

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0176a.2
(342 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
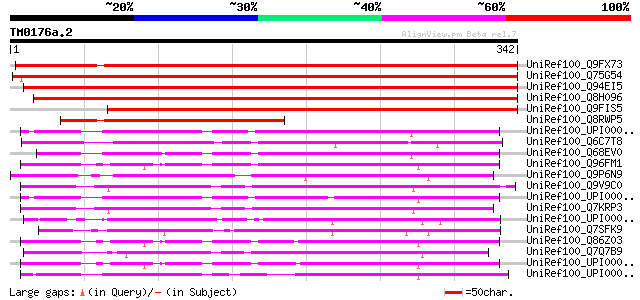
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9FX73 F19K19.12 protein [Arabidopsis thaliana] 532 e-150
UniRef100_Q75G54 Hypothetical protein B1003C08.3 [Oryza sativa] 489 e-137
UniRef100_Q94EI5 AT5g62130/mtg10_150 [Arabidopsis thaliana] 436 e-121
UniRef100_Q8H096 Hypothetical protein OSJNBa0096G08.20 [Oryza sa... 384 e-105
UniRef100_Q9FIS5 Arabidopsis thaliana genomic DNA, chromosome 5,... 366 e-100
UniRef100_Q8RWP5 Hypothetical protein At1g16560 [Arabidopsis tha... 250 5e-65
UniRef100_UPI0000365B72 UPI0000365B72 UniRef100 entry 198 2e-49
UniRef100_Q6C7T8 Yarrowia lipolytica chromosome D of strain CLIB... 197 3e-49
UniRef100_Q68EV0 MGC84367 protein [Xenopus laevis] 191 2e-47
UniRef100_Q96FM1 CAB2 protein [Homo sapiens] 189 1e-46
UniRef100_Q9P6N9 SPAC823.07 protein [Schizosaccharomyces pombe] 180 4e-44
UniRef100_Q9V9C0 CG3271-PB, isoform B [Drosophila melanogaster] 174 2e-42
UniRef100_UPI0000365B73 UPI0000365B73 UniRef100 entry 171 3e-41
UniRef100_Q7KRP3 CG3271-PA, isoform A [Drosophila melanogaster] 169 8e-41
UniRef100_UPI000023E7E4 UPI000023E7E4 UniRef100 entry 167 4e-40
UniRef100_Q7SFK9 Hypothetical protein [Neurospora crassa] 167 5e-40
UniRef100_Q86Z03 CAB2 [Homo sapiens] 164 2e-39
UniRef100_Q7Q7B9 ENSANGP00000021025 [Anopheles gambiae str. PEST] 163 6e-39
UniRef100_UPI000036AC80 UPI000036AC80 UniRef100 entry 159 1e-37
UniRef100_UPI000027CC32 UPI000027CC32 UniRef100 entry 157 4e-37
>UniRef100_Q9FX73 F19K19.12 protein [Arabidopsis thaliana]
Length = 342
Score = 532 bits (1371), Expect = e-150
Identities = 242/338 (71%), Positives = 276/338 (81%), Gaps = 4/338 (1%)
Query: 5 YVFAFLLVLSWSVEVIDASAGDVDPRYRDCIRQCQETGCVAQSCFPHCKFPSDGEHFDRP 64
Y A L+L + +ASAGD DP YR C+ +C+ +GCV Q CFP C SDG P
Sbjct: 5 YWTALFLLLPCLFCISNASAGDADPDYRTCVSECEISGCVGQLCFPQCNSSSDGG----P 60
Query: 65 WYMQEPLYLQWKKWDCQNDCRYHCMLDREKERKLLNLVPVKYHGKWPFARIYGMQEAASV 124
WY+QEPLYLQWKKW CQ DCRY CM++RE ER+ L PVKYHGKWPF R+ G+QE ASV
Sbjct: 61 WYIQEPLYLQWKKWGCQGDCRYQCMVNRETERETLGQAPVKYHGKWPFKRVLGIQEPASV 120
Query: 125 AFSALNLAMHFHGWVSFFILLYYKLPLKDGKEAYYEYAGLWHIYGLLSLNSWFWSAVFHS 184
AFS LNLAMHFHGW+SFFI++YYKLPLK + AYYEY GLWHIYGLLS+NSWFWSAVFHS
Sbjct: 121 AFSVLNLAMHFHGWLSFFIMIYYKLPLKQDRTAYYEYVGLWHIYGLLSMNSWFWSAVFHS 180
Query: 185 RDVDLTETLDYSSAVILLGYSLILAILRSFNVRDEATRVMVSAPLIAFVITHVMYINFYK 244
RDVDLTE LDYSSAV +LG+SLILAILR+F++R EA RVMVSAP++AFV TH++YINFYK
Sbjct: 181 RDVDLTERLDYSSAVAILGFSLILAILRTFDIRVEAARVMVSAPILAFVTTHILYINFYK 240
Query: 245 LDYGWNMIVCVVMAVVQLVIWAVWAGLSGHPSRWKLWLVVIAGGLAMLLEIYDFPPYEGL 304
LDYGWNMIVCV M V QL +WA WA +S HPS WKLW+VVIAGGLAMLLEIYDFPPYEG
Sbjct: 241 LDYGWNMIVCVAMGVSQLFLWARWAAVSSHPSNWKLWVVVIAGGLAMLLEIYDFPPYEGY 300
Query: 305 LDAHALWHATTIPLTYIWWSFIRDDAEFRTSIRVKKAK 342
DAH++WHA TIPLT +WWSFIRDDAEFRTS +KK K
Sbjct: 301 FDAHSIWHAATIPLTILWWSFIRDDAEFRTSSLLKKTK 338
>UniRef100_Q75G54 Hypothetical protein B1003C08.3 [Oryza sativa]
Length = 349
Score = 489 bits (1259), Expect = e-137
Identities = 219/344 (63%), Positives = 276/344 (79%), Gaps = 4/344 (1%)
Query: 3 GGYV--FAFLLVLSWSVEVIDASAGDVDPRYRDCIRQCQETGCVAQSCFPHCKFP-SDGE 59
GG+V A LLV+ + + +DAS GDVDP+YR C+ +C TG + ++ HC+ P +D
Sbjct: 6 GGWVARLAALLVVGFVLGSVDASLGDVDPQYRTCVEECHTTGIIGENIISHCQSPGNDDA 65
Query: 60 HFDRPWYMQEPLYLQWKKWDCQNDCRYHCMLDREKERKLLNLVPVKYHGKWPFARIYGMQ 119
WY QEPLY+QWK+ +C NDCRY+CM+ RE ER+ L PVKYHGKWPF R+ Q
Sbjct: 66 SVGSSWYTQEPLYMQWKQLNCMNDCRYYCMMQREGERQSRGLNPVKYHGKWPFIRVSVFQ 125
Query: 120 EAASVAFSALNLAMHFHGWVSFFILLYYKLPLK-DGKEAYYEYAGLWHIYGLLSLNSWFW 178
E S A SA+NL MHF GW+SFF+L+ YKLP++ K YYEY GLWHIY +LS+N+WFW
Sbjct: 126 EPLSAALSAVNLLMHFTGWLSFFLLVNYKLPVRPQTKRTYYEYTGLWHIYAILSMNAWFW 185
Query: 179 SAVFHSRDVDLTETLDYSSAVILLGYSLILAILRSFNVRDEATRVMVSAPLIAFVITHVM 238
S++FH+RD+DLTE LDYSSAV LLGYSLIL++LR+FNV+DEATRVM +AP++AFV TH++
Sbjct: 186 SSIFHTRDIDLTEKLDYSSAVALLGYSLILSLLRTFNVKDEATRVMFAAPILAFVTTHIL 245
Query: 239 YINFYKLDYGWNMIVCVVMAVVQLVIWAVWAGLSGHPSRWKLWLVVIAGGLAMLLEIYDF 298
Y+NFY+LDYGWNM VCVVMAVVQL+ WA+WAG++ HPSR+KLW+VV G LAMLLE+YDF
Sbjct: 246 YLNFYELDYGWNMKVCVVMAVVQLLAWAIWAGITQHPSRFKLWVVVFGGALAMLLEVYDF 305
Query: 299 PPYEGLLDAHALWHATTIPLTYIWWSFIRDDAEFRTSIRVKKAK 342
PPY+G DAH+LWHA+TIPLTY+WWSFI+DDAEFRTS +KKAK
Sbjct: 306 PPYKGYADAHSLWHASTIPLTYLWWSFIKDDAEFRTSTLIKKAK 349
>UniRef100_Q94EI5 AT5g62130/mtg10_150 [Arabidopsis thaliana]
Length = 343
Score = 436 bits (1122), Expect = e-121
Identities = 187/333 (56%), Positives = 256/333 (76%)
Query: 10 LLVLSWSVEVIDASAGDVDPRYRDCIRQCQETGCVAQSCFPHCKFPSDGEHFDRPWYMQE 69
++V+S V ++AS GD D Y+ C+ QCQ+TGCV +CF HCKF +DG+ D PWYMQE
Sbjct: 11 IIVVSCLVSTLEASEGDSDSLYKSCVDQCQKTGCVGDTCFQHCKFSADGKAIDGPWYMQE 70
Query: 70 PLYLQWKKWDCQNDCRYHCMLDREKERKLLNLVPVKYHGKWPFARIYGMQEAASVAFSAL 129
PLYL+WK+WDCQ+DC+Y CM+ RE+ERK P KY GKWP +YG+QE SVAFSAL
Sbjct: 71 PLYLRWKQWDCQSDCQYECMMTREEERKRNGERPTKYFGKWPLKHVYGIQEPVSVAFSAL 130
Query: 130 NLAMHFHGWVSFFILLYYKLPLKDGKEAYYEYAGLWHIYGLLSLNSWFWSAVFHSRDVDL 189
+LAM F GWVS+FIL+YYKLPL+ ++ YYEY G+ HIY ++ +NS FWS++ HSRDV+L
Sbjct: 131 DLAMQFQGWVSYFILVYYKLPLQPNRKTYYEYNGIVHIYAIIVMNSLFWSSICHSRDVEL 190
Query: 190 TETLDYSSAVILLGYSLILAILRSFNVRDEATRVMVSAPLIAFVITHVMYINFYKLDYGW 249
TE LDYSSA +L G+SLILAILRSF+++D++ ++MV+AP++A V TH++Y+NFY LD G
Sbjct: 191 TERLDYSSATVLAGFSLILAILRSFSIQDQSVKIMVTAPILAVVATHILYLNFYNLDEGL 250
Query: 250 NMIVCVVMAVVQLVIWAVWAGLSGHPSRWKLWLVVIAGGLAMLLEIYDFPPYEGLLDAHA 309
+ V + ++LV+W +WA L+ HPS+WKL +I+ L + L ++DFPPY+G +DAHA
Sbjct: 251 HWKVIFGIGGIELVVWGLWAALTSHPSKWKLRAFLISSILTLCLRMFDFPPYKGYIDAHA 310
Query: 310 LWHATTIPLTYIWWSFIRDDAEFRTSIRVKKAK 342
LW IPL+Y+WWSF+ DDA FRT++ +KK+K
Sbjct: 311 LWRGAGIPLSYLWWSFVCDDAVFRTTVNLKKSK 343
>UniRef100_Q8H096 Hypothetical protein OSJNBa0096G08.20 [Oryza sativa]
Length = 347
Score = 384 bits (985), Expect = e-105
Identities = 170/327 (51%), Positives = 233/327 (70%), Gaps = 1/327 (0%)
Query: 17 VEVIDASAGDVDPRYRDCIRQCQETGCVAQSCFPHCKFPSDGEHFDRPWYMQEPLYLQWK 76
V + AS GD DP YR C+ +C++TG + ++ HC+ P+D D+ WY EPLYLQWK
Sbjct: 21 VATLGASEGDADPLYRACVDECEKTGSLRETSVRHCQVPTDDHPADKSWYAHEPLYLQWK 80
Query: 77 KWDCQNDCRYHCMLDREKERKLLNLVPVKYHGKWPFARIYGMQEAASVAFSALNLAMHFH 136
+W+C+++CRYHCM++RE ER+ L L VKYHGKWP R QE S A SAL+L + F+
Sbjct: 81 EWNCKSECRYHCMMERESEREQLGLGSVKYHGKWPMKRASVFQEPISAALSALSLLVQFN 140
Query: 137 GWVSFFILLYYKLPLK-DGKEAYYEYAGLWHIYGLLSLNSWFWSAVFHSRDVDLTETLDY 195
GW+SFF+LL YKLPL+ + + YYEY GLWHIYGLL++N+WFW A++HS D TE L Y
Sbjct: 141 GWLSFFLLLSYKLPLRPETQMTYYEYTGLWHIYGLLAMNAWFWRAIYHSCDTVWTEKLYY 200
Query: 196 SSAVILLGYSLILAILRSFNVRDEATRVMVSAPLIAFVITHVMYINFYKLDYGWNMIVCV 255
SS +GYSLILAILR+ N++DEA+RVMV+AP++AF TH++Y+NFY+LD G N VC
Sbjct: 201 SSFAAFIGYSLILAILRTLNLKDEASRVMVAAPILAFTTTHILYLNFYELDKGLNTKVCT 260
Query: 256 VMAVVQLVIWAVWAGLSGHPSRWKLWLVVIAGGLAMLLEIYDFPPYEGLLDAHALWHATT 315
++ Q ++WAVWA ++ HPS +K+ V+I +++LE YD PP G +D A +
Sbjct: 261 AASLAQFLLWAVWAVMTKHPSCFKILFVIIGNVFSIVLETYDIPPRWGYVDGRVFCVAIS 320
Query: 316 IPLTYIWWSFIRDDAEFRTSIRVKKAK 342
IPLTY+WW F ++DAE RTS +KK +
Sbjct: 321 IPLTYLWWKFAKEDAEMRTSTIIKKTR 347
>UniRef100_Q9FIS5 Arabidopsis thaliana genomic DNA, chromosome 5, P1 clone:MTG10
[Arabidopsis thaliana]
Length = 276
Score = 366 bits (939), Expect = e-100
Identities = 158/276 (57%), Positives = 216/276 (78%)
Query: 67 MQEPLYLQWKKWDCQNDCRYHCMLDREKERKLLNLVPVKYHGKWPFARIYGMQEAASVAF 126
MQEPLYL+WK+WDCQ+DC+Y CM+ RE+ERK P KY GKWP +YG+QE SVAF
Sbjct: 1 MQEPLYLRWKQWDCQSDCQYECMMTREEERKRNGERPTKYFGKWPLKHVYGIQEPVSVAF 60
Query: 127 SALNLAMHFHGWVSFFILLYYKLPLKDGKEAYYEYAGLWHIYGLLSLNSWFWSAVFHSRD 186
SAL+LAM F GWVS+FIL+YYKLPL+ ++ YYEY G+ HIY ++ +NS FWS++ HSRD
Sbjct: 61 SALDLAMQFQGWVSYFILVYYKLPLQPNRKTYYEYNGIVHIYAIIVMNSLFWSSICHSRD 120
Query: 187 VDLTETLDYSSAVILLGYSLILAILRSFNVRDEATRVMVSAPLIAFVITHVMYINFYKLD 246
V+LTE LDYSSA +L G+SLILAILRSF+++D++ ++MV+AP++A V TH++Y+NFY LD
Sbjct: 121 VELTERLDYSSATVLAGFSLILAILRSFSIQDQSVKIMVTAPILAVVATHILYLNFYNLD 180
Query: 247 YGWNMIVCVVMAVVQLVIWAVWAGLSGHPSRWKLWLVVIAGGLAMLLEIYDFPPYEGLLD 306
G + V + ++LV+W +WA L+ HPS+WKL +I+ L + L ++DFPPY+G +D
Sbjct: 181 EGLHWKVIFGIGGIELVVWGLWAALTSHPSKWKLRAFLISSILTLCLRMFDFPPYKGYID 240
Query: 307 AHALWHATTIPLTYIWWSFIRDDAEFRTSIRVKKAK 342
AHALW IPL+Y+WWSF+ DDA FRT++ +KK+K
Sbjct: 241 AHALWRGAGIPLSYLWWSFVCDDAVFRTTVNLKKSK 276
>UniRef100_Q8RWP5 Hypothetical protein At1g16560 [Arabidopsis thaliana]
Length = 156
Score = 250 bits (638), Expect = 5e-65
Identities = 108/151 (71%), Positives = 122/151 (80%), Gaps = 4/151 (2%)
Query: 35 IRQCQETGCVAQSCFPHCKFPSDGEHFDRPWYMQEPLYLQWKKWDCQNDCRYHCMLDREK 94
I +C+ +GCV Q CFP C SDG PWY+QEPLYLQWKKW CQ DCRY CM++RE
Sbjct: 10 ITECEISGCVGQLCFPQCNSSSDGG----PWYIQEPLYLQWKKWGCQGDCRYQCMVNRET 65
Query: 95 ERKLLNLVPVKYHGKWPFARIYGMQEAASVAFSALNLAMHFHGWVSFFILLYYKLPLKDG 154
ER+ L PVKYHGKWPF R+ G+QE ASVAFS LNLAMHFHGW+SFFI++YYKLPLK
Sbjct: 66 ERETLGQAPVKYHGKWPFKRVLGIQEPASVAFSVLNLAMHFHGWLSFFIMIYYKLPLKQD 125
Query: 155 KEAYYEYAGLWHIYGLLSLNSWFWSAVFHSR 185
+ AYYEY GLWHIYGLLS+NSWFWSAVFHSR
Sbjct: 126 RTAYYEYVGLWHIYGLLSMNSWFWSAVFHSR 156
>UniRef100_UPI0000365B72 UPI0000365B72 UniRef100 entry
Length = 306
Score = 198 bits (504), Expect = 2e-49
Identities = 114/325 (35%), Positives = 163/325 (50%), Gaps = 29/325 (8%)
Query: 8 AFLLVLSWSVEVIDASAGDVDPRYRDCIRQCQETGCVAQSCFPHCKFPSDGEHFDRPWYM 67
A LL L V + +S GD +P YRDC++ C T C R +
Sbjct: 5 ALLLAL---VTTVQSSQGDKEPVYRDCVKLCVRTNCTGARL--------------RGFQS 47
Query: 68 QEPLYLQWKKWDCQNDCRYHCMLDREKERKLLNLVPVKYHGKWPFARIYGMQEAASVAFS 127
+P Y+ W C++DCRY CM + ++HGKWPFAR +E AS S
Sbjct: 48 AQPHYMALTGWTCRDDCRYQCMWTTVGLYQAEGYRVPQFHGKWPFARFLCFEEPASALAS 107
Query: 128 ALNLAMHFHGWVSFFILLYYKLPLKDGKEAYYEYAGLWHIYGLLSLNSWFWSAVFHSRDV 187
LN G +LL Y+ + Y+ + + L+SLN+WFWS VFH+RD
Sbjct: 108 LLN------GLACLLMLLRYRSTVPRQSPMYHTI----NAFSLVSLNAWFWSTVFHTRDT 157
Query: 188 DLTETLDYSSAVILLGYSLILAILRSFNVRDEATRVMVSAPLIAFVITHVMYINFYKLDY 247
LTE +DY A ++ YS+ L +R+ +R A +V LI +HV Y+ F DY
Sbjct: 158 YLTEKMDYFCATAVILYSIYLCCVRTLGLRRPAVSSIVGVFLILAFTSHVSYLTFVSFDY 217
Query: 248 GWNMIVCVVMAVVQLVIWAVWA--GLSGHPSRWKLWLVVIAGGLAMLLEIYDFPPYEGLL 305
G+NM + +V L+ W W P WK LVV+ LLE+ DFPP +L
Sbjct: 218 GYNMAANATIGLVNLLWWLCWCWQNRGTLPYWWKCGLVVLLLHGLALLELLDFPPMLWIL 277
Query: 306 DAHALWHATTIPLTYIWWSFIRDDA 330
DAHA+WH +TIP+ ++++SF+ DD+
Sbjct: 278 DAHAVWHLSTIPVHFLFYSFLIDDS 302
>UniRef100_Q6C7T8 Yarrowia lipolytica chromosome D of strain CLIB99 of Yarrowia
lipolytica [Yarrowia lipolytica]
Length = 313
Score = 197 bits (501), Expect = 3e-49
Identities = 110/332 (33%), Positives = 172/332 (51%), Gaps = 37/332 (11%)
Query: 9 FLLVLSWSVEVIDASAGDVDPRYRDCIRQCQETGCVAQSCFPHCKFPSDGEHFDRPWYMQ 68
F ++L ++ AS GD P +R+C+ C C Q P
Sbjct: 3 FYVILVLLTTLVLASVGDRSPDFRNCVTNCIRHTCQTQKYVP------------------ 44
Query: 69 EPLYLQWKKWDCQNDCRYHCMLDREKERKLLNLVPVKYHGKWPFARIYGMQEAASVAFSA 128
PL + WDC +C Y C R V++HGKWPF R +G+QE ASV FS
Sbjct: 45 -PLMHRLLLWDCPQECDYRCQQIITFARLNQGQEIVQFHGKWPFFRFFGIQELASVVFSL 103
Query: 129 LNLAMHFHGWVSFFILLYYKLPLKDGKEAYYEYAGLWHIYGLLSLNSWFWSAVFHSRDVD 188
N H+ GW L+ L + Y G + L+ +NSW WSAVFH+RD
Sbjct: 104 ANFVPHYRGW-----LMLKHLNQRKPNPLIPYYIG----FALVGMNSWIWSAVFHTRDFP 154
Query: 189 LTETLDYSSAVILLGYSLILAILRSFNV-RD--EATRVMVSAPLIAFVITHVMYINFYKL 245
+TE LDY SA + + Y A +R F + RD E TR+++++ + + HV Y++F K
Sbjct: 155 VTEKLDYFSAGLSVLYGFFFATVRIFRLDRDSRETTRLVLASVCVTLFLAHVSYLSFIKF 214
Query: 246 DYGWNMIVCVVMAVVQLVIWAVWAGLSGHPSRWKLWLVVIAG-----GLAMLLEIYDFPP 300
DYG+NM VV+ +QL++W+V++ + + W ++ G AM LE++DFPP
Sbjct: 215 DYGYNMTANVVVGALQLIMWSVYS-FTQFAKTHQWWSLMPFGLCVTISAAMGLELFDFPP 273
Query: 301 YEGLLDAHALWHATTIPLTYIWWSFIRDDAEF 332
++ +DAH+LWHA T+ ++W+++++ D ++
Sbjct: 274 WKFFIDAHSLWHAATVIPCFLWYTWMKKDLQY 305
>UniRef100_Q68EV0 MGC84367 protein [Xenopus laevis]
Length = 317
Score = 191 bits (486), Expect = 2e-47
Identities = 109/315 (34%), Positives = 160/315 (50%), Gaps = 28/315 (8%)
Query: 19 VIDASAGDVDPRYRDCIRQCQETGCVAQSCFPHCKFPSDGEHFDRPWYMQEPLYLQWKKW 78
V+ AS GD +P YRDC+ C + C R + Q+PLY++ W
Sbjct: 12 VVSASRGDREPVYRDCVTVCDQNNCTGFRL--------------RDFRAQQPLYMRLTGW 57
Query: 79 DCQNDCRYHCM-LDREKERKLLNLVPVKYHGKWPFARIYGMQEAASVAFSALNLAMHFHG 137
C +DCRY CM K + VP ++HGKWPF+R QE AS S LN G
Sbjct: 58 TCLDDCRYKCMWYTVSLYLKEGHEVP-QFHGKWPFSRFLFFQEPASALASFLN------G 110
Query: 138 WVSFFILLYYKLPLKDGKEAYYEYAGLWHIYGLLSLNSWFWSAVFHSRDVDLTETLDYSS 197
S +L Y+ + + Y + ++S+N+WFWS +FH+RD LTE +DY
Sbjct: 111 VASLLMLFRYRSSVPSSCQMYRTCLA----FSMVSVNAWFWSTIFHTRDTALTEKMDYFC 166
Query: 198 AVILLGYSLILAILRSFNVRDEATRVMVSAPLIAFVITHVMYINFYKLDYGWNMIVCVVM 257
A ++ +S+ L +R+F ++ + A L+ H+ Y+ + DY +NM
Sbjct: 167 ASSVILHSIYLCCMRTFGLQYPSIANAFGAFLVLLFACHISYLTLGRFDYSYNMAANTSF 226
Query: 258 AVVQLVIWAVWA--GLSGHPSRWKLWLVVIAGGLAMLLEIYDFPPYEGLLDAHALWHATT 315
+V L+ W W P WK LVV+ LLE+ DFPP +LDAHALWH +T
Sbjct: 227 GIVNLMWWLAWCMWRRFHQPYLWKCVLVVVLLQSLALLELLDFPPVMWILDAHALWHFST 286
Query: 316 IPLTYIWWSFIRDDA 330
IPL ++++SF+RDD+
Sbjct: 287 IPLHFLFYSFLRDDS 301
>UniRef100_Q96FM1 CAB2 protein [Homo sapiens]
Length = 320
Score = 189 bits (479), Expect = 1e-46
Identities = 111/330 (33%), Positives = 173/330 (51%), Gaps = 36/330 (10%)
Query: 8 AFLLVLSWSVEVIDASAGDVDPRYRDCIRQCQETGCVAQSCFPHCKFPSDGEHFDRPWYM 67
A L++L+ + + S GD +P YRDC+ QC+E C + HF
Sbjct: 6 ARLVLLAGAAALASGSQGDREPVYRDCVLQCEEQNCSGGAL----------NHFRS---- 51
Query: 68 QEPLYLQWKKWDCQNDCRYHCM-----LDREKERKLLNLVPVKYHGKWPFARIYGMQEAA 122
++P+Y+ W C++DC+Y CM L ++ K VP ++HGKWPF+R QE A
Sbjct: 52 RQPIYMSLAGWTCRDDCKYECMWVTVGLYLQEGHK----VP-QFHGKWPFSRFLFFQEPA 106
Query: 123 SVAFSALNLAMHFHGWVSFFILLYYKLPLKDGKEAYYEYAGLWHIYGLLSLNSWFWSAVF 182
S S LN G S +L Y+ + Y+ + +SLN+WFWS VF
Sbjct: 107 SAVASFLN------GLASLVMLCRYRTFVPASSPMYHTCVA----FAWVSLNAWFWSTVF 156
Query: 183 HSRDVDLTETLDYSSAVILLGYSLILAILRSFNVRDEATRVMVSAPLIAFVITHVMYINF 242
H+RD DLTE +DY A ++ +S+ L +R+ ++ A A L+ + HV Y++
Sbjct: 157 HTRDTDLTEKMDYFCASTVILHSIYLCCVRTVGLQHPAVVSAFRALLLLMLTVHVSYLSL 216
Query: 243 YKLDYGWNMIVCVVMAVVQLVIWAVWAGLSGH--PSRWKLWLVVIAGGLAMLLEIYDFPP 300
+ DYG+N++ V + +V +V W W + P K +VV+ LLE+ DFPP
Sbjct: 217 IRFDYGYNLVANVAIGLVNVVWWLAWCLWNQRRLPHVRKCVVVVLLLQGLSLLELLDFPP 276
Query: 301 YEGLLDAHALWHATTIPLTYIWWSFIRDDA 330
+LDAHA+WH +TIP+ +++SF+ DD+
Sbjct: 277 LFWVLDAHAIWHISTIPVHVLFFSFLEDDS 306
>UniRef100_Q9P6N9 SPAC823.07 protein [Schizosaccharomyces pombe]
Length = 331
Score = 180 bits (457), Expect = 4e-44
Identities = 109/333 (32%), Positives = 167/333 (49%), Gaps = 33/333 (9%)
Query: 1 MIGGYVFAFLLVLSWSVEVIDASAGDVDPRYRDCIRQCQETGCVAQSCFPHCKFPSDGEH 60
++ + FL I ASAGD+ P Y C+ +C E C PSD
Sbjct: 3 VLRNFTIFFLFTALSLFRQISASAGDLHPVYVSCVNRCIENKCHGN--------PSDTSK 54
Query: 61 FDRPWYMQEPLYLQWKKWDCQNDCRYHCMLDREKERKLLNLVPVKYHGKWPFARIYGMQE 120
PL L+ +WDC ++C Y C + E NL +YHGKW F R++G+QE
Sbjct: 55 L--------PLDLKLFRWDCGSNCGYECEITAENYFAAHNLPSQQYHGKWYFIRVFGIQE 106
Query: 121 AASVAFSALNLAMHFHGWVSFFILLYYKLPLKDGKEAYYEYAGLWHIYGLLSLNSWFWSA 180
SV FS LN +H++G+ + + P K L + ++ +N+W WS+
Sbjct: 107 LFSVFFSMLNFMIHYNGYHIMRRCIPDEHPAK----------RLCLSWAIVGMNAWVWSS 156
Query: 181 VFHSRDVDLTETLDYSSA---VILLGYSLILAILRSFNV-RDEATRVMVSAPLIAFVITH 236
VFH RD +TE LDY SA V+ Y ++ +LR + + ++ IA I H
Sbjct: 157 VFHIRDTPITEKLDYFSAGAFVLFGSYCTLILMLRLDQLPGGKLLCWIIGVIFIAAFIAH 216
Query: 237 VMYINFYKLDYGWNMIVCVVMAVVQLVIWAVWAGLSGHPS-RWKLW--LVVIAGGLAMLL 293
V Y++FY DYG+NM V + +VQ ++W ++ + + W W +V + LA L
Sbjct: 217 VSYLSFYSFDYGYNMKANVAVGLVQNILWYYYSWSNRNSGLYWTRWPAYIVTSLMLATSL 276
Query: 294 EIYDFPPYEGLLDAHALWHATTIPLTYIWWSFI 326
E++DF P L+DAHALWH +T+P+T+ + F+
Sbjct: 277 ELFDFSPIANLIDAHALWHLSTVPITHYLYGFV 309
>UniRef100_Q9V9C0 CG3271-PB, isoform B [Drosophila melanogaster]
Length = 330
Score = 174 bits (442), Expect = 2e-42
Identities = 107/338 (31%), Positives = 170/338 (49%), Gaps = 28/338 (8%)
Query: 8 AFLLVLSWSVEVIDASAGDVDPRYRDCIRQCQETGCVAQSCFPHCKFPSDGEHFDRPW-- 65
A +L+L V AS GD + +C + C+ T C A DG
Sbjct: 8 AIVLLLGALVTACLASNGDRTQFFHNCRQNCERTNCSA-----------DGLEIQEQAVK 56
Query: 66 YMQEPLYLQWKKWDCQNDCRYHCMLDREKERKLLNLVPVKYHGKWPFARIYGMQEAASVA 125
+ Q+ ++ + +W C ++C+Y CM +++GKWPF R+ GMQE ASV
Sbjct: 57 FYQQSVFDRLFQWSCADECQYGCMWRTVFAFFERGWPIPQFYGKWPFLRLLGMQEPASVI 116
Query: 126 FSALNLAMHFHGWVSFFILLYYKLPLKDGKEAYYEYAGLWHIYGLLSLNSWFWSAVFHSR 185
FS LN +H +L ++ ++ Y L HI+ + SLN W WSA+FH+R
Sbjct: 117 FSCLNFVVHLR------LLRKFRREVRPDSPCYM----LTHIFAVTSLNGWIWSAIFHTR 166
Query: 186 DVDLTETLDYSSAVILLGYSLILAILRSFNVRDEATRVMVSAPLIAFVITHVMYINFYKL 245
D LTE LDY+ A ++ SL + ++R + R +++ +++ I + Y++ +
Sbjct: 167 DFPLTELLDYAFAYSIILCSLYVMVMRMLHRYSLFLRGVITLAFLSYYINYFAYLSVGRF 226
Query: 246 DYGWNMIVCVVMAVVQLVIWAVWAGL--SGHPSRWKLWLVVIAGGLAMLLEIYDFPPYEG 303
+Y +NM+V V V+ V W VW + P ++ I LAM LE+ DFPP
Sbjct: 227 NYAFNMMVNVATGVIAAVGWFVWCHFVRTRRPYFRRILRFYILMALAMSLELLDFPPILW 286
Query: 304 LLDAHALWHATTIPLTYIWWSFIRDDAEFRTSIRVKKA 341
+LDAHALWH TIPL +++ F+ +D ++R +KA
Sbjct: 287 ILDAHALWHLATIPLASLYYDFMIEDCR---TLRKEKA 321
>UniRef100_UPI0000365B73 UPI0000365B73 UniRef100 entry
Length = 303
Score = 171 bits (433), Expect = 3e-41
Identities = 106/325 (32%), Positives = 155/325 (47%), Gaps = 32/325 (9%)
Query: 8 AFLLVLSWSVEVIDASAGDVDPRYRDCIRQCQETGCVAQSCFPHCKFPSDGEHFDRPWYM 67
A LL L V + +S GD +P YRDC++ C T C R +
Sbjct: 5 ALLLAL---VTTVQSSQGDKEPVYRDCVKLCVRTNCTGARL--------------RGFQS 47
Query: 68 QEPLYLQWKKWDCQNDCRYHCMLDREKERKLLNLVPVKYHGKWPFARIYGMQEAASVAFS 127
+P Y+ W C++DCRY CM + ++HGKWPFAR +E AS S
Sbjct: 48 AQPHYMALTGWTCRDDCRYQCMWTTVGLYQAEGYRVPQFHGKWPFARFLCFEEPASALAS 107
Query: 128 ALNLAMHFHGWVSFFILLYYKLPLKDGKEAYYEYAGLWHIYGLLSLNSWFWSAVFHSRDV 187
LN G +LL Y+ + Y+ + + L+SLN+WFWS VFH+RD
Sbjct: 108 LLN------GLACLLMLLRYRSTVPRQSPMYHTI----NAFSLVSLNAWFWSTVFHTRDT 157
Query: 188 DLTETLDYSSAVILLGYSLILAILRSFNVRDEATRVMVSAPLIAFVITHVMYINFYKLDY 247
L T+ + +++ L + A +V LI +HV Y+ F DY
Sbjct: 158 YLKWTISVQQQSSSTQSTCVVSELWACG---PAVSSIVGVFLILAFTSHVSYLTFVSFDY 214
Query: 248 GWNMIVCVVMAVVQLVIWAVWAGLSGH--PSRWKLWLVVIAGGLAMLLEIYDFPPYEGLL 305
G+NM + +V L+ W W + P WK LVV+ LLE+ DFPP +L
Sbjct: 215 GYNMAANATIGLVNLLWWLCWCWQNRGTLPYWWKCGLVVLLLHGLALLELLDFPPMLWIL 274
Query: 306 DAHALWHATTIPLTYIWWSFIRDDA 330
DAHA+WH +TIP+ ++++SF+ DD+
Sbjct: 275 DAHAVWHLSTIPVHFLFYSFLIDDS 299
>UniRef100_Q7KRP3 CG3271-PA, isoform A [Drosophila melanogaster]
Length = 326
Score = 169 bits (429), Expect = 8e-41
Identities = 102/323 (31%), Positives = 161/323 (49%), Gaps = 25/323 (7%)
Query: 8 AFLLVLSWSVEVIDASAGDVDPRYRDCIRQCQETGCVAQSCFPHCKFPSDGEHFDRPW-- 65
A +L+L V AS GD + +C + C+ T C A DG
Sbjct: 8 AIVLLLGALVTACLASNGDRTQFFHNCRQNCERTNCSA-----------DGLEIQEQAVK 56
Query: 66 YMQEPLYLQWKKWDCQNDCRYHCMLDREKERKLLNLVPVKYHGKWPFARIYGMQEAASVA 125
+ Q+ ++ + +W C ++C+Y CM +++GKWPF R+ GMQE ASV
Sbjct: 57 FYQQSVFDRLFQWSCADECQYGCMWRTVFAFFERGWPIPQFYGKWPFLRLLGMQEPASVI 116
Query: 126 FSALNLAMHFHGWVSFFILLYYKLPLKDGKEAYYEYAGLWHIYGLLSLNSWFWSAVFHSR 185
FS LN +H +L ++ ++ Y L HI+ + SLN W WSA+FH+R
Sbjct: 117 FSCLNFVVHLR------LLRKFRREVRPDSPCYM----LTHIFAVTSLNGWIWSAIFHTR 166
Query: 186 DVDLTETLDYSSAVILLGYSLILAILRSFNVRDEATRVMVSAPLIAFVITHVMYINFYKL 245
D LTE LDY+ A ++ SL + ++R + R +++ +++ I + Y++ +
Sbjct: 167 DFPLTELLDYAFAYSIILCSLYVMVMRMLHRYSLFLRGVITLAFLSYYINYFAYLSVGRF 226
Query: 246 DYGWNMIVCVVMAVVQLVIWAVWAGL--SGHPSRWKLWLVVIAGGLAMLLEIYDFPPYEG 303
+Y +NM+V V V+ V W VW + P ++ I LAM LE+ DFPP
Sbjct: 227 NYAFNMMVNVATGVIAAVGWFVWCHFVRTRRPYFRRILRFYILMALAMSLELLDFPPILW 286
Query: 304 LLDAHALWHATTIPLTYIWWSFI 326
+LDAHALWH TIPL +++ +
Sbjct: 287 ILDAHALWHLATIPLASLYYECV 309
>UniRef100_UPI000023E7E4 UPI000023E7E4 UniRef100 entry
Length = 331
Score = 167 bits (423), Expect = 4e-40
Identities = 105/331 (31%), Positives = 172/331 (51%), Gaps = 36/331 (10%)
Query: 10 LLVLSWSVEVIDASAGDVDPRYRDCIRQCQETGCVAQSCFPHCKFPSDGEHFDRPWYMQE 69
+LVL++++ V DAS GD P ++DC++ C A++C P+ +P
Sbjct: 17 VLVLTFAITV-DASTGDRLPEFKDCLKVCN-----AENCAPN-----------KP-QTPI 58
Query: 70 PLYLQWKKWDCQNDCRYHCMLDREKERKLLNLVPVKYHGKWPFARIYGMQEAASVAFSAL 129
P+ + W+C ++C Y C +R L +++GKWPF R GMQE SV FS
Sbjct: 59 PVLHRLLLWNCASECDYACQHIVTGQRMATGLSVEQFYGKWPFYRFLGMQEPFSVLFSLG 118
Query: 130 NLAMHFHGWVSFFILLYYKLPLKDGKEAYYEYAGLWHIYGLLSLNSWFWSAVFHSRDVDL 189
NL H++G + + ++P +Y+ W Y + + SW +S++FH+RD +
Sbjct: 119 NLWAHWYGLKT---MDQARIPKSYSMRIFYD----WLAY--IGIASWTFSSIFHTRDFHV 169
Query: 190 TETLDYSSAVILLGYSLILAILRSFNV-----RDEATRVMVSAPLIAFVITHVMYINFYK 244
TE LDY +A + Y L ++R F + R T + S + ++HV Y+ F +
Sbjct: 170 TEELDYFAAGASVLYGLYYTVVRVFRLDKRTPRRRTTLRLWSLLCASLFLSHVSYLKFVR 229
Query: 245 LDYGWNMIVCVVMAVVQLVIWAVWAGLSGHPSR--WKLWLVVIAGGL--AMLLEIYDFPP 300
DY +NM V +VQ V+W+ ++ SR W W + + AM +E++DFPP
Sbjct: 230 WDYTYNMAANVAAGIVQHVLWSWFSFNRYRESRRIWAAWPGFVVAWIIFAMSMELFDFPP 289
Query: 301 YEGLLDAHALWHATTIPLTYIWWSFIRDDAE 331
+ G +DAH+LWH TI T +W++F+ DA+
Sbjct: 290 WLGCIDAHSLWHLMTIGPTILWYNFLVKDAQ 320
>UniRef100_Q7SFK9 Hypothetical protein [Neurospora crassa]
Length = 345
Score = 167 bits (422), Expect = 5e-40
Identities = 109/327 (33%), Positives = 155/327 (47%), Gaps = 40/327 (12%)
Query: 20 IDASAGDVDPRYRDCIRQCQETGCVAQSCFPHCKFPSDGEHFDRPWYMQEPLYLQWKKWD 79
+ AS GD P +++CIR C+ C D EH PL+ + W
Sbjct: 33 VAASIGDRLPEFQECIRVCERENC-----------GPDAEH-----QTPIPLHRRLLLWS 76
Query: 80 CQNDCRYHCMLDREKERKLLNLVP-----VKYHGKWPFARIYGMQEAASVAFSALNLAMH 134
C ++C Y C R + P V+YHGKWPF R GMQE SV FS N H
Sbjct: 77 CPSECDYTCQHLTTSSRLSQSPPPLPHPVVQYHGKWPFIRFLGMQEPLSVLFSLGNFWAH 136
Query: 135 FHGWVSFFILLYYKLPLKDGKEAYYEYAGLWHIYGLLSLNSWFWSAVFHSRDVDLTETLD 194
+ G LY K+ Y + + + + SWF+SAVFH+RD +TE LD
Sbjct: 137 YQG-------LYTKI--LPNIPPSYPLRKWYILLSYVGMASWFFSAVFHTRDFPVTEQLD 187
Query: 195 YSSAVILLGYSLILAILRSFNV------RDEATRVMVSAPLIAFVITHVMYINFYKLDYG 248
Y +A + Y L ++R F + R E+ + +A I + HV Y+ + DY
Sbjct: 188 YFAAGANVLYGLYYTVVRIFRLDKKDTPRRESLLRLWTALCILMYVAHVTYLKMWAWDYT 247
Query: 249 WNMIVCVVMAVVQLVIWA--VWAGLSGHPSRWKLW--LVVIAGGLAMLLEIYDFPPYEGL 304
+NM V + +Q ++W+ W W W +VV +AM LE+ DFPP G
Sbjct: 248 YNMAANVAVGAIQNLLWSWYSWTRYREQKKGWAAWPGIVVAWVLVAMSLELLDFPPLWGS 307
Query: 305 LDAHALWHATTIPLTYIWWSFIRDDAE 331
+DAH+LWHA TI T IW++F+ DA+
Sbjct: 308 VDAHSLWHAGTIVPTIIWYNFLVKDAQ 334
>UniRef100_Q86Z03 CAB2 [Homo sapiens]
Length = 319
Score = 164 bits (416), Expect = 2e-39
Identities = 105/331 (31%), Positives = 166/331 (49%), Gaps = 39/331 (11%)
Query: 8 AFLLVLSWSVEVIDASAGDVDPRYRDCIRQCQETGCVAQSCFPHCKFPSDGEHFDRPWYM 67
A L++L+ + + S GD +P YRDC+ QC+E C + HF
Sbjct: 6 ARLVLLAGAAALASGSQGDREPVYRDCVLQCEEQNCSGGAL----------NHFRS---- 51
Query: 68 QEPLYLQWKKWDCQNDCRYHCM-----LDREKERKLLNLVPVKYHGKWPFARIYGMQEAA 122
++P+Y+ W C++DC+Y CM L ++ K VP ++HGKWPF+R QE A
Sbjct: 52 RQPIYMSLAGWTCRDDCKYECMWVTVGLYLQEGHK----VP-QFHGKWPFSRFLFFQEPA 106
Query: 123 SVAFSALNLAMHFHGWVSFFILLYYKLPLKDGKEAYYEYAGLWHIYGLLSLNSWFWSAVF 182
S S LN G S +L Y+ + Y+ + +SLN+WFWS VF
Sbjct: 107 SAVASFLN------GLASLVMLCRYRTFVPASSPMYHTCVA----FAWVSLNAWFWSTVF 156
Query: 183 HSRDVDLTETLDYSSAVILLGYSL-ILAILRSFNVRDEATRVMVSAPLIAFVITHVMYIN 241
H+RD DL ++++V + Y+ A + + A L+ + HV Y++
Sbjct: 157 HTRDTDLQRK--WTTSVPPVSYTQSTCAASGPWGCSTQLWSSAFRALLLLMLTVHVSYLS 214
Query: 242 FYKLDYGWNMIVCVVMAVVQLVIWAVWAGLSGH--PSRWKLWLVVIAGGLAMLLEIYDFP 299
+ DYG+N++ V + +V +V W W + P K +VV+ LLE+ DFP
Sbjct: 215 LIRFDYGYNLVANVAIGLVNVVWWLAWCLWNQRRLPHVRKCVVVVLLLQGLSLLELLDFP 274
Query: 300 PYEGLLDAHALWHATTIPLTYIWWSFIRDDA 330
P +LDAHA+WH +TIP+ +++SF+ DD+
Sbjct: 275 PLFWVLDAHAIWHISTIPVHVLFFSFLEDDS 305
>UniRef100_Q7Q7B9 ENSANGP00000021025 [Anopheles gambiae str. PEST]
Length = 302
Score = 163 bits (413), Expect = 6e-39
Identities = 96/324 (29%), Positives = 148/324 (45%), Gaps = 39/324 (12%)
Query: 10 LLVLSWSVEVIDASAGDVDPRYRDCIRQCQETGCVAQSCFPHCKFPSDGEHFDRPWYMQE 69
+++L + +I AS GD Y++C++ C C Y +E
Sbjct: 8 IVLLIYFFRLIFASGGDRSQFYQNCLKFCTLDNCTQSGVS----------------YKKE 51
Query: 70 PLYLQWKK--------WDCQNDCRYHCMLDREKERKLLNLVPVKYHGKWPFARIYGMQEA 121
LQWK W C ++C Y CM N +++GKWPF R GMQE
Sbjct: 52 ---LQWKHDPINKLLLWTCYDECGYDCMWRTTAAFHNRNWTTPQFYGKWPFVRFLGMQEP 108
Query: 122 ASVAFSALNLAMHFHGWVSFFILLYYKLPLKDGKEAYYEYAGLWHIYGLLSLNSWFWSAV 181
ASV FS N A H+ +L ++ ++ Y G W + + LN+W WSA
Sbjct: 109 ASVLFSVANFATHYK------MLQRFRREVRTDSPMY----GTWRAFSYICLNAWIWSAF 158
Query: 182 FHSRDVDLTETLDYSSAVILLGYSLILAILRSFNVRDEATRVMVSAPLIAFVITHVMYIN 241
FH+RD +TE LDY+ A ++ S ++R + R S + F + H Y++
Sbjct: 159 FHTRDFPVTELLDYTFAYSMVLASFHCMVMRMIHRSSIVVRGAFSCLCVLFFVNHFSYLS 218
Query: 242 FYKLDYGWNMIVCVVMAVVQLVIWAVWAGLSGHPSR--WKLWLVVIAGGLAMLLEIYDFP 299
+ DY +NM +V + + W +W L R WK + ++ ++LLEI DFP
Sbjct: 219 VGRFDYSYNMKANIVTGMSGALGWILWCFLQRKKRRYVWKCFTFIVLATSSLLLEINDFP 278
Query: 300 PYEGLLDAHALWHATTIPLTYIWW 323
P DAH++WH T PLT +++
Sbjct: 279 PILWTFDAHSIWHLVTAPLTILFY 302
>UniRef100_UPI000036AC80 UPI000036AC80 UniRef100 entry
Length = 318
Score = 159 bits (401), Expect = 1e-37
Identities = 104/330 (31%), Positives = 160/330 (47%), Gaps = 38/330 (11%)
Query: 8 AFLLVLSWSVEVIDASAGDVDPRYRDCIRQCQETGCVAQSCFPHCKFPSDGEHFDRPWYM 67
A L++L+ + + S GD +P YRDC+ QC+E C + HF
Sbjct: 6 ARLVLLAGAAALASGSQGDREPVYRDCVLQCEEQNCSGGAL----------NHFRS---- 51
Query: 68 QEPLYLQWKKWDCQNDCRYHCM-----LDREKERKLLNLVPVKYHGKWPFARIYGMQEAA 122
++P+Y+ W C++DC+Y CM L ++ K VP ++HGKWPF+R QE A
Sbjct: 52 RQPIYMSLAGWTCRDDCKYECMWVTVGLYLQEGHK----VP-QFHGKWPFSRFLFFQEPA 106
Query: 123 SVAFSALNLAMHFHGWVSFFILLYYKLPLKDGKEAYYEYAGLWHIYGLLSLNSWFWSAVF 182
S S LN G S +L Y+ + Y+ + +SLN+WFWS VF
Sbjct: 107 SAVASFLN------GLASLVMLCRYRTFVPASSPMYHTCVA----FAWVSLNAWFWSTVF 156
Query: 183 HSRDVDLTETLDYSSAVILLGYSLILAILRSFNVRDEATRVMVSAPLIAFVITHVMYINF 242
H+RD D + +S L Y+ A L+ + HV Y++
Sbjct: 157 HTRDTDQRKWT--TSVPPLSSYTQSTCAASGPWGCSTQLWSAFRALLLLMLTVHVSYLSL 214
Query: 243 YKLDYGWNMIVCVVMAVVQLVIWAVWAGLSGH--PSRWKLWLVVIAGGLAMLLEIYDFPP 300
+ DYG+N++ V + +V +V W W + P K +VV+ LLE+ DFPP
Sbjct: 215 IRFDYGYNLVANVAIGLVNVVWWLAWCLWNQRRLPHVRKCVVVVLLLQGLSLLELLDFPP 274
Query: 301 YEGLLDAHALWHATTIPLTYIWWSFIRDDA 330
+LDAHA+WH +TIP+ +++SF+ DD+
Sbjct: 275 LFWVLDAHAIWHISTIPVHVLFFSFLEDDS 304
>UniRef100_UPI000027CC32 UPI000027CC32 UniRef100 entry
Length = 318
Score = 157 bits (397), Expect = 4e-37
Identities = 100/332 (30%), Positives = 153/332 (45%), Gaps = 49/332 (14%)
Query: 8 AFLLVLSWSVEVIDASAGDVDPRYRDCIRQCQETGCVAQSCFPHCKFPSDGEHFDRPWYM 67
A +++L+W+ + +S GD +P YRDC++ C T C R +
Sbjct: 3 AVVILLAWT-STVQSSPGDKEPVYRDCVKLCVRTNCTGARL--------------RGFQS 47
Query: 68 QEPLYLQWKKWDCQNDCRYHCMLDREKERKLLNLVPVKYHGKWPFARIYGMQEAASVAFS 127
+P Y+ W C++DCRY CM + ++HGKWPFAR +E AS S
Sbjct: 48 AQPQYMALTGWTCRDDCRYQCMWTTVGLYQAEGYRVPQFHGKWPFARFLCFEEPASALAS 107
Query: 128 ALNLAMHFHGWVSFFILLYYKLPLKDGKEAYYEYAGLWHIYGLLSLNSWFWSAVFHSRDV 187
LN G +LL Y+ A + ++H SL
Sbjct: 108 LLN------GLACLLMLLRYR-------SAVPRQSPMYHTINAFSLK------------- 141
Query: 188 DLTETLDYSSAVILLGYSLILAILRSFNVRDEATRVMVSAPLIAFVITHVMYINFYKLDY 247
+DY A ++ YS+ L +R+ +R A +V LI +HV Y+ F DY
Sbjct: 142 -----MDYFCATAVILYSIYLCCVRTLGLRRPAVSSIVGVFLILAFTSHVSYLTFVSFDY 196
Query: 248 GWNMIVCVVMAVVQLVIWAVWAGLSGH--PSRWKLWLVVIAGGLAMLLEIYDFPPYEGLL 305
G+NM + +V L+ W W + P WK LVV+ LLE+ DFPP +L
Sbjct: 197 GYNMAANTSIGLVNLLWWLCWCWQNRGTLPYWWKCGLVVLLLHGLALLELLDFPPMLWVL 256
Query: 306 DAHALWHATTIPLTYIWWSFIR-DDAEFRTSI 336
DAHA+WH +TIP+ ++++ F+ ++ EF TS+
Sbjct: 257 DAHAVWHLSTIPVHFLFYRFVEVEEREFATSL 288
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.328 0.142 0.485
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 612,780,656
Number of Sequences: 2790947
Number of extensions: 25857225
Number of successful extensions: 83359
Number of sequences better than 10.0: 59
Number of HSP's better than 10.0 without gapping: 43
Number of HSP's successfully gapped in prelim test: 18
Number of HSP's that attempted gapping in prelim test: 83161
Number of HSP's gapped (non-prelim): 86
length of query: 342
length of database: 848,049,833
effective HSP length: 128
effective length of query: 214
effective length of database: 490,808,617
effective search space: 105033044038
effective search space used: 105033044038
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.8 bits)
S2: 75 (33.5 bits)
Lotus: description of TM0176a.2


