

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
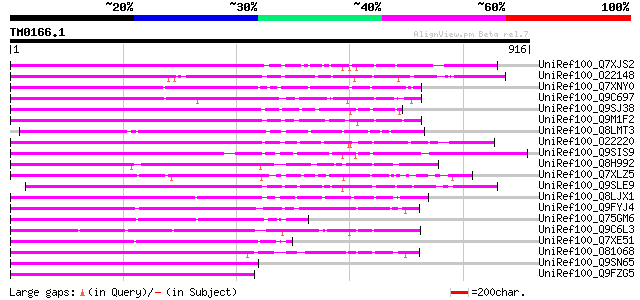
Query= TM0166.1
(916 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q7XJS2 Putative non-LTR retroelement reverse transcrip... 339 3e-91
UniRef100_O22148 Putative non-LTR retroelement reverse transcrip... 338 6e-91
UniRef100_Q7XNY0 OSJNBb0015N08.11 protein [Oryza sativa] 333 1e-89
UniRef100_Q9C697 Reverse transcriptase, putative; 16838-20266 [A... 317 8e-85
UniRef100_Q9SJ38 Putative non-LTR retroelement reverse transcrip... 312 3e-83
UniRef100_Q9M1F2 Hypothetical protein F9K21.130 [Arabidopsis tha... 298 4e-79
UniRef100_Q8LMT3 Putative retroelement [Oryza sativa] 298 4e-79
UniRef100_O22220 Putative non-LTR retroelement reverse transcrip... 298 7e-79
UniRef100_Q9SIS9 Putative non-LTR retroelement reverse transcrip... 298 7e-79
UniRef100_Q8H992 Oryza sativa (japonica cultivar-group) retrotra... 294 7e-78
UniRef100_Q7XLZ5 OSJNBa0086O06.14 protein [Oryza sativa] 290 2e-76
UniRef100_Q9SLE9 Putative non-LTR retroelement reverse transcrip... 280 1e-73
UniRef100_Q8LJX1 Putative reverse transcriptase [Sorghum bicolor] 280 2e-73
UniRef100_Q9FYJ4 F17F8.5 [Arabidopsis thaliana] 278 7e-73
UniRef100_Q75GM6 Putative non-LTR retroelement reverse transcrip... 273 2e-71
UniRef100_Q9C6L3 Hypothetical protein F2J7.11 [Arabidopsis thali... 271 5e-71
UniRef100_Q7XE51 Putative non-LTR retroelement reverse transcrip... 268 7e-70
UniRef100_O81068 Putative non-LTR retroelement reverse transcrip... 267 9e-70
UniRef100_Q9SN65 Hypothetical protein F25I24.40 [Arabidopsis tha... 266 2e-69
UniRef100_Q9FZG5 T2E6.4 [Arabidopsis thaliana] 266 2e-69
>UniRef100_Q7XJS2 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 1344
Score = 339 bits (869), Expect = 3e-91
Identities = 262/883 (29%), Positives = 420/883 (46%), Gaps = 67/883 (7%)
Query: 1 PGPDGFNFTFFREFWHLIKEEIMQMFEEFYEGGKLVKGLNAGFIALIPKRQTPEMVTDYR 60
PGPDGF FF++ W L+K +I+ F+E G L + N I LIPK +P+ ++D R
Sbjct: 424 PGPDGFTALFFQQHWDLVKHQILTEIFGFFETGVLPQDWNHTHICLIPKITSPQRMSDLR 483
Query: 61 LISLIGSIYKLLAKVLAARLQGVMPYLLSENQFSFTKGRQIANCLLIGNEVVDTMLSSGR 120
ISL +YK+++K+L RL+ +P ++S Q +F R I++ +L+ +E++ ++ ++ R
Sbjct: 484 PISLCSVLYKIISKILTQRLKKHLPAIVSTTQSAFVPQRLISDNILVAHEMIHSLRTNDR 543
Query: 121 ---GGFLLKLDFAKAYDNIEWDFLMNMMEDLGFGPRWRCWIHSCVSTATLAVLVNGSPTE 177
K D +KAYD +EW FL MM LGF +W WI +CV++ + +VL+NG P
Sbjct: 544 ISKEHMAFKTDMSKAYDRVEWPFLETMMTALGFNNKWISWIMNCVTSVSYSVLINGQPYG 603
Query: 178 FFEIEKGLRQGDPLSPLLFNLCVQGLSVMWNRLAIRERPCGVVL-GPGVCLNHLQFADDT 236
+G+RQGDPLSP LF LC + L + N+ + G+ V +NHL FADDT
Sbjct: 604 HIIPTRGIRQGDPLSPALFVLCTEALIHILNKAEQAGKITGIQFQDKKVSVNHLLFADDT 663
Query: 237 LVFCDPECAQMEELFDVMTAFLWGSGLKLNYAKSELV-GCNVDSAIVNQLAVELGVVVGT 295
L+ C + EEL ++ + SG +N KS + G NVD I + + G+ +
Sbjct: 664 LLMCKATKQECEELMQCLSQYGQLSGQMINLNKSAITFGKNVDIQIKDWIKSRSGISLEG 723
Query: 296 LPLPYLGSVLGGDPRRRGFWAPVIEKVRVALAS*GSNSVSLSGRLVVLRSVASAIPIYHM 355
YLG +R + + EK++ L + ++S G+ V+L+S+A A+P+Y M
Sbjct: 724 GTGKYLGLPECLSGSKRDLFGFIKEKLQSRLTGWYAKTLSQGGKEVLLKSIALALPVYVM 783
Query: 356 AVYKAPLGVVADIEKLMRCFLWGRSGGGRRIAWVAWEQVCKRKEIGGLGVEFLKFRNKAM 415
+ +K P + + +M F W R+I W++W+++ K+ GG G + L+ N+A+
Sbjct: 784 SCFKLPKNLCQKLTTVMMDFWWNSMQQKRKIHWLSWQRLTLPKDQGGFGFKDLQCFNQAL 843
Query: 416 LLKWAWRFGQEPEALWRRVLCAKYGLNSIPLVLHRVLEGLRHPSRYMFDIVNVVLEDSVL 475
L K AWR QE +L+ RV ++Y NS L R SR + +++ +L
Sbjct: 844 LAKQAWRVLQEKGSLFSRVFQSRYFSNSDFLSATR-------GSRPSYAWRSILFGRELL 896
Query: 476 GKTFRDQCFCVVGDGLRISFWRDPWV--ASKTLAVLYPRFYAMASNQDIMVSEAGVFSNL 533
+ R V+G+G + W D W+ S + RF N D+ VS+ +
Sbjct: 897 MQGLR----TVIGNGQKTFVWTDKWLHDGSNRRPLNRRRFI----NVDLKVSQ--LIDPT 946
Query: 534 GWEWNLGFRRRFFGWEEQVYDELIARCNGVVPSREKDSLIWLGDTNGRYTVKT----FCM 589
WNL R F W++ E+I + + ++DS WL NG Y+VKT
Sbjct: 947 SRNWNLNMLRDLFPWKDV---EIILKQRPLF--FKEDSFCWLHSHNGLYSVKTGYEFLSK 1001
Query: 590 EVELQLYG----PPSWVVPSCVKKL----APQKVVLFLWQAAENRVAVMVNLHNRGVLVD 641
+V +LY PS V S K+ K+ +FLW+A + V L RG+ D
Sbjct: 1002 QVHHRLYQEAKVKPS--VNSLFDKIWNLHTAPKIRIFLWKALHGAIPVEDRLRTRGIRSD 1059
Query: 642 AEAVCVLCGQAEESVTHLILVCKFAWVLWSKVLLREGVVWCVPASILSLLKEWDSLRKGS 701
C++C E++ H++ C A +W+ L S+ + + L + +
Sbjct: 1060 DG--CLMCDTENETINHILFECPLARQVWAITHLSSAGSE-FSNSVYTNMSRLIDLTQQN 1116
Query: 702 DLV-LWRLIPYALAWSVWMARNGLIFKGEEVSVETVWDIHITRMYWWIKAGKPHCSFSLC 760
DL R + + W +W RN L+F+G+ T+ D + W
Sbjct: 1117 DLPHHLRFVSPWILWFLWKNRNALLFEGKGSITTTLVDKAYEAYHEW------------- 1163
Query: 761 DFLQNFGAVTVGQRVERCRQDVWTCPP-EGILKWNVDGAARGAPGMGGIGGVVRDATGEI 819
F A T Q E+ + CPP G LK N+ A G VVRD+ G++
Sbjct: 1164 -----FSAQTHMQNDEKHLKITKWCPPLPGELKCNIGFAWSKQHHFSGASWVVRDSQGKV 1218
Query: 820 RGFFSMSLGVVFA-YEAEVKAIRTALEFSKEFGLSRVVVESDS 861
S V + Y A++++ ALE RV+ S +
Sbjct: 1219 LLHSRRSFNEVHSPYSAKIRSWEWALESMTHHHFDRVIFASST 1261
>UniRef100_O22148 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 1374
Score = 338 bits (866), Expect = 6e-91
Identities = 268/911 (29%), Positives = 424/911 (46%), Gaps = 84/911 (9%)
Query: 1 PGPDGFNFTFFREFWHLIKEEIMQMFEEFYEGGKLVKGLNAGFIALIPKRQTPEMVTDYR 60
PGPDG N +++FW + ++I +M + F+ G + +G+N I LIPK E +TD+R
Sbjct: 446 PGPDGMNGFLYQQFWETMGDQITEMVQAFFRSGSIEEGMNKTNICLIPKILKAEKMTDFR 505
Query: 61 LISLIGSIYKLLAKVLAARLQGVMPYLLSENQFSFTKGRQIANCLLIGNEVVDTMLSSGR 120
ISL IYK++ K++A RL+ ++P L+SE Q +F KGR I++ +LI +E++ + S+ +
Sbjct: 506 PISLCNVIYKVIGKLMANRLKKILPSLISETQAAFVKGRLISDNILIAHELLHALSSNNK 565
Query: 121 GG---FLLKLDFAKAYDNIEWDFLMNMMEDLGFGPRWRCWIHSCVSTATLAVLVNGSPTE 177
+K D +KAYD +EW FL M LGF W I CV + VL+NG+P
Sbjct: 566 CSEEFIAIKTDISKAYDRVEWPFLEKAMRGLGFADHWIRLIMECVKSVRYQVLINGTPHG 625
Query: 178 FFEIEKGLRQGDPLSPLLFNLCVQGLSVMWNRLAIRERPCGVVLGPGV-CLNHLQFADDT 236
+GLRQGDPLSP LF +C + L M + + G+ + G ++HL FADD+
Sbjct: 626 EIIPSRGLRQGDPLSPYLFVICTEMLVKMLQSAEQKNQITGLKVARGAPPISHLLFADDS 685
Query: 237 LVFCDPECAQMEELFDVMTAFLWGSGLKLNYAKSEL-VGCNVDS----AIVNQLAVEL-- 289
+ +C + ++ ++ + SG ++NY KS + G ++ + +L +E
Sbjct: 686 MFYCKVNDEALGQIIRIIEEYSLASGQRVNYLKSSIYFGKHISEERRCLVKRKLGIEREG 745
Query: 290 --GVVVGTLPLPYLGSVLGGDPRRRGFWAPVIEKVRVALAS*GSNSVSLSGRLVVLRSVA 347
GV +G LP + GS + + + +++ + SN +S G+ ++L++VA
Sbjct: 746 GEGVYLG-LPESFQGS-------KVATLSYLKDRLGKKVLGWQSNFLSPGGKEILLKAVA 797
Query: 348 SAIPIYHMAVYKAPLGVVADIEKLMRCFLWGRSGGGRRIAWVAWEQVCKRKEIGGLGVEF 407
A+P Y M+ +K P + IE +M F W GR + W AW + + K +GGLG +
Sbjct: 798 MALPTYTMSCFKIPKTICQQIESVMAEFWWKNKKEGRGLHWKAWCHLSRPKAVGGLGFKE 857
Query: 408 LKFRNKAMLLKWAWRFGQEPEALWRRVLCAKYGLNSIPLVLHRVLEGLRHPSRYMFDIVN 467
++ N A+L K WR E ++L +V ++Y S PL SR F +
Sbjct: 858 IEAFNIALLGKQLWRMITEKDSLMAKVFKSRYFSKSDPL-------NAPLGSRPSFAWKS 910
Query: 468 VVLEDSVLGKTFRDQCFCVVGDGLRISFWRDPWVASK--TLAVLYPRFYAM---ASNQDI 522
+ ++ + R V+G+G I+ W DPW+ +K A R + + A+N
Sbjct: 911 IYEAQVLIKQGIR----AVIGNGETINVWTDPWIGAKPAKAAQAVKRSHLVSQYAANSIH 966
Query: 523 MVSEAGVFSNLGWEWNLGFRRRFFGWEEQVYDELIARCNGVVPSREKDSLIWLGDTNGRY 582
+V + + W WNL + + + ++A G +R D W +G Y
Sbjct: 967 VVKDLLLPDGRDWNWNL----VSLLFPDNTQENILALRPGGKETR--DRFTWEYSRSGHY 1020
Query: 583 TVKT---FCMEVELQLYGPPSWVVPSC------VKKL-APQKVVLFLWQAAENRVAVMVN 632
+VK+ E+ Q P + PS + KL P K+ FLW+ N ++V N
Sbjct: 1021 SVKSGYWVMTEIINQRNNPQEVLQPSLDPIFQQIWKLDVPPKIHHFLWRCVNNCLSVASN 1080
Query: 633 LHNRGVLVDAEAVCVLCGQAEESVTHLILVCKFAWVLW--SKVLLREGVVWCVPA----- 685
L R + E CV C E+V HL+ C FA + W S + G W
Sbjct: 1081 LAYRHLA--REKSCVRCPSHGETVNHLLFKCPFARLTWAISPLPAPPGGEWAESLFRNMH 1138
Query: 686 SILSLLKEWDSLRKGSDLVLWRLIPYALAWSVWMARNGLIFKGEEVSVETVWDIHITRMY 745
+LS+ K L+ W + W +W RN L+FKG E + V M
Sbjct: 1139 HVLSVHKSQPEESDHHALIPW------ILWRLWKNRNDLVFKGREFTAPQVILKATEDMD 1192
Query: 746 WWIKAGKPHCSFSLCDFLQNFGAVTVGQRVERCRQDVWTCPPEGILKWNVDGAARGAPGM 805
W +P VT R +RC + W P G +K N DGA G
Sbjct: 1193 AWNNRKEPQ------------PQVTSSTR-DRCVK--WQPPSHGWVKCNTDGAWSKDLGN 1237
Query: 806 GGIGGVVRDATGEIRGFFSMSL-GVVFAYEAEVKAIRTALEFSKEFGLSRVVVESDSTLA 864
G+G V+R+ TG + +L E EV+A+R A+ F RV+ ESDS
Sbjct: 1238 CGVGWVLRNHTGRLLWLGLRALPSQQSVLETEVEALRWAVLSLSRFNYRRVIFESDSQYL 1297
Query: 865 VGWVSKRANRP 875
V + + P
Sbjct: 1298 VSLIQNEMDIP 1308
>UniRef100_Q7XNY0 OSJNBb0015N08.11 protein [Oryza sativa]
Length = 1026
Score = 333 bits (854), Expect = 1e-89
Identities = 231/731 (31%), Positives = 361/731 (48%), Gaps = 30/731 (4%)
Query: 1 PGPDGFNFTFFREFWHLIKEEIMQMFEEFYEGGKLVKGLNAGFIALIPKRQTPEMVTDYR 60
PGPDG F+ FW LIK +++ + +F + LN G I LIPK + + +R
Sbjct: 257 PGPDGMPGEFYIHFWDLIKHDLLDLINDFQRNSVNIDRLNFGVITLIPKTKDASQIQKFR 316
Query: 61 LISLIGSIYKLLAKVLAARLQGVMPYLLSENQFSFTKGRQIANCLLIGNEVVDTMLSSGR 120
I L+ +K++ KVL RL G M Y++S+ Q +F K R I ++I +EV++++ +
Sbjct: 317 PICLLNVSFKIITKVLMNRLNGTMAYIISKQQTAFLKNRFIMEGVVILHEVLNSIHQKKQ 376
Query: 121 GGFLLKLDFAKAYDNIEWDFLMNMMEDLGFGPRWRCWIHSCVSTATLAVLVNGSPTEFFE 180
G L K+DF KAYD + W F+ M++ GF +W WI V +AV VN FF+
Sbjct: 377 SGILFKVDFEKAYDKVNWVFIYRMLKAKGFPDQWCDWIMKVVMGGKVAVKVNDQIGSFFK 436
Query: 181 IEKGLRQGDPLSPLLFNLCVQGLSVMWNRLAIRERPCGVVLGPGVCLNHLQFADDTLVFC 240
KGLRQGDPLSPLLFNL + L+++ R G+ + LQ+ADDT+
Sbjct: 437 THKGLRQGDPLSPLLFNLAAEALTLLVQRAEENSLIEGLGTNGDNKIAILQYADDTIFLI 496
Query: 241 DPECAQMEELFDVMTAFLWGSGLKLNYAKSELVGCNVDSAIVNQLAVELGVV-VGTLPLP 299
+ + + L ++ F SGLK+N+ KSE V C ++ L + VG+LPL
Sbjct: 497 NDKLDHAKNLKYILCLFEQLSGLKINFNKSE-VFCFGEAKEKQDLYSNIFTCKVGSLPLK 555
Query: 300 YLGSVLGGDPRRRGFWAPVIEKVRVALAS*GSNSVSLSGRLVVLRSVASAIPIYHMAVYK 359
YLG + W K+ L S+ GRL++L S S++P+Y ++ Y+
Sbjct: 556 YLGIPIDQKRILNKDWKLAENKMEHKLGCWQGRLQSIGGRLILLNSTLSSVPMYMISFYR 615
Query: 360 APLGVVADIEKLMRCFLWGRSGGGRRIAWVAWEQVCKRKEIGGLGVEFLKFRNKAMLLKW 419
P GV I+ + FLW G R+ V W VC ++ GGLGV L+ NKAML KW
Sbjct: 616 LPKGVQERIDYFRKRFLWQEDQGIRKYHLVNWPLVCSPRDQGGLGVLDLEAMNKAMLGKW 675
Query: 420 AWRFGQEPEALWRRVLCAKYGLNSIPLVLHRVLEGLRHPSRYMFDIVNVVLEDSVLGKTF 479
WR E E W+ ++ AKY + PL R+ G S + + V V +D F
Sbjct: 676 IWRLENE-EGWWQEIIYAKY-CSDKPLSGLRLKAG----SSHFWQGVMEVKDD------F 723
Query: 480 RDQCFCVVGDGLRISFWRDPWVASKTLAVLYPRFYAMASNQDIMVSEAGVFSNLGWEWNL 539
C +VG+G + FW D W+ K LA+ +P Y + + I +++ + G + +
Sbjct: 724 FSFCTKIVGNGEKTLFWEDSWLGGKPLAIQFPSLYGIVITKRITIAD---LNRKGIDC-M 779
Query: 540 GFRRRFFGWEEQVYDELIARCNGV-VPSREKDSLIWLGDTNGRYTVKTFCMEVELQLYGP 598
FRR G + + + +++ G+ + KD L W +G++TV++F ++LQ
Sbjct: 780 KFRRDLHGDKLRDWRKIVNSWEGLNLVENCKDKLWWTLSKDGKFTVRSFYRALKLQQTSF 839
Query: 599 PSWVVPSCVKKLAPQKVVLFLWQAAENRVAVMVNLHNRGVLVDAEAVCVLCGQAEESVTH 658
P+ K P K+ +F+W +N++ NL RG + C C + E+V H
Sbjct: 840 PN---KKIWKFRVPLKIRIFIWFFTKNKILTKDNLLKRG-WRKGDNKCQFCDKV-ETVQH 894
Query: 659 LILVCKFAWVLWSKVLLREGVVWCVPASIL--SLLKEWDSLRKGSDLVLWRLIPYALAWS 716
L C A ++W+ + V + L S ++ D K +LV+ + A+ WS
Sbjct: 895 LFFDCPLARLIWNIIACALNVKPVLSRQDLFGSWIQSMDKFTK--NLVIVGIA--AVLWS 950
Query: 717 VWMARNGLIFK 727
+W RN F+
Sbjct: 951 IWKCRNKACFE 961
>UniRef100_Q9C697 Reverse transcriptase, putative; 16838-20266 [Arabidopsis thaliana]
Length = 1142
Score = 317 bits (813), Expect = 8e-85
Identities = 226/752 (30%), Positives = 362/752 (48%), Gaps = 62/752 (8%)
Query: 1 PGPDGFNFTFFREFWHLIKEEIMQMFEEFYEGGKLVKGLNAGFIALIPKRQTPEMVTDYR 60
PGPDG FF++ W +IK +++ + F + G K LN I LIPK + P +T+ R
Sbjct: 230 PGPDGMTALFFQKSWAIIKSDLLSLVNSFLQEGVFDKRLNTTNICLIPKTERPTRMTELR 289
Query: 61 LISLIGSIYKLLAKVLAARLQGVMPYLLSENQFSFTKGRQIANCLLIGNEVVDTML--SS 118
ISL YK+++K+L RL+ V+P L+SE Q +F GR I++ +LI E+ + SS
Sbjct: 290 PISLCNVGYKVISKILCQRLKTVLPNLISETQSAFVDGRLISDNILIAQEMFHGLRTNSS 349
Query: 119 GRGGFL-LKLDFAKAYDNIEWDFLMNMMEDLGFGPRWRCWIHSCVSTATLAVLVNGSPTE 177
+ F+ +K D +KAYD +EW+F+ ++ +GF +W WI C++T VL+NG P
Sbjct: 350 CKDKFMAIKTDMSKAYDQVEWNFIEALLRKMGFCEKWISWIMWCITTVQYKVLINGQPKG 409
Query: 178 FFEIEKGLRQGDPLSPLLFNLCVQGLSVMWNRLAIRERPCGV-VLGPGVCLNHLQFADDT 236
E+GLRQGDPLSP LF LC + L + + G+ V P ++HL FADD+
Sbjct: 410 LIIPERGLRQGDPLSPYLFILCTEVLIANIRKAERQNLITGIKVATPSPAVSHLLFADDS 469
Query: 237 LVFCDPECAQMEELFDVMTAFLWGSGLKLNYAKSELVGCNVDSAIVNQLAVELGVVVGTL 296
L FC Q + +++ + SG ++N++KS + + + + ++ +++G
Sbjct: 470 LFFCKANKEQCGIILEILKQYESVSGQQINFSKSSI---QFGHKVEDSIKADIKLILGIH 526
Query: 297 PLPYLGSVLGGDPRRRGFWAPVIEKVRVALAS*----GSNSVSLSGRLVVLRSVASAIPI 352
L +GS LG G V VR L S + +S G+ V+++SVA+ +P
Sbjct: 527 NLGGMGSYLGLPESLGGSKTKVFSFVRDRLQSRINGWSAKFLSKGGKEVMIKSVAATLPR 586
Query: 353 YHMAVYKAPLGVVADIEKLMRCFLWGRSGGGRRIAWVAWEQVCKRKEIGGLGVEFLKFRN 412
Y M+ ++ P + + + + F W +G R + W+AW+++C K GGLG + N
Sbjct: 587 YVMSCFRLPKAITSKLTSAVAKFWWSSNGDSRGMHWMAWDKLCSSKSDGGLGFRNVDDFN 646
Query: 413 KAMLLKWAWRFGQEPEALWRRVLCAKYGLNSIPLVLHRVLEGLRHPSRYMFDIVNVVLED 472
A+L K WR P++L+ +V +Y S PL + + R M ++V +
Sbjct: 647 SALLAKQLWRLITAPDSLFAKVFKGRYFRKSNPLDSIKSYSP-SYGWRSMISARSLVYKG 705
Query: 473 SVLGKTFRDQCFCVVGDGLRISFWRDPWVASKTLAVLYPRFYAMASN-QDIMVSEAGVFS 531
+ VG G IS W DPW+ ++ +PR + D + +
Sbjct: 706 LIKR----------VGSGASISVWNDPWIPAQ-----FPRPAKYGGSIVDPSLKVKSLID 750
Query: 532 NLGWEWNLGFRRRFFGWEEQVYDELIARCNGVVPSREKDSLIWLGDTNGRYTVKTFCMEV 591
+ WN+ + F E+ + N P+ E D+L W G YTVK+
Sbjct: 751 SRSNFWNIDLLKELFDPEDVPLISALPIGN---PNME-DTLGWHFTKAGNYTVKSGYHTA 806
Query: 592 ELQL------YGPPSWVVPSCVKKL-APQKVVLFLWQAAENRVAVMVNLHNRGVLVDAEA 644
L L GP + + + K+ P K+ FLWQ V V NL RG+L D
Sbjct: 807 RLDLNEGTTLIGPDLTTLKAYIWKVQCPPKLRHFLWQILSGCVPVSENLRKRGILCDKG- 865
Query: 645 VCVLCGQAEESVTHLILVCKFAWVLW--SKVLLREGVVWCVPASILSLLKEWDSLRKGSD 702
CV CG +EES+ H + C A +W S++ G+ P+ +S+ D
Sbjct: 866 -CVSCGASEESINHTLFQCHPARQIWALSQIPTAPGI---FPS---------NSIFTNLD 912
Query: 703 LVLWRL------IPYA-LAWSVWMARNGLIFK 727
+ WR+ PY + W +W ARN +F+
Sbjct: 913 HLFWRIPSGVDSAPYPWIIWYIWKARNEKVFE 944
>UniRef100_Q9SJ38 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 1229
Score = 312 bits (799), Expect = 3e-83
Identities = 219/707 (30%), Positives = 350/707 (48%), Gaps = 41/707 (5%)
Query: 1 PGPDGFNFTFFREFWHLIKEEIMQMFEEFYEGGKLVKGLNAGFIALIPKRQTPEMVTDYR 60
PGPDGF F+ +WH+I ++ + F+ + +N I LIPK P V DYR
Sbjct: 341 PGPDGFTAGFYHSYWHIISTDVGREIRLFFTSKNFPRRMNETHIRLIPKDLGPRKVADYR 400
Query: 61 LISLIGSIYKLLAKVLAARLQGVMPYLLSENQFSFTKGRQIANCLLIGNEVVDTMLSSG- 119
I+L YK++AK++ R+Q ++P L+SENQ +F GR I++ +LI +EV+ + +S
Sbjct: 401 PIALCNIFYKIVAKIMTKRMQLILPKLISENQSAFVPGRVISDNVLITHEVLHFLRTSSA 460
Query: 120 --RGGFLLKLDFAKAYDNIEWDFLMNMMEDLGFGPRWRCWIHSCVSTATLAVLVNGSPTE 177
+K D +KAYD +EWDFL +++ GF W W+ CV++ + + L+NG+P
Sbjct: 461 KKHCSMAVKTDMSKAYDRVEWDFLKKVLQRFGFHSIWIDWVLECVTSVSYSFLINGTPQG 520
Query: 178 FFEIEKGLRQGDPLSPLLFNLCVQGLSVMWNRL-AIRERPCGVVLGPGVCLNHLQFADDT 236
+GLRQGDPLSP LF LC + LS + R +R+ P V G +NHL FADDT
Sbjct: 521 KVVPTRGLRQGDPLSPCLFILCTEVLSGLCTRAQRLRQLPGVRVSINGPRVNHLLFADDT 580
Query: 237 LVFCDPECAQMEELFDVMTAFLWGSGLKLNYAKSELV-GCNVDSAIVNQLAVELGVVVGT 295
+ F + +L ++++ + SG +N+ KS + ++ Q+ L +
Sbjct: 581 MFFSKSDPESCNKLSEILSRYGKASGQSINFHKSSVTFSSKTPRSVKGQVKRILKIRKEG 640
Query: 296 LPLPYLGSVLGGDPRRRGFWAPVIEKVRVALAS*GSNSVSLSGRLVVLRSVASAIPIYHM 355
YLG R+R + +I+K+R S S +S +G+ V+L++V +++P+Y M
Sbjct: 641 GTGKYLGLPEHFGRRKRDIFGAIIDKIRQKSHSWASRFLSQAGKQVMLKAVLASMPLYSM 700
Query: 356 AVYKAPLGVVADIEKLMRCFLWGRSGGGRRIAWVAWEQVCKRKEIGGLGVEFLKFRNKAM 415
+ +K P + I+ L+ F W R+ +WVAW ++ K GGLG ++ N ++
Sbjct: 701 SCFKLPSALCRKIQSLLTRFWWDTKPDVRKTSWVAWSKLTNPKNAGGLGFRDIERCNDSL 760
Query: 416 LLKWAWRFGQEPEALWRRVLCAKYGLNSIPLVLHRVLEGLRHPSRYMFDIVNVVLEDSVL 475
L K WR PE+L R+L KY +S + PS+ +++ +L
Sbjct: 761 LAKLGWRLLNSPESLLSRILLGKYCHSS-------SFMECKLPSQPSHGWRSIIAGREIL 813
Query: 476 GKTFRDQCFCVVGDGLRISFWRDPWVA-SKTLAVLYPRFYAMASNQDIMVSEAGVFSNLG 534
++ ++ +G ++S W DPW++ SK L + P A+ +QD+ VS + L
Sbjct: 814 ----KEGLGWLITNGEKVSIWNDPWLSISKPLVPIGP---ALREHQDLRVSALINQNTLQ 866
Query: 535 WEWNLGFRRRFFGWEEQVYDELIARCNGVVPSREKDSLIWLGDTNGRYTVKTFCMEVELQ 594
W+WN Y+ LI + SR D L WL +G+YT ++ +
Sbjct: 867 WDWNK------IAVILPNYENLIKQL-PAPSSRGVDKLAWLPVKSGQYTSRSGYGIASVA 919
Query: 595 LYGPP----SWVVPSCVKKLAPQKVVLFLWQAAENRVAVMVNLHNRGVLVDAEAVCVLCG 650
P +W + P K+ +W+AA + V + L R + A C CG
Sbjct: 920 SIPIPQTQFNWQSNLWKLQTLP-KIKHLMWKAAMEALPVGIQLVRRH--ISPSAACHRCG 976
Query: 651 QAEESVTHLILVCKFAWVLWSKVLLREGVVWCVPAS----ILSLLKE 693
A ES THL C+FA +W L+E V P S LSLLK+
Sbjct: 977 -APESTTHLFFHCEFAAQVWELAPLQETTV--PPGSSMLDALSLLKK 1020
>UniRef100_Q9M1F2 Hypothetical protein F9K21.130 [Arabidopsis thaliana]
Length = 851
Score = 298 bits (764), Expect = 4e-79
Identities = 221/752 (29%), Positives = 369/752 (48%), Gaps = 58/752 (7%)
Query: 1 PGPDGFNFTFFREFWHLIKEEIMQMFEEFYEGGKLVKGLNAGFIALIPKRQTPEMVTDYR 60
PGPDG F+++ W ++ ++++ + F+E + +N I +IPK Q P+ ++DYR
Sbjct: 67 PGPDGLTARFYKQCWDIVGNDVIKEVKLFFESSHMKTSVNHTNICMIPKIQNPQTLSDYR 126
Query: 61 LISLIGSIYKLLAKVLAARLQGVMPYLLSENQFSFTKGRQIANCLLIGNEVVDTMLSSGR 120
I+L +YK+++K + RL+ + ++S++Q +F GR I + ++I +E++ ++ R
Sbjct: 127 PIALCNVLYKVISKCMVNRLKAHLNSIVSDSQAAFIPGRIINDNVMIAHEIMHSLKVRKR 186
Query: 121 GG---FLLKLDFAKAYDNIEWDFLMNMMEDLGFGPRWRCWIHSCVSTATLAVLVNGSPTE 177
+K D +KAYD +EWDFL M GF +W WI + V + +VL+NGSP
Sbjct: 187 VSKTYMAVKTDVSKAYDRVEWDFLETTMRLFGFCDKWIGWIMAAVKSVHYSVLINGSPHG 246
Query: 178 FFEIEKGLRQGDPLSPLLFNLCVQGLSVMWNRLAIRERPCGVVLGPGV-CLNHLQFADDT 236
+ +G+RQGDPLSP LF LC LS + A GV +G G + HLQFADD+
Sbjct: 247 YISPTRGIRQGDPLSPYLFILCGDILSHLIKVKASSGDIRGVRIGNGAPAITHLQFADDS 306
Query: 237 LVFCDPECAQMEELFDVMTAFLWGSGLKLNYAKSELV-GCNVDSAIVNQLAVELGVVVGT 295
L FC + L DV + + SG K+N KS + G V + +L L +
Sbjct: 307 LFFCQANVRNCQALKDVFDVYEYYSGQKINVQKSLITFGSRVYGSTQTRLKTLLNIPNQG 366
Query: 296 LPLPYLGSVLGGDPRRRGFWAPVIEKVRVALAS*GSNSVSLSGRLVVLRSVASAIPIYHM 355
YLG +++ + +I++V+ AS + +S +G+ ++L+SVA A+P+Y M
Sbjct: 367 GGGKYLGLPEQFGRKKKEMFNYIIDRVKERTASWSAKFLSPAGKEILLKSVALAMPVYAM 426
Query: 356 AVYKAPLGVVADIEKLMRCFLWGRSGGGRRIAWVAWEQVCKRKEIGGLGVEFLKFRNKAM 415
+ +K P G+V++IE L+ F W ++ R I WVAW+++ K+ GGLG L N A+
Sbjct: 427 SCFKLPQGIVSEIESLLMNFWWEKASNKRGIPWVAWKRLQYSKKEGGLGFRDLAKFNDAL 486
Query: 416 LLKWAWRFGQEPEALWRRVLCAKYGLNSIPLVLHRVLEGLRHPSRYMFDIVNVVLEDSVL 475
L K AWR Q P +L+ RV+ A+Y ++ + + S+ + +++ ++L
Sbjct: 487 LAKQAWRIIQYPNSLFARVMKARYFKDNSII-------DAKTRSQQSYGWSSLLSGIALL 539
Query: 476 GKTFRDQCFCVVGDGLRISFWRDPWVASKTLAVLYPRFYAMASNQDIMVSEAGVFSNLGW 535
K R V+GDG I D V S +P + Q +S +F + G
Sbjct: 540 RKGTR----YVIGDGKTIRLGIDNVVDS------HPPRPLLTDEQHNGLSLDNLFQHRGH 589
Query: 536 E--WNLGFRRRFFGWEEQVYDELIARCNGVVPSREK-DSLIWLGDTNGRYTVKT-FCMEV 591
W+ + F + Y + I + +R K D LIW ++ G YTV++ + +
Sbjct: 590 SRCWDNAKLQTFVDQSDHDYIKRI-----YLSTRSKTDRLIWSYNSTGDYTVRSGYWLST 644
Query: 592 ELQLYGPPSWVVPSCVKKLAP-------------QKVVLFLWQAAENRVAVMVNLHNRGV 638
PS +P+ K K+ FLW+ + L RG+
Sbjct: 645 H-----DPSNTIPTMAKPHGSVDLKTKIWNLPIMPKLKHFLWRILSKALPTTDRLTTRGM 699
Query: 639 LVDAEAVCVLCGQAEESVTHLILVCKFAWVLW---SKVLLREGVVWC-VPASILSLLKEW 694
+D C C + ES+ H + C FA + W L R ++ + +I ++L
Sbjct: 700 RIDPG--CPRCRRENESINHALFTCPFATMAWRLSDTPLYRSSILSNNIEDNISNILL-- 755
Query: 695 DSLRKGSDLVLWRLIPYALAWSVWMARNGLIF 726
L+ + +LIP+ L W +W ARN ++F
Sbjct: 756 -LLQNTTITDSQKLIPFWLLWRIWKARNNVVF 786
>UniRef100_Q8LMT3 Putative retroelement [Oryza sativa]
Length = 764
Score = 298 bits (764), Expect = 4e-79
Identities = 210/716 (29%), Positives = 335/716 (46%), Gaps = 29/716 (4%)
Query: 18 IKEEIMQMFEEFYEGGKLVKGLNAGFIALIPKRQTPEMVTDYRLISLIGSIYKLLAKVLA 77
+KE++ +M E YEG ++ LN G I LIPK + + +R I L+ +K L K+L
Sbjct: 10 LKEQLREMVELLYEGKLDMRRLNYGVITLIPKVKDANTIKSFRPICLLNVCFKFLTKILT 69
Query: 78 ARLQGVMPYLLSENQFSFTKGRQIANCLLIGNEVVDTMLSSGRGGFLLKLDFAKAYDNIE 137
RL + ++ E+Q +F GRQ + ++I +EV+ + + G + KLDF KAYD ++
Sbjct: 70 RRLSEIAQKVIGESQTAFIPGRQNLDGVVILHEVLHELKKEKKSGIIFKLDFEKAYDKVQ 129
Query: 138 WDFLMNMMEDLGFGPRWRCWIHSCVSTATLAVLVNGSPTEFFEIEKGLRQGDPLSPLLFN 197
W FL +++ GF RW W+ +AV +NG +FF+ +G+RQGDPLSPLLFN
Sbjct: 130 WSFLFDVLHKKGFSDRWIQWVKMATIGGKMAVNINGEVKDFFKTYRGVRQGDPLSPLLFN 189
Query: 198 LCVQGLSVMWNRLAIRERPCGVVLGPGVCLNHLQFADDTLVFCDPECAQMEELFDVMTAF 257
L LS M N + L PG L HLQ+ADDT++F + + ++ +
Sbjct: 190 LVADALSEMLN----NAKQAVPHLVPG-GLTHLQYADDTILFMTNTEENIVTVKFLLYCY 244
Query: 258 LWGSGLKLNYAKSELVGCNVDSAIVNQLAVELGVVVGTLPLPYLGSVLGGDPRRRGFWAP 317
SGLK+NY KSE++ D ++A G +P YLG + +
Sbjct: 245 EAMSGLKINYQKSEIMVIGGDEMETQRVADLFNCQAGKMPFTYLGIPISMNKLTNADLDI 304
Query: 318 VIEKVRVALAS*GSNSVSLSGRLVVLRSVASAIPIYHMAVYKAPLGVVADIEKLMRCFLW 377
K+ LA+ +S G+ +++ S S+IP+Y M VY P GV ++ + F W
Sbjct: 305 PPNKIEKRLATWKCGYLSYGGKAILINSCLSSIPLYMMGVYLLPEGVHNKMDSIRARFFW 364
Query: 378 GRSGGGRRIAWVAWEQVCKRKEIGGLGVEFLKFRNKAMLLKWAWRFGQEPEALWRRVLCA 437
R+ + WE +C+ KE GGLG + N A+L KW +R E +L
Sbjct: 365 EGLEKKRKYHMIKWEALCRPKEFGGLGFIDTRKMNIALLCKWIYRLESGKEDPCCVLLRN 424
Query: 438 KYGLNSIPLVLHRVLEGLRHPSRYMFDIVNVVLEDSVLGKTFRDQCFCVVGDGLRISFWR 497
KY + + E S + ++ V + LG +++ VG+G +FW
Sbjct: 425 KYMKDGGGFFQSKAEE-----SSQFWKGLHEVKKWMDLGSSYK------VGNGKATNFWS 473
Query: 498 DPWVASKTLAVLYPRFYAMASNQDIMVSEAGVFSNLGWEWNLGFRRRFFGWEEQVYDELI 557
D W+ L YP Y M ++++ VS+ L +W + RR + +++L
Sbjct: 474 DVWIGETPLKTQYPNIYRMCADKEKTVSQ----MCLEGDWYIELRRSLGERDLNEWNDLH 529
Query: 558 ARCNGVVPSREKDSLIWLGDTNGRYTVKTFCMEVELQLYGPPSWVVPSCVKKLAPQKVVL 617
+ E+D +IW NG Y KT + L G V+ + P KV +
Sbjct: 530 NTPREIHLKEERDCIIWKLTKNGFYKAKT--LYQALSFGGVKDKVMQDLWRSSIPLKVKI 587
Query: 618 FLWQAAENRVAVMVNLHNRGVLVDAEAVCVLCGQAEESVTHLILVCKFAWVLWSKVLLRE 677
W + R+ L + + C LCGQ E+ V HL+ C + +W R+
Sbjct: 588 LFWLMLKGRIQAAGQL--KKMKWSGSPNCKLCGQTED-VDHLMFRCHISQFMW--CCYRD 642
Query: 678 GVVW-CVPASILSLLKEWDSLRKGSDLVLWRLIPYALAWSVWMARNGLIFKGEEVS 732
W +P+S LL++ + + G VL + A W++W+ RN +F G+ S
Sbjct: 643 AFGWDNIPSSREDLLEKLNIQKDGQKSVLLACLA-AGTWAIWLMRNDWVFNGKLTS 697
>UniRef100_O22220 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 1094
Score = 298 bits (762), Expect = 7e-79
Identities = 237/881 (26%), Positives = 403/881 (44%), Gaps = 76/881 (8%)
Query: 1 PGPDGFNFTFFREFWHLIKEEIMQMFEEFYEGGKLVKGLNAGFIALIPKRQTPEMVTDYR 60
PGPDGF FF+ W L+K +I+ E F++ G L + N + LIPK P + D R
Sbjct: 175 PGPDGFTALFFQRQWPLVKNQIISDIELFFQTGILPEDWNHTHLCLIPKITKPARMADIR 234
Query: 61 LISLIGSIYKLLAKVLAARLQGVMPYLLSENQFSFTKGRQIANCLLIGNEVVDTMLSS-- 118
ISL +YK+++K+L+ARL+ +P ++S Q +F R +++ +++ +E+V + ++
Sbjct: 235 PISLCSVMYKIISKILSARLKKYLPVIVSPTQSAFVAERLVSDNIILAHEIVHNLRTNEK 294
Query: 119 -GRGGFLLKLDFAKAYDNIEWDFLMNMMEDLGFGPRWRCWIHSCVSTATLAVLVNGSPTE 177
+ + K D +KAYD +EW FL ++ LGF W W+ +CVS+ + +VL+NG P
Sbjct: 295 ISKDFMVFKTDMSKAYDRVEWPFLKGILLALGFNSTWINWMMACVSSVSYSVLINGQPFG 354
Query: 178 FFEIEKGLRQGDPLSPLLFNLCVQGLSVMWNRLAIRERPCGVVL-GPGVCLNHLQFADDT 236
+GLRQGDPLSP LF LC + L + N+ + G+ G G +NHL FADDT
Sbjct: 355 HITPHRGLRQGDPLSPFLFVLCTEALIHILNQAEKIGKISGIQFNGTGPSVNHLLFADDT 414
Query: 237 LVFCDPECAQMEELFDVMTAFLWGSGLKLNYAKSELV-GCNVDSAIVNQLAVELGVVVGT 295
L+ C + E+ ++ + SG +N KS + G V+ + G+
Sbjct: 415 LLICKASQLECAEIMHCLSQYGHISGQMINSEKSAITFGAKVNEETKQWIMNRSGIQTEG 474
Query: 296 LPLPYLGSVLGGDPRRRGFWAPVIEKVRVALAS*GSNSVSLSGRLVVLRSVASAIPIYHM 355
YLG ++ + + EK++ L+ + ++S G+ ++L+S+A A P+Y M
Sbjct: 475 GTGKYLGLPECFQGSKQVLFGFIKEKLQSRLSGWYAKTLSQGGKDILLKSIAMAFPVYAM 534
Query: 356 AVYKAPLGVVADIEKLMRCFLWGRSGGGRRIAWVAWEQVCKRKEIGGLGVEFLKFRNKAM 415
++ + + +M F W ++I W+ +++ K +GG G + L+ N+A+
Sbjct: 535 TCFRLSKTLCTKLTSVMMDFWWNSVQDKKKIHWIGAQKLMLPKFLGGFGFKDLQCFNQAL 594
Query: 416 LLKWAWRFGQEPEALWRRVLCAKYGLNSIPLVLHRVLEGLRHPSRYMFDIVNVVLEDSVL 475
L K A R + ++L ++L ++Y +NS L +G R + + L S L
Sbjct: 595 LAKQASRLHTDSDSLLSQILKSRYYMNSDFL---SATKGTRPSYAWQSILYGRELLVSGL 651
Query: 476 GKTFRDQCFCVVGDGLRISFWRDPWVASKTLAVLYPRFYAMASNQDIMVSEAGVFSNLGW 535
K ++G+G W D W+ R ++ DI + + +
Sbjct: 652 KK--------IIGNGENTYVWMDNWIFDDKPR----RPESLQIMVDIQLKVSQLIDPFSR 699
Query: 536 EWNLGFRRRFFGWEEQVYDELIARCNGVVPSREKDSLIWLGDTNGRYTVKT---FC-MEV 591
WNL R F W+ E+ C + +DS W G +G YTVK+ C +V
Sbjct: 700 NWNLNMLRDLFPWK-----EIQIICQQRPMASRQDSFCWFGTNHGLYTVKSEYDLCSRQV 754
Query: 592 ELQLY----GPPS--------WVVPSCVKKLAPQKVVLFLWQAAENRVAVMVNLHNRGVL 639
Q++ PS W + S AP K+ +FLW+ + VAV L RGVL
Sbjct: 755 HKQMFKEAEEQPSLNPLFGKIWNLNS-----AP-KIKVFLWKVLKGAVAVEDRLRTRGVL 808
Query: 640 VDAEAVCVLCGQAEESVTHLILVCKFAWVLWSKVLLR---EGVVWCVPASILSLLKEWDS 696
+ E C +C + E++ H++ C A +W+ ++ G + ++ ++ +
Sbjct: 809 I--EDGCSMCPEKNETLNHILFQCPLARQVWALTPMQSPNHGFGDSIFTNVNHVIGNCHN 866
Query: 697 LRKGSDLVLWRLIPYALAWSVWMARNGLIFKG-EEVSVETVWDIHITRMYWWIKAGKPHC 755
L R + + W +W RN +F+G VS+ V + W+KA + C
Sbjct: 867 TELSPHL---RYVSPWIIWILWKNRNKRLFEGIGSVSLSIVGKA-LEDCKEWLKAHELIC 922
Query: 756 SFSLCDFLQNFGAVTVGQRVERCRQDVWTCPPEGILKWNVDGAARGAPGMGGIGGVVRDA 815
S E + W P LK N+ A M G+ VVR+
Sbjct: 923 S------------------KEPTKDLTWIPPLMNELKCNIGIAWSKKHQMAGVSWVVRNW 964
Query: 816 TGEIRGFFSMSLGVVFA-YEAEVKAIRTALEFSKEFGLSRV 855
G + S + + ++A++K A+E +F RV
Sbjct: 965 KGRVLLHSRCSFSQISSHFDAKIKGWNYAVESMDQFKFDRV 1005
>UniRef100_Q9SIS9 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 1715
Score = 298 bits (762), Expect = 7e-79
Identities = 256/935 (27%), Positives = 418/935 (44%), Gaps = 78/935 (8%)
Query: 1 PGPDGFNFTFFREFWHLIKEEIMQMFEEFYEGGKLVKGLNAGFIALIPKRQTPEMVTDYR 60
PG DGF F+ W LI ++ M F+E + +N I LIPK P+ ++DYR
Sbjct: 808 PGFDGFTAAFYHHLWDLIGNDVCLMVRHFFESDVMDNQINQTQICLIPKIIDPKHMSDYR 867
Query: 61 LISLIGSIYKLLAKVLAARLQGVMPYLLSENQFSFTKGRQIANCLLIGNEVVDTMLSSGR 120
ISL + YK+++K+L RL+ + ++S++Q +F G+ I++ +L+ +E++ ++ S
Sbjct: 868 PISLCTASYKIISKILIKRLKQCLGDVISDSQAAFVPGQNISDNVLVAHELLHSLKSRRE 927
Query: 121 ---GGFLLKLDFAKAYDNIEWDFLMNMMEDLGFGPRWRCWIHSCVSTATLAVLVNGSPTE 177
G +K D +KAYD +EW+FL +M LGF PRW WI +CV++ + VL+NGSP
Sbjct: 928 CQSGYVAVKTDISKAYDRVEWNFLEKVMIQLGFAPRWVKWIMTCVTSVSYEVLINGSPYG 987
Query: 178 FFEIEKGLRQGDPLSPLLFNLCVQGLSVMWNRLAIRERPCGV-VLGPGVCLNHLQFADDT 236
+G+RQGDPLSP LF C + LS M + + ++ G+ + + ++HL FADD+
Sbjct: 988 KIFPSRGIRQGDPLSPYLFLFCAEVLSNMLRKAEVNKQIHGMKITKDCLAISHLLFADDS 1047
Query: 237 LVFCDPECAQMEELFDVMTAFLWGSGLKLNYAKSELV-GCNVDSAIVNQLAVELGVVVGT 295
L FC +E+L + + SG K+NYAKS ++ G + + +L LG+
Sbjct: 1048 LFFCRASNQNIEQLALIFKKYEEASGQKINYAKSSIIFGQKIPTMRRQRLHRLLGIDNVR 1107
Query: 296 LPLPYLGSVLGGDPRRRGFWAPVIEKVRVALAS*GSNSVSLSGRLVVLRSVASAIPIYHM 355
YLG R+ + ++ KV+ N +S +G+ +V++++A A+P+Y M
Sbjct: 1108 GGGKYLGLPEQLGRRKVELFEYIVTKVKERTEGWAYNYLSPAGKEIVIKAIAMALPVYSM 1167
Query: 356 AVYKAPLGVVADIEKLMRCFLWGRSGGGRRIAWVAWEQVCKRKEIGGLGVEFLKFRNKAM 415
+ P + +I L+ F WG ++ G LG + L N+A+
Sbjct: 1168 NCFLLPTLICNEINSLITAFWWG------------------KENEGDLGFKDLHQFNRAL 1209
Query: 416 LLKWAWRFGQEPEALWRRVLCAKYGLNSIPLVLHRVLEGLRHPSRYMFDIVNVVLEDSVL 475
L K AWR P++L R+ Y N+ L R +G H S Y ++ +
Sbjct: 1210 LAKQAWRILTNPQSLLARLYKGLYYPNTTYL---RANKG-GHAS-YGWNSIQE------- 1257
Query: 476 GKTFRDQCFCV-VGDGLRISFWRDPWVASKTLAVLYPRFYAMASNQDIMVSEAGVFSNLG 534
GK Q V +GDG W DPW L L PR A D + A ++
Sbjct: 1258 GKLLLQQGLRVRLGDGQTTKIWEDPW-----LPTLPPR-PARGPILDEDMKVADLWRENK 1311
Query: 535 WEWNLGFRRRFFGWEEQVYDELIARCNGVVPSREKDSLIWLGDTNGRYTVK------TFC 588
EW+ E+Q + + N +DS W N +YTV+ T
Sbjct: 1312 REWDPVIFEGVLNPEDQQLAKSLYLSNYAA----RDSYKWAYTRNTQYTVRSGYWVATHV 1367
Query: 589 MEVELQLYGPPSWVVPSCVK----KLAPQKVVLFLWQAAENRVAVMVNLHNRGVLVDAEA 644
E ++ P VP + K+ P K+ F+W+ ++ L NR + A+
Sbjct: 1368 NLTEEEIINPLEGDVPLKQEIWRLKITP-KIKHFIWRCLSGALSTTTQLRNRN--IPADP 1424
Query: 645 VCVLCGQAEESVTHLILVCKFAWVLWSKVLLREGVVWCVPASILSLLKEWDSLRKGSDL- 703
C C A+E++ H+I C +A V+W C ++ ++ +K +L
Sbjct: 1425 TCQRCCNADETINHIIFTCSYAQVVWRSANFSGSNRLCFTDNLEENIRLILQGKKNQNLP 1484
Query: 704 VLWRLIPYALAWSVWMARNGLIFKGEEVSVETVWDIHITRMYWWIKAGKPHCSFSLCDFL 763
+L L+P+ + W +W +RN +F+ + R W + + + +
Sbjct: 1485 ILNGLMPFWIMWRLWKSRNEYLFQ------------QLDRFPWKVAQKAEQEATEWVETM 1532
Query: 764 QNFGAVT--VGQRVER--CRQDVWTCPPEGILKWNVDGAARGAPGMGGIGGVVRDATGEI 819
N A++ Q +R R W+ PPEG LK N D G ++RD G +
Sbjct: 1533 VNDTAISHNTAQSNDRPLSRSKQWSSPPEGFLKCNFDSGYVQGRDYTSTGWILRDCNGRV 1592
Query: 820 RGFFSMSLGVVF-AYEAEVKAIRTALEFSKEFGLSRVVVESDSTLAVGWVSKRANRPWKL 878
L + A +AE AL+ G V E D+ ++K + L
Sbjct: 1593 LHSGCAKLQQSYSALQAEALGFLHALQMVWIRGYCYVWFEGDNLELTNLINKTEDH-HLL 1651
Query: 879 LNDLHMIDILRNEVECLRIYHVFREINCQPDFLAK 913
L+ I ++ I +V RE N D L K
Sbjct: 1652 ETLLYDIRFWMTKLPFSSIGYVNRERNLAADKLTK 1686
>UniRef100_Q8H992 Oryza sativa (japonica cultivar-group) retrotransposon Karma DNA,
complete sequence [Oryza sativa]
Length = 1197
Score = 294 bits (753), Expect = 7e-78
Identities = 219/776 (28%), Positives = 355/776 (45%), Gaps = 75/776 (9%)
Query: 1 PGPDGFNFTFFREFWHLIKEEIMQMFEEFYEGGKLVKGLNAGFIALIPKRQTPEMVTDYR 60
PGPDGF +F+ W+ IKE+I+++ F ++ +N I LIPK V YR
Sbjct: 459 PGPDGFGPSFYNSAWNSIKEDIIKLAASFCSNSVQLERINRAHIVLIPKPGKENTVDGYR 518
Query: 61 LISLIGSIYKLLAKVLAARLQGVMPYLLSENQFSFTKGRQIANCLLIGNEVVDTMLSSGR 120
ISL K+L+KVLA RLQ V+ ++ +Q F KGR I+ L+ E++ +
Sbjct: 519 PISLQNCSVKILSKVLANRLQKVLTRMIHLDQTGFLKGRCISENLIYATELIQACHARKC 578
Query: 121 GGFLLKLDFAKAYDNIEWDFLMNMMEDLGFGPRWRCWIHSCVSTATLAVLVNGSPTEFFE 180
++KLDFAKA+D++ W L ++ GF W WI S + T+ AVL+NG P ++
Sbjct: 579 KTLIIKLDFAKAFDSVLWTSLFQILAVRGFPNNWISWIKSLLQTSKSAVLLNGIPGKWIN 638
Query: 181 IEKGLRQGDPLSPLLFNLCVQGLS-VMWNRLAIR-----ERPCGVVLGPGVCLNHLQFAD 234
+KGLRQGDPLSP LF L L ++ IR +RPC + Q+AD
Sbjct: 639 CKKGLRQGDPLSPYLFILVADVLQRLLEKNFQIRHPIYQDRPCATI----------QYAD 688
Query: 235 DTLVFCDPECAQMEELFDVMTAFLWGSGLKLNYAKSELVGCNVDSAIVNQLAVELGVVVG 294
DTLV C E + L + F +GL++N+AKS ++ ++D + + L+ L +
Sbjct: 689 DTLVICRAEEDDVLALRSTLLQFSKATGLQINFAKSTMISLHIDRSKESSLSELLQCKLE 748
Query: 295 TLPLPYLGSVLGGDPRRRGFWAPVIEKVRVALAS*GSNSVSLSGRLVVLRSVASAIPIYH 354
+LP+ YLG L P++ KV L ++ +S + RL+++ +V S++P+Y
Sbjct: 749 SLPMSYLGLPLSLHKLTNNDLQPIVVKVDSFLTGWEASLLSQAERLILVNAVLSSVPVYA 808
Query: 355 MAVYKAPLGVVADIEKLMRCFLW-GRSGGGRRIAWVAWEQVCKRKEIGGLGVEFLKFRNK 413
M+ +K P V+ I+K R F W G VAW++VC KE GGLG++ LK +N+
Sbjct: 809 MSAFKLPPKVIEAIDKRRRAFFWTGDDTCSGAKCLVAWDEVCTAKEKGGLGIKSLKTQNE 868
Query: 414 AMLLKWAWRFGQEPEALWRRVLCAKYG----LNSIPLVLHRVLEGLRHPSRYMFDIVNVV 469
A+LLK + + + W + ++ L S+PL H + P + V+
Sbjct: 869 ALLLKRLFNLFSD-NSSWTNWIWKEFDGRSLLKSLPLGQHWSVFQKLLPELFKITTVH-- 925
Query: 470 LEDSVLGKTFRDQCFCVVGDGLRISFWRDPWVASKTLAVLYPRFYAMASNQDIMVSEA-- 527
+GDG R SFW D W + TLA + ++ +++ V +
Sbjct: 926 -----------------IGDGSRTSFWHDRWTGNMTLAAQFESLFSHSTDDLATVEQILS 968
Query: 528 -GVFSNLGWEWNLGFRRRFFGWEEQVYDELIARCNGVVPSREKDSLIWLGDTNGRYTVKT 586
G++ L R + EL + E D+ L + G T KT
Sbjct: 969 FGIYDIL--------TPRLSSAAQNDLQELQTILQNFSLTNESDTR--LNNRLGNLTTKT 1018
Query: 587 FCMEVELQLYGPPSWVVPS---CVKKLAPQKVVLFLWQAAENRVAVMVNLHNRGVLVDAE 643
L PP + P+ P K+ LF W +R++ +NL + ++
Sbjct: 1019 I-----YDLRSPPGFPSPNWKFIWDCRMPLKIKLFAWLLVRDRLSTKLNLLKKKIV--QT 1071
Query: 644 AVCVLCGQAEESVTHLILVCKFAWVLWSKVLLREGVVWCVPASILSLLKEWDSLRKGSDL 703
A C +C +E+ HL C FA W + ++ I++ K L+ + +
Sbjct: 1072 ATCDICSTTDETADHLSFNCPFAISFWQALHIQ---------PIINETKYLSQLKAPATI 1122
Query: 704 VLWRLIPYALA--WSVWMARNGLIFKGEEVSVETVWDIHITRMYWWIKAGKPHCSF 757
+ + W +W R+ ++F+G S+ + I+ W + P F
Sbjct: 1123 PTRHFQSFFMLCFWVLWNHRHDVVFRGRPPSIASCLQRGISESSLWAETFTPDDRF 1178
>UniRef100_Q7XLZ5 OSJNBa0086O06.14 protein [Oryza sativa]
Length = 1205
Score = 290 bits (741), Expect = 2e-76
Identities = 252/853 (29%), Positives = 391/853 (45%), Gaps = 117/853 (13%)
Query: 1 PGPDGFNFTFFREFWHLIKEEIMQMFEEFYEGGKLVKGLNAGFIALIPKRQTPEMVTDYR 60
PGPDGF FF+ W ++K ++++ EF+E G+ +G+N I +IPK P + D+R
Sbjct: 434 PGPDGFPARFFQRNWGVLKRDVIEGVREFFETGEWKEGMNDTVIVMIPKTNAPVEMKDFR 493
Query: 61 LISLIGSIYKLLAKVLAARLQGVMPYLLSENQFSFTKGRQIANCLLIGNEVVDTMLSSGR 120
+SL IYK++AK L RL+ ++ ++SE Q +F GR I + L+ E ++ R
Sbjct: 494 PVSLCNVIYKVVAKCLVNRLRPLLQEIISETQSAFVPGRMITDNALVAFECFHSIHKCTR 553
Query: 121 GG---FLLKLDFAKAYDNIEWDFLMNMMEDLGFGPRWRCWIHSCVSTATLAVLVNGSPTE 177
LKLD +KAYD ++W FL ++ LGFG WR WI SCV++ +V +NG+ E
Sbjct: 554 ESQDFCALKLDLSKAYDRVDWGFLDGALQKLGFGNIWRKWIMSCVTSVRYSVRLNGNMLE 613
Query: 178 FFEIEKGLRQGDPLSPLLFNLCVQGLS-VMWNRLAIRE-RPCGVVLG-PGVCLNHLQFAD 234
F +GLR+GDPL+P LF GLS ++ R R+ +P V PGV +HL FAD
Sbjct: 614 PFYPTRGLREGDPLNPYLFLFIADGLSNILQRRRDERQIQPLKVCRSAPGV--SHLLFAD 671
Query: 235 DTLVFCDPECAQMEELFDVMTAFLWGSGLKLNYAKSELVGCNVDSAIVNQ-----LAVEL 289
D+L+F E Q + + + + +G +N + L+ SA+ Q + L
Sbjct: 672 DSLLFFKAEVIQATRIKEALDLYERCTGQLINPKECSLLF----SALCPQERQDGIKAVL 727
Query: 290 GVVVGTLPLPYLGSVLGGDPRRRGFWAPVIEKVRVALAS*GSNSVSLSGRLVVLRSVASA 349
V LG + + P+ E+ L +SL+G+ +++SVA A
Sbjct: 728 QVERTCFDDKCLGLPTPDGRMKAEQFQPIKERFEKRLTDWSERFLSLAGKEALIKSVAQA 787
Query: 350 IPIYHMAVYKAPLGVVADIEKLMRCFLWGRSGGGRRIAWVAWEQVCKRKEIGGLGVEFLK 409
+P Y M V+K P + E+L+R F WG G +++ W+AWE++ K +GGLG ++
Sbjct: 788 LPTYTMGVFKMPERFCEEYEQLVRNFWWGHEKGEKKVHWIAWEKLTSPKLLGGLGFRDIR 847
Query: 410 FRNKAMLLKWAWRFGQEPEALWRRVLCAKYGLN------SIPLVLHRVLEGLRHPSRYMF 463
N+A+L + AWR + P++L RVL AKY N + P V +G+ H
Sbjct: 848 CFNQALLARQAWRLIESPDSLCARVLKAKYYPNGTITDTAFPSVSSPTWKGIVHG----- 902
Query: 464 DIVNVVLEDSVLGKTFRDQCFCVVGDGLRISFWRDPWVA-SKTLAVLYPRFYAMASNQDI 522
LE G +R +GDG + WR+ WVA + L +L + + N+ I
Sbjct: 903 ------LELLKKGLIWR------IGDGSKTKIWRNHWVAHGENLKILEKKTW----NRVI 946
Query: 523 MVSEAGVFSNLGWEWNLGFRRRFFGWEEQVYDELIARCNGVVPSREKDSL-IWLGDTNGR 581
V E V W L R EE + L R +P RE++ W + G
Sbjct: 947 YVRELIVTDTKTWNEPL---IRHIIREEDADEILKIR----IPQREEEDFPAWHYEKTGI 999
Query: 582 YTVKTF-----------CMEVELQLYGPPSWVVPSCVKKLAPQ-KVVLFLWQAAENRVAV 629
++V++ + G + V K Q KV +F W+ A++R+A
Sbjct: 1000 FSVRSVYRLAWNLARKTSEQASSSSGGADGRKIWDNVWKANVQPKVRVFAWKLAQDRLAT 1059
Query: 630 MVNLHNRGVLVDAEAVCVLCGQAEESVTHLILVCKFAWVLWSKVLLREGVVWCVP-ASIL 688
N R ++ C +CGQ EE+ H + C A L + LRE W +P S+
Sbjct: 1060 WENKKKR--KIEMFGTCPICGQKEETGFHATVECTLAKAL--RASLREH--WTLPDESLF 1113
Query: 689 SLLKEWDSLRKGSDLVLWRLIPYALAWSVWMARNGLIFKGEEVSVETVWDIHITRMYWWI 748
S+ G D +L V + R K + + + + DI
Sbjct: 1114 SM--------TGPDWLL-----------VLLDRLSSEKKAQLPARKHMEDI--------- 1145
Query: 749 KAGKPHCSFSLCDFLQNFGAVTVGQRVERCR----QDVWTCPPEGILKWNVDGAARGAPG 804
GK C Q+ + C+ ++ W+CPP+G K NVD A R G
Sbjct: 1146 -KGKGPMFQDPC------------QKEQTCQLNAEKEKWSCPPDGSAKLNVDAAYRTETG 1192
Query: 805 MGGIGGVVRDATG 817
G ++RD G
Sbjct: 1193 EASAGIIIRDCRG 1205
>UniRef100_Q9SLE9 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 1319
Score = 280 bits (717), Expect = 1e-73
Identities = 221/851 (25%), Positives = 400/851 (46%), Gaps = 60/851 (7%)
Query: 29 FYEGGKLVKGLNAGFIALIPKRQTPEMVTDYRLISLIGSIYKLLAKVLAARLQGVMPYLL 88
F+E G L + N + LIPK P+ ++D R ISL +YK+++K+L+ +L+ +P ++
Sbjct: 426 FFETGVLPQEWNHTHLYLIPKFTNPQRMSDIRPISLCSVLYKIISKILSFKLKKHLPSIV 485
Query: 89 SENQFSFTKGRQIANCLLIGNEVVDTMLSSGRGG---FLLKLDFAKAYDNIEWDFLMNMM 145
S +Q +F R I++ +LI +E+V ++ ++ + + K D +KAYD +EW FL ++
Sbjct: 486 SPSQSAFFAERLISDNILIAHEIVHSLRTNDKISKEFMVFKTDMSKAYDRVEWSFLQEIL 545
Query: 146 EDLGFGPRWRCWIHSCVSTATLAVLVNGSPTEFFEIEKGLRQGDPLSPLLFNLCVQGLSV 205
LGF +W WI CV++ T +VL+NG E+G+RQGDP+SP LF LC + L
Sbjct: 546 VALGFNDKWISWIMGCVTSVTYSVLINGQHFGHITPERGIRQGDPISPFLFVLCTEALIH 605
Query: 206 MWNRLAIRERPCGVVL-GPGVCLNHLQFADDTLVFCDPECAQMEELFDVMTAFLWGSGLK 264
+ + ++ G+ G G +NHL F DDT + C + E++ ++ + SG
Sbjct: 606 ILQQAENSKKVSGIQFNGSGPSVNHLLFVDDTQLVCRATKSDCEQMMLCLSQYGHISGQL 665
Query: 265 LNYAKSELV-GCNVDSAIVNQLAVELGVVVGTLPLPYLGSVLGGDPRRRGFWAPVIEKVR 323
+N KS + G VD + G+ + YLG ++ + + EK++
Sbjct: 666 INVEKSSITFGVKVDEDTKRWIKNRSGIHLEGGTGKYLGLPENLSGSKQDLFGYIKEKLQ 725
Query: 324 VALAS*GSNSVSLSGRLVVLRSVASAIPIYHMAVYKAPLGVVADIEKLMRCFLWGRSGGG 383
L+ ++S G+ ++L+S+A A+P+Y M ++ P G+ + +M F W
Sbjct: 726 SHLSGWYDKTLSQGGKEILLKSIALALPVYIMTCFRLPKGLCTKLTSVMMDFWWNSMEFS 785
Query: 384 RRIAWVAWEQVCKRKEIGGLGVEFLKFRNKAMLLKWAWRFGQEPEALWRRVLCAKYGLNS 443
+I W+ +++ K +GG G + L+ N+A+L K AWR + +++ ++ ++Y +N+
Sbjct: 786 NKIHWIGGKKLTLPKSLGGFGFKDLQCFNQALLAKQAWRLFSDSKSIVSQIFKSRYFMNT 845
Query: 444 IPLVLHRVLEGLRHPSRYMFDIVNVVLEDSVLGKTFRDQCFCVVGDGLRISFWRDPWVAS 503
L R +R + +++ +L + ++G+G + + W D W+
Sbjct: 846 DFL-------NARQGTRPSYTWRSILYGRELLNGGLKR----LIGNGEQTNVWIDKWLFD 894
Query: 504 KTLAVLYPRFYAMASNQDIMVSEAGVFSNLGWEWNLGFRRRFFGWEEQVYDELIARCNGV 563
R + S +I + + + L WNL F E+ V +LI +
Sbjct: 895 GHSR----RPMNLHSLMNIHMKVSHLIDPLTRNWNLKKLTELF-HEKDV--QLIMHQRPL 947
Query: 564 VPSREKDSLIWLGDTNGRYTVK---------TF-CMEVELQLYGPPSWVVPSCVKKLAPQ 613
+ S +DS W G NG YTVK TF + E +Y + +
Sbjct: 948 ISS--EDSYCWAGTNNGLYTVKSGYERSSRETFKNLFKEADVYPSVNPLFDKVWSLETVP 1005
Query: 614 KVVLFLWQAAENRVAVMVNLHNRGVLVDAEAVCVLCGQAEESVTHLILVCKFAWVLWSKV 673
K+ +F+W+A + +AV L +RG+ A+ C+ C + E++ HL+ C FA +W+
Sbjct: 1006 KIKVFMWKALKGALAVEDRLRSRGIRT-ADG-CLFCKEEIETINHLLFQCPFARQVWALS 1063
Query: 674 LLREGVVWCVPASILSLLKE--WDSLRKGSDLVLWRLIPYALAWSVWMARNGLIFKGEEV 731
L++ SI S + +S G + + P+ L W +W RN +F+G +
Sbjct: 1064 LIQAPATG-FGTSIFSNINHVIQNSQNFGIPRHMRTVSPW-LLWEIWKNRNKTLFQGTGL 1121
Query: 732 SVETVWDIHITRMYWWIKAGKPHCSFSLCDFLQNFGAVTVGQRVERCRQDVWTCPPEGIL 791
+ + WI A + ++ G V+ + W PP G L
Sbjct: 1122 TSSEIVAKAYEECNLWINAQE-----------KSSGGVSPSEH-------KWNPPPAGEL 1163
Query: 792 KWNVDGAARGAPGMGGIGGVVRDATGEIRGFFSMSLGVVFA-YEAEVKAIRTALEFSKEF 850
K N+ A + G+ V+RD+ G++ S V++ ++A++K+ ALE F
Sbjct: 1164 KCNIGVAWSRQKQLAGVSWVLRDSMGQVLLHSRRSYSQVYSLFDAKIKSWDWALESMDHF 1223
Query: 851 GLSRVVVESDS 861
+V + S
Sbjct: 1224 HFDKVTFAATS 1234
>UniRef100_Q8LJX1 Putative reverse transcriptase [Sorghum bicolor]
Length = 1998
Score = 280 bits (715), Expect = 2e-73
Identities = 205/743 (27%), Positives = 352/743 (46%), Gaps = 36/743 (4%)
Query: 1 PGPDGFNFTFFREFWHLIKEEIMQMFEEFYEGGKLVKGLNAGFIALIPKRQTPEMVTDYR 60
PGPDGF+ F + W +K +I + F+ ++ +N FI LIPK P V DYR
Sbjct: 1260 PGPDGFSNEFMKASWQTVKVDIFNLCHAFHNHNVCLRSINTSFITLIPKSHNPVSVNDYR 1319
Query: 61 LISLIGSIYKLLAKVLAARLQGVMPYLLSENQFSFTKGRQIANCLLIGNEVVDTMLSSGR 120
ISL+ + KL+ K+LA RLQ + L+ +NQ+ F + R I +CL E + S +
Sbjct: 1320 PISLLNTSMKLITKLLANRLQMKITDLVHKNQYGFIRQRTIQDCLAWSFEYLHLCHHSRK 1379
Query: 121 GGFLLKLDFAKAYDNIEWDFLMNMMEDLGFGPRWRCWIHSCVSTATLAVLVNGSPTEFFE 180
++KLDF KA+D + ++ +M+ GFG +W W+ S ++ T VL+NG+P +
Sbjct: 1380 AIVIIKLDFEKAFDKMNHKAMLTIMKAKGFGQKWLNWMKSIFTSGTSNVLLNGTPGKTIH 1439
Query: 181 IEKGLRQGDPLSPLLFNLCVQGLSVMWNRLAIRE-RPCGVVLGPGVCLNHLQFADDTLVF 239
+G+RQGDPLSPLLF L L + N+ + + + +Q+ADDTL+
Sbjct: 1440 CLRGVRQGDPLSPLLFVLAADFLQSLINKAKCMDLLKLPIPMQTNQDFPIIQYADDTLII 1499
Query: 240 CDPECAQMEELFDVMTAFLWGSGLKLNYAKSELVGCNVDSAIVNQLAVELGVVVGTLPLP 299
+ + Q+ L ++ F +GLK+N KS ++ N+D ++ LA G G+ P
Sbjct: 1500 AEGDTRQLLILKSIINTFSEATGLKVNLQKSMMLPINMDEERLDTLARTFGCSKGSFPFT 1559
Query: 300 YLGSVLGGDPRRRGFWAPVIEKVRVALAS*GSNSVSLSGRLVVLRSVASAIPIYHMAVYK 359
YLG LG + P+I K L S + +S +GRL + +V +A+P ++M
Sbjct: 1560 YLGLPLGITKPTIQDYQPLINKCEARLGS-VATFLSEAGRLELTNAVFTALPTFYMCTLA 1618
Query: 360 APLGVVADIEKLMRCFLWGRSG-GGRRIAWVAWEQVCKRKEIGGLGVEFLKFRNKAMLLK 418
P V+ I+K + LW +G R+ AW+ VCK K GGLGV ++ +N+A+L+K
Sbjct: 1619 IPKSVIHKIDKFRKHCLWRGNGINARKPPKAAWKLVCKPKNEGGLGVIDIEKQNEALLMK 1678
Query: 419 WAWRFGQEPEALWRRVLCAK-YGLNSIPLVLHRVLEGLRHPSRYMFDIVNVVLEDSVLGK 477
+F + + W +++ K Y +P +R S + DI+ ++
Sbjct: 1679 NLDKFFNKKDTPWVQMIWDKHYAKGKLP-------GQIRKGSFWWRDILKLL-------D 1724
Query: 478 TFRDQCFCVVGDGLRISFWRDPWVASKTLAVLYPRFYAMASNQDIMVSEAGVFSNLGWEW 537
+++ + G I FW+D W K L + P ++ ++ + +E +
Sbjct: 1725 PYKEMSLIQMKGGESIVFWQDKWGMQK-LRLEMPELFSFVQHKFLSAAEVLGQEDFTRLL 1783
Query: 538 NLGFRRRFFGWEEQVYDELIARCNGVVPSREKDSLIWLGDTNGRYTVKTFCMEVELQLYG 597
L F +Q L R + + ++E+D + N T + Q++
Sbjct: 1784 QLPISETAFHQLQQ----LTTRIDNLERTQEQDQWKYKWGNNFSATKAYSELMGHQQIHE 1839
Query: 598 PPSWVVPSCVKKLAPQKVVLFLWQAAENRVAVMVNLHNRGVLVDAEAVCVLCGQ-AEESV 656
W+ K K +F W ++R++ L + + + + CVLC Q +EE+
Sbjct: 1840 VHKWI----WKSFCQPKHKVFFWLLIKDRLSTCNILKRKNMQLQSYN-CVLCLQNSEETT 1894
Query: 657 THLILVCKFAWVLWSKVLLREGVVWCVPASILSLLKEWDSLRKGSDLVLWRLIPYALAWS 716
+HL L C +A W + + +P + + D + G + + + L W+
Sbjct: 1895 SHLFLHCAYARQCWQLLDMD------IPPN-SDFPEITDHFKHGFNNQFFMVTTILLCWA 1947
Query: 717 VWMARNGLIFKGEEVSVETVWDI 739
+W RN LIF+G + ++E +I
Sbjct: 1948 IWSTRNNLIFRGIQPTIEGTKEI 1970
>UniRef100_Q9FYJ4 F17F8.5 [Arabidopsis thaliana]
Length = 872
Score = 278 bits (710), Expect = 7e-73
Identities = 212/738 (28%), Positives = 348/738 (46%), Gaps = 46/738 (6%)
Query: 1 PGPDGFNFTFFREFWHLIKEEIMQMFEEFYEGGKLVKGLNAGFIALIPKRQTPEMVTDYR 60
PGPDG+ F++ W +I +E + F++ G L KG+N+ +ALIPK+ + + DYR
Sbjct: 118 PGPDGYTSEFYKATWDIIGQEFTLPVQSFFQKGFLPKGINSIILALIPKKLAAKEMRDYR 177
Query: 61 LISLIGSIYKLLAKVLAARLQGVMPYLLSENQFSFTKGRQIANCLLIGNEVV-DTMLSSG 119
IS +YK+++K++A RL+ ++P ++ENQ +F K R + LL+ E+V D S
Sbjct: 178 PISCCNVLYKVISKIIANRLKLLLPRFIAENQSAFVKDRLLIENLLLATELVKDYHKDSI 237
Query: 120 RGGFLLKLDFAKAYDNIEWDFLMNMMEDLGFGPRWRCWIHSCVSTATLAVLVNGSPTEFF 179
+K+D +KA+D+++W FL N + + F P + WI+ C++TA+ +V VNG +F
Sbjct: 238 SARCAIKIDISKAFDSVQWSFLTNTLVAMNFSPTFIHWINLCITTASFSVQVNGDLVGYF 297
Query: 180 EIEKGLRQGDPLSPLLFNLCVQGLSVMWNRLA-IRE---RPCGVVLGPGVCLNHLQFADD 235
+ ++GLRQG LSP LF +C+ LS M ++ A +R+ P LG L HL FADD
Sbjct: 298 QSKRGLRQGCSLSPYLFVICMDVLSKMLDKAAGVRKFGFHPKCQRLG----LTHLSFADD 353
Query: 236 TLVFCDPECAQMEELFDVMTAFLWGSGLKLNYAKSELVGCNVDSAIVNQLAVELGVVVGT 295
+V D + +E + +V F SGL+++ KS L V I ++A + VG
Sbjct: 354 LMVLSDGKTRSIEGILEVFDEFCKRSGLRISLEKSTLYMAGVSPIIKQEIAAKFLFDVGQ 413
Query: 296 LPLPYLGSVLGGDPRRRGFWAPVIEKVRVALAS*GSNSVSLSGRLVVLRSVASAIPIYHM 355
LP+ YLG L ++P++E+++ +A+ S +GR +++SV +I + +
Sbjct: 414 LPVRYLGLPLVTKRLTSADYSPLLEQIKKRIATWTFRFFSFAGRFNLIKSVLWSICNFWL 473
Query: 356 AVYKAPLGVVADIEKLMRCFLWGRSGGGRRIAWVAWEQVCKRKEIGGLGVEFLKFRNKAM 415
A ++ P + +I+KL FLW S A ++W+ VCK K GGLG+ LK N
Sbjct: 474 AAFRLPRQCIREIDKLCSSFLWSGSEMSSHKAKISWDIVCKPKAEGGLGLRNLKEANDVS 533
Query: 416 LLKWAWRFGQEPEALWRRVLCAKYGLNSIPLVLHRVLEGLRHPSRYMFDIVNVVLEDSVL 475
LK WR +LW + + A+Y L+ + + L+ + I +L+ +
Sbjct: 534 CLKLVWRIISNSNSLWTKWV-AEY------LIRKKSIWSLKQSTSMGSWIWRKILKIRDV 586
Query: 476 GKTFRDQCFCVVGDGLRISFWRDPWVASKTLAVLYPRFYAMASNQDIMVSEAGVFSNLGW 535
K+F VG+G SFW D W A + R ++ + +++
Sbjct: 587 AKSFSR---VEVGNGESASFWYDHWSA-------HGRLIDTVGDKGTIDLGIPREASVAD 636
Query: 536 EWNLGFRRRFFGWEEQVYDELIARCNGVVPSREKDSLIWLG--DTNGRYTVKTFCMEVEL 593
W RRR +E++A + S +D+++W G D + +
Sbjct: 637 AWTRRSRRRHRTSLLNEIEEMMA-YQRIHHSDAEDTVLWRGKNDVFKPHFSTRDTWHLIK 695
Query: 594 QLYGPPSWVVPSCVKKLAPQKVVLFLWQAAENRVAVMVNLHNRGVLVDAEAVCVLCGQAE 653
SW + P K L W A NR+ + CVLC
Sbjct: 696 ATSSTVSWHKGVWFRHATP-KYALCTWLAIHNRLPTGDRMLKWNSSGSVSGNCVLCTNNS 754
Query: 654 ESVTHLILVCKFAWVLWSKVLLREGVVWCVPASILSLLKEWDSL--------RKGSDLVL 705
+++ HL C +A +W+ L +G +W S W L + + L
Sbjct: 755 KTLEHLFFSCSYASTVWA--ALAKG-IWKTRYS-----TRWSHLLTHISTHFQDRVEGFL 806
Query: 706 WRLIPYALAWSVWMARNG 723
R I A + VW RNG
Sbjct: 807 TRYIFQATIYHVWRERNG 824
>UniRef100_Q75GM6 Putative non-LTR retroelement reverse transcriptase [Oryza sativa]
Length = 1614
Score = 273 bits (698), Expect = 2e-71
Identities = 175/527 (33%), Positives = 270/527 (51%), Gaps = 16/527 (3%)
Query: 2 GPDGFNFTFFREFWHLIKEEIMQMFEEFYEGGKLVKGLNAGFIALIPKRQTPEMVTDYRL 61
GPDGF+ F+R FW ++ ++ +M Y +K LN G I+LIPK P + +R
Sbjct: 1077 GPDGFSIPFYRAFWPQLRPDLFEMLLMLYNEELDLKRLNFGVISLIPKNSNPTDIKQFRP 1136
Query: 62 ISLIGSIYKLLAKVLAARLQGVMPYLLSENQFSFTKGRQIANCLLIGNEVVDTMLSSGRG 121
I ++ +K ++K + RL + ++S Q +F GR I +I +EV+ M
Sbjct: 1137 ICVLNDCFKFISKCVCNRLTEIARDVISPTQTAFIPGRFILEGCVIIHEVLHEMNRKNLE 1196
Query: 122 GFLLKLDFAKAYDNIEWDFLMNMMEDLGFGPRWRCWIHSCVSTATLAVLVNGSPTEFFEI 181
G +LK+DF KAYD + WDFL+ +M GF +W WI +CV + + +NG T+FF
Sbjct: 1197 GIILKIDFEKAYDKVSWDFLIEVMVRKGFPSKWVNWIKTCVMGGRVCININGERTDFFRT 1256
Query: 182 EKGLRQGDPLSPLLFNLCVQGLSVMWNRLAIRERPCGVV--LGPGVCLNHLQFADDTLVF 239
+GLRQGDPLSPLLFNL L+ M + G+V + PG + HLQ+ADDT++F
Sbjct: 1257 FRGLRQGDPLSPLLFNLISDALAAMLDSAKREGVLSGLVPDIFPG-GITHLQYADDTVLF 1315
Query: 240 CDPECAQMEELFDVMTAFLWGSGLKLNYAKSELVGCNVDSAIVNQLAVELGVVVGTLPLP 299
+ Q+ ++ F + LK+NY KSE+ + N++A+ VG P+
Sbjct: 1316 VANDDKQIVATKFILYCFEEMAVLKVNYHKSEIFTLGLSDNDTNRVAMMFNCPVGQFPMK 1375
Query: 300 YLGSVLGGDPRRRGFWAPVIEKVRVALAS*GSNSVSLSGRLVVLRSVASAIPIYHMAVYK 359
YLG +G D + + +K+ L S G N++S +GR V + + S+IP Y M Y+
Sbjct: 1376 YLGLPIGPDKILNLGFDFLGQKLEKRLNSWG-NNLSHAGRAVQINTCLSSIPSYAMCFYQ 1434
Query: 360 APLGVVADIEKLMRCFLWGRSGGGRRIAWVAWEQVCKRKEIGGLGVEFLKFRNKAMLLKW 419
P GV + + W R+ + V WE + K+ GGLG + N A+L KW
Sbjct: 1435 LPEGVHQKFGSIRGRYYWARNRLKGKYHMVKWEDLAFPKDYGGLGFTETRRMNTALLAKW 1494
Query: 420 AWRFGQEPEALWRRVLCAKYGLNSIPLVLHRVLEGLRHPSRYMFDIVNVVLEDSVLGKTF 479
+ E ++L +L KY + R+ S++ ++N+ S LG +
Sbjct: 1495 IMKIESEDDSLCIELLRRKYLQDGGFFQCKE-----RYASQFWKGLLNIRRWLS-LGSVW 1548
Query: 480 RDQCFCVVGDGLRISFWRDPWVASKTLAVLYPRFYAMASNQDIMVSE 526
+ VGDG ISFWRD W L L+P +A+ + QDI+VS+
Sbjct: 1549 Q------VGDGSHISFWRDVWWGQCPLRTLFPAIFAVCNQQDILVSQ 1589
>UniRef100_Q9C6L3 Hypothetical protein F2J7.11 [Arabidopsis thaliana]
Length = 1213
Score = 271 bits (694), Expect = 5e-71
Identities = 210/741 (28%), Positives = 333/741 (44%), Gaps = 51/741 (6%)
Query: 2 GPDGFNFTFFREFWHLIKEEIMQMFEEFYEGGKLVKGLNAGFIALIPKRQTPEMVTDYRL 61
GPDGF FF + W ++ E+ +EF+ G L+K NA I LIPK P +D+R
Sbjct: 466 GPDGFTAEFFIDSWSIVGAEVTDAIKEFFSSGCLLKQWNATTIVLIPKIVNPTCTSDFRP 525
Query: 62 ISLIGSIYKLLAKVLAARLQGVMPYLLSENQFSFTKGRQIANCLLIGNEVV---DTMLSS 118
IS + ++YK++A++L RLQ ++ ++S Q +F GR +A +L+ ++V + S
Sbjct: 526 ISCLNTLYKVIARLLTDRLQRLLSGVISSAQSAFLPGRSLAENVLLATDLVHGYNWSNIS 585
Query: 119 GRGGFLLKLDFAKAYDNIEWDFLMNMMEDLGFGPRWRCWIHSCVSTATLAVLVNGSPTEF 178
RG +LK+D KA+D++ W+F++ + L ++ WI C+ST T V +NG F
Sbjct: 586 PRG--MLKVDLKKAFDSVRWEFVIAALRALAIPEKFINWISQCISTPTFTVSINGGNGGF 643
Query: 179 FEIEKGLRQGDPLSPLLFNLCVQGLSVMWNRLAIRERPCGVVLGP---GVCLNHLQFADD 235
F+ KGLRQGDPLSP LF L ++ S N L R + P + ++HL FADD
Sbjct: 644 FKSTKGLRQGDPLSPYLFVLAMEAFS---NLLHSRYESGLIHYHPKASNLSISHLMFADD 700
Query: 236 TLVFCDPECAQMEELFDVMTAFLWGSGLKLNYAKSELVGCNVDSAIVNQLAVELGVVVGT 295
++F D + + + + F SGLK+N KS L ++ N A G +GT
Sbjct: 701 VMIFFDGGSFSLHGICETLDDFASWSGLKVNKDKSHLYLAGLNQLESNANAA-YGFPIGT 759
Query: 296 LPLPYLGSVLGGDPRRRGFWAPVIEKVRVALAS*GSNSVSLSGRLVVLRSVASAIPIYHM 355
LP+ YLG L R + P++EK+ S + +S +GR+ ++ SV + M
Sbjct: 760 LPIRYLGLPLMNRKLRIAEYEPLLEKITARFRSWVNKCLSFAGRIQLISSVIFGSINFWM 819
Query: 356 AVYKAPLGVVADIEKLMRCFLWGRSGGGRRIAWVAWEQVCKRKEIGGLGVEFLKFRNKAM 415
+ + P G + IE L FLW + + V+W +C K GGLG+ L NK +
Sbjct: 820 STFLLPKGCIKRIESLCSRFLWSGNIEQAKGIKVSWAALCLPKSEGGLGLRRLLEWNKTL 879
Query: 416 LLKWAWRFGQEPEALWRRVLCAKYGLNSIPLVLHRVLEGLRHPSRYMFDIVNVVLEDSVL 475
++ WR ++LW + L H SR F V DS
Sbjct: 880 SMRLIWRLFVAKDSLWAD------------------WQHLHHLSRGSFWAVEGGQSDSWT 921
Query: 476 GKTF-------RDQCFCVVGDGLRISFWRDPWVASKTLAVLYPRFYAMASNQDIMVSEAG 528
K C VG+GL+ +W D W + L + + ++ A
Sbjct: 922 WKRLLSLRPLAHQFLVCKVGNGLKADYWYDNWTSLGPLFRIIGDIGPSSLRVPLLAKVAS 981
Query: 529 VFSNLGWEWNLGFRRRFFGWEEQVYDELIARCNGVVPSREKDSLIWLGDTNGRYTVKTFC 588
FS GW + G ++D L C VPS ++ + + + + F
Sbjct: 982 AFSEDGWRLPVSRSAPAKG----IHDHL---CTVPVPSTAQEDVDRYEWSVNGFLCQGFS 1034
Query: 589 MEVELQLYGP----PSWVVPSCVKKLAPQKVVLFLWQAAENRVAVMVNLHNRGVLVDAEA 644
+ P SW K P K +W + NR+ L + G +
Sbjct: 1035 AAKTWEAIRPKATVKSWASSIWFKGAVP-KYAFNMWVSHLNRLLTRQRLASWGHI--QSD 1091
Query: 645 VCVLCGQAEESVTHLILVCKFAWVLWSKVLLREGVVWCVPASILSLLKEWDSLRKGSDLV 704
CVLC A ES HL+L+C+F+ +W V R + +S LL + +
Sbjct: 1092 ACVLCSFASESRDHLLLICEFSAQVWRLVFRRICPRQRLFSSWSELLSWVRQSSPEAPPL 1151
Query: 705 LWRLIPYALAWSVWMARNGLI 725
L +++ + +++W RN L+
Sbjct: 1152 LRKIVSQVVVYNLWRQRNNLL 1172
>UniRef100_Q7XE51 Putative non-LTR retroelement reverse transcriptase [Oryza sativa]
Length = 1652
Score = 268 bits (684), Expect = 7e-70
Identities = 165/501 (32%), Positives = 262/501 (51%), Gaps = 17/501 (3%)
Query: 1 PGPDGFNFTFFREFWHLIKEEIMQMFEEFYEGGKLVKGLNAGFIALIPKRQTPEMVTDYR 60
PGPDGF F++ FW +IK+++M +F +F+EG + L G I L+PK++ + YR
Sbjct: 1165 PGPDGFPAEFYQVFWDVIKDDLMAVFCDFHEGTLPLHRLKFGIITLLPKQKDASRIQQYR 1224
Query: 61 LISLIGSIYKLLAKVLAARLQGVMPYLLSENQFSFTKGRQIANCLLIGNEVVDTMLSSGR 120
I L+ +K+ KV+A R+ V ++ +Q +F GR I ++I +E + + +
Sbjct: 1225 PICLLNVSFKIFTKVMANRIALVAQKVIKPSQTTFLSGRNIMEGVVILHETLHELHKKKK 1284
Query: 121 GGFLLKLDFAKAYDNIEWDFLMNMMEDLGFGPRWRCWIHSCVSTATLAVLVNGSPTEFFE 180
G +LKLDF KAYD ++W FL + GF +W WI S V ++AV VN +F+
Sbjct: 1285 NGVILKLDFEKAYDKVDWKFLQQSLRMKGFSSKWCDWIDSIVRGGSVAVKVNDEIGSYFQ 1344
Query: 181 IEKGLRQGDPLSPLLFNLCVQGLSVMWNRLAIRERPCGVVLGPGVC---LNHLQFADDTL 237
KGLRQGDPLSP+LFNL V L+++ R + R GVV P + L+ LQ+ADDT+
Sbjct: 1345 TRKGLRQGDPLSPILFNLVVDMLAILIQRAKDQGRFKGVV--PHLVDNGLSILQYADDTI 1402
Query: 238 VFCDPECAQMEELFDVMTAFLWGSGLKLNYAKSELVGCNVDSAIVNQLAVELGVVVGTLP 297
+F D + + +L V++ F S LK+N+ KSEL + ++ G P
Sbjct: 1403 LFMDHDLDEARDLKLVLSTFEKLSSLKINFYKSELFCYGKAKDVEHEYVKLFGCDTEDYP 1462
Query: 298 LPYLGSVLGGDPRRRGFWAPVIEKVRVALAS*GSNSVSLSGRLVVLRSVASAIPIYHMAV 357
YLG + W V E+++ L+S +S+ GRLV++ SV S + I+ ++
Sbjct: 1463 FKYLGIRMHHKRINNKDWQGVEERIQKKLSSWKGKFLSVGGRLVLINSVLSNLAIFMLSF 1522
Query: 358 YKAPLGVVADIEKLMRCFLWGRSGGGRRIAWVAWEQVCKRKEIGGLGVEFLKFRNKAMLL 417
++ P G++ ++ F W ++ W +CK KE GGLG++ L+ +NK +L
Sbjct: 1523 FEIPKGILKKLDYYRSRFFWQCDEHKKKYRLARWSVLCKPKECGGLGIQNLEIQNKCLLS 1582
Query: 418 KWAWRFGQEPEALWRRVLCAKYGLNSIPLVLHRVLEGLRHPSRYMFDIVNVVLEDSVLGK 477
KW ++ E E +W+ +L KY L + G H + + +V + G
Sbjct: 1583 KWLYKLINE-EGVWQDILRNKY-LTKKTITQVEKCPGDSHFWAGLMGVRDVFFK----GG 1636
Query: 478 TFRDQCFCVVGDGLRISFWRD 498
F+ VGDG +I FW D
Sbjct: 1637 AFK------VGDGSQIRFWED 1651
>UniRef100_O81068 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 1529
Score = 267 bits (683), Expect = 9e-70
Identities = 214/757 (28%), Positives = 328/757 (43%), Gaps = 87/757 (11%)
Query: 1 PGPDGFNFTFFREFWHLIKEEIMQMFEEFYEGGKLVKGLNAGFIALIPKRQTPEMVTDYR 60
PGPDG+ FF+ W L + + + F+ G L KGLNA +ALIPK+ + DYR
Sbjct: 768 PGPDGYTSEFFKATWSLTGPDFIAAIQSFFVKGFLPKGLNATILALIPKKDEAIEMKDYR 827
Query: 61 LISLIGSIYKLLAKVLAARLQGVMPYLLSENQFSFTKGRQIANCLLIGNEVV-DTMLSSG 119
IS +YK+++K+LA RL+ ++P + +NQ +F K R + +L+ E+V D S
Sbjct: 828 PISCCNVLYKVISKILANRLKLLLPSFILQNQSAFVKERLLMENVLLATELVKDYHKESV 887
Query: 120 RGGFLLKLDFAKAYDNIEWDFLMNMMEDLGFGPRWRCWIHSCVSTATLAVLVNGSPTEFF 179
+K+D +KA+D+++W FL+N +E L F +R WI C+STAT +V VNG FF
Sbjct: 888 TPRCAMKIDISKAFDSVQWQFLLNTLEALNFPETFRHWIKLCISTATFSVQVNGELAGFF 947
Query: 180 EIEKGLRQGDPLSPLLFNLCVQGLSVMWNRLAIRERPCGVVLGPGVCLNHLQFADDTLVF 239
+GLRQG LSP LF +C+ LS M + A+ + L HL FADD +VF
Sbjct: 948 GSSRGLRQGCALSPYLFVICMNVLSHMIDEAAVHRNIGYHPKCEKIGLTHLCFADDLMVF 1007
Query: 240 CDPECAQMEELFDVMTAFLWGSGLKLNYAKSELVGCNVDSAIVNQLAVELGVVVGTLPLP 299
D +E + +V F SGL+++ KS + V ++ Q G LP+
Sbjct: 1008 VDGHQWSIEGVINVFKEFAGRSGLQISLEKSTIYLAGVSASDRVQTLSSFPFANGQLPVR 1067
Query: 300 YLGSVLGGDPRRRGFWAPVIEKVRVALAS*GSNSVSLSGRLVVLRSVASAIPIYHMAVYK 359
YLG L ++P+IE V+ ++S + S+S +GRL +L SV +I + M+ Y+
Sbjct: 1068 YLGLPLLTKQMTTADYSPLIEAVKTKISSWTARSLSYAGRLALLNSVIVSIANFWMSAYR 1127
Query: 360 APLGVVADIEKLMRCFLWGRSGGGRRIAWVAWEQVCKRKEIGGLGVEFLKFRNKAMLLK- 418
P G + +IEKL FLW + A +AW +C+ K+ GGLG++ L NK LK
Sbjct: 1128 LPAGCIREIEKLCSAFLWSGPVLNPKKAKIAWSSICQPKKEGGLGIKSLAEANKVSCLKL 1187
Query: 419 --------------WAWRFGQEPEALWRRVLCAKYGLNSIPLVLHRVLEGLRHPSRYMFD 464
W W F W + +S+ + + L R ++ M
Sbjct: 1188 IWRLLSTQPSLWVTWIWTFIIRKGTFW-----SANERSSLGSWMWKKLLKYRELAKSMHK 1242
Query: 465 IVNVVLEDSVLGKTFRDQCFCVVGDGLRISFWRDPWVASKTLAVLYPRFYAMASNQDIMV 524
+ V +G SFW D W L DI
Sbjct: 1243 VE--------------------VRNGSSTSFWYDHWSHLGRLL-------------DITG 1269
Query: 525 SEAGVFSNLGWEWNLGFRRRFFGWEEQ---VYDELIARCNGVVPSREK---DSLIWLG-- 576
+ + + E NL R + +Y+ + A + + D +W
Sbjct: 1270 TRRVIDLGIPLETNLETVLRTHQHRQHRAAIYNRINAEIQRLQQQEREAGPDISLWRSLK 1329
Query: 577 -DTNGRYTVKTFCMEVELQLYGPPSWVVPSCVKKLAPQKVVLFLWQAAENRVAV--MVNL 633
D N R+ K V +W P K LW +NR++ +
Sbjct: 1330 NDFNKRFITKVTWNNVRTH-QPQQNWYKGVWFPYSTP-KYSFLLWLTVQNRLSTGDRIKA 1387
Query: 634 HNRGVLVDAEAVCVLCGQAEESVTHLILVCKFAWVLWSKVLLREGVVWCVPASILSLLKE 693
N G LV C LC AEE+ HL C++ +W + R + ++
Sbjct: 1388 WNSGQLV----TCTLCNNAEETRDHLFFSCQYTSYVWEALTQR--------LLSTNYSRD 1435
Query: 694 WDSL--------RKGSDLVLWRLIPYALAWSVWMARN 722
W+ L L L+R + A + +W RN
Sbjct: 1436 WNRLFTLLCTSNLPRDHLFLFRYVFQASIYHIWRERN 1472
>UniRef100_Q9SN65 Hypothetical protein F25I24.40 [Arabidopsis thaliana]
Length = 1294
Score = 266 bits (680), Expect = 2e-69
Identities = 149/444 (33%), Positives = 247/444 (55%), Gaps = 5/444 (1%)
Query: 1 PGPDGFNFTFFREFWHLIKEEIMQMFEEFYEGGKLVKGLNAGFIALIPKRQTPEMVTDYR 60
PGPDG F++ W ++ ++++ + F+ + + +N I +IPK PE ++DYR
Sbjct: 808 PGPDGLTARFYKSCWEIVGPDVIKEVKIFFRTSYMKQSINHTNICMIPKITNPETLSDYR 867
Query: 61 LISLIGSIYKLLAKVLAARLQGVMPYLLSENQFSFTKGRQIANCLLIGNEVVDTMLSSGR 120
I+L +YK+++K L RL+G + ++S++Q +F GR + + ++I +E++ ++ + R
Sbjct: 868 PIALCNVLYKIISKCLVERLKGHLDAIVSDSQAAFIPGRLVNDNVMIAHEMMHSLKTRKR 927
Query: 121 GG---FLLKLDFAKAYDNIEWDFLMNMMEDLGFGPRWRCWIHSCVSTATLAVLVNGSPTE 177
+K D +KAYD +EW+FL M GF W WI V + +VLVNG P
Sbjct: 928 VSQSYMAVKTDVSKAYDRVEWNFLETTMRLFGFSETWIKWIMGAVKSVNYSVLVNGIPHG 987
Query: 178 FFEIEKGLRQGDPLSPLLFNLCVQGLSVMWNRLAIRERPCGVVLGPGVC-LNHLQFADDT 236
+ ++G+RQGDPLSP LF LC L+ + G+ +G GV + HLQFADD+
Sbjct: 988 TIQPQRGIRQGDPLSPYLFILCADILNHLIKNRVAEGDIRGIRIGNGVPGVTHLQFADDS 1047
Query: 237 LVFCDPECAQMEELFDVMTAFLWGSGLKLNYAKSELV-GCNVDSAIVNQLAVELGVVVGT 295
L FC + L DV + + SG K+N +KS + G V N+L LG+
Sbjct: 1048 LFFCQSNVRNCQALKDVFDVYEYYSGQKINMSKSMITFGSRVHGTTQNRLKNILGIQSHG 1107
Query: 296 LPLPYLGSVLGGDPRRRGFWAPVIEKVRVALAS*GSNSVSLSGRLVVLRSVASAIPIYHM 355
YLG ++R + +IE+V+ +S + +S +G+ ++L+SVA ++P+Y M
Sbjct: 1108 GGGKYLGLPEQFGRKKRDMFNYIIERVKKRTSSWSAKYLSPAGKEIMLKSVAMSMPVYAM 1167
Query: 356 AVYKAPLGVVADIEKLMRCFLWGRSGGGRRIAWVAWEQVCKRKEIGGLGVEFLKFRNKAM 415
+ +K PL +V++IE L+ F W ++ R I W+AW+++ K+ GGLG L N A+
Sbjct: 1168 SCFKLPLNIVSEIEALLMNFWWEKNAKKREIPWIAWKRLQYSKKEGGLGFRDLAKFNDAL 1227
Query: 416 LLKWAWRFGQEPEALWRRVLCAKY 439
L K WR P +L+ R++ A+Y
Sbjct: 1228 LAKQVWRMINNPNSLFARIMKARY 1251
>UniRef100_Q9FZG5 T2E6.4 [Arabidopsis thaliana]
Length = 740
Score = 266 bits (680), Expect = 2e-69
Identities = 151/432 (34%), Positives = 234/432 (53%), Gaps = 1/432 (0%)
Query: 1 PGPDGFNFTFFREFWHLIKEEIMQMFEEFYEGGKLVKGLNAGFIALIPKRQTPEMVTDYR 60
PGPDG+ FF+ W + ++ + + F+ G L KGLNA +ALIPK+ ++ DYR
Sbjct: 44 PGPDGYTSEFFKATWSITGQDFIAAIKSFFIKGFLPKGLNATILALIPKKDEATLMRDYR 103
Query: 61 LISLIGSIYKLLAKVLAARLQGVMPYLLSENQFSFTKGRQIANCLLIGNEVV-DTMLSSG 119
IS IYK+++K++A RL+ ++P + +NQ +F + R + +L+ E+V D S
Sbjct: 104 PISCCNVIYKVISKIIANRLKVMLPTFILQNQSAFVRERLLIENVLLATELVKDYHKDSI 163
Query: 120 RGGFLLKLDFAKAYDNIEWDFLMNMMEDLGFGPRWRCWIHSCVSTATLAVLVNGSPTEFF 179
+K+D +KA+D+++W FL+N +E L F + WI C+STAT +V VNG FF
Sbjct: 164 SPRCAMKIDISKAFDSVQWQFLLNTLEALNFPENFCHWIKLCISTATFSVQVNGELAGFF 223
Query: 180 EIEKGLRQGDPLSPLLFNLCVQGLSVMWNRLAIRERPCGVVLGPGVCLNHLQFADDTLVF 239
++GLRQG LSP LF +C+ LS M + A+ + L HL FADD +VF
Sbjct: 224 GSKRGLRQGCALSPYLFVICMNVLSHMIDVAAVHRNIGYHPKCKKLSLTHLCFADDLMVF 283
Query: 240 CDPECAQMEELFDVMTAFLWGSGLKLNYAKSELVGCNVDSAIVNQLAVELGVVVGTLPLP 299
D + +E + ++ F SGL ++ KS L V N + G LP+
Sbjct: 284 IDGQQRSVEGVINIFKEFAGKSGLHISLEKSTLYLAGVSELNRNNILSAFPFASGQLPVR 343
Query: 300 YLGSVLGGDPRRRGFWAPVIEKVRVALAS*GSNSVSLSGRLVVLRSVASAIPIYHMAVYK 359
YLG L ++P+++KVR ++S + S+S +GRL ++ SV ++ + M+ Y+
Sbjct: 344 YLGLPLLTKQMTTADYSPLLDKVRSKISSWTARSLSYAGRLALINSVIVSLSNFWMSAYR 403
Query: 360 APLGVVADIEKLMRCFLWGRSGGGRRIAWVAWEQVCKRKEIGGLGVEFLKFRNKAMLLKW 419
P G + +IEKL FLW + A + W +CK K+ GGLG++ L NK LK
Sbjct: 404 LPAGCIKEIEKLCSAFLWSGPELNPKKAKITWTSLCKLKQEGGLGIKSLLEANKVSCLKL 463
Query: 420 AWRFGQEPEALW 431
WR +LW
Sbjct: 464 IWRLVSRQSSLW 475
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.326 0.142 0.462
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,601,650,832
Number of Sequences: 2790947
Number of extensions: 68935484
Number of successful extensions: 180924
Number of sequences better than 10.0: 1235
Number of HSP's better than 10.0 without gapping: 705
Number of HSP's successfully gapped in prelim test: 530
Number of HSP's that attempted gapping in prelim test: 177971
Number of HSP's gapped (non-prelim): 1839
length of query: 916
length of database: 848,049,833
effective HSP length: 137
effective length of query: 779
effective length of database: 465,690,094
effective search space: 362772583226
effective search space used: 362772583226
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 80 (35.4 bits)
Lotus: description of TM0166.1


