 355 2e-96
UniRef100_Q5VR06 Helicase-like protein [Oryza sativa] 353 1e-95
UniRef100_Q9LIU0 Similar to Arabidopsis thaliana chromosome II B... 347 6e-94
UniRef100_Q6AUR0 Hypothetical protein OSJNBa0077L08.8 [Oryza sat... 341 6e-92
UniRef100_Q6YTQ6 Helicase-like protein [Oryza sativa] 337 8e-91
UniRef100_Q6YSD5 Helicase-like protein [Oryza sativa] 337 8e-91
>UniRef100_Q8RV60 Hypothetical protein At2g05640 [Arabidopsis thaliana]
Length = 1308
Score = 412 bits (1058), Expect = e-113
Identities = 205/384 (53%), Positives = 271/384 (70%), Gaps = 13/384 (3%)
Query: 390 KASLIIWDETPMLNKHCFEALDRSLNDIIKTQSTHGYDIPFGGIVVVLGGDFRQILPVIS 449
+A LIIWDE PML++HC+EALDR+L DI++ + PFGG + GGDFRQILPVI+
Sbjct: 927 RAKLIIWDEAPMLHRHCYEALDRTLKDIVQADNHK----PFGGKTITFGGDFRQILPVIT 982
Query: 450 KGSRSEIVGSAINSSYLWKHCKVMKLTVNMILQNATSTSSPAE---IKEFADWLLQVGDG 506
KGSR +I+ +++ SS LW CKV+ LT NM L T+ P E IKEF+DW+L++GDG
Sbjct: 983 KGSREQIIHASLTSSRLWNSCKVLTLTKNMRL-----TADPTEKDNIKEFSDWILKLGDG 1037
Query: 507 TVKTIDEEETLIEIPPNLLIEQCKEPLLELVNFAYPKLAHNLQKNSFFQERAILAPTLES 566
+ ++ E I+IP ++L+ P+ + N YP L NL +FF+ERAIL PT +
Sbjct: 1038 KLSEPNDGEAAIDIPEDMLLLDSLHPIDSIANCVYPNLLQNLNDQTFFRERAILCPTNDD 1097
Query: 567 VEEINNFMLAMIPGDETEYLSYDTLCKSDEDSGVNAEWFTSEFLNDFKCSEIPNHAIKLK 626
V E+NN ++ ++PG+ EY S D +C D +A T EFLN KCS +PNH ++LK
Sbjct: 1098 VSEVNNHIMDLLPGEVKEYFSSDKICDFDTSVERDANMST-EFLNAIKCSGVPNHVLRLK 1156
Query: 627 VGVPIMLIRNIDQAAGLCNDTRMIVNALTKYIIVATILNGNKMGETTFIPRMSLTPSNSD 686
+GVP+MLIRN+DQ GLCN TR+ V L +I A +L G+ G ++PR+ LTP++
Sbjct: 1157 LGVPVMLIRNLDQKYGLCNGTRLQVTQLGDRVIEAKVLTGSNAGNKVYLPRLVLTPADFR 1216
Query: 687 IPFKFQRRQFPVTLCFAMTINKSQGQSLSHVGLYLPRPVFTHGQLYVALSRVKSRKGLKM 746
IPF+FQRRQFPV CF MTINKSQGQSLSHVG+YLPRPVF+HGQLYVA+SRVKSR+GLK+
Sbjct: 1217 IPFRFQRRQFPVVPCFGMTINKSQGQSLSHVGIYLPRPVFSHGQLYVAVSRVKSRRGLKI 1276
Query: 747 LIIDDEGVVSNTTRNVMYQEVFDN 770
LIID+EG TT NV+++EVF N
Sbjct: 1277 LIIDEEGNRGKTTTNVVFKEVFQN 1300
>UniRef100_Q9LTU4 Helicase-like protein [Arabidopsis thaliana]
Length = 1428
Score = 392 bits (1007), Expect = e-107
Identities = 198/385 (51%), Positives = 268/385 (69%), Gaps = 8/385 (2%)
Query: 389 KKASLIIWDETPMLNKHCFEALDRSLNDIIKTQSTHGYDIPFGGIVVVLGGDFRQILPVI 448
K+ASLIIWDE PM++K CFEALD+S +DIIK FGG V+V GGDFRQ+LPVI
Sbjct: 1048 KEASLIIWDEAPMMSKFCFEALDKSFSDIIKRVDNK----VFGGKVMVFGGDFRQVLPVI 1103
Query: 449 SKGSRSEIVGSAINSSYLWKHCKVMKLTVNM-ILQNATSTSSPAEIKEFADWLLQVGDGT 507
+ R+EIV S++N+SYLW HCKV++LT NM +L N S EI+EF+DWLL VGDG
Sbjct: 1104 NGAGRAEIVMSSLNASYLWDHCKVLRLTKNMRLLNNDLSVDEAKEIQEFSDWLLAVGDGR 1163
Query: 508 VKTIDEEETLIEIPPNLLIEQCKEPLLELVNFAY--PKLAHNLQKNSFFQERAILAPTLE 565
V ++ E +I+IP LLI++ P+ + Y P H + FFQ RAILAP E
Sbjct: 1164 VNEPNDGEVIIDIPEELLIQEADNPIEAISREIYGDPTKLHEISDPKFFQRRAILAPKNE 1223
Query: 566 SVEEINNFMLAMIPGDETEYLSYDTLCKSDEDSGVNAEWFTSEFLNDFKCSEIPNHAIKL 625
V IN +ML + +E YLS D++ SD DS N T +FLN K S +P+H+++L
Sbjct: 1224 DVNTINQYMLEHLDSEERIYLSADSIDPSDSDSLKNPV-ITPDFLNSIKVSGMPHHSLRL 1282
Query: 626 KVGVPIMLIRNIDQAAGLCNDTRMIVNALTKYIIVATILNGNKMGETTFIPRMSLTPSNS 685
KVG P+ML+RN+D GLCN TR+ + L +I+ A ++ G+++G+ +IP +++TPS++
Sbjct: 1283 KVGAPVMLLRNLDPKGGLCNGTRLQITQLCSHIVEAKVITGDRIGQIVYIPLINITPSDT 1342
Query: 686 DIPFKFQRRQFPVTLCFAMTINKSQGQSLSHVGLYLPRPVFTHGQLYVALSRVKSRKGLK 745
+PFK +RRQFP+++ F MTINKSQGQSL VGLYLP+PVF+HGQLYVALSRV S+ GLK
Sbjct: 1343 KLPFKMRRRQFPLSVAFVMTINKSQGQSLEQVGLYLPKPVFSHGQLYVALSRVTSKTGLK 1402
Query: 746 MLIIDDEGVVSNTTRNVMYQEVFDN 770
+LI+D EG + T NV+++EVF N
Sbjct: 1403 ILILDKEGKIQKQTTNVVFKEVFQN 1427
>UniRef100_Q9LX60 Hypothetical protein F4M19_60 [Arabidopsis thaliana]
Length = 1752
Score = 386 bits (991), Expect = e-105
Identities = 196/388 (50%), Positives = 268/388 (68%), Gaps = 8/388 (2%)
Query: 386 NY*KKASLIIWDETPMLNKHCFEALDRSLNDIIKTQSTHGYDIPFGGIVVVLGGDFRQIL 445
N K+ASLIIWDE PM++K CFE+LD+S DI+ + FGG VVV GGDFRQ+L
Sbjct: 1368 NLIKEASLIIWDEAPMMSKFCFESLDKSFYDILNNKDNK----VFGGKVVVFGGDFRQVL 1423
Query: 446 PVISKGSRSEIVGSAINSSYLWKHCKVMKLTVNM-ILQNATSTSSPAEIKEFADWLLQVG 504
PVI+ R EIV S++N+SYLW HCKV+KLT NM +L S+ EI++F+DWLL VG
Sbjct: 1424 PVINGAGRVEIVMSSLNASYLWDHCKVLKLTKNMRLLSGGLSSEEAKEIQQFSDWLLAVG 1483
Query: 505 DGTVKTIDEEETLIEIPPNLLIEQCKEPLLELVNFAY--PKLAHNLQKNSFFQERAILAP 562
DG + ++ E LI+IP LLI++ P+ + Y P H + FFQ RAILAP
Sbjct: 1484 DGRINEPNDGEALIDIPEELLIKEAGNPIEAISKEIYGDPSELHMINDPKFFQRRAILAP 1543
Query: 563 TLESVEEINNFMLAMIPGDETEYLSYDTLCKSDEDSGVNAEWFTSEFLNDFKCSEIPNHA 622
T E V IN +ML + +E YLS D++ +D DS N T +FLN + + +P+HA
Sbjct: 1544 TNEDVNTINQYMLEHLKSEERIYLSADSIDPTDSDSLANPV-ITPDFLNSIQLTGMPHHA 1602
Query: 623 IKLKVGVPIMLIRNIDQAAGLCNDTRMIVNALTKYIIVATILNGNKMGETTFIPRMSLTP 682
++LKVG P+ML+RN+D GLCN TR+ + L K ++ A ++ +++G+ IP ++LTP
Sbjct: 1603 LRLKVGAPVMLLRNLDPKGGLCNGTRLQITQLAKQVVQAKVITRDRIGDIVLIPLINLTP 1662
Query: 683 SNSDIPFKFQRRQFPVTLCFAMTINKSQGQSLSHVGLYLPRPVFTHGQLYVALSRVKSRK 742
S++ +PFK +RRQFP+++ FAMTINKSQGQSL VGLYLP+PVF+HGQLYVALSRV S+K
Sbjct: 1663 SDTKLPFKMRRRQFPLSVAFAMTINKSQGQSLEQVGLYLPKPVFSHGQLYVALSRVTSKK 1722
Query: 743 GLKMLIIDDEGVVSNTTRNVMYQEVFDN 770
GLK+LI+D +G + T NV+++EVF N
Sbjct: 1723 GLKILILDKDGNMQKQTTNVVFKEVFQN 1750
>UniRef100_Q5W673 Putative helicase [Oryza sativa]
Length = 1634
Score = 385 bits (988), Expect = e-105
Identities = 190/381 (49%), Positives = 261/381 (67%), Gaps = 2/381 (0%)
Query: 389 KKASLIIWDETPMLNKHCFEALDRSLNDIIKTQSTHGYDIPFGGIVVVLGGDFRQILPVI 448
KK SLI+WDE PM N+ CFEALDRSL DI++++ PFGG+ VVLGGDFRQILPV+
Sbjct: 1255 KKTSLILWDEAPMANRICFEALDRSLRDILRSKGEDNSTKPFGGMTVVLGGDFRQILPVV 1314
Query: 449 SKGSRSEIVGSAINSSYLWKHCKVMKLTVNMILQNATSTSSPAE-IKEFADWLLQVGDGT 507
KG R++IV ++I SYLW+H + KLT NM L + + +FA W+L +GDG
Sbjct: 1315 RKGRRTQIVNASIKRSYLWQHFHIFKLTRNMRLSCISRDEDEQKRTADFAQWILNIGDGK 1374
Query: 508 VKTIDEEETLIEIPPNLLIEQCKEPLLELVNFAYPKLAHNLQKNSFFQERAILAPTLESV 567
+ D EE IEIP +L++++ +P E+V YP L N +K F ++RAIL P E+
Sbjct: 1375 TTSADGEEW-IEIPDDLILKKGGDPKEEIVKSIYPNLVQNYKKRDFLEQRAILCPRNETA 1433
Query: 568 EEINNFMLAMIPGDETEYLSYDTLCKSDEDSGVNAEWFTSEFLNDFKCSEIPNHAIKLKV 627
EIN F++ MI G+E YLS DT+CK+ + + +EFLN +PNH +KLKV
Sbjct: 1434 REINEFIMNMIEGEEITYLSCDTVCKATTNDSETDVLYPTEFLNSLNFPGMPNHVLKLKV 1493
Query: 628 GVPIMLIRNIDQAAGLCNDTRMIVNALTKYIIVATILNGNKMGETTFIPRMSLTPSNSDI 687
G+P+ML+RNI+Q++GLCN TRM + L K I A I+ G +GE +IPR+ +TP+ S
Sbjct: 1494 GLPVMLLRNINQSSGLCNGTRMTITQLGKRFIEAQIITGTHVGEKVYIPRIIMTPTESGW 1553
Query: 688 PFKFQRRQFPVTLCFAMTINKSQGQSLSHVGLYLPRPVFTHGQLYVALSRVKSRKGLKML 747
PF +RRQ+P+++CFAMTINKSQGQSL+ VGLYLP+ VFTHGQLYVA SRV R GL+++
Sbjct: 1554 PFLLKRRQYPLSVCFAMTINKSQGQSLNMVGLYLPKQVFTHGQLYVAFSRVTRRDGLRIM 1613
Query: 748 IIDDEGVVSNTTRNVMYQEVF 768
+ D+E + RN++Y+E+F
Sbjct: 1614 LDDNESPGEHMVRNIVYKEIF 1634
>UniRef100_Q65XV4 Hypothetical protein P0016H04.14 [Oryza sativa]
Length = 1525
Score = 382 bits (981), Expect = e-104
Identities = 188/381 (49%), Positives = 261/381 (68%), Gaps = 2/381 (0%)
Query: 389 KKASLIIWDETPMLNKHCFEALDRSLNDIIKTQSTHGYDIPFGGIVVVLGGDFRQILPVI 448
KK SLI+WDE PM N+ CFEALDRSL DI++++ PFGG+ VVLGGDFRQILPV+
Sbjct: 1146 KKTSLILWDEAPMANRICFEALDRSLRDILRSKGEDNSTKPFGGMTVVLGGDFRQILPVV 1205
Query: 449 SKGSRSEIVGSAINSSYLWKHCKVMKLTVNMILQNATSTSSPAE-IKEFADWLLQVGDGT 507
KG R++IV ++I SYLW+H + KLT NM L + + +FA W+L +GDG
Sbjct: 1206 RKGRRTQIVNASIKRSYLWQHFHIFKLTRNMRLSCISRDEDEQKRTADFAQWILNIGDGK 1265
Query: 508 VKTIDEEETLIEIPPNLLIEQCKEPLLELVNFAYPKLAHNLQKNSFFQERAILAPTLESV 567
+ D EE IEIP +L++++ +P E+V YP L N +K F ++RAIL P E+
Sbjct: 1266 TTSADGEEW-IEIPDDLILKKGGDPKEEIVKSIYPNLVQNYKKRDFLEQRAILCPRNETA 1324
Query: 568 EEINNFMLAMIPGDETEYLSYDTLCKSDEDSGVNAEWFTSEFLNDFKCSEIPNHAIKLKV 627
+IN F++ MI G+E YLS DT+CK+ + + +EFLN +PNH +KLK+
Sbjct: 1325 RKINEFIMNMIEGEEITYLSCDTVCKATTNDSETDVLYPTEFLNSLNFPGMPNHVLKLKL 1384
Query: 628 GVPIMLIRNIDQAAGLCNDTRMIVNALTKYIIVATILNGNKMGETTFIPRMSLTPSNSDI 687
G+P+ML+RNI+Q++GLCN TRM + L K I A I+ G +GE +IPR+ +TP+ S
Sbjct: 1385 GLPVMLLRNINQSSGLCNGTRMTITQLGKRFIEAQIITGTHVGEKVYIPRIIMTPTESGW 1444
Query: 688 PFKFQRRQFPVTLCFAMTINKSQGQSLSHVGLYLPRPVFTHGQLYVALSRVKSRKGLKML 747
PF +RRQ+P+++CFAMTINKSQGQSL+ VGLYLP+ VFTHGQLYVA SRV R GL+++
Sbjct: 1445 PFLLKRRQYPLSVCFAMTINKSQGQSLNMVGLYLPKQVFTHGQLYVAFSRVTRRDGLRIM 1504
Query: 748 IIDDEGVVSNTTRNVMYQEVF 768
+ D+E + RN++Y+E+F
Sbjct: 1505 LDDNESPGEHMVRNIVYKEIF 1525
>UniRef100_Q9ZQR0 Putative helicase [Arabidopsis thaliana]
Length = 1265
Score = 371 bits (952), Expect = e-101
Identities = 187/371 (50%), Positives = 257/371 (68%), Gaps = 8/371 (2%)
Query: 386 NY*KKASLIIWDETPMLNKHCFEALDRSLNDIIKTQSTHGYDIPFGGIVVVLGGDFRQIL 445
N KKASL+IWDE PM+++ CFEALD+S +DIIK + FGG VVV GGDFRQ+
Sbjct: 898 NLVKKASLVIWDEAPMMSRFCFEALDKSFSDIIKNTD----NTVFGGKVVVFGGDFRQVF 953
Query: 446 PVISKGSRSEIVGSAINSSYLWKHCKVMKLTVNM-ILQNATSTSSPAEIKEFADWLLQVG 504
PVI+ R+EIV S++N+SYLW +CKV+KLT N +L N S + EI+EF+DWLL VG
Sbjct: 954 PVINGAGRAEIVMSSLNASYLWDNCKVLKLTKNTRLLANNLSETEAKEIQEFSDWLLAVG 1013
Query: 505 DGTVKTIDEEETLIEIPPNLLIEQCKEPLLELVNFAY--PKLAHNLQKNSFFQERAILAP 562
DG + ++ +I+IP +LLI +P+ + N Y PK+ H + FFQ RAILA
Sbjct: 1014 DGRINESNDGVAIIDIPEDLLITNADKPIESITNEIYGDPKILHEITDPKFFQGRAILAS 1073
Query: 563 TLESVEEINNFMLAMIPGDETEYLSYDTLCKSDEDSGVNAEWFTSEFLNDFKCSEIPNHA 622
E V IN ++L + +E YLS D++ +D DS N T +FLN K +PNH+
Sbjct: 1074 KNEDVNTINEYLLDQLHAEERIYLSADSIDPTDSDSLSNPV-ITPDFLNSIKLPGLPNHS 1132
Query: 623 IKLKVGVPIMLIRNIDQAAGLCNDTRMIVNALTKYIIVATILNGNKMGETTFIPRMSLTP 682
++LKVG P++L+RN+D GLCN TR+ + L I+ A ++ G+++G IP ++LTP
Sbjct: 1133 LRLKVGAPVLLLRNLDPKGGLCNGTRLQITQLCTQIVEAKVITGDRIGHIILIPTVNLTP 1192
Query: 683 SNSDIPFKFQRRQFPVTLCFAMTINKSQGQSLSHVGLYLPRPVFTHGQLYVALSRVKSRK 742
+N+ +PFK +RRQFP+++ F MTINKS+GQSL HVGLYLP+PVF+HGQLYVALSRV S+K
Sbjct: 1193 TNTKLPFKMRRRQFPLSVAFVMTINKSEGQSLEHVGLYLPKPVFSHGQLYVALSRVTSKK 1252
Query: 743 GLKMLIIDDEG 753
GLK+LI+D +G
Sbjct: 1253 GLKILILDKDG 1263
>UniRef100_Q8LRG5 Putative helicase [Oryza sativa]
Length = 1453
Score = 369 bits (946), Expect = e-100
Identities = 189/386 (48%), Positives = 263/386 (67%), Gaps = 5/386 (1%)
Query: 389 KKASLIIWDETPMLNKHCFEALDRSLNDIIKTQSTHGYDIPFGGIVVVLGGDFRQILPVI 448
+K SLIIWDE PM ++ CFEALDR+L D++ + +PFGG VVLGGDFRQILPVI
Sbjct: 1054 QKTSLIIWDEAPMTHRRCFEALDRTLRDLLSEHNPSNSVLPFGGKFVVLGGDFRQILPVI 1113
Query: 449 SKGSRSEIVGSAINSSYLWKHCKVMKLTVNM-ILQNATSTSSPAEIKEFADWLLQVGDGT 507
KG+R+ IV ++I +S LW+H ++KLTVNM + Q+ S ++++FA W+L +GDG
Sbjct: 1114 KKGTRNSIVDASITNSPLWQHVVLLKLTVNMRLFQSGLSEGRRHDLEQFARWVLALGDGM 1173
Query: 508 V---KTIDEEE-TLIEIPPNLLIEQCKEPLLELVNFAYPKLAHNLQKNSFFQERAILAPT 563
+ K IDE E T I+IP +LLI + + +VN +P H +S+ RAI+ P
Sbjct: 1174 LPVSKRIDESEATWIDIPDDLLIRASDDKIYSIVNEVFPCYVHRYTDSSYLASRAIVCPN 1233
Query: 564 LESVEEINNFMLAMIPGDETEYLSYDTLCKSDEDSGVNAEWFTSEFLNDFKCSEIPNHAI 623
+V+EIN++M+AMIPG+ EYLS DT+ K+ E + +EFLN + P H +
Sbjct: 1234 NSTVDEINDYMVAMIPGEMKEYLSCDTISKTSEHIPDFDILYPTEFLNSINANNFPTHRL 1293
Query: 624 KLKVGVPIMLIRNIDQAAGLCNDTRMIVNALTKYIIVATILNGNKMGETTFIPRMSLTPS 683
LK G +ML+RN++Q+ GLCN TR++V +L ++ IL G+ +GE FIPR+ L+ +
Sbjct: 1294 ALKKGATVMLLRNLNQSLGLCNGTRLLVLSLGHRLLECVILTGSNVGERAFIPRIVLSTT 1353
Query: 684 NSDIPFKFQRRQFPVTLCFAMTINKSQGQSLSHVGLYLPRPVFTHGQLYVALSRVKSRKG 743
+S PF QRRQFPV +C+AMTINKSQGQ+LS VG+YL + VFTHGQLYVA+SR SR G
Sbjct: 1354 SSKWPFVLQRRQFPVRVCYAMTINKSQGQTLSRVGVYLKKAVFTHGQLYVAVSRSTSRDG 1413
Query: 744 LKMLIIDDEGVVSNTTRNVMYQEVFD 769
L++LI DD+G S+ TRNV+Y EV +
Sbjct: 1414 LRILIKDDDGACSSKTRNVVYHEVLE 1439
>UniRef100_Q7XW15 OSJNBb0013O03.3 protein [Oryza sativa]
Length = 2052
Score = 367 bits (943), Expect = e-100
Identities = 183/384 (47%), Positives = 260/384 (67%), Gaps = 5/384 (1%)
Query: 389 KKASLIIWDETPMLNKHCFEALDRSLNDIIKTQSTHGYDIPFGGIVVVLGGDFRQILPVI 448
+K LIIWDE PM ++ CFEALDR+L D++ +PFGG V+VLGGDFRQILPVI
Sbjct: 1031 QKTLLIIWDEAPMTHRLCFEALDRTLRDLLSEHDPANAIVPFGGKVIVLGGDFRQILPVI 1090
Query: 449 SKGSRSEIVGSAINSSYLWKHCKVMKLTVNM-ILQNATSTSSPAEIKEFADWLLQVGDGT 507
KGSR+ I ++I +S LW+H K++ L +NM +L++ + + E+ FA W+L +G+G
Sbjct: 1091 QKGSRASIDDASITNSPLWRHVKLLSLKINMRLLRSGLTQTKKDELDNFAKWVLHIGNGD 1150
Query: 508 VKTIDEEE----TLIEIPPNLLIEQCKEPLLELVNFAYPKLAHNLQKNSFFQERAILAPT 563
V E T +EIP +LLI+ + + L++ +P L HN ++ RAI+ P
Sbjct: 1151 VPATQRERETEPTWVEIPQDLLIKTDGDKIPALIDEVFPDLLHNHTDPTYLSCRAIVCPN 1210
Query: 564 LESVEEINNFMLAMIPGDETEYLSYDTLCKSDEDSGVNAEWFTSEFLNDFKCSEIPNHAI 623
+V++INN+++ ++PG+E EYLS DT+ KS E + +EF N + PNH +
Sbjct: 1211 NGTVDDINNYVVGLLPGEEKEYLSCDTIAKSSEHIPDLDLLYPTEFPNSINVNNFPNHRL 1270
Query: 624 KLKVGVPIMLIRNIDQAAGLCNDTRMIVNALTKYIIVATILNGNKMGETTFIPRMSLTPS 683
LK GV IML+RN++Q+ GLCN TR+++N L ++++ TIL G+K+GE F+PR+SL +
Sbjct: 1271 VLKKGVIIMLLRNLNQSMGLCNGTRLLINVLGEWVLQRTILTGSKIGEIVFVPRISLNTT 1330
Query: 684 NSDIPFKFQRRQFPVTLCFAMTINKSQGQSLSHVGLYLPRPVFTHGQLYVALSRVKSRKG 743
NS PF QRRQFPV +C+AMTINKSQGQ+LSHVG+YL +PVFTHGQLYV +SR SR G
Sbjct: 1331 NSKWPFTLQRRQFPVRVCYAMTINKSQGQTLSHVGVYLKKPVFTHGQLYVVISRATSRSG 1390
Query: 744 LKMLIIDDEGVVSNTTRNVMYQEV 767
LK+LI DD ++ T NV+Y E+
Sbjct: 1391 LKILIEDDNESCASETSNVVYHEI 1414
>UniRef100_Q94LS7 Putative helicase [Oryza sativa]
Length = 1573
Score = 367 bits (941), Expect = e-100
Identities = 180/380 (47%), Positives = 260/380 (68%), Gaps = 2/380 (0%)
Query: 390 KASLIIWDETPMLNKHCFEALDRSLNDIIKTQSTHGYDIPFGGIVVVLGGDFRQILPVIS 449
K SLI+WDE PM N++CFEALD+SL D+ + ++ + Y PFGG+ VVLGGDFRQILP++
Sbjct: 1194 KTSLILWDEAPMANRNCFEALDKSLRDVQRFRNENSYQKPFGGMTVVLGGDFRQILPIVP 1253
Query: 450 KGSRSEIVGSAINSSYLWKHCKVMKLTVNMILQNATSTSSPAE-IKEFADWLLQVGDGTV 508
KG R V ++I SYLW+H +V LT NM L + + + + EFA+W+L++G+G
Sbjct: 1254 KGRREHTVNASIKFSYLWQHFEVFNLTKNMRLNSVSKDQAEHQKTAEFAEWILRIGNGDT 1313
Query: 509 KTIDEEETLIEIPPNLLIEQCKEPLLELVNFAYPKLAHNLQKNSFFQERAILAPTLESVE 568
+DE+ + +P +LL+++ +P ++V+ YP L +N K + +ERAIL PT + V
Sbjct: 1314 ILLDEKGW-VSMPSDLLLQKGDDPKAQIVDSTYPGLQYNCCKPKYLEERAILCPTNDDVN 1372
Query: 569 EINNFMLAMIPGDETEYLSYDTLCKSDEDSGVNAEWFTSEFLNDFKCSEIPNHAIKLKVG 628
E+N +++ I GD+ YLS+D++ KS S + +EFLN K IPNH +KLKVG
Sbjct: 1373 ELNEYIMDQIQGDKVTYLSHDSVSKSMSYSHEMEMLYPTEFLNSLKHPGIPNHQLKLKVG 1432
Query: 629 VPIMLIRNIDQAAGLCNDTRMIVNALTKYIIVATILNGNKMGETTFIPRMSLTPSNSDIP 688
+P+ML+RNI+Q AGLCN TRM + K +I A I+ G +G+ IP++ ++P+ P
Sbjct: 1433 LPVMLLRNINQNAGLCNGTRMRITRFGKRVIEAEIITGTHIGDMVCIPQIIMSPNERKWP 1492
Query: 689 FKFQRRQFPVTLCFAMTINKSQGQSLSHVGLYLPRPVFTHGQLYVALSRVKSRKGLKMLI 748
F R+QFP+++CFAMTINKSQGQ+L+ VGLYLPR VFTHGQLYVA+SRV SR GLK++I
Sbjct: 1493 FVLNRKQFPLSVCFAMTINKSQGQTLNKVGLYLPRQVFTHGQLYVAVSRVTSRDGLKIMI 1552
Query: 749 IDDEGVVSNTTRNVMYQEVF 768
D E +N++Y+E+F
Sbjct: 1553 ADKECPGEGMVKNIVYKEIF 1572
>UniRef100_Q7XDN7 Hypothetical protein [Oryza sativa]
Length = 1477
Score = 363 bits (931), Expect = 1e-98
Identities = 179/379 (47%), Positives = 256/379 (67%), Gaps = 2/379 (0%)
Query: 390 KASLIIWDETPMLNKHCFEALDRSLNDIIKTQSTHGYDIPFGGIVVVLGGDFRQILPVIS 449
K SLI+WDE PM N++CFEALD+SL D+ + ++ + PFGG+ VVLGGDFRQILP++
Sbjct: 1098 KTSLILWDEAPMANRNCFEALDKSLRDVQRCRNENSCQKPFGGMTVVLGGDFRQILPIVP 1157
Query: 450 KGSRSEIVGSAINSSYLWKHCKVMKLTVNMILQNATSTSSPAEIK-EFADWLLQVGDGTV 508
KG R V + I SYLW+H +V LT NM L + + + EFA+W+LQ+G+G
Sbjct: 1158 KGRREHTVNATIKCSYLWQHFEVFNLTKNMRLNYVSKDQTEHQKSAEFAEWILQIGNGDT 1217
Query: 509 KTIDEEETLIEIPPNLLIEQCKEPLLELVNFAYPKLAHNLQKNSFFQERAILAPTLESVE 568
++DE+ + +P +LL+++ +P +++ YP L N K ++ +ERAIL P E+V
Sbjct: 1218 ISLDEKGW-VRMPSDLLLQKGDDPKAQIIESTYPDLQDNCCKQNYLEERAILCPVNENVN 1276
Query: 569 EINNFMLAMIPGDETEYLSYDTLCKSDEDSGVNAEWFTSEFLNDFKCSEIPNHAIKLKVG 628
E+N +++ I GD+ YLS D++ KS S + +EFLN S IPNH +KLKVG
Sbjct: 1277 ELNEYIMDQIQGDKVTYLSRDSVSKSVSYSHEMEMLYPTEFLNSLNHSGIPNHQLKLKVG 1336
Query: 629 VPIMLIRNIDQAAGLCNDTRMIVNALTKYIIVATILNGNKMGETTFIPRMSLTPSNSDIP 688
+P+ML+RNI+Q+AGLCN TRM + L +I A I+ G G+ IP++ ++P+ P
Sbjct: 1337 LPVMLLRNINQSAGLCNGTRMTITRLGNKVIEAQIITGTHSGDMVCIPQIIMSPTEPKWP 1396
Query: 689 FKFQRRQFPVTLCFAMTINKSQGQSLSHVGLYLPRPVFTHGQLYVALSRVKSRKGLKMLI 748
F R+QFP+++CFAMTINKSQGQ+L+ VGLYLPR VFTHGQLYVA+SRV SR GLK+LI
Sbjct: 1397 FMLNRKQFPLSVCFAMTINKSQGQTLNKVGLYLPRQVFTHGQLYVAVSRVTSRDGLKILI 1456
Query: 749 IDDEGVVSNTTRNVMYQEV 767
D+E +N++Y+E+
Sbjct: 1457 ADEECPGEGMVKNIVYKEI 1475
>UniRef100_Q94LS3 Putative helicase [Oryza sativa]
Length = 1501
Score = 363 bits (931), Expect = 1e-98
Identities = 179/379 (47%), Positives = 256/379 (67%), Gaps = 2/379 (0%)
Query: 390 KASLIIWDETPMLNKHCFEALDRSLNDIIKTQSTHGYDIPFGGIVVVLGGDFRQILPVIS 449
K SLI+WDE PM N++CFEALD+SL D+ + ++ + PFGG+ VVLGGDFRQILP++
Sbjct: 1122 KTSLILWDEAPMANRNCFEALDKSLRDVQRCRNENSCQKPFGGMTVVLGGDFRQILPIVP 1181
Query: 450 KGSRSEIVGSAINSSYLWKHCKVMKLTVNMILQNATSTSSPAEIK-EFADWLLQVGDGTV 508
KG R V + I SYLW+H +V LT NM L + + + EFA+W+LQ+G+G
Sbjct: 1182 KGRREHTVNATIKCSYLWQHFEVFNLTKNMRLNYVSKDQTEHQKSAEFAEWILQIGNGDT 1241
Query: 509 KTIDEEETLIEIPPNLLIEQCKEPLLELVNFAYPKLAHNLQKNSFFQERAILAPTLESVE 568
++DE+ + +P +LL+++ +P +++ YP L N K ++ +ERAIL P E+V
Sbjct: 1242 ISLDEKGW-VRMPSDLLLQKGDDPKAQIIESTYPDLQDNCCKQNYLEERAILCPVNENVN 1300
Query: 569 EINNFMLAMIPGDETEYLSYDTLCKSDEDSGVNAEWFTSEFLNDFKCSEIPNHAIKLKVG 628
E+N +++ I GD+ YLS D++ KS S + +EFLN S IPNH +KLKVG
Sbjct: 1301 ELNEYIMDQIQGDKVTYLSRDSVSKSVSYSHEMEMLYPTEFLNSLNHSGIPNHQLKLKVG 1360
Query: 629 VPIMLIRNIDQAAGLCNDTRMIVNALTKYIIVATILNGNKMGETTFIPRMSLTPSNSDIP 688
+P+ML+RNI+Q+AGLCN TRM + L +I A I+ G G+ IP++ ++P+ P
Sbjct: 1361 LPVMLLRNINQSAGLCNGTRMTITRLGNKVIEAQIITGTHSGDMVCIPQIIMSPTEPKWP 1420
Query: 689 FKFQRRQFPVTLCFAMTINKSQGQSLSHVGLYLPRPVFTHGQLYVALSRVKSRKGLKMLI 748
F R+QFP+++CFAMTINKSQGQ+L+ VGLYLPR VFTHGQLYVA+SRV SR GLK+LI
Sbjct: 1421 FMLNRKQFPLSVCFAMTINKSQGQTLNKVGLYLPRQVFTHGQLYVAVSRVTSRDGLKILI 1480
Query: 749 IDDEGVVSNTTRNVMYQEV 767
D+E +N++Y+E+
Sbjct: 1481 ADEECPGEGMVKNIVYKEI 1499
>UniRef100_Q9LW42 Helicase-like protein [Arabidopsis thaliana]
Length = 1669
Score = 360 bits (925), Expect = 7e-98
Identities = 183/380 (48%), Positives = 256/380 (67%), Gaps = 5/380 (1%)
Query: 391 ASLIIWDETPMLNKHCFEALDRSLNDIIKTQSTHGYDIPFGGIVVVLGGDFRQILPVISK 450
A LIIWDE PM++K+CFE+LD+SL DI+ T D+PFGG +++ GGDFRQILPVI
Sbjct: 1294 AKLIIWDEAPMMSKYCFESLDKSLKDILSTPE----DMPFGGKLIIFGGDFRQILPVILA 1349
Query: 451 GSRSEIVGSAINSSYLWKHCKVMKLTVNMILQNATSTSSPAEIKEFADWLLQVGDGTVKT 510
R IV S++NSS+LW++CKV KLT NM L + EI++F+ W+L VG+G +
Sbjct: 1350 AGRELIVKSSLNSSHLWQYCKVFKLTKNMRLLQDIDINEAREIEDFSKWILAVGEGKLNQ 1409
Query: 511 IDEEETLIEIPPNLLIEQCKEPLLELVNFAYPKLAHNLQKNSFFQERAILAPTLESVEEI 570
++ T I+I ++LI + P+ ++ Y + FFQ+RAIL PT + V I
Sbjct: 1410 PNDGVTQIQIRDDILIPEGDNPIESIIKAVYGTSFDEERDPKFFQDRAILCPTNDDVNSI 1469
Query: 571 NNFMLAMIPGDETEYLSYDTLCKSDEDSGVNAEWFTSEFLNDFKCSEIPNHAIKLKVGVP 630
N+ ML+ + G+E Y S D++ SD + N +T +FLN K S +PNH + LKVG P
Sbjct: 1470 NDHMLSKLTGEEKIYRSSDSIDPSDTRADKNPV-YTPDFLNKIKISGLPNHLLWLKVGCP 1528
Query: 631 IMLIRNIDQAAGLCNDTRMIVNALTKYIIVATILNGNKMGETTFIPRMSLTPSNSDIPFK 690
+ML+RN+D GL N TR+ + L ++ IL G ++G+ IPRM LTPS+ +PFK
Sbjct: 1529 VMLLRNLDSHGGLMNGTRLQIVRLGDKLVQGRILTGTRVGKLVIIPRMPLTPSDRRLPFK 1588
Query: 691 FQRRQFPVTLCFAMTINKSQGQSLSHVGLYLPRPVFTHGQLYVALSRVKSRKGLKMLIID 750
+RRQFP+++ FAMTINKSQGQSL +VG+YLP+PVF+HGQLYVA+SRVKS+ GLK+LI D
Sbjct: 1589 MKRRQFPLSVAFAMTINKSQGQSLGNVGIYLPKPVFSHGQLYVAMSRVKSKGGLKVLITD 1648
Query: 751 DEGVVSNTTRNVMYQEVFDN 770
+G N T NV+++E+F N
Sbjct: 1649 SKGKQKNETTNVVFKEIFRN 1668
>UniRef100_Q9C8B0 Hypothetical protein F10O5.11 [Arabidopsis thaliana]
Length = 1678
Score = 358 bits (920), Expect = 3e-97
Identities = 182/365 (49%), Positives = 251/365 (67%), Gaps = 8/365 (2%)
Query: 409 ALDRSLNDIIKTQSTHGYDIPFGGIVVVLGGDFRQILPVISKGSRSEIVGSAINSSYLWK 468
+LD+S +DIIK + FGG VVV GGDFRQ+LPVI+ R+EIV S++N+SYLW
Sbjct: 1317 SLDKSFSDIIKNTNNK----VFGGKVVVFGGDFRQVLPVINGAGRAEIVMSSLNASYLWD 1372
Query: 469 HCKVMKLTVNM-ILQNATSTSSPAEIKEFADWLLQVGDGTVKTIDEEETLIEIPPNLLIE 527
HCKV+KLT NM +L N S + EI+EF+DWLL V DG + ++ I+IP +LLI
Sbjct: 1373 HCKVLKLTKNMRLLANNLSATEAKEIQEFSDWLLAVSDGRINEPNDGVATIDIPEDLLIT 1432
Query: 528 QCKEPLLELVNFAY--PKLAHNLQKNSFFQERAILAPTLESVEEINNFMLAMIPGDETEY 585
+P+ + N Y PK+ H + FFQ RAILAP E V IN ++L + +E Y
Sbjct: 1433 NADKPIETITNEIYGDPKILHEITDPKFFQGRAILAPKNEDVNTINEYLLEQLDAEERIY 1492
Query: 586 LSYDTLCKSDEDSGVNAEWFTSEFLNDFKCSEIPNHAIKLKVGVPIMLIRNIDQAAGLCN 645
LS D++ +D DS +N T +FLN K +PNH++ LKVG P+ML+RN+D GLCN
Sbjct: 1493 LSADSIDPTDSDS-LNNPVITPDFLNSIKLPGLPNHSLCLKVGAPVMLLRNLDPKGGLCN 1551
Query: 646 DTRMIVNALTKYIIVATILNGNKMGETTFIPRMSLTPSNSDIPFKFQRRQFPVTLCFAMT 705
TR+ + L I+ A ++ G+++G IP ++LTP+++ +PFK +RRQFP+++ FAMT
Sbjct: 1552 GTRLQITQLCTQIVEAKVITGDRIGNIVLIPTVNLTPTDTKLPFKMRRRQFPLSVAFAMT 1611
Query: 706 INKSQGQSLSHVGLYLPRPVFTHGQLYVALSRVKSRKGLKMLIIDDEGVVSNTTRNVMYQ 765
INKSQGQSL H+GLYLP+PVF+HGQLYVALSRV S+KGLK+LI+D +G + T NV+++
Sbjct: 1612 INKSQGQSLEHIGLYLPKPVFSHGQLYVALSRVTSKKGLKILILDKDGKLQKQTTNVVFK 1671
Query: 766 EVFDN 770
EVF N
Sbjct: 1672 EVFQN 1676
>UniRef100_Q9C8V4 Hypothetical protein T22A15.15 [Arabidopsis thaliana]
Length = 729
Score = 358 bits (920), Expect = 3e-97
Identities = 182/365 (49%), Positives = 251/365 (67%), Gaps = 8/365 (2%)
Query: 409 ALDRSLNDIIKTQSTHGYDIPFGGIVVVLGGDFRQILPVISKGSRSEIVGSAINSSYLWK 468
+LD+S +DIIK + FGG VVV GGDFRQ+LPVI+ R+EIV S++N+SYLW
Sbjct: 368 SLDKSFSDIIKNTNNK----VFGGKVVVFGGDFRQVLPVINGAGRAEIVMSSLNASYLWD 423
Query: 469 HCKVMKLTVNM-ILQNATSTSSPAEIKEFADWLLQVGDGTVKTIDEEETLIEIPPNLLIE 527
HCKV+KLT NM +L N S + EI+EF+DWLL V DG + ++ I+IP +LLI
Sbjct: 424 HCKVLKLTKNMRLLANNLSATEAKEIQEFSDWLLAVSDGRINEPNDGVATIDIPEDLLIT 483
Query: 528 QCKEPLLELVNFAY--PKLAHNLQKNSFFQERAILAPTLESVEEINNFMLAMIPGDETEY 585
+P+ + N Y PK+ H + FFQ RAILAP E V IN ++L + +E Y
Sbjct: 484 NADKPIETITNEIYGDPKILHEITDPKFFQGRAILAPKNEDVNTINEYLLEQLDAEERIY 543
Query: 586 LSYDTLCKSDEDSGVNAEWFTSEFLNDFKCSEIPNHAIKLKVGVPIMLIRNIDQAAGLCN 645
LS D++ +D DS +N T +FLN K +PNH++ LKVG P+ML+RN+D GLCN
Sbjct: 544 LSADSIDPTDSDS-LNNPVITPDFLNSIKLPGLPNHSLCLKVGAPVMLLRNLDPKGGLCN 602
Query: 646 DTRMIVNALTKYIIVATILNGNKMGETTFIPRMSLTPSNSDIPFKFQRRQFPVTLCFAMT 705
TR+ + L I+ A ++ G+++G IP ++LTP+++ +PFK +RRQFP+++ FAMT
Sbjct: 603 GTRLQITQLCTQIVEAKVITGDRIGNIVLIPTVNLTPTDTKLPFKMRRRQFPLSVAFAMT 662
Query: 706 INKSQGQSLSHVGLYLPRPVFTHGQLYVALSRVKSRKGLKMLIIDDEGVVSNTTRNVMYQ 765
INKSQGQSL H+GLYLP+PVF+HGQLYVALSRV S+KGLK+LI+D +G + T NV+++
Sbjct: 663 INKSQGQSLEHIGLYLPKPVFSHGQLYVALSRVTSKKGLKILILDKDGKLQKQTTNVVFK 722
Query: 766 EVFDN 770
EVF N
Sbjct: 723 EVFQN 727
>UniRef100_Q9C925 Hypothetical protein F14G24.23 [Arabidopsis thaliana]
Length = 996
Score = 355 bits (912), Expect = 2e-96
Identities = 177/357 (49%), Positives = 244/357 (67%), Gaps = 5/357 (1%)
Query: 389 KKASLIIWDETPMLNKHCFEALDRSLNDIIKTQSTHGYDIPFGGIVVVLGGDFRQILPVI 448
K+A+LIIWDETPM++KHCFE+LDR+L DI+ D PFGG +V GGDFRQ+LPVI
Sbjct: 643 KEANLIIWDETPMMSKHCFESLDRTLRDIMNNPG----DKPFGGKGIVFGGDFRQVLPVI 698
Query: 449 SKGSRSEIVGSAINSSYLWKHCKVMKLTVNMILQNATSTSSPAEIKEFADWLLQVGDGTV 508
+ R EIV +A+NSSY+W+HCKV++LT NM L S +I+ F+ W+L VGDG +
Sbjct: 699 NGAGREEIVFAALNSSYIWEHCKVLELTKNMRLLANISEHEKRDIEYFSKWILDVGDGKI 758
Query: 509 KTIDEEETLIEIPPNLLIEQCKEPLLELVNFAYPKLAHNLQKNSFFQERAILAPTLESVE 568
++ LI+IP LI +P+ ++ Y + FFQ RAIL PT E V
Sbjct: 759 SQPNDGIALIDIPEEFLINGDNDPVESIIEAVYGNTFMEEKDPKFFQGRAILCPTNEDVN 818
Query: 569 EINNFMLAMIPGDETEYLSYDTLCKSDEDSGVNAEWFTSEFLNDFKCSEIPNHAIKLKVG 628
IN M++M+ G+E YLS D++ +D S N + ++++FLN + S +PNH ++LKVG
Sbjct: 819 SINEHMMSMLDGEERIYLSSDSIDPADTSSA-NNDAYSADFLNSVRVSGLPNHCLRLKVG 877
Query: 629 VPIMLIRNIDQAAGLCNDTRMIVNALTKYIIVATILNGNKMGETTFIPRMSLTPSNSDIP 688
P+ML+RN+D GLCN TR+ V + +I A + GN++G+ IPRM +TPS++ +P
Sbjct: 878 CPVMLLRNMDPNKGLCNGTRLQVTQMADTVIQARFITGNRVGKIVLIPRMLITPSDTRLP 937
Query: 689 FKFQRRQFPVTLCFAMTINKSQGQSLSHVGLYLPRPVFTHGQLYVALSRVKSRKGLK 745
FK +RRQFP+++ FAMTINKSQGQ+L VGLYLPRPVF+HGQLYVA+SRV S+ G K
Sbjct: 938 FKMRRRQFPLSVAFAMTINKSQGQTLESVGLYLPRPVFSHGQLYVAISRVTSKTGTK 994
>UniRef100_Q5VR06 Helicase-like protein [Oryza sativa]
Length = 1427
Score = 353 bits (905), Expect = 1e-95
Identities = 181/389 (46%), Positives = 255/389 (65%), Gaps = 8/389 (2%)
Query: 390 KASLIIWDETPMLNKHCFEALDRSLNDIIKTQSTHGYDIPFGGIVVVLGGDFRQILPVIS 449
+ SLIIWDE M N+ CFEALDR+L DI+ + + D PFGG VVVLGGD +QILPVI
Sbjct: 1036 ETSLIIWDEALMTNRQCFEALDRTLRDILSEKYINAIDKPFGGKVVVLGGDPKQILPVIE 1095
Query: 450 KGSRSEIVGSAINSSYLWKHCKVMKLTVNMILQNATS-TSSPAEIKEFADWLLQVGDGTV 508
S+ EI+ ++I SYLW + K + L NM LQ S T EI +F +W+L +G+G +
Sbjct: 1096 NASKLEIINASIVKSYLWGYVKKIFLFENMRLQKTKSNTLEYKEINDFNNWILDIGNGKI 1155
Query: 509 KTI-------DEEETLIEIPPNLLIEQCKEPLLELVNFAYPKLAHNLQKNSFFQERAILA 561
T D + T+I +P NLLI + L ELV F YP ++ ++ + RAILA
Sbjct: 1156 NTKQSTAQNEDTDTTIILVPENLLINTGENKLEELVKFTYPDFKNSFFNPNYLKNRAILA 1215
Query: 562 PTLESVEEINNFMLAMIPGDETEYLSYDTLCKSDEDSGVNAEWFTSEFLNDFKCSEIPNH 621
T E V+E+NN++L+++P E EY S DTL + + + + E+LN + P H
Sbjct: 1216 TTNEIVDEVNNYILSLVPNQEKEYYSADTLSQCMDTTNDADILYPVEYLNSLNANNFPTH 1275
Query: 622 AIKLKVGVPIMLIRNIDQAAGLCNDTRMIVNALTKYIIVATILNGNKMGETTFIPRMSLT 681
+KLKVGVPI+L+RN++Q GLCN TR+I+ L II I+ G +GE +IPR++LT
Sbjct: 1276 VLKLKVGVPIILLRNLNQNLGLCNGTRLIITNLGDNIIEGIIITGTHIGEKAYIPRINLT 1335
Query: 682 PSNSDIPFKFQRRQFPVTLCFAMTINKSQGQSLSHVGLYLPRPVFTHGQLYVALSRVKSR 741
+ PF RR FP+ +C++MTINKSQGQ+LS+VGLYL +PVFTHGQLYVA+SRV +
Sbjct: 1336 TRGNQWPFTLCRRHFPIKVCYSMTINKSQGQTLSNVGLYLKKPVFTHGQLYVAISRVSNS 1395
Query: 742 KGLKMLIIDDEGVVSNTTRNVMYQEVFDN 770
KGLK+LI +++G + T+N++Y+E+ D+
Sbjct: 1396 KGLKILIENEDGTCATQTKNIVYREILDS 1424
>UniRef100_Q9LIU0 Similar to Arabidopsis thaliana chromosome II BAC T13P21 genomic
sequence [Oryza sativa]
Length = 1278
Score = 347 bits (891), Expect = 6e-94
Identities = 179/380 (47%), Positives = 251/380 (65%), Gaps = 3/380 (0%)
Query: 389 KKASLIIWDETPMLNKHCFEALDRSLNDIIKTQSTHGYDIPFGGIVVVLGGDFRQILPVI 448
+K SLIIWDE PM ++ CFEALDR+L D++ + +PFGG VVVLGGDFRQILPV+
Sbjct: 862 QKTSLIIWDEAPMTHRRCFEALDRTLRDLLSEHAPSNGLVPFGGKVVVLGGDFRQILPVV 921
Query: 449 SKGSRSEIVGSAINSSYLWKHCKVMKLTVNM-ILQNATSTSSPAEIKEFADWLLQVGDGT 507
KGSR+ I+ ++I +S LW H ++KLTVNM +LQ E+++FA W+L +GDG
Sbjct: 922 RKGSRASIIDASITNSPLWSHAVLLKLTVNMRLLQCNLGEQQQQELEKFAQWVLALGDG- 980
Query: 508 VKTIDEEETLIEIPPNLLIEQCKEPLLELVNFAYPKLAHNLQKNSFFQERAILAPTLESV 567
K + E T I+IP +LLI+ + + +VN +P A S+ AI+ P +V
Sbjct: 981 -KNDESEATWIDIPDDLLIKTTGDKIHSIVNEVFPGFASKYTDPSYLASCAIVCPNNSTV 1039
Query: 568 EEINNFMLAMIPGDETEYLSYDTLCKSDEDSGVNAEWFTSEFLNDFKCSEIPNHAIKLKV 627
++IN+ M+ M+PG+ EYLS DT+ KS E + +EFLN + P H + LK
Sbjct: 1040 DDINDRMVDMVPGEVKEYLSCDTISKSSEHIPDFDLLYPTEFLNSISANNFPAHKLALKK 1099
Query: 628 GVPIMLIRNIDQAAGLCNDTRMIVNALTKYIIVATILNGNKMGETTFIPRMSLTPSNSDI 687
GV +ML+RN++Q+ GLCN TR++ +L + ++ IL G +G+ FIPR++LT ++
Sbjct: 1100 GVTVMLLRNLNQSMGLCNGTRLLALSLGQRLLECQILTGTNIGDRVFIPRIALTTTSPKW 1159
Query: 688 PFKFQRRQFPVTLCFAMTINKSQGQSLSHVGLYLPRPVFTHGQLYVALSRVKSRKGLKML 747
PF QRRQFPV +C+AMTINKSQGQ+L VG+YL + VFTHGQLYVA+SR SR GL++L
Sbjct: 1160 PFTLQRRQFPVRVCYAMTINKSQGQTLKRVGVYLKKAVFTHGQLYVAVSRSTSRDGLRIL 1219
Query: 748 IIDDEGVVSNTTRNVMYQEV 767
D+ S+ TRNV+Y EV
Sbjct: 1220 TEGDDEACSSKTRNVVYHEV 1239
>UniRef100_Q6AUR0 Hypothetical protein OSJNBa0077L08.8 [Oryza sativa]
Length = 807
Score = 341 bits (874), Expect = 6e-92
Identities = 182/386 (47%), Positives = 245/386 (63%), Gaps = 5/386 (1%)
Query: 389 KKASLIIWDETPMLNKHCFEALDRSLNDIIKTQSTHGYDIPFGGIVVVLGGDFRQILPVI 448
+K SLI+WDE P+ +K+CFE+LDR+L DI+ + + D FGGI VVLGGDFRQ LPVI
Sbjct: 415 QKTSLIVWDEAPVNHKYCFESLDRTLRDILSETNPNSLDKQFGGITVVLGGDFRQTLPVI 474
Query: 449 SKGSRSEIVGSAINSSYLWKHCKVMKLTVNMILQNAT-STSSPAEIKEFADWLLQVGDGT 507
++ +I+ S I +SYLW C +++LT NM L +A+ S E++ F++WLL++G+GT
Sbjct: 475 QNATKQQILRSCIVNSYLWNKCILIELTENMRLTSASISAQDREELRNFSNWLLKIGNGT 534
Query: 508 VKTIDEEETL----IEIPPNLLIEQCKEPLLELVNFAYPKLAHNLQKNSFFQERAILAPT 563
+D E L IE+P +LL+ L L++F Y S+ ERAILAPT
Sbjct: 535 EPFVDVPEQLTNMFIEMPQSLLLSPDCRNLDGLISFVYNLGCQPSNLTSYLCERAILAPT 594
Query: 564 LESVEEINNFMLAMIPGDETEYLSYDTLCKSDEDSGVNAEWFTSEFLNDFKCSEIPNHAI 623
E V EINN M+A + E Y S D++ S + + +EFLN + +P H +
Sbjct: 595 NEVVSEINNRMIAQLEASEMSYYSSDSIDDSSTNCTAIEALYPTEFLNTISINGLPEHVL 654
Query: 624 KLKVGVPIMLIRNIDQAAGLCNDTRMIVNALTKYIIVATILNGNKMGETTFIPRMSLTPS 683
LK+GVPIML+RN+D + GLCN TR+IV LT +I I G G +IPR+ T +
Sbjct: 655 HLKIGVPIMLLRNLDPSIGLCNGTRLIVTQLTSRVIEGEINTGKAKGTKAYIPRIVTTLT 714
Query: 684 NSDIPFKFQRRQFPVTLCFAMTINKSQGQSLSHVGLYLPRPVFTHGQLYVALSRVKSRKG 743
S PFK +RRQFP+ L +AMTINKSQGQ+LS VG+YLP PVF+HGQLYVA SRV S G
Sbjct: 715 QSKWPFKLRRRQFPIHLSYAMTINKSQGQTLSRVGVYLPSPVFSHGQLYVAFSRVTSPNG 774
Query: 744 LKMLIIDDEGVVSNTTRNVMYQEVFD 769
LK+LI + N T NV+Y EVF+
Sbjct: 775 LKVLIENSPASYENCTHNVVYSEVFN 800
>UniRef100_Q6YTQ6 Helicase-like protein [Oryza sativa]
Length = 1430
Score = 337 bits (864), Expect = 8e-91
Identities = 171/383 (44%), Positives = 244/383 (63%), Gaps = 5/383 (1%)
Query: 390 KASLIIWDETPMLNKHCFEALDRSLNDIIKTQSTHGYDIPFGGIVVVLGGDFRQILPVIS 449
+ +LIIWDE PM ++ CFEALDR+L DI+ IPFGG +VLGGDF+QILPVI
Sbjct: 1005 ETALIIWDEAPMTHRRCFEALDRTLRDILSETCPSNSIIPFGGKPIVLGGDFKQILPVIP 1064
Query: 450 KGSRSEIVGSAINSSYLWKHCKVMKLTVNMILQNAT-STSSPAEIKEFADWLLQVGDGTV 508
KGSR I+ ++I +S LWKH ++ L +NM L N + E+ +F+ W+L +G+GT+
Sbjct: 1065 KGSRQAIINASITNSELWKHVALLSLNINMRLLNPMLPDNQKKELHDFSQWVLAIGNGTL 1124
Query: 509 KTIDEE----ETLIEIPPNLLIEQCKEPLLELVNFAYPKLAHNLQKNSFFQERAILAPTL 564
+E I IP +LL+ + + +V+ Y + + RAI+ PT
Sbjct: 1125 PMTAKEGENYPAWITIPDDLLVMTSGDKIAAIVHEVYSDFLTCYRDIEYLASRAIVCPTN 1184
Query: 565 ESVEEINNFMLAMIPGDETEYLSYDTLCKSDEDSGVNAEWFTSEFLNDFKCSEIPNHAIK 624
+V+EIN++++ ++PGD YLS DT+ KS E + EFLN + P H +
Sbjct: 1185 TTVDEINDYIIGLVPGDSRVYLSCDTISKSSEQIPDFDLLYPPEFLNSINATNFPTHKLV 1244
Query: 625 LKVGVPIMLIRNIDQAAGLCNDTRMIVNALTKYIIVATILNGNKMGETTFIPRMSLTPSN 684
LK GV +ML+RN++Q+ GLCN TR++V L I+ +L G+ +GET +IPR++L +
Sbjct: 1245 LKEGVVVMLLRNLNQSIGLCNGTRLLVTVLGDRILQCIVLTGSNIGETVYIPRITLGTTK 1304
Query: 685 SDIPFKFQRRQFPVTLCFAMTINKSQGQSLSHVGLYLPRPVFTHGQLYVALSRVKSRKGL 744
PF QRRQFPV +C++MTINKSQGQ+L VG+YL +PVFTHGQLYVA+SRV SR GL
Sbjct: 1305 MKWPFTLQRRQFPVRVCYSMTINKSQGQTLQRVGVYLRKPVFTHGQLYVAISRVTSRSGL 1364
Query: 745 KMLIIDDEGVVSNTTRNVMYQEV 767
K+LI +D+G T+N++Y EV
Sbjct: 1365 KILIENDDGSCGTQTKNIVYSEV 1387
>UniRef100_Q6YSD5 Helicase-like protein [Oryza sativa]
Length = 1516
Score = 337 bits (864), Expect = 8e-91
Identities = 171/383 (44%), Positives = 244/383 (63%), Gaps = 5/383 (1%)
Query: 390 KASLIIWDETPMLNKHCFEALDRSLNDIIKTQSTHGYDIPFGGIVVVLGGDFRQILPVIS 449
+ +LIIWDE PM ++ CFEALDR+L DI+ +PFGG +VLGGDF+QILPVI
Sbjct: 1091 ETALIIWDEAPMTHRRCFEALDRTLRDILSETCPSNSIVPFGGKPIVLGGDFKQILPVIP 1150
Query: 450 KGSRSEIVGSAINSSYLWKHCKVMKLTVNMILQNAT-STSSPAEIKEFADWLLQVGDGTV 508
KGSR I+ ++I +S LWKH ++ L +NM L N + E+ +F+ W+L +G+GT+
Sbjct: 1151 KGSRQAIINASITNSELWKHVALLSLNINMRLLNPMLPDNQKKELHDFSQWVLAIGNGTL 1210
Query: 509 KTIDEE----ETLIEIPPNLLIEQCKEPLLELVNFAYPKLAHNLQKNSFFQERAILAPTL 564
+E I IP +LL+ + + +V+ Y + + RAI+ PT
Sbjct: 1211 PMTAKEGENYPAWITIPDDLLVMTSGDKIAAIVHEVYSDFLTCYRDIEYLASRAIVCPTN 1270
Query: 565 ESVEEINNFMLAMIPGDETEYLSYDTLCKSDEDSGVNAEWFTSEFLNDFKCSEIPNHAIK 624
+V+EIN++++ ++PGD YLS DT+ KS E + EFLN + P H +
Sbjct: 1271 TTVDEINDYIIGLVPGDSRVYLSCDTISKSSEQIPDFDLLYPPEFLNSINATNFPTHKLV 1330
Query: 625 LKVGVPIMLIRNIDQAAGLCNDTRMIVNALTKYIIVATILNGNKMGETTFIPRMSLTPSN 684
LK GV +ML+RN++Q+ GLCN TR++V L I+ IL G+ +GET +IPR++L +
Sbjct: 1331 LKEGVVVMLLRNLNQSIGLCNGTRLLVTVLGDRILQCIILTGSNIGETVYIPRITLGTTK 1390
Query: 685 SDIPFKFQRRQFPVTLCFAMTINKSQGQSLSHVGLYLPRPVFTHGQLYVALSRVKSRKGL 744
PF QRRQFPV +C++MTINKSQGQ+L VG+YL +PVFTHGQLYVA+SRV SR GL
Sbjct: 1391 MKWPFTLQRRQFPVRVCYSMTINKSQGQTLQRVGVYLRKPVFTHGQLYVAISRVTSRSGL 1450
Query: 745 KMLIIDDEGVVSNTTRNVMYQEV 767
K+LI +D+G T+N++Y EV
Sbjct: 1451 KILIENDDGSCGTQTKNIVYSEV 1473
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.347 0.154 0.513
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,116,969,183
Number of Sequences: 2790947
Number of extensions: 43812039
Number of successful extensions: 178905
Number of sequences better than 10.0: 277
Number of HSP's better than 10.0 without gapping: 146
Number of HSP's successfully gapped in prelim test: 131
Number of HSP's that attempted gapping in prelim test: 178163
Number of HSP's gapped (non-prelim): 499
length of query: 770
length of database: 848,049,833
effective HSP length: 135
effective length of query: 635
effective length of database: 471,271,988
effective search space: 299257712380
effective search space used: 299257712380
T: 11
A: 40
X1: 14 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.7 bits)
S2: 79 (35.0 bits)
Lotus: description of TM0178.5


